Abstract
Bacillus thuringiensis is the most widely used biopesticide, targets a diversity of insect pests belonging to several orders. However, information regarding the B. thuringiensis strains and toxins targeting Zeugodacus cucurbitae is very limited. Therefore, in the present study, we isolated and identified five indigenous B. thuringiensisstrains toxic to larvae of Z. cucurbitae. However, of five strains NBAIR BtPl displayed the highest mortality (LC50 = 37.3 μg/mL) than reference strain B. thuringiensis var. israelensis (4Q1) (LC50 = 45.41 μg/mL). Therefore, the NBAIR BtPl was considered for whole genome sequencing to identify the cry genes present in it. Whole genome sequencing of our strain revealed genome size of 6.87 Mb with 34.95% GC content. Homology search through the BLAST algorithm revealed that NBAIR BtPl is 99.8% similar to B. thuringiensis serovar tolworthi, and gene prediction through Prokka revealed 7406 genes, 7168 proteins, 5 rRNAs, and 66 tRNAs. BtToxin_Digger analysis of NBAIR BtPl genome revealed four cry gene families: cry1, cry2, cry8Aa1, and cry70Aa1. When tested for the presence of these four cry genes in other indigenous strains, results showed that cry70Aa1 was absent. Thus, the study provided a basis for predicting cry70Aa1 be the possible reason for toxicity. In this study apart from novel genes, we also identified other virulent genes encoding zwittermicin, chitinase, fengycin, and bacillibactin. Thus, the current study aids in predicting potential toxin-encoding genes responsible for toxicity to Z. cucurbitae and thus paves the way for the development of B. thuringiensis-based formulations and transgenic crops for management of dipteran pests.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus thuringiensis is a Gram-positive, ubiquitous aerobic or facultative anaerobe spore-forming, rod-shaped bacteria belonging to the Bacillus cereus group (Vilas-Bôas et al. 2007). It is primarily known for its insecticidal activity against insect pests belonging to orders like Lepidoptera, Diptera, Hemiptera, Coleoptera, Hymenoptera, and Orthoptera (Höfte and Whiteley 1989; Manjunatha et al. 2023). The insecticidal activity of B. thuringiensis is predominantly due to the production of crystalline inclusions consisting of one or more proteins called δ- endotoxins (commonly called Cry and Cyt proteins). To date, nearly 700 cry genes and 40 cyt genes belong to 75 different families (Cry1-Cry75), and three families (Cyt1-Cyt3), respectively were identified (Crickmore et al. 2021).
The activity of cry genes is highly specific to one or a few closely related insect species. Hence, the characterization of B. thuringiensis strains with target-specific cry genes is very essential for the efficient management of insects of economically important insect pests. However, characterization of such strains is not an easy task and largely depends on the traditional PCR strategy using cry gene specific primer. The target cry gene in the strain can also be confirmed using DNA-microarray techniques and DNA hybridization (Letowski et al. 2005). However, these techniques are limited to the detection of previously known cry genes, and it is very hard to find novel cry genes present in a strain. Previous studies suggest that whole genome sequencing is the widely used coherent strategy for discovering novel bioinsecticidal toxin-related genes, as many strains have been sequenced so far using PacBio strategies and Illumina (Cao et al. 2018). No attempts have been made so far to identify the cry genes encoding the potential toxins targeting melon fruit fly, Zeugodacus cucurbitae (Coquillett) (Tephritidae: Diptera).
The melon fruit fly is an economically important insect pest, inflicting damage to around 130 plant species belonging to 30 families, including pumpkin, cucumber, bitter gourd, eggplant, and tomato (Ahmad et al. 2023). Crop losses attributed to this pest varied from 30 to 100% in various fruit and vegetable crops (Dhillon et al. 2005). Maggots are the damaging stages, and their concealed feeding behaviour inside fruits makes management extremely challenging. Hence, the management practices are targeted against the adult stages that largely depend on synthetic pesticides, mainly as cover sprays, due to their rapid knockdown effect (De Bon et al. 2014). However, these chemicals have adverse effects on the environment and non-target organisms. Therefore, alternative control methods are warranted against this pest. In this regard, the use of microbials like Metarhizium anisopliae and Beauveria bassiana found promising alternative method in management of different fruit fly species (Onsongo et al. 2022). Among the microbial biopesticides, B. thuringiensis based biopesticides are highly effective and B. thuringiensis based biopesticides successfully used for the management of insects pests in the field condition (Kumar et al. 2021). However, information regarding the B. thuringiensis strains and the Cry toxins effective against Z. cucurbitae is very limited. Accordingly, in this study, we aimed to isolate and characterize the B. thuringiensis strains effective against maggots of Z. cucurbitae, we further focused on whole genome sequence of virulent strains to identify the insecticidal toxin-related genes responsible for causing mortality in larvae of Z. cucurbitae.
Materials and methods
Sample collection
A total of 50 samples, including soil and insect cadavers, were collected aseptically from different parts of Karnataka. Details of collection sites are given in Table 1.
