Abstract
Key Message
Maize group II LEA protein ZmDHN11 could protect protein activity and confer resistance to osmotic stress on transgenic yeast and tobacco.
Abstract
Late embryogenesis abundant (LEA) proteins are widely assumed to play crucial roles in environmental stress tolerance, but their function has remained obscure. Dehydrins are group II LEA proteins, which are highly hydrophilic plant stress proteins. In the present study, a novel group II LEA protein, ZmDHN11, was cloned and identified from maize. The expression of ZmDHN11 was induced by high osmotic stress, low temperature, salinity, and ABA (abscisic acid). The ZmDHN11 protein specifically accumulated in the nuclei and cytosol. Further study indicated that ZmDHN11 is phosphorylated by the casein kinase CKII. ZmDHN11 protected the activity of LDH under water-deficit stress. The overexpression of ZmDHN11 endows transgenic yeast and tobacco with tolerance to osmotic stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Due to global climate change, drought stress has had the most significant impact on the morphological, physiological, and biochemical characteristics of plants (Jiang et al. 2013; Cai et al. 2014; Pan et al. 2012; Li et al. 2018). With the loss of cellular homeostasis under drought conditions, the increased production of reactive oxygen species (ROS) and the accumulation of toxic metabolites could cause further damage to plants (Xiong et al. 2002; Miller et al. 2010). During the evolutionary adaptation process, plants have developed a series of molecular and physiological mechanisms to reduce the damage caused by water deficit, including structural changes and the synthesis of hydrophilic proteins (Wei et al. 2016).
Studies have implicated late embryogenesis abundant (LEA) proteins in water-deficit stress (Goyal et al. 2005). LEA proteins were first found in cotton seeds and accumulated in the late stage of plant seed development (Dure et al. 1981). LEA proteins are mainly involved in the protection of desiccation during seed dehydration, or vegetative tissues under stress conditions by acting as cellular dewatering protectants (Wise et al. 2004). According to their conserved motifs and amino acid sequence homology, LEA proteins can be divided into seven families (Battaglia et al. 2008). Group II LEA proteins, also known as dehydrins (dehydration proteins), are a large group of highly hydrophilic proteins. Dehydrins contain a low fraction of hydrophobic and nonpolar residues, whereas others contain a high proportion of polar and charged amino acids (Garay et al. 2000; Liu et al. 2017a). To adapt to severe environmental conditions, the accumulation of dehydrins is prominent in plants. The high levels of dehydrin expression enhance the ability of plants to survive abiotic stress (Malik et al. 2017).
Although significant similarity has not been found among the members of the different LEA protein families, recent studies have provided experimental evidence demonstrating the function of LEA proteins under various environmental stress conditions (Sun et al. 2013; Amara et al. 2013; Liu et al. 2014). For example, overexpression of the barley LEA protein HVA1 improved drought tolerance in wheat and rice (Sivamani et al. 2000). A previous study examined the LEA gene family in cucumbers and found that the DHN gene CsLEA54 might facilitate the adaptation of cucumbers to drought stress (Celik et al. 2016). The overexpression of cold-induced dehydrins CuCOR19 and WCOR410 in tobacco and strawberry plants, respectively, resulted in improved cold tolerance of transgenic plants (Houde et al. 2004; Hara et al. 2003). The maize group III LEA protein ZmLEA3 conferred transgenic yeast and tobacco tolerance to low temperature, drought, and oxidative stresses (Liu et al. 2013a, 2016). The accumulation of Arabidopsis thaliana LEA protein AtLEA4 was positively correlated with its drought stress resistance (Olvera et al. 2011).
There are three conserved segments in dehydrins: the K-, S-, and Y-segments. All dehydrins possess one to 11 of copies conserved K-segment(s) whose typical sequence is the Lys-rich 15-residue motif EKKGIMEKIKEKLPG. Dehydrin functions are believed to be related to the K-segment, which has been predicted to form an amphipathic helix (Koag et al. 2009). Dehydrins may also possess three other segments: a track of Ser residues (the S-segment), which in some proteins can be phosphorylated, and less conserved motifs (φ-segments) are usually rich in polar amino acids and lay interspersed between K-segments. A consensus motif, [V/T]D[E/Q]YGNP (the Y-segment), has been found near the N-terminus (Close et al. 1997). Based on the distribution and the number of the three segments, dehydrins are classified into five subclasses, namely SKn, YnSKn, YnKn, KnS, and Kn (Tanmoy et al. 2017).
