Abstract
Bacillus thuringiensis (Bt) is a biological alternative to the indiscriminate use of chemical insecticides in agriculture. Due to resistance development on insect pests to Bt crops, isolating novel Bt strains is a strategy for screening new pesticidal proteins or strains containing toxin profile variety that can delay resistance. Besides, the combined genomic and proteomic approaches allow identifying pesticidal proteins and virulence factors accurately. Here, the genome of a novel Bt strain (Bt TOL651) was sequenced, and the proteins from the spore–crystal mixture were identified by proteomic analysis. Toxicity bioassays with the spore–crystal mixture against larvae of Diatraea saccharalis and Anticarsia gemmatalis, key pests of sugarcane and soybean, respectively, were performed. The toxicity of Bt TOL651 varies with the insect; A. gemmatalis (LC50 = 1.45 ng cm−2) is more susceptible than D. saccharalis (LC50 = 73.77 ng cm−2). Phylogenetic analysis of the gyrB gene indicates that TOL651 is related to Bt kenyae strains. The genomic analysis revealed the presence of cry1Aa18, cry1Ac5, cry1Ia44, and cry2Aa9 pesticidal genes. Virulence factor genes such as phospholipases (plcA, piplc), metalloproteases (inhA), hemolysins (cytK, hlyIII, hblA, hblC, hblD), and enterotoxins (nheA, nheB, nheC) were also identified. The combined use of the genomic and proteomic data indicated the expression of Cry1Aa18, Cry1Ac5, and Cry2Aa9 proteins, with Cry1Ac5 being the most abundant. InhA1 also was expressed and may contribute to Bt TOL651 pathogenicity. These results provide Bt TOL651 as a new tool for the biocontrol of lepidopteran pests.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane (Saccharum officinarum), corn (Zea mays), and soybean (Glycine max L) are high-value crops and applied to different purposes such as food and biofuel production (de Matos et al. 2020; Heinrichs et al. 2017). The sugarcane borer, Diatraea saccharalis (Fabricius, 1794) (Lepidoptera: Crambidae), is a key pest of sugarcane and corn. The velvetbean caterpillar, Anticarsia gemmatalis (Hübner 1818) (Lepidoptera: Noctuidae), is a key pest of soybean (Horikoshi et al. 2022; Mendonça et al. 2020). These insects are among Brazil’s most relevant Lepidoptera pests causing damage to the raw material used in food and biofuel processing (Silva 1995; Praça et al. 2004; Moscardi et al. 2012; Dinardo-Miranda et al. 2013).
Biocontrol is a safe alternative to reduce the use of chemical insecticides in crop pest management. Entomopathogenic microorganisms have been considered important agents for this purpose with relevant importance for the bacteria Bacillus thuringiensis (Bt) (Bel et al. 2017; Daquila et al. 2019; Sanahuja et al. 2011). Biological control products based on Bt are being used as biopesticides for decades and currently have the majority of the biological control market share in the world (Arthur and Dara 2019; Lacey et al. 2015; Sena da Silva et al. 2022). Bt is a Gram-positive, spore-forming bacterium that can produce parasporal crystal proteins during the sporulation phase (Cry and Cyt proteins) (Pohare et al. 2021). These proteins when ingested by a susceptible insect are solubilized in the alkaline midgut environment, acquiring an oligomeric form, binding to the midgut cell membrane, leading to the destruction of these cells and their insecticidal proprieties (Frankenhuyzen 2009; Bravo et al. 2007, 2011; Pinheiro and Valicente 2021). Bt can produce pesticidal proteins during the vegetative phase (Vip proteins) (Pohare et al. 2021). Virulence factors such as metalloproteases, chitinases, hemolysins, and enterotoxins also represent Bt pathogenicity (Malovichko et al. 2019; Palma et al. 2014).
Bt is an important biopesticide used against lepidopteran pests as spray formulations and Bt crops (transgenic plants that express Cry and/or Vip3 proteins) (Castro et al. 2019; Daquila et al. 2019; Horikoshi et al. 2022; Srikanth et al. 2011). However, the evolution of resistance to Bt crops in lepidopteran pests has been reported, including in D. saccharalis (de Oliveira et al. 2022; Huang et al. 2015) and A. gemmatalis (Pezenti et al. 2021). Therefore, the isolation of novel Bt strains is an important strategy for the discovery of new pesticidal proteins or strains containing a range of toxin profiles that can delay the target insect’s resistance.
Characterizing novel Bt strains and studying the genome and proteome is important to understand their pathogenicity. Genome sequencing technology has accelerated the discovery of novel pesticidal proteins, secondary metabolites, and virulence factors in Bt (Zghal et al. 2018; Liu et al. 2017; Cardoso et al. 2020; Day et al. 2014; Jeong et al. 2017; Jia et al. 2016; Liu et al. 2015). However, considering not all of the coding regions predicted from the annotated genome sequence are expressed, there are cryptic pesticidal proteins (Quan et al. 2016; Rang et al. 2015). So, the pesticidal proteins expression profile could be explored using proteomic analysis. Thus, in combination with genomic studies, proteomic analysis allows the accurate identification of pesticidal proteins and virulence factors in different Bt strains (Baragamaarachchi et al. 2019; Gomis-Cebolla et al. 2018; Khorramnejad et al. 2020; Wu et al. 2011). Furthermore, from the genomic–proteomic analysis it is also possible to estimate the protein abundance in purified parasporal crystals and spore–crystal mixtures (Baragamaarachchi et al. 2019; Huang et al. 2012; Khorramnejad et al. 2020).
This study sequenced the genome of a novel strain Bt TOL651, toxic against A. gemmatalis and D. saccharalis, and the entomopathogenic characteristics were explored. Additionally, an LC–MS/MS analysis of the spore–crystal mixture was performed to determine the expressed proteins.
Materials and methods
Culturing of TOL651 strain
Bacillus-like colonies were isolated from soil samples collected in Tocantins State (Brazil) (11º43′45′′S; 49º04′07′′W) according to the previously described protocol (Monnerat et al. 2001). To screen for Bt strains, Petri dishes with a selective NYSM medium (nutrient yeast extract salt medium) [8 g/l of nutrient broth (Difco, USA), 0.103 g/l of CaCl2·2H2O, 0.01 g/l of MnCl2·4H2O, 0.203 g/l of MgCl2·6H2O] (Kalfon et al. 1983) containing 100 mg/l penicillin G were used and the culture grown for 24 h at 30 ± 0.5 °C at 180 rpm. Then, each colony was individually analyzed and identified by phase-contrast microscopy (× 1000) to verify the presence of inclusion bodies and crystals (Frankland and Frankland 1887). The Bt TOL651 was selected among 87 crystal-forming Bt strains (87 strains from 2.445 Bacillus-like colonies), due to being the most toxic among different isolated Bt strains and tested simultaneously against Diatrea saccharalis and Anticarsia gemmatalis in the selective bioassays, according to Monnerat et al. (2007). Bt HD-1was isolated from the commercial sample (Dipel WP 32 g/kg, Sumitomo Chemical do Brasil Representações Ltda., SP) and used as a reference strain (Cerqueira et al. 2017; Lazart et al. 2021; Sathyan et al. 2022). Bt HD-1 has been designated as the primary US reference standard strain for toxicity against lepidopteran insects (Dulmage 1973). Bt TOL651 strain was cultured at 28 °C for 12 h on solid Luria–Bertani medium (LB) (10 gL−1 tryptone, 5 gL−1 yeast extract, 10 gL−1 NaCl, and 20 gL−1 agar). A single bacterial colony was inoculated in the LB liquid medium at 28 °C, 200 rpm for 16 h, which was used as a starter culture for spore–crystal mixture production and in the genomic DNA extraction step.