Isolation of B. thuringiensis from soil and insect cadaver
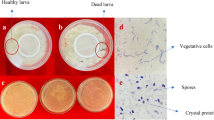
Approximately, 10 g of each soil sample was grounded under aseptic conditions separately in a porcelain mortar for 10 min followed by washing with sterile distilled water to remove debris. One gram of each soil sample is added to 125 mL of flask containing 10 mL of Luria–Bertani broth buffered with 0.25 M sodium acetate (pH 6.8) was After incubation at 30 °C for 4 hours (h) the cultures were heated at 80 °C for 15 min to kill vegetative cells. From each sample, 100 μL were spread on T3 agar plates then incubated for 72 h at 30 °C. This technique abled the purchase of spores that germinated, multiplied, and entered in sporulation phase. To confirm the presence of spores and crystals related to the presence of B. thuringiensis isolates, pure colonies obtained on T3 agar plates were investigated under light microscope. Then, each parasporal crystal forming isolate was stored and considered as B. thuringiensis (Travers et al. 1987). Similarly, to isolate B. thuringiensis from insect cadaver, insect body was sterilized by immersing in 0.85% sterile saline solution to eliminate the surface microflora of the body and it is washed twice with distilled water. Later, the insect was crushed in a 1.5 mL Eppendorf tube containing PBS buffer (1×), and the contents were heated to 80 °C in a water bath for 40 min. Further, 1 mL of the aliquot was used to isolate B. thuringiensis following the procedure used in isolation of B. thuringiensis from soil.
Microscopic examination of crystal proteins
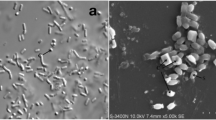
All isolates were cultured in LB broth at room temperature (28 ± 2 °C) with agitation on an orbital shaker at 250 rpm until reaching the autolysis phase. A single drop of the culture was spread onto a microscope glass slide then air-dried and heat-fixed. Then the slide was stained with a 0.25% solution of Coomassie Brilliant Blue (CBB) in 50% acetic acid for less than 2 min (Rampersad and Ammons 2005). Subsequently, it was destained for about 2 min with 70% ethanol, rinsed with distilled water, dried, and examined under a light microscope (Olympus BX41, Microscope Central, Pennsylvania, USA) using a 100× oil immersion objective lens. Isolates positive for crystal proteins were given a code for further identification. Further, investigation for parasporal crystal presence was done using SEM as described by Loutfi et al. (2020). About 50 μL of the spore-crystal mixture was placed on a microscopic glass slide and fixed by drying in an oven at 37 °C, and then sputter coated with gold. Finally, the slide containing crystals was viewed with SEM at 2500× (Quanta 250, FEI, Oregon, USA).
Amplification and sequencing of 16Sr RNA gene
The 16S rRNA gene of each B. thuringiensis isolate was amplified using primers 27F (5ʹ-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-TACGGYTACCTTGTTACGACTT-3′) (Dos Santos et al. 2019). The amplified product is further sequenced by Sanger sequencing (Eurofins India Private Limited, Bengaluru, India). The sequences were further analysed using NCBI Blastn and further deposited in GenBank, and accession numbers were obtained.
Insect rearing
Melon fruit fly culture maintained at ICAR-National Bureau of Agriculture Insect Resources was used for the present study. Ten pairs of males and females were collected from the mother culture and stocked in a glass cage covered with a net on three sides of the cages for rearing to get the homogenous culture. Culture was maintained at a temperature of 25 ± 2 °C with a relative humidity of 70–80% at the Insect Bacteriology Lab, ICAR-NBAIR, Bengaluru, Karnataka, India. For rearing, adult flies were offered yeast for feeding and bananas for egg-laying. The eggs were collected gently using a camel brush and seeded over the artificial diet for further larval development according to the composition given by Liu et al. (2020).
Biotoxicity assay of B. thuringiensisstrains against melon fruit fly
The toxicity of five indigenous strains, along with the reference B. thuringiensis svar. israelensis 4Q1was assessed against 3rd instar larvae of melon fruit fly under in vitro conditions. All strains, including 4Q1, inoculated separately on LB broth for 5 to 6 days. The inoculated broth was centrifuged at 10,000 rpm (Sigma 3-30KHS; 22096), and the pellet containing the spore-crystal mixture was resuspended with double distilled water. The protein concentration was estimated by Lowry’s methods using bovine serum albumin (BSA) as a standard following the most commonly used protocol (Lowry et al. 1951). Exactly, 100 µL of spore crystal homogenate was mixed with 10 g of artificial diet in a small plastic container (3 × 3 cm), and each container was considered a replicate. Three replications were given for each treatment. The reference strain 4Q1 and diet without spore crystal suspension of B. thuringienis treatment were used as negative control. The bioassay was carried out under controlled conditions of 25 ± 2 °C and 70 ± 5% relative humidity. The percentage mortality was calculated at 24 h intervals till the fifth day of the experiment. The confirmation of death is due to B. thuringiensis strains was done by crushing the individual dead larvae in PBS (1X) then suspension was spread on nutrient agar plates. Finally, bacterial colonies were obtained and tested for the presence of spores and crystals. The concentration–response relationships for larval mortality data were used to calculate lethal concentration (LC50) of each B. thuringiensis strain corresponding 95% fiducial limit (95% FL) by using PROC PROBIT in SAS software (version 9.3, 2011; SAS Institute, Cary, NC, USA).
Whole genome sequencing of Bacillus thuringiensis strain NBAIR BtPl
Bacillus thuringiensis strain NBAIR BtPl was selected for whole genome sequencing to identify the insecticidal toxin-related genes responsible for causing mortality in larvae of Z. cucurbitae. The NBAIR BtPl strain caused significantly greater mortality compared to the five B. thuringiensis strains isolated in this study and the reference 4Q1 strain.
