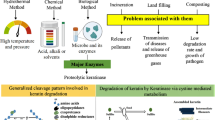
Abstract
Since the dawn of century, tons of keratin bio-waste is generated by the poultry industry annually, and they end up causing environmental havoc. Keratins are highly flexible fibrous proteins which exist in α- and β- forms and provide mechanical strength and stability to structural appendages. The finding of broad-spectrum protease, keratinase, from thermophilic bacteria and fungi, has provided an eco-friendly solution to hydrolyze the peptide bonds in highly recalcitrant keratinous substances such as nails, feathers, claws, and horns into valuable amino acids. Microorganisms produce these proteolytic enzymes by techniques of solid-state and submerged fermentation. However, solid-state fermentation is considered as a yielding approach for the production of thermostable keratinases. This review prioritized the molecular and biochemical properties of microbial keratinases, and the role of keratinases in bringing prodigious impact for the sustainable progress of the economy. It also emphasizes on the current development in keratinase production with the focus to improve the biochemical properties related to enzyme’s catalytic activity and stability, and production of mutant and cloned microbial strains to improve the yield of keratinases. Recently, multitude molecular approaches have been employed to enhance enzyme’s productivity, activity, and thermostability which makes them suitable for pharmaceutical industry and for the production of animal feed, organic fertilizers, biogas, clearing of animal hides, and detergent formulation. Hence, it can be surmised that microbial keratinolytic enzymes are the conceivable candidates for numerous commercial and industrial applications.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Keratin waste comprises 85% crude protein and is the third most abundant biopolymer waste after cellulose and chitin. Protein processing industries, including slaughterhouses, meat packing, and leather processing plants, produce a substantial amount of keratin-containing wastes. Of these industries, about seven billion tons of keratinous waste is added to the world’s solid waste biomass annually due to the poultry slaughter industry (da Silva 2018). Therefore, management strategies to mitigate the environmental impact of keratin waste are urgently required. Keratins are fibrous proteins with a high degree of cross-linking hydrophobic interactions, disulfide, and hydrogen bonds. The supercoiling and condensed aggregation of proteins, i.e., α-keratins and β-keratins, are responsible for the stability and toughness of keratins. The α-keratins in wool and hair consist of complex supercoiling due to the presence of cysteine and tyrosine in α-helical chains (Daroit and Brandelli 2014), while β-keratins have β-sheets, and 38% of these keratins present in feathers (Rouse and Van Dyke 2010). In addition, keratins are resistant to proteolytic degradation by typical proteases because of the great mechanical stability and disulfide bonds cross-linking of protein chains.
General proteases like papain, trypsin, and pepsin cannot degrade keratin into its constituents due to the polymer’s rigid and complex structure (Gopinath et al. 2015). Therefore, the application of microorganisms that produce keratinases (EC 3.4.21; 3.4.24; 3.4.99.11) would provide a sustainable alternative for the removal of keratin. Usually, keratinases contain serine in their active site and are called serine proteases or serine metalloproteases (Gurunathan et al. 2021). Due to their unique property of degrading keratinous waste, keratinases are an encouraging substitute for other proteases. Furthermore, keratinases degrade keratinous waste into industrially valuable products like livestock feed, bio-hydrogen, bio-composite films, and nitrogenous fertilizers; thus; waste material can be converted into valuable by-products (Tamreihao et al. 2019). Therefore, keratinase are commonly known as “green chemicals” due to their eco-friendly nature (Akram et al. 2021).
Currently, more than 30 classes of microorganisms, including bacteria (e.g., actinomyces) and fungi, have been described to have effective keratinolytic properties (He et al. 2018). However, keratinases produced by mesophilic bacteria are mostly pathogenic and are not suitable for pilot-scale applications (Gupta and Ramnani 2006). In contrast, those microbial keratinases which withstand high temperature, pH, and harsh environmental conditions can degrade complex structure of proteins like nails, azokeratin, stratum corneum of eyes, and elastin. Most of these noteworthy keratinase-producing microbes reside on feathers and can be isolated from the surface of feathers (Javůrková et al. 2019). Therefore, thermostable keratinases are suitable for degrading complex keratinous waste because of ameliorated stability and elevated reactivity under extreme conditions. However, low enzyme yield and prolonged fermentations are the critical factors in determining the effectiveness of these keratinases. Recently, it has been observed that the genome sequencing might play a significant role in determining the potential of thermophilic microorganisms, but the detailed mechanism is still unclear (Kang et al. 2021).
Moreover, thermostable microbial keratinases are industrially significant enzymes have been extensively used in various industrial processes (leather, detergent, organic fertilizer, animal feed, textile, cosmetics, and pharmacy, etc.) due to their unique property to withstand harsh processing conditions (Akram et al. 2020, 2021; Hassan et al. 2020a). However, it is believed that the production of microbial keratinases in wild microbes can be enhanced by employing various protein engineering techniques (Fang et al. 2019). Similarly, the hydrolytic efficiency of microbial keratinases can be increased by the metabolic engineering of living microbes (Peng et al. 2020). This review presents an overview on microbial keratinase and their biochemical characteristics. Here, various metabolic and genetic engineering approaches including promoter and propeptide engineering, chromosomal integration, signal peptide, codon optimization, and glycoengineering are discussed to improve the stability and catalytic efficiency of keratinase. It also emphasizes on keratinases feasible industrial and biotechnological applications in diverse disciplines.
Keratinases from different domains of life
Keratinases are omnipresent and produced by numerous actinomycetes, bacteria, and fungi. MEROPS database provides the amino acid sequence and conserved domains among keratinases from different microbes and increase diversity among keratinases for their classification. Keratinolytic proteases are largely classified as serine or metalloproteases including S1, S8, S9, S10, S16, M3, M4, M14, M16, M28, M32, M36, M38, and M55 (Qiu et al. 2020). Figure 1 shows the keratinolytic activity of enzymes from above mentioned families. S8 family of proteases contain more than half of currently characterized keratinases but all of them cannot degrade keratin, while a keratinolytic strain, i.e., Streptomyces pactum DSM 40,530 produces a combination of serine proteases and effectively degrade insoluble substrates like keratins (Böckle et al. 1995). Serine proteases are produced by numerous microbial strains such as Bacillus pseudofirmus (Raval et al. 2014), Blakeslea trispora, Arthrobacter sp., Streptomyces sp., Flavobacterium sp., Bacillus licheniformis, Saccharomyces cerevisiae, Conidiobolus sp., Neurospora sp., and Aspergillus sp. (Arya et al. 2021), and Coprothermobacter proteolyticus (Toplak et al. 2013). Based on sequence and structural homology, most keratinases are reported as subtilisin-like protease. Intermittently, keratinases from Nocardiopsis sp. TOA-1B and S. albidoflavus are structurally categorized as chymotrypsin-like protease (Su et al. 2020).
Keratin biodegradation by various keratinolytic enzymes (modified from Qiu et al. 2020)
However, with ease in production and specific properties to degrade keratin effectively, keratinases from microbial sources are getting immense attention for large-scale production (Hamiche et al. 2019). Thermophilic strains (Table 1) with keratinase production, i.e., Thermoanaerobacter sp. and Clostridium sporogenes, have been isolated from volcanic areas, geothermal vents, hot springs, and solfataric muds. Furthermore, some alkaliphilic strains, i.e., Nocardiopsis sp. TOA-1 and Nesternkonia sp., have been reported for keratinase production in alkaline pH (Brandelli et al. 2010). Keratinase thus produced from these types of bacteria can exhibit a wide pH (5.8–11) and temperature (28–90 °C) stability, making the use of this enzyme in harsh industrial conditions possible (Tamreihao et al. 2019). Fervidobacterium pennivorans also produced keratinases which are stable at high temperature and alkaline pH but it is not clear which structural unit of enzyme is responsible for their stability in harsh conditions (Kim et al. 2004).
Besides, keratinase from an archaea Desulfurococcales is stable at pH 6 and 70 °C and Vibrio is gram-negative thermophilic actinobacteria which can also degrade keratin by producing thermostable keratinases (Nnolim and Nwodo 2020). Moreover, thermostable keratinolytic enzymes have also been reported from several Bacillus sp. like B. licheniformis, B. cereus, and B. subtilis (Huang et al. 2020). Keratinases from Bacillus spp. are either serine proteases or metalloproteases (Brandelli et al. 2010). B. subtilis KD-N2 is efficient in feather degradation after B. licheniformis RG1 which degrades feathers efficiently in 24 h (Ramnani and Gupta 2004). Furthermore, some salt-tolerant and thermophilic keratinases have also been reported from several bacterial groups that mainly include actinomyces.
Numerous actinomycetes have been also reported for the production of keratinolytic enzymes (Wang et al. 2015) and Thermoactinomyces sp. strain YT06 from poultry compost produced keratinase that completely degrade chicken feathers at 60 °C with the formation of 3.24 mg/mL amino acids from the culture medium (50 mL) containing chicken feather (10 g/L) (Wang et al. 2017). Under natural settings, keratinolytic fungi recycle the carbon, nitrogen, and sulphur of the keratins. Their prevalence and distribution appear to be influenced by the availability of keratin, particularly in areas where humans and animals exert substantial selective pressure on the environment (Brandelli et al. 2010). Majorly, keratinolytic fungi include species of Acremonium, Aphanoascus, Aspergillus, Chrysosporium, Cladosporium, Doratomyces, Fusarium, Lichtheimia, Microsporum, Paecilomyces, Scopulariopsis, Trichoderma, and Trichophyton (Alwakeel et al. 2021). A rich diversity of keratinolytic enzymes were produced by dermatophytic species of fungus and saprotrophic species growing on hooves and feathers, i.e., Onygena corvina (Mercer and Stewart 2019).
Molecular and biochemical properties of keratinases
Keratinase commercialization has opened new opportunities to identify keratinolytic microbes and produce keratinase on a pilot scale. Multiple proteases are required to degrade complex keratin material as different enzymes work on different sites of material and degrade into respective amino acids. Keratinase in general consist of catalytic domains, signal peptide, N-terminal pro-peptide, and C-terminal extension (Wu et al. 2017). Keratinase from Deinococcus gobiensis I-0 (DgoKerA) consist of N-propeptide, a mature region having a catalytic site, and signal peptide (Meng et al. 2022). The N-propeptide of keratinase functions as an intramolecular chaperone to facilitate the folding of keratinase and is then cleaved by mature keratinases for the effective function of this enzyme.