SDS-PAGE analysis of cry proteins
For sporulation and crystal production, a starter culture was transferred to the CCY medium (30 ml) (13 mM KH2P04, 26 mM K2HP04, 0.002% [w/v] L-glutamine, 0.1% [w/v] casein hydrolysate, 0.1% [w/v] Bacto Casitone, 0.04% Bacto Yeast extract, 0.6% [w/v] glycerol, 0.05 M ZnCl2, 0.5 M MgCl2, 0.01 M MnCI2, 0.2 M CaCl2, 0.05 M FeCl3) and incubated at 28 °C, 200 rpm for 72 h. For SDS-PAGE analysis, the crystals were purified using hexane and low-speed centrifugation, according to Mounsef et al. (2014). The spores and crystals were collected for centrifugation at 6000 rpm, 4 °C for 10 min, and the pellet was washed twice by suspending it in saline solution (1 M NaCl containing 0.01% Triton X-100) by centrifugation (6000 rpm, 4 °C for 10 min). Then, the pellet was suspended in a 50 ml centrifuge tube with 27 ml of saline solution and sonicated at 100 W of potency for 10 min. Then, 3 ml of hexane was added to the suspension following the centrifugation at 6000 rpm, 4 °C for 10 min. This procedure was repeated three times. The pellet was washed twice with cold distilled water by centrifuging. Then, the crystals were solubilized using 50 mM NaOH buffer at 30 °C and quantified using Bradford reagent (Bio-Rad protein assay). Following this, 7 μg of solubilized crystals was analyzed by 12% sodium dodecyl sulfate-–polyacrylamide gel electrophoresis (SDS-PAGE) and stained with Coomassie Brilliant Blue (R250, 0.4%).
Microscopy
The spore–crystal mixture of TOL651 was collected and diluted in sterile water. Then, 100 µL of this dilution was deposited over metallic supports to be dried for 24 h at 37 °C, covered with gold for 180 s using an Emitech apparatus (model K550), and observed in a Zeiss scanning electron microscope (model DSM 962) at 10 or 20 kV.
Toxicity against Diatraea saccharalis and Anticarsia gemmatalis
Eggs from D. saccharalis and A. gemmatalis were obtained from Biocontrole (biocontrole.com.br) and Embrapa Recursos Genéticos e Biotecnologia—CERNAGEN (Brasília, DF, Brazil), respectively. The insect’s eggs were maintained under ideal rearing conditions (i.e., 26 °C ± 1 °C and 70 ± 10% RH and 10:14 h (L: D) photoperiod) (Schmidt et al. 2001), on specific artificial diets prepared for D. saccharalis (Hensley and Hammond 1968) and A. gemmatalis (Greene et al. 1976).
Bioassays against D. saccharalis were performed using 24-well cell culture plate (TPP, Techno Plastic Products AG). 1.5 ml of diet was poured into each well and, after solidification, aliquots 35 µL of ten spore–crystals dilutions (from 0.1 to 1000 ng/cm2) were spread on the diet surface. Subsequently, 1-day-old second instar larvae were placed in each plate. Trays were closed with acrylic lids, keeping them under controlled conditions (26 °C ± 1 °C and 70 ± 10% RH and 10:14 [L: D] h). After 48 h, the surviving larvae were individually transferred to six-well cell culture plates containing a rearing diet. Larvae mortality was evaluated again on day 7 (Praça et al. 2004). Larvae were considered dead when stimulated and no movement was detected.
In the A. gemmatalis bioassays, a total volume of 3 ml artificial diet was poured into 30 ml plastic cups. After solidification, aliquots (150 μl) of ten spore–crystal concentrations (from 0.1 to 1000 ng/cm2) were applied on the diet surface and dried at room temperature. Then, ten 1-day-old second instar larvae of A. gemmatalis were added to each cup. The cups were covered with plastic lids and kept under the same conditions described above. After 48 h, the surviving larvae were placed in 30 ml cups containing a rearing diet and the mortality was assessed. Larval mortality was evaluated again on day 5 as described (da Silva et al. 2004).
All the bioassays were performed in triplicate. The commercial strain HD-1 was used as a reference and sterile water, pH 7.0, was added as the control group. The lethal concentrations (LC50 and LC95) were determined by Probit analysis (Finney 1971) using the PoloPlus 1.0 (LeOra Software Berkeley, CA, USA).
Genome sequencing, data assembly, and annotation
Total DNA from the Bt TOL651 strain was extracted and purified by the Wizard® Genomic DNA Purification Kit (Promega, Madison, WI, USA) following the manufacturer’s instructions. The DNA concentration and purity were checked by the NanoDrop™ 8000 apparatus (ThermoFisher Scientific, Waltham, MA, USA) and then stored at − 20 °C until further use. Genome sequencing was performed using Illumina MiSeq technologies (paired-end application, 2x ~ 80 bp, average insert size of 200 bp), and the coverage of 137 X. FastQC v.0.11.9 (Andrews 2015) was used for the reads libraries quality analyses, with reads being trimmed using Geneious v.10.2.6 (Kearse et al. 2012) (Workflow Trim and Filter, Error Probability: 0.05). The de novo assembly was performed using SPAdes v.3.10.0, using default parameters (Bankevich et al. 2012). The assembled contigs (= > 500 bp) were run through MeDuSa v.1.6 (Bosi et al. 2015) for scaffolding, using complete genomes Bt YBT-1520 and Bt HD-1 (NCBI RefSeq NZ_CP004858 and CP004870 respectively) strains as a reference dataset. The quality of the final assembly was assessed using Quast v.5.0.2 (Gurevich et al. 2013) in standard mode, and completeness percentage, and N50 and L50 values were obtained. The annotation and CDS prediction of scaffolds were performed using RASTtk 2.0 (Brettin et al. 2015).
Average nucleotide identity (ANI) and phylogenetic relationships
The genome similarity was assessed through ANI using JSpeciesWS (Richter et al. 2015) on Tetra Correlation Search (TCS) function for selecting related genomes. The Heatmap dendrogram was created using the Morpheus tool (https://software.broadinstitute.org/morpheus). The phylogenetic relationship analysis was performed using the gyrB gene (DNA gyrase subunit B) including other genes of the closely related Bacillus spp. strains retrieved from GenBank. The sequences were aligned using ClustalW, and a phylogenetic tree was created using MEGA 11 (Kumar et al. 2018), using the neighbor-joining method and bootstrap percentages based on 1000 replications.
Gene identification of pesticidal proteins
Putative pesticidal proteins were determined using Blastx, through the Btoxin_Digger tool (scaffolds as a query) (Liu et al. 2021) and a customized database (CDS predicted as a query). The customized database was created through Geneious, using Add/Remove Database tool, from the Bt pesticidal proteins list available at the Bt nomenclaturewebsite (http://www.lifesci.sussex.ac.uk/home/Neil_Crickmore/Bt/toxins2.html). CDS with homology to the Bt pesticidal proteins was filtered using E-value 0.001 and word size 6 parameters. To check the presence of novel putative Cry sequences in TOL651, the sequences obtained from the non-redundant protein database NCBI were used (Lazart et al. 2021).
Identification of virulence factors and antibiotic resistance genes
The potential virulence factors were predicted using the bacterial virulence factor database (VFDB) (Liu et al. 2019). The TOL651 genome was screened for antibiotic resistance factors, using the Resistance Gene Identifier (RGI), within Comprehensive Antibiotic Resistance Database (CARD) (Alcock et al. 2023), according to the parameters: perfect, strict, complete genes only and 95% identity nudge.
Proteomic analysis
The proteins in the spore–crystal mixture of the Bt TOL651 strain were identified by LC–MS/MS at the Biotech Company Veritas /Life Sciences at the University of São Paulo (USP, Ribeirão Preto, SP, Brazil). The sample of the spore–crystal mixture was washed three times in 1× PBS (phosphate-buffered saline) and resuspended in 750 µL of solubilization buffer (8 M urea, 0.5% Octyl-glucopiranoside (OG) and 0.05 M Tris–HCL, pH 8.8). Then, the sample was sonicated (three cycles of 60 s, 30% amplitude, and shut off for 2 s) and maintained on ice. The solubilized proteins were quantified through the Bradford method using Protein Assay Dye Reagent Concentrate (Bio-Rad Laboratories), according to the manufacturer’s instructions.
The preparation of the sample for advanced mass spectrometry consisted of three main steps: (i) reduction and alkylation of proteins, (ii) enzymatic digestion using trypsin, and (iii) cleanup/desalting. Briefly, 50 µg of the sample was subjected to disulfide bridge reduction using 50 µg of DTT (dithiothreitol) and 60 min of incubation at 37 °C. Then, the process was followed by alkylation using 250 µg of I.A (iodoacetamide) and 60 min at room temperature in the dark. Finally, the sample was diluted five times in Tris–hydrochloride (0.05 M Tris–HCl, pH 8.8) and incubated with 2 µg of trypsin (Promega, V511A) at 37 °C overnight.
