Abstract
Small non-coding RNAs (sRNAs) are a class of regulatory RNAs 20–500 nucleotides in length, which have recently been discovered in prokaryotic organisms. sRNAs are key regulators in many biological processes, such as sensing various environmental changes and regulating intracellular gene expression through binding target mRNAs or proteins. Bacterial sRNAs have recently been rapidly mined, thus providing new insights into the regulatory network of biological functions in prokaryotes. Although most bacterial sRNAs have been discovered and studied in Escherichia coli and other Gram-negative bacteria, sRNAs have increasingly been predicted and verified in Gram-positive bacteria in the past decade. The genus Streptococcus includes many commensal and pathogenic Gram-positive bacteria. However, current understanding of sRNA-mediated regulation in Streptococcus is limited. Most known sRNAs in Streptococcus are associated with the regulation of virulence. In this review, we summarize recent advances in understanding of the functions and mechanisms of sRNAs in Streptococcus, and we discuss the RNA chaperone protein and synthetic sRNA-mediated gene regulation, with the aim of providing a reference for the study of microbial sRNAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Streptococcus, comprising a group of commensal and pathogenic low GC Gram-positive bacteria, typically has spherical or ovoid cells with homofermentative, anaerobic/aerotolerant, non-motile, catalase negative, and non-spore forming characteristics [1]. More than 55 species of Streptococcus are found in a wide variety of habitats including the mouth, respiratory tract, and skin surfaces of animals and humans, as well as in various environments such as soil and plants [2]. According to its hemolytic properties, Streptococcus is divided into three categories: (I) α-hemolytic Streptococcus e.g., S. mutans; (II) β-hemolytic Streptococcus, mainly including S. pyogenes (also called group A Streptococcus or GAS) [3] and S. agalactiae; and (III) γ-hemolytic Streptococcus, which are also called non-hemolytic Streptococcus and are generally not pathogenic [1]. Some species of Streptococcus significantly affect both humans and animals. For example, S. pyogenes, S. pneumoniae, and S. agalactiae are human pathogens causing serious acute infections [1,2,3], whereas S. thermophilus is an important probiotic lactic acid bacterium and the only “generally recognized as safe” species of Streptococcus [4].
Small non-coding RNAs (sRNAs) are a novel class of regulatory RNAs that fine-tune biological function networks, e.g., responses to environmental changes [5, 6]. Many sRNAs have been discovered [7, 8], and their detailed regulatory functions have been studied in bacteria [7]. However, compared with model bacteria such as Escherichia coli and Bacillus subtilis, current knowledge of sRNAs and sRNA-mediated regulation in Streptococcus is limited. In this review, we provide a comprehensive overview of the functions and regulatory mechanisms of known sRNAs in Streptococcus.
Small Non-coding RNAs
As an important class of regulators, sRNAs 20–500 nucleotides (nt) in length exist in bacteria but are generally not translated into proteins [7, 9]. sRNAs are transcribed from the intergenic regions (IGRs) located between open reading frames, or in the 5′ or 3′ non-coding regions (UTRs) of coding genes [10]. sRNAs play regulatory roles at the post-transcriptional level in the bacterial response to environmental stress, through complementary base-pairing with mRNA molecules or interaction with the corresponding protein molecules [11, 12]. sRNAs are important nodes in many signaling pathways and physiological metabolic pathways [5, 13]. Most known functional sRNAs, in the form of antisense RNAs, specifically bind target mRNAs and regulate target gene expression. Most known bacterial sRNAs act as regulatory elements that respond to extracellular environment stresses and play key roles in intracellular physiological processes [14].
The process of sRNA recognition is not easy to identify through experimental methods. First, sRNAs themselves do not encode proteins, their lengths are short, and they are not easily inhibited by single nucleotide mutations. Second, sRNAs are not translated, and therefore their sequences cannot be obtained by simply recognizing open reading frames. Hence, many sRNAs are discovered and identified through a combination of bioinformatics and experimental methods [15, 16]. For example, Rath et al. [15] have created an RNase III null mutant and used RNA sequencing (RNA-Seq) to analyze the differential expression between this mutant and the wild-type of S. pyogenes. Twelve significantly differential transcripts have been obtained using the analysis of Cufflinks and Stringtie softwares, and six putative sRNAs have been visually verified by Artemis and Bamview genome viewers.
The targets of bacterial sRNAs is commonly predicted by bioinformatics methods, mainly on the basis of sequence length, base matching, RNA secondary structure, and sequence conservation and location [17]. Currently used software programs for predicting sRNA target mRNAs include RNAPredator [18], sRNATarBase [19], CopraRNA [20], RNAup [21], RNAhybrid [22], TarPicker [23], IntaRNA [24], and Target RNA2 [25]. The main difference of these in silico sRNA target discovery programs is due to the use of different prediction models, including models for general RNA–RNA interaction and models specifically designed for sRNA–target mRNA interactions in bacteria [17]. Many experimental methods have been used to verify sRNA target genes, among which genetic methods, affinity technology, immune co-precipitation methods, and microarray technology are the most mature and widely used [16, 26]. Ribosome profiling (Ribo-seq) technology is a recently developed method that simultaneously determines transcriptional and translational levels in vivo [27]. Wang et al. [28] have used Ribo-seq to fully validate the known targets of sRNA RyhB from E. coli and have identified many novel sRNA targets. Guo et al. [29] have also employed Ribo-seq to discover the target of a new σE-dependent sRNA MicL. The experimental data have indicated that Ribo-seq is an effective method for identifying sRNA targets and detecting the extent of regulation by sRNA.