DNA extraction, library preparation, and sequencing
One-day-old culture of the NBAIR BtPl strain was used for the extraction of DNA using the DNeasy Blood and Tissue Kit following the manufacturer’s protocol (Qiagen, Hilden, Germany). The quality and quantity of isolated DNA were checked using Nanodrop and agarose gel electrophoresis, respectively. From QC-passed DNA samples, paired-end sequencing libraries were prepared using the Illumina TruSeq Nano DNA Library prep Kit. Approximately 100 ng of DNA was subjected to ultrasonication (Covaris M220, Massachusetts, USA) to generate the mean fragments of 350 bp dsDNA fragments containing 3ʹ and 5ʹ overhangs. These overhangs were removed by subjecting fragments to end-repair using the 3ʹ and 5ʹ exonuclease enzymes and then to adapter ligation using the 5ʹ and 3ʹ polymerase enzymes. The ligated products were size-selected using AMPure XP beads, and the products were PCR amplified with index primers. The ends of DNA fragments were ligated with Indexing adapters to, prepare them for hybridization onto flow cell. The analysis of these PCR-enriched libraries was done using a 4200 tape station system with high-sensitivity D10000 screen tape following the manufacturer’s protocol. After obtaining the Qubit concentration for the libraries and mean peak sizes from the alignment tape station profile, the PE Illumina libraries were loaded onto NextSeq500 for cluster generation and sequencing.
Genome assembly and functional annotation
The obtained raw data of the NBAIR BtPl strain was processed to remove low-quality reads, ambiguous reads, adapter sequences, and then the high-quality reads were obtained using Trimmomatic version 0.36 (Bolger et al. 2014). The filtered, high-quality reads were assembled into scaffolds using the SPAdes-assembler (Bankevich et al. 2012). Then assembled scaffolds were further analyzed to identify the most closely related organisms using a homology-based approach. The genome of NBAIR BtPl was used for reference-based scaffolding for the scaffolds using GFinisher (Kremer et al. 2017). Gene prediction was performed by Prokka (Seeman 2014), and the sequences of predicted genes of NBAIR BtPl were searched against the NCBI nonreductant protein database using the Basic Local Alignment Search Tool (BlastX) (E-Value: 1e−05) through the Diamond tool (Buchfink et al. 2015). Gene ontology annotations of the genes were determined by the Blast2GO platform (Götz et al. 2008). Gene ontology assignments were used to classify the functions of predicted genes. Gene ontology mapping was done, and the gene products were grouped into three main domains. The pathway annotation of identified genes was carried out against the KEGG GENES database using KAAS (KEGG Automatic Annotation Server) (Moriya et al. 2007). Further, the gene clusters encoding for potential secondary metabolites in NBAIR BtPl were predicted with antiSMASH version 3.0 using default parameters as well as the ClusterFinder algorithm (Weber et al. 2015).In addition, the genome similarity between the B. thuringiensis NBAIR BtPl with other B. thuringiensis strains such as B. thuringiensis serovar kurstaki strain HD-1 (CP010005.1), B. thuringiensis serovar tolworthi (AP014864.1), B. thuringiensis serovar israelensis strain AR23 (NZ_JABXXM010000014.1) was done using Proksee (https://proksee.ca).
Identification and validation of insecticidal toxicity-related genes (ITRGs)
BT Toxin_Digger was used for the identification of ITRGs present in NBAIR BtPl (Liu et al. 2022). The identified ITRGs were further validated in the PCR using the gene-specific primers that were designed using Primer3Plus. All PCR reactions were carried out with 12.5 µL of PCR master mix (EmeraldAmp GT PCR master mix, TakaRa, Japan) with 2 µL of 50 ng/µL template DNA and 1 µL of 10 pmol from each primer in a thermal cycler (T100, Bio-Rad, California, USA). The cycle started with an initial denaturation for 5 min at 95 °C, 30 cycles of denaturation for 30 s at 95 °C, and annealing for 1 min at a temperature specific to the primer, a 72 °C extension step was carried out for 1 min, and then the final extension for 10 min at 72 °C. The amplified product was subjected to gel electrophoresis, and the product was visualized under a 1.2% agarose gel and documented using the MiniLumi gel documentation system (DNR, Israel).
PCR amplification of cry genes
Total DNA was extracted from all B. thuringiensis isolates using the DNeasy Blood and Tissue Kit as per the manufacturer’s protocol. The DNA of each strain was individually used as a template for the amplification of the dipteran active cry gene (Valtierra-de-Luis et al. 2020). The reaction was set up at 94 °C as pre-denaturation for 4 min and followed by 35 cycles of denaturation at 94 °C for 1 min, annealing for 1 min at the temperature specific to each primer, extension for 1 min at 72 °C, and a 10 min final extension at 72 °C. The annealing temperatures for each primer are given in Supplemental Table 1. The amplified products were visualized using 1.2% agarose gel, and documented under the gel documentation system. Further, the crystals proteins present in the individual strain were analyzed following Alves et al. (2023).
Analysis of aminoacid sequence and evolutionary analysis of cry70Aa1
The aminoacid sequence homology of cry70 family members was identified using NCBI nucleotide–nucleotide BLAST and protein–protein BLAST (http://www.ncbi.nlm.nih.gov/BLAST). The aminoacid sequence alignment was done using ClustalW software. Phylogenetic analysis of cry70 family members was done using MEGA 11.0 software.
Results
Isolation and identification of B. thuringiensis isolates
Out of 50 samples examined, five were identified as B. thuringiensis NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, and NBAIR BtPl on the basis of morphological and molecular characteristics. We observed bipyramidal shaped crystal proteins for NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, and NBAIR BtPl under light microscopy. Whereas the spherical-shaped crystal proteins were observed in only NBAIR BtPl. In scanning electron microscopic, we also observed spores, bipyramidal, spherical, and cuboidal-shaped crystal proteins very distinctly for NBAIR BtPl (Fig. 1). Further, plasmid profiling and the taxonomic characterization of all five isolates was confirmed by sequencing the 16S rRNA gene, which displayed an amplicon size of 1300–1500 bp on agarose gel (Supplemental Fig. 1). The amplified 16S rRNA gene product was sequenced and compared with international standard B. thuringiensis strains collected from the National Centre for Biotechnology Information (NCBI, USA) database. The nucleotide sequence of five isolates showed 99–100% homology with internationally standard B. thuringiensis strains through blastn analysis. Therefore, these all five isolates were considered B. thuringiensis, then sequences finally deposited in the NCBI database and accession numbers were obtained (Table 1).
