Abstract
Arsenic trioxide (ATO) treatment effectively prolongs the overall survival of patients with acute promyelocytic leukemia (APL). Mutations in the oncogene PML::RARA were found in patients with ATO-resistant and relapsed APL. However, some relapsed patients do not have such mutations. Here, we performed microarray analysis of samples from newly diagnosed and relapsed APL, and found different microRNA (miRNA) expression patterns between these two groups. Among the differentially expressed miRNAs, miR-603 was expressed at the lowest level in relapsed patients. The expression of miR-603 and its predicted target tropomyosin-related kinase B (TrkB) were determined by PCR and Western blot. Proliferation was measured using an MTT assay, while apoptosis, cell cycle and CD11b expression were analyzed using flow cytometry. In APL patients, the expression of miR-603 was negatively correlated with that of TrkB. miR-603 directly targeted TrkB and downregulated TrkB expression in the APL cell line NB4. miR-603 increased cell proliferation by promoting the differentiation and inhibiting the apoptosis of NB4 cells. This study shows that the miR-603/ TrkB axis may be a potent therapeutic target for relapsed APL.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute promyelocytic leukemia (APL) accounts for approximately 10–15% of adult acute myeloid leukemias and is characterized by a specific reciprocal translocation, t(15;17)(q22;q21), which results in the formation of the promyelocytic leukemia (PML) and retinoic acid receptor-α (RARα) fusion gene [1]. The PML::RARA fusion protein leads to differentiation blockade at the promyelocytic stage. Arsenic trioxide (ATO) acts on the PML::RARA fusion protein and reverses the inhibition of cellular differentiation mediated by this oncoprotein. The clinical application of ATO and all-trans retinoic acid (ATRA) has converted APL from a fatal to a highly curable hematological malignancy [2]. However, notably, approximately 5–6% of newly diagnosed APL patients cannot achieve complete remission, and approximately 5–20% experience relapse [3]. Several studies have shown additional genetic and epigenetic processes that accompany the expression of the PML::RARA protein [4, 5].
MicroRNAs (miRNAs) are endogenous, non-protein-coding small RNAs that mediate the regulation of target gene expression at the posttranscriptional level by inducing translational repression or mRNA degradation [6]. miRNAs have been shown to play key roles in hematopoietic differentiation as well as in the formation, maintenance and progression of leukemia [7, 8]. Several studies have shown that some miRNAs, such as miR-223, miR-34 and miR-30c, play important roles in myeloid differentiation [9,10,11,12]. miR-223 has been implicated in APL differentiation and tumorigenesis [13]. However, knowledge about the expression and function of other miRNAs during disease progression, such as relapse after treatment with ATO, remains lacking.
Tropomyosin-related kinase B (TrkB), a member of the TRK family, is encoded by the neurotrophic tyrosine kinase receptor type 2 (NTRK2) gene and acts as a receptor for brain-derived neurotrophic factor (BDNF) [14]. The binding of BDNF to TrkB activates several downstream signaling pathways, including the PI3K/AKT and JAK/STAT pathways [15]. There is increasing evidence for involvement of TrkB in normal hematopoiesis and leukemogenesis [16,17,18].TrkB expression is greatest in precursor CD4−CD8− thymocytes and progressively declines throughout the T-cell differentiation pathway [16]. TrkB plays a crucial role in B-cell chronic lymphocytic leukemia cell survival by interacting with neurotension receptor 2, and then activates survival signaling pathways and expression of the anti-apoptotic proteins [17]. Coexpression of TrkB/BDNF in murine hematopoitic cells induced leukemia [18]. TrkB is highly expressed in acute leukemia, and coexpression of TrkB and BDNF is related to poor prognosis [18]. However, the expression of TrkB have not been reported in relapsed APL.
In this study, we showed different miRNA expression patterns in patients with relapsed APL compared with newly diagnosed patients by microarray analysis. Among the differentially expressed miRNAs, miR-603 was the most significant. TrkB was indicated by bioinformatic analysis and a luciferase activity assay to be a target of miR-603. Furthermore, we found that miR-603 increased cell proliferation by promoting differentiation and inhibiting apoptosis and G1 arrest in the APL cell line NB4.