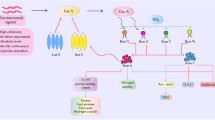
With the progress of bacterial sRNAs, the regulatory mechanisms of sRNAs are gradually being elucidated (Fig. 1A and B). Bacterial sRNAs regulate the virulence of bacteria in host cells and initiate internal mechanisms in bacterial cells, thus allowing bacteria to respond to changes in growth conditions, adapt to the environmental changes in infected cells [30], and control bacterial density by regulating the expression of quorum sensing system-regulated proteins [31]. In the absence of nutrients in the growth environment, bacterial sRNAs such as CsrB and CsrC [32] play important regulatory roles in anti-nutritional stress. Trace elements are indispensable in cell metabolism, and sRNAs regulate the relative balance of trace elements in bacteria. For example, sRNA RyhB regulates the gene expression of Fur (an iron-uptake regulator) in E. coli and YdeP (which is responsible for regulation of acidity) in Shigella flexneri [33]. In addition to regulating acid tolerance, sRNAs regulate other environmental stress responses, such as MicA, RybB, and MicL to Membrance stress [34], SgrS to phosphosugar stress [34], TisB to SOS stress [34], DsrA, ArcZ, RprA, and OxyS to oxygen stress [35]. sRNAs may have different regulatory mechanisms in different bacteria, thus leading to different functional effects [36].
Regulatory mechanisms of sRNAs. A Mechanisms of inhibition of translation. a sRNAs prevent the binding of 30S ribosomes and the initiation of translation, through incomplete complementary base-pairing with the ribosome binding site (RBS) located at the 5′-UTR of the target mRNA. b sRNAs bind the mRNA being translated with the aid of Hfq, thus resulting in structural recombination of mRNA. The RBS region undergoes complementary base-pairing to form a secondary structure that prevents ribosome binding, thus leading to translational inhibition. c After the sRNA base-pairs with the mRNA, the mRNA is degraded by RNase E. In target mRNA degradation, RNase E specifically acts on AU-rich single-stranded RNA, and can simultaneously degrade target mRNA and sRNA. B Regulatory mechanisms of activation of translation. a Some sRNAs activate translation by binding to the 5′-untranslated region (5′-UTR) of the target mRNA, making the RBS becomes available, allowing initiation of translation. b Some sRNAs directly and stringently base-pair with target mRNA with the help of Hfq, thus forming a double-stranded structure, thereby increasing the stability of mRNA and protecting it from degradation by RNase E
The Biological Functions of sRNAs in Streptococcus
sRNAs from S. pyogenes
Streptococcus pyogenes is a bacterial pathogen that causes diseases ranging from superficial infections (e.g., impetigo and pharyngitis) to systemic diseases (e.g., necrotizing fasciitis and toxic shock) [3]. Several sRNAs have been identified in S. pyogenes (Table 1), most of which are involved in virulence regulation [37].
sRNA FasX
FasX is an sRNA (~ 200 nt) that affects the expression of proteins (e.g., streptokinase) involved in the conversion of plasminogen to plasmin [37]. The mechanism of FasX regulation has been well described elsewhere [38,39,40,41]. FasX is part of an operon including genes encoding two histidine kinases (FasBC) and one response regulator (FasA) [37]. FasX influences the expression of virulence factors through three steps [38,39,40,41]: (I) FasX enhances the expression of streptokinase (a virulence factor) by stabilizing the mRNA of ska, encoding streptokinase; (II) FasX regulates pilus biosynthesis and adherence through destabilizing the mRNA of the pilus biosynthesis operon through base-pairing; and (III) FasX inhibits the translation of cpa, which encodes a pilin protein (Fig. 2a). FasX regulation is serotype specific and depends on the presence of target genes [38,39,40,41]. For example, FasX can posttranscriptionally regulate the expression of the adhesion- and internalization-promoting, fibronectin-binding proteins PrtF1 and PrtF2. The mechanism is through base-pairing to the prtF1 and prtF2 mRNAs within their 5′ untranslated regions, where blocks ribosome access and leads to an inhibition of mRNA translation [41]. As a molecular switch, FasX governs the transition of S. pyogenes between the colonization and dissemination stages of infection [41].
Regulation of sRNAs in Streptococcus. a FasX regulates the expression of virulence in S. pyogenes. FasX is a main effector of the FasBCA three-component system. FasB and FasC are histidine kinases that phosphorylate the response regulator FasA after signal sensing and subsequently upregulate expression of the fas operon. FasX inhibits the expression of sag (encoding streptolysin S, SLS), fbp54 (encoding a fibronectin-binding protein), and mrp (encoding M-related protein, a fibrinogen-binding protein). FasX increases the stability of ska (encoding streptokinase) mRNA through direct targeting, thereby increasing streptokinase synthesis. b Pel controls the expression of virulence factors in S. pyogenes. The direction of the arrow of ralp3 represents its direction of transcription. c RivX and rivR activate the expression of an operon encoding Mga, M protein (encoded by emm), and Sic (streptococcal inhibitor of complement) in S. pyogenes. d MarS activates the expression of mga (encoding multiple gene activator) mRNA
sRNAs Pel/sagA and RivX
Pleiotropic effect locus/streptolysin-associated gene A (Pel/sagA) regulates M and M-related proteins (virulence factor streptolysin S, SLS) in S. pyogenes [8]. Pel activates virulence gene expression with strain specificity. RofA-like protein IV regulator X (RivX) is an sRNA that is derived from a longer mRNA encoding the transcription regulator RivR (RALP4, a RofA-like protein family of transcriptional regulators) and corresponds to a processed form of the rivRX transcript [8]. Both RivR and RivX positively regulate the mga regulon (encoding multiple gene activator), which activates the expression of virulence-associated regulators and factors (Fig. 2b and c).