Although previous studies have been carried out on group II LEA proteins, their physiological functions have not been revealed. As the lowest molecular weight dehydrin family member, the K-segment of KS-type dehydrins begins with the sequence HKEG (in others, a similar site contains of EKKG), which demonstrates that the KS-type dehydrin may have different functions. In this study, to investigate the functional mechanism and the molecular properties of group II LEA proteins in response to abiotic stresses, we cloned a novel group II LEA protein, the KS-type dehydrin ZmDHN11 from maize. ZmDHN11 is phosphorylated by the casein kinase CKII. In vitro experiments showed that ZmDHN11 could protect the activity of LDH under water-deficit stress. Additional studies indicated that overexpression of ZmDHN11 conferred water-deficit stress tolerance in transgenic yeast and tobacco.
Materials and methods
Plant materials and growth conditions
Maize (Zea mays L. cv. Zhengdan 958) and tobacco (Nicotiana benthamiana) were cultured with Hoagland's solution (pH 6.0) in a growth chamber at 26 °C/22 °C (day/night) for 2 weeks. The nutrient solution was replaced daily. The photoperiod of the growth chamber was 14 h/10 h (day/night), and the photosynthetic effective radiation was 600 mol−2 s−1.
Amplification and sequence analysis of ZmDHN11
First-strand cDNA was obtained using 5 μg of total RNA, which was extracted from young maize leaves using Plant RNA Reagent according to the manufacturer’s instructions (TIANGEN, China). The entire coding sequence of ZmDHN11 was amplified with specific primers (forward GGATCCCAGTCCATTATTGCCGTC, BamHI site underlined and reverse GAGCTCGTGTTGCACCTGTGCT, SacI site underlined). The hydropathic profiles of the ZmDHN11 protein were constructed by the ProtScale method (http://expasy.org/tools/protscale.html).
Subcellular localization
The ZmDHN11 gene was inserted into the reconstructed binary vector pBI121-GFP, which generated a C-terminal fusion protein with the green fluorescent protein (GFP) gene controlled by the cauliflower mosaic virus (CaMV) 35S promoter. Constructs were transformed into the Agrobacterium tumefaciens strain LBA4404, and the transformation of tobacco plants was performed by the leaf disk transformation method (Liu et al. 2013b).
Gene expression analyses of ZmDHN11 under different stress conditions using real-time PCR
Two-week-old maize was treated with 20% PEG6000 (w/v), 100 μM ABA, 250 mM NaCl, and low temperature (4℃). Leaves collected from the different treated plants at the indicated time points were immediately frozen in liquid nitrogen and stored at − 80 °C. First-strand cDNA synthesis was performed as described above. All cDNAs were used in real-time PCRs (qRT-PCR) performed using primers (forward 5'-GAAGAGCAGTCGCCATGTCT-3' and reverse 5'-CTCCACGATGCCTTCCTTGT-3') and SYBR Green qRT-PCR SuperMix (TransGene, China). The maize actin gene was amplified along with the ZmDHN11 gene, which was used to normalize the amount of template.
Protein purification
The expression and purification of ZmDHN11 was performed as described with minor changes (Liu et al. 2016). The open-reading frame (ORF) of ZmDHN11 was ligated into the expression system (Novagen) of pET30 Escherichia coli strain BL21 (DE3). The recombinant protein 6 × His tag was expressed at the N-terminus according to the protocol of the manufacturer (pET system). The ZmDHN11 fusion protein was purified by affinity chromatography using a Ni- column and then exchanged in low–medium salt buffer (100 mM NaCl, 50 mM Tris–HCl at pH 8.0) using a HiPrep desalting column (GE Healthcare). After determining the purity of the fusion protein on an SDS-PAGE gel, the samples were stored at − 20 °C.
Water-deficit inactivation of lactate dehydrogenase
LDH (lactate dehydrogenase, Roche, U.K.) in rabbit muscle was diluted with 25 mM Tris–HCl at pH 7.5. Water-deficit treatment was assayed according to a previously published method (Liu et al. 2014). For water-deficit treatment, LDH was treated by vacuum drying for 1 h. The enzymatic activity of the untreated LDH was referred to as 100%. The values represent the average of three independent experiments. The hydrophilic proteins poly-lysine (Invitrogen) and lysozyme (Amresco) were used as controls. For LDH activity, the buffer was 25 mM Tris–HCl (pH 7.5), which contained 0.15 mM NADH (Roche) and 2 mM pyruvate (sigma). At 25 °C, NADH transformed into NAD, and the activity of LDH was the rate of decrease in absorbance at 340 nm over 1 min using a spectrophotometer.