Materials and methods
Patient samples
From May 2015 to December 2017, bone marrow samples were collected from patients with newly diagnosed and hematologic relapsed APL (according to the WHO diagnostic criteria) admitted to the Department of Hematology at the First Affiliated Hospital of Harbin Medical University. Written informed consent and approval from the Ethical Committee of Human Experimentation in Harbin were obtained in accordance with the current version of the Declaration of Helsinki. Mononuclear cells were harvested by Ficoll-Hypaque density gradient centrifugation.
miRNA evaluation by quantitative PCR-based array analyses
Using the mirVana miRNA Isolation Kit, total RNA enriched with miRNA was isolated and reverse transcribed using the TaqMan MicroRNA Reverse Transcription Kit (Life Technologies, USA). A TaqMan Low-Density Array (TLDA) was used for miRNA analysis in a 7900HT Real-Time PCR system. The expression of miRNAs was quantified using TaqMan Array Human MicroRNA A + B Cards v3.0 (Life Technologies, USA) with a total of 384 miRNAs and controls per card. The relative expression levels were calculated from the measured Ct values using the 2−ΔΔCT method [19]. The P value (P < 0.05) and fold change (FC > 2) thresholds were set for the comparison of miRNA expression between different groups of samples.
PCA; GO and KEGG functional enrichment analyses
Principal component analysis (PCA) was performed as previously reported [20]. The principal components are represented by the 26 miRNAs identified as differentially expressed by array analysis listed in Table 1. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses were performed using DIANA-miRPath v. 3.0. Analyses were performed using the human orthologs of the miRNAs with an expression fold change of at least ± 10 based on the microarray analysis. Predicted gene targets with significant enrichment for these miRNAs based on TarBase v. 7.0 were used. Heatmaps were generated to show the clusters with significant pathways or overlapping categories defined using Fisher's exact test. The level of significance was set at P < 0.05.
Cell culture and transfection
Human APL cell lines: NB4 cells were kindly provided by Shanghai Cell Bank, Chinese Academy of Sciences (Shanghai, China). Cells were maintained in RPMI 1640 culture medium containing 10% fetal bovine serum (FBS), 100 U/mL penicillin and 100 U/ml streptomycin and were incubated in a humidified incubator containing 5% CO2 at 37 °C. Cells were plated in growth medium without antibiotics for approximately 24 h before transfection. Transient transfection of precursor miRNAs (Ambion)/siRNAs (Origene, USA) was carried out using Lipofectamine 2000 (Invitrogen) according to the manufacturer's protocol. Cells were harvested at different time points. Transfection reagent alone was used as a control. For the luciferase activity assay, the 293 T cell line was maintained in DMEM containing 10% fetal bovine serum.
Luciferase activity assay
To construct the luciferase reporter plasmid, the target fragment (TrkB) was inserted into the psiCHECK™-2 vector (Promega, WI, USA). 293 T cells were seeded in 24-well plates at a density of 2 × 104 cells/well. The next day, the culture medium was replaced with 300 µL of Opti-MEM. In each well, 1 µL of wild-type (WT) or mutated (MUT) miRNA-603 (20 µM) was cotransfected into cells along with 0.5 µg of luciferase reporter plasmid using Lipofectamine 2000 transfection reagent according to the manufacturer's instructions (Invitrogen, Grand Island, NY, USA). Forty-eight hours after transfection, luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega). The relative luciferase intensity was determined. Each experimental condition was analyzed in triplicate.
PCR
Quantitative reverse transcription PCR (q-RT‒PCR) and quantitative PCR (Q-PCR) were used to validate the miRNA and mRNA expression data. The reactions were performed in an ABI 7900 HT Thermal Cycler (Applied Biosystems, Stockholm, Sweden). Relative miRNA expression levels (miRNA vs. U6 and mRNA vs. GAPDH) were calculated with the 2−ΔΔCT method [16]. All primers were obtained from Life Technologies (Grand Island, NY) and are listed in Table 2. All reactions were performed in triplicate.
