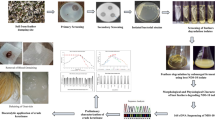
Abstract
Enormous aggregates of keratinous wastes are produced annually by the poultry and leather industries which cause environmental degradation globally. To combat this issue, microbially synthesized extracellular proteases known as keratinase are used widely which is effective in degrading keratin found in hair and feathers. In the present work, keratinolytic bacteria were isolated from poultry farm soil and feather waste, and various cultural conditions were optimized to provide the highest enzyme production for efficient keratin waste degradation. Based on the primary and secondary screening methods, the potent keratinolytic strain (HFS_F2T) with the highest enzyme activity 32.65 ± 0.16 U/mL was genotypically characterized by 16S rRNA sequencing and was confirmed as Bacillus velezensis HFS_F2T ON556508. Through one-variable-at-a-time approach (OVAT), the keratinase production medium was optimized with sucrose (carbon source), beef extract (nitrogen source) pH-7, inoculum size (5%), and incubation at 37 °C). The degree of degradation (%DD) of keratin wastes was evaluated after 35 days of degradation in the optimized keratinase production medium devoid of feather meal under submerged fermentation conditions. Further, the deteriorated keratin wastes were visually examined and the hydrolysed bovine hair with 77.32 ± 0.32% degradation was morphologically analysed through Field Emission Scanning Electron Microscopy (FESEM) to confirm the structural disintegration of the cuticle. Therefore, the current study would be a convincing strategy for reducing the detrimental impact of pollutants from the poultry and leather industries by efficient keratin waste degradation through the production of microbial keratinase.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As the human population expands one of the gravest dangers to the human race is the disposal and proper handling of solid waste. Among the solid wastes, keratin wastes are produced in mammoth quantities from pecuniary poultry processing plants, the leather industry, the wool industry, the textile industry, slaughterhouses, and barber shops [1]. Keratin debris in the form of bird feathers, strands of hair, horns, nails, and hooves, is primarily generated via animal body parts [2]. This waste contains over 90% of keratin that is highly inert, water-insoluble, and non-biodegradable by most proteolytic enzymes such as trypsin, pepsin, and papain [3, 4].
In 2020, India would consume around 3.9 million metric tonnes of chicken meat, with each person consuming 2.6 kg of the meat annually. Poultry is one of the industries that contributes the most to the global economy and is still growing. The annual production of feathers by poultry processing facilities poses a significant challenge for the management of solid waste. One such waste is chicken feathers, which make up 7% of the bird's weight overall. Each bird can have up to 125 g of feathers, and the amount of feather waste produced worldwide each week is thought to be over 3000 tonnes [5]. The dumping of poultry feathers might involve issues with handling, storage, emissions management, and ash disposal. They can also be burned, buried, or utilised as fill for land [6]. These wastes may pose a potential hazard to human health or the environment (soil, air, water) [7]. Additionally, discarded feathers can lead to a number of human illnesses, such as mycoplasmosis, chlorosis, and fowl cholera [6]. Thus, the disposal of keratin waste is quite challenging.
Keratin wastes are biodegraded by keratinophilic bacteria or their enzymes (keratinases), which overcomes the limitations of chemical and thermal treatments [8]. These keratinases with the ability to break down keratins from numerous sources, will play a key role in agricultural and environmental chemistry to tackle the disposal issue [9]. Biological degradation of keratinous waste through microbial keratinases is an effective and feasible mitigation technique that comprises handling of waste, human and ecological protection, and extraction of resources such as amino acids, peptides, and non-protein nitrogenous compounds [10]. They are distinctive enzymes with proteolytic activity that can hydrolyze some insoluble and extremely stable proteins, such as feathers, hairs, and wool, by destroying disulfide bonds that are part of keratin-rich substrates [11, 12]. In addition to increasing the commercial value of keratin wastes, this biotechnological and environmentally beneficial alternative for hydrolysis and recycling also provides comfortable conditions for the production of products that are useful. In terms of cost and environmental considerations, hydrothermal and chemical keratin waste degradation are less efficient than the more recent biological method. This process yields a toxin-free substance called keratin hydrolysate, which has commercial potential. Consequently, keratin wastes can be degraded and used as a useful biomaterial through the innovative process of bioconversion, or biological degradation of keratin wastes, which is both economical and ecologically benign [8].
In the keratin waste degradation process, peptide bond cleavage in compact structures, like keratins, is problematic because the target peptide bonds are insoluble and difficult to approach. The following mechanisms are necessary for the multiple-step procedure that involves keratin enzymatic degradation: (i) The keratinases bind to the macromolecule's surface through hydrophobic and electrostatic contacts, and then (ii) catalysis takes place. Sulfitolysis, or the elimination of disulfide bonds, and proteolysis are the two main steps in the multistage keratin degradation process. Only in the presence of reducing agents, such as disulfide reductases, glutathione, sodium sulphide, dithiothreitol (DTT), mercaptoethanol, cysteine, thioglycolic acid, or cysteine, does sulfitolysis occur. These agents work in tandem with keratinases to break down keratin molecules [13,14,15].
Many microorganisms can degrade such wastes by secreting keratinolytic and proteolytic enzymes [16, 17]. The microbes have been isolated from different environments that are rich in keratin and have been applied to degrade keratin-containing wastes from different resources [18]. The microorganisms include bacteria, actinomycetes, and fungi [19]. Keratinases come from bacteria such as Bacillus licheniformis, B. subtilis and Stenotrophomonas maltophilia. Moreover, the actinobacteria Streptomyces albidoflavus and Streptomyces fradiae also secrete keratinases. Fungal keratinases are mainly from Trichophyton rubrum and Microsporum canis [15].
Bacillus licheniformis is the most effective keratin-degrading bacterium in the genus [20]. Other bacteria including Bacillus, Pseudomonas, Brevibacillus, Stenotrophomonas, Fusarium, Chryseobacterium, Xanthomonas, Geobacillus, Serratia and Nesterenkonia, can produce keratin-degrading enzymes. Among these, only ample numbers of organisms have been exploited for commercial use. The most prevalent and easy-to-handle microorganism among them is Bacillus sp. Similarly, Bacillus pumilus, Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus cereus are also capable of producing keratinase enzyme that has many potential industrial and medical applications [12]. Shih and his colleagues in North Carolina have developed a source of keratinase enzyme named ‘Versazyme’ from Bacillus licheniformis to be the first keratinase enzyme [21]. Hence, Bacillus sp. is always a highly preferred microbial strain for keratin degradation by many researchers.