The phosphorylation analyses of ZmDHN11
Phosphorylation was performed as previously described (Liu et al. 2017b). Casein kinase (CKII, New England Biolabs) was used to phosphorylate ZmDHN11. The 5 μg of recombinant proteins, 100 U of casein kinase CKII, 1X reaction buffer (20 mM Tris–HCl, 50 mM KCl, 10 mM MgCl2), and 200 μM ATP were mixed in a total reaction volume of 25 μl. The reaction mixtures were incubated for 2 h at 30 °C and then analyzed by SDS-PAGE. A Pro-Q Diamond Phosphoprotein Gel Staining Kit (Invitrogen) was used to detect phosphoproteins according to the manufacturer's instructions. The details of the experiment are as follows. The gel was incubated the in 100 mL of fix solution (50% methanol and 10% acetic acid) at room temperature with gentle agitation for 30 min. Incubate the gel in 100 mL of ultrapure water with gentle agitation for 10 min. Next, the gel was incubated in a volume of Pro-Q Diamond phosphoprotein gel stain that was equivalent to 10 times the volume of the gel. Then, the gel was incubated in 100 mL of destain solution (50 mL of 1 M sodium acetate, pH 4.0, 750 mL of ultrapure water, and 200 mL of acetonitrile) with gentle agitation for 30 min at room temperature. The gel was washed twice with ultrapure water at room temperature for 5 min per wash. Finally, the stained gels were observed using a UV transilluminator.
Identification of transgenic tobacco transcription levels
The transcription levels of the ZmDHN11 transcripts in transgenic tobacco plants were analyzed by qRT-PCR. The tobacco actin gene was amplified by using primers (forward 5'-GATGAAGATACTCACAGA-3' and reverse 5'-ATAGTCAAGAGCAATGTA-3') along with the ZmDHN11 gene to normalize the expression of the target gene.
The expression of ZmDHN11 in Pichia yeast GS115
The correct sequences of ZmDHN11 and the AOXI promoter were connected to plasmid Ppic3.5 K (Invitrogen, USA). Five micrograms of linearized plasmid Ppic3.5 K -ZmDHN11 was transformed into the Pichia GS115 strain by the LiCl method. The same strain transformed with the empty body of Ppic3.5 K was used as a negative control. All yeast cells were transformed and transferred to MD plates (4 × 10–5 biotin, 1.34% YNB and 2% glucose), and then identified by PCR. The accumulation of the ZmDHN11 in transgenic yeasts was performed as previously described (Liu et al. 2014).
Osmotic tolerance assays of yeast transformants
The osmotic tolerance assays of yeast were assayed according to a previously published method (Liu et al. 2014). The recombinant colonies were inoculated in 25 mL of BMGY medium (2% peptone, 1% yeast extract, 1.34% YNB, 10 mM K3PO4, 4 × 10–5 biotin, and 1% glycerine). After incubation at 28℃ for 18 h, the yeast cells were collected by centrifugation, resuscitated in 200 mL induction BMMY medium (2% peptone, 1% yeast extract, 1.34% YNB, 10 mM K3PO4, 0.5% methanol and 4 × 10–5 biotin), and incubated at 28℃ for 4 days. Methanol was added every 24 h for a final concentration of 0.5%. The yeast transformants Ppic3.5 K-ZmDHN11 and Ppic3.5 K were cultured to OD600 = 0.8, and then 2 mL of culture was inoculated into 150 mL of BMGY medium supplemented with 800 mM mannitol. At each time point, the OD600 of 3 mL of culture was measured by a spectrophotometer. Growth was measured at least three times.
Histochemical detection of O2 −
O2− accumulation was detected by NBT staining methods. The methods were performed as previously described (Liu et al. 2013a). Two-week-old control lines and transgenic lines were treated with 250 mM mannitol for 7 days, and then, the seedlings were infiltrated with 0.5 mg/mL nitroblue tetrazolium (NBT) for 24 h in the dark to detect O2−. Then, the seedlings were decolorized by boiling in ethanol (95%) for 15 min. After cooling, the seedlings were extracted at room temperature with fresh ethanol and then photographed using a stereomicroscope. The experiment was repeated three times.
Osmotic stress treatments and assays in plants
Six-week-old control lines and transgenic lines were treated with 20% PEG6000 for 5 days, and then, 0.5 g of leaves was collected for MDA (malondialdehyde) measurements. Relative electrolytic leakage and MDA content were determined as described by Liu et al. 2014. Peroxidase (POD) and superoxide dismutase (SOD) were measured as described in Liu et al. 2017b. The experiments were repeated more than three times.
