Abstract
In this study, we present the characterization of the BNO1T bacterial strain isolated from the deep subsurface saline spring at the Baksan Neutrino Observatory INR RAS (Kabardino-Balkaria, Russia). The complete genome sequence of the strain BNO1T is 5,347,902 bp, with a GC content 41 and 49%. The cell wall peptidoglycan contains meso-diaminopimelic acid. The major isoprenoid quinone is MK-7 and the polar lipids are diphosphatidylglycerol, phosphatidylglycerol, and phosphatidylethanolamine. The major fatty acids are anteiso-C15:0 (23.34%), iso-C15:0 (20.10%), C16:0 (11.96%), iso-C16:0 (10.88%), and anteiso-C17:0 (10.79%). The 16S rRNA gene sequence clearly demarcated the strain as belonging to Cytobacillus genera. Based on the phylogenetic analysis, ANI (average nucleotide identity) and dDDH (digital DNA-DNA hybridization) assessments we propose to assign the strain BNO1T and other related strains to new species and to name it Cytobacillus pseudoceanisediminis sp. nov. (The values of ANI and dDDH between BNO1T and Cytobacillus oceanisediminis CGMCC 1.10115 T are 80.65% and 24.7%, respectively; values of ANI and dDDH between BNO1T and Cytobacillus firmus NCTC 10335 T are 89% and 38%, respectively). Genomic analysis of strain BNO1T revealed pathways for C1 compounds oxidation and two pathways for C1 compounds assimilation: serine and ribulose monophosphate pathways. In addition, strain BNO1T contains a plasmid (342,541 bp) coding multiple genes involved in heavy metal ion balance. Moreover, heavy metal toxicity testing confirmed the high potential of the strain BNO1T as a source of metal resistance genes and enzymes. The type strain is BNO1T (= BIM B-1921 T = VKM B-3664 T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Life in remote corners of the Earth with extreme conditions has always aroused great interest among scientists. The harsh living conditions in such places force their inhabitants to create special biological adaptations in order to remain active and stable. A range of adaptive enzymes and their pathways may pave way for the potential applications in biotechnology, for instance, enzymes that are stable at wide ranges of temperature and salinity [1, 2] or mechanisms of adaptation to elevated levels of heavy metals that can be used for the development of biosensors and bioremediation techniques [3]. Furthermore, studies of extremophiles is an important area of research in astrobiology due to the similarity of extremophile habitats on Earth to conditions on other planets [4]. Thus, extremophiles are important models for studies related to the search of extraterrestrial life and the development of life-sustaining mechanisms during spaceflights [4].
Unusual habitat conditions, such as highly mineralized underground hydrothermal springs, are of interest in the search for extremophilic organisms. One of these springs is located in an unused part of Baksan Neutrino Observatory (BNO) INR RAS, used for the conduct of studies in nuclear physics and astrophysics [5,6,7]. Here we report about sequencing and characterization of a bacterial strain BNO1T isolated from the underground highly mineralized spring of BNO located 2 km beneath the mountain Andyrchy in Elbrus region, Kabardino-Balkaria, Russia. In this work, we present the complete genome sequence and phylogenetic analysis of strain BNO1T, propose its central carbohydrate metabolism, and identify abundant heavy metal ion resistance genes. Based on phylogenetic data we propose the name Cytobacillus pseudoceanisediminis sp. nov. for Cytobacillus oceanisediminis 2691-related strains including strain BNO1T with strain BNO1T as type strain.
To the best of our knowledge, this and previous work [8] are the first studies in the field of biology carried out at Baksan Neutrino Observatory and our work reveals the unique experimental potential of BNO for biophysical, radiobiological, and astrobiological studies [9].
Materials and Methods
Bacterial Strain, Culture Conditions, and Biochemical Tests
The strain was obtained from samples collected in September 2020 in an unused part of the underground tunnel of Baksan Neutrino Observatory INR RAS (Russia) from a spring (41 °C) at a distance of 4 km from the entrance (43°16′32.5″N 42°41′30.3″E) and 2 km under the mountain Andyrchy surface, Baksan Gorge, Kabardino-Balkaria, Russia (Supplementary Fig. 1). The strain was isolated using dilution-plating technique on LB agar at 37 °C. Biochemical characteristics were screened with API® 20E and API® 50CH microbiological tests (BioMérieux®, France), according to Logan and Berkeley [10]. Nitrate reduction test was performed with sodium nitrate and Griess reagent (Lenreactiv®, Russia), according to Skerman [11]. For biochemical profiling, three independent repeats were performed.