Feathers have become one source of pollutants because of their recalcitrant nature [22]. Untreated feather waste can sustain many pathogenic microorganisms and emit various pollutants such as nitrous oxide, ammonia, and hydrogen sulfide, which are a threat to the environment and people’s health. Therefore, converting feathers into value-added products using economic methods is of great interest to many researchers [23]. Accumulated studies have shown that feathers can be efficiently degraded by various microorganisms. Microbial conversion of feathers into value-added products such as biofertilizers and animal feeds should be used in poultry industries [16].
The present research work explores the potential use of feather meal to be turned into a useful substrate for keratinase enzyme production using the potent bacterial strain Bacillus velezensis isolated from poultry feather waste. The investigation further extends the possibility of manipulating the degradative efficacy of keratin-rich wastes such as whole feathers, chicken nails, and bovine hair in the optimized keratinase production medium. As a result, the current research would be a persuasive approach for overcoming the environmental effect of poultry and leather sector pollutants through microbial keratinase production during the fermentation process for efficient keratin waste degradation.
Materials and Methods
Sampling and Isolation of Bacteria
A total of three different samples-Poultry feather soil (fresh), Poultry feather soil (old), and Raw Chicken feather were collected from feather waste dumping area in and around Palakkarai, Erode District, Tamil Nadu, India (11.322879,77.544983). The soil samples were collected at 10–15 cm depth from the surface of the soil containing feather waste in sterile plastic bags and brought to the laboratory for further processing. Bacteria were isolated by serial dilution and plating methods on a nutrient agar medium. 10 g of homogenized soil sample was taken with 90 mL of sterilized distilled water and mixed well correctly and serially diluted to 10–7. The spread plate method was performed and the plates were kept at 37 °C for 48 h. From raw chicken feathers, the bacteria were isolated by the direct plating method in nutrient agar plates, and the plates were kept at 37 °C for 48 h. Pure cultures of the bacterial isolates were maintained on nutrient agar slants at 4 °C.
Preliminary Screening of Keratinolytic Bacteria
Screening on Skim Milk Agar
For screening of keratinase-producing bacteria, the bacterial isolates were inoculated on skim milk agar plates (Himedia M 763) [24]. The media plates were then incubated at 37 °C for a period of 24 h. After 24 h, plates were observed for the zone of hydrolysis. The diameter of each zone was measured. The bacterial colony which shows the zone of hydrolysis was selected for secondary screening and was maintained on nutrient agar slants at 4 °C for further study.
Screening on Casein Agar
The proteolytic activity of the isolates was confirmed by the casein hydrolysis method. The keratinolytic bacteria were streaked on casein agar (Himedia, India) plates and maintained at 37 °C for 48 h. The selection of isolates was done by the zone of clearance in this medium [25].
Preparation of Feather Meal Substrate
The feather meal was prepared from native chicken feathers as described earlier [26]. Briefly, the finely fragmented feather was defatted in chloroform to methanol (1:1 v/v) for 48 h, followed by chloroform to acetone to methanol (4:1:3 v/v/v) for another 48 h, and then rinsed in sterile water, dried for 24 h at 37 °C, and ground in a mortar-pestle to obtain a powdered feather.
Screening for the Highest Keratinase Producer
Nutrient broth of 50 mL was inoculated with a single colony of the selected isolate, under sterile conditions. The culture was then incubated overnight (16 h), at 37 °C using 150 rpm. After 16 to 18 h 1 ml of the bacterial inoculum was added in modified basal salt medium, enriched with feather meal under aseptic conditions (Feather Meal Broth Composition: NaCl-0.05%, MgCl2. 6H2O- 0.01% KH2PO4- 0.03%, K2HPO4- 0.04%, Glucose- 0.5%, Yeast Extract- 0.2%, Feather meal- 1% Distilled water- 100 mL, pH-7.5). The medium was incubated for a period of 72 h at 37 °C in a rotary shaking incubator, at 150 rpm. Following 72 h of incubation, centrifugation was done at 6000 rpm for 10 min and the supernatant was collected to be used as a crude enzyme and subjected to keratinase activity [27].
Determination of Proteolytic Activity
Keratinase enzyme activity was determined using the procedure described by MacDonald & Chen [28], using casein as substrate [29]. A reaction mixture was prepared by adding 1 mL of 1% casein and 1 mL of enzyme extract in the test tube and was then incubated at 40 °C temperature, for 30 min. The reaction between an enzyme and its substrate was stopped by adding 5 mL of 5% TCA (Trichloroacetic acid). The untreated casein was separated by centrifuging at 6000 rpm for 10 min. 5 ml of alkaline reagent, 1 mL of 1N NaOH was then added in 1 mL of supernatant, and 1 mL of Folin & Ciocalteau (FC) reagent was added to produce a dark blue colour. A control tube was also prepared in which 5 mL of 5% TCA was added, before incubation. The optical density of the mixture was measured at 660 nm by using a UV-spectrophotometer.
Protein Estimation
The amount of protein was determined in the culture supernatants according to Lowry’s method using bovine serum albumin as the standard. Readings were carried out in a spectrophotometer at 660 nm [30].
Identification of Potent Keratinolytic Bacterial Isolate
Phenotypic and Biochemical Characterization
The bacterial strain showing maximum keratinolytic activity was selected for further studies. The keratinolytic bacterial strain was identified based on the methods described in Bergey's Manual of Determinative Bacteriology and its morphological, cultural, and biochemical characteristics [31]. Biochemical characterization was done by performing specific tests such as Indole, Methyl red, Voges Proskauer and Citrate tests, Catalase, Oxidase, Triple Sugar Iron agar test, Carbohydrate fermentation tests, Urease test, Starch hydrolysis test. Further, molecular characterization was done to identify the potent strain at the species level.