Biotoxicity assay of B. thuringiensis strains against melon fruit fly
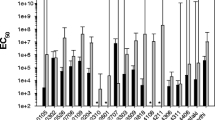
The third instar larvae were found to be lethargic, and their gut region turned black after feeding the artificial diet treated with indigenous B. thuringiensis strains and reference strain 4Q1. The present study results showed that all five tested B. thuringiensis strains caused morality in third instar of Z. cucurbitae. But the NBAIR BtPl caused greater mortality compared to the other four indigenous B. thuringiensis strains and reference strain 4Q1. The average mortality in third instar larvae of Z. cucurbitae was between 40 and 90% due to B. thuringiensis strains. No larval mortality was observed in untreated control treatment. The results of the probit analysis showed that the LC50 values for all the tested B. thuringiensis strains in this study were between 37.30 and 106.70 μg/mL. The LC50 value recorded for the NBAIR BtPl was 37.30 μg/mL, where as 45.41 μg/mL for reference strain (4Q1). These results showed that the NBAIR BtPl was more virulent than 4Q1 strain (Table 2).
Genome analysis of NBAIR BtPl
Sequencing of NBAIR BtPl revealed a total number of 1,14,07,645 reads (PE) and 3,37,10,34,441 bases and a total of 5 rRNAs and 66 tRNAs were identified. The assemblage of these filtered high-quality reads using the SPAdes assembler showed a total number of 726 scaffolds with 77,05,163 bp and an average scaffold length of 9121 bp. We also recorded the maximum scaffold length of 1,70,233 bp and minimum scaffold length 534 bp for the genome of NBAIR Btpl. Reference-based scaffolding showed the genome having a total length of 6.87 Mb and a GC content of 34.95% with scaffolds 44,203, 368,302, and 412 bp as average, maximum, and minimum scaffolds lengths, respectively. The gene prediction by Prokka identified genome of NBAIR BtPl as having 7406 genes and 7168 protein-coding sequences, with a maximum and maximum length of 12,442 and 39 bp, respectively. The average length of NBAIR BtPl gene was 761 bp. (Table 3). The top blast hit species distribution of the genome of NBAIR BtPl showed that most of the genes found were homologous to Bacillus thuringiensis serovar tolworthi (Fig. 2). Moreover, the average nucleotide identity (ANI) between B. thuringiensis NBAIR BtPl with other B. thuringiensis strains revealed that NBAIR BtPl has 99.8% similarity with strain B. thuringiensis serovar tolworthi. Similarly, B. thuringiensis NBAIR BtPl has ANI of 96.9 and 96.1% with B. thuringiensis HD1 and B. thuringiensis israelensis, respectively (Fig. 3).
Average Nuclotide Identity (ANI) analysis of B. thuringiensis NBAIR BtPl with other B. thuringiensis strains. a ANI between B. thuringiensis NBAIR BtPl and B. thuringiensis HD1; b ANI between B. thuringiensis NBAIR BtPl and B. thuringiensis serovar tolworthi; c ANI between B. thuringiensis NBAIR BtPl and B. thuringiensis israelensis
Gene ontology (GO) analysis of NBAIR BtPl genes using the Blast2GO platform classified the predicted genes into three main domains, such as molecular function, biological process, and cellular components. Genes annotated with gene ontology revealed the maximum number of genes present in the genome of NBAIR BtPl involved in molecular function (3037), followed by genes involved in biological processes (2347) and cellular components (2022).
The results of gene ontology analysis of genes present in NBAIRBtPl genome are represented in pie charts and WEGO plot (Fig. 4). The pathway annotation of predicted genes using KASS is done, and the genes are categorized into five main categories, such as metabolism, genetic information processing, environmental information processing, cellular processes, and organismal system. A total of 1985 KEGG-annotated genes with 23 KEGG categories are present in NBAIR BtPl (Table 4). This strain also carrying two gene clusters encoding non-ribosomal peptide synthetase (NRPS). Out of these two gene clusters, one was found to have 85% similarity with bacillibactin and 81% with zwittermicin A. In this study, we also found that the NBAIR BtPl gene cluster encoding for molybdenum co-factor, lanthipeptide class ii, and Ripp (Fig. 5).
Gene Ontology analysis of genes present in the genome of Bacillus thuringiensis strain NBAIR BtPl. a Pie chart describing the category of genes of NBAIR BtPl falling under biological process; b Pie chart describing the category of genes of NBAIR BtPl falling under cellular components; c Pie chart describing the category of genes of NBAIR BtPl falling under molecular functions; d WEGO plot of assembled genes of NBAIR BtPl
Profiling and validation of insecticidal toxicity-related genes (ITRGs)
BTtoxin_Digger was used to mine the insecticidal toxicity-related genes present in B. thuringiensis strain NBAIR BtPl. A total of sixteen ITRGs were identified (Table 5) and out of these, 12 genes validated using PCR (Table 6; Fig. 6).