Nevertheless, thermostable keratinases from the Bacillus genus have been reported to effectively degrade keratin (Kshetri et al. 2020). Keratinase produced by Bacillus subtilis consists of two domains, i.e., the first domain has 59 amino acids (from 19 to 77 amino acids sequence) and encodes for inhibitor-I9; the second domain has 243 amino acids (from 103 to 345 amino acids sequence) and encodes peptidase S8. The enzyme also has a calcium ion metal-binding site and indicate calcium ions as their cofactors for keratinase from this bacterium (Haq and Akram 2018). S8 family of proteases consist of both α and β structures, as confirmed by crystal structures of rMtaKer from Meiothermus taiwanensis WR-220 and Fervidolysin from Fervidobacterium pennivorans (Li 2021). In spite of different folding structures, these enzymes have a catalytic triad for effective degradation of peptide bonds and have Ser, Asp, and His residues in them. Moreover, the molecular basis of protease and substrate interactions can also be studied by rMtaKer structure (Wu et al. 2017).
Distinct protease families have different structures of proteases as the M32 family of proteases have altered secondary structures, i.e., the crystal structure of protease FisCP from Fervidobacterium islandicum AW-1 have a short β-sheet near the active site with main helical structures. The active site has a Co2+ metal-binding site with His253, Glu254, His257, and Glu283 amino acids for effective substrate binding (Lee et al. 2015). A significant number of microbial strains generate monomeric enzymes but multimeric keratinases have also been reported. Usually, metalloproteases and the enzymes produced from thermophiles have high molecular mass (Bernal et al. 2006). High molecular weight keratinases have been identified in Bacillus cereus 1268 (∼200 kDa), Fervidobacterium islandicum (200 kDa), and Kocuria rosea (240 kDa) (Mazotto et al. 2011). Furthermore, microbial keratinases possess exclusive cleave preferences as Thermoanaerobacter sp. produces keratinase which have four cleavage sites located between Phe24-Phe25, Leu15-Tyr-16, Leu11-Val12, and Cys7-Gly8 (Kublanov et al. 2009). Therefore, keratinase prefer hydrophobic amino acids, i.e., leucine, valine, and phenylalanine at the catalytic site for keratinase degradation.
Most of the keratinolytic enzymes show active catalysis within a range of neutral to basic pH, i.e., 7 and 9. Some keratinases are highly alkaline in nature that work optimum at a pH of 10 and 13 (Nnolim and Nwodo 2020). Bacillus megaterium produce keratinase that show its catalytic activity at a broad range of pH 7 to 11 (Park and Son 2009). Alkali-tolerant keratinases produced by Bacillus pumilis and Microcuccus luteus exhibit the maximum catalytic efficiency at pH 9 to 11.5 (Laba et al. 2015). Mostly, microbial keratinases show their maximum catalytic activity at an optimum temperature between 37 to 65 °C, but some isolated from thermophilic microorganisms work best at 70 to 100 °C (Haq and Akram 2018). The thermotolerant strains such as Streptomyces gulbargensis produce keratinase that exhibits the maximum enzyme activity at 45 to 90 °C (Syed et al. 2009). Keratinases produced from mesophilic microorganisms such as Stenotrophomonas maltophila shows the optimum catalytic efficiency at 40 °C (Cao et al. 2009). Keratinases belonging to the group of metalloproteases require metal ions for their catalytic activity. They may require either one or two metallic ions such as zinc (Zn2+), cobalt (Co2+), and magnesium (Mg2+) for optimum activity (Supuran et al. 2002). Some metallic ions like calcium (Ca2+), cadmium (Cd2+), manganese (Mn2+), and sodium (Na+) enhance the catalytic efficiency of thermostable keratinases (Akram et al. 2020, 2021), but some cations such as cupper (Cu2+), silver (Ag2+), and mercury (Hg2+) act as inhibitors for keratinase activity (Pawar et al. 2018).
Factors affecting thermostability and catalytic activity of keratinases
Thermostability in keratinase can contribute to its utilization for myriad of industrial sectors and several facets like random genetic drift, extensive ionic network, and additional surface charge might be responsible for inducing thermostability in proteases. It was also observed that extra stability to protein structure at high temperature was provided by a decrease of salt bridge desolation at high temperature and resist conformational changes in protein structure (Tiberti and Papaleo 2011). Divalent cations act as a cofactor for microbial keratinase and are responsible for the thermostability of these enzymes. These metal ions might act as ion or salt bridges to stabilize the conformational folding of keratinase or act as an ion to stabilize the enzyme–substrate interaction for effective catalysis (Tork et al. 2013). Thermostable metalloprotease from Bacillus subtilis KT004404 demonstrate increased catalytic efficiency and stability in the presence of Zn2+ cation, making this enzyme a member of the zinc-dependent metalloprotease family (Rehman et al. 2017). Metal ions also enhance the substrate hydrolysis by stimulating water molecules to increase the nucleophilic attack during catalysis (Wu and Chen 2011). In addition, experimental analysis has revealed that subtilases (subtilisin superfamily of proteases) contain Ca2+ ion binding sites that play a significant role in stabilizing enzyme against auto-degradation. A large number of serine proteases carry calcium binding site in their autolysis loop that majorly stabilize these enzymes by providing a more compact structure to the enzyme (Fakhfakh et al. 2009). Apart from metal ion binding sites, aliphatic residues also play an important role in defining the thermostability of keratinases as the number of these residues increase the aliphatic index of the protein. Keratinase from Bacillus sp. Nnolim-K1 have a high aliphatic index and this enzyme is highly thermostable (Nnolim et al. 2020).
Keratinases can degrade complex substances in the presence of reducing agents as keratin can easily be hydrolyzed in the presence of reducing reagents during in vitro conditions (Pandey et al. 2019). Keratinase produced from Thermoactinomyces sp. RM4 appeared to be organic solvent tolerant, thermostable, and these bacteria produce keratinase in culture medium in the presence of organic solvents (Verma et al. 2016). Bacillus licheniformis produces thermotolerant and stable keratinase, and its activity was increased by 6.25-folds in the presence of mercaptoethanol in the reaction mixture (Tiwary and Gupta 2010). Usually, low concentrations of reducing agents can increase keratinolytic activity, mainly because they contribute to the sulfitolysis of the keratin substrate. Cleavage of disulfide bonds facilitates keratinase access to degrade keratin. In nutshell, keratinase belonging to the metallo or serine protease family have a wide range of pH, temperature, and substrate stability with efficient activation in the presence of thiol reagents.
Fermentative biosynthesis of thermostable keratinases
Due to the extreme shortage of protein resources, scientists are working hard to find economical and sustainable alternatives to produce new functional materials in an environmental friendly conduit. As a by-product of various meat and poultry processing operations, keratin is a valuable but difficult-to-recycle fibrous protein (Wang et al. 2016). Currently, the identification of feed additives for the replacement of expensive fishmeal requires acidic or alkaline hydrolysis of keratin to release amino acids (Kornillowicz-Kowalska and Bohacz 2010) These methods do not address the problems of process complexity, energy expenditure and poor product absorption, as well as the utilization of organic reagents. Except for a few thermophilic bacteria and fungi, keratinase has been produced under submerged shaking conditions (da Gioppo et al. 2009). With chicken feathers as sole carbon and nitrogen source, Bacillus sp. MD24 has also been reported to produce keratinase via submerged fermentation (SmF). However, this method appeared to be an ineffective due to its requirement for substantial quantity of water and involves the inadequate degradation of chicken feather (Andriyani et al. 2021).
Recently, solid state fermentation (SSF) has emerged as a critical method for the synthesis of thermostable keratinases (Table 2), owing to its high productivity, particularly when microorganisms are grown on insoluble substrates, and high concentration of end product (Nurkhasanah and Suharti 2019). The critical element affecting microbial growth and product output in SSF systems is the initial moisture content of the substrate (Chitturi and Lakshmi 2016). Mazotto et al (2013) indicated that SSF might be used to investigate the production of keratinases by A. niger, particularly with the 3T5B8 strain, which demonstrated a sevenfold increase in keratinolytic activity in SSF compared to SmF. In keeping with these findings, De Azeredo et al (2006) revealed that Streptomyces sp. 594 produced twice as much keratinase in SSF than it did in SmF. As a result, the fermentation environment has a significant impact on the production of extracellular enzymes. However, the benefits of SSF over SmF are not entirely known, and so this process should be given further consideration (Belmessikh et al. 2013). Myriad of molecular strategies have been formulated to improve the thermostability and productivity of keratinases in this domain.
Metabolic engineering of living cells to enhance the hydrolytic ability of keratinase
Research has revealed that keratin can be broken down into peptides and amino acids by keratinases released by microbes (Reddy et al. 2017). Nevertheless, the efficacy of separated and purified keratinase to hydrolyze keratin is relatively low, halting keratinase applications in industrial sector. The study of keratinase appeared to be entangled until it was discovered that sulphite can significantly improve hydrolytic ability of keratinases (Peng et al. 2020). This indicates that keratinase may require substantial factors from the environment or living cells in order to accomplish the hydrolysis.
For the biogenesis of Fe–S clusters, the sulfur formation (Suf) mechanism of most extremophilic anaerobes is essential. Additionally, the Suf system may play a crucial part in the degrading of keratin by Fervidobacterium islandicum AW-1, as well as its vital involvement in the redox chemistry and stress responses of Fe–S cluster proteins. There is a full Suf-like machinery (SufCBDSU) that is significantly expressed in cells growing on native feathers in the absence of elemental sulphur (S0) in the order Thermotogales, according to comparative genomics of the order Thermotogales (Jin et al. 2021). Sulfite is essential to activate the catalytic procedure of keratinase, and cysteine acts as a connection for live cells to govern this process, as recently disclosed by Peng et al (2021). Cysteine catabolism results in the production of sulfite as a by-product, which is released to help keratinase in hydrolysis of keratin. Cysteine produced during keratin hydrolysis enters the live cell and continues to be catabolized into sulfite as it enters the living cell (Fig. 2). The two processes must be linked in a chain reaction to ensure that keratin hydrolysis is enough. This self-circulation synergistic catalysis mechanism has also been shown to dramatically boost the keratin hydrolysis capability of cells that secrete keratinase. It may be summarized as follows: this work found a critical link in the keratin hydrolysis chain and produced modified strains that had a higher hydrolysis capability. Transcriptomic study of a feather degrading bacterium was recently published. Sulfite metabolism pathway genes have been upregulated in Streptomyces SCUT-3 (Li et al. 2020).