mga-Activating Regulatory sRNA (MarS)
Recently, Pappesch et al. [42] have reported the novel sRNA MarS in S. pyogenes M49591, which modulates the expression of Mga-dependent virulence factor and affects capsule biosynthesis and the fate of S. pyogenes. Deletion of MarS results in downregulation of the expression of mga and Mga-activated genes. The mode of action of MarS is shown in Fig. 2d.
sRNAs from S. pneumoniae
Streptococcus pneumoniae is an opportunistic pathogen associated with a wide range of human diseases [43]. It can cause serious infections such as sepsis, endocarditis, and meningitis. Furthermore, it is a leading cause of pneumonia, which kills more young children than any other infectious diseases [44].
sRNAs F20 and F32
The sRNAs F20 (also called as srn157) and F32 (previously identified as a tmRNA) are associated primarily with the virulence regulation of S. pneumoniae. F20 and F32 deletion mutants show decreased adhesion and invasion of nasopharyngeal or endothelial cells, thus resulting in a lack of fitness and competitive index in the environment of the nasopharynx and lungs [44]. Through proteomic analysis, Acebo et al. [45] have found that purine metabolism is significantly downregulated, whereas the pathways of DNA synthesis and repair are strongly upregulated, in an F20 deletion mutant. Kumar et al. [46] have reported that metabolic pathways encompassing the lactose transport system and multiple phosphoglucose transferase systems are depressed in an F32 mutant. F20 deletion thus has pleiotropic effects on virulence attenuation. The F32 mutant also shows a strong effect on bacterial pathogenesis. F32 is associated with deficiencies in the stress response and pathogenicity and has a core role in the trans-translation mechanism, which resolves stalled ribosomes on non-stop mRNAs [47]. However, the specific regulatory mechanisms of these two sRNAs and their target genes remains unclear.
cia-Dependent Small sRNAs (csRNAs)
csRNAs are a family of Cia, competence induction and altered cefotaxime susceptibility (CiaR)-regulated sRNAs in Streptococcus. CiaR-targeted promoters have been identified through mapping the genome of S. pneumoniae [48]. CiaR is a response regulator in the CiaRH two-component system (TCS), which is responsible for β-lactam susceptibility, competence, bacteriocin biosynthesis, autolysis and virulence in bacteria. Sixty-one csRNAs with lengths of 51–202 nt have been identified and classified into 40 different types through a search for CiaR-activated promoters in the genomes of 14 Streptococcus species [49]. Among these csRNAs, five pneumococcal csRNAs (csRNA1 to 5) are transcribed by cia-controlled non-coding RNA (ccnA-E) in the CiaR regulon. Halfmann et al. [48] have reported that csRNA4 and csRNA5 are involved in stationary-phase autolysis in S. pneumoniae R6. Microarray analysis of csRNA1 inactivation has uncovered a cis-acting effect on the mRNAs of adjacent genes, thus suggesting that csRNA1 is co-transcribed in strain D39 [50]. However, csRNA1 inactivation or activation does not affect cell growth, stress responses, global transcription or virulence.
sRNAs from S. mutans
Streptococcus mutans is a major clinical pathogen in dental caries that possesses various virulence factors and consequently can accumulate and colonize the surfaces of teeth [51]. It can utilize diverse carbohydrate sources, and produce and tolerate various acids; therefore, it can survive at low pH. It adheres to the surfaces of teeth, thereby resulting in demineralization of tooth surfaces [52]. Currently, sRNAs are known to regulate adherence and polysaccharide biosynthesis in S. mutans. Moreover, sRNAs in S. mutans are induced by different concentrations (1–5%) of sucrose during planktonic growth [53]. An array of sRNAs has been found to be induced under acid stress, in comparison with the sRNAs reported by Lee et al.[54]. The study of sRNAs that regulate bacterial virulence in S. mutans may be applied to solving the problem of dental caries (Table 1).
sRNA L10-Leader
Xia et al. [55] have predicted 334 sRNAs and verified the existence of L10-Leader in S. mutans UA159. The upregulation of L10-Leader in an acidic environment may indicate that L10-Leader binding to target mRNA improves the stability and translation of target mRNA. Consequently, cells can quickly adapt to acidic stress, and correct DNA matching is ensured under various environment stresses. However, the mechanism of L10-Leader in acidic environments remains unclear in S. mutans.
sRNA133474
Streptococcus mutans harbors diverse stress response pathways including TCSs to mitigate acid stress in the oral cavity [56]. Nine acid tolerance-associated sRNAs have been validated in S. mutans UA159 and other clinical strains cultured at pH 5.5 and 7.5. Compared with S. mutans culture at pH 7.5, in culture at pH 5.5, sRNA133474 is the most significantly downregulated among these sRNAs [56]. The mechanism of action of sRNA133474 involves regulation of the expression of LiaSR, CiaRH, and CovRS—TCSs responsible for acid tolerance.
sRNA srn225147
The bacteriocin mutacin IV from S. mutans antagonizes numerous non-mutans streptococcal species (e.g., S. gordonii). A target of sRNA srn225147 has been found to be the mutacin IV formation-associated gene comD, on the basis of RNAhybrid and RNAPredator prediction [57]. As compared with a negative control condition, comD expression has been found to significantly increase with 1400-fold increased srn225147 expression but to decrease with 400-fold increased srn225147 expression, thus suggesting that srn225147 has a two-way regulatory effect on comD expression. However, srn225147′s regulatory effect on mutacin IV biosynthesis is weak [57].
sRNA0426
The virulence of S. mutans is dependent on the formation of biofilms, which provide better conditions for bacteria to adapt to the changing environment in the oral cavity [58]. According to bioinformatics analyses, qRT-PCR, and crystal violet staining assays, Yin et al. [58] have found that sRNA0426 has a positive relationship with dynamic biofilm formation and exopolysaccharide production. In the synthesis of exopolysaccharide, sRNA0426 expression is positively correlated with the expression of three target mRNAs (GtfB, GtfC, and CcpA). Therefore, sRNA0426 has been found to play an important positive role in biofilm formation, thus providing novel insights into the biofilm regulatory network in S. mutans.