Gel picture showing the insecticidal gene profile of B. thuringiensis strain NBAIR BtPl. Lane M Ladder (100 bp). Lane 1 cry1Ab9, Lane 2 cry1Ab14, Lane 3 cry1Ea10, Lane 4 cry1Gc1, Lane 5 cry1AaIa44, Lane 6 cry2Aa9, Lane 7 cry2Ab4, Lane 8 cry8Aa1, lane9 cry70Aa1, Lane10 cyt1Da2, Lane 11 vip3Aa59, Lane 12 Vpb4Aa1, Lane 13 Zwa5A, Lane 14 Zwa5B, Lane 15 Zwa6
PCR amplification of cry genes
All the identified cry genes in a strain of NBAIR BtPl belong to four families: cry1, cry2, cry70Aa1, and cry8Aa1. To dwell into the cry gene profiles of each strain and correlate them with mortality percentage, we tested the remaining four indigenous strains for amplification of the four cry genes mentioned above. Then cry gene profile revealed that cry1 tested positive for all five strains but cry2 tested positive in only two strains (NBAIR Bt154 and NBAIR BtPl). Similarly, cry8Aa1 is present only in NBAIR Bt151 and NBAIR BtPl. None of the isolates tested positive for cry70Aa1, except NBAIR BtPl. Further SDS PAGE analysis confirmed the presence of crystal proteins in all five indigenous B. thuringiensis strains (Supplemental Fig. 2).
Analysis of aminoacid sequence and evolutionary analysis of cry70Aa1
Amino acid sequence comparison between cry70 family members with cry70 member present in the NBAIR BtPl revealed that there is a variation of 328 aminoacids with maximum variation observed at 701–780 bp length of cry70Aa1 of NBAIR BtPl (Fig. 7). Further phylogenetic tree analysis revealed that higher branch length for cry70Aa1 present in NBAIR BtPl. This depicts that greater genetic diversity exists in cry70Aa1 in comparison to other cry70 members of study Supplemental Fig. 3).
Amino acid sequence similarity relationships of the Bacillus thuringiensis NBAIR BtPl cry70Aa1 with other cry70 family members. The * mark indicates the conserved aminoacid among cry70 family members the arrow indicates the variation in aminoacid observed in Bacillus thuringiensis NBAIR BtPl cry70Aa1
Discussion
In the current study, we isolated and characterized five indigenous B. thuringiensis strains. Of the five strains, three were isolated from soil and one from naturally infected larvae of P. xylostella. All five indigenous strains and reference strain (4Q1) were tested for virulence against the third instar larvae of Z. cucurbitae. Bioassay results revealed that out of five strains tested for toxicity, only NBAIR BtPl displayed greater toxicity compared to 4Q1. Hence, NBAIR BtPl strain was considered for the whole genome sequencing to identify the insecticidal toxin related genes responsible for the toxicity against Z. cucurbitae. The genomic size of NBAIR BtPl is nearly 6.87 Mb with a GC content of 34.95%. Similarly, Yilmaz et al. (2022) also found a genome size and GC content of 6.32 Mb and 34.68%, respectively in Bacillus thuringiensis strain SY49.1 toxic to mosquitoes.
As the toxicity of B. thuringiensis is mainly because of cry genes, we further delved into the cry gene profile of NBAIR BtPl and compared it with 4Q1. The reference strain 4Q1 harbors a combination of cry genes, including cry4Aa, cry4Ba, cry10Aa, cry11Aa, cyt1Aa, and cyt2Ba located on plasmid pBtoxis (Ben-Dov et al. 1999; Guerchicoff et al. 1997). When we checked the presence of these cry genes in NBAIR BtPl using genes-specific primers, we found that NBAIR BtPl strain showed negative for these genes. Nevertheless, in NBAIR BtPl we also found cry70Aa1 along with other cry genes, the presence of cry70Aa1 may have had a significant impact on toxicity to Z. cucurbitae. These results were in consistent with the results of Fayad and co-workers. They have identified a novel cry gene, such as cry70B, which was identified in strain B. thuringiensis H3 and reported to have an anti-dipteran activity (Fayad et al. 2021). The same gene is absent in the 4Q1 and in all indigenous strains except for NBAIR BtPl, which contains cry70Aa1, showing 78% similarity with cry70B. Taking this into consideration, this suggests that cry70Aa1 can be a potential cry toxin of NBAIR BtPl responsible for causing toxicity to Z. cucurbitae. Hence, further studies like cloning of cry70Aa1 followed by bioassay can reveal the actual role of cry70Aa1 in causing toxicity against Z. cucurbitae and other dipteran insects.
Even though cry2 is not specific to dipteran insect pests, it is effective against dipterans along with lepidopterans. For instance, the study conducted by Da Silva and co-workers, reported that cry2Ab toxin present in S701 and S764 B. thuringiensis strains could be responsible for the toxicity against mosquitoes. However, the correlation between cry gene and mortality percentage reveals that cry2Ab causes mortality only in combination with other cry genes (Da Silva et al. 2004). This suggests that the presence of identical toxin cry2Ab4 in NBAIR BtPl could also be a potential toxin to cause mortality in Z. cucurbitae either alone or in combination cry70Aa1. Similarly, genes such as cry1Ia44, cry2Aa9 are effective in the management of lepidopteran insect pests (Alves et al. 2023). Apart from cry genes, few other genes such as vegetative insecticidal proteins (Vip) help in supplementing the insecticidal activity of Cry proteins, especially in overcoming the resistance (Boukedi et al. 2020). NBAIR BtPl strain also harbors a Vip like genes such as vip3Aa59 this belongs to vip3 family. Several studies conducted by Boukedi and co-workers, vip3 is effective in the management lepidopteran insects (Boukedi et al. 2016, 2018). So, the presence of these genes in NBAIR BtPl has a great potential for the management of lepidopteran pests as well. Apart from insecticidal toxicity-related genes, NBAIR BtPl strain also produces chitinases that have greater role in the management of plant pathogenic fungi (Gomaa 2012) and also degrades the laminated chitin in the intestinal peritrophic membrane of insect pests (Malovichko et al. 2019).