Degradation of keratin by cysteine mediated self-circulation cascade. Cysteine is a by-product of keratin. After being taken up by the cell, it is converted into sulfite by the enzymatic action of cysteine dioxygenase (Cdo1) and aspartate amino transferase (Ast1). The transportation of cysteine and sulfite is usually controlled by transporter (ydeD and SSU1) genes (modified from Peng et al. 2021)
Codon optimization strategy to increase production of keratinases
Codon optimization is a genetic strategy for optimizing the expression of a foreign gene in a host cell system. It is accomplished by substituting a specie’s current codons with a set of more appropriate host codons. Numerous studies indicate that codon biases lower metabolic burden by reducing the variability of isoacceptor tRNAs, hence improving heterologous gene expression in the host (Yahaya et al. 2021). A two-codon optimization technique is used to increase output of keratinase (kerA) from Bacillus licheniformis S90 and its expression in Pichia pastoris in order to enhance enzyme output compared to preparations using the native kerA gene. The recombinant keratinase was capable of degrading both α-keratin (azure keratin) and β-keratin (chicken feather meal) effectively and remained optimally active at pH 7.5 and 50 °C. These characteristics make P. pastoris pPICZA-kerAopti1 an attractive choice for commercial keratinase production (Hu et al 2013).
Promoter engineering to enhance the expression of keratinases
Because induction of keratinase using the native promoter is still inadequate for large-scale demands, the choice of promoter is also critical in providing increased expression of recombinant keratinase in Bacillus sp. (Nnolim et al. 2020). Promoters such as aprE, sigX, and srfA conferred substantial expressions of recombinant keratinase in Bacillus sp. by 10-–16-folds compared to the native control (Gong et al. 2020). A second approach would be to modify the Pgrac promoter to be auto-inducible so that recombinant keratinase may be initiated in the presence of a particular signal from the bacterial host (Thi et al. 2020). The quorum-sensing competence signal was activated by codon optimization of the PsrfA during the late log phase of Bacillus sp. growth. Having PsrfA eliminate the redundant sequence enabled the corresponding signal molecule to bind more firmly to the promoter, which resulted in a stronger activation of the promoter.
Chromosomal integration to enhance keratinase production
This technique involves the insertion of desired keratinase genes into bacterial plasmid or chromosome to enhance the expression, production and catalytic efficiency of keratinase protein. The kerA gene encodes for feather degrading keratinase isolated from B. licheniformis PWD-1 and integrated in an expression vector P43-pUB18 followed by propagation in B. subtilis DB104 to enhance the production of keratinase, but it decreased the stability of enzyme (Lin et al. 1997). A pLAT vector of Bacillus was also used to integrate the promoter and kerA gene the in chromosome of B. licheniformis T399D. In order to produce a keratinase variant with better degradation ability, kerA gene isolated from B. licheniformis ATCC 53,757 was combined with SPlip gene of pHIS1525 plasmid (B. megaterium) to create a SPK gene cassette. Finally, this was cloned in pMUTIN-GFP + vector and used to transform B. megaterium ATCC 14,945 (Jalendran and Baygi 2011).
Glycoengineering to maximize keratinase production
Glycoengineering means the enhancement of polypeptides (enzymes or/proteins) properties by glycosylation. This technique involving improved post translational modification processes is used to increase the keratinase production in P. pastoris (Karbalaei 2020). This strategy is mainly for eukaryotes due to high efficiency of glycosylation process and presence of glycosyltransferases which is not a feature of prokaryotes. Keratinase isolated from B. licheniformis MKU3 containing 5 N-glycosylation sites was glycosylated during expression in P. Pastoris X33 which showed noteworthy temperature and pH stability (Radha and Gunasekaran 2009). Same keratinase was non-glycosylated during expression in B. megaterium MS941 showing decreased structural stability and thermostability (Radha and Gunasekaran 2009). On the other hand, de-glycosylation can also be used to enhance keratinase production. Keratinase (having 4 N-glycosylation sites) isolated from Pseudomonas aeruginosa was not glycosylated during expression in P. pastoris, and even showed better production and thermostability. This means glycosylation sites present in recombinant keratinase of P. pastoris should either be glycosylated or de-glycosylated to maximize the keratinase production (Juturu and Wu 2018).
Molecular approaches to enhance keratinase efficiency
Keratinase, in particular, has shown considerable promise in catalysing keratin hydrolysis and is therefore regarded as a suitable biocatalyst for the conversion of keratin waste (Verma et al. 2017). However, the present usage of keratinase to hydrolyse feathers is hampered by low enzyme activity and limited processing capacity; consequently, the real practical applicability of keratinase remains unknown. As a result, protein engineering is required to enhance the enzymatic capabilities of certain enzymes.
Rational protein engineering approach
Rational protein engineering strategies have emerged as a promising option for obtaining exceptional keratinase variants with better thermostability and activity for the breakdown of feather waste. Fang et al (2017) revealed that by employing site-directed mutagenesis on a keratinase variant of FDD from Stenotrophomonas maltophilia, two variants (Y94F and Y215F) demonstrated greater extracellular keratinolytic activity in Escherichia coli expression system. The C-terminal fusion of keratinase DDF led to the production of a new variant DDFD with the highest substrate selectivity and keratinolytic activity. Based on model structure analysis, site-directed mutagenesis and C-terminus fusion are exceptionally effective methods for acquiring novel keratinases, and they allow the commercialization of new keratinases.
ΔGu is the most significant of the protein thermodynamic properties, in addition to providing a general index of protein thermostability. To forecast the impact of mutations on ΔGu, several computer algorithms have been proposed, and ΔGu may be simulated using a bioinformatics approach such as the PoPMuSiC algorithm. PoPMuSiC algorithm is a fast web server to determine the thermodynamic stability changes induced by point mutations in proteins in less than a minute. It has an excellent prediction performance when compared to other metaheuristics techniques, with a correlation coefficient of 0.8 between predicted and measured stability changes in cross-validation, following the deletion of 10% outliers (Dehouck et al. 2011). Liu et al (2013) used the PoPMuSiC algorithm to estimate the folding free energy change (G) of amino acid substitutions to improve the thermostability of B. licheniformis BBE11-1 keratinase. Using the algorithm in conjunction with the molecular modification of homologous subtilisin, four amino acid changes (N122Y, N217S, A193P, and N160C) were introduced into the enzyme by site-directed mutagenesis, and mutant genes were produced in B. subtilis WB600. With an 8.6-fold increase in the t1/2 value at 60 °C, the quadruple mutant demonstrated synergistic or additive effects. The N122Y substitution also resulted in a 5.6-fold increase in catalytic efficiency relative to wild-type keratinase. These findings add to our understanding of keratinase’s thermostability and point to further potential industrial uses.
PPC-Domain exchange and truncation
It has been appeared that some enzymes such as amylases and dextranases have unnecessary amino acids or domains at C-terminus, whose truncation reveal improved catalytic efficiency and other enzymatic properties (Yang et al. 2013; Kim et al. 2011). Recently, few proteases including keratinases have appeared to carry pre-peptidase C-terminal (PPC) domains that are associated with keratinase maturation and their cleavage activate enzyme, since the PPC domain may participate in activation before being cleaved off after peptidase secretion. To form beta-strands, the PPC domain always contains a single alpha-helix packed against an antiparallel beta-sheet (Yan et al. 2009; Fang et al. 2016a). Ribitsch et al. (2010) reported that serine protease (HP70) from Xanthomonadales having C-terminal domain truncation appeared to be considered as a good contender for detergent industry. The keratinase from S. maltophilia (KerSMD) is renowned for its high activity and pH stability in keratin degradation (Fang et al. 2016b). However, in order to use these enzymes in industrial applications, catalytic efficiency and detergent tolerance must be enhanced. As a result, it was revealed that truncation of PPC domain in keratinase had no influence on alkaline stability but significantly reduced collagenase activity, indicating that it may be used in leather treatment. When compared to the wild type, the variants of KerSMD (V380, V370, and V355) appeared to be thermophilic, with 1.7-fold increase in keratinolytic efficiency at 60 °C. The variant V355 was obtained by truncating the whole PPC domain, which enhanced resistance to alkalinity, salt, chaotropic chemicals, and detergents.
Because the pre-peptidase C-terminal (PPC) domain is thought to be associated with thermostability and substrate specificity (Fang et al. 2016a), it has been predicted that modifying the PPC domain would affect the keratinase’s stability and catalytic efficacy. In order to break down feather debris, Fang et al (2016b) improved the catalytic effectiveness and thermostability of the keratinase KerSMD by substituting its N/C-terminal domains with those of a related protease, KerSMF. Replacement of both N- and C-terminal domains produced a more stable mutant protein with a Tm of 64.60 ± 0.65 °C and a half-life of 244.6 ± 2 min at 60 °C, while deletion of the C-terminal domain from KerSMD or KerSMF produced mutant proteins with significant activity under mesophilic conditions. These results suggest that the C-terminal domain and N-propeptide of the pre-peptidase are critical not only for substrate selectivity, proper folding, and thermostability but also for the enzyme’s potential to convert feather waste into feed additives.
Non-canonical amino acid-based approach
Protein engineering approaches such as rational design and directed evolution, for example, rely greatly on mutual mutagenesis of the 20 canonical amino acids (cAAs). The use of just 20 cAAs as building blocks limits the ability to enhance protein properties via protein engineering. As a result, ncAA-based protein engineering might lead to increased activity and stability, as well as novel activities (Pagar et al. 2021). Although cAA-based engineering has enhanced the enzymatic properties of keratinases, engineering keratinases with distinct ncAAs may considerably increase the capacity ability to handle enzymatic structure and function. Genetic code expansion is superior to other approaches for incorporating ncAAs into proteins because it can include diverse ncAAs site-specific and at multiple sites into target proteins in all living species (Chin 2014).
Pan et al (2021) used genetic code expansion to engineer KerPA (M4 family) from Pseudomonas aeruginosa with non-canonical amino acids (ncAAs). The triple Y21pBpF/Y70pBpF/Y114pBpF variant exhibited a 1.3-fold increase in activity, and this suggests that when alteration of ncAAs at several sites may have cumulative impact on the enzyme’s properties and combining these mutations in one version may enhance the enzyme’s properties drastically. The findings revealed that pBpF mutations at particular enzyme sites might fill gaps, generate new interactions, and modify the local structure of the enzyme’s active region. Yi et al (2020) studied the keratinase from B. licheniformis WHU (KerBL) and used ncAA-based proximity-triggered chemical crosslinking to increase its autolytic resistance and thermostability under decreasing circumstances. Two variants, N159C/Y260BprY and N159C/Y260BbtY, with improved keratinolytic activity, were found after screening a series of non-canonical amino acid (ncAA)-based variants generated by rational design. The findings showed that covalent bonds between BprY-Cys and BbtY-Cys in the N159C/Y260TAG variant could greatly reduce the long loop’s flexibility and fluctuations, proving that stabilizing loop regions may effectively increase protein stability. However, the biggest barrier to the widespread use of ncAAs in enzyme engineering and directed evolution is that the medium must be supplemented with exogenous, often costly ncAAs during protein expression, which could raise production costs (Pan et al. 2021). The biosynthesis of ncAA through metabolic pathway engineering and in situ genetic code expansion in host cells might eliminate the requirement for exogenous ncAA replenishment and save money.