Previously to the mass spectrometry application, the cleanup/desalting of the sample was performed using C18 resin (Supleco). The column was calibrated using 2% acetonitrile containing 0.1% formic acid, and the elution was performed with 50% acetonitrile. The sample was then dried in a speed vac and applied in a mass spectrometer (Termo Fisher Orbitrap Eclipse), coupled to a nanoflow nano LC–MS/MS chromatography system (Dionex Ultimate 3000 RLSCnano System, Thermofisher). Peptides were separated in nanoEase MZ peptide BEH C18 column (130A, 1.7 µm, 75 µm × 250 mm, Waters) 300 nL/min using a 4–50% acetonitrile gradient for 90 min. The data were obtained on MS1 in the range of M/Z 375–1500 (120,000 resolution, AGC target 1E6, maximum time injection of 100 ms). The most abundant ions were submitted to MS/MS (30% collision energy, 1.2 m/z, AGC target 1E5, 15000 resolution).
The raw data were converted to mzXML format and processed using PatternLabV (Santos et al. 2022). The data were analyzed against the database created using CDS translated to the TOL651 genome (Generate Search DB option). The contaminant library content of common MS contaminant sequences (e.g., trypsin, keratins, and albumin) was included in the database. The modifications selected in the search were carbamidomethyl (C), deamination (NQ, variable), and oxidation (M, variable). Enzyme trypsin (fully specific), two maximum missed cleavages, and initial precursor mass tolerance of 10 ppm were set as Comet parameters. The acceptable FDR (false discovery rate) estimates of 3% at spectral, 2% at peptide, and 1% at protein levels and MS and MS/MS tolerance errors of 10 ppm were added as parameters in the Filtering (SEPro) options.
The functional annotation of the identified proteins was performed using UniProtKB/Swiss-Prot database, and the graphical summary of functional classification was created using GO terms through the WEGO 2.0 tool (Web Gene Ontology Annotation Plot) (Ye et al. 2018).
Data availability
This whole genome shotgun project has been deposited at DDBJ/ENA/GenBank under the accession JANVFA000000000. The version described in this paper is JANVFA010000000.
Results
Protein profile and morphological and toxicity analysis
Cry protein profile of Bt TOL651 revealed the presence of two major protein bands of approximately 130 and 65 kDa size, indicating the presence of both Cry1 and Cry2 proteins, also observed in the reference strain (Fig. 1A). The morphological analysis revealed the presence of spores and bipyramidal and cuboidal crystals in this strain, also indicating the expression of Cry1 (bipyramidal crystals) and Cry2 (cuboidal crystals) proteins (Fig. 1B).
SDS-PAGE analysis of Cry proteins and scanning electron microscopic of spore–crystal mixture of Bt TOL651. a Protein profile: Lane 1—molecular mass markers; Lane 2—HD-1; Lane 3—TOL651. Arrows indicate likely protein band size. b The view of spores (S) and bipyramidal (B) and cuboidal (C) crystals enlarged 2,000 (×) times and approximated 10,000 times (×)
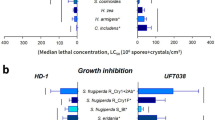
The insect bioassays using spore–crystal mixtures of Bt TOL651 and the reference Bt HD-1 strain showed that both strains were toxic to D. saccharalis and A. gemmatalis (Table 1). However, the Bt TOL651 showed significantly higher toxicity to D. saccharalis and A. gemmatalis when compared to the Bt HD-1 strain (Table 1). The RT50 estimate showed that the Bt TOL651 strain was 1.97- and 1.75-fold more toxic than the Bt HD-1 strain against D. sacharalis and A. gemmatalis, respectively (Table 1).
Genomic characterization
The draft genome of Bt TOL651 was obtained and consists of ~ 6.17 Mb with 35.3% GC content (Table 2). A total of 7003 coding sequences (CDS) were found, out of which 4812 proteins had functional assignments and 2191 were considered hypothetical proteins. Sixty-three tRNA and 5 rRNA genes were also annotated (Table 2). In the subsystem class distribution, most of the genes were involved, in decreasing order: amino acids and derivatives metabolism (387); carbohydrate (265); cofactors, vitamins, prosthetic groups, pigments metabolism subsystems (161); protein metabolism (155); and nucleosides/nucleotides metabolism (118) (Fig. 2).
Bt TOL651 relationships with other Bt strains
In the search for Bt genomes highly correlated to Bt TOL651, different Bt strains’ genomes were compared with Bt TOL651. ANI values were obtained and showed that Bt TOL651 was highly similar to other Bt strains (ANI ≥ 94%). High ANI values were observed for Bt TOL651 and other Bt strains such as Bt kurstaki T03a001 (99.06%), Bt kurstaki HD73 (99.00%), Bacillus sp. G3 (98.88%), Bt YC-10 (98.77%), Bt mexicanensis 27 (98.63%), Bt NBIN-66 (98.41%), Bt galleriae HD-29 (98.28%), and Bt aizawai Leap01 (98.04%). However, Bacillus cereus (Bc) B4158 genome also was highly correlated with Bt TOL651 (ANI = 98.16%) (Fig. 3). The phylogenetic analysis using gyrB gene indicated that Bt TOL651 was clustered with Bt kenyae BGSC-4F2 and IEBC-T04B001 strains (Fig. 4).
Genes related to pesticidal proteins, virulence factors, and antibiotic resistance
Based on the Bt database and Btoxin_Digger, a total of four cry1 and cry2-type genes were found in two scaffolds (18 and 21) in the genome of Bt TOL651. The CDS regions peg.5608, peg.5616, and peg.6113 showed high homology to the cry2Aa9, cry1Ia44, and cryAc5 genes, respectively. On the other hand, the peg.5617 CDS region showed high homology to the cry1Aa18 gene in the custom Bt database, but was not found in the Btoxin_Digger (Table 3, Fig. 5). Finally, the peg.3270 CDS region was retrieved using Btoxin_Digger and custom Bt database, demonstrating high homology to the spp1Aa1 gene (~ 80%).
Virulence factor genes from different classes (adherence, enzyme, immune evasion, iron acquisition, regulation, secretion system, toxins, acid resistance, magnesium uptake, surface protein anchoring, and others) were also found in the genome sequence of Bt TOL651 (Table S1). Among these genes, we can point out enzymes such as phospholipases (plcA, piplc) and metalloproteases (inhA), and other toxins such as hemolysins (cytK, hlyIII, hblA, hblC, hblD) and enterotoxins (nheA, nheB, nheC). (Table S1). Four putative antibiotic resistance genes were identified in the genome of Bt TOL651, including BcII gene (subclass B1 beta-lactamase) (% identity: 90.12; % length of reference sequence: 122.66), BcI, and two BcIII genes (class A beta-lactamase) (% identities: 95.42, 86.83 and 76.58, respectively, % length of reference sequences: 100.65, 100.32 and 100, respectively).
Proteomic of spores–crystal mixture
The general functional classification carried out by LC–MS/MS analysis of the spore–crystal mixture of Bt TOL651 revealed that detected proteins sequences were involved in 11 GO terms related to cellular components, 10 GO terms related to molecular functions, and 19 terms related to biological processes (Fig. 6). In the cellular component groups, most proteins were related mainly to cell and membrane components. Furthermore, the molecular function classification was represented by proteins with catalytic and binding activities; next, in the biological process category, most proteins belonged to metabolic and cellular processes.
The comparison among genomic and proteomic data indicated that 24 coding sequences were identified in the proteins’ sequences and, based on unique peptide count, the most abundant proteins were: Cry1Ac5, Cry2Aa9, and Cry1Aa18 (Table 4) (Table S2). Except for Cry1Ia44 (peg.5616), which had no unique peptide, all Cry proteins identified in the genome were expressed in the sporulation phase (72 h of incubation). Cry1Ac5 (peg.6113) showed the highest number of peptides detected, with 85 in total and 82 unique sequences (Table 4) (Table S2). Additionally, regarding the pesticidal proteins, the inhibitor A metalloprotease (InhA1), a potential pathogenic factor, was also found in the proteome of Bt TOL651 (Table 4) (Table S2). Other spore-associated proteins (spore coat and forespore-specific proteins), peptides associated with protein metabolism (elongation factor Tu, shock protein Hsp20 family, and chaperone protein DnaK), and other functions (aminopeptidase, glycerophosphoryl phosphodiesterase, choline-binding protein, DUF3915 domain containing, acid endopeptidase, neutral protease B, and DNA-binding protein) were also detected (Table 4).