sRNAs from S. suis
Although S. suis serotype 2 (S. suis 2) is the main cause of porcine diseases, it can directly infect humans and cause hearing loss, meningitis, and septic shock [59, 60]. Zhang et al. [59] have performed RNA-seq in three strains of S. suis 2: P1/7 (reference strain) as well as 05ZYH33 and 98HAH33 (highly virulent isolates). Fifty-six sRNAs were predicted in three isolates, of which 12, particularly rliD and RatA, in all three isolates as well as cspA and rli38 in 05ZYH33 and 98HAH33 were associated with bacterial virulence. For the remaining sRNAs, over 27% (12/44) were found to be involved in riboswitches. Recently, sRNA rss04 has been confirmed to repress capsular polysaccharide biosynthesis in P1/7 [60], thus increasing adhesion and invasion of mouse brain microvascular endothelial cells. Moreover, rss04 regulates catabolite control protein A (CcpA) and the virulence factor LuxS, thereby affecting biofilm formation, capsular polysaccharide biosynthesis, and meningitis progression [61, 62]. In addition, Gong et al. [63] have verified that sRNA34 regulates the virulence of S. suis 05ZYH33. Transcriptomic analysis has indicated that deletion of sRNA34 represses the expression of HPA2 (encoding histone acetyltransferase) and 05SSU0013 (a cell cycle control gene) as well as 05SSU0003 and 05SSU1265, which are involved in the synthesis of lipoteichoic acids and contribute to cellular chain formation and elongation. Thus, the inactivation of sRNA34 results in longer cellular chains and attenuated virulence [63]. However, the mechanism of action of sRNAs in S. suis remains to be elucidated.
sRNAs from S. sanguinis
Streptococcus sanguinis, such as S. mutans, has been recognized as a pathogen involved in dental caries and periodontal diseases. Choi et al. [9] have predicted 219 sRNAs and characterized five putative sRNAs by RNA-seq, northern blotting, and qRT-PCR in S. sanguinis SK36. In the genome of SK36, among these five characterized sRNAs, the sRNA S.S-1964 is located to the right of the gene SSA_0513, encoding a putative ATP:cob (I) alamin denosyltransferase, and it may be involved in the conversion of vitamin B12 to coenzyme B12 through regulation of SSA_0513 expression [9]. Additionally, Ota et al. [64] have used computational target prediction and luciferase reporter assays to identify csRNA1-1 (a csRNA) as a negative regulator of the expression of pilT (a constituent of the type IV pilus operon) in S. anguinis. csRNA1-1 directly binds pilT mRNA, on the basis of RNA–RNA electrophoretic mobility shift assays. Moreover, csRNA1-1 and csRNA1-2 have been found to negatively regulate biofilm formation, thus suggesting that csRNAs are involved in the colonization process in S. sanguinis [64].
Perspectives
Although several sRNAs have been predicted and identified in Streptococcus, most current studies on sRNAs have focused on those involved in the regulation of virulence, such as FasX, Pel, RivX, and MarS in S. pyogenes, as well as F20 and F32 in S. pneumoniae. Understanding the roles of sRNAs in other cellular functions, particularly in the control of toxin–antitoxin systems and the responses to environmental changes, such as those in pH, oxygen, and nutrients, is very limited in Streptococcus but abundant in other bacteria [65]. Moreover, the current understanding of sRNA-mediated regulation, including modes of action and target identification, is largely insufficient in Streptococcus. sRNAs and their regulatory mechanisms reported in other bacteria have not yet been studied in Streptococcus. Thus, several questions regarding sRNAs in Streptococcus remain to be answered.
RNA chaperone protein Hfq (also known as HF-1) is required for the interaction between sRNA and target mRNAs in most sRNAs in Gram-negative bacteria [66]. For example, at least one-third of the experimentally verified E. coli sRNAs are Hfq-binding sRNAs [26]. Hfq is also at the core of sRNA-mediated post-transcriptional regulation, although the need for Hfq in the regulation of sRNA depends on multiple factors. The presence of Hfq promotes the pairing of most bacterial sRNAs with target mRNAs [26]. Hfq is involved in regulating the stability, polyadenylation, and translation of many RNAs, as well as in RNA processing. Deletion of the Hfq gene results in growth defects, decreased resistance to environmental stress, and changes in toxicity [67]. Hfq homologues also exist in some Gram-positive bacteria. However, the only known Hfq-dependent sRNA in Gram-positive bacteria is LhrA from L. monocytogenes [68]. On the basis of our comparative genome analysis, Hfq and its homologues are not found in Streptococcus. Currently known Streptococcus sRNAs do not require Hfq to function [66]. It suggests that Hfq and its homologues are not necessary for sRNA regulation in Streptococcus. How are sRNAs stabilized and annealed to the targets in Streptococcus? Generally, sRNAs are more likely to be associated with RNA-binding proteins than naked in the cellular niche [16]. Thus, whether other RNA chaperone proteins are necessary or helpful for sRNA regulation in Streptococcus needs further study.
sRNA-mediated gene silencing with an sRNA-assisted framework can provide a basis to construct artificial synthetic sRNA. The rational design of synthetic sRNAs based on natural sRNAs, can achieve high-throughput knockdown without genetic alterations; this method has been widely applied in metabolic engineering of model bacteria such as E. coli and B. subtilis [69]. Na et al. [70] have used synthetic sRNAs for combinatorial repression of four genes in 14 strains of E. coli. Repression of the expression of tyrR and csrA caused the mutant strain to produce 2 g/L of tyrosine. Moreover, repression of murE led to a 55% increase in the cadaverine titer. Meng et al. [71] have used synthetic sRNA to increase 6-deoxyerythronolide B production in E. coli, thus achieving the highest reported titer for heterologous polyketide biosynthesis. Liu et al. [72] have used an sRNA scaffold from E. coli to design synthetic sRNA to increase the yield of N-acetylglucosamine in B. subtilis. Thus, synthetic sRNA might potentially be used for gene regulation in Streptococcus. With deeper understanding of sRNAs and their regulation in Streptococcus, we believe that sRNAs could be designed and engineered as a genetic toolkit for metabolic engineering and synthetic biology in Streptococcus and other prokaryotes.