Molecular Characterization of the Potent Strain
Bacterial DNA was isolated using phenol–chloroform isoamyl alcohol (25:24:1) from an overnight culture. Until it was needed again, extracted DNA was kept at -20 °C [32]. Using universal primers 27F (5′-AGA GTT TGA TCC TGG CTG AG-3) and 1492R (5′-GGC TAC CTT GTT ACG ACT T-3′), bacterial 16S rRNA was amplified by PCR. To 25 µl of PCR reaction solution (1.5 µl of Forward and Reverse primers, 5 µl of deionized water, and 12 µl of Taq Master Mix), 5 µl of extracted DNA was added. The steps involved in PCR amplification were as follows: two minutes of initial denaturation at 95˚C, 20 min of annealing at 50 ˚C, and two minutes of extension at 72 ˚C. For 10 min, the final extension is carried out at 72 ˚C. With the use of a Montage PCR Clean-up kit (Millipore), the PCR product was refined. The primers were used to sequence the PCR result. ABI PRISM® BigDyeTM Terminator Cycle Sequencing Kits with AmpliTaq® DNA polymerase (FS enzyme) (Applied Biosystems) was used to carry out the sequencing operations. The samples underwent electrophoresis on an ABI 3730xl sequencer (Applied Biosystems) after being resuspended in distilled water. The closest phylogenetic neighbour with > 99% sequence similarity was determined to be the species level after the sequences were compared to known sequences in the Gene Bank nucleotide database. 16S rRNA Sequence analysis was carried out using the BLAST algorithms (National Centre for Biotechnology Information [http://www.ncbi.nlm.nih.gov]) to ensure accuracy in identification. The alignment software Clustal X (version 1.81), which is freely available, was used to perform multiple sequence alignment approaches [33, 34].
Optimization of Culture Conditions for Maximum Keratinase Production
The optimization of media and the fermentation condition for keratinase production was done by the one-variable at a time (OVAT) method [35]. The effect of pH on the keratinase enzyme production was studied by varying the pH of the fermentation medium from 6 to 10. The pH was adjusted using 0.1N HCl and 0.1N NaOH in the production medium. The effect of temperature on keratinase enzyme production was studied by varying the temperature (25 °C, 37 °C, 40 °C, and 45 °C) to find out the optimum temperature for maximum keratinase production. The effects of different volumes of inoculum were investigated using 1%, 2%, 3%, 4%, 5%, and 6% in fermentation medium for maximum production of keratinase. The effect of carbon sources on the keratinase enzyme production was analysed by using 5 different media prepared by replacing the carbon source (0.5% Glucose) in the fermentation medium to find out the favourable carbon on the supplement. The different carbon sources such as Fructose, Lactose, Maltose, Galactose, and Sucrose. The effect of nitrogen sources on keratinase enzyme production was analysed by using 5 different media prepared by replacing the nitrogen source in the fermentation medium to find out the favourable nitrogen source on the supplement. Yeast extract (0.2%) in the basal media was replaced with Gelatin, Casein, Urea, Ammonium nitrate, and Beef extract at the same concentration, individually.
Degradation of Keratin Waste Samples in Optimized Liquid Medium
Preparation of Inoculum for Keratin Degradation
An inoculum had been prepared in order to present findings on keratin waste deterioration [36]. The physiochemically optimized keratinase production broth (100 mL) was sterilized for 20 min at 121 °C. A loopful of the sterile broth was added. A sterilised bacterial culture that produced keratinase was inoculated. The broth culture was incubated at 37 °C for 14 h using a rotary shaker set at 150 rpm. This acted as an inoculum for the upcoming degradation studies.
Degradation of Keratin Wastes in Shake Flask Fermentation Method
The efficacy of degradation of keratin-rich wastes (whole feather, bovine hair, bovine hide and chicken nail) was tested on an optimized medium as per the method [36]. Degradation of keratin substrates was visually inspected and aliquots were removed for keratinase activity and degree of degradation percentages were determined.
Microscopic Examination of Degraded Keratin Wastes
Light Microscopy
The disintegrated keratin wastes (whole feather, chicken nail) were collected and carefully cleansed to remove debris after 35 days of incubation. They were then examined under a light microscope. Microscopic examinations were performed on disintegrated whole feathers and chicken nail remnants at a magnification of 100 x [37, 38].
FESEM Analysis
To check for keratinase activity, the structural changes of biodegraded keratin waste (bovine hair) were examined by Field emission scanning electron microscopy (FESEM) as described by [21, 39]. FESEM analysis was done at the Department of Nanoscience and Technology, Bharathiar University, Coimbatore.
Statistical Analysis
All experiments were carried out in triplicates. IBM SPSS Statistics 25 software was used for the statistical analysis (ANOVA) and the means were compared by Duncan’s test. When P < 0.05, differences were deemed significant. Values are expressed as means ± standard deviation (SD). In graphs, the standard error values are represented as Y error bars. Means with the different letters are significantly different.
Results
Isolation and Screening of Potent Keratinase Producing Bacterial Strain
The bacteria were isolated from three different samples- Poultry feather soil (fresh), Poultry feather soil (old), and Raw chicken feathers in and around areas of poultry feather dumping sites. Nutrient agar plates upon serial dilution showed morphologically different colonies at 10–5 dilution. A total of 19 bacterial colonies were isolated from Poultry feather soil (fresh)-9, Poultry feather soil (old)-7. Through the direct plating method from raw chicken feathers 3 different colonies were isolated. The isolates were maintained in nutrient agar slants for further studies (Figure S1).
The bacterial isolates were screened for their proteolytic activity on skim milk and casein agar plates. Amongst the 19 bacterial strains plated on skim milk agar plates, a total of 9 strains showed proteolytic activity (Figure S2.A). The strain isolated from raw feather (F2) and poultry feather soil (HFSO7) actively exhibited clear larger zones of inhibition of about 4.5 mm and 3 mm. Next to them, HFSO6, HFSO1, HFSO4, HFSF1, and HFSF4 showed comparatively limited hydrolytic activity with a moderate zone of clearance. A very mild zone of clearance was observed in F3 and HFSF9. The noteworthy nature of F2 and HFSO7 would be due to the secretion of casein (proteolytic enzyme), thereby resulting in a clear zone of hydrolysis (Figure S2 B). The less proteolytic activity in F3 and HFSF9 points to their insufficiency to secrete enough proteolytic enzymes. Hence, F3 and HFSF9 were neglected for further screening studies. The isolates F2, HFSO7, HFSO1, and HFSO6 showed a significant zone of clearance on casein agar plates. Among them, F2 and HFSO7 showed maximum zones of hydrolysis of 4.8 mm and 3.3 mm respectively. From, the obtained results it is evident that the isolate can hydrolyse keratin. The 4 positive isolates were taken for further secondary screening to select the highest keratinase producer. The comparative growth activity of the keratinolytic strains in screening on different media is tabulated below (Table 1).
