Abstract
Species tree estimation faces many significant hurdles. Chief among them is that the trees describing the ancestral lineages of each individual gene—the gene trees—often differ from the species tree. The multispecies coalescent is commonly used to model this gene tree discordance, at least when it is believed to arise from incomplete lineage sorting, a population-genetic effect. Another significant challenge in this area is that molecular sequences associated to each gene typically provide limited information about the gene trees themselves. While the modeling of sequence evolution by single-site substitutions is well-studied, few species tree reconstruction methods with theoretical guarantees actually address this latter issue. Instead, a standard—but unsatisfactory—assumption is that gene trees are perfectly reconstructed before being fed into a so-called summary method. Hence much remains to be done in the development of inference methodologies that rigorously account for gene tree estimation error—or completely avoid gene tree estimation in the first place. In previous work, a data requirement trade-off was derived between the number of loci m needed for an accurate reconstruction and the length of the locus sequences k. It was shown that to reconstruct an internal branch of length f, one needs m to be of the order of \(1/[f^{2} \sqrt{k}]\). That previous result was obtained under the restrictive assumption that mutation rates as well as population sizes are constant across the species phylogeny. Here we further generalize this result beyond this assumption. Our main contribution is a novel reduction to the molecular clock case under the multispecies coalescent, which we refer to as a stochastic Farris transform. As a corollary, we also obtain a new identifiability result of independent interest: for any species tree with \(n \ge 3\) species, the rooted topology of the species tree can be identified from the distribution of its unrooted weighted gene trees even in the absence of a molecular clock.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Modern sequencing technologies have provided a wealth of data to assist biologists in the inference of evolutionary relationships between species. It is now common to sequence thousands of genes, or entire genomes, simultaneously across a range of species. With this abundance of data comes new algorithmic and statistical challenges. One such challenge arises because phylogenomic inference entails dealing with the interplay of two processes. While a species phylogeny depicts the history of speciation of extant organisms, each gene within the genomes of these organisms has its own history. That history is captured by a gene tree. In practice, by contrasting the molecular sequences of a gene (or other genomic region) across species, one can reconstruct the corresponding gene tree. Indeed the accumulation of mutations along the gene tree reflects, if imperfectly, the underlying history. Much is known about the reconstruction of single-gene trees, a subject with a long history. See Steel (2016), Warnow (2017) for an overview.
But a gene tree does not necessarily coincide with the species phylogeny. In particular, various mechanisms lead to discordance between gene trees. These include the transfer of genetic material between species, hybrid speciation events and a population-genetic effect known as incomplete lineage sorting (Maddison 1997). The wide availability of genomic datasets has brought to the fore the major impact these discordances have on phylogenomic inference (Degnan and Rosenberg 2009). As a result, in addition to the stochastic process governing the evolution of molecular sequences on a fixed gene tree, one is led to model the structure of the gene tree itself, in relation to the species phylogeny, through a separate stochastic process. The inference of these complex, two-level evolutionary models is an active area of research.
We focus on incomplete lineage sorting (ILS) and consider the problem of reconstructing a species phylogeny from multiple genes under the multispecies coalescent (Rannala and Yang 2003), a standard population-genetic model. The problem is of great practical interest in computational evolutionary biology and is currently the subject of intense study; see e.g. Nakhleh (2013), Kapli et al. (2020), Scornavacca et al. (2020). There is in particular a growing body of theoretical results in this area (Degnan and Rosenberg 2006; Degnan et al. 2009; DeGiorgio and Degnan 2010; Mossel and Roch 2010; Liu et al. 2010; Allman et al. 2011; Roch 2013; Roch and Steel 2015; DeGiorgio and Degnan 2014; Dasarathy et al. 2015; Chifman and Kubatko 2015; Roch and Warnow 2015; Mossel and Roch 2017; Shekhar et al. 2017; Long and Kubatko 2017; Roch 2018; Roch et al. 2019; Allman et al. 2018, 2019; Long and Kubatko 2019; Rhodes 2019). A significant portion of prior rigorous work on species phylogeny estimation in the presence of ILS has been aimed at the case where true gene trees are assumed to be available. In reality, one needs to estimate gene trees from molecular sequences, leading to reconstruction errors, and indeed there has been a recent thrust towards understanding the effect of this important source of error in phylogenomic inference, both from empirical (Kubatko and Degnan 2007; Bayzid and Warnow 2013; Mirarab et al. 2014, 2016) and theoretical (DeGiorgio and Degnan 2014; Roch and Steel 2015; Bayzid et al. 2015; Roch and Warnow 2015) standpoints. Another option—which we further explore here—is to bypass the reconstruction of gene trees altogether and infer the species history directly from sequence data (Dasarathy et al. 2015; Mossel and Roch 2017; Chifman and Kubatko 2015; Rusinko and McPartlon February 2017; Allman et al. 2019; Long and Kubatko 2019).
In previous work (Mossel and Roch 2017), a surprising trade-off was derived between the number of genes m needed to accurately reconstruct a species phylogeny and the length of the genes k. Specifically, it was shown that m needs to scale like \(1/[f^{2} \sqrt{k}]\), where f is the length of the shortest branch in the tree (measured in expected number of substitutions per unit of time per site; see Sect. 2 for formal definitions). This result was obtained under a restrictive molecular clock assumption where the leaves are “equidistant” from the root; i.e., it was assumed that the mutation rates and population sizes do not vary across species.
In this work, we make progress towards designing species tree estimation methods that provably achieve the theoretical limit by relaxing the above assumption. Our key contribution is of independent interest. We show how to transform sequence data to appear as though it was generated under the multispecies coalescent with a molecular clock. We achieve this through a novel reduction which we call a stochastic Farris transform. Our construction relies on an identifiability result which is partly new: for any species phylogeny with \(n \ge 3\) species, the rooted topology of the species tree can be identified from the distribution of the unrooted weighted gene trees even in the absence of a molecular clock. We state our results in Sect. 2 and describe our reduction in Sect. 3. The main proofs are in Sects. 4 and 5.
Related work A common approach to species tree estimation that bypasses gene trees is to (1) concatenate the aligned gene sequences and (2) apply a standard phylogenetic reconstruction method (under the incorrect assumption that all sites have evolved on a fixed tree), such as maximum likelihood or a distance-based method, to the concatenated data. That approach has been shown to have serious theoretical drawbacks. Indeed, in Roch and Steel (2015), it was proved that, under the multispecies coalescent with a standard site substitution model (see Sect. 2), maximum likelihood on a concatenated alignment is statistically inconsistent, that is, it can converge on the wrong phylogeny even as the amount of data grows to infinity. The result in Roch and Steel (2015) allows for gene sequence lengths to vary with the number of genes. In follow-up work (Roch et al. 2019), it was shown that fully partitioned concatenated maximum likelihood (that is, maximizing the likelihood of the sequence data under the assumption that the tree topology is fixed across genes but allowing for branch lengths to vary) is also statistically inconsistent. That result was established under bounded gene sequence lengths.
On the other hand, positive results have also been obtained for some concatenation-based approaches under the multispecies coalescent. In Dasarathy et al. (2015), a notion of distance between taxa defined over a concatenated alignment was shown to satisfy the four-point condition (see, e.g., Semple and Steel (2003)) in the limit of infinitely many genes, for any fixed gene sequence length. The latter ensures that the unrooted topology of the species phylogeny is identifiable from the sequence data, that is, that species phylogenies with distinct unrooted topologies necessarily produce distinct data distributions. The results in Dasarathy et al. (2015) allow for varying population sizes and mutation rates across branches and were proved under the Jukes-Cantor substitution model. They also come with a data requirement guarantee: using an appropriate distance-based method, for any gene sequence length \(k \ge 1\), the correct unrooted species tree topology is guaranteed to be recovered with high probability provided the number of genes m scales roughly like \(\propto e^{C \Delta } \log n\) where \(\Delta \) is the depth of the tree (see, e.g., Erdos et al. 1999), n is the number of leaves and C is a universal constant.
The distance-based pipeline introduced in Dasarathy et al. (2015), specifically in combination with the reconstruction method Neighbor Joining (Saitou and Nei 1987) (but under a more general model of site substitution), was tested on simulated datasets in Rusinko and McPartlon (February 2017). Moreover, the theoretical identifiability results of Dasarathy et al. (2015) were extended to a significantly broader class of site substitution models in Allman et al. (2019) using the concept of log-det distance (see, e.g., Semple and Steel 2003). The model considered in Allman et al. (2019) allows for an arbitrary mixture of general time-reversible rate matrices across the genome and population sizes that vary on each branch of the species tree as a function of time.
There has also been closely related work on single nucleotide polymorphism (SNP)-based approaches (that is, the case \(k=1\)). In particular, it was shown in Chifman and Kubatko (2015) that the unrooted topology of the species phylogeny is identifiable given observed data at the leaves of the tree that are assumed to have arisen from the multispecies coalescent under a time-reversible substitution process with site-specific rate variation modeled by the discrete gamma distribution and a proportion of invariable sites. The results of Chifman and Kubatko (2015), which led to a practical SNP-based approach (Chifman and Kubatko 2014), were extended in Long and Kubatko (2019) to a modified model that allows for varying population sizes and mutation rates.
Finally, the data requirement bounds in Dasarathy et al. (2015) were substantially improved in Mossel and Roch (2017) using a different reconstruction approach, but under more restrictive assumptions (see Sect. 2). One of our main contributions here is to relax these assumptions while preserving roughly the same data requirements.
2 Background and results
We begin with a description of our modeling assumptions. More details on these standard models can be found for example in Steel (2016).
Species phylogeny v. gene trees A species phylogeny is a graphical depiction of the evolutionary history of a set of species. The leaves of the tree correspond to extant species while internal vertices indicate a speciation event. Each edge (or branch) corresponds to an ancestral population and will be described here by two numbers: one that indicates the amount of time that the corresponding population lived, and a second one that specifies the rate of genetic mutation in that population. Formally, we define the species phylogeny as follows. Throughout, we use the notation \([n] = \{1,\ldots ,n\}\).
Definition 1
(Species phylogeny). A species phylogeny \(S = (V_s, E_s; r, \vec {\tau },\vec {\mu })\) is a directed tree rooted at \(r\in V_s\) with vertex set \(V_s\), edge set \(E_s\), and n labelled leaves \(L = [n]\) such that (a) the degree of all internal vertices is 3 except for the root r which has degree 2, and (b) each edge \(e\in E_s\) is associated with a length \(\tau _e\in (0,\infty )\) and a mutation rate \(\mu _e \in (0,\infty )\).
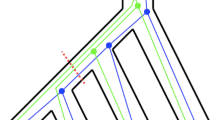
Our model involves several natural time scales, each of which can be used as a time unit. See, e.g., Allman et al. (2019) for a discussion. Here we choose to measure the length \(\tau _e\) of a branch \(e\in E_s\) in coalescent time units, which is the duration of the branch \(t_e\) in number of generations divided by twice its population size \(N_e\). That is, \(\tau _e = t_e/2 N_e\). The mutation rates \(\mu _e\) are expressed in expected number of substitutions per site per unit of time, where time is measured in coalescent units again. We pictorially represent species phylogenies as thick shaded trees; see Fig. 1. The goal in our applications will be to reconstruct the rooted topology of the species phylogeny \(S = (V_s, E_s; r, \vec {\tau },\vec {\mu })\), that is, the rooted tree \((V_s, E_s, r)\).
While a species phylogeny describes the history of speciation, each gene has its own history captured by a gene tree.
Definition 2
(Gene trees). Gene tree \(G^{(i)} = (V^{(i)}, E^{(i)}; {R}, \vec {\delta }^{(i)})\) of gene i is a directed tree rooted at R with vertex set \(V^{(i)}\) and edge set \(E^{(i)}\), and the same labeled leaf set \(L = [n]\) as S such that (a) the degree of each internal vertex is 3, except the root R whose degree is 2, and (b) each branch \(e\in E^{(i)}\) is associated with a branch length \(\delta _e^{(i)}\in (0,\infty )\).
The branch lengths \(\delta _e^{(i)}\) are expressed in expected number of substitutions per site. The precise relationship between the branch lengths of the species phylogeny and those of the gene trees under our modeling assumptions will be given below in (2).
Multispecies coalescent and Jukes-Cantor models While the species phylogeny is assumed to be fixed (but unknown), the gene trees are random. Their distribution depends on the species phylogeny. Specifically, we assume that a multispecies coalescent (MSC) process produces m independent gene trees \(G^{(1)}\), \(G^{(2)}\), \(\ldots ,\) \(G^{(m)}\). This process is parametrized by the species phylogeny S. In words, proceeding backwards in time, in each population, every pair of lineages entering from descendant populations merges at exponential rate one in coalescent time units. One key feature of the gene trees is the following: their topology may be distinct from that of the species phylogeny. This discordance, which in this context is referred to as incomplete lineage sorting, is a major challenge for species tree estimation from multiple genes. See again Figure 1 for an illustration.