Western blot analysis
Total protein was extracted from cells after lysis with RIPA lysis buffer (Beyotime, China). Protein concentration was determined using a BCA Protein Assay Kit (Beyotime, China). Equal amounts of protein samples were subjected to SDS-PAGE and transferred onto polyvinylidene fluoride membranes that were blocked with 5% milk powder containing 0.1% TBS-Tween-20, and incubated with specific primary antiboides against TrkB (1:2000, ab187041; Abcam, Shanghai, China). An anti-GAPDH antibody (1:5000, ab128915; Abcam) obtained from Sigma‒Aldrich was used as a loading control.
Cell proliferation assay
To evaluate cell proliferation, cells were seeded into a 96-well plate at a density of 1 × 104 cells/well. A total of 10 μL of CCK-8 solution was added to 100 μL of culture medium. The absorbance of the culture medium was measured at 450 nm (A450) using a scanning microplate spectrophotometer (Multiscan MK3, Thermo Fisher Scientific).
Flow cytometry analysis
For cell cycle analysis, cells were fixed overnight in chilled methanol prior to staining with 50 μg/ml propidium iodide (PI, Sigma‒Aldrich) in the presence of 1 mg/ml RNase (100 units/ml; Sigma‒Aldrich) and 0.1% NP40 (Sigma‒Aldrich). For apoptosis analysis, samples were incubated with Annexin V-fluorescein isothiocyanate (FITC)/PI according to the manufacturer’s protocol (Sigma‒Aldrich). For CD11b expression, samples were incubated with anti-CD11b-FITC antibody (Sigma-Aldrich). Cellular fluorescence was analyzed on a FACSCalibur flow cytometer (Becton Dickinson, CA, USA) with Cellquest software.
Statistical analysis
Significant differences in miRNA expression between samples from patients with newly diagnosed and relapsed APL were assessed using the Wilcoxon rank–sum test with correction for multiple comparisons with the Benjamini–Hochberg false discovery rate (FDR). SPSS 17.0 was used for statistical analysis. All results were obtained from at least three separate experiments. The data are expressed as the means ± SDs. The expression levels of miRNA-603 and TrkB in samples were analyzed by ANOVA or a t test. Two-tailed tests were used for univariate comparisons. P < 0.05 was considered statistically significant. Correlation analysis between miR-603 and TrkB mRNA expression was performed by Spearman correlation analysis.
Results
miRNAs are differentially expressed in samples from newly diagnosed patients and patients with relapse
To investigate the involvement of miRNAs in relapsed APL after treatment with ATO, TaqMan miRNA assays were performed to analyze the miRNA expression profiles in samples from 3 patients with relapsed APL(without PML::RARA mutation) and 5 patients with newly diagnosed APL. miRNAs with a fold changes ≥ 2.0 and P values ≤ 0.05 were considered significantly differentially expressed. In patients with relapsed APL compared with newly diagnosed patients, miR-214 was highly expressed, and twenty-five other miRNAs were expressed at low levels, with miR-603 exhibiting the lowest level (Table 1). The hierarchical clustering heatmap revealed homogenous but distinctive expression patterns for each sample, with all samples from patients with relapse assigned to a cluster distinct from that of newly diagnosed APL (Fig. 1A). These data were consistent with the PCA data, in which the newly diagnosed and relapsed groups were completely separated (Fig. 1B).
Different expression patterns of miRNAs and enriched GO and KEGG pathways in APL. A Hierarchical clustering of miRNA expression in APL patients. miRNA expression profiles of samples from 3 relapsed (R) and 5 newly diagnosed (N) patients. The samples are shown in the columns; the miRNAs, in the rows. ΔCT values were used in the analysis. B PCA of miRNA expression in APL patients. The blue dots indicate samples from relapsed patients; the yellow dots, samples from newly diagnosed patients. The significant major GO processes C and KEGG pathways D predicted for the gene targets of the 10 most altered miRNAs. Log P values are plotted, with greater significance indicated in red. GO: Gene Ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes
GO term and KEGG pathway enrichment analyses
To clarify the biological process related to APL recurrence, GO and KEGG pathway enrichment analyses were performed with the top 10 dysregulated miRNAs in this study. The GO analysis results revealed that these miRNAs participated in vital biological processes, including small metabolic process and neurotrophin TRK receptor signaling pathway (Fig. 1C). The most significant KEGG pathway was prion diseases (Fig. 1D). For miR-603, the most significant GO biological process term was neurotrophin TRK receptor signaling pathway, and the most significant KEGG pathway was hippo signaling pathway.