Chemotaxonomic Characterization
To determine 2,6-diaminopimelic acid stereoisomers, the cells were hydrolyzed with 6-M HCl at 100 °C for 18 h, the acid was evaporated in air, and the hydrolysis products were separated in the system methanol/water/HCl/pyridine (80:17.5:2.5:10) using paper chromatography. The meso-diaminopimelic acid was detected with the ninhydrin spray reagent [12]. Respiratory quinones were extracted from freeze-dried cells with chloroform/methanol (2:1 v/v) and analyzed by mass spectrometer LCQ Advantage MAX (Thermo Finnigan) [13]. The fatty acid composition was analyzed using a gas chromatography-mass spectrometer 7890B + 5977B (Agilent Technologies, USA). The cell biomass was dried, saponificated (3.75 M NaOH/MeOH, 100 °C, 30 min), and subjected to acidic methanolysis (6 N HCl/MeOH, 80 °C, 10 min). The products of methanolysis were extracted with hexane:methyl tert-butyl ether (1:1 w/w) and processed with alkali (0.3 M NaOH, 5 min). The obtained products were separated on a 5% phenyl methyl silicone capillary column HP-5MS (0.25 mm × 30 m) in the temperature gradient from 45 to 300 °C at 40 °C min−1. Fatty acids and other lipid components were ionized by electron impact and analyzed in the scan mode. The compounds were identified using the NIST17 mass spectrometer library (https://chemdata.nist.gov/dokuwiki/doku.php?id=chemdata:nist17). The fatty acid content was determined as a percentage of the total ion current peak area [14]. Polar lipids were extracted from freeze-dried cells with chloroform/methanol (2:1, by vol.) and separated on HPTLC Silica gel 60F plates (Merck, Germany) using the following solvent systems: chloroform/methanol/water (65:25:4, by vol.) in the horizontal dimension and chloroform/methanol/acetic acid/water (80:12:15:5, by vol.) in the vertical dimension for two-dimensional TLC. Polar lipids were identified using spraying reagents for visualizing the spots: molybdenum blue for phospholipids, ninhydrin for aminolipids, and alpha-naphthol solution for glycolipids [15].
DNA isolation, Sequencing, and Genome Assembly
Total DNA was extracted with QIAGEN Genomic-tip 20/G (QIAGEN, Germany), according to manufacturer’s protocol. DNA was sequenced without amplification using Oxford Nanopore® MinION with r9.4.1 flow cell. Sequencing data were basecalled using Guppy® 5.0.14. Basecalling ended with 1.39 million of sequences (3.256 billion of bases), N50 of 3.6 kb, quality score of 11.7. Assembly was performed with Canu 2.2 [16]. Genome sequence is available in NCBI databases under accession GCA_023516215.1.
Phylogenetic Analysis and Gene Annotation
Initial classification of the strain BNO1T was performed using GTDB-Tk v2.0.0 [17]. That included gene calling for 120 bacterial genes [18], multiple sequence alignment to GTDB genomes, trimming of alignment to approximately 5000 positions, and mapping on GTDB preconstructed tree (release 207). A dataset containing all available C. oceanisediminis genomes (from NCBI genome database), except for low-quality and incomplete ones (strains DE0166, DE0392, MN36-4, 1T3), as well as representative genomes of all Cytobacillus species and also S. globispora genome were used for further phylogeny analysis. Same as in the initial classification gene calling, multiple sequence alignment and trimming of alignment were performed using GTDB-Tk v2.0.0 [17]. Based on this alignment tree was inferred in MEGA 11 [19] using maximum-likelihood approach under WAG model [20], 1000 bootstrap resamplings were performed. Representation of the tree was polished using ETE3 Toolkit v3.1.2 [21] and Treemmer v0.3 [22]. Average nucleotide identity (ANI) values were calculated using webservice (https://www.ezbiocloud.net/tools/ani) [23]. Digital DNA–DNA Hybridization values were calculated using webservice (https://ggdc.dsmz.de/ggdc.php) [24]. The 16S rRNA gene tree was inferred from Clustal Omega sequence alignment [25] in MEGA 11 [19] using the maximum-likelihood approach under Tamura 3-parameter model [26], with 1000 bootstrap resamples.
Gene annotation was performed using the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) (build 5742) as well as the RAST v2.0 server [27]. Annotations produced by RAST contained 7124 coding sequences and 144 RNAs, and annotations produced by PGAP contained 5830 coding sequences and 150 RNAs. The genes (28%) were assigned to curated subsystems using RAST annotator. Attention has been paid to methylotrophy due to huge number of genes assigned to subsystem “One-carbon Metabolism” (80 genes) as opposed to number of genes from “Central Carbohydrate Metabolism” subsystem (142 genes). Enzymes associated with methylotrophy were added to the metabolic circuit if their genes were annotated with at least one of the two annotation tools.