Results
Isolation and bioinformatic analysis of ZmDHN11
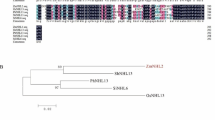
To elucidate the role of group II LEA proteins, ZmDHN11 (GenBank: AY103822.1) was isolated from maize. The open-reading frame (ORF) of ZmDHN11 is 303 bp and encodes a protein of 100 amino acids with a predicted molecular mass of 11 kDa (http://www.expasy.org/tools/pi_tool.html) (Fig. 1a). Analysis of ZmDHN11 with the maizeGDB database (http://www.maizegdb.org/) showed that ZmDHN11 is located on chromosome 1. The ZmDHN11 protein is rich in Lys (23%), His (16%), and Glu (16%), while the Leu, Thr, and Val residue contents are only 1%. As shown by hydropathy plots, the ZmDHN11 protein is highly hydrophilic (Fig. 1b). ZmDHN11 displays diverse homology with other KS-type dehydrins, and its motifs broadly match similar segments in related dehydrins, indicating a close evolutionary relationship between these proteins (Fig. 1c). ZmDHN11 possesses one K-segment (HKEGIVEKIKDKITG) and one S-segment (SSSSSDSD). According to the conserved domains, ZmDHN11 is a KS-type dehydrin.
Sequence analysis of ZmDHN11. a Nucleotide sequence of ZmDHN11 cDNA together with its predicted amino acid sequence. b Hydropathic predictions from the Kyte–Doolittle algorithm. Regions with hydropathic indices below 0 are considered to be hydrophilic. c Multiple sequence alignment of ZmDHN11 with other KS-type dehydrin proteins
The transcription of ZmDHN11could be induced by different transcript stress treatments
To study the function of ZmDHN11 in the regulation of abiotic stress resistance in maize, the expression of ZmDHN11 in maize under drought, low temperature, ABA, and salt stress was detected by fluorescence quantitative PCR (Fig. 2). Under the 20% PEG6000 and low-temperature treatment, the expression of ZmDHN11 reached its maximum value in 6 h, and then decreased to below the normal level. Under ABA treatment, the expression of ZmDHN11 reached a maximum at 6 h then decreased to the normal level. Under salt treatment, the expression of ZmDHN11 reached its maximum value at 12 h and then returned to the normal level. The results demonstrated that the transcript accumulation of ZmDHN11 could be induced by high osmotic stress, low temperature, NaCl, and ABA.
Transcript accumulation of ZmDHN11 in response to abiotic stresses as determined by qRT-PCR. The maize seedlings were incubated with Hoagland’s solution for 2 weeks; Uniformly sized plants at similar growth stages were chosen for further study. Maize seedlings were treated with 20% PEG6000 (w/v), low temperature (4 °C), 100 μM ABA, 150 mM NaCl, and water (control). Total RNA was isolated from maize leaves at the indicated times after treatment
Subcellular localization of ZmDHN11
The subcellular localization of proteins often determines their function. To determine the cellular localization of ZmDHN11, leaf epidermal cells of transgenic tobacco plants expressing the ZmDHN11-GFP fusion protein were examined by a Leica confocal laser scanning microscope. As shown in Fig. 3, the ZmDHN11–GFP fusion protein specifically accumulated in the nuclei and cytosol.
ZmDHN11 could be phosphorylated by casein kinase II
Casein kinase II (CKII) is a constitutively active serine/threonine protein kinase composed of two catalytic α-subunits (44 kDa) and two regulatory β-subunits (26 kDa) in an α2β2 configuration that forms stable heterotetramers (Liu et al. 2017b). In the present study, the signal strength of the phosphorylated protein ZmDHN11 was stained with the Pro-Q Diamond Phosphoprotein Gel Staining Kit in vitro in phosphorylation assays (Fig. 4). The results showed that the ZmDHN11 protein could be phosphorylated by CKII.
ZmDHN11 protected LDH (lactate dehydrogenase) activity under water-deficit stress
We tested the ability of ZmDHN11 to prevent damage to LDH activity under drought conditions with or without ZmDHN11. After three cycles, LDH (alone) retained 30% of its initial activity, while when ZmDHN11 was added, LDH retained 50% of its initial activity. Highly hydrophilic proteins, such as poly-lysine and lysozyme, did not affect the deactivation rate of LDH under drought stress (Fig. 5). These results suggest that ZmDHN11 protects LDH activity under drought conditions.