miR-603 and TrkB expression in APL patients
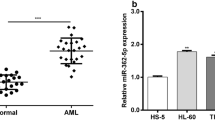
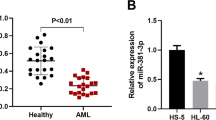
TrkB, a member of the TRK family, plays an important role in the initiation and progression of solid tumors and leukemias [16,17,18, 21, 22]. It was identified as one of the potential targets of miR-603 in the neurotrophin TRK receptor signaling pathway by several algorithms (miRDB, miRanda and TargetScan). First, we measured miR-603 and TrkB mRNA expression in array samples by PCR. Compared with that in patients with newly diagnosed APL, miR-603 expression in relapsed patients was low, while TrkB expression in relapsed patients was high (Fig. 2A and B). Then, miR-603 and TrkB mRNA expression was measured in three healthy volunteers, in ten patients with newly diagnosed APL and in seven patients with relapsed APL treated with ATO. Compared with that in the healthy volunteers, miR-603 expression was low in APL patients, especially in those with relapse. TrkB was highly expressed in samples from patients with relapse compared with samples from healthy volunteers and newly diagnosed patients but did not differ significantly between the control volunteers and newly diagnosed patients (Fig. 2C and D). Spearman correlation analysis showed that miR-603 expression was negatively correlated with TrkB expression in control volunteers, newly diagnosed patients and patients with relapsed APL (Fig. 2E and F).
miR-603 and TrkB mRNA expression levels in APL, as evaluated by PCR. A and B miR-603 expression was low (A) and TrkB expression was high (B) in samples from patients with relapsed APL compared with samples from newly diagnosed patients in the array. C and D Compared with that in healthy volunteers, miR-603 expression was low in APL patients, especially relapsed patients (C). TrkB was highly expressed in APL patients, especially in relapsed patients (D). E and F The miR-603 and TrkB mRNA expression levels were negatively correlated among the healthy volunteers, newly diagnosed patients and relapsed patients overall (E) and in each comparison (F). * indicates P < 0.05. # indicates P < 0.05 compared with newly diagnosed patients (C and D)
miR-603 targets TrkB by binding the 3' UTR
The relationship between miR-603 and TrkB was subsequently demonstrated by a luciferase activity assay. Because miR-603 was predicted to bind to the 3'-untranslated region (UTR) of TrkB by miRanda (Fig. 3A), we constructed firefly luciferase reporter vectors containing either the entire WT TrkB 3'-UTR or a mutant version (MUT) of the TrkB 3'-UTR. Cotransfection of miR-603 and the luciferase reporter with the WT TrkB 3'-UTR greatly reduced luciferase activity (Fig. 3B). No significant difference in luciferase activity was found in cells transfected with the MUT TrkB 3'-UTR. Then, we measured the TrkB mRNA and protein expression levels in NB4 cells transfected with either a miR-603 mimic or a miR-603 inhibitor. The miR-603 mimic significantly decreased the TrkB mRNA and protein levels, whereas the miR-603 inhibitor markedly increased TrkB expression in these cells (Fig. 3C and D). This increased expression of TrkB was inhibited by si-TrkB. These data indicate that miR-603 directly binds to the 3'-UTR of TrkB and significantly downregulates its mRNA and protein expression.