Secondary screening of the isolates was carried out in submerged conditions for which chicken feather meal was used as a sole source of carbon and nitrogen. The isolates HFSO1, HFSO6, HFSO7, and F2 were subjected to keratinase production. The results revealed that the bacterial strain HFSO6 and HFSO1 exhibited the lowest keratinase production of 7.39 ± 0.14 U/mL and 9.02 ± 0.17 U/mL with protein concentrations of 0.65 ± 0.07 mg/mL and 1.06 ± 0.08 mg/mL. However, a marked increase of 32.65 ± 0.16 U/mL and 24.59 ± 0.09 U/mL with protein concentrations of 4.54 ± 0.14 mg/mL and 3.34 ± 0.09 mg/mL was observed with F2 and HFSO7 (Fig. 1). The isolate F2 which exhibited the highest enzyme production was selected as the potent keratinolytic bacterial strain for further studies.
Identification of Potent Keratinolytic Bacterial Strain
The primary identification of the selected bacteria is provided by the Gram staining technique. When observed under the microscope the bacteria appear as endospores forming gram-positive rods (Figure S3 A &B). The morphological and biochemical characteristics of the selected strain were tabulated (Table 2). From the results of the biochemical test the selected bacterial strain comes under the genus Bacillus. For species level identification, molecular characterization is done further. After morphological and biochemical characterization, the selected keratinase-producing bacterial strain was identified by 16S rRNA sequencing using the 27F and 1492R universal primers. Bacterial identifications were based on 16S rRNA gene sequence similarity. Neighbours joining the phylogenetic tree were generated using sequence data from the gene bank for strains that showed a high percentage of similarities with our strain. The NCBI database disclosed that the alignment findings of the 16S rRNA sequences from the strain HFS_F2T were homologous with numerous references for the Bacillus genus. The results revealed that the 16S rRNA partial gene sequence of the isolate HFS_F2T showed the highest similarity (> 99%) with Bacillus velezensis SM-10 MT377875.1 T. The sequences were submitted in gene bank with accession number: ON556508 (Fig. 2). The results of morphological, biochemical, and molecular characterization confirmed that the potent keratinase producing bacterial strain was Bacillus velezensis.
Optimization of Bioprocess Parameters for Maximum Keratinase Production
Effect of pH
Keratinase activity at various pH was determined. Maximum activity of about 32.56 ± 0.10 U/mL was recorded at pH-7, followed by pH-6 (26.53 ± 0.13 U/mL) and pH-8 (23.61 ± 0.05 U/mL). Considerably lower enzyme activity was observed at pH-9 (24.29 ± 0.04 U/mL) and pH-10 (19.73 ± 0.05 U/mL). As the pH increased beyond 7, keratinase activity decreased Fig. 3a. This reflected the neutral nature of the produced enzyme.
Optimization of bioprocess variables for maximizing keratinase production. a Effect of pH on keratinase production. b Effect of temperature on keratinase production. c Effect of inoculum size on keratinase production. d Effect of carbon source on keratinase production. e Effect of nitrogen source on keratinase production. The standard deviation and mean of the values are displayed. Significant differences are indicated by values above the bar that do not have the same superscript letters according to Duncan’s at P < 0.05
Effect of Temperature
Keratinase production by Bacillus velezensis at different temperatures such as 25 °C, 37 °C, 40 °C, and 45 °C was determined to study the effect of temperature on keratinase activity. After incubation, the highest keratinase activity of 100.25 ± 0.27 U/mL was achieved at 37 °C followed by 67.76 ± 1.13 U/mL at 40 °C. Rise in temperature resulted in a decline of enzyme activity to 32.40 ± 1.26 U/mL at 45 °C. Thus, the optimal temperature required by Bacillus velezensis for keratinase production was 37 °C Fig. 3b.
Effect of Inoculum Size
The effect of inoculum size on the production of keratinase by Bacillus velezensis was studied for inoculum sizes of 1 to 6% (v/v) as presented in Fig. 3c. From the results, it was observed that the maximum production (44.28 ± 0.07 U/mL) was obtained at 5% of inoculum size. Bacillus velezensis showed higher production of keratinase as the inoculum size was increased above 2%. Minimum enzyme activity of (18.86 ± 0.22 U/mL) was observed at 1% of inoculum size.
Effect of Carbon Source
The keratinase production medium (feather meal broth) compromised of 0.5% glucose was replaced with various commercial carbon sources such as lactose, galactose, fructose, maltose, and sucrose were evaluated for the keratinase enzyme activity. Maximum extracellular enzyme activity of about 38.72 ± 0.09 U/mL was recorded with sucrose as a carbon source Fig. 3d, which was relatively higher than that of other sugars employed – maltose (33.5 ± 0.14 U/mL) > fructose (29.75 ± 0.10 U/mL) > lactose (26.96 ± 0.15 U/mL) > galactose (24.47 ± 0.24 U/mL). From the results, 0.5% glucose was replaced with 0.5% sucrose for further studies.
Effect of Nitrogen Source
About 0.2% of distinct nitrogen sources of commercial grade (beef extract, casein, gelatin, urea, and ammonium nitrate) were exploited for the production medium for evaluating the keratinase activity. The highest keratinase production of about 32.33 ± 0.11 U/mL was achieved with beef extract as a nitrogen source, followed by casein (26.23 ± 0.32 U/mL), ammonium nitrate (24.26 ± 0.10 U/mL) and gelatin (23.76 ± 0.30 U/mL) Fig. 3e. Lower keratinase activity was observed with urea (19.77 ± 0.35 U/mL). Hence, 0.2% of yeast extract was replaced with beef extract for further use.
Degradation of keratin waste samples
The efficacy of Bacillus velezensis in degrading keratin wastes (whole feather, chicken nails, bovine hair, and bovine hide) by the shake flask method has been evaluated. Bacillus velezensis strain inoculated into the modified keratinase production medium (devoid of feather meal) served as an inoculum for degradation studies. The incubation period was 35 days and aliquots were visualized for degradation every week. After 35 days of incubation, about 77.32 ± 0.32% of degradation with bovine hair was achieved. Next to bovine hair, the whole feathers with 71.40 ± 0.21% degradation were notable. The chicken nails and bovine hide with 63.56 ± 0.12% and 58.94 ± 0.09% degradation respectively was comparatively the least degraded keratin waste. Maximum degradation of bovine hair and chicken feathers would be attributed to the keratinolytic nature of Bacillus velezensis. The enzyme secreted by the microbe would have exhibited proteolytic and sulfitolytic properties disintegrating the keratin structure into finer peptide units. Bovine hair and chicken feather would have served as a favourable substrate of microbial growth and produced keratinolytic enzymes adequately to degrade the keratin samples. The keratinase enzyme activity was checked after the degradation of keratin waste with maximum activity in bovine hair (142.55 ± 0.26 U/mL) followed by whole feather (112.39 ± 0.15 U/mL). The comparative minimum keratinase activity was seen in chicken nails (104.12 ± 0.21 U/mL) followed by bovine hide (96.74 ± 0.10 U/mL) (Fig. 4).
