Abstract
Bombyx mori bidensovirus (BmBDV) VD1-ORF4 consists of 3,318 nucleotides, which codes for a predicted protein with molecular weight of about 127 kDa. However, the authentic proteins encoded by VD1-ORF4 in silkworm midguts infected with BmBDV and their interacting proteins are still unclear. In this study, Western blot analysis revealed that a 127-kDa protein was confirmed to be translated from the VD1-ORF4 transcript using polyclonal antibodies and monoclonal antibodies against VD1-ORF4 deduced amino acid. Moreover, four smaller proteins with molecular weight of about 70, 60, 53, and 42 kDa were also examined in the infected midguts. Transient expression assay indicated that the expression amount of VD1-ORF4 fused with egfp was at least 30-fold lower than that of egfp gene, and immunofluorescence staining result indicated that these proteins encoded by VD1-ORF4 were located in both the cytoplasm and nucleus. Co-immunoprecipitation result showed that Aminopeptidase and Heat shock protein 90 can be captured by these proteins encoded by VD1-ORF4. In conclusion, multiple proteins were produced from the transcripts of VD1-ORF4 gene by an uncertain expression strategy, which may play important roles in viral replication and assembly.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A history of Chinese raising silkworms has been there for at least 5,000 years. Now, about thirty million farmer households in ten Chinese provinces go in for sericultural production, which plays an important role in increasing income of farmers [6]. Additionally, silkworm has become a model organism in the study of lepidopteran and arthropod biology due to its miniature size and ease of culture, and it is also a good model for studying the kinetic interaction between pathogen and host [2, 11, 14].
However, silkworms are easily suffered from the attack of bacteria, fungi, and protozoa, especially infection of some viruses, which severely impairs the production of silkworm cocoon [8, 12]. To date, Bombyx mori bidensovirus (BmBDV) remains a major pathogen resulting in flacherie disease in the silkworm, which causes great economic loss and badly contuses the enthusiasm of silkworm raisers in China [4].
BmBDV is a non-enveloped spherical DNA virus with 20–24 nm in diameter. It has narrow host range and tissue specificity (i.e., solely invades the columnar cells of silkworm midgut epithelium). Silkworm infected with BmBDV showed thorax transparency in body, anorexia, and diarrhea. Evidence showed that a gene encoding a 12-pass transmembrane protein only expressed in midgut was responsible for the binding of BmBDV to silkworm midgut [7]. However, how BmBDV interacts with this membrane protein and how host factors regulate the response to BmBDV infection remain reclusive.
The genome of BmBDV comprises two single DNA molecules VD1 (about 6.6 nts) and VD2 (about 6.0 nts), and their sequences have been published in GenBank, referred to NCBI (Accession No: DQ017268, DQ017269) since 2007 [15, 19]. Due to its genome split into two molecules and containing its own DNA polymerase motif, BmBDV was excluded from the family Parvoviridae and it was the unique representative species of the new family Bidnaviridae according to the latest version of virus taxonomy [1]. Analysis of the sequence indicated that a common 53 nts located in terminal sequence of the VD1 and VD2, and four predicted ORFs (open reading frame) were contained in the VD1 genome and two predicted ORFs were contained in the VD2 genome. RACE result indicated that VD1-ORF4 (open reading frame 4, ORF4) consists of 3,318 nucleotides encoding a predicted 1,105 amino acids protein [10]. Alignment result showed that a conserved DNA polymerase motif was contained in VD1-ORF4, which was thus regarded as a putative DNA polymerase involved in viral replication. The determination of proteins encoded by VD1-ORF4 and interactions between VD1-ORF4 proteins and host proteins are helpful to disclose the role of VD1-ORF4 gene in viral life cycle.
Materials and Methods
Plasmids, Antibodies, Viruses, and Cell Lines
Plasmids HTB-Pie1-3315(VD1-ORF4)-egfp, encoding VD1-ORF4 lacking TAA codon fused with an enhanced green fluorescent protein (EGFP) was under the control of the AcMNPV ie1 promoter, which was constructed as described in the following procedure. Briefly, ie1-F and ie1-R were used to amplify the sequence from AcMNPV ie1 promoter. The purified DNA fragment of ie1 promoter replaced the polyhedrin promoter of plasmid HTB to generate HTB-Pie1 using SnaBI and EcoRI. Primers 3315-F and 3315-R were used to amplify the sequence of VD1-ORF4 lacking TAA termination codon, and the purified fragment was ligated into HTB-Pie1 to generate HTB-Pie1-3315(VD1-ORF4) using EcoRI and PstI. The egfp was amplified with egfp-F and egfp-R, which was ligated into the plasmid HTB-Pie1-3315(VD1-ORF4) to generate the final plasmid HTB-Pie1-3315(VD1-ORF4)-egfp. Additionally, the amplified fragment of egfp was ligated into HTB-Pie1 to generate HTB-Pie1-egfp. These primers are listed in Table 1 and all plasmids were verified by sequencing.