References
Zhou X, Li Y (2015) Chapter 3—supragingival microbes. In: Zhou X, Li Y (eds) Atlas of oral microbiology. Academic Press, Oxford, pp 41–65. https://doi.org/10.1016/B978-0-12-802234-4.00003-3
Le Rhun A, Charpentier E (2012) Small RNAs in streptococci. RNA Biol 9(4):414–426. https://doi.org/10.4161/rna.20104
Cunningham MW (2008) Pathogenesis of group A streptococcal infections and their sequelae. Adv Exp Med Biol 609:29–42. https://doi.org/10.1007/978-0-387-73960-1_3
Xiong ZQ, Kong LH, Meng HL, Cui JM, Xia YJ, Wang SJ, Ai LZ (2019) Comparison of gal-lac operons in wild-type galactose-positive and -negative Streptococcus thermophilus by genomics and transcription analysis. J Ind Microbiol Biotechnol 46(5):751–758. https://doi.org/10.1007/s10295-019-02145-x
Ahmed W, Hafeez MA, Ahmed R (2019) Advances in engineered trans-acting regulatory RNAs and their application in bacterial genome engineering. J Ind Microbiol Biotechnol 46:819–830. https://doi.org/10.1007/s10295-019-02160-y
Lalaouna D, Simoneau-Roy M, Lafontaine D, Masse E (2013) Regulatory RNAs and target mRNA decay in prokaryotes. Biochem Biophys Acta 182(6–7):742–747. https://doi.org/10.1016/j.bbagrm.2013.02.013
Dutta T, Srivastava S (2018) Small RNA-mediated regulation in bacteria: a growing palette of diverse mechanisms. Gene 656:60–72. https://doi.org/10.1016/j.gene.2018.02.068
Brantl S, Bruckner R (2014) Small regulatory RNAs from low-GC Gram-positive bacteria. RNA Biol 11(5):443–456. https://doi.org/10.4161/rna.28036
Choi JW, Kwon TY, Hong SH, Lee HJ (2018) Isolation and characterization of a microRNA-size secretable small RNA in Streptococcus sanguinis. Cell Biochem Biophys 76(1–2):293–301. https://doi.org/10.1007/s12013-016-0770-5
Hoekzema M, Romilly C, Holmqvist E, Wagner EGH (2019) Hfq-dependent mRNA unfolding promotes sRNA-based inhibition of translation. EMBO J 38(7):e101199. https://doi.org/10.15252/embj.2018101199
Leistra AN, Curtis NC, Contreras LM (2019) Regulatory non-coding sRNAs in bacterial metabolic pathway engineering. Metab Eng 52:190–214. https://doi.org/10.1016/j.ymben.2018.11.013
Nguyen HTB, Romero AD, Amman F, Sorger-Domenigg T, Tata M, Sonnleitner E, Blasi U (2018) Negative control of RpoS synthesis by the sRNA real in Pseudomonas aeruginosa. Front Microbiol 9:2488. https://doi.org/10.3389/fmicb.2018.02488
Mihailovic MK, Vazquez-Anderson J, Li Y, Fry V, Vimalathas P, Herrera D, Lease RA, Powell WB, Contreras LM (2018) High-throughput in vivo mapping of RNA accessible interfaces to identify functional sRNA binding sites. Nat Commun 9(1):4084. https://doi.org/10.1038/s41467-018-06207-z
Qi JK, Caiyin QG, Wu H, Tian KR, Wang BB, Li YN, Qiao JJ (2017) The novel sRNA s015 improves nisin yield by increasing acid tolerance of Lactococcus lactis F44. Appl Microbiol Biotechnol 101(16):6483–6493. https://doi.org/10.1007/s00253-017-8399-x
Rath EC, Pitman S, Cho KH, Bai Y (2017) Identification of streptococcal small RNAs that are putative targets of RNase III through bioinformatics analysis of RNA sequencing data. BMC Bioinform 18(Suppl 14):540. https://doi.org/10.1186/s12859-017-1897-0
Mraheil MA, Billion A, Kuenne C, Pischimarov J, Kreikemeyer B, Engelmann S, Hartke A, Giard JC, Rupnik M, Vorwerk S, Beier M, Retey J, Hartsch T, Jacob A, Cemic F, Hemberger J, Chakraborty T, Hain T (2010) Comparative genome-wide analysis of small RNAs of major Gram-positive pathogens: from identification to application. Microb Biotechnol 3(6):658–676. https://doi.org/10.1111/j.1751-7915.2010.00171.x
Li W, Ying X, Lu Q, Chen L (2012) Predicting sRNAs and their targets in bacteria. Genom Proteom Bioinform 10(5):276–284. https://doi.org/10.1016/j.gpb.2012.09.004
Eggenhofer F, Tafer H, Stadler PF, Hofacker IL (2011) RNApredator: fast accessibility-based prediction of sRNA targets. Nucleic Acids Res 39:W149–W154. https://doi.org/10.1093/nar/gkr467
Wang J, Liu T, Zhao B, Lu Q, Wang Z, Cao Y, Li W (2016) sRNATarBase 3.0: an updated database for sRNA-target interactions in bacteria. Nucleic Acids Res 44(D1):D248–D253. https://doi.org/10.1093/nar/gkv1127
Wright PR, Georg J, Mann M, Sorescu DA, Richter AS, Lott S, Kleinkauf R, Hess WR, Backofen R (2014) CopraRNA and IntaRNA: predicting small RNA targets, networks and interaction domains. Nucleic Acids Res 42(W1):W119–W123. https://doi.org/10.1093/nar/gku359
Muckstein U, Tafer H, Hackermuller J, Bernhart SH, Stadler PF, Hofacker IL (2006) Thermodynamics of RNA-RNA binding. Bioinformatics 22(10):1177–1182. https://doi.org/10.1093/bioinformatics/btl024