Gene trees are not observed directly. They are inferred from sequence data evolving on the gene trees. We model this sequence generation process according to the standard Jukes-Cantor (JC) model. Given a gene tree \(G^{(i)} = (V^{(i)},E^{(i)}; r, \vec {\delta }^{(i)})\), we associate to each \(e\in E^{(i)}\), a probability \(p_{e}^{(i)} = \frac{3}{4}\left( 1 - e^{-\frac{4}{3}\delta _{e}^{(i)}} \right) \). In words, the corresponding gene i is a sequence of length k in \(\left\{ \mathtt{A, T, G, C}\right\} ^k\). Each position in the sequence evolves independently, starting from a uniform state in \(\left\{ \mathtt{A, T, G, C}\right\} \) at the root. Moving away from the root, a substitution occurs on edge e with probability \(p_{e}^{(i)}\), in which case the state changes uniformly at random. Repeating this process for each position produces a sequence of length k for all leaves of \(G^{(i)}\), for each \(i \in [m]\)—our input.
Tree metrics It remains to describe the relationship between the branch lengths of the species phylogeny and those of the gene trees. For this purpose, we recall some notions on tree metrics.
Definition 3
(Species metric). A species phylogeny \(S = (V_s, E_s; r, \vec {\tau }, \vec {\mu })\) induces the following metric on the leaf set L. For any pair of leaves \(a,b\in L\), we let
where \(\pi (a,b;S)\) is the unique path connecting a and b in S interpreted as a set of edges. We will refer to \(\left\{ \mu _{ab} \right\} _{a,b\in L}\) as the species metric induced by S.
The species metric \(\mu \) can be naturally extended to the entire set \(V_s\) by using (1) for an arbitrary pair of vertices. The metric \(\left\{ \mu _{ab} \right\} _{a,b\in L}\) is said to be ultrametric when \(\mu _{ra} = \mu _{rb}\) for all \(a, b \in L\), that is, when the distance from the root to every leaf is the same. We do not make this assumption here. In particular, we allow mutation rates and population sizes to vary across branches of the species phylogeny. Instead, the key to our contribution is a transformation of the sequence data that mimics an ultrametric case. Details on this transformation are given below in Algorithm 1 and Defintion 5.
Recall from Definition 2 that each gene tree has an associated set of branch lengths. Under the MSC, a single branch of a gene tree may span across multiple branches of the species phylogeny; this can also be seen in Fig. 1. Let \(t_{\tilde{e}}\) denote the length of the branch \(\tilde{e}\in E^{(i)}\) in coalescent time units. For any species phylogeny branch \(e\in E_s\), let \(t_{\tilde{e}\cap e}\) denote the length of the branch \(\tilde{e}\) that overlaps with e. Then, \(\delta _{\tilde{e}}\) and \(t_{\tilde{e}}\) satisfy the following relationship
This set of weights defines a different metric on the leaves. Note that, under the MSC, these weights are random. They also lead to a metric which will play a central role in our approach.
Definition 4
(Gene metric). A gene tree \(G^{(i)} = (V^{(i)}, E^{(i)}; {R^{(i)}}, \vec {\delta }^{(i)})\) induces the following metric on the leaf set L. For any pair of leaves \(a,b\in L\), we let \(\delta _{ab}^{(i)} = \sum _{e\in \pi (a,b;G^{(i)})}\delta _e^{(i)}\) where, again, \(\pi (a,b;G^{(i)})\) is the unique path connecting a and b in \(G^{(i)}\) interpreted as a set of edges. We will refer to \(\{ \delta _{ab}^{(i)} \}_{a,b\in L}\) as the gene metric induced by \(G^{(i)}\).
The gene metric \( \delta ^{(i)}\) can be extended to the entire set \(V_s\) and we say that it is ultrametric if \(\delta _{ra}^{(i)} = \delta _{rb}^{(i)}\) for all \(a, b \in L\). Throughout, the \(\mu _{ab}\)’s are deterministic (but unknown) while the \(\delta _{ab}\)’s are random.
Inference problem For gene i, we let \(\left\{ \xi ^{ij}_x \,:\, j\in [k], x\in L \right\} \) denote the data generated at the leaves L of the tree \(G^{(i)}\) by the Jukes-Cantor process, the superscript j runs across the positions of the gene sequence. To simplify the notation, we denote \(\xi ^{ij} = (\xi ^{ij}_x)_{x \in L}\). The species phylogeny estimation problem can then be stated as:
The \(n\times m\times k\) data array \(\left\{ \xi ^{ij} \right\} _{i\in [m], j\in [k]}\) is generated according to the Jukes-Cantor process on the m gene trees, each of which is generated by the multispecies coalescent on S. The goal is to recover the topology of the species phylogeny S from \(\left\{ \xi ^{ij} \right\} _{i\in [m], j\in [k]}\).
We refer to this two-step process as the MSC-JC(m, k) process on S.
A “stochastic” Farris transform Previous work in Mossel and Roch (2017) on tight data requirement trade-offs under the MSC-JC model was restricted to the case where mutation rates and population sizes do not vary across the species phylogeny. This results in the species metric \(\mu _{ab}\) being in fact ultrametric, as defined above. That, in turn, implies that the gene metrics \(\delta ^{(i)}\) are also ultrametric. That property produces symmetries that are useful in the design and analysis of reconstruction algorithms. Our main contribution here is a reduction to this ultrametric case.

That is, we transform the input sequences to appear as though they were generated (approximately) by a species phylogeny with an ultrametric species metric. This ultrametric reduction, inspired by a classical technique known as the Farris transform, may be of independent interest as it could be used to generalize other reconstruction algorithms. Further details and intuition about the Farris transform are given in Sect. 3.1. At a high level, this transform relates a general tree metric to an ultrametric over the same topology. Here we split the data and use one piece to estimate quantities necessary to perform a randomized version of the transform.
Although our reduction could be applied to a dataset of arbitrary size, for ease of presentation we fix a triple of leaves \(\mathcal {X} = [3]\). Specifically, Algorithm 1 takes as input a set of genes [m] divided into two disjoint subsets, \(\mathcal {M}_{\mathrm{R}}\) and \(\mathcal {M}_{\mathrm{Q}}\). The set \(\mathcal {M}_{\mathrm{R}}\) is used to estimate parameters needed for the reduction. The reduction is subsequently performed on \(\mathcal {M}_{\mathrm{Q}}\). For \(\phi >0\), we say that two metrics \(\mu '\) and \(\mu ''\) over \(\mathcal {X}\) are \(\phi \)-close if \(\left| \mu '_{xy} - \mu ''_{xy} \right| \le \phi \), for all \(x, y\in \mathcal {X}\). For convenience, we also say that two species phylogenies are \(\phi \)-close if their species metrics are. In essence, we show that \(m \ge 1/(k\phi ^2)\) genes suffice to output a dataset that is \(\phi \)-close to ultrametric. Given that \(1/\phi ^{2}\) independent sites are required to estimate distances within \(\phi \) Steel and Székely (2002), our bound on the total number of sites mk likely cannot be improved. Throughout, for \(a, b \in \mathbb {R}\), we use the notation \(a \vee b = \max \{a,b\}\).
Theorem 1
(Ultrametric reduction). Suppose we have sequence data \(\left\{ \xi ^{ij} \right\} _{i\in [m], j\in [k]}\) generated under the MSC-JC(m, k) process on a three-species phylogeny \(S = (V_s, E_s; r, \vec {\tau },\vec {\mu })\). The mutation rates, leaf-edge lengths and internal-edge lengths are respectively in the intervals \((\mu _L, \mu _U)\), \((f', g')\) and (f, g). Then, there are constants \(c_{}', c_{}'' > 0\) such that, for any \(\varepsilon , \phi \in (0,1)\) satisfying
with probability at least \(1-\varepsilon \), the output of Algorithm 1 is distributed according to the MSC-JC process on a species phylogeny \(S'\) that is \(\phi \)-close to a species phylogeny with an ultrametric species metric and a rooted topology identical to that of S, provided
Application: Data requirement trade-off Our main application of the ultrametric reduction is an extension of the data requirement trade-off in Mossel and Roch (2017) without the assumption that mutation rates and population sizes do not vary across the species phylogeny. After applying our reduction, we use the quantile triplet test developed in Mossel and Roch (2017). Roughly speaking this test, which is detailed in Algorithm 2 in the appendix, compares a well-chosen quantile of the sequence-based estimates of gene metrics \(\left\{ \delta _{ab}^{(i)} \right\} _{a,b\in \mathcal {X}}\) in order to determine which pair of leaves is closest.
For any leaves \(x, y, z \in L\), the species phylogeny S restricted to these three leaves has one of three possible rooted topologies: xy|z, xz|y, or yz|x. It is a classical phylogenetic result that if one is able to correctly reconstruct the topology of all triples of leaves in L, then the topology of the full species phylogeny can be correctly reconstructed as well (see e.g., Steel 2016). Hence, we restrict to the case \(\mathcal {X} = [3]\). Our data requirement applies to an unknown species phylogeny in the following class. We assume that: mutation rates are in the interval \((\mu _L, \mu _U)\); leaf-edge lengths are in \((f', g')\); and internal-edge lengths are in (f, g). We suppress the dependence on \(\mu _L, \mu _U, f', g', g\), which we think of as constants, and focus here on the role of f, which is known to play a critical role. Specifically, we answer the following question: as \(f \rightarrow 0\), how many genes m of length k are needed for a correct reconstruction with high probability? We obtain the same trade-off between m and k as in Mossel and Roch (2017), whose proof only applies to the ultrametric case.
Theorem 2
(Data requirement). Suppose that we have sequence data \(\left\{ \xi ^{ij} \right\} _{i\in [m], j\in [k]}\) generated according to the MSC-JC(m, k) process on a species phylogeny \(S = (V_s, E_s; r, \vec {\tau },\vec {\mu })\) . The mutation rates, leaf-edge lengths and internal-edge lengths are respectively in \((\mu _L, \mu _U)\), \((f', g')\) and (f, g). We assume further that there are \(C, C' > 0\) such that \(k = f^{-C}\) and \(\varepsilon , \phi \in (0,1)\) satisfy (3) with \(\phi = C' f/ \log f^{-1}\). Then, there exists a universal constant \(c''' > 0\) such that Algorithm 2 correctly identifies the rooted topology of S restricted to \(\mathcal {X} = [3]\) with probability at least \(1-\varepsilon \) provided that
3 Main steps of the proof of Theorem 1
The goal of the ultrametric reduction step, Algorithm 1, is to transform the sequence data to appear statistically as though it is the output of an MSC-JC process on an ultrametric species phylogeny with the same topology as S restricted to \(\mathcal {X}\). Here we provide an overview of the key ideas behind this step.
3.1 Preliminary step: an identifiability result
Before diving into the description of Algorithm 1, we provide some insights into the algebra of our reduction by first proving an identifiability result, which is partly new.
It was shown in (Allman et al. 2011, Theorem 9) that the distribution of the unrooted topologies of the gene trees suffices to identify the rooted topology of the species phylogeny when the number of leaves exceeds 4. In fact, even more was shown in that case: the species metric (in coalescent time units, that is, taking \(\mu _e = 1\) for all e in our notation) can be recovered from the same information. On the other hand, it was also proved in (Allman et al. 2011, Proposition 3) that, when \(n=4\), the gene tree topologies are not enough to locate the root of the species phylogeny.
We complement these previous results by revisiting the cases \(n=3, 4\) when information about gene tree branch lengths is available. Here we show that, already with three species (and therefore when \(n \ge 3\)), this extra information allows to recover the rooted topology of the species phylogeny. Branch length information plays a critical role in achieving the data requirement in Theorem 2. We give a constructive proof of Theorem 3 below, leading to Algorithm 1.
Theorem 3
(Identifiability of rooted topology of species phylogeny from gene metrics). Let \(S = (V_s,E_s; r, \vec {\tau }, \vec {\mu })\) be a species phylogeny with \(n \ge 3\) leaves and root r and let \(G=(V,E; {R}, \vec {\delta })\) be a gene tree sampled from the MSC with corresponding (random) gene metric \(\{ \delta _{ab}\}_{a,b\in L}\). Then the rooted topology of the species phylogeny is identifiable from the distribution of the gene metric.