Discussion
Bt TOL651 strain analyses presented a Cry protein profile in SDS-PAGE gel with two major protein bands of approximately 130 and 65 kDa in size, which is also associated with Cry1 and Cry2 proteins, respectively (Ganesh et al. 2018; Monnerat et al. 2007; Schnepf et al. 1998; Singh et al. 2021). In concordance with the protein profile, the crystal morphology of Bt TOL651 revealed bipyramidal forms associated with Cry1 proteins and cuboids formed by Cry2 proteins similar to the HD-1 strain (Monnerat et al. 2007; Schnepf et al. 1998).
The TOL651 genome similarity analysis confirmed that this isolate is a Bt species, since ANI values ≥ 95% (98–99%) were assigned in comparison with other genomes of this species (Richter and Rosselló-Móra 2009). However, a high ANI value between Bt TOL651 and the B. cereus (Bc B4158) was also found, which corroborates the complex separation between Bt and B. cereus at the genomic level (Helgason et al. 2000; Lechuga et al. 2020; Zhou et al. 2022). However, in the subspecies classification, the phylogenetic study showed that the Bt TOL651 strain might have a close relationship with Bt kenyae strains.
Bt strains identified as kenyae subspecies have been reported to harbor cry1Ab, cry1Ac, cry1E, and cry2Aa genes and showed toxic activity against lepidopteran insects (Chang et al. 1999; Hire et al. 2008, 2009). Bt TOL651 harbors cry2Aa9, cry1Ia44, and cryAc5 genes. HD-1 strain, a kurstaki subspecies, harbors cry1Aa, cry1Ab, cry1Ac, cry1Ia, cry2Aa, cry2Ab, and vip3Aa genes of pesticidal proteins (Zhu et al. 2015).
In addition, Bt TOL651 harbored enterotoxins such as hemolysins genes (hemolysins I, III, and IV) presented in other species of Bacillus cereus sensu lato group and detected in new Bt strains (Kim et al. 2015; Ma et al. 2020; Lechuga et al. 2020). Although Bt is considered a safe bioinsecticide for non-target organisms (Pohare et al. 2021), the presence of these genes in Bt TOL651 requires their validation for safe use in crops, as well as this toxicity for humans, since we propose its use as a spore–crystal mixture. Considering the potential antibiotic resistance of Bt TOL651, the putative antibiotic resistance genes were screened in the genome sequence, indicating the presence of genes resistant to beta-lactam antibiotics, corroborating other studies (Luna et al. 2007; Kaze et al. 2021). Banik et al. (2019) have demonstrated the sensitivity of Bt strains against clinically important antibiotics, indicating it is a safe biocontrol agent for crop application without any harm to consumers. However, additional experimental antibiogram tests need to be conducted to validate the resistance or sensitivity of the TOL651 to antibiotics.
The combined genomic and proteomic analysis described in this work revealed that the spore–crystal mixture of the Bt TOL651 has a high proportion of the pesticidal proteins of Cry1Ac5, Cry2Aa9, and Cry1Aa18. These Cry proteins were shown in other studies to be toxic to lepidopteran insect pests (Dammak et al. 2015; dos Santos et al. 2009; Sun et al. 2022). Similarly, genomic and proteomic analysis of the Bt 4.0718 strain indicated the expression of Cry2Aa, Cry1Aa, and Cry1Ac (Rang et al. 2015; Huang et al. 2012). The Bt HD-1 strain in Caballero et al.’s (2020) study using the same omics approach indicated the expression of Cry1Aa, Cry1Ac, Cry1Ab, and Cry2Aa.
Although present in the genome of Bt TOL651, the Cry1Ia44 protein was not detected in the spore–crystal mixture by proteomic analysis. Despite harboring the cry1Ia gene sequence, in other Bt strains the expression of this protein was also not detected, based on the LC–MS/MS approach (Huang et al. 2012; Quan et al. 2016). This result, corroborating with other studies, has suggested the cryptic nature of the cry1I genes based on their absence in parasporal crystals (Gleave et al. 1993; Tailor et al. 1992). However, the genomic and proteomic study of the Bt AB1 strain revealed three unique peptides of Cry1Ia, their expression being considered at a trace level (Baragamaarachchi et al. 2019).
Cry1Ac5 was the most abundant spore–crystal of Bt TOL651. The gene coding Cry1Ac5 has been described in Bt isolates from warehouses in China (Hongyu et al. 2000) and in Bt strain isolated from India, toxic to Spodoptera frugiperda (Sathyan et al. 2022). In contrast to TOL651, another wild-type Bt isolates and commercial HD-1 strain expressed Cry2Aa in greater proportion (Huang et al. 2012; Caballero et al. 2020), indicating that the abundance of Cry-like proteins may change according to the strain.
Bioassays with the spore–crystal mixture showed, in comparison with the commercial strain Bt HD-1, that Bt TO651 was more toxic for both insects, with A. gemmatalis being the most susceptible. Studies showed that the toxicity level of each Cry protein varies with the insect species. For example, Bel et al. (2017) found that Cry1Ac was more toxic than Cry1Aa against A. gemmatalis, but the opposite occurred when these proteins were tested on Chrysodeixis includens (Walker 1858) (Noctuidae: Lepidoptera). D. saccharalis was susceptible to Cry1Aa and Cry1Ac, of which Cry1Ac was more effective (Davolos et al. 2015; Tan et al. 2011). However, this insect was more susceptible to a mixture of Cry1 and Cry2 proteins than when tested separately (Macedo et al. 2012). This can happen because Cry1Ac and Cry2Aa proteins do not share the same midgut receptor binding sites; therefore, a synergic effect can be expected when both proteins are present (Macedo et al. 2012). Cry1Aa, Cry1Ab, and Cry1Ac proteins share binding sites (Davolos et al. 2015). In contrast to HD-1 (Caballero et al. 2020), TOL651 expressed only Cry1Aa18 and Cry1Ac5. This finding has suggested that Cry1Ac has a high affinity in comparison to Cry1Aa for the shared binding site, due to divergences in domain II of proteins (Hernández-Rodríguez et al. 2013). So, the presence of Cry1Aa18 and Cry1Ac5 in the spore–crystal mixture of TOL651 may not affect the action of Cry1Ac5. Thus, our results suggest that a major proportion of Cry1Ac5 protein in the spore–crystal mixture along Cry1Aa18 and Cry2Aa9 proportions may explain the toxicity of Bt TOL651 against A. gemmatalis and D. saccharalis.
In addition to Cry proteins, the inhibitor A metalloprotease (InhA1) was also detected in the spore–crystal mixture of Bt TOL651. InhA1 produced early in the sporulation phase allows the neutralization of the host immune system by specifically degrading the insect attacin and cecropin proteins (Miyoshi and Shinoda 2000; Pohare et al. 2021). Besides, Dammak et al. (2015) mentioned that InhA1 within a spore–crystal mixture can enhance the pathogenic effect of Cry1–Cry2 proteins, since it can cause disorganization of the intestinal epithelium and delay a possible resistance caused by the intensive use of Cry proteins. Although InhA1 expression has been reported in a spore mixture of Bt strains (Banik et al. 2019; Khorramnejad et al. 2020), researches have indicated the absence of InhA1 in the late sporulation phase (Li et al. 2012), and different levels in transcription and expression of InhA1 between Bt strains, suggesting a possible strategy to adapt to various hosts (Zhu et al. 2015). The deficiency in the expression of the virulence factor camelysin also produced for Bt could be involved in the lack of expression of InhA (Yin et al. 2011).
Neutral protease B (NprB) (also named NprA and Npr99) was also present in the spore–crystal mixture of Bt TOL651 and associated with the virulence of Bacillus cereus, degrading host tissues and resulting in increasing tissue permeability to the pathogen (Chung et al. 2006). Heat shock protein Hsp20 and the elongation factor Tu were also detected in the spore–crystal mixture of Bt TOL651 and are necessary for the formation of crystals in Bt strains (Ding et al. 2009). The Hsp20 protein supports other proteins in refolding and preventing protein degradation (Xie et al. 2019).