Biotechnological applications of thermostable keratinase
Keratinases have significant uses in a variety of industries, including leather (tanneries) production, detergent formulation, pharmaceuticals, and biomedicine (Fig. 3), not only owing to their catalytic effectiveness, but also due to their cost-effective synthesis on a renewable resource (Srivastava et al. 2020).
Biofuel production from keratinases
Bioconversion of keratinous wastes can also lead to the production of biogas by anaerobic digestion of keratinase producing microorganisms. The anaerobic degradation of feathers result in the production of amino acids which are further converted into ammonia, carbon dioxide, hydrogen, and organic acids (de Menezes et al. 2021). Mostly, methane gas is produced from the degradation of keratinous substances. Keratinases from B. megaterium can be used to degrade chicken feathers for the production of methane gas (Forgács et al. 2011). Production of methane can be increased using a mixture of feather hydrolysates and manure by anaerobic digestion of keratinolytic microorganisms. The cloning of keratinase gene from B. licheniformis in B. megaterium is known to increase biodegradation of poultry feathers. This recombinant strain results in about 80% production of methane gas. The process of bio-gas formulation usually involves B. licheniformis KK1 that convert feather waste into hydrolysate rich in amino acids and peptides. This hydrolysate is then utilized by Thermococcus litoralis for bio-hydrogen production (Vidmar and Vodovnik 2018). Keratinases produced from strains of Clostridium can also be used to produce butanol from chicken feathers and wheat straw hydrolysates (Branska et al. 2020).
Other eminent applications of keratinases
Some agro-industry companies have developed an endergonic–mechanical process for converting feathers into feedstuffs (feather meal) to add economic value to the protein-rich chicken feathers (Khumalo et al. 2020). Feather hydrolysates produced from keratinolytic B. subtilis AMR have been used with maize meal to provide a high-amino-acid-content cattle feed (Mazotto et al. 2017). The animal feed produced from keratinolytic waste also accelerates animal growth and boosts the digestibility of animals. In this regard, production of commercial keratinase (Versazyme) from B. licheniformis PWD-1 has been employed to produce feed additive to enhance the digestibility of animals (de Menezes et al. 2021). Recently, keratinase has been used for removal of dag from livestock animals. Dag is composed of organic matter such as feces, hair, straw, and soil (Navone and Speight 2020). Keratinase breaks the interaction of dag with hair of livestock animals which is then completely removed after washing. This process will reduce the chances of animal meat from getting contaminated.
Nowadays, bio-additive-based laundry detergents are more preferred over synthetic detergents because of their cleaning properties like low-temperature washing, removal of recalcitrant dirt, and maintenance of cloth fibers. Alkaline proteases are also used in detergent formulations to remove proteinaceous stains. For example, one of the alkaline keratinases, isolated from Paenibacillus woosongensis TKB2, is used in the laundry industry to remove the stains from the clothes without affecting the texture of fabric (Paul et al. 2013). Microbial keratinases are added in both liquid and solid detergent formulations because they remain stable even in presence of surface-active agents. Thermostable keratinases isolated from B. subtilis (Paul et al. 2016), Bacillus sp. NKSP-7 (Akram et al. 2020), and Bacillus sp. NDS-10 (Akram et al. 2021) have perceptible potential to remove the stains of blood from cotton blood-stained fabrics.
Keratinases are also used as biological pesticides. In this regard, keratinases produced from Bacillus sp. have appeared to kill Meloidogyne incognita which is a root-knot nematode (Yue et al. 2011). Apart from industrial applications, keratinases carry an utmost importance in pharmaceutical industry. Alzheimer’s disease (AD) is a common age-related neurological disease disorder characterized by the progressive degradation of cognitive abilities, resulting in poor performance in routine activities. β-sheet-rich, soluble amyloid-beta (Aβ1-42) oligomers have been established as the primary pathological hallmark of Alzheimer’s disease (Mroczko et al. 2018). The propensity of these oligomeric aggregates to produce insoluble amyloid deposits in the brain, especially in the cerebrum and hippocampus, is profoundly damaging. Rajput et al (2019) used amyloid-degrading keratinase (kerA) enzyme as a framework for identifying five keratinase-guided peptides (KgPs) capable of interacting with and changing Aβ1-42 amyloidogenic conversion. The KgPs exhibited a micromolar affinity for Aβ1-42 and inhibited its sigmoidal amyloidogenic transition, hence preventing fibrillogenesis. Overall, these latest findings provide a novel method for designing possible anti-amyloid molecules, which might open the way for the development of effective therapies for Alzheimer’s disease and other amyloid illnesses. They have been recognized for various other medical applications such as drug delivery system, skin ailments (psoriasis and acne) treatment, earwax removal, prion degradation, transfer accelerators, human callus removal, against dermatophytosis, and fungal infections (onychomycosis) (Vidmar and Vodovnik 2018; Haq and Akram 2018).
Thermostable microbial keratinases have been prominently used in leather and tannery industry for dehairing the hides of animals which are the best alternatives to the conventional injurious chemicals method. Recently, two thermostable keratinases from Bacillus sp. NKSP-7 (Akram et al. 2020), and Bacillus sp. NDS-10 (Akram et al. 2021) effectively dehaired goat’s hides deprived of any damage after 8 h and 6 h, respectively. It has proved them notable environmental friendly candidates for hide-depilation in leather industries. Keratinolytic enzyme are also employed for the bioprocessing of agro-industrial wastes, and wastewater treatment (Verma et al. 2017). Moreover, keratinase are extensively used in the manufacturing of bio-organic fertilizers, cosmetic products (hair-removing lotions and creams, etc.); protein supplements; biodegradable films, foils, glue, and pearl bleaching; and processing of edible bird’s nests (Tamreihao et al. 2019; Akram et al. 2020; Hassan et al. 2020a).
Future perspectives
Keratin is the amplest recalcitrant protein on earth surface which can be efficiently degraded by keratinolytic microorganisms. Various microorganisms produce keratinase that has vast applications in industrial and pharmaceutical processes. Nevertheless, a combination of limiting features, such as a long fermentation duration and limited enzyme output, may jeopardize the commercial prospects of these keratinases. Despite the fact, many researchers have investigated various yield enhancement tactics at the laboratory scale to improve extracellular keratinase production by wild microorganisms. For instance, cloning and overexpression in relation to molecular optimization of keratinase expression in industrially appropriate hosts presage the potential to improve the productivity of wild microbial producers. Still, a number of challenges in proteomics, transcriptomic, metagenomic, and genetic engineering of keratinases exist. In addition to engineering C-terminal and pro-peptide domains of keratinases, the amalgamation of numerous mutation approaches may have a superposition effect on enzyme through cooperative action.
Furthermore, employing many advanced in silico forecasting and analytic methods, computer-aided rational or semi-rational alterations of keratinases will become an important topic of research. A large number of microbial enzymes have been discovered from decomposition sites of keratinous biomass. But little is known about the action of non-keratinase metabolites (e.g., lipases and cellulases) in mechanistic keratin degradation. In this regard, investigating the role of these biocatalyst may act as a viable approach in valorization of keratin-rich agro-industrial wastes by microbial enzyme cocktail strategy. Therefore, the need of an hour is to employ metabolic and protein engineering techniques to increase the yield and thermostability of keratinases. The keratinolytic activity of various microorganisms is considered as a green process throughout the world to deal with recalcitrant keratinous wastes. It is necessary to develop novel probe-based techniques to detect keratinase production from these keratinolytic microorganisms. Biosensors should also be developed for detecting novel keratinases. Most of the keratinases are produced from mesophilic microorganisms but the industrial demand is more for thermostable keratinases. So, novel thermophilic and halophilic microorganisms should be studied and focused to discover novel keratinolytic enzymes.
In addition to pharmaceutical and industrial applications, recently a novel keratinolytic specie of B. cereus has been discovered that can effectively carry out keratin hydrolysis and develop keratin-based bio-plastic films (Alshehri et al. 2021). The research revealed that keratin-based bio-plastics possess superior crystalline morphology over synthetic plastic but requires further investigations to substitute fossil oil-based materials. For effective industrial applications, optimization strategies should also be employed to increase the simultaneous yield of keratinases and bio-plastic films. Recently, keratinases have been used in waste-water treatment as they can be immobilized on bagasse cellulose to decolorize the molasses wastewater. Due to abundance of sulphur amino acids in keratin hydrolysate, they are efficiently involved to promote the growth of plants. Such green fertilizers are in high demand to produce chemical free fruits and vegetable crops.
Nowadays, research is being driven by the notion of poultry bio refinery — a method of managing and processing chicken manure by keratinases for energy and nutrient recovery as well as for the formulation of added value goods. However, more research is needed to validate and upscale this paradigm under a variety of economic conditions, farm typologies, and regulatory and environmental requirements. Recent studies have shown use of keratinase for prion decontamination which has greatly advanced in medical field regarding sterilization of surgical instruments. No green product other than keratinase is present in leather industry for safe dehairing of animal hides. However, there is no recognized product in the market for the green processing of hides and skins at the pilot scale. In this light, the development of effective keratinase-based dehairing products for industrial processing of hides will greatly minimize the environmental pollution and help to achieve the sustainable development goals. They are truly considered as green biocatalysts due to their promising applications so more research is required to find out novel varieties of keratinases having high catalytic efficiency and broad substrate specificity. Hence, thermo-efficient microbial keratinases have great potential for the development of greener, clean, and hygiene technology.