Our proof is inspired by the Farris transform (also related to the Gromov product; see e.g. Semple and Steel 2003), a classical technique to transform a general tree metric into an ultrametric. In a typical application of the Farris transform, one “roots” the species phylogeny S at an “outgroup” o (i.e., a species that is “far away” from the leaves of S) and then uses the quantities \(\mu _{ox}, x\in L\) to implicitly stretch the leaf edges appropriately, so as to make all inter-species distances to o equal, without changing the underlying topology. More specifically, let S be a species phylogeny. Suppose \(\mathcal {X} = [3]\) and let \(o \in L-\mathcal {X}\) be any leaf of S outside \(\mathcal {X}\). Assume that \( \mu _{o1} \ge \max \{\mu _{o2}, \mu _{o3}\} \) (the other cases being similar) and define the Farris transform
A standard phylogenetic result (proved for instance in Semple and Steel 2003, Lemma 7.2.2) states that \(\{\dot{\mu }_{xy}\}_{x,y \in \mathcal {X}}\) is an ultrametric on \(\mathcal {X}\) consistent with the topology of S re-rooted at o and, then, restricted to \(\mathcal {X}\).
In the multi-gene context, however, we cannot apply a Farris transform in this manner. In particular, we do not have direct access to \(\{\mu _{xy}\}\); rather, we only estimate the gene metrics \(\{\delta _{xy}^{(i)}\}\). Moreover the latter vary across genes under the MSC.
Key idea 1: We artificially fix rooted gene tree topologies through conditioning. Doing so allows us to relate species and gene metrics.
We prove Theorem 3 for \(n=3\), which suffices.Footnote 1 Let S be a species phylogeny with three leaves and recall that r is the root of S. Let \(G=(V,E; {R}, \vec {\delta })\) be a gene tree sampled from the MSC with corresponding (random) gene metric \(\{ \delta _{ab}\}_{a,b\in L}\). Unlike the classical Farris transform above, we do not use an outgroup. Instead, we show how to achieve the same outcome by using only the distribution of G and, in particular, of \(\{ \delta _{ab}\}_{a,b\in L}\). Notice from (6) that we only need the differences \( \Delta _{xy} := \mu _{rx} - \mu _{ry}. \) It is these quantities that we derive from the distribution of the gene metric. The high-level idea is to:
-
1.
Condition on an event such that the rooted topology of the gene tree is guaranteed to be equal to xy|z. Intuitively, we achieve this by considering an event where one pair of leaves is “somewhat close” while the other two pairs are “somewhat far.” We make the latter condition precise in Proposition 1 below.
-
2.
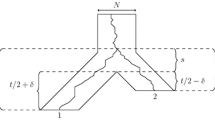
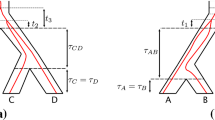
Recover the species-based difference \(\Delta _{xy}=\mu _{rx} - \mu _{ry}\) from the gene-based difference \(\delta _{xz} - \delta _{yz}\). Indeed, when the rooted topology of G is xy|z, then the difference \(\delta _{xz} - \delta _{yz}\) is equal to \(\Delta _{xy}\) This is established in Proposition 2 below. See Fig. 2 for an illustration.
More formally, we establish the following two propositions, whose proofs are in Sect. 4. For \(x,y \in L\) and \(\beta \in [0,1]\), let \(\delta _{xy}^{(\beta )}\) be the \(\beta \)th quantile of \(\delta _{xy}\). That is, \(\delta _{xy}^{(\beta )}\) is the smallest number \(a\in [0,1]\) such that \(\mathbb {P}\left[ \delta _{xy} \le a \right] \ge \beta .\) Note that this quantile is a function of the distribution of G.
Proposition 1
(Fixing the rooted topology of the gene tree). Let (x, y, z) be an arbitrary permutation of (1, 2, 3). The following event has positive probability and implies that the rooted topology of G is xy|z:
Conditioning on \(\mathscr {E}_{I}\), we then show how to recover the difference \(\Delta _{xy}\) from \(\delta _{xz} - \delta _{yz}\).
Proposition 2
(A formula for the height difference). We have that, conditioned on \(\mathscr {E}_{I}\), almost surely
The quantity on the l.h.s. of (8) is a function of the distribution of G. From the \(\Delta _{xy}\) s, we can solve for \(\mu _{rx}\)’s. Combining the properties of the Farris transform with Propositions 1 and 2 leads to Theorem 3. The Proof of Theorem 3 can also be found in Sect. 4.
3.2 Algorithm 1: the reduction step
We are now ready to describe the reduction algorithm (Algorithm 1). Recall that we are restricting our attention to the case of three leaves \(\mathcal {X} = \left\{ 1,2,3 \right\} \) with rooted species tree topology 12|3. The main idea underlying the reduction algorithm is based on the proof of the identifiability result (Theorem 3). That is, we find a set of genes whose topology is highly likely to be a fixed triplet, we estimate the height differences on this set using (8), and we perform a type of Farris transform. However, given that we do not have access to the actual gene tree distribution but only to sequence data, there are several differences with the identifiability proof that make the analysis and the algorithm more involved. We explain these next in details.
A first challenge is that, in the regime where sequence length is “short,” i.e., when \(k \ll f^{-2}\), the sequence-based estimate of the gene tree metric is much less accurate than what is needed for our reduction step.
Key idea 2: We show how to combine genes satisfying a condition related to (7) to produce a much more accurate estimate of distance differences.
Fixing gene tree topologies Here we only have access to sequence data. In particular the \(\delta \)s are unknown. So, we work instead with the p-distances for gene i and \(x,y \in \mathcal {X}\), and their empirical quantiles \(\widehat{p}^{(\beta )}_{xy}\).Footnote 2 Similarly to Proposition 1, we then consider those genes for which the event
holds for some chosen permutation (x, y, z) of (1, 2, 3). We will call this set of genes I. We show that this set has a “non-trivial” size and that, with high probability, the genes satisfying (9) have topology xy|z (see Proposition 3).Footnote 3 In particular, letting \(p(x) = \frac{3}{4}\left( 1 - e^{-4x/3} \right) \), the analysis of this construction accounts for the “sequence noise” around the expected values
Estimating distance differences Because we work with p-distances, we adapt formula (8) for the difference \(\Delta _{xy}\) as follows. Using \( \widehat{p}^I_{xz} = \frac{1}{\left| I \right| }\sum _{i\in I}\widehat{p}_{xz}^i\) and \(\widehat{p}^I_{yz} = \frac{1}{\left| I \right| }\sum _{i\in I}\widehat{p}_{yz}^i, \) our estimate of the distance differences is given by
Recall from Proposition 2 and Fig. 2 that, for this formula to be valid, we need to ensure that the topology of the gene trees used is xy|z. The logarithmic transforms in the curly brackets are the usual distance corrections in the Jukes-Cantor sequence model (see e.g. Semple and Steel (2003)). Note, however, that we perform an average over I before the correction; this is important to obtain the needed statistical power of our estimator. The non-trivial part of the analysis of this step is to bound the estimation error. Indeed, unlike the identifiability result, we have a finite amount of gene data and, moreover, we must account for the sequence noise. This is done using concentration inequalities in Proposition 4.
In Algorithm 1, we compute \(\widehat{\Delta }_{xy}\) with the formula above for two of the pairs in \(\mathcal {X}\), say (1, 2) and (1, 3), and then derive the third quantity consistently, i.e., \(\widehat{\Delta }_{23} = \widehat{\Delta }_{13} - \widehat{\Delta }_{12}\). We also set \(\widehat{\Delta }_{xy} = - \widehat{\Delta }_{yx}\) and \(\widehat{\Delta }_{xx} = 0\) for all x, y.
Stochastic Farris transform The quantile test (see Section 1) is not a distance-based method in the traditional sense of the term. Indeed we do not define a pairwise distance matrix on the leaves. Instead, we use the empirical distribution of the p-distances across genes. It is for this reason that we do not simply apply the classical Farris transform of (6) to the estimated distances. Rather, we perform what we call a “stochastic” Farris transform to ensure that we properly mimic the contributions from both the multispecies coalescent and the Jukes-Cantor model to the distribution of p-distances.
Key idea 3: We transform the sequence data itself to mimic the distribution under an ultrametric species phylogeny. This is done by adding the right amount of noise to the sequence data at each gene, as detailed next.
We will let \(\oplus \) denote addition mod-4 and identify \(\mathtt{A,T,G,C}\) with 0, 1, 2, 3 respectively in that order when doing this addition. For instance, this means that \(\mathtt{A} \oplus 1 = \mathtt{T}\) and \(\mathtt{G} \oplus 2 = \mathtt{A}\).
Definition 5
(Stochastic Farris transform). For a gene i, let \(\{\xi _x^{i}\}_{x\in \mathcal {X}}\) be a sequence dataset over the species \(\mathcal {X} = [3]\) and let \(\Delta _{xy} = \mu _{rx} - \mu _{ry}, x, y \in \mathcal {X}\). Assume without loss of generality that \(\min \{\Delta _{12},\Delta _{13}\} \ge 0\). The stochastic Farris transform defines a new set of sequences \(\{\xi ^i_{x,N}\}_{x \in \mathcal {X}}\) such that \(\xi _{x,N}^i = \xi _x^i \oplus \epsilon _x^i\), where \(\epsilon _x^i\in \{0,1,2,3\}^k\) is an independent random sequence whose jth coordinate is drawn according to: \(\epsilon _x^{ij} = 0\), w.p. \(1- p(\Delta _{1x})\); otherwise it is chosen uniformly among [3].
We write this as \(\{\xi _{x,N}^i\}_{x\in \mathcal {X}} = \mathcal {F}(\{\xi _{x}^i\}_{x\in \mathcal {X}}; \{\Delta _{xy}\}_{x,y\in \mathcal {X}})\).
By the Markov property, for \(x,y \in \mathcal {X}\), the “noisy” sequence data above satisfy
Notice that \(\delta _{xy}^i\), the random gene tree distance between x and y under gene i, can be decomposed as \(\mu _{xy} + \Gamma _{xy}^i\), where \(\Gamma _{xy}^i\) is the random component contributed by the multispecies coalescent. On the other hand, the set of distances \(\mu _{xy} + \Delta _{1x} + \Delta _{1y}\) is ultrametric by the properties of the classical Farris transform (see the Proof of Theorem 3). As a result, the stochastic Farris transform modifies the sequence data so that it appears as though it was generated from an ultrametric MSC-JC process.
In reality, we do not have access to the true differences \(\Delta _{xy}, x,y\in \mathcal {X}\). Instead, we employ our estimates \(\widehat{\Delta }_{xy}\) for all \(x,y \in \mathcal {X}\) in the previous step to obtain the following approximate stochastic Farris transform:
This is the output of the reduction. See Algorithm 1 for details. We prove Theorem 1 in Sect. 5.
4 Identifiability: detailed proofs
Proof
(Proposition 1) Recall the definition of the event \(\mathscr {E}_{I}\) from (7). Our goal is to show that it has positive probability and that it implies that the rooted topology of G is xy|z. This makes sense on an intuitive level because the event \(\mathscr {E}_{I}\) requires that \(\delta _{xy}\) is “somewhat small” and that \(\delta _{xz}\), \(\delta _{yz}\) are “somewhat large.” To prove this, we make crucial use of the following symmetry. Let w be the most recent common ancestor of x, y and z in S. By “above (respectively, below) w,” we refer to the times prior to (respectively, following) the species divergence at w (forward in time). Let
and let \(\mathscr {B}_{xy}\) denote the event that the coalescence between the lineages of x and y occurs below w (which is only possible if x and y happen to be sister populations in the species phylogeny). For \(\beta \in [0,1]\), let \(\Gamma _{xy}^{(\beta )}\) be the \(\beta \)th quantile of \(\Gamma _{xy}\). We define the quantities and event above similarly for the other pairs. Under \(\mathscr {B}_{xy}^{c}\), note that \(\Gamma _{xy}\) is the contribution to \(\delta _{xy}\) coming from the path above w on G, and the same holds for the other pairs. Hence, by the exchangeability of the coalescent process above w, we have
That observation will facilitate the comparison of quantiles.