Evaluation of Culture Growth
The strain BNO1T was grown at 37 °C in personal bioreactors RTS-1C (Biosan®, Latvia) using Buffered Peptone Water (Bio-Rad, USA) and Buffered Peptone Water containing 4% of methanol. The devices were calibrated on the same set of culture dilutions and ran simultaneously; cell growth kinetics were registered every 30 min. Obtained growth curves were fitted with Gompertz function [28].
Chemical Analysis of Water
Chemical analysis of water was performed in Dubna Ecoanalytical Laboratory of Federal State Budgetary Water Management Institution “Centrregionvodhoz.”
Heavy Metal Toxicity Tests
Heavy metal toxicity tests were performed on LB agar plates supplemented with heavy metal ion in concentrations: Cu2+−300 µM, 1 mM, 3 mM, and 5 mM; Pb2+−300 µM, 1 mM, 3 mM, 5 mM, and 9 mM; Zn2+−1 mM, 3 mM, 5 mM, 9 mM, and 27 mM; and Cd2+−100 µM, 300 µM, 500 µM, and 1 mM. Stock solutions were prepared from CuSO4, Pb(NO3)2, Cd(NO3)2·4H2O, and Zn(NO3)2·6H2O (Sigma-Aldrich®). Experiments were done in duplicate. Plates were checked for growth after 24 h of incubation at 37 °C (Supplementary Fig. 2). The lowest heavy metal concentration that inhibited strain growth was recorded as minimum inhibitory concentration (MIC) of Cu, Pb, Zn, and Cd. The highest heavy metal concentration that did not prevent strain growth was recorded as survival limit. The test for polymetallic resistance was performed with a mixture of metals at a concentration of half the survival limit: copper (1.5 mM), lead (2.5 mM), and cadmium (250 μM).
The Sequence Accession Number
Genome sequence is available in NCBI databases under accession number GCA_023516215.1.
Results
Genome Sequence and Phylogenetic Analysis of the Strain BNO1T
Genome coverage was × 94.9. As a result of the assembly, two circular contigs were formed, representing a chromosome and a plasmid. The chromosome consists of 5.347.902 bp with a G + C content of 41.49% and the plasmid consists of 342.541 bp (Table 1).
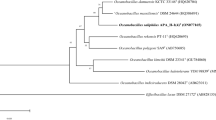
In the initial classification using GTDB-Tk, the isolate was placed into the clade of Cytobacillus strains and all three indels specific to the genus Cytobacillus were identified, namely in genes coding histidinol dehydrogenase, transcription repair conjugation factor, and serine protease (PDZ-domain) [29]. The BLAST search [30] of the 16S rRNA gene sequence through the NCBI nucleotide database revealed the 100% similarity of the 16S rRNA gene sequence of the strain BNO1T with Cytobacillus oceanisediminis 2691 strain (Fig. 1).
Phylogenetic tree constructed on 16S rRNA gene sequences of type strains of Cytobacillus genera. RefSeq 16S rRNA gene sequences are used when available. *—S. globispora genome should be reclassified, as discussed in Fig. 2 description
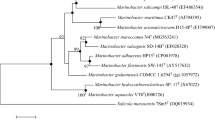
The result of subsequent phylogenetic analysis performed on the dataset of Cytobacillus genomes (described in the Materials and Methods section) and the ANI and dDDH values between strains of this dataset and the strain BNO1T are represented in Fig. 2. Surprisingly, phylogenetic analysis revealed that C. oceanisediminis strains presented in the NCBI genome database form two separate phylogenetic clades—groups A and B, with the group A more closely related to Cytobacillus firmus than the group B. This placement can be explained by the erroneous assignment of isolated from sedimentary layer of warm sea (South Korea) in 2012 strain 2691 to C. oceanisediminis [31]. The type strain of C. oceanisediminis CGMCC 1.10115 T was first described in 2010 (according to biochemical analysis and phylogenetic analysis based on 16S rRNA gene sequences the strain CGMCC 1.10115 T was classified as a novel species C. oceanisediminis [32]). However, the complete genome of C. oceanisediminis CGMCC 1.10115 T was not sequenced until 2015 [33]. Therefore, the strain 2691 was assigned to C. oceanisediminis in 2012 solely on the basis of the high similarity of 16S rRNA gene sequences between 2691 and CGMCC 1.10115 T strains, which is not a suitable criterion for identifying members of this family [32, 34].