miR-603 downregulates TrkB. A Predicted binding sites for miR-603 and TrkB. B Luciferase assay results showing the decrease in relative luciferase activity in 293 T cells cotransfected with miR-603 and TrkB. C and D The TrkB mRNA (C) levels and protein expression levels (D-E) changed after transfection with the miR-603 mimic or miR-603 inhibitor. D The graphs show the fold changes in protein expression relative to the untreated control, as measured by densitometry. The data are presented as the mean ± SD of three different experiments. WT: wild-type; MUT: mutated; NC: negative control. *: P < 0.05; **: P < 0.01
miR-603 promotes cell proliferation
Subsequently, we evaluated the effect of miR-603 on NB4 cell proliferation. As shown in Fig. 4A, the miR-603 mimic promoted but the miR-603 inhibitor suppressed the proliferation of NB4 cells, and these effects were partially reversed by si-TrkB. To explore the mechanism(s) by which miR-603 affects the growth of NB4 cells, apoptosis, the cell cycle and differentiation were examined. The number of apoptotic cells decreased in response to miR-603 mimic treatment but increased in response to miR-603 inhibitor treatment, and these effects were decreased by si-TrkB (Fig. 4B). The proportions of NB4 cells in the G1 and G2 phases were increased by the miR-603 inhibitor but were not affected by the miR-603 mimic (Fig. 4C). CD11b expression was increased by the miR-603 mimic but decreased by the miR-603 inhibitor (Fig. 4D). These results indicated that miR-603 promotes the growth of the NB4 cells by increasing cell differentiation and inhibiting apoptosis.
Influence of miR-603 and TrkB expression on cell proliferation, apoptosis, the cell cycle and differentiation. A The miR-603 mimic increased cell proliferation, and the miR-603 inhibitor decreased cell proliferation; these effects were partially reversed by si-TrkB. B/E The miR-603 mimic decreased apoptosis, while the miR-603 inhibitor increased the number of apoptotic NB4 cells; these effects were partially decreased by si-TrkB. C/F The miR-603 inhibitor significantly increased the proportion of NB4 cells in G1 phase. D/G The miR-603 mimic increased CD11b expression in NB4 cells. Nontransfected cells were used as controls. Three independent experiments were performed. The data are presented as the means ± SDs. *: P < 0.05; **: P < 0.01
Discussion
As ATO has become a first-line therapeutic strategy, identification of the mechanisms underlying APL relapse in patients treated with ATO is urgently needed. Studies have focused on mutations in the oncogenic PML::RARA gene, which can result in acquired resistance to ATO and have been found in relapsed APL [23,24,25,26]. However, some patients who relapse after treatment with ATO-based regimens do not have such mutations. Accumulating have shown the importance of miRNAs in the initiation, maintenance and progression of leukemia. For example, a specific miRNA signature of APL blasts at diagnosis that differs from that of normal promyelocytes has been reported [27]. In addition, several recent studies have shown that some miRNAs are promising indicators of relapse and correlate with poor prognosis in acute leukemia [28,29,30]. In the present study, we characterized a small number of miRNAs differentially expressed in patients with relapsed APL who were treated with ATO compared with newly diagnosed patients.
Among these differentially expressed miRNAs, miR-603 had the most significant difference in expression. miR-603 has been revealed to play different roles in different tumors. miR-603 has been reported to act as a tumor suppressor in breast cancer [31] but as an oncomiR in glioma, osteosarcoma, and hepatocellular carcinoma [32, 33]. In this study, we verified that miR-603 induced malignant behaviors in an APL cell line by increasing cell proliferation and inhibiting apoptosis. However, miR-603 also promoted NB4 cell differentiation by increasing CD11b expression. In addition, in NB4 cells, miR-603 silencing led to G1 arrest and inhibition of differentiation, which is a main mechanism by which ATO cures APL.
TrkB was identified as a target of miR-603 in this study. As a member of TRK family, TrkB is an important regulator of cell proliferation, differentiation and survival [15]. Accumulating evidence highlights the role of TrkB in chemoresistance and recurrence of both solid and liquid maligancies [17, 34, 35]. Consistent with these previous studies, we also found that TrkB is overexpressed in relapsed APL patients. However, there was no significant difference in the expression of TrkB between newly diagnosed APL patients and healthy volunteers in this study. These results are consistent with the a previous published report and TCGA data [data not show] [18]. This may be partially due to the small number of patients and the different gene expression patterns between newly diagnosed and relapsed APL [36]. A recent report showed that TrkB was a target of miR-204 and involved in differentiation and proliferation of neuroblastoma [37]. We also found that TrkB was a target of miR-603 and involved in the differentiation and proliferation of NB4 cells. Although there have been no previous reports of TrkB involvement in APL cell differentiation, activation of TrkB protects myeloid cells from apoptosis and supports proliferation [18]. Accumulated evidence suggests that some promising differentiation inducers, such as epigenetic modifiers, glycosylation modifiers, cytokines, chemotherapeutic agents, and some kinase inhibitors [38, 39]. We also found that inhibition of TrkB by si-TrkB was shown to promote differentiation of NB4 cells. Inducing cellular differentiation is the key treatment of APL, suggesting that TrkB might be beneficial in the treatment of relapsed APL.