We break up the proof into 3 cases depending on the rooted topology of S:
-
Rooted topology of S is yz|x. In that case, the lineages from the pairs (x, y) and (x, z) coalesce only above w. That is, the events \(\mathscr {B}_{xy}\) and \(\mathscr {B}_{xz}\) occur almost surely. By (13), we then get \(\Gamma _{xy} {\mathop {=}\limits ^\mathrm{d}} \Gamma _{xz}\) which implies that \(\Gamma _{xy}^{(1/2)} = \Gamma _{xz}^{(1/2)} =: \gamma \). Because \(\mu _{xy}\) and \(\mu _{xz}\) are deterministic, (12) guarantees further that
$$\begin{aligned} \delta _{xy}^{(1/2)} = \mu _{xy} + \gamma , \qquad \delta _{xz}^{(1/2)} = \mu _{xz} + \gamma . \end{aligned}$$As a result, the event \(\mathscr {E}_{I}\) means that
$$\begin{aligned} \mu _{xy} + \Gamma _{xy} = \delta _{xy} \le \delta _{xy}^{(1/2)} = \mu _{xy} + \gamma , \end{aligned}$$and
$$\begin{aligned} \mu _{xz} + \Gamma _{xz} = \delta _{xz} > \delta _{xz}^{(1/2)} = \mu _{xz} + \gamma . \end{aligned}$$The last two inequalities are equivalent to \(\Gamma _{xy} \le \gamma < \Gamma _{xz}\) which is only possible if the rooted topology of G is xy|z. It remains to show that the event \(\mathscr {E}_{I}\) occurs with positive probability. What we have shown implies also that \(\Gamma _{yz} = \Gamma _{xz}\) almost surely. Moreover, conditioned on \(\mathscr {B}_{yz}\), the random variable \(\Gamma _{yz}\) is non-positive; hence, \(\Gamma _{yz}^{(1/2)} \le \gamma < \Gamma _{xz} = \Gamma _{yz}\). So \(\mathscr {E}_{I}\) is indeed possible and occurs with positive probability under the MSC.
-
Rooted topology of S is xz | y . This is case is similar to the previous one, and is therefore omitted.
-
Rooted topology of S is xy | z . In that case, the lineages from the pairs (x, z) and (y, z) coalesce only above w. On the other hand, conditioned on \(\mathscr {B}_{xy}\), the random variable \(\Gamma _{xy}\) is non-positive. Hence, combining these observations with (13), we get \(\Gamma _{xy}^{(1/2)} \le \Gamma _{xz}^{(1/2)} = \Gamma _{yz}^{(1/2)} =: \gamma \). The event \(\mathscr {E}_{I}\) then means that
$$\begin{aligned} \mu _{xy} + \Gamma _{xy} = \delta _{xy} \le \delta _{xy}^{(1/2)} = \mu _{xy} + \Gamma _{xy}^{(1/2)} \le \mu _{xy} + \gamma , \end{aligned}$$and
$$\begin{aligned} \mu _{xz} + \Gamma _{xz} = \delta _{xz} > \delta _{xz}^{(1/2)} = \mu _{xz} + \gamma , \end{aligned}$$and again \(\Gamma _{xy} \le \gamma < \Gamma _{xz}\), implying that the rooted topology of G is xy|z. The equality \(\Gamma _{xz} = \Gamma _{yz}\) holds as well almost surely. So \(\mathscr {E}_{I}\) has positive probability.
That proves the claim. \(\square \)
‘
Proof
(Proposition 2) We know from Proposition 1 that, conditioned on \(\mathscr {E}_{I}\), coalescence between the lineages of x and z necessarily occurs in the common ancestral population of x, y and z, irrespective of the species tree topology. The same holds for y and z. Further \(\Gamma _{xz} = \Gamma _{yz}\) almost surely, where the \(\Gamma \)s were defined in the proof of Proposition 1. Using (12) it follows that, conditioned on \(\mathscr {E}_{I}\), we have \( \delta _{xz} - \delta _{yz} = \mu _{xz} - \mu _{yz} = \Delta _{xy}\) with probability 1 as claimed.\(\square \)
Proof
(Theorem 3) By Proposition 2, conditioning on the event \(\mathscr {E}_{I}\)—which depends only on the gene metric—we have \( \delta _{xz} - \delta _{yz} = \mu _{xz} - \mu _{yz} = \Delta _{xy}\). Hence, from this information, we can construct
assuming that \(\min \{\Delta _{12}, \Delta _{13}\}\ge 0\) (the other cases follow similarly). Recalling that \(\Delta _{xy} := \mu _{rx} - \mu _{ry}\), where r is the root of S, it follows that \(\dot{\mu }_{r1} = \dot{\mu }_{r2} = \dot{\mu }_{r3}\). That is, \(\dot{\mu }\) is ultrametric. From this, the rooted topology of the species tree can be reconstructed. See Fig. 2 for an illustration and, for instance (Semple and Steel 2003, Lemma 7.2.2) for more details on this last step. \(\square \)
5 Ultrametric reduction: detailed proofs
Before proving Theorem 1, we define \(S'\) formally. Given estimates \(\widehat{\Delta }_{xy}\) for all \(x,y \in \mathcal {X}\), and supposing that \(\min \{\widehat{\Delta }_{12}, \widehat{\Delta }_{13}\}\ge 0\) or equivalently that the quantity \(-\frac{3}{4}\log \left( 1 - \frac{4}{3}\widehat{p}_{xy}^I \right) \) is minimized for the pair \((x,y) = (2,3)\) (the other cases follow similarly), we let
Note that this formula is consistent with the definition of \(\dot{\mu }_{xy}\) in (6). Observe further that we do not have access to \(\widehat{\dot{\mu }}_{xy}\), as \(\mu _{xy}\) is unknown—it is only used to define \(S'\) in the statement of the theorem.
Let \(S' = (V_s, E_s, r, \vec {\tau },\vec {\hat{\dot{\mu }}})\) be a species phylogeny with the same topology and branch lengths as S restricted to \(\mathcal {X}\), and with mutation rates \(\{\widehat{\dot{\mu }}\}_{e\in E_s}\) that are chosen such that:
-
(a)
If \(e\in E_s\) is an internal branch, we let \(\widehat{\dot{\mu }}_e = \mu _e\) ; and
-
(b)
Otherwise, that is, if \(e\in E_s\) is incident to a leaf x of S, we let \(\widehat{\dot{\mu }}_e\) be chosen to satisfy
$$\begin{aligned} \widehat{\dot{\mu }}_e \tau _e = \mu _e \tau _e + \widehat{\Delta }_{1x}. \end{aligned}$$
The goal here is to “stretch” the leaf edges so that the modified species metric between any pair of leaves \(x, y \in \mathcal {X}\) is given by \(\widehat{\dot{\mu }}_{xy}\). Alternatively, we could have modified the branch lengths.
5.1 Proof of Theorem 1
Proof
(Theorem 1) Define two disjoint subsets \(\mathcal {M}_{\mathrm{R}1}, \mathcal {M}_{\mathrm{R}2}\) of [m] satisfying:
for a constant \(c_{2}\) to be determined below, and let \(\mathcal {M}_{\mathrm{R}}\) and \(\mathcal {M}_{\mathrm{Q}}\) be such that \(\mathcal {M}_{\mathrm{R}} = \mathcal {M}_{\mathrm{R}1}\sqcup \mathcal {M}_{\mathrm{R}2}\) and \([m] = \mathcal {M}_{\mathrm{R}} \sqcup \mathcal {M}_{\mathrm{Q}}\). For a gene i and leaves \(x,y \in \mathcal {X}\), recall that In fact, we split this last average into two to avoid unwanted correlations as we explain below. Assuming k is even for simplicity, we denote these as

For \(\beta \in [0,1]\), let \(\widehat{p}^{(\beta )}_{xy}\) be the corresponding empirical quantiles computed based on the set \(\{\widehat{{p}}_{xy}^i: i\in \mathcal {M}_{\mathrm{R}1}\}\). Fix a permutation (x, y, z) of (1, 2, 3). Consider the following subset of genes in \(\mathcal {M}_{\mathrm{R}2}\):
The proof of the theorem closely tracks that of the identifiability result (Theorem 3). We first show that the rooted topologies in I are highly likely to be xy|z and prove some technical claims that will be useful in the proof of Proposition 4 (proof in Sect. 5.2).\(\square \)
Proposition 3
(Fixing gene tree topologies) There are \(c_{9}, c_{10}, c_{10}', \varepsilon _0 > 0\) such that with probability at least
the following hold:
-
(a)
The rooted topology of all gene trees in I is xy|z,
-
(b)
For all \(i\in I\), \(p^i_{xy} \le p^{(7/24)}_{xy}\), \(p^{(17/24)}_{xz} \le p^i_{xz} \le p^{(19/24)}_{xz}\), \(p^{(17/24)}_{yz} \le p^i_{yz} \le p^{(19/24)}_{yz}\),
-
(c)
The size of I is greater than \(c_{10}' |\mathcal {M}_{\mathrm{R}2}|\).
Let \(\mathscr {I}\) be the event that the conclusion of Proposition 3 holds. To simplify the notation, we use \(\widetilde{\mathbb {P}}\) and \(\widetilde{\mathbb {E}}\) to denote the probability and expectation operators conditioned on \(\mathscr {I}\). Using
let
Recall that \( \Delta _{xy} = \mu _{rx} - \mu _{ry}. \) We next show that \(\widehat{\Delta }_{xy}\) is a good approximation to \(\Delta _{xy}\) (proof in Sect. 5.2).
Proposition 4
(Estimating differences). There is a \(c_{11} \in (0,1)\) such that with \(\widetilde{\mathbb {P}}\)-probability at least \( 1 - 4\exp \left( - c_{11} k|\mathcal {M}_{\mathrm{R}2}| \phi ^2\right) , \) it holds that \( \left| \widehat{\Delta }_{xy} - \Delta _{xy} \right| \le \phi /2. \)
We repeat the height difference estimation above for all pairs in \(\mathcal {X}\). Therefore, by a union bound, we get the above guarantee for all pairs with probability at least \(1 - 12\exp \left( - c_{11} k|\mathcal {M}_{\mathrm{R}2}| \phi ^2\right) \).
Without loss of generality, assume that \( \mu _{r1} \ge \max \{\mu _{r2}, \mu _{r3}\}, \) and recall the Farris transform
which defines an ultrametric, and consider the approximation
Assuming that the conclusion of Proposition 4 holds for all \(x,y \in \mathcal {X}\), we have shown that \((\widehat{\dot{\mu }}_{xy})\) is \(\phi \)-close to the ultrametric \((\dot{\mu }_{xy})\). As we explained in Sect. 3.2, we produce a new sequence dataset using an approximate stochastic Farris transform \( \{\xi _{x,N}^i\}_{x\in \mathcal {X}} = \mathcal {F}(\{\xi _x^{i}\}_{x\in \mathcal {X}};\{\widehat{\Delta }_{xy}\}_{x,y\in \mathcal {X}}). \)
Hence, again by a union bound, we get the claim of Theorem 1 except with probability
We get the data requirement by asking for the conditions under which the above quantity is less than \(\varepsilon \), which involves choosing \(c_{2}, c_{}', c_{}''\) large enough in (14) and in the statement of the theorem. \(\square \)
5.2 Proofs of Propositions 3 and 4
Proof
(Proposition 3) First fix \(0< \varepsilon _1 < 1/24\) and let \(\varepsilon _0 > 0\) be the largest value such that
The fact that these inequalities hold is guaranteed by (27) in the Appendix which characterizes the behavior of the quantile functions of the random variables associated with the MSC. Note that \(\varepsilon _0\) depends on the parameters \(\mu _U\), \(g'\) and g.
Let (x, y, z) be an arbitrary permutation of the leaves (1, 2, 3). The idea of the proof is to rely on Proposition 1, which we rephrase in terms of p-distances. For a gene \(G_i\), let
And, for \(\beta \in [0,1]\), the corresponding \(\beta \)th quantile is given by
similarly for the other pairs. Then, by Proposition 1, the event
implies that the rooted topology of \(G_i\) is xy|z.
Hence, our main goal is to show that
implies \(\mathscr {E}^i_{I}\) with high probability. We do this by controlling the deviations of \(\widehat{p}_{uw}^{(\beta )}\) (Lemma 1 below) and \(\widehat{p}^{i\downarrow }_{uw}\) (Lemma 2 below). (The upper bounds on \(\widehat{p}^{i\downarrow }_{xz}\) and \(\widehat{p}^{i\downarrow }_{yz}\) in (16) are included for technical reasons that will be useful in proving Proposition 4.)
Recall that we use only the genes in \(\mathcal {M}_{\mathrm{R}}\) for the reduction step and this set in turn is divided into disjoint subsets \(\mathcal {M}_{\mathrm{R}1}\) and \(\mathcal {M}_{\mathrm{R}2}\). The quantiles are estimated using \(\mathcal {M}_{\mathrm{R}1}\), while \(\mathcal {M}_{\mathrm{R}2}\) is used to compute I. We do not argue about the deviation of \(\widehat{p}_{uw}^{(\beta )}\) from the true \(\beta \)th quantile of the distribution of \(\widehat{p}_{uw}^i\). Instead we show that \(\widehat{p}_{uw}^{(\beta )}\) is close to the \(\beta \)th quantile \(p_{uw}^{(\beta )}\) of the disagreement probability under the MSC, that is, the quantile without the sequence noise. We argue this way because the events that we are ultimately interested in (whether a certain coalescence event has occured in a particular population) are expressed in terms of the MSC. Note that, in order to obtain a useful bound of this type, we must assume that the sequence length is sufficiently long, that is, that the sequence noise is reasonably small. Hence this is one of the steps of our argument where we require a lower bound on k.