Visual and Microscopic Examination of Degraded Keratin Wastes
Observation of Whole Feather, Chicken Nail, and Bovine Hide Degradation
The efficiency of feather degradation and chicken nails was observed under the light microscope. The maximum degeneration of feather barbules was observed from the rachis region of the feather. The residual feather barbules were observed within the media. The bacteria adhered to the surface penetrated the feather and finally broke it into a powdered mass. The bacteria penetrated the quill region of the feather first and with the separation of the barb which hydrolyse into a gelatinous mass. Complete degradation was achieved in 21 days of incubation. Hollow structures were observed in chicken nails under 100 × after 28 days of incubation which confirms the microbial keratinase activity. The chicken nail keratin was hydrolysed and the disintegration of the structure was confirmed visually. The dehairing activity was observed with bovine hide. After 14 days of incubation partially dehaired bovine hide with damaged skin was visually inspected (Figure S4, S5).
FESEM Analysis of Degraded Bovine Hair
Since, the degradation of bovine hair cannot be visually inspected; FESEM analysis was performed with the microscopic examination of the control bovine hair (Fig. 5 A- i, ii, iii, iv) and degraded bovine hair (Fig. 5 B- i, ii, iii, iv) for comparison. The normal bovine hair shaft is made up of overlapping scales of dead cells containing the protein keratin (control). The structural changes of the hair after 35 days of degradation were analysed. Complete degradation of the cuticle was observed with exposure of the cortex and damaged medulla.
Discussion
The upsurge accumulation of keratin feather residues produced by poultry and leather industries and lacking of ecologically sound remedies have resulted to the screening of prospective keratin-degrading microbes. Bacillus sp NDS-10 from soil [40]; Pseudomonas aeruginosa from slaughterhouses [41]; Bacillus cereus from poultry dump soil [42]; Bacillus thuringiensis strain MT1 from cattle-yard [42]; Bacillus pumilus AR57 from the slaughterhouse soil [43], Bacillus velezensis strain ZBE1 from deep forest soil [44] etc. In most of the studies, potential keratinolytic bacteria have been isolated from chicken feather dumping sites since most Bacillus sp. are feather keratin degraders [45]. According previous literature study, the first keratinolytic species of Bacillus velezensis was identified which degrades keratin chicken feathers [45]. Nutritional and environmental variables possess a substantial effect on the biosynthesis of keratinase enzyme [46]. Numerous bioprocess variables, including temperature, pH, and the types of carbon and nitrogen sources in the medium, might influence the keratinases that microorganisms produce. The present study is correlated with [47], where maximum keratinase production was achieved at pH-7 by most of the Bacillus strains. In a recent study, S. netropsis and B. subtilis produced the highest concentrations of keratinase (47.9 ± 1.3 and 57.8 ± 1.4 U/mL) at pH 7 and 7.5, respectively. Generally, the most favourable pH for keratinase production from bacteria, actinomycetes, and fungi, was found to be in the neutral to alkaline range [48]. On day 3 (72 h), there was a significant increase in keratinase activity at pH 7 compared to previous days which was similar to the results of recent study with Bacillus cereus FD2 keratinolytic strain. The shake flask feather media had a milky appearance with digested feather debris, with neutral pH being the most favourable pH range. Acidic and basic pH showed significant results when compared to neutral pH [49]. This study revealed that the enzyme produced by the bacterial species was considerably thermo-stable up to 40 °C, but was able to exhibit maximum activity at 37 °C which was most commonly maintained temperature for most of the fermentation processes. The results were correlated with previous reports by [50]. According to Kainoor and Naik (2010), Bacillus berevis produces a considerable amount of keratinase at pH 7.5 and 37 °C. The results coincided with their findings [51]. The results of the present study were found similar to the previous studies as many workers have described that higher keratinase production is obtained at a higher percentage of inoculum sizes, for instance [52] observed maximum keratinase production at a 5% concentration of inoculum size with B. cereus LAU08 strain. Similarly, in a previous study, 5% of the initial inoculum was optimal for keratinase production by Bacillus sp. FK 46 [53]. The further increase in inoculum size decreased the keratinase enzyme activity, so 5% of inoculum is fixed for further studies. Cai et al., 2009 similarly reported that sucrose stimulated keratinase production the most and was selected as an extra carbon source in media optimization [54]. Thus, sucrose serves as a vital carbon source next to glucose supporting the metabolism of distinct keratinolytic microorganisms. Numerous findings on beef extract as a potent nitrogen source for Bacillus strains have been reported earlier. Similarly, Singh et al., 2017 reported beef extract as the best additional nitrogen source for keratinase production by Bacillus subtilis [55]. Following optimization, it is evident that the production of keratinase is stimulated by extra supplies of carbon and nitrogen. The optimized factors were subsequently incorporated into the keratinase production broth, altering the current medium for maximum keratinase production. Further degradation studies of keratin wastes were performed with the optimized keratinase production medium consisting of—sucrose (carbon source), beef extract (nitrogen source) with pH-7, inoculum size (5%), and incubation at 37 °C). Keratinase activity was detected during growth, but the complete degradation of these substrates was not always achieved. From the results, it is evident that keratinase activity influenced the degradation rate of different keratin substrates. Because of the long-term stationary phase, which is characterised by continuous nutrient starvation in the media and constant bacterial cell viability, the degradation process is only evaluated after 35 days. During the course of this period, keratinase enzyme activity gradually decreased until it eventually dissipated and then the degraded samples were undergone for further microscopic analysis. Similarly, B. licheniformis S23, one of the strains investigated by Lal et al.,1999 had a remarkable capacity for decomposing human hair, particularly in long-term cultures. After a month of cultivation, the strain's maximum keratinase activity was reached, and its concentration of minimised thiols surpassed 0.3 mM, revealing its keratinolytic action on hair as compared to bovine hoove or horn and human nail matter [56]. The results correlate with previous reports where 58% of chicken feather degradation with Bacillus mycoides was achieved [57]. The results were correlated with reports of [58]. B. licheniformis K18102, comparative cultures on chicken feathers and other materials like bovine hair, wool, human hair, and nails were investigated. The activity on bovine hair was 69.2% which was lower than the present study with 77.32 ± 0.32% degradation using Bacillus velezensis. From the result, it was observed that keratin protease can hydrolyse bovine collagen (skin) since most of the Bacillus species are soluble protein degraders [59]. The results indicated that the structural integrity of the bovine hair is lost after degradation by 77% which is comparatively lower with previous investigation with the Brevibacterium luteolum strain degraded sheep's wool and goat hair by almost 85%. The results were correlated with [60]. Our bacterial strain’s degradation efficacy is compared to S. albidoflavus, which could only break down 10% of the keratin in hair [61]. In a prior study, the thermophilic Bacillus sp. PA-001A hydrolysed 90%, 60%, and 50% of the sheep skin, feather, and horn that were used in the medium. Whereas, the hair was not well supported by the medium, compared to our keratinolytic strain's ability to hydrolyse all of the keratin wastes including bovine hair in a single optimized fermented medium [62].From the FESEM micrograph results, it was evident that Bacillus velezensis HFS_F2T can hydrolyse bovine hair keratin leading to the complete disintegration of hair structure could be used as the potent keratinolytic bacterial strain in the degradation of bovine hair waste from leather industries at large scale. The outcomes unambiguously demonstrate the overall effectiveness of the isolated potent keratinolytic strain Bacillus velezensis HFS_F2T in the biodegradation of all environmentally hazardous keratin wastes in one optimized keratinase production medium, and it exhibited an acceptable degree of degradation efficacy for potential utilization of these microbial keratinases in direct environmental applications further.