Conclusion
Keratinases are proteolytic enzymes which are mainly extracted from microbial sources. Due to their conceivable applications, they are widely produced at commercial level. The key function of keratinase is to degrade “keratin” protein into valuable bio-products. Keratin containing substrates are easily available as a waste material in our environment, which are utilized for the production of keratinases. Keratinolytic enzymes originated from microbial sources (both from bacteria and fungi), are considered as worthwhile green biocatalysts, which are able to do biotransformation of keratin rich raw materials into valuable and beneficial products, although the enzyme can be produced by fermentation techniques (both SmF and SSF). Various protein and metabolic engineering techniques have recently been applied to increase the thermostability, productivity and activity of microbial keratinases. Due to the vast industrial, biotechnological, and pharmacological applications, the enzyme has been used in almost every industry at commercial level. In addition, use of keratinases for skin (acne) and nails treatment, prion degradation, wastewater treatment, and pearl bleaching are novel applications. This enzyme is available in limited quantity whereas their demand is globally high. In view of this, more emphasis should be made on research regarding screening of novel keratinases and their easy and manageable production strategies to meet the high demand of industrial sector.
Data availability
All data generated or analyzed during this study are included in this published article [and its supplementary information file].
References
Abu-Tahon MA, Arafat HH, Isaac GS (2020) Laundry detergent compatibility and dehairing efficiency of alkaline thermostable protease produced from Aspergillus terreus under solid-state fermentation. J Oleo Sci 69(3):241–254. https://doi.org/10.5650/jos.ess19315
Akram F, Haq IU, Jabbar Z (2020) Production and characterization of a novel thermo-and detergent stable keratinase from Bacillus sp. NKSP-7 with perceptible applications in leather processing and laundry industries. Int J Biol Macromol 164:371–383. https://doi.org/10.1016/j.ijbiomac.2020.07.146
Akram F, Haq IU, Hayat AK, Ahmed Z, Jabbar Z, Baig IM, Akram R (2021) Keratinolytic enzyme from a thermotolerant isolate Bacillus sp. NDS-10: An efficient green biocatalyst for poultry waste management, laundry and hide-dehairing applications. Waste Biomass Valor 12:5001–5018. https://doi.org/10.1007/s12649-021-01369-2
Alshehri WA, Khalel A, Elbanna K, Ahmad I, Abulreesh HH (2021) Bio-plastic films production from feather waste degradation by keratinolytic bacteria Bacillus cereus. J Pure Appl Microbiol 15:681–688. https://doi.org/10.22207/JPAM.15.2.17
Alwakeel SS, Ameen F, Al Gwaiz H, Sonbol H, Alghamdi S, Moharram AM, Al-Bedak OA (2021) Keratinases produced by Aspergillus stelliformis, Aspergillus sydowii, and Fusarium brachygibbosum isolated from human hair: yield and activity. J Fungi (basel, Switzerland) 7(6):471. https://doi.org/10.3390/jof7060471
Andriyani A, Wongkar FT, Suharti S (2021) Production of keratinase under solid-state fermentation (SSF) by Bacillus sp. MD24 and potential of its liquid by-product as organic fertilizers. International conference on life sciences and technology (ICoLiST 2020). https://doi.org/10.1063/5.0052647
Anitha TS, Palanivelu P (2013) Purification and characterization of an extracellular keratinolytic protease from a new isolate of Aspergillus parasiticus. Protein Expr Purif 88(2):214–220. https://doi.org/10.1016/j.pep.2013.01.007
Arya PS, Yagnik SM, Rajput KN, Panchal RR, Raval VH (2021) Understanding the basis of occurrence, biosynthesis, and implications of thermostable alkaline proteases. Appl Biochem Biotechnol 193(12):4113–4150. https://doi.org/10.1007/s12010-021-03701-x
Belmessikh A, Boukhalfa H, Mechakra-Maza A, Gheribi-Aoulmi Z, Amrane A (2013) Statistical optimization of culture medium for neutral protease production by Aspergillus oryzae. Comparative study between solid and submerged fermentations on tomato pomace. J Taiwan Inst Chem Eng 44:377–385. https://doi.org/10.1016/j.jtice.2012.12.011
Ben Elhoul M, Jaouadi ZN, Bouacem K, Allala F, Rekik H, Mechri S, Ezzine KH, Miled N, Jaouadi B (2021) Heterologous expression and purification of keratinase from Actinomadura viridilutea DZ50: feather biodegradation and animal hide dehairing bioprocesses. Environ Sci Pollut Res 28(8):9921–9934. https://doi.org/10.1007/s11356-020-11371-1
Bernal C, Cairó J, Coello N (2006) Purification and characterization of a novel exocellular keratinase from Kocuria Rosea. Enzyme Microb Technol 38(1–2):49–54. https://doi.org/10.1016/j.enzmictec.2005.02.021
Bhange K, Chaturvedi V, Bhatt R (2016) Simultaneous production of detergent stable keratinolytic protease, amylase and biosurfactant by Bacillus subtilis PF1 using agro industrial waste. Biotechnol Rep 10:94–104. https://doi.org/10.1016/j.btre.2016.03.007
Böckle B, Galunsky B, Müller R (1995) Characterization of a keratinolytic serine proteinase from Streptomyces pactum DSM 40530. Appl Environ Microbiol 61(10):3705–3710. https://doi.org/10.1128/aem.61.10.3705-3710.1995
Bokveld A, Nnolim NE, Nwodo UU (2021) Chryseobacterium aquifrigidense FANN1 produced detergent-stable metallokeratinase and amino acids through the abasement of chicken feathers. Front Bioeng Biotechnol 9:720176. https://doi.org/10.3389/fbioe.2021.720176
Brandelli A, Daroit DJ, Riffel A (2010) Biochemical features of microbial keratinases and their production and applications. Appl Microbiol Biotechnol 85:1735–1750. https://doi.org/10.1007/s00253-009-2398-5
Branska B, Fořtová L, Dvořáková M, Liu H, Patakova P, Zhang J, Melzoch M (2020) Chicken feather and wheat straw hydrolysate for direct utilization in biobutanol production. Renew Energy 145:1941–1948. https://doi.org/10.1016/j.renene.2019.07.094
Cao ZJ, Zhang Q, Wei DK, Chen L, Wang J, Zhang XQ, Zhou MH (2009) Characterization of a novel Stenotrophomonas isolate with high keratinase activity and purification of the enzyme. J Ind Microbiol Biotechnol 36(2):181–188. https://doi.org/10.1007/s10295-008-0469-8
Chaya E, Suzuki T, Karita S, Hanya A, Y-Yasuda S, Kitamoto N (2014) Sequence Analysis and heterologous expression of the wool cuticle-degrading enzyme encoding genes in Fusarium oxysporum 26–1. J Biosci Bioeng 117(6):711–714. https://doi.org/10.1016/j.jbiosc.2013.11.012
Chen X, Zhou B, Xu M, Huang Z, Jia G, Zhao H (2015) Prokaryotic expression and characterization of a keratinolytic protease from Aspergillus niger. Biologia 70(2):157–164. https://doi.org/10.1515/biolog-2015-0031
Chimbekujwo KI, Ja’afaru MI, Adeyemo OM (2020) Purification, characterization and optimization conditions of protease produced by Aspergillus brasiliensis strain BCW2. Sci Afr 20:8. https://doi.org/10.1016/j.sciaf.2020.e00398
Chin JW (2014) Expanding and reprogramming the genetic code of cells and animals. Annu Rev Biochem 83:379–408. https://doi.org/10.1146/annurev-biochem-060713-035737
Chitturi Ch, Lakshmi VV (2016) Development of semi-solid state fermentation of Keratinase and optimization of process by cheaper and alternative agricultural wastes. European J Biotechnol Biosci 4:2321–9122
da Silva RR (2018) Keratinases as an alternative method designed to solve keratin disposal on the environment: its relevance on agricultural and environmental chemistry. J Agric Food Chem 66:7219–7221. https://doi.org/10.1021/acs.jafc.8b03152
da Gioppo NM, Moreira-Gasparin FG, Costa AM, Alexandrino AM, de Souza CG, Peralta RM (2009) Influence of the carbon and nitrogen sources on keratinase production by Myrothecium verrucaria in submerged and solid-state cultures. J Ind Microbiol Biotechnol 36(5):705–711. https://doi.org/10.1007/s10295-009-0540-0 (PMID: 19229574)
Dagnaw M, Andualem B (2019) Solid state fermentation of keratinolytic proteases production using Bacillus spp. isolated from hair and mud sample of traditional leather processing ponds in North Gondar, Ethiopia. J Med Plants Stud 7:127–138
Damare S, Mishra A, D’Souza-Ticlo-Diniz D, Krishnaswamy A, Raghukumar C (2020) A deep-sea hydrogen peroxide-stable alkaline serine protease from Aspergillus favus. 3 Biotech 10(12):1–9. https://doi.org/10.1007/s13205-020-02520-x
Daroit DJ, Brandelli A (2014) A current assessment on the production of bacterial keratinases. Crit Rev Biotechnol 34(4):372–384. https://doi.org/10.3109/07388551.2013.794768
De Azeredo LA, De Lima MB, Coelho RR, Freire DM (2006) Thermophilic protease production by Streptomyces sp. 594 in submerged and solid-state fermentations using feather meal. J Appl Microbiol 100(4):641–7. https://doi.org/10.1111/j.1365-2672.2005.02791.x (PMID: 16553718)
de Menezes CLA, Santos RDC, Santos MV, Boscolo M, da Silva R, Gomes E, da Silva RR (2021) Industrial sustainability of microbial keratinases: production and potential applications. World J Microbiol Biotechnol 37(5):86. https://doi.org/10.1007/s11274-021-03052-z
Dehouck Y, Kwasigroch JM, Gilis D, Rooman MP (2011) PMuSiC 2.1: a web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinformatics 12:151. https://doi.org/10.1186/1471-2105-12-151