Three mouse polyclonal antiserums against the corresponding residues from 1 to 326, 706 to 1004, 906 to 1105 of VD1-ORF4 were prepared by immunizing mice with the purified antigens. Additionally, three mouse monoclonal antibodies (directed against epitopes contained between residues 30 and 41, 977 and 988, 1,080 and 1,091 of VD1-ORF4) were prepared in Abmart (Shanghai, China). Alkaline Phosphatase horse anti-mouse immunoglobulin G (IgG) was purchased from ZSBIO corporation (Beijing, China). BmBDV was propagated in the 4th instar larvae of 306 silkworm (susceptible strain to BmBDV), which were purified by sucrose gradient centrifugation. Spodoptera frugiperda (Sf-9) cells were grown at 27 °C in Grace’s medium supplemented with 10 % fetal calf serum, penicillin (50 U/ml), and streptomycin (50 μg/ml).
Silkworm Cultivation, Virus Infection, and Preparation of BmBDV-Infected Midgut Protein Sample
Strain 306 of silkworm is highly susceptible to BmBDV, which was used in this study. After hatching, larvae of the silkworms were reared on mulberry leaves at 27 °C and 60 % relative humidity under a 12 h light/12 h dark photoperiod. When larvae were grown to the 4th instar, these larvae were fed with fresh mulberry leaves contaminated with BmBDV, and silkworms orally ingested with the virus showed some typical infection symptoms. Hundreds of BmBDV-infected silkworm were dissected and the midguts were washed three times with PBS. Then, the midguts were ground to the powder under liquid nitrogen, and RIPA lysis buffer (Beyotime, China) was added to extract the total proteins according to the manufacturer’s instructions.
Western Blotting
The protein concentrations of silkworm midgut lysates were determined by the BCA method (Pierce, USA). Thirty micrograms of each protein sample was separated by 12 % SDS-PAGE under reducing condition (1 % β-mercaptoethanol) and transferred onto a PVDF membrane (Millipore Cat. No. IPVH00010) at 250 mA for 60 min at 4 °C. Three polyclonal antibodies and three specific monoclonal antibodies against different epitopes of VD1-ORF4 deduced amino acid were respectively used as primary antibodies. After incubating with these antibodies for 1 h, these PVDF membranes were washed three times with PBST at 10 min interval, followed by incubation with 1:5,000 horseradish peroxidase (HRP) conjugated to goat anti-rat IgG (Abmart) for 1 h. Blots were then washed three times with PBST, the specific proteins were visualized by DAB kit (CWBIO, China) according to the manufacturer’s instructions.
Transient Transfection of Sf-9 Cells and Fluorescence Microscopy
Sf-9 cells (1.5 × 105) were plated into a 35-mm tissue culture dish (Corning) and allowed to attach for 24 h at 27 °C. Approximately 2 μg DNA was mixed with 6 μl Cellfectin Reagent (Invitrogen) for 45 min, then 800 μl non-serum Grace’s medium was added to the mixed DNA–cellfectin solution and overlaid onto the seeded Sf-9 cells for 6 h at 27 °C. The supernatant solution was discarded and the cells were washed twice with non-serum Grace’s medium. Finally, the cells were cultured at 27 °C in 2 ml Grace’s medium supplemented with 10 % fetal bovine serum and 100 U/ml penicillin/streptomycin. The GFP expression was examined through fluorescence microscopy after transfection of plasmid HTB-Pie1-3315(VD1-ORF4)-egfp or HTB-Pie1-egfp into Sf-9 cells.