Biopesticides are commonly used in multi-strain consortia and represent better cost-effectiveness than constructing recombinant or purified toxins for the development of products (Sreshty et al. 2011). The synergism between different crystalline proteins produced by two Bt strains that do not compete for the same binding site has shown enhanced activity against lepidopteran pests (Konecka et al. 2012). The genes and proteins identified in the genome–proteomic step study of Bt TOL651 will experimentally facilitate the determination of the potential of synergism between TOL651 and other strains.
In conclusion, our findings showed the potential use of the Brazilian Bt TOL651 strain in the control of D. saccharalis and A. germmatalis, of which A. germmatalis was most susceptible. Bt TOL651 was closely related to kenyae subspecies and expressed mainly Cry1Aa18, Cry1Ac5, and Cry2Aa9 pesticidal proteins in the spore–crystal mixture, with Cry1Ac5 being the most abundant protein. The virulence factor InhA1 may contribute to the pathogenicity of Bt TOL651. The genomic–proteomic approach used in this study allowed a better understanding of Bt TOL651 pathogenicity, representing an important step for the development and monitoring of potential new bioinsecticides.
Data availability
All data and code generated appear in the submitted article.
References
Alcock BP, Huynh W, Chalil R, Smith KW, Raphenya AR, Wlodarski MA, McArthur AG (2023) CARD 2023: expanded curation, support for machine learning, and resistome prediction at the comprehensive antibiotic resistance database. Nucleic Acids Res 51:D690–D699
Andrews S (2015) FastQC: a quality-control tool for high-throughput sequence. Retrieved October from https://www.bioinformatics.babraham.ac.uk/projects/fastqc. Accessed 18 June 2020
Arthur S, Dara SK (2019) Microbial biopesticides for invertebrate pests and their markets in the United States. J Invertebr Pathol 165:13–21. https://doi.org/10.1016/j.jip.2018.01.008
Banik A, Chattopadhyay A, Ganguly S, Mukhopadhyay SK (2019) Characterization of a tea pest specific Bacillus thuringiensis and identification of its toxin by MALDI-TOF mass spectrometry. Ind Crops Prod 137:549–556. https://doi.org/10.1016/j.indcrop.2019.05.051
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Baragamaarachchi RY, Samarasekera JK, Weerasena OV, Lamour K, Jurat-Fuentes JL (2019) Identification of a native Bacillus thuringiensis strain from Sri Lanka active against Dipel-resistant Plutella xylostella. PeerJ 7:e7535. https://doi.org/10.7717/peerj.7535
Bel Y, Sheets JJ, Tan SY, Narva KE, Escriche B (2017) Toxicity and binding studies of Bacillus thuringiensis Cry1Ac, Cry1F, Cry1C, and Cry2A proteins in the soybean pests Anticarsia gemmatalis and Chrysodeixis (Pseudoplusia) includens. Appl Environ Microbiol 83:e00326-e317. https://doi.org/10.1128/AEM.00326-17
Bosi E, Donati B, Galardini M, Brunetti S, Sagot MF, Lió P, Crescenzi P, Fani R, Fondi M (2015) MeDuSa: a multi-draft based scaffolder. Bioinformatics 31:2443–2451. https://doi.org/10.1093/bioinformatics/btv171
Bravo A, Gill SS, Soberón M (2007) Mode of action of Bacillus thuringiensis cry and cyt toxins and their potential for insect control. Toxicon 49:423–435. https://doi.org/10.1016/j.toxicon.2006.11.022
Bravo A, Likitvivatanavong S, Gill SS, Soberón M (2011) Bacillus thuringiensis: a story of a successful bioinsecticide. Insect Biochem Mol Biol 41:423–431. https://doi.org/10.1016/j.ibmb.2011.02.006
Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Thomason JA, Stevens R, Vonstein V, Wattam AR, Xia F (2015) RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5:8365. https://doi.org/10.1038/srep08365
Caballero J, Jiménez-Moreno N, Orera I, Williams T, Fernández AB, Villanueva M, Ferré J, Caballero P, Ancín-Azpilicueta C (2020) Unraveling the composition of insecticidal crystal proteins in Bacillus thuringiensis: a proteomics approach. Appl Environ Microbiol 86:e00476-e520. https://doi.org/10.1128/AEM.00476-20
Cardoso P, Fazion F, Perchat S, Buisson C, Vilas-Bôas G, Lereclus D (2020) Rap-Phr systems from plasmids pAW63 and pHT8–1 act together to regulate sporulation in the Bacillus thuringiensis serovar kurstaki HD73 strain. Appl Environ Microbiol 86:e01238-e1220. https://doi.org/10.1128/AEM.01238-20
Castro BMdC, Martinez LC, Barbosa SG, Serrão JE, Wilcken CF, Soares MA, da Silva AA, de Carvalho AG, Zanuncio JC (2019) Toxicity and cytopathology mediated byBacillus thuringiensis in the midgut of Anticarsia gemmatalis (Lepidoptera: Noctuidae). Sci Rep 9:6667. https://doi.org/10.1038/s41598-019-430740
Cerqueira FB, Alves GB, Corrêa RFT, Martins ES, Barbosa LCB, do Nascimento IR, Aguiar RDS (2016) Selection and characterization of Bacillus thuringiensis isolates with a high insecticidal activity against Spodoptera frugiperda (Lepidoptera: Noctuidae). Biosci J 32:1522–1536
Chang JH, Je YH, Roh JY, Park HW, Jin BR, Lee DW, Kim S-H, Yang-W KSK (1999) Isolation and characterization of a strain of Bacillus thuringiensis serovar kenyae encoding olny δ-endotoxin Cry1E. Appl Entomol Zool 34:379–382. https://doi.org/10.1303/aez.34.379
Chung MC, Popova TG, Millis BA, Mukherjee DV, Zhou W, Liotta LA, Petricoin EF, Chandhoke V, Bailey C, SG P (2006) Secreted neutral metalloproteases of Bacillus anthracis as candidate pathogenic factors. J Biol Chem 281:31408–31418. https://doi.org/10.1016/S0021-9258(19)84053-X
da Silva SM, Silva-Werneck JO, Falcão R, Gomes AC, Fragoso RR, Quezado MT, Neto OB, Aguiar JB, de Sá MF, Bravo A, Monnerat RG (2004) Characterization of novel Brazilian Bacillus thuringiensis strains active against Spodoptera frugiperda and other insect pests. J Appl Entomol 128:102–107. https://doi.org/10.1046/j.1439-0418.2003.00812.x
da Silva IHS, de Freitas MM, Polanczyk RA (2022) Bacillus thuringiensis, a remarkable biopesticide: from lab to the field. In: Rakshit A, Meena VS, Abhilash PC, Sarma BK, Singh HB, Fraceto L, Parihar M, Singh AK (eds) Biopesticides. Woodhead Publishing, Cambridge, pp 117–131. https://doi.org/10.1016/B978-0-12-823355-9.00021-3
Dammak I, Dammak M, Tounsi S (2015) Histopathological and combinatorial effects of the metalloprotease InhA1 and Cry proteins of Bacillus thuringiensis against Spodoptera littoralis. Int J Biol Macromol 81:759–762. https://doi.org/10.1016/j.ijbiomac.2015.09.006
Daquila BV, Scudeler EL, Dossi FC, Moreira DR, Pamphile JA, Conte H (2019) Action of Bacillus thuringiensis (Bacillales: Bacillaceae) in the midgut of the sugarcane borer Diatraea saccharalis (Fabricius, 1794) (Lepidoptera: Crambidae). Ecotoxicol Environ Saf 184:109642. https://doi.org/10.1016/j.ecoenv.2019.109642
Davolos CC, Hernández-Martinez P, Crialesi-Legori PC, Desidério JA, Ferré J, Escriche B, Lemos MV (2015) Binding analysis of Bacillus thuringiensis Cry1 proteins in the sugarcane borer, Diatraea saccharalis (Lepidoptera: Crambidae). J Invertebr Pathol 127:32–34. https://doi.org/10.1016/j.jip.2015.01.013