Dhiva S, Ranjith KR, Prajisya P, Sona KP, Narendrakumar G, Prakash P, Emilin Renitta R, Samrot AV (2020) Optimization of keratinase production using Pseudomonas aeruginosa SU-1 having feather as substrate. Biointerface Res Appl Chem 10:6540–6549. https://doi.org/10.33263/BRIAC105.65406549
Duffeck CE, de Menezes C, Boscolo M, da Silva R, Gomes E, da Silva RR (2020) Citrobacter diversus-derived keratinases and their potential application as detergent-compatible cloth-cleaning agents. Braz J Microbiol: [publication of the Brazilian Society for Microbiology] 51(3):969–977. https://doi.org/10.1007/s42770-020-00268-3
El-Ayouty YM, El-Said A, Salama AM (2012) Purification and characterization of a keratinase from the feather-degrading cultures of Aspergillus flavipes. Afr J Biotechnol 11:2313–2319. https://www.ajol.info/index.php/ajb/article/view/100603/89822
El-Gendy MM (2010) Keratinase production by endophytic Penicillium spp. Morsy1 under solid-state fermentation using rice straw. Appl Biochem Biotechnol 162(3):780–794. https://doi.org/10.1007/s12010-009-8802-x
Emran MA, Ismail SA, Abdel-Fattah AM (2020a) Valorization of feather via the microbial production of multi-applicable keratinolytic enzyme. Biocatal Agric Biotechnol 27:101674. https://doi.org/10.1016/j.bcab.2020.101674
Fakhfakh N, Kanoun S, Manni L, Nasri M (2009) Production and biochemical and molecular characterization of a keratinolytic serine protease from chicken feather-degrading Bacillus licheniformis RPk. Can J Microbiol 55(4):427–436. https://doi.org/10.1139/w08-143
Fang Z, Zhang J, Du G, Chen J (2016a) Improved catalytic efficiency, thermophilicity, anti-salt and detergent tolerance of keratinase KerSMD by partially truncation of PPC domain. Sci Rep 6:27953. https://doi.org/10.1038/srep27953
Fang Z, Zhang J, Liu B, Du G, Chen J (2016b) Enhancement of the catalytic efficiency and thermostability of Stenotrophomonas sp. keratinase KerSMD by domain exchange with KerSMF. Microbial Biotechnol 9(1):35–46. https://doi.org/10.1111/2F1751-7915.12300
Fang Z, Zhang J, Du G, Chen J (2017) Rational protein engineering approaches to further improve the keratinolytic activity and thermostability of engineered keratinase KerSMD. Biochem Eng J 127:147–153. https://doi.org/10.1016/j.bej.2017.08.010
Fang Z, Sha C, Peng Z, Zhang J, Du G (2019) Protein engineering to enhance keratinolytic protease activity and excretion in Escherichia coli and its scale-up fermentation for high extracellular yield. Enzyme Microb Technol 121:37–44. https://doi.org/10.1016/j.enzmictec.2018.11.003
Forgács G, Alinezhad S, Mirabdollah A, Feuk-Lagerstedt E, Sárvári HI (2011) Biological treatment of chicken feather waste for improved biogas production. J Environ Sci (china) 23:1747–1753. https://doi.org/10.1016/S1001-0742(10)60648-1
Gong JS, Ye JP, Tao LY, Su C, Qin J, Zhang YY, Li HH, Li HH, Xu ZH, Shi JS (2020) Efficient keratinase expression via promoter engineering strategies for degradation of feather wastes. Enzym Microb Technol 137:109550. https://doi.org/10.1016/j.enzmictec.2020.109550
Gopinath SC, Anbu P, Lakshmipriya T, Tang TH, Chen Y, Hashim U, Ruslinda AR, Arshad MK (2015) Biotechnological aspects and perspective of microbial keratinase production. Biomed Res Int. https://doi.org/10.1155/2015/140726
Gupta R, Ramnani P (2006) Microbial keratinases and their prospective applications: an overview. Appl Microbiol Biotechnol 70(1):21–33. https://doi.org/10.1007/s00253-005-0239-8
Gurunathan R, Huang B, Ponnusamy VK, Hwang JS, Dahms HU (2021) Novel recombinant keratin degrading subtilisin like serine alkaline protease from Bacillus cereus isolated from marine hydrothermal vent crabs. Sci Rep 11(1):12007. https://doi.org/10.1038/s41598-021-90375-4
Habbeche A, Saoudi B, Jaouadi B, Haberra S, Kerouaz B, Boudelaa M, Badis A, Ladjama A (2014) Purification and biochemical characterization of a detergent-stable keratinase from a newly thermophilic actinomycete Actinomadura keratinilytica strain Cpt29 isolated from poultry compost. J Biosci Bioeng 117(4):413–421. https://doi.org/10.1016/j.jbiosc.2013.09.006
Hamiche S, Mechri S, Khelouia L, Annane R, El Hattab M, Badis A, Jaouadi B (2019) Purification and biochemical characterization of two keratinases from Bacillus amyloliquefaciens S13 isolated from marine brown alga Zonaria tournefortii with potential keratin-biodegradation and hide-unhairing activities. Int J Biol Macromol 122:758–769. https://doi.org/10.1016/j.ijbiomac.2018.10.174
Haq IU, Akram F (2018) Striking applications of keratinase enzyme isolated from various natural sources: a review. Proc Pak Acad Sci: B. Life Environ Sci 55(4):1–17
Haq IU, Akram F, Jabbar Z (2020) Keratinolytic enzyme-mediated biodegradation of recalcitrant poultry feathers waste by newly isolated Bacillus sp. NKSP-7 under submerged fermentation. Folia Microbiologica 65(5):823–834. https://doi.org/10.1007/s12223-020-00793-6
Hassan MA, Abol-Fotouh D, Omer AM, Tamer TM, Abbas E (2020a) Comprehensive insights into microbial keratinases and their implication in various biotechnological and industrial sectors: A review. Int J Biol Macromol 1(154):567–583. https://doi.org/10.1016/j.ijbiomac
Hassan MA, Taha TH, Hamad GM, Hashem M, Alamri S, Mostafa YS (2020b) Biochemical characterisation and application of keratinase from Bacillus thuringiensis MT1 to enable valorisation of hair wastes through biosynthesis of vitamin B-complex. Inter J Biol Macromol 153:561–572. https://doi.org/10.1016/j.ijbiomac.2020.03.032
He Z, Sun R, Tang Z, Bu T, Wu Q, Li C, Chen H (2018) Biodegradation of feather waste keratin by the keratin-degrading strain Bacillus subtilis 8. J Microbiol Biotechnol 28(2):314–322. https://doi.org/10.4014/jmb.1708.08077
Hu H, Gao J, He J, Yu B, Zheng P, Huang Z, Mao X, Yu J, Han G, Chen D (2013) Codon optimization significantly improves the expression level of a keratinase gene in Pichia pastoris. PLoS ONE 8(3):e58393. https://doi.org/10.1371/journal.pone.0058393
Huang Y, Busk PK, Herbst FA, Lange L (2016) Genome and secretome analyses provide insights into keratin decomposition by novel proteases from the non-pathogenic fungus Onygena corvina. Appl Microbiol Biotechnol 99(22):9635–9649. https://doi.org/10.1007/s00253-015-6805-9
Huang Y, Łężyk M, Herbst FA, Busk PK, Lange L (2020) Novel keratinolytic enzymes, discovered from a talented and efficient bacterial keratin degrader. Sci Rep 10(1):10033. https://doi.org/10.1038/s41598-020-66792-2
Jalendran E, Baygi SJD (2011) Chromosomal integration of KerA gene in Bacillus megaterium for stable keratinase production. https://www.diva-portal.org/smash/get/diva2:1312301/FULLTEXT01.pdf
Javůrková VG, Kreisinger J, Procházka P, Požgayová M, Ševčíková K, Brlík V, Adamík P, Heneberg P, Porkert J (2019) Unveiled feather microcosm: feather microbiota of passerine birds is closely associated with host species identity and bacteriocin-producing bacteria. ISME J 13(9):2363–2376. https://doi.org/10.1038/s41396-019-0438-4
Jin HS, Dhanasingh I, Sung JY, La JW, Lee Y, Lee EM, Kang Y, Lee DY, Lee SH, Lee DW (2021) The sulfur formation system mediating extracellular cysteine-cystine recycling in Fervidobacterium islandicum AW-1 is associated with keratin degradation. Microb Biotechnol 14(3):938–952. https://doi.org/10.1111/1751-7915.13717
Juturu V, Wu JC (2018) Heterologous protein expression in Pichia pastoris: latest research progress and applications. ChemBioChem 19:7–21. https://doi.org/10.1002/cbic.201700460
Kalaikumari SS, Vennila T, Monika V, Chandraraj K, Gunasekaran P, Rajendhran J (2019) Bioutilization of poultry feather for keratinase production and its application in leather industry. J Clean Prod 208:44–53. https://doi.org/10.1016/j.jclepro.2018.10.076
Kang D, Huang Y, Nesme J, Herschend J, Jacquiod S, Kot W et al (2021) Metagenomic analysis of a keratin-degrading bacterial consortium provides insight into the keratinolytic mechanisms. Sci Total Environ 761:143281. https://doi.org/10.1016/j.scitotenv.2020.143281
Karbalaei M (2020) Pichia pastoris: a highly successful expression system for optimal synthesis of heterologous proteins. J Cell Physiol 2020:1–15. https://doi.org/10.1002/jcp.29583
Khumalo M, Sithole B, Tesfaye T (2020) Valorization of waste chicken feathers: optimization of keratin extraction from waste chicken feathers by sodium bisulphite, sodium dodecyl sulphate and urea. J Environ Manag 262:110329. https://doi.org/10.1016/j.jenvman.2020.110329
Kim JS, Kluskens LD, de Vos WM, Huber R, van der Oost J (2004) Crystal structure of fervidolysin from Fervidobacterium pennivorans, a keratinolytic enzyme related to subtilisin. J Mol Biol 335(3):787–797. https://doi.org/10.1016/j.jmb.2003.11.006 (PMID: 14687574)
Kim YM, Shimizu R, Nakai H, Mori H, Okuyama M, Kang MS, Fujimoto Z, Funane K, Kim D, Kimura A (2011) Truncation of N- and C-terminal regions of Streptococcus mutans dextranase enhances catalytic activity. Appl Microbiol Biotechnol 91(2):329–339. https://doi.org/10.1007/s00253-011-3201-y
Kornillowicz-Kowalska T, Bohacz J (2010) Dynamics of growth and succession of bacterial and fungal communities during composting of feather waste. Bioresour Technol 101:1268–1276. https://doi.org/10.1016/j.biortech.2009.09.053