ZmDHN11 protein could protect LDH (lactate dehydrogenase) from inactivation by water -deficit stress. One drying cycle corresponded to vacuum drying for 1 h in a modified tray freeze-dryer followed by immediate rehydration in water to the original volume. The poly-lysine and lysozyme proteins were used as controls. The enzymatic activity of untreated LDH is referred to as 100%. Each curve/column represents an average of three replicates, and error bars represent the standard deviation. Statistical significance of the difference was confirmed by Student’s t test, **P < 0.01, *P < 0.05
Overexpression of ZmDHN11 enhanced yeast cell tolerance to osmotic stress
To determine the function of the ZmDHN11 protein on the survival ratio of yeast recombinants under osmotic stresses, the growth curves of the yeast cell lines transformed with the pPI3.5 k-ZmDHN11 vector and the control lines containing the empty vector (pPI3.5 k) were measured under osmotic stress conditions (Fig. 6a). There was no significant difference between the control and the transformed yeast under optimal conditions. However, the lag phase of the transformed yeasts was shorter than that of the control, and the transformed yeast displayed higher growth compared to the control under osmotic stress conditions (Fig. 6b). According to these results, we could conclude that overexpression of ZmDHN11 could enhance the tolerance of transformed yeast to water-deficit stress.
Overexpression of ZmDHN11 enhances tolerance to copper stress in transformant yeast (GS115) tolerant to osmotic stress. a The transformant yeast was grown in nonstress BMGY medium. b The transformant yeast was grown in BMGY medium supplemented with 800 mM mannitol. c Accumulation of the ZmDHN11 in transgenic yeasts. Each curve/column represents an average of three replicates, and error bars represent the standard deviations
Overexpression of the ZmDHN11 fusion protein enhanced transgenic tobacco tolerance to osmotic stress
Osmotic stress can cause ROS accumulation, which could lead to damage to DNA, protein and plant cells (Liu et al. 2019a, b). Antioxidant enzymes such as POD and SOD can protect plants from damage caused by the accumulation of ROS. To investigate the osmotic stress tolerance of transgenic tobacco, three independent lines (ZmDHN11-1, ZmDHN11-2, and ZmDHN11-3) with ZmDHN11 overexpression were selected for analysis (Fig. 7). After drought treatment, the tobacco was stained with NBT. Under optimal growing conditions, there were no significant differences in the accumulation of ROS between transgenic plants and the control. However, overexpression of ZmDHN11 reduced the accumulation of ROS under drought conditions (Fig. 8). The relative electrolyte leakage and MDA content can represent membrane injury. There were no significant differences in the relative electrolyte leakage and MDA between transgenic plants and the control under optimal conditions. However, the relative electrolyte leakage and the MDA content of transgenic plants and control plants increased significantly under drought stress, and the increase in the relative electrolyte leakage and the MDA content in transgenic tobacco plants was less than that of the control plants (Fig. 8).
The antioxidant enzymes of SOD and POD can reduce the accumulation of ROS. There were no significant differences between transgenic plants and the control under optimal conditions. However, the activity of these antioxidant enzymes in transgenic plants and control plants increased significantly, and the increase in the transgenic tobacco plants was higher than that in control plants (Fig. 9).
Water deficit tolerance assay in ZmDHN11-overexpressing transgenic tobacco plants. MDA, relative electrolyte leakage and the activity of antioxidant enzymes (SOD and POD) in 6-week-old transgenic and control tobacco plants were measured at the indicated times after treatment with 20% PEG6000. Statistical significance of the difference was confirmed by Student’s t test, **P < 0.01, *P < 0.05
Lipid peroxidation was measured by analyzing MDA levels. Electrolyte leakage always occurs following membrane damage. We analyzed MDA content and relative electrolyte leakage in WT and transgenic plants under osmotic stress conditions, and found that the levels of relative electrolyte leakage and MDA in the overexpression plants were remarkably lower than those in WT plants. The SOD and POD activities in the three overexpressing lines were significantly higher than those in the WT after 1 day of drought stress. These results suggest that overexpression of ZmDHN11 improves transgenic tobacco tolerance to osmotic stress.
Discussion
As sessile organisms, environmental stress, such as water deficit, has a serious impact on plants (Zhang et al. 2012; Dou et al. 2015; Li et al. 2019). To respond to abiotic stress, plants produce a series of hydrophilic proteins to protect cell metabolism. Late embryogenesis abundant (LEA) proteins are hydrophilic proteins that were first identified as proteins abundant in the later stages of seed development (Liu et al. 2016). Environmental stresses, such as drought, low temperature, and high salinity, affect plant growth by disturbing the water balance. Many studies have demonstrated that LEA proteins play crucial roles in reducing the damage caused by environmental stress (Xing et al. 2011; Chen et al. 2016; Yu et al. 2019). ZmDHN11 belongs to the group II LEA protein family, also known as dehydrin. In the present study, the transcription of ZmDHN11 could be induced by drought, low temperature, high salt, and the application of ABA.
It is well known that some dehydrins are phosphorylated and that this modification influences their interaction with other proteins and membranes and might affect subcellular localization (Rorat 2006). Phosphorylation has been widely found in dehydrins containing the S-segment. Maize dehydrins Rab17 and ZmDHN13 can be phosphorylated by CKII, which can influence the dis distribution between the cytoplasm and nucleus (Riera et al. 2004; Liu et al. 2017b; Liu et al. 2019a, b).