Conclusion
In the present study, Bacillus velezensis strain (HFS_F2T) was identified as a potent keratinolytic bacterial strain which exhibited the highest keratinase activity with 32.65 ± 0.16 U/mL than the other stains. Enhanced keratinase production and keratin waste degradation were achieved through optimization of basal media at pH (7), temperature (37 °C), inoculum level (5%), carbon source (sucrose), and nitrogen source (beef extract). After 35 days of incubation of keratinous wastes in optimized keratinase production broth, an elevated degradation of bovine hair with 77.32 ± 0.32% followed by whole feather with 71.40 ± 0.21%, chicken nail with 63.56 ± 0.12%, and bovine hide 58.94 ± 0.09% were observed with partially degraded keratin debris in the medium. In light of the present results, it could be concluded that Bacillus velezensis HFS_F2T was found to be a potential candidate for keratinase production and in addition, they were able to grow and display keratinolytic activity in diverse keratin wastes resulting in effective degradation which was confirmed through weight loss and morphological changes. Consequently, this research raises hope for the successful implementation of a robust keratinolytic bacterial strain for the commercialization of microbial keratinases and the biological degradation of industrial leather and poultry pollutants. Recycling keratinous wastes with limited resources would address the problem of disposing of trash and detrimental impacts on ecology which perhaps yield economic and biotechnological benefits. Conversely, research prospects for the future hinge on the ability to convert by-products from the biodegradation process into value-added products, such as animal feed, biofertilizers, cosmetic products, composites, detergents, etc.
Availability of Data and Materials
Not applicable.
References
Anbesaw MS (2022) Bioconversion of keratin wastes using keratinolytic microorganisms to generate value-added products. Int J Biomater. https://doi.org/10.1155/2022/2048031
Sharma S, Gupta A (2016) Sustainable management of keratin waste biomass: applications and future perspectives. Braz Arch Biol Technol 59:1–14. https://doi.org/10.1590/1678-4324-2016150684
Akhter M, Wal Marzan L, Akter Y, Shimizu K (2020) Microbial bioremediation of feather waste for keratinase production: an outstanding solution for leather detailing in tanneries. Microbiol Insights 13:1178636120913280. https://doi.org/10.1177/1178636120913280
Onifade A, Al-Sane N, Al-Musallam A, Al-Zarban S (1998) A review: potentials for biotechnological applications of keratin-degrading microorganisms and their enzymes for nutritional improvement of feathers and other keratins as livestock feed resources. Bioresour Technol 66:1–11. https://doi.org/10.1016/S0960-8524(98)00033-9
Kokilan R, Priya BD (2022) A sustainable approach for processing organic waste in India. Mater Today 64:1069–1074. https://doi.org/10.1016/j.matpr.2022.05.291
Williams CM, Lee CG, Garlich JD, Shih JC (1991) Evaluation of a bacterial feather fermentation product, feather-lysate, as a feed protein. Poult Sci 70(1):85–94. https://doi.org/10.3382/ps.0700085
Saber WIA, El-Metwally MM, El-Hersh MS (2010) Keratinase production and biodegradation of some keratinous wastes by Alternaria tenuissima and Aspergillus nidulans. Res J Microbiol 5(1):21–35
Kumawat TK, Sharma A, Sharma V, Chandra S (2018) Keratin waste: the biodegradable polymers. In Keratin. IntechOpen
Da Silva, R. R. (2018). Keratinases as an alternative method designed to solve keratin disposal on the environment: its relevance on agricultural and environmental chemistry, 7219–7221. https://doi.org/10.1021/acs.jafc.8b03152
Sharma I, Pranaw K, Soni H et al (2022) Parametrically optimized feather degradation by Bacillus velezensis NCIM 5802 and delineation of keratin hydrolysis by multi-scale analysis for poultry waste management. Sci Rep 12:17118. https://doi.org/10.1038/s41598-022-21351-9
Sharma R, Devi S (2018) Versatility and commercial status of microbial keratinases: a review. Rev Environ Sci Bio/Technol 17(1):19–45. https://doi.org/10.1007/s11157-017-9454-x
Li Q (2019) Progress in microbial degradation of feather waste. Front Microbiol 10:2717. https://doi.org/10.3389/fmicb.2019.02717
Gupta R, Ramnani P (2006) Microbial keratinases and their prospective applications: an overview. Appl Microbiol Biotechnol 70:21–33. https://doi.org/10.1007/s00253-005-0239-8
Yamamura S, Morita Y, Hasan Q, Yokoyama K, Tamiya E (2002) Keratin degradation: a cooperative action of two enzymes from Stenotrophomonas sp. Biochem Biophys Res Commun 294(5):1138–1143. https://doi.org/10.1016/s0006-291x(02)00580-6
Vidmar B, Vodovnik M (2018) Microbial keratinases: enzymes with promising biotechnological applications. Food Technol Biotechnol 56(3):312–328. https://doi.org/10.17113/ftb.56.03.18.5658
Tamreihao K, Mukherjee S, Khunjamayum R, Devi LJ, Asem RS, Ningthoujam DS (2019) Feather degradation by keratinolytic bacteria and biofertilizing potential for sustainable agricultural production. J Basic Microbiol 59(1):4–13. https://doi.org/10.1002/jobm.201800434
Williams CM, Shih JCH (1989) Enumeration of some microbial groups in thermophilic poultry waste digesters and enrichment of a feather-degrading culture. J Appl Bacteriol 67(1):25–35. https://doi.org/10.1111/j.1365-2672.1989.tb04951.x
Chaturvedi V, Bhange K, Bhatt R, Verma P (2014) Production of kertinases using chicken feathers as substrate by a novel multifunctional strain of Pseudomonas stutzeri and its dehairing application. Biocatal Agric Biotechnol 3(2):167–174. https://doi.org/10.1016/j.bcab.2013.08.005