Krueger J, Rehmsmeier M (2006) RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res 34:W451–W454. https://doi.org/10.1093/nar/gkl243
Ying X, Cao Y, Wu J, Liu Q, Cha L, Li W (2011) sTarPicker: a method for efficient prediction of bacterial sRNA targets based on a two-step model for hybridization. PLoS ONE 6(7):e22705. https://doi.org/10.1371/journal.pone.0022705
Busch A, Richter AS, Backofen R (2008) IntaRNA: efficient prediction of bacterial sRNA targets incorporating target site accessibility and seed regions. Bioinformatics 24(24):2849–2856. https://doi.org/10.1093/bioinformatics/btn544
Kery MB, Feldman M, Livny J, Tjaden B (2014) TargetRNA2: identifying targets of small regulatory RNAs in bacteria. Nucleic Acids Res 42(W1):W124–W129. https://doi.org/10.1093/nar/gku317
Guillier M, Gottesman S (2008) The 5’ end of two redundant sRNAs is involved in the regulation of multiple targets, including their own regulator. Nucleic Acids Res 36(21):6781–6794. https://doi.org/10.1093/nar/gkn742
Ingolia NT, Ghaemmaghami S, Newman JR, Weissman JS (2009) Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324(5924):218–223. https://doi.org/10.1126/science.1168978
Wang J, Rennie W, Liu C, Carmack CS, Prevost K, Caron M-P, Masse E, Ding Y, Wade JT (2015) Identification of bacterial sRNA regulatory targets using ribosome profiling. Nucleic Acids Res 43(21):10308–10320. https://doi.org/10.1093/nar/gkv1158
Guo MS, Updegrove TB, Gogol EB, Shabalina SA, Gross CA, Storz G (2014) MicL, a new sigmaE-dependent sRNA, combats envelope stress by repressing synthesis of Lpp, the major outer membrane lipoprotein. Genes Dev 28(14):1620–1634. https://doi.org/10.1101/gad.243485.114
Khan MA, Gopel Y, Milewski S, Goerke B (2016) Two Small RNAs conserved in enterobacteriaceae provide intrinsic resistance to antibiotics targeting the cell wall biosynthesis enzyme glucosamine-6-phosphate synthase. Front Microbiol. https://doi.org/10.3389/fmicb.2016.00908
Rutherford ST, van Kessel JC, Shao Y, Bassler BL (2011) AphA and LuxR/HapR reciprocally control quorum sensing in vibrios. Gene Dev 25(4):397–408. https://doi.org/10.1101/gad.2015011
Jonas K, Melefors O (2009) The Escherichia coli CsrB and CsrC small RNAs are strongly induced during growth in nutrient-poor medium. FEMS Microbiol Lett 297(1):80–86. https://doi.org/10.1111/j.1574-6968.2009.01661.x
Oglesby AG, Murphy ER, Iyer VR, Payne SM (2005) Fur regulates acid resistance in Shigella flexneri via RyhB and ydeP. Mol Microbiol 58(5):1354–1367. https://doi.org/10.1111/j.1365-2958.2005.04920.x
Holmqvist E, Wagner EGH (2017) Impact of bacterial sRNAs in stress responses. Biochem Soc Trans 45(6):1203–1212. https://doi.org/10.1042/BST20160363
Froehlich KS, Gottesman S (2018) Small regulatory RNAs in the enterobacterial response to envelope damage and oxidative stress. Microbiol Spectr 6(4):1–16. https://doi.org/10.1128/microbiolspec.RWR-0022-2018
Pitman S, Cho KH (2015) The Mechanisms of virulence regulation by small noncoding RNAs in low GC Gram-positive pathogens. Int J Mol Sci 16(12):29797–29814. https://doi.org/10.3390/ijms161226194
Kreikemeyer B, Boyle MDP, Buttaro BA, Heinemann M, Podbielski A (2001) Group A streptococcal growth phase-associated virulence factor regulation by a novel operon (Fas) with homologies to two-component-type regulators requires a small RNA molecule. Mol Microbiol 39(2):392–406. https://doi.org/10.1046/j.1365-2958.2001.02226.x
Ramirez-Pena E, Trevino J, Liu Z, Perez N, Sumby P (2010) The group A Streptococcus small regulatory RNA FasX enhances streptokinase activity by increasing the stability of the ska mRNA transcript. Mol Microbiol 78(6):1332–1347. https://doi.org/10.1111/j.1365-2958.2010.07427.x
Liu Z, Trevino J, Ramirez-Pena E, Sumby P (2012) The small regulatory RNA FasX controls pilus expression and adherence in the human bacterial pathogen group A Streptococcus. Mol Microbiol 86(1):140–154. https://doi.org/10.1111/j.1365-2958.2012.08178.x
Danger JL, Cao TN, Cao TH, Sarkar P, Trevino J, Pflughoeft KJ, Sumby P (2015) The small regulatory RNA FasX enhances group A Streptococcus virulence and inhibits pilus expression via serotype-specific targets. Mol Microbiol 96(2):249–262. https://doi.org/10.1111/mmi.12935
Danger JL, Makthal N, Kumaraswami M, Sumby P (2015) The FasX small regulatory RNA negatively regulates the expression of two fibronectin-binding proteins in group A Streptococcus. J Bacteriol 197(23):3720–3730. https://doi.org/10.1128/JB.00530-15