There are several limitations in our study. As the dramatically effects of ATO treatment in APL and genetic mutations were found in some patients with relapsed APL, only 10 relapsed APL patients (3 cases in microarray and 7 cases in PCR) without reported genetic mutations were included in our study [23,24,25, 36]. Because of high expression of PML::RARA in relapsed APL, we chose the NB4 cell line that contains a constitutively active PML::RARA rearrangementfor functional experiments [40].
In conclusion, we observed different miRNA expression patterns in patients with relapsed APL who were treated with ATO compared with newly diagnosed patients. In particular, miR-603 expression was extremely low and negatively correlated with TrkB expression. In NB4 cells, we found that miR-603/TrkB may participate in cell differentiation. These results may show that differentiation arrest may participate in APL relapse and suggest that the miR-603/TrkB axis is a novel therapeutic target for relapsed APL.
Data Availability
The data are available from the corresponding author on reasonable request.
References
Mistry AR, Pedersen EW, Solomon E, Grimwade D (2003) The molecular pathogenesis of acute promyelocytic leukaemia: implications for the clinical management of the disease. Blood Rev 17(2):71–97
Wang ZY, Chen Z (2008) Acute promyelocytic leukemia: from highly fatal to highly curable. Blood 111(5):2505–2515
Coombs CC, Tavakkoli M, Tallman MS (2015) Acute promyelocytic leukemia: where did we start, where are we now, and the future. Blood Cancer J 5(4):e304
Carbone R, Botrugno OA, Ronzoni S, Insinga A, Di Croce L, Pelicci PG, Minucci S (2006) Recruitment of the histone methyltransferase SUV39H1 and its role in the oncogenic properties of the leukemia-associated PML-retinoic acid receptor fusion protein. Mol Cell Biol 26(4):1288–1296
Hoemme C, Peerzada A, Behre G, Wang Y, McClelland M, Nieselt K, Zschunke M, Disselhoff C, Agrawal S, Isken F et al (2008) Chromatin modifications induced by PML-RARalpha repress critical targets in leukemogenesis as analyzed by ChIP-Chip. Blood 111(5):2887–2895
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136(2):215–233
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86
Havelange V, Garzon R, Croce CM (2009) MicroRNAs: new players in acute myeloid leukaemia. Br J Cancer 101(5):743–748
Fazi F, Rosa A, Fatica A, Gelmetti V, De Marchis ML, Nervi C, Bozzoni I (2005) A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPalpha regulates human granulopoiesis. Cell 123(5):819–831
Pulikkan JA, Dengler V, Peramangalam PS, Peer Zada AA, Müller-Tidow C, Bohlander SK, Tenen DG, Behre G (2010) Cell-cycle regulator E2F1 and microRNA-223 comprise an autoregulatory negative feedback loop in acute myeloid leukemia. Blood 115(9):1768–1778
Pulikkan JA, Peramangalam PS, Dengler V, Ho PA, Preudhomme C, Meshinchi S, Christopeit M, Nibourel O, Müller-Tidow C, Bohlander SK et al (2010) C/EBPα regulated microRNA-34a targets E2F3 during granulopoiesis and is down-regulated in AML with CEBPA mutations. Blood 116(25):5638–5649
Katzerke C, Madan V, Gerloff D, Bräuer-Hartmann D, Hartmann JU, Wurm AA, Müller-Tidow C, Schnittger S, Tenen DG, Niederwieser D et al (2013) Transcription factor C/EBPα-induced microRNA-30c inactivates Notch1 during granulopoiesis and is downregulated in acute myeloid leukemia. Blood 122(14):2433–2442