The cyt1Aa is the most widely studied group from the cyt1 family, and it is a major component in the crystal proteins of 4Q1 (Valtierra-de-luis et al. 2020). The toxic activity of cyt1Aa against the larvae of various dipteran species has been studied earlier (Wu et al. 1994; Chilcott and Ellar 1998). Although cyt1Aa is less toxic alone, it is proven to be the strongest synergist and known to interact synergically with some dipteran-specific cry genes like cry4Ba and cry11A (González-Villarreal et al. 2020). In contrast, NBAIR BtPl do not contain cyt1Aa but contains another cyt gene cyt1Da2, which belongs to the same cyt1 family. So, it can be interpreted that cyt1Da2 might have acted synergistically with the dipteran-specific cry gene; cry70Aa1, which resulted to cause greater mortality in larvae of Z. cucurbitae.
NBAIR BtPl strain also contains the gene clusters encoding secondary metabolites, which encode principal bioactive molecules. One among them is a gene showing 40% similarity with fengycin, which has biocontrol characteristics capable of preventing the spread of various kinds of fungal diseases in plants (Toure et al. 2004). Additionally, NBAIR BtPl also harbors clusters for bacillibactin, a siderophore that could function as a biocontrol agent against fungi in plants by chelating iron and decreasing its bioavailability (Dimopoulou et al. 2021). The clusters encoding the gene show 81% similarity with zwittermicin A, which is highly active against the numerous plant pathogenic fungi such as Fusarium, Alternaria, Ustilago, and Helminthosporium.
Conclusion
With unique insecticidal toxicity-related genes, NBAIR BtPl is a potentially potent B. thuringiensis strain that exhibits considerable toxicity against third instar larvae of Z. cucurbitae. This helps for the development of an environmentally friendly bioformulation in anticipation of its possible application in the field. Our work also sets the stage for future research on the discovery of novel B. thuringiensis toxins that are specific to dipterans, as well as the mechanism of action and interactions between different toxins. Future experiments like, cloning of individual genes, to study the synergistic effect of cry genes with other genes and to identify the efficiency of individual Cry proteins targeting the Z. cucurbitae and also other Dipteran insect pests are required to develop this strain as a new bioinsecticide.
Data availability
The data underlying this article is available in the NCBI database at Bioproject ID PRJNA1098464, SRA data SRR28606278, Biosample Accession SAMN40909047 (https://www.ncbi.nlm.nih.gov/sra/PRJNA1098464).
References
Ahmad S, Jamil M, Jaworski CC, Luo Y (2023) Comparative transcriptomics of the irradiated melon fly (Zeugodacus cucurbitae) reveal key developmental genes. Front Physiol 14:1112548. https://doi.org/10.3389/fphys.2023.1112548
Alves GB, De Oliveira EE, Jumbo LOV, Dos Santos GR, Dos Santos MM, Ootani MA, Aguiar RWDS (2023) Genomic–proteomic analysis of a novel Bacillus thuringiensis strain: toxicity against two lepidopteran pests, abundance of Cry1Ac5 toxin, and presence of INHA1 virulence factor. Arch Microbiol 205:143. https://doi.org/10.1007/s00203-023-03479-y
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Ben-Dov E, Nissan G, Pelleg N, Manasherob R, Boussiba S, Zaritsky A (1999) Refined, circular restriction map of the Bacillus thuringiensis subsp. israelensis plasmid carrying mosquito larvicidal genes. Plasmid 42:186–191. https://doi.org/10.1006/plas.1999.1415
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Boukedi H, Tounsi S, Abdelkefi-Mesrati L (2016) Abiotic factors affecting the larvicidal activity of the Bacillus thuringiensis Vip3Aa16 toxin against the lepidopteran pest Ephestia kuehniella. J Plant Dis Protect 123:59–64. https://doi.org/10.1007/s41348-016-0004-5
Boukedi H, Tounsi S, Abdelkefi-Mesrati L (2018) Insecticidal activity, putative binding proteins and histopathological effects of Bacillus thuringiensis Vip3 (459) toxin on the lepidopteran pest Ectomyelois ceratoniae. Acta Trop 182:60–63. https://doi.org/10.1016/j.actatropica.2018.02.006
Boukedi H, Hman M, Khedher SB, Tounsi S, Abdelkefi-Mesrati L (2020) Promising active bioinsecticides produced by Bacillus thuringiensis strain BLB427. J Adv Res Rev 8:026–035. https://doi.org/10.30574/wjarr.2020.8.1.0358
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
Cao ZL, Tan T, Jiang K, Mei SQ, Hou XY, Cai J (2018) Complete genome sequence of Bacillus thuringiensis L-7601, a wild strain with high production of melanin. J Biotech 275:40–43. https://doi.org/10.1016/j.jbiotec.2018.03.020
Chilcott C, Ellar D (1988) Comparative toxicity of Bacillus thuringiensis var. israelensis crystal proteins in vivo and in vitro. J Gen Microbiol 134:2551–2558. https://doi.org/10.1099/00221287-134-9-2551
Crickmore N, Berry C, Panneerselvam S, Mishra R, Connor TR, Bonning BC (2021) A structure-based nomenclature for Bacillus thuringiensis and other bacteria-derived pesticidal proteins. J Invertebr Pathol 186:107438. https://doi.org/10.1016/j.jip.2020.107438