Co-immunoprecipitation (Co-IP) Assay
The Co-IP experiment was performed to explore the interacting proteins with VD1-ORF4 in silkworm midgut. Briefly, fifty midguts were dissected out from BmBDV-infected silkworms. These midguts were washed with cold PBS (pH 7.2) and ground to a fine powder under the liquid nitrogen. RIPA lysis buffer (Beyotime, China) containing protease inhibitors was added to the powder and the resulting mixture was homogenized by ultrasonication at 2 W for 30 s on ice. The suspension was centrifuged at 10,000×g for 15 min, and the supernatant was then transferred to a fresh eppendorf tube. For immunoprecipitation, the cleared supernatant was incubated with anti-(VD1-ORF4) polyclonal antibodies (against amino acid 706–1,004 or 906–1,105 of VD1-ORF4) for 1 h, followed by incubation with 100 μl of protein A beads (Santa Cruz Biotechnology, CA, USA) for 4 h at 4 C. Additionally, the supernatant incubated with negative mice serum coupled with protein A beads was used as blank control. After centrifugation, the immunoprecipitate was washed four times with RIPA buffer, and resuspended in 50 μl of SDS-PAGE loading buffer and heated for 5 min at 100 °C. Finally, the treated suspension was subjected to 12 % SDS-PAGE analysis and visualized by Coomassie blue staining.
In Gel Digestion and Peptide Analysis by MALDI-TOF
To identify the proteins interacting with VD1-ORF4, the protein strips were extracted from the gel and digested with sequencing grade trypsin. The resulting peptides were then extracted twice in 100 μl of 60 % (v/v) ACN and 0.1 % (v/v) Trifluoroacetic acid (TFA), which was subsequently analyzed by liquid chromatography-tandem mass spectrometry (LC–MS/MS) (AB SCIEX). Masses of tryptic peptides were searched against the database NCBInr, via the program MASCOT.
Results
Identification of Proteins Translated from VD1-ORF4
According to the RACE result reported by Li [10], VD1-ORF4 was regarded to be consisted of 3,318 nucleotides. Sequence analysis indicated that there were 18 potential translation initiation condons of ATG in the nucleotide sequence of VD1-ORF4. So, the VD1-ORF4 transcript was postulated to translate multiple proteins by leaky ribosomal scanning. To determine the exact expression of VD1-ORF4 transcript in BmBDV-infected midgut, the lysates from BmBDV-infected midgut were subjected to Western blotting analysis using three different polyclonal antibodies (respectively directed against an epitope contained residues between 1 and 326, or 706 and 1,004, or 906 and 1,105). The results showed that a specific protein with molecular weight of about 127 kDa was detected in the lysate of BmBDV-infected midgut using each of these three antibodies. Moreover, another specific protein about 42 kDa was also examined in the lysate of BmBDV-infected midgut using the antiserum against the epitope contained between amino acids 1 and 326, and three other proteins with molecular weight of about 70, 60, and 53 kDa were detected in the lysate of BmBDV-infected midgut using the antiserum against the antigen between amino acids 706 and 1,004.
To further confirm the expression of VD1-ORF4 in BmBDV-infected midgut, three mouse monoclonal anti-VD1-ORF4 (directed against an epitope contained between residues 30 and 41, 977 and 988, 1,080 and 1,091) were used as primary antibodies to incubate with bolt membrane. The results were similar to those obtained with the three polyclonal antiserums. Besides the specific 127-kDa protein, another specific protein about 53 kDa was examined in the lysate of BmBDV-infected midgut using the monoclonal antibodies against the epitope contained between residues 1,080 and 1,091 (Fig. 1).
Western blot analysis of the BmBDV-infected silkworm midgut. a Three polyclonal antibodies were used as primary antibodies. Lane 1, The midgut protein from healthy silkworm was used as blank control, lane 2–4, Specific proteins were examined using antibodies against the epitope corresponding to amino acids 1–326, 706–1,004, 906–1,105 of VD1-ORF4 deduced amino acid, respectively. b Three mouse monoclonal antibodies were used as primary antibodies. Lane 1–3: Specific proteins were examined using antibodies against the epitope corresponding to amino acids 30–41, 977–988, 1,080–1,091 of VD1-ORF4, respectively. The migration of molecular mass markers (in kilodaltons), as well as specified polypeptides, is indicated
Analysis of Transient Expression of VD1-ORF4 Fused with egfp
To further reveal the expression strategy of VD1-ORF4, we attempted to examine the expression product of the VD1-ORF4 expressed in Sf-9 cells. The recombinant virus harboring VD1-ORF4 under the control of polyhedrin promoter was prepared and used to infect Sf-9 cells. However, target proteins were not examined in the total protein of virus-infected Sf-9 cells using antibodies against the epitope of VD1-ORF4 deduced amino acid. Therefore, it was urgent to determine whether VD1-ORF4 was expressed in cultured insect cells or not. For this purpose, we constructed the transient expression plasmid of HTB-Pie1-3315(VD1-ORF4)-egfp, in which egfp fused with the 3′-terminus of VD1-ORF4 deleting TAA was under the control of AcMNPV ie1 promoter.