Phylogenetic tree of NCBI representative species for the genera Cytobacillus along with all strains of Cytobacillus oceanisediminis and the strain BNO1T. Four strains of C. oceanisediminis (DE0166, DE0392, MN36-4, 1T3) are not represented due to the low quality of assembly. Cytobacillus formosensis and Cytobacillus purgationiresistens are not represented due to the absence of their genome assemblies in NCBI database. NCBI IDs are shown in brackets, and branch support numbers greater than 50% are shown on nodes. The ANI values of the strains compared to the strain BNO1T are represented on the right. *—deposited genome of Sporosarcina globispora (GCF_001274725.1) apparently belongs to the genera Cytobacillus. It contains three indels specific to the genera Cytobacillus [23] and does not contain Sporosarcina-specific indels [32]. In addition, the genome size of S. globispora is significantly larger than that of other Sporosarcina species (by 1–3 Mb more) and shares less than 70% of ANI with them. Thus, as suggested by Gupta and Patel [32], S. globispora should be reclassified
The DNA–DNA hybridization (DDH) method has been proposed as a reliable way to distinguish between species with high 16S rRNA gene sequences similarity [35] and its complementary digital approaches, Average Nucleotide Identity (ANI) and digital DDH (dDDH) have been shown to correlate with DDH values [36, 37]. In this work, evidence provided by ANI values (using an ANI threshold of 95% to distinguish between species [35]), dDDH values (threshold of 70% to distinguish between species [27]), and phylogenetic analysis suggest that groups A and B belong to different species (Fig. 2). The results of metabolic tests also revealed differences in metabolism of the type strain C. oceanisediminis CGMCC 1.10115 T (group B) and the strain BNO1T (group A) (Table 2). The presence of methylotrophy genes as well indicates the similarity of strains from group B to each other, in contrast to strain C. oceanisediminis CGMCC 1.10115 T (group A) (Supplementary Table 1).
Thus, we suggest to attribute group A to a different species than group B (that contains the type strain of C. oceanisediminis CGMCC 1.10115 T discovered by Zhang et al. 2010 [32]) and propose to name it Cytobacillus pseudoceanisediminis sp. nov. with type strain BNO1T [38].
Chemotaxonomy Characteristics of the Strain BNO1T
Analysis of 2,6-diaminopimelic acid showed the presence of meso-stereoisomer. The major menaquinone of strain BNO1—MK-7. The major fatty acids are anteiso-C15:0 (23.34%), iso-C15:0 (20.10%), C16:0 (11.96%), iso-C16:0 (10.88%), and anteiso-C17:0 (10.79%) (Supplementary table 2). The major polar lipids were identified as diphosphatidylglycerol, phosphatidylglycerol, and phosphatidylethanolamine. All chemotaxonomic properties are typical of the genus Cytobacillus [32, 39,40,41,42].
One-Carbon Metabolism of the Strain BNO1
Genome annotation using Prokaryotic Genome Annotation Pipeline (PGAP) [43] revealed that the strain BNO1T contains 5470 protein-coding genes (Table 1). Annotation using RAST [27] resulted in the categorization of genes to curated subsystem categories with a huge number of genes assigned to subsystem “One-carbon Metabolism” (80 genes) as opposed to the number of genes from “Central Carbohydrate Metabolism” subsystem (142 genes). Thus, methylotrophy has been proposed as the strain BNO1T central carbon metabolism. Throughout the characterization, we followed the approach proposed by Chistoserdova [44] where she considered the phenomenon of methylotrophy as the sum of moduli for each step of substrate oxidation and carbon fixation. The scheme of methylotrophic metabolism of the strain BNO1T is depicted in Fig. 3. Numbers for enzymes in the text are given in parentheses corresponding Fig. 3.
Scheme of the strain BNO1T methylotrophy-related pathways. Only methylotrophy pathways with a considerable number of annotated enzymes are shown. Reactions for which enzyme genes were not annotated are shown in red. The names of enzymes and metabolites are listed in Supplementary Tables 5 and 6. RuMP ribulose monophosphate pathway, H4F tetrahydrofolate pathway, GRC/EMCP glyoxylate regeneration cycle/ethylmalonyl-CoA pathway, TCA tricarbonic acid cycle (Color figure online)
Primary Oxidation
Genome analysis of the strain BNO1T revealed that it does not contain any known methane-oxidizing genes. For methanol oxidation, the strain BNO1T has three copies of the gene encoding Fe-containing NAD(P)-dependent alcohol dehydrogenases. Although these genes demonstrated quite low similarity to Bacillus methanolicus genes encoding methanol dehydrogenases (about 25%), they have 40% similarity to Gram-negative Cupriavidus necator gene coding methanol dehydrogenase (NAD(P)-dependent) [45]. Moreover, the strain BNO1T also possesses NUDIX hydrolase gene, which is 73% identical to B. methanolicus methanol dehydrogenase activator protein ACP. It is also important to note that alcohol dehydrogenases are able to oxidize a number of alcohols, but with different specificities. For example, even classic methanol dehydrogenases of Gram-positive B. methanolicus have a higher specificity for oxidation of ethanol rather than methanol [46]. Thus, these data suggest that the strain BNO1T can have an enzyme with methanol dehydrogenase activity.