Garzon R, Pichiorri F, Palumbo T, Visentini M, Aqeilan R, Cimmino A, Wang H, Sun H, Volinia S, Alder H et al (2007) MicroRNA gene expression during retinoic acid-induced differentiation of human acute promyelocytic leukemia. Oncogene 26(28):4148–4157
Serafim Junior V, Fernandes GMM, Oliveira-Cucolo JG, Pavarino EC, Goloni-Bertollo EM (2020) Role of Tropomyosin-related kinase B receptor and brain-derived neurotrophic factor in cancer. Cytokine 136:155270
Malekan M, Nezamabadi SS, Samami E, Mohebalizadeh M, Saghazadeh A, Rezaei N (2023) BDNF and its signaling in cancer. J Cancer Res Clin Oncol 149(6):2621–2636
Maroder M, Bellavia D, Meco D, Napolitano M, Stigliano A, Alesse E, Vacca A, Giannini G, Frati L, Gulino A et al (1996) Expression of trKB neurotrophin receptor during T cell development. Role of brain derived neurotrophic factor in immature thymocyte survival. J Immunol 157(7):2864–2872
Abbaci A, Talbot H, Saada S, Gachard N, Abraham J, Jaccard A, Bordessoule D, Fauchais AL, Naves T, Jauberteau MO (2018) Neurotensin receptor type 2 protects B-cell chronic lymphocytic leukemia cells from apoptosis. Oncogene 37(6):756–767
Li Z, Beutel G, Rhein M, Meyer J, Koenecke C, Neumann T, Yang M, Krauter J, von Neuhoff N, Heuser M et al (2009) High-affinity neurotrophin receptors and ligands promote leukemogenesis. Blood 113(9):2028–2037
Guo M, Zhang X, Wang G, Sun J, Jiang Z, Khadarian K, Yu S, Zhao Y, Xie C, Zhang K et al (2015) miR-603 promotes glioma cell growth via Wnt/β-catenin pathway by inhibiting WIF1 and CTNNBIP1. Cancer Lett 360(1):76–86
Sell SL, Widen SG, Prough DS, Hellmich HL (2020) Principal component analysis of blood microRNA datasets facilitates diagnosis of diverse diseases. PLoS ONE 15(6):e0234185
Cocco E, Scaltriti M, Drilon A (2018) NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol 15(12):731–747
Yuzugullu H, Von T, Thorpe LM, Walker SR, Roberts TM, Frank DA, Zhao JJ (2016) NTRK2 activation cooperates with PTEN deficiency in T-ALL through activation of both the PI3K-AKT and JAK-STAT3 pathways. Cell Discov 2:16030
Liu J, Zhu HH, Jiang H, Jiang Q, Huang XJ (2016) Varying responses of PML-RARA with different genetic mutations to arsenic trioxide. Blood 127(2):243–250
Lehmann-Che J, Bally C, de Thé H (2014) Resistance to therapy in acute promyelocytic leukemia. N Engl J Med 371(12):1170–1172
Zhu HH, Qin YZ, Huang XJ (2014) Resistance to arsenic therapy in acute promyelocytic leukemia. N Engl J Med 370(19):1864–1866
Madan V, Shyamsunder P, Han L, Mayakonda A, Nagata Y, Sundaresan J, Kanojia D, Yoshida K, Ganesan S, Hattori N et al (2016) Comprehensive mutational analysis of primary and relapse acute promyelocytic leukemia. Leukemia 30(12):2430
Careccia S, Mainardi S, Pelosi A, Gurtner A, Diverio D, Riccioni R, Testa U, Pelosi E, Piaggio G, Sacchi A et al (2009) A restricted signature of miRNAs distinguishes APL blasts from normal promyelocytes. Oncogene 28(45):4034–4040
Zamani A, Fattahi Dolatabadi N, Houshmand M, Nabavizadeh N (2021) miR-324-3p and miR-508-5p expression levels could serve as potential diagnostic and multidrug-resistant biomarkers in childhood acute lymphoblastic leukemia. Leuk Res 109:106643
Cao Y, Liu Y, Shang L, Chen H, Yue Y, Dong W, Guo Y, Yang H, Yang X, Gu W et al (2022) Overexpression of miR-17 predicts adverse prognosis and disease recurrence for acute myeloid leukemia. Int J Clin Oncol 27(7):1222–1232
Zhang TJ, Lin J, Zhou JD, Li XX, Zhang W, Guo H, Xu ZJ, Yan Y, Ma JC, Qian J (2018) High bone marrow miR-19b level predicts poor prognosis and disease recurrence in de novo acute myeloid leukemia. Gene 640:79–85