Day M, Ibrahim M, Dyer D, Bulla L (2014) Genome sequence of Bacillus thuringiensis subsp. kurstaki strain HD-1. Genome Announc 2:e00613-00614. https://doi.org/10.1128/genomeA.00613-14
de Matos M, Santos F, Eichler P (2020) Chapter 1—Sugarcane world scenario. In: Santos F, Rabelo SC, De Matos M, Eichler P (eds) Sugarcane biorefinery, technology and perspectives. Academic Press, Cambridge, pp. 1–19. https://doi.org/10.1016/B978-0-12-814236-3.00001-9
de Oliveira WS, Sakuno CIR, Miraldo LL, Tavares MA, Komada KM, Teresani D, Santos JL, Huang F (2022) Varied frequencies of resistance alleles to Cry1Ab and Cry1Ac among Brazilian populations of the sugarcane borer, Diatraea saccharalis (F.). Pest Manage Sci 1:14. https://doi.org/10.1002/ps.7133
Dinardo-Miranda LL, Fracasso JV, Da Costa VP, Dos Anjos IA, Lopes DOP (2013) Reação de cultivares de cana-de-açúcar à broca do colmo. Bragantia Campinas 72:29–34
Ding X, Huang J, Xia L, Li X, Yuan C, Dan S (2009) A proteomic analysis approach to study insecticidal crystal proteins from different strains of Bacillus thuringiensis. Biocontrol Sci Technol 19:289–299. https://doi.org/10.1080/09583150902749984
dos Santos KB, Neves P, Meneguim AM, dos Santos RB, dos Santos WJ, Boas GV, Dumas V, Martins E, Praça LB, Queiroz P, Berry C, Monnerat R (2009) Selection and characterization of the Bacillus thuringiensis strains toxic to Spodoptera eridania (Cramer), Spodoptera cosmioides (Walker) and Spodoptera frugiperda (Smith) (Lepidoptera: Noctuidae). Biol Control 50:157–163. https://doi.org/10.1016/j.biocontrol.2009.03.014
Dulmage HTB (1973) Thuringiensis US assay standard: report on the adoption of a primary US reference standard for assay of formulations containing the delta-endotoxin of Bacillus thuringiensis. Bull Entomol Soc Am 19:200–202
Finney DJ (1971) Probit analysis. Cambridge University Press, Cambridge
Frankenhuyzen KV (2009) Insecticidal activity of Bacillus thuringiensis crystal proteins. J Invertebr Pathol 101:1–16. https://doi.org/10.1016/j.jip.2009.02.009
Frankland GC, Frankland PF (1887) XI. Studies on some new micro-organisms obtained from air. Philos Trans R Soc B 178:257–287. https://doi.org/10.1098/rstb.1887.0011
Ganesh KN, Reyaz A, Balakrishnan N (2018) Molecular characterization of an indigenous lepidopteran toxic Bacillus thuringiensis strain T532. J Biol Control 32:246–251. https://doi.org/10.18311/jbc/2018/21604
Gleave AP, Williams R, Hedges RJ (1993) Screening by polymerase chain reaction of Bacillus thuringiensis serotypes for the presence of cryV-like insecticidal protein genes and characterization of a cryV gene cloned from B. thuringiensis subsp. kurstaki. Appl Environ Microbiol 59:1683–1687. https://doi.org/10.1128/aem.59.5.1683-1687.1993
Gomis-Cebolla J, Scaramal Ricietto AP, Ferré J (2018) A genomic and proteomic approach to identify and quantify the expressed Bacillus thuringiensis proteins in the supernatant and parasporal crystal. Toxins (basel) 10:193. https://doi.org/10.3390/toxins10050193
Greene GL, Leppla NC, Dickerson WA (1976) Velvetbean caterpillar: a rearing procedure and artificial medium. J Econ Entomol 69:487–488. https://doi.org/10.1093/jee/69.4.487
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29:1072–1075. https://doi.org/10.1093/bioinformatics/btt086
Heinrichs R, Otto R, Magalhães A, Meirelles GC (2017) Importance of sugarcane in Brazilian and world bioeconomy. In: Dabbert S, Lewandowski I, Weiss J, Pyka A (eds) Knowledge-driven developments in the bioeconomy: technological and economic perspectives. Springer International Publishing, Cham, pp 205–217. https://doi.org/10.1007/978-3-319-58374-7_11
Helgason E, Økstad OA, Caugant DA, Johansen HA, Fouet A, Mock M, Hegna I, Kolstø A (2000) Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis; one Ssecies on the basis of genetic evidence. Appl Environ Microbiol 66:2627–2630. https://doi.org/10.1128/AEM.66.6.2627-2630.2000
Hensley SD, Hammond AM Jr (1968) Laboratory techniques for rearing the sugarcane borer on an artificial diet. J Econ Entomol 61:1742–1743. https://doi.org/10.1093/jee/61.6.1742
Hernández-Rodríguez CS, Hernández-Martínez P, Van Rie J, Escriche B, Ferré J (2013) Shared midgut binding sites for Cry1A. 105, Cry1Aa, Cry1Ab, Cry1Ac and Cry1Fa proteins from Bacillus thuringiensis in two important corn pests, Ostrinia nubilalis and Spodoptera frugiperda. PLoS ONE 8:e68164. https://doi.org/10.1371/journal.pone.0068164
Hire RS, Makde RD, Dongre TK, D’souza SF (2008) Characterization of the cry1Ac17 gene from an indigenous strain of Bacillus thuringiensis subsp. kenyae. Curr Microbiol 57:570–574. https://doi.org/10.1007/s00284-008-9244-3
Hire RS, Makde RD, Dongre TK, D’souza SF (2009) Expression, purification and characterization of the Cry2Aa14 toxin from Bacillus thuringiensis subsp. kenyae. Toxicon 54:519–524. https://doi.org/10.1016/j.toxicon.2009.05.022
Hongyu Z, Ziniu Y, Wangxi D (2000) Composition and ecological distribution of cry proteins and their genotypes of Bacillus thuringiensis isolates from warehouses in China. J Invertebr Pathol 76:191–197. https://doi.org/10.1006/jipa.2000.4970
Horikoshi RJ, Ferrari G, Dourado PM, Climaco JI, Vertuan HV, Evans A, Pleau M, Morrell K, José MO, Anderson H, Martinelli S, Ovejero RF, Berger GU, Head G (2022) MON 95379 Bt maize as a new tool to manage sugarcane borer (Diatraea saccharalis) in South America. Pest Manag Sci 78:3456–3466. https://doi.org/10.1002/ps.6986
Huang S, Ding X, Sun Y, Yang Q, Xiao X, Cao Z, Xia L (2012) Proteomic analysis of Bacillus thuringiensis at different growth phases by using an automated online two-dimensional liquid chromatography-tandem mass spectrometry strategy. Appl Environ Microbiol 78:5270–5279. https://doi.org/10.1128/AEM.00424-12
Huang F, Chen M, Gowda A, Clark TL, McNulty BC, Yang F, Niu Y (2015) Identification, inheritance, and fitness costs of Cry2Ab2 resistance in a field-derived population of sugarcane borer, Diatraea saccharalis (F.) (Lepidoptera: Crambidae). J Invertebr Pathol 130:116–123. https://doi.org/10.1016/j.jip.2015.07.007
Jeong H, Choi SK, Park SH (2017) Genome sequences of Bacillus thuringiensis Serovar kurstaki strain BP865 and B. thuringiensis, Serovar aizawai strain HD-133. Genome Announ 5:e01544-e1516. https://doi.org/10.1128/genomeA.01544-16
Jia N, Ding MZ, Gao F, Yuan YJ (2016) Comparative genomics analysis of the companion mechanisms of Bacillus thuringiensis Bc601 and Bacillus endophyticus Hbe603 in bacterial consortium. Sci Rep 6:28794. https://doi.org/10.1038/srep28794
Kalfon A, Larget-Thiéry I, Charles JF, Barjac H (1983) Growth, sporulation and larvicidal activity of Bacillus sphaericus. Appl Microbiol Biotechnol 18:68–173. https://doi.org/10.1007/BF00498040
Kaze M, Brooks L, Sistrom M (2021) Antibiotic resistance in Bacillus-based biopesticide products. bioRxiv. https://doi.org/10.1101/2021.03.15.435560