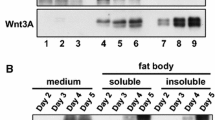
The recombinant plasmid HTB-Pie1-3315(VD1-ORF4)-egfp was extracted and transfected into Sf-9 cells. The results indicated that little green fluorescence were visualized in very few cells from 24 to 96 h p.t. At the same time, the transfection of HTB-Pie1-egfp into Sf-9 cells was used as control. The result showed that green fluorescent signal appeared in some Sf-9 cells at 24 h p.t., and the green fluorescence signal was sharply intensified from 48 to 96 h p.t. (Fig. 2).
Subcellular Localization of Proteins Encoded by VD1-ORF4 in Sf-9 Cells
The expression and the subcellular localization of proteins encoded by VD1-ORF4 were examined by immunofluorescence staining. The staining results showed that fluorescence was observed in the Sf-9 cell cytoplasm at 24, 48, or 72 h p.t., and a relatively little fluorescence was also present in the nucleus from 48 to 72 h p.t. In control experiment, no obvious fluorescene was observed but blue nucleus stained with DAPI. The above results revealed that these proteins encoded by VD1-ORF4 localized both in the cytoplasm and nucleus (Fig. 3).
Subcellular localization of proteins encoded by VD1-ORF4 in Sf-9 cells transfected with pFastHTB-Pie1-3318(VD1-ORF4). Sf-9 cells were treated with monoclonal antibodies against amino acid 977–988 of VD1-ORF4 deduced amino acid, followed by treatment with secondary antibody conjugated-FITC and the nucleus was treated with DAPI, and examined by confocal microscope. From the left to the right: DAPI-treated nucleus, green fluorescence for FITC-treated proteins encoded by VD1-ORF4 and the merged images. The times after transfection were indicated on the left
Identification of the Interacting Proteins with VD1-ORF4
To identify the interacting proteins with the proteins encoded by VD1-ORF4, Co-IP experiment was performed to pull down the target protein using antibodies against VD1-ORF4. The protein–protein complexes were subjected to 12 % SDS-PAGE analysis for separation of target proteins. The result indicated that three specific protein bands were visualized by staining the gel with Coomassie brilliant blue. LC–MS/MS results revealed that No. 1 protein was the 127-kDa protein encoded by VD1-ORF4, No. 2 protein was Aminopeptidase, and No. 3 protein was Heat shock protein 90 (Fig. 4). At the same time, no protein was pulled down in the negative serum, which was used as negative control.
Midgut lysate from BmBDV-infected silkworms was subjected to Co-IP using anti-VD1-ORF4 antibodies. Precipitated proteins were separated by 12 % SDS-PAGE and visualized by Coomassie blue staining. Lane 1, Negative serum was used as blank control; Lane 2–3, Positive antibodies against 706–1,004, 906–1,105 of VD1-ORF4 deduced amino acid was used in the study, respectively. The potential proteins interacting with proteins encoded by VD1-ORF4 were indicated with arrow
Discussion
Study of the interactions between BmBDV and silkworm is helpful for discovering unique antiviral defense and providing valuable insight into the evolution of immune system in insects. It becomes important to determine the exact expression of viral genes in viral life cycle. For this purpose, this article focuses on the identification of authentic expression of BmBDV VD1-ORF4 gene and the interactions between proteins encoded by VD1-ORF4 and the host.