Da Silva SMB, Silva-Werneck JO, Falcão R, Gomes AC, Fragoso RR, Quezado MT, Monnerat RG (2004) Characterization of novel Brazilian Bacillus thuringiensis strains active against Spodoptera frugiperda and other insect pests. J Appl Entomol 128:102–107. https://doi.org/10.1046/j.1439-0418.2003.00812.x
De Bon H, Huat J, Parrot L, Sinzogan A, Martin T, Malezieux E, Vayssieres JF (2014) Pesticide risks from fruit and vegetable pest management by small farmers in sub-Saharan Africa. A review. Agron Sustain Dev 34:723–736. https://doi.org/10.1007/s13593-014-0216-7
Dhillon MK, Singh R, Naresh JS, Sharma HC (2005) The melon fruit fly, Bactrocera cucurbitae: A review of its biology and management. J Insect Sci 5:40. https://doi.org/10.1093/jis/5.1.40
Dimopoulou A, Theologidis I, Benaki D, Koukounia M, Zervakou A, Tzima A, Diallinas G, Skandalis N (2021) Direct antibiotic activity of Bacillibactin broadens the biocontrol range of Bacillus amyloliquefaciens MBI600. Msphere 6:10–1128. https://doi.org/10.1128/msphere.00376-21
Dos Santos HRM, Argolo CS, Argôlo-Filho RC, Loguercio LL (2019) A 16S rDNA PCR-based theoretical to actual delta approach on culturable mock communities revealed severe losses of diversity information. BMC Microbiol 19:1–14. https://doi.org/10.1186/s12866-019-1446-2
Fayad N, Kambris Z, El Chamy L, Mahillon J, Kallassy Awad M (2021) A novel antidipteran Bacillus thuringiensis strain: unusual Cry toxin genes in a highly dynamic plasmid environment. Appl Environ Microbiol 87:1–20. https://doi.org/10.1128/AEM.02294-20
Gomaa EZ (2012) Chitinase production by Bacillus thuringiensis and Bacillus licheniformis: their potential in antifungal biocontrol. J Microbiol 50:103–111. https://doi.org/10.1007/s12275-012-1343-y
González-Villarreal SE, García-Montelongo M, Ibarra JE (2020) Insecticidal activity of a Cry1Ca toxin of Bacillus thuringiensis Berliner (Firmicutes: Bacillaceae) and its synergism with the Cyt1Aa toxin against Aedes aegypti (Diptera: Culicidae). J Med Entomol 57:1852–1856. https://doi.org/10.1093/jme/tjaa116
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435. https://doi.org/10.1093/nar/gkn176
Guerchicoff A, Ugalde RA, Rubinstein CP (1997) Identification and characterization of a previously undescribed cyt gene in Bacillus thuringiensis subsp. israelensis. Appl Environ Microbiol 62:2716–2721. https://doi.org/10.1128/aem.63.7.2716-2721.1997
Höfte H, Whiteley H (1989) Insecticidal crystal proteins of Bacillus thuringiensis. Microbiol Rev 53:242–255. https://doi.org/10.1128/mr.53.2.242-255.1989
Kremer FS, McBride AJA, Pinto LDS (2017) Approaches for in silico finishing of microbial genome sequences. Genet Mol Biol 40:553–576. https://doi.org/10.1590/1678-4685-GMB-2016-0230
Kumar J, Ramlal A, Mallick D, Mishra V (2021) An overview of some biopesticides and their importance in plant protection for commercial acceptance. Plants 10:1185. https://doi.org/10.3390/plants10061185
Letowski J, Bravo A, Brousseau R, Masson L (2005) Assessment of cry1 gene contents of Bacillus thuringiensis strains by use of DNA microarrays. Appl Environ Microbiol 71:5391–5398. https://doi.org/10.1128/aem.71.9.5391-5398.2005
Liu X, Lin X, Li J, Li F, Cao F, Yan R (2020) A novel solid artificial diet for Zeugodacus cucurbitae (Diptera: Tephritidae) larvae with fitness parameters assessed by two-sex life table. J Insect Sci 20:1–21. https://doi.org/10.1093/jisesa/ieaa058
Liu H, Zheng J, Bo D, Yu Y, Ye W, Peng D, Sun M (2022) BtToxin_Digger: a comprehensive and high-throughput pipeline for mining toxin protein genes from Bacillus thuringiensis. Bioinformatics 38:250–251. https://doi.org/10.1093/bioinformatics/btab506
Loutfi H, Fayad N, Pellen F, Le Jeune B, Chakroun M, Benfarhat D, Lteif R, Kallassy M, Le Brun G, Abboud M (2020) Morphological study of Bacillus thuringiensis crystals and spores. Appl Sci 11:155. https://doi.org/10.3390/app11010155
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Malovichko YV, Nizhnikov AA, Antonets KS (2019) Repertoire of the Bacillus thuringiensis virulence factors unrelated to major classes of protein toxins and its role in specificity of host-pathogen interactions. Toxins 11:347. https://doi.org/10.3390/toxins11060347
Manjunatha C, Velavan V, Rangeshwaran R, Mohan M, Kandan A, Sivakumar G, Sushil SN (2023) Assessment of bio-formulations of indigenous strains of Bacillus thuringiensis, Metarhizium robertsii and Metarhizium majus for management of the rhinoceros beetle, Oryctes rhinoceros L., in field. Egypt J Biol 33:1–15. https://doi.org/10.1186/s41938-023-00715-x
Moriya Y, Itoh M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:182–185. https://doi.org/10.1093/nar/gkm321
Onsongo SK, Mohamed SA, Akutse KS, Gichimu BM, Dubois T (2022) The entomopathogenic fungi Metarhizium anisopliae and Beauveria bassiana for Management of the Melon Fly Zeugodacus cucurbitae: pathogenicity, horizontal transmission, and compatability with cuelure. InSects 13:859. https://doi.org/10.3390/insects13100859
Rampersad J, Ammons D (2005) A Bacillus thuringiensis isolation method utilizing a novel stain, low selection, and high throughput produced atypical results. BMC Microbiol 5:1–11. https://doi.org/10.1186/1471-2180-5-52