The phosphorylated dehydrins COR47 and ERD10 could regulate calcium binding (Alsheikh et al. 2005). Dehydrin ERD14 can be phosphorylated by the protein kinase SnRK2.10, and the phosphorylation of ERD14 within the S-segment is involved in the regulation of dehydrin subcellular localization in response to stress (Maszkowska et al. 2019). In this study, protein kinase CKII phosphorylated ZmDHN11. Further study indicated that the ZmDHN11–GFP fusion protein accumulated in the nuclei and cytosol in transgenic tobacco leaf epidermal cells. We speculated that the location of ZmDHN11 might be influenced by phosphorylation.
As the reporter enzyme, lactate dehydrogenase (LDH) is sensitive to water loss (Goyal et al. 2005). LDH has been widely used to test the protective activity of LEA proteins (Liu et al. 2013b, 2014; Yang et al. 2015; Hara et al. 2016). LEA proteins have been proposed to be involved in the stabilization and protection of cell structures and macromolecules by ion sequestration, stabilizing the structure of membranes and proteins. Here, ZmDHN11 protected the activity of LDH from damage caused by water-deficit in vitro. There was a significantly reduced increase in endogenous ROS, MDA, and relative electrolyte leakage levels in transgenic tobacco than in the WT plants. Overexpression of ZmDHN11 in the transgenic plants increased the activities of the antioxidant enzymes POD and SOD compared those in control plants under drought stress conditions. Further study indicated that overexpression of ZmDHN11 enhanced transgenic yeast and tobacco tolerance to drought stress. It is reasonable to speculate that the ZmDHN11 protein could stabilize the structure of macromolecules such as membranes and protect the activity of antioxidant enzymes that can decrease the accumulation of reactive oxygen species and prevent damage to proteins and membranes.
In conclusion, this study identified a novel group II LEA gene ZmDHN11, which is inducible in response to a variety of abiotic stresses. ZmDHN11 is a KS- type dehydrin. In vitro experiments showed that ZmDHN11 can be phosphorylated by the protein kinase CKII. Further study indicated that the ZmDHN11 protein could protect LDH activity from damage caused by water-deficit stress in vitro. Overexpression of ZmDHN11 could enhance transgenic yeast and tobacco tolerance to osmotic stress.
References
Alsheikh MK, Svensson JT, Randall SK (2005) Phosphorylation regulated ion-binding is a property shared by the acidic subclass dehydrins. Plant Cell Environ 28(9):1114–1122. https://doi.org/10.1111/j.1365-3040.2005.01348.x
Amara I, Capellades M, Ludevid MD et al (2013) Enhanced water stress tolerance of transgenic maize plants over- expressing LEA Rab28 gene. J Plant Physiol 170(9):864–873. https://doi.org/10.1016/j.jplph.2013.01.004
Battaglia M, Olvera-Carrillo Y, Garciarrubio A et al (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148(1):6–24. https://doi.org/10.1104/pp.108.120725
Cai G, Wang G, Wang L et al (2014) A maize mitogen-activated protein kinase kinase, ZmMKK1, positively regulated the salt and drought tolerance in transgenic Arabidopsis. J Plant Physiol 171(12):1003–1016. https://doi.org/10.1016/j.jplph.2014.02.012
Celik AY, Baloglu P, Yer EN et al (2016) Identification and expression analysis of LEA gene family members in cucumber genome. Plant Growth Regul 80(2):225–241. https://doi.org/10.1007/s10725-016-0160-4
Chen J, Fan L, Du Y et al (2016) Temporal and spatial expression and function of TaDlea3 in Triticum aestivum during developmental stages under drought stress. Plant Sci 252:290–299. https://doi.org/10.1016/j.plantsci.2016.08.010
Close TJ (1997) Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiol Plant 100(2):291–296. https://doi.org/10.1111/j.1399-3054.1997.tb04785.x
Dou H, Xv K, Meng Q, Li G, Yang X (2015) Potato plants ectopically expressing Arabidopsis thaliana CBF 3 exhibit enhanced tolerance to high-temperature stress. Plant Cell Environ 38(1):61–72. https://doi.org/10.1111/pce.12366
Dure L, Greenway SC, Galau GA (1981) Developmental biochemistry of cottonseed embryogenesis and germination: changing messenger ribonucleic acid populations as shown by in vitro and in vivo protein synthesis. Biochemistry 20(14):4162–4168. https://doi.org/10.1021/bi00517a033