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Khorramnejad A, Gomis-Cebolla J, Talaei-Hassanlouei R, Bel Y, Escriche B (2020) Genomics and proteomics analyses revealed novel candidate pesticidal proteins in a lepidopteran-toxic Bacillus thuringiensis strain. Toxins (basel) 12:673. https://doi.org/10.3390/toxins12110673
Kim MJ, Han JK, Park JS, Lee JS, Lee SH, Cho JI, Kim KS (2015) Various enterotoxin and other virulence factor genes widespread among Bacillus cereus and Bacillus thuringiensis strains. J Microbiol Biotechnol 25:872–879. https://doi.org/10.4014/jmb.1502.02003
Konecka E, Baranek J, Kaznowski A, Ziemnicka J, Ziemnicki K (2012) Interaction between crystalline proteins of two Bacillus thuringiensis strains against Spodoptera exigua. Entomol Exp Appl 143:148–154. https://doi.org/10.1111/j.1570-7458.2012.01254.x
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547. https://doi.org/10.1093/molbev/msy096
Lacey LA, Grzywacz D, Shapiro-Ilan DI, Frutos R, Brownbridge M, Goettel MS (2015) Insect pathogens as biological control agents: back to the future. J Invertebr Pathol 132:1–41. https://doi.org/10.1016/j.jip.2015.07.009
Lazarte JN, Valacco MP, Moreno S, Salerno GL, Berón CM (2021) Molecular characterization of a Bacillus thuringiensis strain from Argentina, toxic against Lepidoptera and Coleoptera, based on its whole-genome and cry protein analysis. J Invertebr Pathol 183:107563. https://doi.org/10.1016/j.jip.2021.107563
Lechuga A, Lood C, Salas M, Van Noort V, Lavigne R, Redrejo-Rodríguez M (2020) Completed Genomic Sequence of Bacillus thuringiensis HER1410 reveals a Cry-containing chromosome, two megaplasmids, and an integrative plasmidial prophage. G3 Genes|genom|genet 10:2927–2939. https://doi.org/10.1534/g3.120.401361
Li X, Ding X, Xia L, Sun Y, Yuan C, Yin J (2012) Proteomic analysis of Bacillus thuringiensis strain 4.0718 at different growth phases. Sci World J. https://doi.org/10.1100/2012/798739
Liu X, Zuo M, Wang T, Sun Y, Liu S, Hu S, He H, Yang Q, Rang J, Quan M, Xia L, Ding X (2015) Proteomic analysis of the influence of Cu2+ on the crystal protein production of Bacillus thuringiensis X022. Microb Cell Factories 14:153. https://doi.org/10.1186/s12934-015-0339-9
Liu J, Li L, Peters BM, Li B, Chen D, Xu Z, Shirtliff ME (2017) Complete genome sequence and bioinformatics analyses of Bacillus thuringiensis strain BM-BT15426. Microb Pathog 108:55–60. https://doi.org/10.1016/j.micpath.2017.05.006
Liu B, Zheng D, Jin Q, Chen L, Yang J (2019) VFDB 2019: a comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res 47:D687–D692. https://doi.org/10.1093/nar/gky1080
Liu H, Zheng J, Bo D, Yu Y, Ye W, Peng D, Sun M (2021) BtToxin_Digger: a comprehensive and high-throughput pipeline for mining toxin protein genes from Bacillus thuringiensis. Bioinformatics 38:250–251. https://doi.org/10.1093/bioinformatics/btab506
Luna VA, King DS, Gulledge J, Cannons AC, Amuso PT, Cattani J (2007) Susceptibility of Bacillus anthracis, Bacillus cereus, Bacillus mycoides, Bacillus pseudomycoides and Bacillus thuringiensis to 24 antimicrobials using Sensititre® automated microbroth dilution and Etest® agar gradient diffusion methods. J Antimicrob Chemothe 60:555–567. https://doi.org/10.1093/jac/dkm213
Ma W, Chen H, Jiang X, Wang J, Gelbič I, Guan X, Zhang L (2020) Whole genome sequence analysis of the mosquitocidal Bacillus thuringiensis LLP29. Arch Microbiol 202:1693–1700. https://doi.org/10.1007/s00203-020-01875-2
Macedo CL, Martins ÉS, Macedo LL, Santos AC, Praça LB, Góis LA, Monnerat RG (2012) Seleção e caracterização de estirpes de Bacillus thuringiensis eficientes contra a Diatraea saccharalis (Lepidoptera: Crambidae). Pesq Agropecu Bras 47:1759–1765. https://doi.org/10.1590/S0100-204X2012001200012
Malovichko YV, Nizhnikov AA, Antonets KS (2019) Repertoire of the Bacillus thuringiensis virulence factors unrelated to major classes of protein toxins and its role in specificity of host-pathogen interactions. Toxins (Basel) 11:347. https://www.mdpi.com/2072-6651/11/6/347
Mendonça EG, de Almeida BR, Cordeiro G, da Silva CR, Campos WG, de Oliveira JA, de Almeida Oliveira MG (2020) Larval development and proteolytic activity of Anticarsia gemmatalis Hübner (Lepidoptera: Noctuidae) exposed to different soybean protease inhibitors. Arch Insect Biochem Physiol 103:e21637. https://doi.org/10.1002/arch.21637
Miyoshi S, Shinoda S (2000) Microbial metalloproteases and pathogenesis. Microb Infect 2:91–98. https://doi.org/10.1016/S1286-4579(00)00280-X
Monnerat RG, Silva SF, Silva-Werneck JO (2001) Catálogo do banco de germoplasma de bactérias do gênero Bacillus. Embrapa Recursos Genéticos e Biotecnologia, Brasília, p 65 (Embrapa Recursos Gene´ticos e Biotecnologia. Documentos, 60)
Monnerat RG, Batista AC, de Medeiros PT, Martins ÉS, Melatti VM, Praça LB, Dumas VF, Morinaga C, Demo C, Gomes ACM, Falcão R, Siqueira CB, Silva-Werneck JO, Berry C (2007) Screening of Brazilian Bacillus thuringiensis isolates active against Spodoptera frugiperda, Plutella xylostella and Anticarsia gemmatalis. Biol Control 41:291–295. https://doi.org/10.1016/j.biocontrol.2006.11.008
Moscardi F, Corrêa-Ferreira BS, Corso IC (2012) Pragas que atacam plântulas, hastes e pecíolos da soja. In: Hoffmann CB, Corrêa-Ferreira BS, Moscardi F (eds) Soja: manejo integrado de insetos e outros artrópodes-praga. Embrapa, Brasília, pp 145–168
Mounsef JR, Salameh D, kallassy Awad M, Chamy L, Brandam C, Lteif R (2014) A simple method for the separation of Bacillus thuringiensis spores and crystals. J Microbiol Methods 107:147–149. https://doi.org/10.1016/j.mimet.2014.10.003
Palma L, Muñoz D, Berry C, Murillo J, Caballero P (2014) Bacillus thuringiensis Toxins: an overview of their biocidal activity. Toxins (Basel) 6:3296–3325. https://www.mdpi.com/2072-6651/6/12/3296
Pezenti LF, Sosa-Gómez DR, de Souza RF, Vilas-Boas LA, Gonçalves KB, da Silva CR, Vilas-Bôas GT, Baranoski A, Mantovani MS, da Rosa R (2021) Transcriptional profiling analysis of susceptible and resistant strains of Anticarsia gemmatalis and their response to Bacillus thuringiensis. Genomics 113:2264–2275. https://doi.org/10.1016/j.ygeno.2021.05.012
Pinheiro DH, Valicente FH (2021) Identification of Bacillus thuringiensis strains for the management of lepidopteran pests. Neotrop Entomol 50:804–811. https://doi.org/10.1007/s13744-021-00896-w
Pohare MB, Wagh SG, Udayasuriyan V (2021) Bacillus thuringiensis as potential biocontrol agent for sustainable agriculture. In: Yadav AN, Singh J, Singh C, Yadav N (eds) Current trends in microbial biotechnology for sustainable agriculture. Springer Singapore, Singapore, pp 439–468. https://doi.org/10.1007/978-981-15-6949-4_18
Praça LB, Batista AC, Martins ÉS, Siqueira CB, Dias DG, Gomes AC, Falcão R, Monnerat RG (2004) Estirpes de Bacillus thuringiensis efetivas contra insetos das ordens Lepidoptera, Coleoptera e Diptera. Pesq Agropec Bras 39:11–16. https://doi.org/10.1590/S0100-204X2004000100002