Kshetri P, Roy SS, Chanu SB, Singh TS, Tamreihao K, Sharma SK, Ansari MA, Prakash N (2020) Valorization of chicken feather waste into bioactive keratin hydrolysate by a newly purified keratinase from Bacillus sp. RCM-SSR-102. J Environ Manage 273:111195. https://doi.org/10.1016/j.jenvman.2020.111195
Kublanov IV, Tsirul’nikov KB, Kaliberda EN, Rumsh LD, Haertle T, Bonch-Osmolovskaia EA (2009) Keratinase of an anaerobic thermophilic bacterium Thermoanaerobacter sp. strain 1004–09 isolated from a hot spring in the Baikal Rift zone. Mikrobiologiia 78(1):79–88
Kumar J, Mahal S (2021) Isolation of chrysosporium indicum from poultry soil for keratinase enzyme, its purification and partial characterization. J Appl Natural Sci 13(2):744–51. https://doi.org/10.31018/jans.v13i2.2609
Laba W, Choinska A, Rodziewicz A, Piegza M (2015) Keratinolytic abilities of Micrococcus luteus from poultry waste. Braz J Microbiol 46:691–700. https://doi.org/10.1590/2FS1517-838246320140098
Lee YJ, Dhanasingh I, Ahn JS, Jin HS, Choi JM, Lee SH, Lee DW (2015) Biochemical and structural characterization of a keratin-degrading M32 carboxypeptidase from Fervidobacterium islandicum AW-1. Biochem Biophys Res Commun 468(4):927–933. https://doi.org/10.1016/j.bbrc.2015.11.058
Li Q (2021) Structure, Application, and Biochemistry of Microbial Keratinases. Front Microbiol 12:674345. https://doi.org/10.3389/fmicb.2021.674345
Li ZW, Liang S, Ke Y, Deng JJ, Zhang MS, Lu DL, Li JZ, Luo XC (2020) The feather degradation mechanisms of a new Streptomyces sp. isolate SCUT-3. Commun Biol 3(1):191. https://doi.org/10.1038/s42003-020-0918-0
Lin X, Wong SL, Miller ES, Shih JC (1997) Expression of the Bacillus licheniformis PWD-1 keratinase gene in B. subtilis. J Ind Microbiol Biotechnol 19(2):134–138. https://doi.org/10.1038/sj.jim.2900440
Liu B, Zhang J, Fang Z, Gu L, Liao X, Du G, Chen J (2013) Enhanced thermostability of keratinase by computational design and empirical mutation. J Ind Microbiol Biotechnol 40(7):697–704. https://doi.org/10.1007/s10295-013-1268-4
Ma Y, Ke X, Li X, Shu W, Yang W, Liu Y, Yan X, Jia L, Yan H (2017) Expression and characterization of a keratinase encoding gene gm2886 in Streptomyces pactum ACT12 strain. Sheng wu gong cheng xue bao (Chinese J Biotechnol) 33:1968–1978. https://doi.org/10.13345/j.cjb.170081
Mazotto AM, de Melo AC, Macrae A, Rosado AS, Peixoto R, Cedrola SM, Couri S, Zingali RB, Villa AL, Rabinovitch L, Chaves JQ, Vermelho AB (2011) Biodegradation of feather waste by extracellular keratinases and gelatinases from Bacillus spp. World J Microbiol Biotechnol 27(6):1355–1365. https://doi.org/10.1007/s11274-010-0586-1
Mazotto A, Courib S, Damasod M, Vermelho A (2013) Degradation of feather waste by Aspergillus niger keratinases: comparison of submerged and solid-state fermentation. Int Biodeterior Biodegradation 7:03. https://doi.org/10.1016/j.ibiod.2013.07.003
Mazotto A, Ascheri JL, Godoy R, Damaso M, Couri S, Vermelho A (2017) Production of feather protein hydrolyzed by B. subtilis AMR and its application in a blend with cornmeal by extrusion. LWT-Food Science and Technology 84. https://doi.org/10.1016/j.lwt.2017.05.077
Meng Y, Tang Y, Zhang X, Wang J, Zhou Z (2022) Molecular identification of keratinase DgokerA from Deinococcus gobiensis for feather degradation. Appl Sci 12(1):464. https://doi.org/10.3390/app12010464
Mercer DK, Stewart CS (2019) Keratin hydrolysis by dermatophytes. Med Mycol 57(1):13–22. https://doi.org/10.1093/mmy/myx160
Mini KD, Sumi MG, Jyothis M (2015) Screening and selection of fungus for keratinase production by solid state fermentation and optimization of conditions of SSF and formulation of low cost medium for the production of keratinase by Aspergillus flavus S125. Int J Curr Microbiol App Sci 4(9):535–548. https://www.ijcmas.com/vol-4-9/K.D.%20Mini,%20et%20al.pdf
Mroczko B, Groblewska M, Litman-Zawadzka A, Kornhuber J, Lewczuk P (2018) Amyloid β oligomers (AβOs) in Alzheimer’s disease. J Neural Transm (vienna) 125(2):177–191. https://doi.org/10.1007/s00702-017-1820-x
Nandini A, Madhusudhan DN, Dayanand A (2015) Enhanced production, purification and characterization of alkaline keratinase from Streptomyces minutiscleroticus DNA38. Int Lett Nat Sci 43:27–37. https://doi.org/10.18052/www.scipress.com/ILNS.43.27
Navone L, Speight R (2020) Enzymatic removal of dags from livestock: an agricultural application of enzyme technology. Appl Microbiol Biotechnol 104(13):5739–5748. https://doi.org/10.1007/s00253-020-10656-2
Nnolim NE, Nwodo UU (2020) Bacillus sp. CSK2 produced thermostable alkaline keratinase using agro-wastes: keratinolytic enzyme characterization. BMC Biotechnol 20(1):65. https://doi.org/10.1186/s12896-020-00659-2
Nnolim NE, Okoh AI, Nwodo UU (2020) Proteolytic bacteria isolated from agro-waste dumpsites produced keratinolytic enzymes. Biotechnol Rep (amst) 27:e00483. https://doi.org/10.1016/j.btre.2020.e00483
Nurkhasanah U, Suharti, (2019) Preliminary study on keratinase fermentation by Bacillus sp. MD24 under solid state fermentation. IOP Confe Ser: Earth Environ Sci 276(1):012016. https://doi.org/10.1088/1755-1315/276/1/012016
Pagar AD, Patil MD, Flood DT, Yoo TH, Dawson PE, Yun H (2021) Recent advances in biocatalysis with chemical modification and expanded amino acid Alphabet. Chem Rev 121(10):6173–6245. https://doi.org/10.1021/acs.chemrev.0c01201
Pan X, Yang J, Xie P, Zhang J, Ke F, Guo X, Liang M, Liu L, Wang Q, Gao X (2021) Enhancement of Activity and thermostability of keratinase from Pseudomonas aeruginosa CCTCC AB2013184 by directed evolution with noncanonical amino acids. Front Bioengin Biotechnol 9:770–907. https://doi.org/10.3389/2Ffbioe.2021.770907
Pandey SC, Pande V, Sati D, Gangola S, Kumar S, Pandey A et al (2019) Microbial keratinase: a tool for bioremediation of feather waste. Smart Bioremediation Technol 217–53. https://doi.org/10.1016/B978-0-12-818307-6.00013-5
Park GT, Son HJ (2009) Keratinolytic activity of Bacillus megaterium F7–1, a feather-degrading mesophilic bacterium. Microbiol Res 164:478–485. https://doi.org/10.1016/j.micres.2007.02.004
Patience N, Abigail O, Ponchang W, Deborah A (2015) Keratinolytic activity of Cladosporium and Trichoderma species isolated from barbers’ landfill. Int J Biosci (IJB). 6(10):104–115. https://doi.org/10.12692/ijb/6.105-115
Paul T, Das A, Mandal A, Jana A, Maity C, Adak A, Mondal KC (2013) Effective dehairing properties of keratinase from Paenibacillus woosongensis TKB2 obtained under solid state fermentation. Waste Biomass Valorization 5(1):97–107. https://doi.org/10.1007/s12649-013-9217-z
Paul T, Jana A, Mandal A, Das Mohapatra P, Mondal K (2016) Bacterial keratinolytic protease, imminent starter for NextGen leather and detergent industries. Sustain Chem Pharm 1:007. https://doi.org/10.1016/j.scp.2016.01.001
Pawar VA, Prajapati AS, Akhani RC, Patel DH, Subramanian RB (2018) Molecular and biochemical characterization of a thermostable keratinase from Bacillus altitudinis RBDV1. Biotech 8(2):107. https://doi.org/10.1007/s13205-018-1130-5
Peng Z, Mao X, Zhang J, Du G, Chen J (2020) Biotransformation of keratin waste to amino acids and active peptides based on cell-free catalysis. Biotechnol Biofuels 13:61. https://doi.org/10.1186/s13068-020-01700-4
Peng Z, Peng X, Yang S, Guocheng D, Zhang J, Chen J (2021) Cysteine-mediated cyclic metabolism drives the microbial degradation of keratin. ACS Sustain Chem Eng 9(29):9861–9870. https://doi.org/10.1021/acssuschemeng.1c02627
Qiu J, Wilkens C, Barrett K, Meyer AS (2020) Microbial enzymes catalyzing keratin degradation: classification, structure, function. Biotechnol Adv 44:107607. https://doi.org/10.1016/j.biotechadv.2020.107607
Qiu J, Barrett K, Wilkens C, Meyer AS (2022) Bioinformatics based discovery of new keratinases in protease family M36. New Biotechnol 68:19–27. Advance online Publication. https://doi.org/10.1016/j.nbt.2022.01.004
Radha S, Gunasekaran P (2009) Purification and characterization of keratinase from recombinant Pichia and Bacillus strains. Protein Expr Purif 64(1):24–31. https://doi.org/10.1016/j.pep.2008.10.008
Rajput RGLB, Srivastava A, Wahi D, Shrivastava N, Kundu B, Grover A (2019) Specific keratinase derived designer peptides potently inhibit A β aggregation resulting in reduced neuronal toxicity and apoptosis. Biochem J 476(12):1817–1841. https://doi.org/10.1042/BCJ20190183
Ramnani P, Gupta R (2004) Optimization of medium composition for keratinase production on feather by Bacillus licheniformis RG1 using statistical methods involving response surface methodology. Biotechnol Appl Biochem 40:191–196. https://doi.org/10.1042/BA20030228
Raval VH, Pillai S, Rawal CM, Singh SP (2014) Biochemical and structural characterization of a detergent-stable serine alkaline protease from seawater haloalkaliphilic bacteria. Process Biochem 49(6):955–62. https://doi.org/10.1016/2Fj.procbio.2014.03.014
Reddy RM, Sathi RK, Ranjita CY, Bee H, Reddy G (2017) Effective feather degradation and keratinase production by Bacillus pumilus GRK for its application as bio-detergent additive. Bioresour Technol 243:254–263. https://doi.org/10.1016/j.biortech.2017.06.067
Rehman R, Ahmed M, Siddique A, Hasan F, Hameed A, Jamal A (2017) Catalytic role of thermostable metalloproteases from Bacillus subtilis KT004404 as Dehairing and destaining agent. Appl Biochem Biotechnol 181(1):434–450. https://doi.org/10.1007/s12010-016-2222-5