Garay A, Colmenero FJM, Garciarrubio A et al (2000) Highly hydrophilic proteins in prokaryotes and eukaryotes are common during conditions of water deficit. J Biol Chem 275(8):5668–5674. https://doi.org/10.1074/jbc.275.8.5668
Goyal K, Walton L, Tunnacliffe A (2005) LEA proteins prevent protein aggregation due to water stress. Biochem J 388:151–157. https://doi.org/10.1042/BJ20041931
Hara M, Terashima S, Fukaya T, Kuboi T (2003) Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta 217(2):290–298. https://doi.org/10.1007/s00425-003-0986-7
Hara M, Monna S, Murata T et al (2016) The Arabidopsis KS-type dehydrin recovers lactate dehydrogenase activity inhibited by copper with the contribution of his residues. Plant Sci 245:135–142. https://doi.org/10.1016/j.plantsci.2016.02.006
Houde M, Dallaire S, N’Dong D et al (2004) Overexpression of the acidic dehydrin WCOR410 improves freezing tolerance in transgenic strawberry leaves. Plant Biotechnol J 2(5):381–387. https://doi.org/10.1111/j.1467-7652.2004.00082.x
Jiang S, Zhang D, Wang L et al (2013) A maize calcium-dependent protein kinase gene, ZmCPK4, positively regulated abscisic acid signaling and enhanced drought stress tolerance in transgenic Arabidopsis. Plant Physiol Biochem 71:112–120. https://doi.org/10.1016/j.plaphy.2013.07.004
Koag MC, Wilkens S, Fenton RD et al (2009) The K-segment of maize DHN1 mediates binding to anionic phospholipid vesicles and concomitant structural changes. Plant Physiol 150(3):1503–1514. https://doi.org/10.1104/pp.109.136697
Li J, Wang Y, Yu B et al (2018) Ectopic expression of StCBF1 and ScCBF1 have different functions in response to freezing and drought stresses in Arabidopsis. Plant Sci 270:221–233. https://doi.org/10.1016/j.plantsci.2018.01.015
Li D, Zhang T, Wang M et al (2019) Genetic engineering of the biosynthesis of glycine betaine modulates phosphate homeostasis by regulating phosphate acquisition in tomato. Front Plant Sci 9:1995. https://doi.org/10.3389/fpls.2018.01995
Liu Y, Wang L, Cai GH et al (2013a) Response of tobacco to the Pseudomonas syringae pv. Tomato DC3000 is mainly dependent on salicylic acid signaling pathway. FEMS Microbiol Lett 344(1):77–85. https://doi.org/10.1111/1574-6968.12157
Liu Y, Wang L, Xing X et al (2013b) ZmLEA3, a multifunctional group 3 LEA protein from maize (Zea mays L.), is involved in biotic and abiotic stresses. Plant Cell Physiol 54(6):944–959. https://doi.org/10.1093/pcp/pct047
Liu Y, Wang L, Jiang SS et al (2014) Group 5 LEA protein, ZmLEA5C, enhanced transgenic tobacco and yeast tolerance to osmotic and low temperature stresses. Plant Physiol Biochem 84:22–31. https://doi.org/10.1016/j.plaphy.2014.08.016
Liu Y, Liang J, Sun L et al (2016) Group 3 LEA protein, ZmLEA3, is involved in protection from low temperature stress. Front Plant Sci 7:1011. https://doi.org/10.3389/fpls.2016.01011
Liu Y, Song QP, Li DX et al (2017a) Multifunctional roles of plant dehydrins in response to environmental stresses. Front Plant Sci 8:1018. https://doi.org/10.3389/fpls.2017.01018
Liu Y, Wang L, Zhang T et al (2017b) Functional characterization of KS-type dehydrin ZmDHN13 and its related conserved domains under oxidative stress. Sci Rep 7(1):7361. https://doi.org/10.1038/s41598-017-07852-y
Liu W, Zhao BG, Chao Q et al (2019a) Function analysis of ZmNAC33, a positive regulator in drought stress response in Arabidopsis. Plant Physiol Biochem 145:174–183. https://doi.org/10.1016/j.plaphy.2019.10.038
Liu Y, Li DX, Song QP, Li D, Yang XH (2019b) The maize late embryogenesis abundant protein ZmDHN13 positively regulates copper tolerance in transgenic yeast and tobacco. Crop J 7(3):403–410. https://doi.org/10.1016/j.cj.2018.09.001
Malik AA, Veltri M, Boddington KF et al (2017) Genome analysis of conserved dehydrin motifs in vascular plants. Front Plant Sci 8:709. https://doi.org/10.3389/fpls.2017.00709
Maszkowska J, Janusz D, Kulik A et al (2019) Phosphoproteomic analysis reveals that dehydrins ERD10 and ERD14 are phosphorylated by SNF1-related protein kinase 2.10 in response to osmotic stress. Plant Cell Environ 42(3):931–946. https://doi.org/10.1111/pce.13465