The resistance of the strain BNO1T to methanol was tested by growing the strain in peptone water alone and in peptone water containing 4% methanol. Fitting the growth data with the Gompertz model curve \({\text{log}}\left\{ {{\text{ N}}\left( {\text{t}} \right)/{\text{N}}\left( 0 \right) \, } \right\} \, = {\text{ C }}*{\text{ exp}}\left( { \, - {\text{ exp}}\left\{ { \, - {\text{ B }}* \, \left[ {{\text{ t }} - {\text{ M }}} \right] \, } \right\} \, } \right)\) produced B-factor values were 1.22 h−1 for peptone water and 1.25 h−1 for peptone water with 4% methanol (Fig. 4). Hence, it can be concluded that the strain BNO1T tolerates a 4% concentration of methanol with slightly decreased value of the optical density at the stationary phase of growth. It is very likely that such resistance of strain BNO1T is associated with the ability to utilize methanol. In addition, during genome analysis the presence of genes coding oxidases of other primary methylated substrates (methylamines, dimethylsulfide, dimethylsulfoniopropionate, methanesulfonic acid) was not detected.
Formaldehyde Oxidation
The genome of the strain BNO1T contains genes involved in four different pathways associated with the formaldehyde oxidation module (enzyme codes corresponding to Fig. 3 are given in parentheses): NAD-dependent formaldehyde dehydrogenase gene (62), glutathione-dependent formaldehyde dehydrogenase gene (61), dissimilatory ribulose monophosphate pathway genes (13–15), and tetrahydrofolate oxidation pathway genes (3–8). In addition, there are two genes coding two formyl-H4F oxidation enzymes: formyl-tetrahydrofolate cycloligase (Ftfl) and formyl-tetrahydrofolate deformylase (PurU), the first acts in both directions, while the second only in oxidative direction [44]. NAD-dependent formate dehydrogenase gene was present. Thus, the strain BNO1T possesses genes for all necessary enzymes for formaldehyde oxidation to carbon dioxide. No trace of tetrahydromethanopterin formate oxidation pathway genes was found.
Carbon Assimilation
The genome of the strain BNO1T contains genes involved in two pathways of carbon assimilation (enzyme codes corresponding to Fig. 3 are given in parentheses): Ribulose Monophosphate Pathway (key enzymes: hexulose-phosphate synthase (10) and hexulose-phosphate isomerase (11) and Serine Pathway. Only one gene in assimilatory RuMP, the gene encoding phosphogluconate dehydratase, has not been found (16). It is possible that the activity of phosphogluconate dehydratase can be compensated by enzymes of the same protein family, such as dihydroxy-acid dehydratase (gene encoding dihydroxy-acid dehydratase has been detected). Glyoxylate regeneration for serine pathway is represented via isocitrate-lyase gene of Tricarboxylic Acid Cycle (39) and a number of genes encoding enzymes participating in Ethyl-Malonyl-CoA and Glyoxylate Regeneration pathways was also identified (genes of enzymes of TCA (33–37), genes of enzymes converting acetyl-CoA to ethylmalonyl-CoA (43–49), and all genes of enzymes facilitating conversion of propionyl-CoA to succinyl-CoA (58–60), suggesting that another glyoxylate regeneration pathway except for glyoxylate shunt may be present.
Metabolic tests indicated that the strain BNO1T can utilize a number of multicarbon substrates (fructose, n-acetylglucosamine, aesculin, maltose, sucrose, trehalose, starch, 5-ketogluconate, l-arginine, citrate, gelatin, d-mannitol) (Table 2). Thus, we can conclude that the strain BNO1T is a facultative methylotroph and C1 compounds used by the strain BNO1T likely can be produced by other members of the BNO spring microbe community.
Metal Resistance
The studied strain was found in the underground spring of the Baksan Neutrino Observatory INR RAS and chemical analysis of water from the BNO spring demonstrated that it is highly mineralized (7.3 g/L), and the most abundant ions are calcium and magnesium (represented in Supplementary Table 3 as hardiness) and chloride. The water also contains a certain amount of lead, cadmium, zinc, copper, manganese, and iron ions (Supplementary Table 3).
In the genome of the strain BNO1T, there are many genes that enable the strain to maintain metal homeostasis. Also about 20% of all plasmid-encoded proteins (that had proper assignments, no “hypothetical protein” assignments) were identified as involved in regulation of concentration of heavy metals; and such a high diversity of genes associated with the homeostasis of various metals (Supplementary table 4) in the strain BNO1T can serve as a potential source for the creation of genetically engineered constructs for the purpose of bioremediation and biosensors development [47]. For instance, CadC-T7 genes from closely related 2691 strain have already been successfully used to construct highly specific and sensitive cadmium and lead biosensors for the heavy metals detection [3].