Lemma 1
(Deviation of \(\widehat{p}_{uw}^{(\beta )}\)). Fix a pair \(u,w \in \mathcal {X}\) and a constant \(\beta \in (0,1)\). For all \(\varepsilon _0 > 0\) and \(0< \varepsilon _1 < \min \{\beta ,1-\beta \}\), there is a constant \(c_9 > 0\) depending on \(\beta \), \(\varepsilon _0\) and \(\varepsilon _1\) such that
provided that k is greater than a constant depending on \(\varepsilon _0\) and \(\varepsilon _1\).
Proof
We prove one side of the first equation (the other inequalities being similar). Define the random variable
and observe that
Our goal is hence to bound the probability on the r.h.s.
For this, we first bound the expectation of M by noting that
by Hoeffding’s inequality Boucheron et al. (2013) applied to \(\widehat{p}^{i}_{uw}\) and the definition of \(p^{(\beta + \varepsilon _1)}_{uw}\); we also used that \(\mathbb {E}[{\widehat{p}^{i}_{uw}}\,|\,p_{uw}] = p_{uw}\).
We then apply Hoeffding’s inequality to M itself. From (19), it follows that
Therefore, letting
we have that
where we used (20) on the first line. Observe moreover that \(c_9\) is strictly positive, provided that k is greater than a constant depending on \(\varepsilon _0\) and \(\varepsilon _1\). Combining with (18) gives the result.\(\square \)
We can also control the deviation of \(\widehat{p}^{i\downarrow }_{uw}\) around the random variable \(p^i_{uw}\).
Lemma 2
(Deviation of \(\widehat{p}^{i\downarrow }_{uw}\)). Fix a pair \(u, w \in \mathcal {X}\). For all i and \(\varepsilon _0 > 0\), almost surely
Proof
Conditioned on \(p^i_{uw}\), the random variable \((k/2) \widehat{p}^{i\downarrow }_{uw}\) is \(\mathrm {Bin}(k/2,p^i_{uw})\). The result follows again from Hoeffding.\(\square \)
Let \(\mathscr {E}_{\mathrm {qu}}\) be the event that the inequality in Lemma 1, i.e., \(p_{uw}^{(\beta -\varepsilon _1)} - \varepsilon _0 \le \widehat{p}_{uw}^{(\beta )} \le p_{uw}^{(\beta + \varepsilon _1)} + \varepsilon _0\), holds for \(\widehat{p}^{(1/3)}_{xy}\), \(\widehat{p}^{(2/3)}_{xz}\), \(\widehat{p}^{(5/6)}_{xz}\), \(\widehat{p}^{(2/3)}_{yz}\) and \(\widehat{p}^{(5/6)}_{yz}\), which occurs with probability at least \(1 - 10\exp (-2 c_0^2 |\mathcal {M}_{\mathrm{R}1}|)\) by a union bound over (17). Let \(\mathscr {D}_i\) be the event that the inequality in Lemma 2, i.e., \(|\widehat{p}^{i\downarrow }_{uw} - p^i_{uw}| {\le } \varepsilon _0\), holds for all pairs (u, w) in \(\mathcal {X}\), an event which occurs with probability at least \(1 - 6\exp (-k \varepsilon _0^2)\) by a union bound over (21).
Recall that our goal is to show that \(\mathscr {Q}_i\) (defined in (16)) implies \(\mathscr {E}^i_{I}\) with high probability. We also condition on \(\mathscr {E}_{\mathrm {qu}}\) and \(\mathscr {D}_i\), which occur with high probability. Given \(\mathscr {E}_{\mathrm {qu}}\), \(\mathscr {D}_i\) and \(\mathscr {Q}_i\), we have
and similarly for the other pairs, where the first inequality follows from \(\mathscr {D}_i\), the second inequality follows from \(\mathscr {Q}_i\), the third inequality follows from \(\mathscr {E}_{\mathrm {qu}}\), and the fourth inequality follows from (15). That is, \(\mathscr {E}^i_{I}\) holds. Finally, the probability that all \(i\in I\) satisfy \(\mathscr {E}^i_{I}\) simultaneously is at least
where we bounded the probability that all i in I satisfy \(\mathscr {D}_i\) with the probability that all i in \(\mathcal {M}_{\mathrm{R}2}\) satisfy \(\mathscr {D}_i\).
It remains to bound the size of I.
Lemma 3
(Size of I). There are constants \(c_{10}, c_{10}'>0\) such that
provided k is greater than a constant depending on \(\varepsilon _0\).
Proof
We show that, under \( \mathscr {E}_{\mathrm {qu}}\), the event \(\mathscr {Q}_i\) has constant probability and we apply Hoeffding’s inequality to |I|, which counts the number of i in \(\mathcal {M}_{\mathrm{R}2}\) satisfying \(\mathscr {Q}_i\).
Observe that, by (15), the events \(\mathscr {D}_i\), \(\{p^i_{xy} \le p^{(7/24)}_{xy}\}\), and \(\mathscr {E}_{\mathrm {qu}}\) (applied in that order) imply
A similar argument shows that, under \( \mathscr {E}_{\mathrm {qu}}\), for \(i \in \mathcal {M}_{\mathrm{R}2}\) the event \( \mathscr {D}_i \cap \mathscr {J}_i, \) implies \(\mathscr {Q}_i\), where we define
This leads to the following lower bound on \(\mathbb {P}[\mathscr {Q}_i \,|\, \mathscr {E}_{\mathrm {qu}}]\):
for some constant \(c_{10}'' > 0\), where we used Lemma 2 to bound the conditional probability of \(\mathscr {D}_i\) on the last line (recalling that Lemma 2 itself is conditioned on the \(p^i_{uw}\)’s). On the third line, we used the fact that \(\mathscr {E}_{\mathrm {qu}}\) depends on \(\mathcal {M}_{\mathrm{R}1}\), and is therefore independent of \(p^i_{xy}, p^i_{xz}, p^i_{yz}\) for \(i \in \mathcal {M}_{\mathrm{R}2}\). The existence of the constant \(c_{10}''\) follows from an argument similar to that leading up to (27) in the Appendix. The expression on the last line is a strictly positive constant provided k is greater than a constant depending on \(\varepsilon _0\).
Finally, applying Hoeffding’s inequality to |I|, we get the result.\(\square \)
Combining (22) and (23) concludes the proof. \(\square \)
Proof
(Proposition 4) Fix \(x,y \in \mathcal {X}\) and let z be the unique element in \(\mathcal {X}- \{x,y\}\). The proof idea is based on Proposition 2. Recall that \(\mathscr {I}\) is the event that the conclusion of Proposition 3 holds and that we use \(\widetilde{\mathbb {P}}\) and \(\widetilde{\mathbb {E}}\) to denote the probability and expectation operators conditioned on \(\mathscr {I}\). Let also \(\mathscr {G}_I\) be the event that the gene trees in I are \(\left\{ G_i \right\} _{i\in I}\). Similarly to the proof of Proposition 2 we note that, conditioned on \(\mathscr {I}\), in all genes in I the coalescences between the lineages of x and z happen in the common ancestral population of x, y and z, irrespective of the species tree topology. The same holds for y and z. That implies that, for \(i\in I\), \(\delta ^i_{xz} = \mu _{rx} + \mu _{rz} + \Gamma ^i_{xz}\), and \(\delta ^i_{yz} = \mu _{ry} + \mu _{rz} + \Gamma ^i_{yz}\), where the \(\Gamma ^i\)s are defined as in the proof of Lemma 1.
Hence
and similarly for the pair (y, z). Letting \(\ell (x) = -\frac{3}{4}\log \left( 1- \frac{4}{3}x\right) \), we get
where in the third equality we used that, conditioned on \(\mathscr {I}\), \(\Gamma ^i_{xz} = \Gamma ^i_{yz}\), for all \(i\in I\). Observe that the computation above relies crucially on the conditioning on \(\mathscr {G}_I\).
It remains to bound the deviation of \( \widehat{\Delta }_{xy} = \ell \left( \widehat{p}_{xz}^I\right) - \ell \left( \widehat{p}_{yz}^I\right) \) around \( \Delta _{xy} = \ell \left( \widetilde{\mathbb {E}}\left[ \widehat{p}_{xz}^I \,|\, \mathscr {G}_I \right] \right) - \ell \left( \widetilde{\mathbb {E}}\left[ \widehat{p}_{yz}^I \,|\, \mathscr {G}_I \right] \right) , \) and take expectations with respect to \(\mathscr {G}_I\). We do this by controlling the error on \(\widehat{p}^I_{xz}\) and \(\widehat{p}^I_{yz}\), conditionally on \(\mathscr {G}_I\). Indeed, observe that the function \(\ell \) satisfies the following Lipschitz property: for \(0 \le x \le y \le M < {3/4}\),
Hence, to control \(|\widehat{\Delta }_{xy} - \Delta _{xy}|\), it suffices to bound \(\left| \widehat{p}_{uz}^I - \widetilde{\mathbb {E}}\left[ \widehat{p}_{uz}^I \,|\, \mathscr {G}_I \right] \right| \) and \(\max \left\{ \widehat{p}_{uz}^I, \widetilde{\mathbb {E}}\left[ \widehat{p}_{uz}^I \,|\, \mathscr {G}_I \right] \right\} \) for \(u = x,y\).\(\square \)
Lemma 4
(Conditional expectation of \(\widehat{p}_{uz}^I\)). Fix \(u = x\) or y. There is a constant \(c_{12}' \in (0,{3/4})\) small enough such that almost surely
Proof
Using \(p_{uz}^i = \widetilde{\mathbb {E}}\left[ \widehat{p}_{uz}^i \,|\, \mathscr {G}_I \right] \) for \(i \in I\), by Proposition 3 (b), we have that
for some constant \(c_{12}' \in (0,{3/4})\). This constant depends on the parameters \(\mu _U\), \(g'\) and g. Hence, the result follows from averaging over i.\(\square \)
Lemma 5
(Conditional deviation of \(\widehat{p}_{uz}^I\)). Fix \(u=x\) or y. For all \(\phi ' > 0\), almost surely
Proof
Observe first that, conditioned on \(\mathscr {G}_I\), the \(k \left| I \right| \) sites that are averaged over in the computation of

are independent. Secondly, each random variable in (25) is bounded by 1. Therefore, the result follows from Hoeffding’s inequality.\(\square \)
We set \(\phi ' = c_{12}' \phi / 6\), which is \(< c_{12}'\) since \(\phi \le 1\). Combining (24) and Lemmas 4 and 5, we get that conditioned on \(\mathscr {G}_I\)
where we used again that \(\phi \le 1\), except with \(\widetilde{\mathbb {P}}\)-probability
by setting \(c_{12} = \frac{1}{8} (c_{12}')^2 c_{10}'\). Taking expectations with respect to \(\mathscr {G}_I\) gives the result.
6 Concluding remarks
Our main contribution is a novel transformation of sequence data under the MSC-JC that produces a new dataset mimicking a molecular clock. We use this reduction, which is of independent interest, to extend a previous data requirement trade-off for species tree estimation beyond the molecular clock case. Our second contribution is a delicate robustness analysis of the quantile-based triplet test of Mossel and Roch (2017). This represents a step towards the design of practical reconstruction algorithms that avoid gene tree estimation and achieve tight rigorous data requirement guarantees. Further issues must be addressed to achieve this goal. In particular, we have assumed here that the mutation rates and sequence lengths are the same across genes. Relaxing these assumptions is important. Identifiability issues may arise however (Matsen and Steel 2007; Steel 2009).
Notes
Actually, the quantiles are estimated from part of the gene set (\(\mathcal {M}_{\mathrm{R}1}\)) to avoid unwanted correlations. The rest of the analysis is done on the other part. See Algorithm 1.
The p-distances in (9) are actually estimated over half the gene length to once again avoid unwanted correlations. That is, we use \(\widehat{p}_{xy}^{i\downarrow }\) defined in Algorithm 1 to compute I. We use the other half to estimate the differences below.