Bohacz J, Korniłłowicz-Kowalska T (2019) Fungal diversity and keratinolytic activity of fungi from lignocellulosic composts with chicken feathers. Process Biochem 80:119–128. https://doi.org/10.1016/j.procbio.2019.02.012
Manczinger L, Rozs M, Vágvölgyi C, Kevei F (2003) Isolation and characterization of a new keratinolytic Bacillus licheniformis strain. World J Microbiol Biotechnol 19(1):35–39. https://doi.org/10.1023/A:1022576826372
Gupta S, Singh R (2014) Hydrolyzing proficiency of keratinases in feather degradation. Indian J Microbiol 54(4):466–470. https://doi.org/10.1007/s12088-014-0477-5
Brandelli A, Sala L, Kalil SJ (2015) Microbial enzymes for bioconversion of poultry waste into added-value products. Food Res Int 73:3–12. https://doi.org/10.1016/j.foodres.2015.01.015
Kang D, Herschend J, Al-Soud WA, Mortensen MS, Gonzalo M, Jacquiod S, Sørensen SJ (2018) Enrichment and characterization of an environmental microbial consortium displaying efficient keratinolytic activity. Biores Technol 270:303–310. https://doi.org/10.1016/j.biortech.2018.09.006
Barman NC, Zohora FT, Das KC, Mowla M, Banu NA, Salimullah M, Hashem A (2017) Production, partial optimization and characterization of keratinase enzyme by Arthrobacter sp. NFH5 isolated from soil samples. AMB Express 7(1):1–8. https://doi.org/10.1186/s13568-017-0462-6
Sekar V, Kannan M, Ganesan R, Dheeba B, Sivakumar N, Kannan K (2016) Isolation and screening of keratinolytic bacteria from feather dumping soil in and around Cuddalore and Villupuram, Tamil Nadu. Proc Natl Acad Sci India Sect B 86:567–575. https://doi.org/10.1007/s40011-014-0483-8
Tork S, Aly M, Nawar L (2010) Biochemical and molecular characterization of a new local keratinase producing Pseudomomanas sp., MS21. Asian J Biotechnol 2:1–13
Rajesh TP, Rajasekar S, Mathan RKH, Anandaraj B (2016) Isolation and identification of feather degrading bacteria from feather-dumped soil. Int J Environ Sustain Dev 15:293–299. https://doi.org/10.1504/IJESD.2016.077393
MacDonald CE, Chen LL (1965) Lowry modification of the Folin reagent for determination of proteinase activity. Anal Biochem 10:175. https://doi.org/10.1016/0003-2697(65)90255-1
Gowdhaman D, Ponnusami V (2014) Production of keratinase from a new strain of Pseudomonas aeruginosa gmp and its application for the removal of dyed keratin waste. Bioresour Technol 9(5):210–217
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin’s phenol reagent. J Biol Chem 193:265–275
Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer K-H, Whitman WB (2011) Bergey’s manual of systematic bacteriology: Volume 3: The Firmicutes. Springer, New York
Butler JM (2011) Advanced topics in forensic DNA typing: methodology. Academic Press, Washington
Abid S, Farid A, Abid R, Rehman MU, Alsanie WF, Alhomrani M, Ghazanfar S (2022) Identification, biochemical characterization, and safety attributes of locally isolated Lactobacillus fermentum from Bubalus bubalis (buffalo) milk as a probiotic. Microorganisms 10(5):954. https://doi.org/10.3390/microorganisms10050954
Sadiqi S, Hamza M, Ali F, Alam S, Shakeela Q, Ahmed S, Zaman W (2022) Molecular characterization of bacterial isolates from soil samples and evaluation of their antibacterial potential against MDRS. Molecules 27(19):6281
Zhou L, Liu Y, Sun H, Li H, Zhang Z, Hao P (2022) Usefulness of enzyme-free and enzyme-resistant detection of complement component 5 to evaluate acute myocardial infarction. Sens Actuators B 369:132315. https://doi.org/10.1016/j.snb.2022.132315
Masih H, Singh S (2014) Degradation of keratinous waste products by keratinolytic bacteria isolated from soil. Int J Eng Comp Sci 3:7588–7595
Ahmad S, Ahmad M, Fawzy Ramadan M, Sultana S, Papini A, Ullah F, Zafar M (2023) Palynological study of fossil plants from miocene murree formation of Pakistan: clues to investigate palaeoclimate and palaeoenvironment. Agronomy 13(1):269. https://doi.org/10.3390/agronomy13010269
Bahadur S, Long W, Ahmad M, Yaseen M, Ullah F, Saqib S (2023) Exploration of pollen traits and their taxonomic relevance in selected taxa of the subfamily Papilionoideae from Hainan Island, China. Palynology 47(2):2144521. https://doi.org/10.1080/01916122.2022.2144521
Madkour FA, Abdelsabour-Khalaf M (2022) Performance scanning electron microscopic investigations and elemental analysis of hair of the different animal species for forensic identification. Microsc Res Tech 85(6):2152–2161. https://doi.org/10.1002/jemt.24073
Akram F, Aqeel A, Shoaib M, Haq IU, Shah FI (2022) Multifarious revolutionary aspects of microbial keratinases: an efficient green technology for future generation with prospective applications. Environ Sci Pollut Res 29(58):86913–86932. https://doi.org/10.1007/s11356-022-23638-w
Pei, X. D., Li, F., Zhang, Y. M., Huang, X. N., Yu, F. T., Su, L. Y., ... & Wang, C. H. (2023). Preparation, Purification, and Identification of Novel Feather Keratin-Derived Peptides with Antioxidative and Xanthine Oxidase Inhibitory Activities. Journal of Agricultural and Food Chemistry. https://doi.org/10.1021/acs.jafc.3c01131
Yadav S, Bumbra P, Laura JS, Khosla B (2022) Optimization of nutritional and physical parameters for enhancing the keratinase activity of Bacillus cereus isolated from soil of poultry dump site in Gurugram, Haryana. Bioresource Technol Rep 18:101108. https://doi.org/10.1016/j.biteb.2022.101108