Pappesch R, Warnke P, Mikkat S, Normann J, Wisniewska-Kucper A, Huschka F, Wittmann M, Khani A, Schwengers O, Oehmcke-Hecht S, Hain T, Kreikemeyer B, Patenge N (2017) The regulatory small RNA MarS supports virulence of Streptococcus pyogenes. Sci Rep (UK) 7:12241. https://doi.org/10.1038/s41598-017-12507-z
Sinha D, Zimmer K, Cameron TA, Rusch DB, Winkler ME, De Lay NR (2019) Redefining the sRNA transcriptome in Streptococcus pneumoniae serotype 2 strain D39. J Bacteriol. https://doi.org/10.1128/jb.00764-18
Mann B, van Opijnen T, Wang J, Obert C, Wang YD, Carter R, McGoldrick DJ, Ridout G, Camilli A, Tuomanen EI, Rosch JW (2012) Control of virulence by small RNAs in Streptococcus pneumoniae. PLoS Pathog 8(7):e1002788. https://doi.org/10.1371/journal.ppat.1002788
Acebo P, Martin-Galiano AJ, Navarro S, Zaballos A, Amblar M (2012) Identification of 88 regulatory small RNAs in the TIGR4 strain of the human pathogen Streptococcus pneumoniae. RNA 18(3):530–546. https://doi.org/10.1261/rna.027359.111
Kumar R, Shah P, Swiatlo E, Burgess SC, Lawrence ML, Nanduri B (2010) Identification of novel non-coding small RNAs from Streptococcus pneumoniae TIGR4 using high-resolution genome tiling arrays. BMC Genom 11:350. https://doi.org/10.1186/1471-2164-11-350
Wilton J, Acebo P, Herranz C, Gomez A, Amblar M (2015) Small regulatory RNAs in Streptococcus pneumoniae: discovery and biological functions. Front Genet 6:126. https://doi.org/10.3389/fgene.2015.00126
Halfmann A, Kovacs M, Hakenbeck R, Brueckner R (2007) Identification of the genes directly controlled by the response regulator CiaR in Streptococcus pneumoniae: five out of 15 promoters drive expression of small non-coding RNAs. Mol Microbiol 66(1):110–126. https://doi.org/10.1111/j.1365-2958.2007.05900.x
Marx P, Nuhn M, Kovacs M, Hakenbeck R, Bruckner R (2010) Identification of genes for small non-coding RNAs that belong to the regulon of the two-component regulatory system CiaRH in Streptococcus. BMC Genom 11:661. https://doi.org/10.1186/1471-2164-11-661
Tsui HCT, Mukherjee D, Ray VA, Sham LT, Feig AL, Winkler ME (2010) Identification and characterization of noncoding small RNAs in Streptococcus pneumoniae serotype 2 strain D39. J Bacteriol 192(1):264–279. https://doi.org/10.1128/Jb.01204-09
Zhu W, Liu S, Liu J, Zhou Y, Lin H (2018) High-throughput sequencing identification and characterization of potentially adhesion-related small RNAs in Streptococcus mutans. J Med Microbiol 67(5):641–651. https://doi.org/10.1099/jmm.0.000718
Liu S, Tao Y, Yu L, Zhuang P, Zhi Q, Zhou Y, Lin H (2016) Analysis of small RNAs in Streptococcus mutans under acid stress—a new insight for caries research. Int J Mol Sci 17(9):1529. https://doi.org/10.3390/ijms17091529
Liu SS, Zhu WH, Zhi QH, Liu J, Wang Y, Lin HC (2017) Analysis of sucrose-induced small RNAs in Streptococcus mutans in the presence of different sucrose concentrations. Appl Microbiol Biotechnol 101(14):5739–5748. https://doi.org/10.1007/s00253-017-8346-x
Lee HJ, Hong SH (2012) Analysis of microRNA-size, small RNAs in Streptococcus mutans by deep sequencing. FEMS Microbiol Lett 326(2):131–136. https://doi.org/10.1111/j.1574-6968.2011.02441.x
Xia L, Xia W, Li SH, Li WJ, Liu JJ, Ding HM, Li J, Li H, Chen Y, Su XT, Wang W, Sun L, Wang CL, Shao NS, Chu BF (2012) Identification and expression of small non-coding RNA, L10-Leader, in different growth phases of Streptococcus mutans. Nucleic Acid Ther 22(3):177–186. https://doi.org/10.1089/nat.2011.0339
Zhu WH, Liu SS, Zhuang PL, Liu J, Wang Y, Lin HC (2017) Characterization of acid-tolerance-associated small RNAs in clinical isolates of Streptococcus mutans: potential biomarkers for caries prevention. Mol Med Rep 16(6):9242–9250. https://doi.org/10.3892/mmr.2017.7751
Liu S, Li H, Guo Z, Guan J, Sun Y, Zhang K (2019) Insight into the effect of small RNA srn225147 on Mutacin IV in Streptococcus mutans. Indian J Microbiol 59(4):445–450. https://doi.org/10.1007/s12088-019-00820-2
Yin L, Zhu W, Chen D, Zhou Y, Lin H (2020) Small noncoding RNA sRNA0426 is involved in regulating biofilm formation in Streptococcus mutans. Microbiologyopen 9(9):e1096. https://doi.org/10.1002/mbo3.1096
Zhang D, Du N, Ma S, Hu Q, Lu G, Chen W, Zeng C (2014) In vitro transcriptome analysis of two Chinese isolates of Streptococcus suis serotype 2. Genom Proteom Bioinform 12(6):266–275. https://doi.org/10.1016/j.gpb.2014.11.001
Xiao GH, Tang HY, Zhang SM, Ren HY, Dai J, Lai LY, Lu CP, Yao HC, Fan HJ, Wu ZF (2017) Streptococcus suis small RNA rss04 contributes to the induction of meningitis by regulating capsule synthesis and by inducing biofilm formation in a mouse infection model. Vet Microbiol 199:111–119. https://doi.org/10.1016/j.vetmic.2016.12.034