References
Allman ES, Degnan JH, Rhodes JA (2011) Identifying the rooted species tree from the distribution of unrooted gene trees under the coalescent. J Math Biol 62(6):833–862
Allman ES, Degnan JH, Rhodes JA (2018) Species tree inference from gene splits by unrooted star methods. IEEE/ACM Trans Comput Biol Bioinf 15(1):337–342
Allman ES, Long C, Rhodes JA (2019) Species tree inference from genomic sequences using the log-det distance. SIAM J Appl Algebra Geom 3(1):107–127 (Publisher: Society for Industrial and Applied Mathematics)
Boucheron S, Lugosi G, Massart P (2013) Concentration inequalities: a nonasymptotic theory of independence. Oxford University Press, Oxford
Bayzid MS, Mirarab S, Boussau B, Warnow T (2015) Weighted statistical binning: enabling statistically consistent genome-scale phylogenetic analyses. PLoS ONE 10(6):e0129183–e0129183 (06)
Bayzid MS, Warnow T (2013) Naive binning improves phylogenomic analyses. Bioinformatics 29(18):2277–2284 (07)
Chifman J, Kubatko L (2014) Quartet inference from SNP data under the coalescent model. Bioinformatics 30(23):3317–3324
Chifman J, Kubatko L (2015) Identifiability of the unrooted species tree topology under the coalescent model with time-reversible substitution processes, site-specific rate variation, and invariable sites. J Theor Biol 374:35–47
DeGiorgio M, Degnan JH (2010) Fast and consistent estimation of species trees using supermatrix rooted triples. Mol Biol Evol 27(3):552–69
DeGiorgio M, Degnan JH (2014) Robustness to divergence time underestimation when inferring species trees from estimated gene trees. Syst Biol 63(1):66
Degnan JH, DeGiorgio M, Bryant D, Rosenberg NA (2009) Properties of consensus methods for inferring species trees from gene trees. Syst Biol 58(1):35–54
Dasarathy G, Nowak R, Roch S (2015) Data requirement for phylogenetic inference from multiple loci: a new distance method. Comput Biol Bioinform IEEE/ACM Trans 12(2):422–432
Degnan JH, Rosenberg NA (2006) Discordance of species trees with their most likely gene trees. PLoS Genetics, 2(5)
Degnan JH, Rosenberg NA (2009) Gene tree discordance, phylogenetic inference and the multispecies coalescent. Trends Ecol Evol 24(6):332–340
Durrett R (1996) Probability: theory and examples, 2nd edn. Duxbury Press, Belmont, CA
Erdos PL, Steel MA, Székely LA, Warnow TJ (1999) A few logs suffice to build (almost) all trees (i). Random Struct Algorithms 14(2):153–184
Kubatko LS, Degnan JH (2007) Inconsistency of phylogenetic estimates from concatenated data under coalescence. Syst Biol 56(1):17–24
Kapli P, Yang Z, Telford MJ (2020) Phylogenetic tree building in the genomic age. Nature Rev Gene
Long C, Kubatko L (2017) Identifiability and reconstructibility of species phylogenies under a modified coalescent. Arxiv publication arXiv:1701.06871
Long C, Kubatko L (2019) Identifiability and reconstructibility of species phylogenies under a modified coalescent. Bull Math Biol 81(2):408–430
Liu L, Yu L, Pearl DK (2010) Maximum tree: a consistent estimator of the species tree. J Math Biol 60(1):95–106
Maddison WP (1997) Gene trees in species trees. Syst Biol 46(3):523–536
Mirarab S, Bayzid Md. S, Boussau B, Warnow T (2014) Statistical binning enables an accurate coalescent-based estimation of the avian tree. Science, 346(6215)
Mirarab S, Bayzid MS, Warnow T (2016) Evaluating summary methods for multilocus species tree estimation in the presence of incomplete lineage sorting. System Biol 65(3):366
Mossel E, Roch S (2010) Incomplete lineage sorting: consistent phylogeny estimation from multiple loci. IEEE/ACM Trans Comput Biol Bioinform 7(1):166–171
Mossel E, Roch S (2017) Distance-based species tree estimation under the coalescent: information-theoretic trade-off between number of loci and sequence length. Ann Appl Probab 27(5):2926–2955
Matsen FA, Steel M (2007) Phylogenetic mixtures on a single tree can mimic a tree of another topology. Syst Biol 56(5):767–775
Nakhleh L (2013) Computational approaches to species phylogeny inference and gene tree reconciliation. Trends Ecol Evol. doi: https://doi.org/10.1016/j.tree.2013.09.004
Rhodes JA (2019) Topological metrizations of trees, and new quartet methods of tree inference. IEEE/ACM Trans Comput Biol Bioinform
Rusinko J, McPartlon M (2017) Species tree estimation using neighbor joining. J Theor Biol 414:5–7
Roch S, Nute M, Warnow T (2019) Long-branch attraction in species tree estimation: inconsistency of partitioned likelihood and topology-based summary methods. System Biol 68(2):281–297
Roch S (2013) An analytical comparison of multilocus methods under the multispecies coalescent: the three-taxon case. In: Biocomputing 2013: proceedings of the pacific symposium, Kohala Coast, Hawaii, USA, January 3-7, 2013, pp 297–306
Roch S (2018) On the variance of internode distance under the multispecies coalescent. In: Comparative genomics - 16th international conference, RECOMB-CG 2018, Magog-Orford, QC, Canada, October 9-12, 2018, Proceedings, pp 196–206
Roos B (2001) Binomial approximation to the poisson binomial distribution: the Krawtchouk expansion. Theory Prob Its Appl 45(2):258–272
Roch S, Steel M (2015) Likelihood-based tree reconstruction on a concatenation of aligned sequence data sets can be statistically inconsistent. Theor Popul Biol 100:56–62
Roch S, Warnow T (2015) On the robustness to gene tree estimation error (or lack thereof) of coalescent-based species tree methods. Syst Biol 64(4):663–676
Rannala B, Yang Z (2003) Bayes estimation of species divergence times and ancestral population sizes using DNA sequences from multiple loci. Genetics 164(4):1645–1656
Scornavacca C, Delsuc F, Galtier N (2020) Phylogenetics in the Genomic Era. No commercial publisher | Authors open access book
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Shekhar S, Roch S, Mirarab S (2017) Species tree estimation using ASTRAL: how many genes are enough? In: RECOMB’17—proceedings of the 21st annual international conference on research in computational molecular biology, pp 393–395
Steel MA, Székely LA (2002) Inverting random functions. II. Explicit bounds for discrete maximum likelihood estimation, with applications. SIAM J. Discrete Math. 15(4):562–575 (electronic)
Semple C, Steel M (2003) Phylogenetics, vol 22. Mathematics and its Applications Series. Oxford University Press, Oxford
Semple C, Steel MA (2003) Phylogenetics, vol 24. Oxford University Press, Oxford
Steel M (2009) A basic limitation on inferring phylogenies by pairwise sequence comparisons. J Theor Biol 256(3):467–472
Steel M (2016) Phylogeny—discrete and random processes in evolution, vol 89 CBMS-NSF Regional Conference Series in Applied Mathematics. Society for Industrial and Applied Mathematics (SIAM), Philadelphia, PA
Warnow T (2017) Computational Phylogenetics: an introduction to designing methods for phylogeny estimation. Cambridge University Press, Cambridge
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
G. Dasarathy: Supported by the NSF grant CCF-2048223 (CAREER) and the NIH grant 1R01GM140468-01. E. Mossel: Supported by Simons-NSF Grant DMS-2031883 and by a Vannevar Bush Faculty Fellowship ONR-N00014-20-1-2826. S. Roch: Supported by NSF Grants DMS-1149312 (CAREER), DMS-1614242, CCF-1740707 (TRIPODS), DMS-1902892 and DMS-2023239 (TRIPODS Phase II), as well as a Simons Fellowship and a Vilas Associates Award.
A Data requirement tradeoff: detailed proofs
A Data requirement tradeoff: detailed proofs
In this section we analyze Algorithm 2, which performs a quantile test on the output of Algorithm 1.

Proof
(Theorem 2) Let \(S_\mathcal {X}\) be the species tree S restricted to \(\mathcal {X} = [3]\) and let \(r'\) denote its root, i.e., the most recent common ancestor of \(\mathcal {X}\).
Define a partition of the set of genes \([m] = \mathcal {M}_{\mathrm{R}1}\sqcup \mathcal {M}_{\mathrm{R}2} \sqcup \mathcal {M}_{\mathrm{Q}1} \sqcup \mathcal {M}_{\mathrm{Q}2}\) such that the following conditions hold:
for a constant \(c_1 > 0\) to be determined later. The reconstruction algorithm on \(\mathcal {X}\) has two steps:
Ultrametric reduction: In this step, we invoke Algorithm 1 with sequence data \(\{\xi _{x}^{ij}: x\in \mathcal {X}, i \in [m], j\in [k]\}\). The algorithm outputs new sequences \(\{\xi _{x,N}^{ij}:x\in \mathcal {X}, i \in \mathcal {M}_{\mathrm{Q}1} \sqcup \mathcal {M}_{\mathrm{Q}2}, j\in [k]\}\), and Theorem 1 guarantees that these new sequences have the same distribution as a multispecies coalescent process on \(S_{\mathcal {X}}'\) with species metric \((\hat{\dot{\mu }}_{xy})\), where \(S_{\mathcal {X}}'\) and S have the same rooted topology, and \((\hat{\dot{\mu }}_{xy})\) and \(({\dot{\mu }}_{xy})\) are \(\mathcal {O}(f /\sqrt{\log k})\)-close with probability at least \( 1- \varepsilon \).
Quantile test: Now, we invoke Algorithm 2 with the sequence data \(\{\xi _{x,N}^{ij}: i \in \mathcal {M}_{\mathrm{Q}1} \sqcup \mathcal {M}_{\mathrm{Q}2}, j\in [k]\}\) output by Step 1. By Propositions 5 and 7 in Sect. 1, it follows that, with probability at least \( 1- \varepsilon \), Algorithm 2 returns the right topology.\(\square \)
1.1 A.1 Quantile test: robustness analysis
Robustness of quantile test Algorithm 1 produces a new sequence dataset \(\left\{ \xi _{x,N}^{ij}: x\in \mathcal {X} \right\} \) that appears close to being distributed according to an ultrametric species phylogeny. The next step is to perform a triplet test of Mossel and Roch (2017), as detailed in Algorithm 2. Roughly speaking, this test is based on comparing an appropriately chosen quantile of the gene metrics. In fact, because we do not have direct access to the latter, we use a sequence-based surrogate, the empirical p-distances  , for each gene \(i \in \mathcal {M}_{\mathrm{Q}}\) in the output of the reduction, whose expectation is a monotone transformation of the corresponding gene metrics. The idea of Algorithm 2 is to use the above p-distances to define a “similarity measure” \(\widehat{s}_{xy}\) between each pair of leaves \(x,y\in \mathcal {X}\) to reveal the underlying species tree topology on \(\mathcal {X}\). It works as follows. The set of genes \(\mathcal {M}_{\mathrm{Q}}\) is divided into two disjoint subsets \(\mathcal {M}_{\mathrm{Q}1}, \mathcal {M}_{\mathrm{Q}2}\) so that \(\left| \mathcal {M}_{\mathrm{Q}1} \right| , \left| \mathcal {M}_{\mathrm{Q}2} \right| \) satisfy the conditions above; this is to avoid unwanted correlations. The set \(\mathcal {M}_{\mathrm{Q}1}\) is used to compute the \(c_3 \alpha \)-quantile \(\widehat{q}^{(c_{3} \alpha )}_{xy}\) of \(\{\widehat{q}^i_{xy}\,:\,i\in \mathcal {M}_{\mathrm{Q}1}\}\), where \(c_3 > 0\) is a constant determined in the proofs and \(\alpha = \max \left\{ \frac{\log m}{m}, \sqrt{\frac{\log k}{k}} \right\} .\) Let \(\widehat{q}_*\) denote the maximum among \(\left\{ \widehat{q}^{(c_{3} \alpha )}_{xy}: x,y\in \mathcal {X} \right\} \). We then use the genes in \(\mathcal {M}_{\mathrm{Q}2}\) to define the similarity measure \(\widehat{s}_{xy} = \frac{1}{\left| \mathcal {M}_{\mathrm{Q}2} \right| }\left| \left\{ i\in \mathcal {M}_{\mathrm{Q}2}: \widehat{q}^i_{xy} \le \widehat{q}_*\right\} \right| .\) Whichever pair \(x, y \in \mathcal {X}\) produces the largest value of \(\widehat{s}_{xy}\) is declared the closest, i.e., the output is xy|z where z is the remaining leaf in \(\mathcal {X}\).