Jagadeesan Y, Meenakshisundaram S, Saravanan V, Balaiah A (2020) Sustainable production, biochemical and molecular characterization of thermo-and-solvent stable alkaline serine keratinase from novel Bacillus pumilus AR57 for promising poultry solid waste management. Int J Biol Macromol 163:135–146. https://doi.org/10.1016/j.ijbiomac.2020.06.219
Revankar AG, Bagewadi ZK, Bochageri NP, Khan TY, Shamsudeen SM (2023) Response surface methodology based optimization of keratinase from Bacillus velezensis strain ZBE1 and nanoparticle synthesis, biological and molecular characterization. Saudi J Biol Sci 30(10):103787. https://doi.org/10.1016/j.sjbs.2023.103787
Sutoyo S, Subandi S, Ardyati T, Suharjono S (2019) Isolation and identification of keratinolytic bacteria from Jember, Indonesia as a biodegradation agent of chicken feather wastes. Asian J Agric Biol 7(4):491–500
Zambare VP, Nilegaonkar SS, Kanekar PP (2007) Production of an alkaline protease by Bacillus cereus MCM B-326 and its application as a dehairing agent. World J Microbiol Biotechnol 23:1569–1574. https://doi.org/10.1007/s11274-007-9402-y
Qadar SA, Shireen E, Iqbal S, Anwar A (2009) Optimization of protease production from newly isolated strain of Bacillus sp. PCSIR EA-3
Abdelmoteleb A, Gonzalez-Mendoza D, Tzintzun-Camacho O, Grimaldo-Juárez O, Mendez-Trujillo V, Moreno-Cruz C, Roumia AF (2023) Keratinases from Streptomyces netropsis and Bacillus subtilis and their potential use in the chicken feather degrading. Fermentation 9(2):96. https://doi.org/10.3390/fermentation9020096
Deba F, Nelofer R, Irfan M (2023) Isolation, identification, and screening of keratinase producing bacteria from soil and production optimization using feather waste as substrate. Punjab Univ J Zool 38(1):109–118. https://doi.org/10.17582/journal.pujz/2023.38.1.109.118
Subugade S, Gupta SG, Mokashe S (2017) Isolation and screening of keratinase producing bacteria from chicken feather dumping site. Int J ChemTech Res 10:900–905
Kainoor PS, Naik GR (2010) Production and characterization of feather degrading keratinase from Bacillus sp. JB 99
Lateef A, Oloke JK, Kana EG, Sobowale BO, Ajao SO, Bello BY (2010) Keratinolytic activities of a new feather-degrading isolate of Bacillus cereus LAU 08 isolated from Nigerian soil. Int Biodeterior Biodegrad 64(2):162–165. https://doi.org/10.1016/j.ibiod.2009.12.007
Suntornsuk W, Suntornsuk L (2003) Feather degradation by Bacillus sp. FK 46 in submerged cultivation. Bioresource Technol 86(3):239–243. https://doi.org/10.1016/S0960-8524(02)00177-3
Cai C, Zheng X (2009) Medium optimization for keratinase production in hair substrate by a new Bacillus subtilis KD-N2 using response surface methodology. J Ind Microbiol Biotechnol 36(7):875–883. https://doi.org/10.1007/s10295-009-0565-4
Singh S, Masih H, Jeyakumar GE, Lawrence R, Ramteke PW (2017) Optimization of fermentative production of keratinase by Bacillus subtilis strain S1 in submerged state fermentation using feather waste. Int J Curr Microbiol App Sci 6(12):1499–1510. https://doi.org/10.20546/ijcmas.2017.612.167
Lal S, Rajak RC, Hasija SK (1999) In vitro degradation of keratin by two species of Bacillus. J Gen Appl Microbiol 45(6):283–287. https://doi.org/10.2323/jgam.45.283
Beryl GP, Thazeem B, Umesh M (2021) Bioconversion of feather composts using proteolytic Bacillus mycoides for their possible application as biofertilizer in agriculture. Waste Biomass Valor 12:6795–6809. https://doi.org/10.1007/s12649-021-01472-4
Desai SS, Hegde S, Inamdar P, Sake N, Aravind MS (2010) Isolation of keratinase from bacterial isolates of poultry soil for waste degradation. Eng Life Sci 10(4):361–367. https://doi.org/10.1002/elsc.200900009
Hassan MA, Haroun BM, Amara AA, Serour EA (2013) Production and characterization of keratinolytic protease from new wool-degrading Bacillus species isolated from Egyptian ecosystem. BioMed Res Intl 2013:1–14. https://doi.org/10.1155/2013/175012
Thankaswamy SR, Sundaramoorthy S, Palanivel S, Ramudu KN (2018) Improved microbial degradation of animal hair waste from leather industry using Brevibacterium luteolum (MTCC 5982). J Clean Prod 189:701–708. https://doi.org/10.1016/j.jclepro.2018.04.095
Bressollier P, Letourneau F, Urdaci M, Verneuil B (1999) Purification and characterization of a keratinolytic serine proteinase from Streptomyces albidoflavus. Appl Environ Microbiol 65(6):2570–2576
Atalo K, Gashe BA (1993) Protease production by a thermophilic Bacillus species (P-001A) which degrades various kinds of fibrous proteins. Biotech Lett 15:1151–1156. https://doi.org/10.1007/BF00131207
Acknowledgements
The authors thank the Department of Science and Technology (DST) for supporting the Department of Microbial Biotechnology, Bharathiar University, Tamil Nadu under the DST-FIST scheme to have the necessary facilities for the successful execution of this research work.
Funding
No funding was received to assist with the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All the authors contributed to the fulfilment of the manuscript. Concept, experiment, material preparation, data collection, and analysis were performed by KS and AV. The first draft of the manuscript was written by KS. The corresponding author, Dr. PK commented on previous versions of the manuscript and proofread and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Not applicable.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saravanan, K., Vijayaveeran, A. & Kathirvel, P. Biodegradation of Keratin Waste by Bacillus velezensis HFS_F2 through Optimized Keratinase Production Medium. Curr Microbiol 81, 179 (2024). https://doi.org/10.1007/s00284-024-03699-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-024-03699-5