Willenborg J, de Greeff A, Jarek M, Valentin-Weigand P, Goethe R (2014) The CcpA regulon of Streptococcus suis reveals novel insights into the regulation of the streptococcal central carbon metabolism by binding of CcpA to two distinct binding motifs. Mol Microbiol 92(1):61–83. https://doi.org/10.1111/mmi.12537
Willenborg J, Fulde M, de Greeff A, Rohde M, Smith HE, Valentin-Weigand P, Goethe R (2011) Role of glucose and CcpA in capsule expression and virulence of Streptococcus suis. Microbiology 157:1823–1833. https://doi.org/10.1099/mic.0.046417-0
Gong X, Zhuge Y, Ding C, Zheng F, Guo X, Zhang Q, Ye F, Wang C, Deng X (2019) A novel small RNA contributes to restrain cellular chain length and anti-phagocytic ability in Streptococcus suis 2. Microb Pathog 137:103730. https://doi.org/10.1016/j.micpath.2019.103730
Ota C, Morisaki H, Nakata M, Arimoto T, Fukamachi H, Kataoka H, Masuda Y, Suzuki N, Miyazaki T, Okahashi N, Kuwata H (2018) Streptococcus sanguinis noncoding cia-dependent small RNAs negatively regulate expression of type IV pilus retraction ATPase PilT and biofilm formation. Infect Immun 86(3):e00894–e00817. https://doi.org/10.1128/IAI.00894-17
Fozo EM, Makarova KS, Shabalina SA, Yutin N, Koonin EV, Storz G (2010) Abundance of type I toxin-antitoxin systems in bacteria: searches for new candidates and discovery of novel families. Nucleic Acids Res 38(11):3743–3759. https://doi.org/10.1093/nar/gkq054
Jousselin A, Metzinger L, Felden B (2009) On the facultative requirement of the bacterial RNA chaperone, Hfq. Trends Microbiol 17(9):399–405. https://doi.org/10.1016/j.tim.2009.06.003
Dos Santos RF, Arraiano CM, Andrade JM (2019) New molecular interactions broaden the functions of the RNA chaperone Hfq. Curr Genet 65:1313–1319. https://doi.org/10.1007/s00294-019-00990-y
Nielsen JS, Lei LK, Ebersbach T, Olsen AS, Klitgaard JK, Valentin-Hansen P, Kallipolitis BH (2010) Defining a role for Hfq in Gram-positive bacteria: evidence for Hfq-dependent antisense regulation in Listeria monocytogenes. Nucleic Acids Res 38(3):907–919. https://doi.org/10.1093/nar/gkp1081
Yoo SM, Na D, Lee SY (2013) Design and use of synthetic regulatory small RNAs to control gene expression in Escherichia coli. Nat Protoc 8(9):1694–1707. https://doi.org/10.1038/nprot.2013.105
Na D, Yoo SM, Chung H, Park H, Park JH, Lee SY (2013) Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat Biotechnol 31(2):170–174. https://doi.org/10.1038/nbt.2461
Meng HL, Xiong ZQ, Song SJ, Wang JF, Wang Y (2016) Construction of polyketide overproducing Escherichia coli strains via synthetic antisense RNAs based on in silico fluxome analysis and comparative transcriptome analysis. Biotechnol J 11(4):530–541. https://doi.org/10.1002/biot.201500351
Liu YF, Zhu YQ, Li JH, Shin HD, Chen RR, Du GC, Liu L, Chen J (2014) Modular pathway engineering of Bacillus subtilis for improved N-acetylglucosamine production. Metab Eng 23:42–52. https://doi.org/10.1016/j.ymben.2014.02.005
Tesorero RA, Yu N, Wright JO, Svencionis JP, Cheng Q, Kim J-H, Cho KH (2013) Novel regulatory small RNAs in Streptococcus pyogenes. PLoS ONE 8(6):e64021. https://doi.org/10.1371/journal.pone.0064021
Slager J, Aprianto R, Veening JW (2018) Deep genome annotation of the opportunistic human pathogen Streptococcus pneumoniae D39. Nucleic Acids Res 46(19):9971–9989. https://doi.org/10.1093/nar/gky725
Liu S, Zhou Y, Tao Y, Zhuang P, Pang L, Zhi Q, Lin H (2019) Effect of different glucose concentrations on small RNA levels and adherence of Streptococcus mutans. Curr Microbiol 76(11):1238–1246. https://doi.org/10.1007/s00284-019-01745-1
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31972101 and 31871776), National Science Fund for Distinguished Young Scholars (Grant No. 32025029), Natural Science Foundation of Shanghai (Grant No. 18ZR1426800), Shanghai Agriculture Applied Technology Development Program (Grant No. 2019-02-08-00-07-F01152), and Shanghai Engineering Research Center of Food Microbiology (Grant No. 19DZ2281100).
Author information
Authors and Affiliations
Contributions
ZQX and ZXL contributed equally to this work. ZQX, ZXL, and LZA conceived and wrote the draft manuscript. XS, XXL, and YJX revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest among the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xiong, ZQ., Lv, ZX., Song, X. et al. Recent Research Advances in Small Regulatory RNAs in Streptococcus. Curr Microbiol 78, 2231–2241 (2021). https://doi.org/10.1007/s00284-021-02484-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02484-y