Miller G, Suzuki N, Ciftci-Yilmaz S et al (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33(4):453–467. https://doi.org/10.1111/j.1365-3040.2009.02041.x
Olvera-Carrillo Y, Campos F, Reyes JL et al (2011) Functional analysis of the group 4 late embryogenesis abundant proteins reveals their relevance in the adaptive response during water deficit in Arabidopsis. Plant Physiol 154(1):373–390. https://doi.org/10.1104/pp.110.158964
Pan J, Zhang M, Kong X et al (2012) ZmMPK17, a novel maize group D MAP kinase gene, is involved in multiple stress responses. Planta 235(4):661–676. https://doi.org/10.1007/s00425-011-1510-0
Riera M, Figueras M, López C et al (2004) Protein kinase CK2 modulates developmental functions of the abscisic acid responsive protein Rab17 from maize. Proceedings of the National Academy of Sciences 101(26):9879–9884. https://doi.org/10.1073/pnas.0306154101
Rorat T (2006) Plant dehydrins—tissue location, structure and function. Cellular and Molecular Biology Letters 11(4):536–556. https://doi.org/10.2478/s11658-006-0044-0
Sivamani E, Bahieldin A, Wraith JM et al (2000) Improved biomass productivity and water use efficiency under water deficit conditions in transgenic wheat constitutively expressing the barley HVA1 gene. Plant Sci 155(1):1–9. https://doi.org/10.1016/s0168-9452(99)00247-2
Sun J, Nie L, Sun G et al (2013) Cloning and characterization of dehydrin gene from Ammopiptanthus mongolicus. Mol Biol Rep 40(3):2281–2291. https://doi.org/10.1007/s11033-012-2291-7
Tanmoy H, Gouranga U, Sudipta R (2017) YSK2 type dehydrin (SbDhn1) from Sorghum bicolor showed improved protection under high temperature and osmotic stress condition. Front Plant Sci 8:918. https://doi.org/10.3389/fpls.2017.00918
Wei F, Lindner H, Robbins NE et al (2016) Growing out of stress: the role of cell- and organ-scale growth control in plant water-stress responses. Plant Cell 28(8):1769–1782. https://doi.org/10.1105/tpc.16.00182
Wise MJ, Tunnacliffe A (2004) POPP the question: what do LEA proteins do? Trends Plant Sci 9(1):13–17. https://doi.org/10.1016/j.tplants.2003.10.012
Xing X, Liu Y, Kong X et al (2011) Overexpression of a maize dehydrin gene, ZmDHN2b, in tobacco enhances tolerance to low temperature. Plant Growth Regul 65(1):109–118. https://doi.org/10.1007/s10725-011-9580-3
Xiong L, Zhu JK (2002) Molecular and genetic aspects of plant responses to osmotic stress. Plant Cell Environ 25(2):131–139. https://doi.org/10.1046/j.1365-3040.2002.00782.x
Yang W, Zhang L, Lv H et al (2015) The K-segments of wheat dehydrin WZY2 are essential for its protective functions under temperature stress. Front Plant Sci 6:406. https://doi.org/10.3389/fpls.2015.00406
Yu Z, Wang X, Tian Y, Zhang D, Zhang L (2019) The functional analysis of a wheat group 3 late embryogenesis abundant protein in Escherichia coli and Arabidopsis under abiotic stresses. Plant Signal Behav 14(11):1667207. https://doi.org/10.1080/15592324.2019.1667207
Zhang M, Pan J, Kong X et al (2012) ZmMKK3, a novel maize group B mitogen-activated protein kinase kinase gene, mediates osmotic stress and ABA signal responses. J Plant Physiol 169(15):1501–1510. https://doi.org/10.1016/j.jplph.2012.06.008
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31701334) and the Shandong Province Natural Science Foundation (ZR2016CQ34).
Author information
Authors and Affiliations
Contributions
YL designed the experiments. HJ and DL performed the experiments and analyzed the data. HJ and YL wrote the article. DL and XY revised the paper and gave positive suggestion. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The author(s) confirm that this article content has no conflicts of interest.
Additional information
Communicated by Chun-Hai Dong.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ju, H., Li, D., Li, D. et al. Overexpression of ZmDHN11 could enhance transgenic yeast and tobacco tolerance to osmotic stress. Plant Cell Rep 40, 1723–1733 (2021). https://doi.org/10.1007/s00299-021-02734-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02734-0