, for each gene \(i \in \mathcal {M}_{\mathrm{Q}}\) in the output of the reduction, whose expectation is a monotone transformation of the corresponding gene metrics. The idea of Algorithm 2 is to use the above p-distances to define a “similarity measure” \(\widehat{s}_{xy}\) between each pair of leaves \(x,y\in \mathcal {X}\) to reveal the underlying species tree topology on \(\mathcal {X}\). It works as follows. The set of genes \(\mathcal {M}_{\mathrm{Q}}\) is divided into two disjoint subsets \(\mathcal {M}_{\mathrm{Q}1}, \mathcal {M}_{\mathrm{Q}2}\) so that \(\left| \mathcal {M}_{\mathrm{Q}1} \right| , \left| \mathcal {M}_{\mathrm{Q}2} \right| \) satisfy the conditions above; this is to avoid unwanted correlations. The set \(\mathcal {M}_{\mathrm{Q}1}\) is used to compute the \(c_3 \alpha \)-quantile \(\widehat{q}^{(c_{3} \alpha )}_{xy}\) of \(\{\widehat{q}^i_{xy}\,:\,i\in \mathcal {M}_{\mathrm{Q}1}\}\), where \(c_3 > 0\) is a constant determined in the proofs and \(\alpha = \max \left\{ \frac{\log m}{m}, \sqrt{\frac{\log k}{k}} \right\} .\) Let \(\widehat{q}_*\) denote the maximum among \(\left\{ \widehat{q}^{(c_{3} \alpha )}_{xy}: x,y\in \mathcal {X} \right\} \). We then use the genes in \(\mathcal {M}_{\mathrm{Q}2}\) to define the similarity measure \(\widehat{s}_{xy} = \frac{1}{\left| \mathcal {M}_{\mathrm{Q}2} \right| }\left| \left\{ i\in \mathcal {M}_{\mathrm{Q}2}: \widehat{q}^i_{xy} \le \widehat{q}_*\right\} \right| .\) Whichever pair \(x, y \in \mathcal {X}\) produces the largest value of \(\widehat{s}_{xy}\) is declared the closest, i.e., the output is xy|z where z is the remaining leaf in \(\mathcal {X}\).
As stated in Theorem 1, the output to the ultrametric reduction is almost—but not perfectly—ultrametric. To account for this extra error, we perform a delicate robustness analysis of the quantile-based triplet test. At a high level, the proof follows Mossel and Roch (2017). After (1) controlling the deviation of the quantiles, we establish that (2) the test works in expectation and then (3) finish off with concentration inequalities. All these steps must be updated to account for the error introduced in the reduction step. Step (2) is particularly involved and requires a delicate analysis of the CDF of a mixture of binomials.
For the rest of the proof, we assume that the rooted topology of \(S_\mathcal {X}\) is 12|3. The other cases are similar.
Control of empirical quantiles In the following proposition, we show that the empirical quantiles are well-behaved, and provide a good estimate of the \(\alpha \)-quantile of the underlying MSC random variables. We define the random variables \(q_{xy}^i\) and \(r_{xy}^i\) associated to a gene tree i:
Also, we need the 0th quantile of these random variables, specifically
We first show that \(\widehat{q}_*= \max \{ \widehat{q}^{(c_{3} \alpha )}_{xy}: x,y\in \mathcal {X} \}\) is close to \(\max \{{r}_{xy}^{(0)} : x,y\in \mathcal {X} \}\).
Proposition 5
(Quantile behaviour). Let \(\alpha = \max \left\{ m^{-1}\log m, k^{-0.5}\sqrt{\log k} \right\} \) and let \(\phi \) be as in Theorem 2. Then, there are \(c_{3}, c_{4}, c_{5} >0\) depending on the parameters \(\mu _U\), \(g'\) and g such that, for each pair of leaves \(x,y \in \mathcal {X}\), the \(c_{3}\alpha \)-quantile satisfies the following with probability at least \(1 - 6\exp \left( -c_{4} \left| \mathcal {M}_{\mathrm{Q}1} \right| \alpha \right) \), provided Theorem 1 holds,
Proof
Proposition 4 guarantees that \(\widehat{\Delta }_{xy}\) and \(\Delta _{xy}\) are close. Therefore, using the fact that \(p(\cdot )\) is a Lipschitz function, we know that there exists a constant \(c_{5}'>0\) such that \(\left| r_{xy}^{(0)} - q_{xy}^{(0)} \right| \le c_{5}' \phi .\) The second containment in the statement of the lemma follows from this after adjusting the constant. The first part is proved as in (Mossel and Roch 2017, Claim 12), except for a small change: we use Bernstein’s inequality (see e.g. Boucheron et al. 2013) in place of Chebyshev’s inequality to obtain an exponential dependence on \(\left| \mathcal {M}_{\mathrm{Q}1} \right| \) (ultimately resulting in a logarithmic dependence in \(\varepsilon \) in the condition on \(\left| \mathcal {M}_{\mathrm{Q}1} \right| \) in (26)).\(\square \)
Expected version of quantile test We will use \(\bar{\mathbb {P}}\), \(\bar{\mathbb {E}}\), and \(\overline{\mathrm {Var}}\) to denote probabilities, expectations and variances conditioned on the event that Theorem 1 and Proposition 5 hold. We use the genes in \(\mathcal {M}_{\mathrm{Q}2}\), which are not affected by the conditioning under \(\bar{\mathbb {P}}\), to define a similarity measure among pairs of leaves in \(\mathcal {X}\), \(\widehat{s}_{xy}\), as above. We next show that this similarity measure has the right behavior in expectation. That is, defining \(s_{xy} := \bar{\mathbb {E}}\left[ \widehat{s}_{xy} \right] \), we show that \(s_{12} > \max \{s_{13}, s_{23}\}\) (proof in Sect. 3).
Proposition 6
(Expected version of quantile test). For any \(C_2 >0\), there exist constants \(c_{6}, c_{7}>0\) such that \(s_{12} - \max \{s_{13}, s_{23}\} \ge c_{6} p(3f/4) > 0\) provided
Sample version of quantile test We finish the proof by showing that the empirical similarity measures are consistent with the underlying species tree with high probability (proof in Sect. 4).
Proposition 7
(Sample version of quantile test). There is a \(c_{8}>0\) such that, provided Proposition 6 holds, the \(\bar{\mathbb {P}}\)-probability that Algorithm 2 fails is less than \(4 \exp \left( - \frac{\left| \mathcal {M}_{\mathrm{Q}2} \right| p(3f/4)^2}{c_{8} \left( p(3f/4) + \alpha \right) } \right) .\)
1.2 A.2 Auxiliary results
We will need a few technical auxiliary claims.
Quantiles We will need an observation about the quantiles of gene tree pairwise distances. For a pair of leaves \(a,b \in L\), \(\delta _{ab}\) is the branch length induced by the random gene tree under the MSC. Notice that, by definition, \(\delta _{ab}\ge \mu _{ab}\) . We will let \(f_{ab}\left( \cdot \right) \) and \(F_{ab}\left( \cdot \right) \) denote respectively the density and the cumulative density function of the random variable \(Z_{ab}\triangleq \frac{\delta _{ab}-\mu _{ab}}{2}\). Because of the memoryless property of the exponential, it is natural to think of the distribution of \(Z_{ab}\) as a mixture of distributions whose supports are disjoint, corresponding to the different branches that the lineages go through before coalescing. We state this more generally as follows. Suppose that \(U_1,U_2,\ldots , U_r\) are subsets of \(\mathbb {R}\) such that they satisfy \(\sup (U_i)\le \inf (U_{i+1})\), \(i =1,2,\ldots , r\). Suppose that f is a probability density function such that \(f(x) = \sum _{i=1}^r \omega _i f_i(x)\), where \(\omega _1,\omega _2,\ldots ,\omega _r\in (0,1)\) are such that \(\sum _{i=1}^r \omega _i = 1\), and the density \(f_i\) is supported on \(U_i\) for \(i=1,\ldots ,r\). Then, the quantile function of f is given as follows

where, we set \(\omega _0 = 0\). Specializing to \(f_{ab}\) under the MSC, it follows that there exists a finite sequence of constants \(\mu _1,\ldots ,\mu _r \in [\mu _L,\mu _U]\) and \(h_0,\ldots ,h_{r-1} \in [f',g'+ng]\) such that
Cancellations lead to \(f_{ab}(x) = \sum _{i=1}^r e^{-\sum _{j=1}^{i-1}\mu _{j}^{-1}(h_j - h_{j-1})}\mu _i^{-1} e^{-\mu _i^{-1} (x - h_{i-1})}\). This formula implies that the density is bounded between positive constants. We will need the following implication. For any \(\alpha \in [0,1)\), we let \(\delta _{ab}^{(\alpha )}\) and \(p_{ab}^{(\alpha )}\) denote the \(\alpha \)-quantile of the \(\delta _{ab}\) and \(p_{ab}\) respectively. Since by definition \(p_{ab} = p(\delta _{ab})\), we have that \(p^{(\alpha )}_{ab} = p(\delta _{ab}^{(\alpha )})\), where \(p(x) = \frac{3}{4}\left( 1 - e^{-4x/3} \right) \). Then, for any \(0< \beta '< \beta < 1\), there are constants \(0< c'< c'' <+\infty \) (depending on \(\mu _L, \mu _U, g, g', n, \beta ', \beta \)) such that for any \(\xi \in (0,1-\beta ')\), we have
CDF lemma Finally, we restate a key bound about the cumulative distribution function from Mossel and Roch (2017).
Lemma 6
(CDF behavior; see Claim 6 in Mossel and Roch (2017)). There exists a constant \(c_{3}'>0\) depending on the parameters \(\mu _U\), \(g'\) and g such that \(\mathbb {P}\left[ \widehat{q}_{xy} \le q^{(0)}_{xy} \right] \le \frac{c_{3}'}{\sqrt{k}}\le c_{3}' \alpha .\)
1.3 A.3 Proof of Proposition 6
Proof
(Proposition 6) In this proof, we are concerned with behavior in expectation. Hence, we fix an \(i\in \mathcal {M}_{\mathrm{Q}2}\) and drop the i in \(\widehat{q}^i_{xy}\), etc. Let \(\mathcal {E}_{12|3}\) be the event that there is a coalescence in the internal branch of the species phylogeny and observe that \(s_{12}\) can be decomposed as follows
In the proof in Mossel and Roch (2017), instead of \(\widehat{q}_{12}\) one deals with \(\widehat{r}_{12}\), and it follows from the symmetries of the MSC that \(\bar{\mathbb {P}}\left[ \widehat{r}_{12} \le \widehat{q}_*| \mathcal {E}_{12|3}^c\right] = \bar{\mathbb {P}}\left[ \widehat{r}_{13} \le \widehat{q}_*\right] \). This turns out to suffice to establish the expected version of the quantile test in Mossel and Roch (2017). In our setting, we must control quantitatively the difference between these two probabilities due to the slack added by the reduction step. The following lemma is proved below.\(\square \)
Lemma 7
(Closeness to symmetry). For \(\widehat{q}_*\) as defined in Algorithm 2 and any constant \(C_2>0\), there exist constants \(c_{7}', c_{7}'' >0\) such that
provided
Using the above lemma in (28), we can now bound \(s_{12}\) from below as follows
This implies that
The expected version of the quantile test succeeds provided the latter quantity is bounded from below by 0. We establish something a bit stronger, which will be useful in the analysis of the sample version of the quantile test. The following lemma is proved below.
Lemma 8
(Tails) There are \(c_{6}', c_{6}'' > 0\) such that \(\bar{\mathbb {P}}\left[ \widehat{q}_{12} \le \widehat{q}_*| \mathcal {E}_{12|3}\right] \ge c_{6}'\) and \(s_{13} = \bar{\mathbb {P}}\left[ \widehat{q}_{13} \le \widehat{q}_*\right] \le c_{6}'' \alpha \).
The first inequality captures the intuition that, conditioned on coalescence in the internal branch of the species phylogeny, the probability of \(\widehat{q}_{12}\) being small is high. The second inequality captures the intuition that since \(\widehat{q}_*\) behaves roughly like \(q_{13}^{(\alpha )} = p(\delta _{13}^{(\alpha )} + \widehat{\Delta }_{13})\), the event that \(\widehat{q}_{13}\le \widehat{q}_*\) is dominated by the event that the underlying MSC random variable satisfies the same inequality, with the deviations of the JC contribution on top of this being of order \(k^{-0.5}\).
Notice that, if we use Lemma 8 in (29), there is a constant \(c_{6}>0\) such that \(s_{12} - s_{13} \ge c_{6}\, \bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] \) provided \(\phi _2 \le c_{6} p(3f/4)\) for a large enough \(c_6 > 0\), where we used that \(\bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] \) is lower bounded by p(3f/4). This, along with a similar argument for \(s_{23}\), concludes the proof of Proposition 6.