Toxicity tests performed with copper, lead, cadmium, and zinc ions showed that the strain BNO1T tolerates high concentrations of cadmium, copper, and lead, but completely lacks tolerance of zinc ions (Table 3, Supplementary Fig. 2). The lead survival limit of at least 5 mM places the strain BNO1T among the most lead-tolerant microorganisms [48,49,50] and the light coloration of the strain BNO1T colonies on Petri dishes containing lead indicates no accumulation of lead on the surface of the cells, because otherwise the colonies would be brown [48]. In addition, the strain BNO1T survives on media containing a combination of copper, cadmium, and lead at half the survival limit concentration and thus the strain also has multi-metal tolerance.
The strain BNO1T also contains a number of genes providing resistances to various antibiotics: fosfomycin, vancomycin, bleomycin, fosmidomycin, puromycin, tosufloxacin, norfloxacin, penicillin, and streptogramin B. As supposed in work of Jianwen et al. [51], antibiotic resistance genes, along with heavy metal resistance genes, often co-occur as an adaptive mechanism in areas that have been contaminated with various heavy metals for a long time. Since the area near Barsan Neutrino Observatory is contaminated with heavy metals [52], our data support this theory.
Description of Cytobacillus pseudoceanisediminis sp. nov.
Cytobacillus pseudoceanisediminis (pseud.o.ce.a.ni.se.di'mi.nis. Gr. masc. adj. pseudes, false; N.L. gen. n. oceanisediminis, a specific epithet; and N.L. gen. n. pseudoceanisediminis, a false (Cytobacillus) oceanisediminis).
Gram-stain-positive rod-shaped aerobic Firmicutes bacterium isolated from deep underground highly mineralized spring of Baksan Neutrino Observatory (North Caucasus). After 24 h at 37 °C on LB agar, colonies are circular, opaque, mud colored, and 2–3 mm in diameter. No diffusible pigment is observed. The result for the presence of oxidase and arginine dihydrolase is positive. H2S production, acetoin production, and indole production are negative. Nitrate is reduced. C. pseudoceanisediminis can hydrolyze gelatin and starch, but not glycogen and urea and can utilize d-maltose, d-mannitol, trehalose, d-fructose, n-acetylglucosamine, aesculin, d-sucrose, 5-ketogluconate, citrate, sucrose, and l-arginine. The genomic DNA G + C content of the C. pseudoceanisediminis is 41.49%. The diagnostic diamino acid of the peptidoglycan is meso-diaminopimelic acid. The major menaquinone is MK-7. The phospholipid profile consists of diphosphatidylglycerol, phosphatidylglycerol and phosphatidylethanolamine. The major fatty acids are anteiso-C15:0 (23.34%), iso-C15:0 (20.10%), C16:0 (11.96%), iso-C16:0 (10.88%), and anteiso-C17:0 (10.79%). Bacterium contains several methylotrophy-related pathways, namely serine, ribulose monophosphate, and tetrahydrofolate pathways, and extensive heavy metal ion resistance genes.
The type strain of C. pseudoceanisediminis is BNO1T (= BIM B-1921 T = VKM B-3664 T).
References
Shaw AJ, Podkaminer KK, Desai SG et al (2008) Metabolic engineering of a thermophilic bacterium to produce ethanol at high yield. Proc Natl Acad Sci USA 105:13769–13774. https://doi.org/10.1073/pnas.0801266105
Bustard MT, Grant Burgess J, Meeyoo V, Wright PC (2000) Novel opportunities for marine hyperthermophiles in emerging biotechnology and engineering industries. J Chem Technol Biotechnol 75:1095–1109. https://doi.org/10.1002/1097-4660(200012)75:12%3c1095::AID-JCTB327%3e3.0.CO;2-3
Kim HJ, Lim JW, Jeong H et al (2016) Development of a highly specific and sensitive cadmium and lead microbial biosensor using synthetic CadC-T7 genetic circuitry. Biosens Bioelectron 79:701–708. https://doi.org/10.1016/j.bios.2015.12.101
Seckbach J, Stan-Lotter H (2020) Extremophiles as astrobiological models. John Wiley & Sons Inc, Hoboken, NJ. https://doi.org/10.1002/9781119593096
Gavriljuk JM, Gangapshev AM, Gezhaev AM et al (2013) Working characteristics of the New Low-Background Laboratory (DULB-4900). Nucl Instruments Methods Phys Res Sect A Accel Spectrometers, Detect Assoc Equip 729:576–580. https://doi.org/10.1016/j.nima.2013.07.090
Kuzminov VV (2018) Research program of the baksan neutrino observatory of the institute for nuclear research of the russian academy of sciences: 50 Years in the Making. Phys Part Nucl 49:473–481. https://doi.org/10.1134/S1063779618040391
Kuzminov VV (2012) The baksan neutrino observatory. Eur Phys J Plus 127:113. https://doi.org/10.1140/epjp/i2012-12113-0