Reis S, Beys-da-Silva WO, Tirloni L, Santi L, Seixas A, Termignoni C, Silva M, Macedo AJ (2020) The extremophile Anoxybacillus sp. PC2 isolated from Brazilian semiarid region (Caatinga) produces a thermostable keratinase. J Basic Microbiol 60(9):809–815. https://doi.org/10.1002/jobm.202000186
Ribitsch D, Karl W, Birner-Gruenberger R, Gruber K, Eiteljoerg I, Remler P, Wieland S, Siegert P, Maurer KH, Schwab H (2010) C-terminal truncation of a metagenome-derived detergent protease for effective expression in E. coli. J Biotechnol 150(3):408–416. https://doi.org/10.1016/j.jbiotec.2010.09.947
Rouse JG, Van Dyke ME (2010) A review of keratin-based biomaterials for biomedical applications. Materials 3:999–1014. https://doi.org/10.3390/2Fma3020999
Sone T, Haraguchi Y, Kuwahara A, Ose T, Takano M, Abe A, Tanaka M, Tanaka I, Asano K (2015) Structural characterization reveals the keratinolytic activity of an Arthrobacter nicotinovorans protease. Protein Pept Lett 22(1):63–72. https://doi.org/10.2174/0929866521666140919100851
Srivastava B, Khatri M, Singh G, Arya SK (2020) Microbial keratinases: an overview of biochemical characterization and its eco-friendly approach for industrial applications. J Clean Prod 252:119847. https://doi.org/10.1016/j.jclepro.2019.119847
Su C, Gong J-S, Qin J, Li H, Li H, Xu Z-H et al (2020) The tale of a versatile enzyme: molecular insights into keratinase for its industrial dissemination. Biotechnol Adv 45:107655. https://doi.org/10.1016/j.biotechadv.2020.107655
Supuran CT, Scozzafava A, Clare BW (2002) Bacterial protease inhibitors. Med Res Rev 22(4):329–372. https://doi.org/10.1002/med.10007
Syed DG, Lee JC, Li WJ, Kim CJ (2009) Production, characterization and application of keratinase from Streptomyces gulbargensis. Bioresour Technol 100:1868–1871. https://doi.org/10.1016/j.biortech.2008.09.047
Tamreihao K, Mukherjee S, Khunjamayum R, Devi LJ, Asem RS, Ningthoujam DS et al (2019) Feather degradation by keratinolytic bacteria and bio-fertilizing potential for sustainable agricultural production. J Basic Microbiol 59:4–13. https://doi.org/10.1002/jobm.201800434
Tang Y, Guo L, Zhao M, Gui Y, Han J, Lu W, Dai Q, Jiang S, Lin M, Zhou Z, Wang J (2021) A Novel Thermostable keratinase from Deinococcus geothermalis with potential application in feather degradation. Appl Sci 11(7):3136. https://doi.org/10.3390/app11073136
Thi D, Tran M, Thi T, Phan P, Thi T, Doan N (2020) Integrative expression vectors with Pgrac promoters for inducer-free overproduction of recombinant proteins in Bacillus subtilis. Biotechnol 10:e3456. https://doi.org/10.1016/j.btre.2020.e00540
Tiberti M, Papaleo E (2011) Dynamic properties of extremophilic subtilisin-like serine-proteases. J Struct Biol 174(1):69–83. https://doi.org/10.1016/j.jsb.2011.01.006
Tiwary E, Gupta R (2010) Medium optimization for a novel 58 kDa dimeric keratinase from Bacillus licheniformis ER-15: biochemical characterization and application in feather degradation and dehairing of hides. Bioresour Technol 101(15):6103–6110. https://doi.org/10.1016/j.biortech.2010.02.090
Toplak A, Wu B, Fusetti F, Quaedflieg PJ, Janssen DB (2013) Proteolysin, a novel highly thermostable and cosolvent-compatible protease from the thermophilic bacterium Coprothermobacter proteolyticus. Appl Environ Microbiol 79(18):5625–5632. https://doi.org/10.1128/AEM.01479-13
Tork SE, Shahein YE, El-Hakim AE, Abdel-Aty AM, Aly MM (2013) Production and characterization of thermostable metallo-keratinase from newly isolated acillus subtilis NRC 3. Int J Biol Macromol 55:169–175. https://doi.org/10.1016/j.ijbiomac.2013.01.002
Uttangi V, Aruna K (2018) Optimization of production and partial characterization of keratinase produced by Bacillus thuringiensis strain BT407 isolated from poultry soil. Int J Curr Microbiol Appl Sci 7(04):596–626. https://doi.org/10.20546/ijcmas.2018.704.069
Verma A, Singh H, Anwar MS, Kumar S, Ansari MW, Agrawal S (2016) Production of thermostable organic solvent tolerant keratinolytic protease from Thermoactinomyces sp. RM4: IAA production and plant growth promotion. Front Microbiol 7:1189. https://doi.org/10.3389/fmicb.2016.01189
Verma A, Singh H, Anwar S, Chattopadhyay A, Tiwari KK, Kaur S, Dhilon GS (2017) Microbial keratinases: industrial enzymes with waste management potential. Crit Rev Biotechnol 37:476–491. https://doi.org/10.1080/07388551.2016.1185388
Vidmar B, Vodovnik M (2018) Microbial keratinases: enzymes with promising biotechnological applications. Food Technol Biotechnol 56(3):312–328. https://doi.org/10.17113/ftb.56.03.18.5658 (PMCID: PMC6233012)
Vijayaraghavan P, Lavanya J, Gnana S, Vincent P (2012) Biosynthesis and characterization of keratinolytic protease from Actinobacterium sp. in solid state culture. Int J Appl Biol Pharm 3(2):149–158. https://pesquisa.bvsalud.org/portal/resource/pt/sea-163843
Vijayaraghavan P, Saranya S, Vincent SG (2014) Cow dung substrate for the potential production of alkaline proteases by Pseudomonas putida strain AT in solid-state fermentation. Hindawi Publishing Corporation Chinese Journal of Biology. Article ID 217434, 7. https://doi.org/10.1155/2014/217434
Wang L, Cheng G, Ren Y, Dai Z, Zhao ZS, Liu F, Li S, Wei Y, Xiong J, Tang XF, Tang B (2015) Degradation of intact chicken feathers by Thermoactinomyces sp. CDF and characterization of its keratinolytic protease. Appl Microbiol Biotechnol 99(9):3949–3959. https://doi.org/10.1007/s00253-014-6207-4
Wang B, Yang W, McKittrick J, Meyers MA (2016) Keratin: Structure, mechanical properties, occurrence in biological organisms, and efforts at bioinspiration. Prog Mater Sci 76:229–318. https://doi.org/10.1016/j.pmatsci.2015.06.001
Wang L, Qian Y, Cao Y, Huang Y, Chang Z, Huang H (2017) Production and characterization of keratinolytic proteases by a chicken feather-degrading thermophilic strain, Thermoactinomyces sp. YT06. J Microbiol Biotechnol 27(12):2190–2198. https://doi.org/10.4014/jmb.1705.05082
Wu JW, Chen XL (2011) Extracellular metalloproteases from bacteria. Appl Microbiol Biotechnol 92(2):253–262. https://doi.org/10.1007/s00253-011-3532-8
Wu WL, Chen MY, Tu IF, Lin YC, Kumar EN, Chen MY, Ho MC, Wu SH (2017) The discovery of novel heat-stable keratinases from Meiothermus taiwanensis WR-220 and other extremophiles. Sci Rep 7(1):4658. https://doi.org/10.1038/s41598-017-04723-4
Yahaya RSR, Normi YM, Phang LY, Ahmad SA, Abdullah JO, Sabri S (2021) Molecular strategies to increase keratinase production in heterologous expression systems for industrial applications. Appl Microbiol Biotechnol 105(10):3955–3969. https://doi.org/10.1007/s00253-021-11321-y
Yan BQ, Chen XL, Hou XY, He H, Zhou BC, Zhang YZ (2009) Molecular analysis of the gene encoding a cold-adapted halophilic subtilase from deep-sea psychrotolerant bacterium Pseudoalteromonas sp. SM9913: cloning, expression, characterization and function analysis of the C-terminal PPC domains. Extremophiles: life under extreme conditions 13(4):725–733. https://doi.org/10.1007/s00792-009-0263-1
Yang H, Liu L, Shin HD, Chen RR, Li J, Du G, Chen J (2013) Integrating terminal truncation and oligopeptide fusion for a novel protein engineering strategy to improve specific activity and catalytic efficiency: alkaline α-amylase as a case study. Appl Environ Microbiol 79(20):6429–6438. https://doi.org/10.1128/AEM.02087-13 (PMID: 23956385; PMCID: PMC3811186)
Yi D, Xing J, Gao Y, Pan X, Xie P, Yang J, Wang Q, Gao X (2020) Enhancement of keratin-degradation ability of the keratinase KerBL from Bacillus licheniformis WHU by proximity-triggered chemical crosslinking. Int J Biol Macromol 163:1458–1470. https://doi.org/10.1016/j.ijbiomac.2020.08.021
Yue XY, Zhang B, Jiang DD (2011) Separation and purification of a keratinase as pesticide against root-knot nematodes. World J Microbiol Biotechnol 27:2147–2153. https://doi.org/10.1007/s11274-011-0680-z
Zheng L, Yu X, Wei C, Qiu L, Yu C, Xing Q (2020) Production and characterization of a novel alkaline protease from a newly isolated Neurospora crassa through solid-state fermentation. LWT (food Science and Technology) 122:108990. https://doi.org/10.1016/j.lwt.2019.108990
Acknowledgements
This work is carried out with the help of prestigious material of the libraries and special thanks to Institute.
Author information
Authors and Affiliations
Contributions
F Akram: designed the study and wrote the manuscript.
A Aqeel: worked on graphics and wrote some part of manuscript.
M Shoaib: help in collecting data.
IU Haq: critically read and supervised all work.
FI Shah: carried out proof reading and help in analyzed the data.
Corresponding author
Ethics declarations
Research involving human participants and/or animals
N/A. This research did not involve human participants and/or animals.
Consent to publish
All authors have given their consent for article publication.
Informed consent
N/A. This research did not involve human participants.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Akram, F., Aqeel, A., Shoaib, M. et al. Multifarious revolutionary aspects of microbial keratinases: an efficient green technology for future generation with prospective applications. Environ Sci Pollut Res 29, 86913–86932 (2022). https://doi.org/10.1007/s11356-022-23638-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-23638-w