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 14:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Touré Y, Ongena MARC, Jacques P, Guiro A, Thonart P (2004) Role of lipopeptides produced by Bacillus subtilis GA1 in the reduction of grey mould disease caused by Botrytis cinerea on apple. J Appl Microbiol 96:1151–1160. https://doi.org/10.1111/j.1365-2672.2004.02252.x
Travers RS, Martin PA, Reichelderfer CF (1987) Selective process for efficient isolation of soil Bacillus spp. Appl Environ Microbiol 53:1263–1266
Valtierra-de-Luis D, Villanueva M, Berry C, Caballero P (2020) Potential for Bacillus thuringiensis and other bacterial toxins as biological control agents to combat dipteran pests of medical and agronomic importance. Toxins 12:773. https://doi.org/10.3390/toxins12120773
Vilas-Bôas GT, Peruca APS, Arantes OMN (2007) Biology and taxonomy of Bacillus cereus, Bacillus anthracis, and Bacillus thuringiensis. Can J Microbiol 53:673–687. https://doi.org/10.1139/W07-02
Weber T, Blin K, Duddela S, Krug D, Kim HU, Bruccoleri R, Medema MH (2015) antiSMASH 3.0—a comprehensive resource for the genome mining of biosynthetic gene clusters. Nucleic Acids Res 43:237–243. https://doi.org/10.1093/nar/gkv437
Wu D, Johnson JJ, Federici BA (1994) Synergism of mosquitocidal toxicity between CytA and CrylVD proteins using inclusions produced from cloned genes of Bacillus thuringiensis. Mol Microbiol 13:965–972. https://doi.org/10.1111/j.1365-2958.1994.tb00488.x
Yılmaz S, Idris AB, Ayvaz A, Temizgül R, Hassan MA (2022) Whole-genome sequencing of Bacillus thuringiensis strain SY49.1 reveals the detection of novel candidate pesticidal and bioactive compounds isolated from Turkey. bioRxiv 3:1–15. https://doi.org/10.1101/2022.03.07.482483
Acknowledgements
The authors would like to express their sincere gratitude to The Director, ICAR-NBAIR, Bangalore, for the invaluable support provided during the successful execution of this research. The current research is funded through the ICAR- NBAIR project “Molecular Studies on Virulence of Bacillus thuringiensis and other entomogenous bacteria against Fall armyworm and White grubs (CRSCNBAIRSIL202100300205).
Funding
This work was supported by the Indian Council of Agricultural Research (ICAR), New Delhi, ICAR- National Bureau of Agricultural Insect Resources, Bengaluru, Karnataka through the project “Molecular Studies on Virulence of Bacillus thuringiensis and other entomogenous bacteria against Fall armyworm and White grubs” (CRSCNBAIRSIL202100300205).
Author information
Authors and Affiliations
Contributions
CM and AN conceptualized the research. AN conducted the experiment and wrote the manuscript. JP has done the statistical analysis of bioassay data. AK, and NS analyzed the whole genome sequence data. RS and KMC performed bioinformatic analysis. KA, ANS, and VKD monitored the experiments and edited the manuscript. All the authors have read the manuscript and approved it for submission.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This is an observational study and no ethical approval is required.
Human and animal rights statement
All procedures performed in the study are in accordance with the ethical standards of the institutional and/or national research committee. We further declare that no animals and humans were harmed during the study.
Informed consent
Informed consent was obtained from all the individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplemental Fig. 1.
Plasmid profiling of indigenous B. thuringiensis isolates. Lane M: 10kb ladder, Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl (JPG 54 KB)
Supplemental Fig. 2.
Analysis of crystal proteins present in indigenous B. thuringiensis isoaltes. a. PCR amplification of cry1 gene in B. thuringiensis isolates Lane M: Ladder (1Kb), Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl. b. PCR amplification of cry2 gene in B. thuringiensis isolates Lane M: Ladder (1Kb), Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl. c. PCR amplification of cry8Aa1 gene in B. thuringiensis isolates Lane M: Ladder (1Kb), Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl. d. PCR amplification of cry70Aa1 gene in B. thuringiensis isolates Lane M: Ladder (1Kb), Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl. d. SDS PAGE analysis of crystal proteins present in B. thuringiensis isolates Lane M: protein marker (245 kDa), Lane 1-5: NBAIR Bt151, NBAIR Bt152, NBAIR Bt153, NBAIR Bt154, NBAIR BtPl (JPG 66 KB)
Supplemental Fig. 3.
A phylogenetic tree constructed based on cry70Aa1 and other cry70 family members constructed by the Maximum Likelihood (ML) method using MEGA software (JPG 33 KB)
Supplemental Table 1.
(DOCX 25 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aarthi, N., Dubey, V.K., Shylesha, A.N. et al. Insights into the whole genome sequence of Bacillus thuringiensis NBAIR BtPl, a strain toxic to the melon fruit fly, Zeugodacus cucurbitae. Curr Genet 70, 13 (2024). https://doi.org/10.1007/s00294-024-01298-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00294-024-01298-2