Previous researches revealed that a conserved motif of DNA polymerase was contained in the VD1-ORF4 deduced amino acid [10, 20], which was regarded to be a novel DNA polymerase involved in viral replication. However, we failed to express the full-length sequence of VD1-ORF4 using baculovirus expression vector system (BEVS), which severely blocked the further studies of its biochemical function in vitro. To facilitate the functional research of VD1-ORF4, some specific antibodies were raised to determine the authentic expression of VD1-ORF4 and examine the subcellular distribution of proteins encoded by VD1-ORF4 in viral life cycle. Interestingly, a 127-kDa protein was detected in BmBDV-infected midguts using each of six antibodies against different epitope of VD1-ORF4 deduced amino acid, and four smaller proteins with molecular weight of about 70, 60, 53, and 42 kDa were also examined in BmBDV-infected midguts using the monoclonal antibodies against the epitope contained between amino acids 977 and 988. Additionally, a small protein with molecular weight of about 42 kDa was detected in BmBDV-infected midguts using the antibodies against the N-terminal epitope, and another small protein with molecular weight of about 53 kDa was detected in BmBDV-infected midguts using the antibodies against the C-terminal epitope. However, the synthesis mechanisms of these proteins and their roles in viral life cycle remain unknown.
Due to the small size and compactness of viral genomes, a lot of viruses have been reported to rely on other strategies to expand their genetic information and optimize usage of the available sequences. For example, there are a number of cases of leaky scanning, including PB1-F2 and N40 translated from influenza A virus segment 2 mRNA [16], 4 capsid proteins of the Junonia coenia densovirus (JcDNV), and VP1–VP4 of GmDNV translated from 2.3 kb vp transcript [13, 17]. Moreover, the viral genome of Hepatitis C virus (HCV) codes for a polyprotein, which is cleaved by viral and cellular proteases to produce at least ten mature viral protein products [18]. Gal-Pol polyprotein of HIV-1 was synthesized by programed ribosomal frameshift, which was posttranslationally cleaved into two functional proteins [9]. Besides that, some viruses were reported to utilize other mechanisms such as internal ribosome entry, non-AUG initiation, ribosome shunting, and reinitiation, etc. [5].
To escape the intense selective pressure, BmBDV had evolved appropriate strategies to deal with some intricate events in viral life cycle. Owing to the unique character and relative compactness of BmBDV genome, we regarded that it utilized multiple unknown mechanisms to maximize genomic coding capacity, especially the large ORF of VD1-ORF4. Therefore, we hypothesized that a large protein translated from the VD1-ORF4 mRNA was digested by viral and cellular proteases to generate these mature proteins in the study. For this purpose, the software of ExPASy http://web.expasy.org/findpept/ was used to analyze whether these protein products were generated from the VD1-ORF4 precursor by digestion. However, the bioinformatic result indicated that it was not possible to generate these proteins in such a way. Therefore, we raised the possibility that these proteins were translated from VD1-ORF4 mRNA by ribosomal leaky scanning or non-AUG initiation.
To confirm the hypothesis, a series of VD1-ORF4 mutants were used for the expression in Sf-9 cells using baculovirus expression vector systemn (BEVS). Unfortunately, we did not examine the expression of VD1-ORF4 using specific antibodies (data not shown). Fluorescence microscopy showed that weak fluorescent signal was in Sf-9 cells transfected with HTB-Pie1-3315(VD1-ORF4)-egfp, but strong fluorescent signal was in Sf-9 cells transfected with HTB-Pie1-egfp. It indicated that the genuine expression of VD1-ORF4 may need the assistance of certain specific factors from BmBDV-infected midguts, which is in agreement with the fact that BmBDV solely infect the epithelium cells of silkworm midgut. Therefore, our next work is to establish the cell line of silkworm midgut, which is necessary to disclose the expression strategy of VD1-ORF4 and their potential roles of these proteins in viral life cycle.
To date, it remains largely unknown about the infection mechanism of BmBDV in its host silkworm. The identification of the specific interactions between host proteins and BmBDV proteins might help elucidate the mechanisms of viral proliferation in host midgut, and it could also shed new lights on how immune mechanisms respond to viral infections in silkworm. Previously it was reported that Trypsin-like protease, 30 kP protease A, Carboxypeptidase, and Translation elongation factor 2 were identified to interact with VD1-ORF4 via a yeast two-hybrid (Y2H) system. However, these interactions were not examined in vivo by bimolecular fluorescence complementation (BiFC) assay [3], which was regarded as false findings. So, we attempted to explore the interactions between these proteins encoded by VD1-ORF4 and silkworm by Co-immunoprecipitation. MS result indicated that the 127-kDa protein encoded by BmBDV ORF4, Aminopeptidase, and Heat shock protein 90 were identified to potentially interact with the proteins encoded by BmBDV ORF4. These interactions need further confirmation by bimolecular fluorescence complementation (BiFC) assay in vivo, and further research was required to examine the functions of these host proteins in antiviral defense.