Zarubin M, Gangapshev A, Gavriljuk Y et al (2021) First transcriptome profiling of D. melanogaster after development in a deep underground low radiation background laboratory. PLoS ONE 16:e0255066. https://doi.org/10.1371/journal.pone.0255066
Zarubin MP, Kuldoshina OA, Kravchenko EV (2021) Biological effects of low background radiation: prospects for future research in the low-background laboratory DULB-4900 of Baksan Neutrino Observatory INR RAS. Phys Part Nucl 52:19–30. https://doi.org/10.1134/S1063779621010056
Logan NA, Berkeley RCW (1984) Identification of Bacillus strains using the API system. J Gen Microbiol 130:1871–1882. https://doi.org/10.1099/00221287-130-7-1871
Skerman VBD (1967) A guide to the identification of the genera of bacteria, 2nd edn. The Williams & Wilkins Co., Baltimore
Kudryashova EB, Karlyshev AV, Ariskina EV et al (2018) Cohnella kolymensis sp. nov., a novel Bacillus isolated from Siberian permafrost. Int J Syst Evol Microbiol 68:2912–2917. https://doi.org/10.1099/ijsem.0.002919
Collins MD, Jones D (1981) Distribution of isoprenoid quinone structural types in bacteria and their taxonomic implication. Microbiol Rev 45:316–354. https://doi.org/10.1128/mr.45.2.316-354.1981
Miller LT (1982) Single derivatization method for routine analysis of bacterial whole-cell fatty acid methyl esters, including hydroxy acids. J Clin Microbiol 16:584–586. https://doi.org/10.1128/jcm.16.3.584-586.1982
Minnikin DE, O’Donnell AG, Goodfellow M et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Koren S, Walenz BP, Berlin K et al (2017) Canu: scalable and accurate long-read assembly via adaptive κ-mer weighting and repeat separation. Genome Res 27:722–736. https://doi.org/10.1101/gr.215087.116
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2020) GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36:1925–1927. https://doi.org/10.1093/bioinformatics/btz848
Parks DH, Rinke C, Chuvochina M et al (2017) Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol 2:1533–1542. https://doi.org/10.1038/s41564-017-0012-7
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699. https://doi.org/10.1093/oxfordjournals.molbev.a003851
Huerta-Cepas J, Serra F, Bork P (2016) ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol Biol Evol 33:1635–1638. https://doi.org/10.1093/molbev/msw046
Menardo F, Loiseau C, Brites D et al (2018) Treemmer: a tool to reduce large phylogenetic datasets with minimal loss of diversity. BMC Bioinformatics 19:164. https://doi.org/10.1186/s12859-018-2164-8
Yoon SH, Ha min S, Lim J et al (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek, Int J Gen Mol Microbiol 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Meier-Kolthoff JP, Carbasse JS, Peinado-Olarte RL, Göker M (2022) TYGS and LPSN: a database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acids Res 50:D801–D807. https://doi.org/10.1093/nar/gkab902
Sievers F, Wilm A, Dineen D et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539. https://doi.org/10.1038/msb.2011.75
Tamura K (1992) Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol Biol Evol 9:678–687. https://doi.org/10.1093/oxfordjournals.molbev.a040752
Overbeek R, Olson R, Pusch GD et al (2014) The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res 42:D206–D214. https://doi.org/10.1093/nar/gkt1226
Micha P, Corradini MG (2011) Microbial growth curves: what the models tell us and what they cannot. Crit Rev Food Sci Nutr 51:917–945. https://doi.org/10.1080/10408398.2011.570463
Patel S, Gupta RS (2020) A phylogenomic and comparative genomic framework for resolving the polyphyly of the genus Bacillus: Proposal for six new genera of Bacillus species, Peribacillus gen. nov., Cytobacillus gen. nov., Mesobacillus gen. nov., Neobacillus gen. nov., Metabacillus gen. nov. and Alkalihalobacillus gen. nov. Int J Syst Evol Microbiol 70:406–438. https://doi.org/10.1099/ijsem.0.003775
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Lee YJ, Lee SJ, Jeong H et al (2012) Draft genome sequence of Bacillus oceanisediminis 2691. J Bacteriol 194:6351–6352. https://doi.org/10.1128/JB.01643-12
Zhang J, Wang J, Fang C et al (2010) Bacillus oceanisediminis sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 60:2924–2929. https://doi.org/10.1099/ijs.0.019851-0
Whitman WB, Woyke T, Klenk HP et al (2015) Genomic encyclopedia of bacterial and archaeal type strains, phase III: The genomes of soil and plant-associated and newly described type strains. Stand Genomic Sci 10:26. https://doi.org/10.1186/s40793-015-0017-x