Wang F, Li J, Li L, Chen Z, Wang N, Zhu M, Mi H, Xiong Y, Guo G, Gu Y (2022) Circular RNA circ_IRAK3 contributes to tumor growth through upregulating KIF2A via adsorbing miR-603 in breast cancer. Cancer Cell Int 22(1):81
Ma C, Zhan C, Yuan H, Cui Y, Zhang Z (2016) MicroRNA-603 functions as an oncogene by suppressing BRCC2 protein translation in osteosarcoma. Oncol Rep 35(6):3257–3264
Lin YX, Wu XB, Zheng CW, Zhang QL, Zhang GQ, Chen K, Zhan Q, An FM (2021) Mechanistic Investigation on the Regulation of FABP1 by the IL-6/miR-603 Signaling in the Pathogenesis of Hepatocellular Carcinoma. Biomed Res Int 2021:8579658
Yin B, Ma ZY, Zhou ZW, Gao WC, Du ZG, Zhao ZH, Li QQ (2015) The TrkB+ cancer stem cells contribute to post-chemotherapy recurrence of triple-negative breast cancers in an orthotopic mouse model. Oncogene 34(6):761–770
Bao L, Wang YT, Lu MQ, Shi L, Chu B, Gao S (2023) BDNF/TrkB confers bortezomib resistance in multiple myeloma by inducing BRINP3. Biochim Biophys Acta Gen Subj 1867(3):130299
Chendamarai E, Ganesan S, Alex AA, Kamath V, Nair SC, Nellickal AJ, Janet NB, Srivastava V, Lakshmi KM, Viswabandya A et al (2015) Comparison of newly diagnosed and relapsed patients with acute promyelocytic leukemia treated with arsenic trioxide: insight into mechanisms of resistance. PLoS ONE 10(3):e0121912
Yang T, Li J, Zhuo Z, Zeng H, Tan T, Miao L, Zheng M, Yang J, Pan J, Hu C et al (2022) TTF1 suppresses neuroblastoma growth and induces neuroblastoma differentiation by targeting TrkA and the miR-204/TrkB axis. iScience 25(7):104655
Takahashi S (2021) Current Understandings of Myeloid Differentiation Inducers in Leukemia Therapy. Acta Haematol 144(4):380–388
Takahashi S (2022) Kinase Inhibitors and Interferons as Other Myeloid Differentiation Inducers in Leukemia Therapy. Acta Haematol 145(2):113–121
Villiers W, Kelly A, He XH, Kaufman-Cook J, Elbasir A, Bensmail H, Lavender P, Dillon R, Mifsud B, Osborne C (2023) Multi-omics and machine learning reveal context-specific gene regulatory activities of PML::RARA in acute promyelocytic leukemia. Nat Commun 14(1):724
Funding
This research was supported by the National Natural Science Foundation of China (81903966) and the Science Foundation of the First Affiliated Hospital of Harbin Medical University (2021M02).
Author information
Authors and Affiliations
Contributions
Huibo Li conceived, carried out experiments and wrote the paper; Jinxiao Hou analyzed data; Yueyue Fu and Yanqiu Zhao carried out experiments and analyzed the data; Jie Liu and Dan Guo performed the experiments; Ruiqi Lei, Yiting Wu and Linqing Tang performed the experiments. Shengjin Fan conceived the experiments and revised the paper. All authors gave final approval of the submitted and published versions.
Corresponding author
Ethics declarations
Ethics approval
All experimental procedures were approved by the First Affiliated Hospital, Harbin Medical University.
Competing interests
The authors declare no competing interests.
Conflict of interest
None of the authors have any financial interest related to this work and therefore declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, H., Hou, J., Fu, Y. et al. miR-603 promotes cell proliferation and differentiation by targeting TrkB in acute promyelocytic leukemia. Ann Hematol 102, 3357–3367 (2023). https://doi.org/10.1007/s00277-023-05441-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00277-023-05441-w