Proof
(Lemma 7) First observe that \(\bar{\mathbb {P}}\left[ \widehat{r}_{13} \le \widehat{q}^*| \mathcal {E}_{12|3}^c \right] = \bar{\mathbb {P}}\left[ \widehat{r}_{13} \le \widehat{q}^*\right] \) since \(\widehat{r}_{13}\) is unaffected by whether coalescence occurs in the internal branch of the species phylogeny. We prove Lemma 7 by arguing that \(\bar{\mathbb {P}}\left[ \widehat{q}_{12} \le \widehat{q}^*| \mathcal {E}_{12|3}^c \right] \) is close to \(\bar{\mathbb {P}}\left[ \widehat{r}_{13} \le \widehat{q}^*| \mathcal {E}_{12|3}^c \right] \), and that \(\bar{\mathbb {P}}\left[ \widehat{q}_{13} \le \widehat{q}^*\right] \) is close to \(\bar{\mathbb {P}}\left[ \widehat{r}_{13} \le \widehat{q}^*\right] \).
We do this by analyzing the effect on the cumulative distribution function (CDF) of a perturbation of the mean. In our case, the distribution of interest is a mixture of binomials. Indeed notice that in the latter case, conditioned on the value of \(\delta _{13}\) and \(q_{13} = p (\delta _{13} + \widehat{\Delta }_{13})\), we are seeking to compare two binomial random variables \(k \,\widehat{q}_{13} \sim \mathrm{Bin}(k, {q_{xy}})\) and \(k \,\widehat{r}_{13} \sim \mathrm{Bin}(k,{q_{xy}} + \beta )\), for some small \(\beta \) (and similarly for the other inequality). The desired result will be implied by the following bound proved at the end of this subsection.\(\square \)
Lemma 9
(Mixture of binomials: CDF perturbation). Let \(\delta _{xy}\) be the distance between x and y on a random gene tree drawn according to the MSC, and let \(p_{xy} = p(\delta _{xy})\) denote the corresponding expected p-distance. Suppose that we have two binomial random variables \(J_1\sim \mathrm{Bin}(k, p_{xy})\) and \(J_2\sim \mathrm{Bin}(k,p_{xy} + \beta )\), for some fixed \(\beta \in (0, 1-p_{xy})\) with \(\beta = \Theta (\phi )\), and that we are given constants \(c_{14},\gamma >0\) such that \(\gamma < p_{xy}^{(c_{14} [\alpha \vee \phi ])}\). Then, for any constant \(C_2>0\) there exist constants \(c_{13},c_{13}' >0\) such that \(\left| \bar{\mathbb {P}}\left[ J_1 \le k \gamma \right] - \bar{\mathbb {P}}\left[ J_2 \le k \gamma \right] \right| \le c_{13}' \beta \sqrt{\log k}\) holds provided
Observe that, although \(J_1\) and \(J_2\) above do not depend on m, the quantity \(\gamma \)—through \(\alpha \)—does.
Notice that this result implies that there exists a constant \(c_{7}'>0\) such that
To see why this is true, first observe that \(\widehat{q}^*\le q^{(0)}_{13} + c_{5} \alpha \le q_{13}^{(c_{5}' \alpha )}\), which follows from Proposition 5 and (27). The bound on \(\widehat{q}^*\) also holds in terms of r-quantiles up to an additive term \(O(\phi )\) per Proposition 5. So we can take \(\gamma = \widehat{q}^*\) in Lemma 9. Using this, and taking \(\beta \) to be \(\Theta (\phi )\), we get the above two inequalities. Note that, for both inequalities, we apply Lemma 9 to \(\widehat{r}_{13}\), \(\widehat{q}_{13}\) or \(\widehat{q}_{12}\) so as to keep \(\beta \) positive as required.
Proof
(Lemma 8) We start with the second inequality in the statement of the lemma. Notice that, from Proposition 5 (on which \(\bar{\mathbb {P}}\) is conditioning) and (27), we know that \(\widehat{q}^*\le q^{(0)}_{13} + c_{5} \alpha \le q_{13}^{(c_{5}' \alpha )}\).
Hence,
The conditional probability on the last line is bounded above by \(c_3' \alpha \) by Lemma 6 and the memoryless property of the exponential. This implies the second inequality of the lemma.
To derive the first one, we make a few observations:
-
(1)
from Proposition 5, \(\widehat{q}_*\ge \widehat{q}_{13}^{(c_{3} \alpha )}\ge {r}_{13}^{(0)}-c_{5}\phi \);
-
(2)
by definition \(q_{12} = p(\delta _{12} + \widehat{\Delta }_{12})\) and, conditioned on \(\delta _{12}\), \(k \,\widehat{q}_{12}\) is distributed as Bin\((k,q_{12})\);
-
(3)
given that \(q_{12} = p(\delta _{12} + \widehat{\Delta }_{12})\) and \(r_{12} = p(\delta _{12} + \Delta _{12})\) and by Proposition 4, it follows that there is \(c_{16} > 0\) such that the event \(\{r_{12} \le {r}_{13}^{(0)} - c_{16} \phi \}\) implies the event \(\{q_{12} \le {r}_{13}^{(0)}-c_{5}\phi \}\) under \( \bar{\mathbb {P}}\);
-
(4)
the event \(\mathcal {E}_{12|3}\) is equivalent to the condition that \(r_{12} = p(\delta _{12} + \Delta _{12}) \le p(\mu _{13} + \Delta _{13}) = r_{13}^{(0)}\).
We use these facts to bound the desired quantity \(\bar{\mathbb {P}}\left[ \widehat{q}_{12} \le \widehat{q}_*| \mathcal {E}_{12|3}\right] \) from below by
where (a) follows from Observation 1 above, (b) follows after conditioning on the event that \(q_{12} \le {r}_{13}^{(0)}-c_{5}\phi \), and (c) follows from Observations 3 and 4. Finally, the last line is greater than some constant \(C'_j > 0\) from the Berry-Esséen theorem (see e.g. Durrett 1996), which gives a constant lower bound on the first term, and (27) together with the assumption that \(\phi \ll p(3f/4)\) and the fact that the probability of \(\mathcal {E}_{12|3}\) is at least p(3f/4), which gives a constant lower bound on the second term.\(\square \)
Proof
(Lemma 9) We need an auxiliary lemma which characterizes the difference between two binomial distributions in terms of the difference of the underlying probabilities. This follows from Roos (2001).\(\square \)
Lemma 10
(Binomial: CDF perturbation). For \(J_1\) and \(J_2\) as above and any \(\gamma \in (0,1)\), we have
Proof
It holds that, if \(\theta (p_{xy}+\beta ) = \frac{\beta ^2 (k+2)}{2 (p_{xy}+\beta ) (1 - p_{xy}-\beta )} < 1\),
where the above inequality comes from (Roos 2001, (15)), by setting \(s = 0\) there, and choosing the Poisson-Binomial distribution to simply be the binomial distribution Bin\((k,p_{xy})\). If \( \beta \le \sqrt{\frac{(p_{xy}+\beta ) (1 -p_{xy}-\beta )}{2 (k+2)}}\), then \(1 - \sqrt{\theta (p_{xy}+\beta )} \ge 0.5\). In this case, we have (30). On the other hand, if \(\beta > \sqrt{\frac{(p_{xy}+\beta ) (1 - p_{xy}-\beta )}{2 (k+2)}}\), since the difference between two probabilities is upper bounded by 2, the upper bound (30) holds trivially.\(\square \)
We cannot directly apply Lemma 10 to prove Lemma 9 since the \(\sqrt{k+2}\) factor in the upper bound is too loose for our purposes. Instead, we employ a more careful argument that splits the domain of \(p_{xy}\).
\({p_{xy}\in I_1 = [ p_{xy}^{(0)}, p_{xy}^{(2c_{14} \sqrt{\frac{\log k}{k}})}]}\) (low substitution regime for small k): In this case, we use the fact that Lemma 10 guarantees that the binomial distributions are \(\mathcal {O}(\beta \sqrt{k})\) apart. That is, there exists a constant \(c_{15}' > 0\) such that
where the last step follows from the quantile definition, after appropriately increasing the constant \(c_{15}'\).
\(p_{xy} \in I_2 = [p_{xy}^{(2c_{14} \sqrt{\frac{\log k}{k}})}, p_{xy}^{({2c_{14} [\alpha \vee \phi ]})}]\) (low substitution regime for large k):
In the case this interval is not empty, there exists a constant \(c_{15}'' >0\) such that
where (a) follows from (27), and (b) follows from the definition of \(\alpha \).
\({p_{xy}\in I_3= [ p_{xy}^{(2c_{14} {[\alpha \vee \phi ]})},0.5]}\) (high substitution regime): In this case observe that, since \(\gamma < p_{xy}^{(c_{14} {[\alpha \vee \phi ]})}\) (i.e., we are looking at a left tail below the mean), we can apply Chernoff’s bound (see e.g. Boucheron et al. 2013) on each of the two terms in the difference individually. For any constant \(C_2 > 0\), we can choose \(c_{14}>0\) so that the following inequality holds for some \(c_{15}'''>0\)
where we used that \(p_{xy}^{(2c_{14} [\alpha \vee \phi ])} - p_{xy}^{(c_{14} [\alpha \vee \phi ])} = \text{\O}mega (\sqrt{\frac{\log k}{k}})\) by (27) and the definition of \(\alpha \).
Putting the three regimes together, there is a constant \(c_{7}'>0\) (not depending on f, m, k) such that
There is a \(c_{13}\) such that the conditions of the lemma imply \(m^{-1}\log m \le \beta \sqrt{\log k}\) and \(k^{-C_2} \le \beta \sqrt{\log k}\).
1.4 A.4 Proof of Proposition 7
Recall that \(\mathcal {E}_{12|3}\) is the event that there is a coalescence in the internal branch of the species phylogeny. First we prove:
Lemma 11
(Variance bound). There is a \(c_{8}'>0\) such that, \(\forall x,y\), \(\overline{\mathrm{Var}}(\widehat{s}_{xy}) \le \frac{c_{8}'}{\left| \mathcal {M}_{\mathrm{Q}2} \right| } \left( \bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] + \phi + \alpha \right) .\)
Proof
Note that the variance of \(\widehat{s}_{12}\) is bounded from above by \(\frac{1}{\left| \mathcal {M}_{\mathrm{Q}2} \right| }\bar{\mathbb {P}}\left[ \widehat{q}_{12}\le \widehat{q}_*\right] \).
Observe further that, by Proposition 5, \(\widehat{q}_*\le r_{13}^{(0)} + c_{5}\phi + c_{5} \alpha \). Moreover, conditioned on the random distance \(\delta _{12}\), \(k\,\widehat{q}_{12}\) is distributed according to Bin\((k, q_{12})\), where \(q_{12} = p\left( \delta _{12} + \widehat{\Delta }_{12} \right) \). Finally, from Lemma 6 and the memoryless property of the exponential, we have that
Therefore, arguing as in the proof of Lemma 8,
From (27), it follows that there is a constant \(c_{8}''>0\) such that
Moreover,
The last inequality follows from the fact that \(\bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] \) is of the order of the length of the internal branch of the species phylogeny; and so is the difference in the middle expression. Notice that this along with (31) and (32) imply that there is a constant \(c_{8}'>0\) (after changing it appropriately) such that \(\bar{\mathbb {P}}\left[ \widehat{q}_{12} \le \widehat{q}_*\right] \le c_{8}'\left( \bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] + \phi + \alpha \right) \).\(\square \)
Proof
(Proposition 7) An error in the quantile test implies that either \(\widehat{s}_{13} > \widehat{s}_{12}\) or \(\widehat{s}_{23} > \widehat{s}_{12}\). Therefore, the probability that the algorithm makes an error is at most
Take the second term on r.h.s. (the other terms being similar). We need two ingredients to invoke Bernstein’s inequality: (1) a lower bound on the “gap” \(\frac{s_{12} - s_{13}}{2}\) (Proposition 6), and (2) an upper bound on the variance of \(\widehat{s}_{12}\) (Lemma 11). We get that \(\bar{\mathbb {P}}\left[ s_{12} - \widehat{s}_{12} \ge \frac{s_{12} - s_{13}}{2} \right] \) is at most
where the last line follows from the fact that \(\bar{\mathbb {P}}\left[ \mathcal {E}_{12|3} \right] \) is bounded from below by p(3f/4) and the assumption that \(\phi \ll p(3f/4)\).\(\square \)
Rights and permissions
About this article
Cite this article
Dasarathy, G., Mossel, E., Nowak, R. et al. A stochastic Farris transform for genetic data under the multispecies coalescent with applications to data requirements. J. Math. Biol. 84, 36 (2022). https://doi.org/10.1007/s00285-022-01731-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00285-022-01731-5