Ko KS, Oh WS, Lee MY et al (2006) Bacillus infantis sp. nov. and Bacillus idriensis sp. nov., isolated from a patient with neonatal sepsis. Int J Syst Evol Microbiol 56:2541–2544. https://doi.org/10.1099/ijs.0.64213-0
Tindall BJ, Rosselló-Móra R, Busse HJ et al (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266. https://doi.org/10.1099/ijs.0.016949-0
Goris J, Konstantinidis KT, Klappenbach JA et al (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Jung J, Jeong H, Kim HJ et al (2016) Complete genome sequence of Bacillus oceanisediminis 2691, a reservoir of heavy-metal resistance genes. Mar Genomics 30:73–76. https://doi.org/10.1016/j.margen.2016.07.002
Kämpfer P (1994) Limits and possibilities of total fatty acid analysis for classification and identification of Bacillus species. Syst Appl Microbiol 17:86–98. https://doi.org/10.1016/S0723-2020(11)80035-4
Albert RA, Archambault J, Rosselló-Mora R et al (2005) Bacillus acidicola sp. nov., a novel mesophilic, acidophilic species isolated from acidic Sphagnum peat bogs in Wisconsin. Int J Syst Evol Microbiol 55:2125–2130. https://doi.org/10.1099/ijs.0.02337-0
Priest FG, Goodfellow M, Todd C (1988) A numerical classification of the genus Bacillus. J Gen Microbiol 134:1847–1882. https://doi.org/10.1099/00221287-134-7-1847
Lim JM, Jeon CO, Kim CJ (2006) Bacillus taeanensis sp. nov., a halophilic Gram-positive bacterium from a solar saltern in Korea. Int J Syst Evol Microbiol 56:2903–2908. https://doi.org/10.1099/ijs.0.64036-0
Tatusova T, Dicuccio M, Badretdin A et al (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Chistoserdova L (2011) Modularity of methylotrophy, revisited. Environ Microbiol 13:2603–2622. https://doi.org/10.1111/j.1462-2920.2011.02464.x
Wu TY, Chen CT, Liu JTJ et al (2016) Characterization and evolution of an activator-independent methanol dehydrogenase from Cupriavidus necator N-1. Appl Microbiol Biotechnol 100:4969–4983. https://doi.org/10.1007/s00253-016-7320-3
Arfman N, Van Beeumen J, De Vries GE et al (1991) Purification and characterization of an activator protein for methanol dehydrogenase from thermotolerant Bacillus spp. J Biol Chem 266:3955–3960. https://doi.org/10.1016/s0021-9258(19)67886-5
Verma S, Kuila A (2019) Bioremediation of heavy metals by microbial process. Environ Technol Innov. https://doi.org/10.1016/j.eti.2019.100369
Utami U, Harianie L, Dunyana NR, Romaidi R (2020) Lead-resistant bacteria isolated from oil wastewater sample for bioremediation of lead. Water Sci Technol 81:2244–2249. https://doi.org/10.2166/wst.2020.281
Abdollahi S, Golchin A, Shahryari F (2020) Lead and cadmium-resistant bacterial species isolated from heavy metal-contaminated soils show plant growth-promoting traits. Int Microbiol 23:625–640. https://doi.org/10.1007/s10123-020-00133-1
Sun F, Shao Z (2007) Biosorption and bioaccumulation of lead by Penicillium sp. Psf-2 isolated from the deep sea sediment of the Pacific Ocean. Extremophiles 11:853–858. https://doi.org/10.1007/s00792-007-0097-7
Chen J, Li J, Zhang H et al (2019) Bacterial heavy-metal and antibiotic resistance genes in a copper tailing dam area in northern China. Front Microbiol 10:1916. https://doi.org/10.3389/fmicb.2019.01916
Bogatikov OA, Bortnikov NS, Dokuchaev AY et al (2014) Primary aspects of technogenic deposits at the modern stage: the example of the Tyrnyauz deposit. Dokl Earth Sci 456:585–589. https://doi.org/10.1134/S1028334X14050237
Author information
Authors and Affiliations
Contributions
EK coordinated the overall study, performed the experiments, and prepared and analyzed the data; AY, MZ, and AG collected the samples; KT performed the experiments and bioinformatic analysis; AY performed the experiments; NP and AA performed the chemotaxonomic characterization; EK, KT, AY, and MZ wrote the first draft; EK and KT edited the final version. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they do not have any conflict of interest to report.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tarasov, K., Yakhnenko, A., Zarubin, M. et al. Cytobacillus pseudoceanisediminis sp. nov., A Novel Facultative Methylotrophic Bacterium with High Heavy Metal Resistance Isolated from the Deep Underground Saline Spring. Curr Microbiol 80, 31 (2023). https://doi.org/10.1007/s00284-022-03141-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03141-8