References
Adams MJ, Carstens EB (2012) Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2012). Arch Virol 157:1411–1422
Arunkumar KP, Tomar A, Daimon T et al (2008) WildSilkbase: an EST database of wild silkmoths. BMC Genom 9:338
Bao YY, Chen LB, Wu WJ et al (2013) Direct interactions between bidensovirus BmDNV-Z proteins and midgut proteins from the virus target Bombyx mori. FEBS J 280:939–949
Bao YY, Li MW, Zhao YP et al (2008) Differentially expressed genes in resistant and susceptible Bombyx mori strains infected with a densonucleosis virus. Insect Biochem Mol Biol 38:853–861
Firth AE, Brierley I (2012) Non-canonical translation in RNA viruses. J Gen Virol 93:1385–1409
Goldsmith MR, Shimada T, Abe H (2005) The genetics and genomics of the silkworm, Bombyx mori. Annu Rev Entomol 50:71–100
Ito K, Kidokoro K, Sezutsu H et al (2008) Deletion of a gene encoding an amino acid transporter in the midgut membrane causes resistance to a Bombyx parvo-like virus. Proc Natl Acad Sci USA 105:7523–7527
Khurad AM, Mahulikar A, Rathod MK et al (2004) Vertical transmission of nucleopolyhedrovirus in the silkworm, Bombyx mori L. J Invertebr Pathol 87:8–15
Kobayashi Y, Zhuang J, Peltz S et al (2010) Identification of a cellular factor that modulates HIV-1 programmed ribosomal frameshifting. J Biol Chem 285:19776–19784
Li G, Hu Z, Guo X et al (2013) Identification of Bombyx mori bidensovirus VD1-ORF4 reveals a novel protein associated with viral structural component. Curr Microbiol 66:527–534
Papanicolaou A, Gebauer-Jung S, Blaxter ML et al (2008) ButterflyBase: a platform for lepidopteran genomics. Nucleic Acids Res 36:582–587
Rahman MM, Gopinathan KP (2004) Systemic and in vitro infection process of Bombyx mori nucleopolyhedrovirus. Virus Res 101:109–118
Tijssen P, Li Y, El-Far M et al (2003) Organization and expression strategy of the ambisense genome of densonucleosis virus of Galleria mellonella. J Virol 77:10357–10365
Taniai K, Lee JH, Lee IH (2006) Bombyx mori cell line as a model of immune-system organs. Insect Mol Biol 15:269–279
Wang YJ, Yao Q, Chen KP et al (2007) Characterization of the genome structure of Bombyx mori densovirus (China isolate). Virus Genes 35:103–108
Wise HM, Foeglein A, Sun J et al (2009) A complicated message: identification of a novel PB1-related protein translated from influenza A virus segment 2 mRNA. J Virol 83:8021–8031
Wang Y, Abd-Alla AM, Bossin H et al (2013) Analysis of the transcription strategy of the Junonia coenia densovirus (JcDNV) genome. Virus Res 174:101–107
Xu Z, Choi J, Yen TS et al (2001) Synthesis of a novel hepatitis C virus protein by ribosomal frameshift. EMBO J 20:3840–3848
ZhaoYang Hu, Li GuoHui, Li GuangTian et al (2013) Bombyx mori bidensovirus: the type species of the new genus Bidensovirus in the new family Bidnaviridae. Chin Sci Bull 58:4528–4532
Zhang J, Li G, Chen H et al (2010) Molecular Cloning and expression of key gene encoding hypothetical DNA polymerase from Bombyx mori parvo-like virus (China isolate). Genet Mol Biol 33:739–744
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31000080, 31270192, 31272507 and 31402016), the National basic Research Program of China under Grant No. 2012CB114604, and the Startup Scientific Research Fund from Jiangsu University for Senior Professionals (No. 09JDG057).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, G., Zhou, Q., Hu, Z. et al. Determination of the Proteins Encoded by BmBDV VD1-ORF4 and Their Interacting Proteins in BmBDV-Infected Midguts. Curr Microbiol 70, 623–629 (2015). https://doi.org/10.1007/s00284-014-0765-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-014-0765-7