Quan M, Xie J, Liu X, Li Y, Rang J, Zhang T, Zhou F, Xia L, Hu S, Sun Y, Ding X (2016) Comparative analysis of genomics and proteomics in the new isolated Bacillus thuringiensis X022 revealed the metabolic regulation mechanism of carbon flux following Cu2+ treatment. Front Microbiol 7:792. https://doi.org/10.3389/fmicb.2016.00792
Rang J, He H, Wang T, Ding X, Zuo M, Quan M, Sun Y, Yu Z, Hu S, Xia L (2015) Comparative analysis of genomics and proteomics in Bacillus thuringiensis 4.0718. PLoS ONE 10:e0119065. https://doi.org/10.1371/journal.pone.0119065
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Richter M, Rosselló-Móra R, Oliver Glöckner F, Peplies J (2015) JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32:929–931. https://doi.org/10.1093/bioinformatics/btv681
Robertson JL, Jones MM, Olguin E, Alberts B (2017) Bioassays with arthropods, 3rd edn. CRC Press, Taylor & Francis Group, Boca Raton
Sanahuja G, Banakar R, Twyman RM, Capell T, Christou P (2011) Bacillus thuringiensis: a century of research, development and commercial applications. Plant Biotechnol J 9:283–300. https://doi.org/10.1111/j.1467-7652.2011.00595.x
Santos MD, Lima DB, Fischer JS, Clasen MA, Kurt LU, Camillo-Andrade AC, Monteiro LC, de Aquino PF, Neves-Ferreira AG, Valente RH, Trugilho MR, Brunoro GV, Souza TA, Santos RM, Batista M, Gozzo FC, Durán R, Yates JR, Barbosa VC, Carvalho PC (2022) Simple, efficient and thorough shotgun proteomic analysis with PatternLab V. Nat Protoc 17:1553–1578. https://doi.org/10.1038/s41596-022-00690-x
Sathyan T, Jayakanthan M, Mohankumar S, Balasubramani V, Kokiladevi E, Ravikesavan R, Sathiah N (2022) Genome profiling of an indigenous Bacillus thuringiensis isolate, T405 toxic against the fall armyworm, Spodoptera frugiperda (JE Smith) (Lepidoptera: Noctuidae). Microb Pathog 173:105820. https://doi.org/10.1016/j.micpath.2022.105820
Schmidt FG, Monnerat RG, Borges M, Carvalho R (2001) Metodologia de criação de insetos para a avaliação de agentes entomopatogênicos. In: e ERG (ed) Biotecnologia. Circular Técnica, Brasília
Schnepf E, Crickmore N, Rie JV, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62:775–806. https://doi.org/10.1128/MMBR.62.3.775-806.1998
Silva MTB (1995) Associação de Baculovirus anticarsia com subdosagem de inseticidas no controle de Anticarsia gemmatalis (Hübner, 1818). Ciência Rural 25:353–358
Singh D, Samiksha Thayil SM, Sohal SK, Kesavan AK (2021) Exploration of insecticidal potential of cry protein purified from Bacillus thuringiensis VIID1. Int J Biol Macromol 174:362–369. https://doi.org/10.1016/j.ijbiomac.2021.01.143
Sreshty MAL, Kumar KP, Murty USN (2011) Synergism between wild-type Bacillus thuringiensis subsp. israelensis and B. sphaericus strains: a study based on isobolographic analysis and histopathology. Act Tropica 118:14–20. https://doi.org/10.1016/j.actatropica.2010.12.012
Srikanth J, Subramonian N, Premachandran MN (2011) Advances in transgenic research for insect resistance in sugarcane. Trop Plant Biol 4:52–61. https://doi.org/10.1007/s12042-011-9077-2
Sun D, Zhu L, Guo L, Wang S, Wu Q, Crickmore N, Zhou X, Bravo A, Soberón M, Guo Z, Zhang Y (2022) A versatile contribution of both aminopeptidases N and ABC transporters to Bt Cry1Ac toxicity in the diamondback moth. BMC Biol 20:33. https://doi.org/10.1186/s12915-022-01226-1
Tailor R, Tippett J, Gibb G, Pells S, Jordan L, Ely S (1992) Identification and characterization of a novel Bacillus thuringiensis δ-endotoxin entomocidal to coleopteran and lepidopteran larvae. Mol Microbiol 6:1211–1217. https://doi.org/10.1111/j.1365-2958.1992.tb01560.x
Tan SY, Cayabyab BF, Alcantara EP, Ibrahim YB, Huang F, Blankenship EE, Siegfried BD (2011) Comparative susceptibility of Ostrinia furnacalis, Ostrinia nubilalis and Diatraea saccharalis (Lepidoptera: Crambidae) to Bacillus thuringiensis Cry1 toxins. Crop Protect 30:1184–1189. https://doi.org/10.1016/j.cropro.2011.05.009
Wu D, He J, Gong Y, Chen D, Zhu X, Qiu N, Sun M, Li M, Yu Z (2011) Proteomic analysis reveals the strategies of Bacillus thuringiensis YBT-1520 for survival under long-term heat stress. Proteomics 11:2580–2591. https://doi.org/10.1002/pmic.201000392
Xie J, Peng J, Yi Z, Zhao X, Li S, Zhang T, Quan M, Yang S, Lu J, Zhou P, Xia L, Ding X (2019) Role of hsp20 in the production of spores and insecticidal crystal proteins in Bacillus thuringiensis. Front Microbiol 10:2059. https://doi.org/10.3389/fmicb.2019.0205
Ye J, Zhang Y, Cui H, Liu J, Wu Y, Cheng Y, Xu H, Huang X, Li S, Zhou A, Zhang X, Bolund L, Chen Q, Wang J, Yang H, Fang L, Shi C (2018) WEGO 2.0: a web tool for analyzing and plotting GO annotations, 2018 update. Nucleic Acids Res 46:W71–W75. https://doi.org/10.1093/nar/gky400
Yin J, Ding X, Xia L, Yu Z, Lv Y, Hu S, Xiao X (2011) Transcription of gene in an acrystalliferous strain of Bacillus thuringiensis XBU001 positively regulated by the metalloprotease camelysin gene at the onset of stationary phase. FEMS Microbiol Lett 318:92–100. https://doi.org/10.1111/j.1574-6968.2011.02247.x
Zghal RZ, Ghedira K, Elleuch J, Kharrat M, Tounsi S (2018) Genome sequence analysis of a novel Bacillus thuringiensis strain BLB406 active against Aedes aegypti larvae, a novel potential bioinsecticide. J Biol Macromol 116:1153–1162. https://doi.org/10.1016/j.ijbiomac.2018.05.119
Zhou H, Zhang J, Shao Y, Wang J, Xu W, Liu Y, Yu S, Ye Q, Pang R, Wu S, Gu Q, Xue L, Zhang J, Li H, Wu Q, Ding Y (2022) Development of a high resolution melting method based on a novel molecular target for discrimination between Bacillus cereus and Bacillus thuringiensis. Food Res Int 151:110845. https://doi.org/10.1016/j.foodres.2021.110845
Zhu L, Peng D, Wang Y et al (2015) Genomic and transcriptomic insights into the efficient entomopathogenicity of Bacillus thuringiensis. Sci Rep 5:14129. https://doi.org/10.1038/srep14129
Funding
This work was supported by grants from the National Council of Scientific and Technological Development (CNPq—process numbers: 313455/2019-8; 427304/2018-0; 163178/2020-8), Tocantins State Foundation for Research Aid (FAPT-SESAU/TO-DECIT/SCTIE/MS_CNPQ/N° 01/2017) and Federal University of Tocantins (PROPESQ)—EDITAL Nº 29/2020 PROPESQ and PPGBIOTEC/UFT/GURUPI- Chamada pública para auxílio de tradução e/ou publicação de artigos científicos-EDITAL Nº 011/2020.
Author information
Authors and Affiliations
Contributions
RWSA, GBA, and EEO designed the experiments. GBA, RWSA, and BMR performed the genomic and proteomic data analysis. GBA, LOVJ, GRS, BMR, and RWSA drafted the manuscript. RWSA, EEO, LOVJ, BMR, MAO, and MMS revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no financial interest.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alves, G.B., de Oliveira, E.E., Jumbo, L.O.V. et al. Genomic–proteomic analysis of a novel Bacillus thuringiensis strain: toxicity against two lepidopteran pests, abundance of Cry1Ac5 toxin, and presence of InhA1 virulence factor. Arch Microbiol 205, 143 (2023). https://doi.org/10.1007/s00203-023-03479-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03479-y