Abstract
Purpose
To develop and externally validate models incorporating a PET radiomics signature (R-signature) obtained by the cross-combination method for predicting the survival of patients with diffuse large B-cell lymphoma (DLBCL).
Methods
A total of 383 patients with DLBCL from two medical centres between 2011 and 2019 were included. The cross-combination method was used on three types of PET radiomics features from the training cohort to generate 49 feature selection-classification candidates based on 7 different machine learning models. The R-signature was then built by selecting the optimal candidates based on their progression-free survival (PFS) and overall survival (OS). Cox regression analysis was used to develop the survival prediction models. The calibration, discrimination, and clinical utility of the models were assessed and externally validated.
Results
The R-signatures determined by 12 and 31 radiomics features were significantly associated with PFS and OS, respectively (P<0.05). The combined models that incorporated R-signatures, metabolic metrics, and clinical risk factors exhibited significant prognostic superiority over the clinical models, PET-based models, and the National Comprehensive Cancer Network International Prognostic Index in terms of both PFS (C-index: 0.801 vs. 0.732 vs. 0.785 vs. 0.720, respectively) and OS (C-index: 0.807 vs. 0.740 vs. 0.773 vs. 0.726, respectively). For external validation, the C-indices were 0.758 vs. 0.621 vs. 0.732 vs. 0.673 and 0.794 vs. 0.696 vs. 0.781 vs. 0.708 in the PFS and OS analyses, respectively. The calibration curves showed good consistency, and the decision curve analysis supported the clinical utility of the combined model.
Conclusion
The R-signature could be used as a survival predictor for DLBCL, and its combination with clinical factors may allow for accurate risk stratification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common type of lymphoma [1]. As a remarkably heterogeneous disease, DLBCL is known for its considerable variation in clinical behaviour, response to therapy, and prognosis [2]. Chemoimmunotherapy with R-CHOP (rituximab, cyclophosphamide, doxorubicin, vincristine, and prednisone) therapy is now used as the standard therapy for DLBCL [3]. However, 30-50% of patients ultimately relapse and die as a result of the disease [4, 5]. Therefore, an accurate identification of patients who will not respond well to standard regimens at diagnosis is needed to guide new therapeutic strategies. The International Prognostic Index (IPI) and its successor (NCCN-IPI) are prognostic scoring systems that predict survival. However, it is difficult to accurately identify DLBCL patients who are likely to relapse, especially those with a poor long-term prognosis, to individualize therapy [6,7,8].
Volume metabolic parameters derived from [18F]-FDG PET/CT, such as metabolic tumour volume(MTV) and total lesion glycolysis (TLG), are powerful indices for assessing tumour burden and risk stratification in DLBCL patients [9,10,11,12,13]. However, these metabolic parameters are limited in their ability to reflect tumour heterogeneity, which ultimately limits their contribution to predicting cancer progression, therapeutic resistance, and survival outcomes [14, 15]. Machine learning advancements have enabled clinicians to use radiomics to extract implicit features from images to assess intratumoural biological heterogeneity and assist in clinical outcome prediction [16, 17]. Several studies [18,19,20,21,22,23] have explored the prognostic value of PET/CT radiomics in lymphoma, including Hodgkin lymphoma (HL), nasal-type extranodal natural killer/T cell lymphoma (ENKTL), mantle cell lymphoma (MCL), and DLBCL. In general, these studies used one machine learning model to select and develop radiomic signatures, e.g. the multilayer perceptron neural network combined with logistic regression analysis [19], the least absolute shrinkage and selection operator regression (LASSO) model [20, 22], and the logistic regression model [23]. However, there is no model that works well for all kinds of data. To address this limitation, cross-combinations of classification and selection methods based on radiomics features have recently appeared and have proven effective in improving classification and diagnostic performance [24, 25]. However, it is still unknown whether these methods can be applied to radiomic images for DLBCL. In this paper, we explored the PET radiomics features using a cross-combination of machine learning models to assess risk in DLBCL patients. Furthermore, we established and externally validated models for predicting survival outcome in routine clinical practice, allowing for its clinical translation.
Materials and methods
Patient data collection
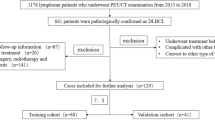
A total of 273 patients from Nanjing Drum Tower Hospital, the Affiliated Hospital of Nanjing University Medical School, were included as the training cohort. To examine the general applicability of nomograms, an independent external validation cohort of 110 patients from Jiangsu Province Hospital, the First Affiliated Hospital of Nanjing Medical University, was included. Ethical approval was obtained for this retrospective analysis, and the requirement for written consent from patients was waived. The workflow of patient selection is shown in Fig. 1. The inclusion criteria were as follows: (I) pathologically confirmed DLBCL, (II) pretreatment [18F]-FDG PET/CT scan was performed, and (III) treatment with R-CHOP-like regimens with curative intent was carried out. Patients with incomplete medical data, those lost to follow-up, and those with a previous history of cancer were excluded. Clinical data (sex, age, B symptoms, Eastern Cooperative Oncology Group performance status (ECOG PS), NCCN-IPI index, lactate dehydrogenase (LDH) level, Ann Arbor stage, and immunohistochemistry result) were obtained from medical records. Participants were divided into two groups: those with a low/low-intermediate risk (NCCN-IPI of <4) and those with a high-intermediate/high risk (NCCN-IPI of ≥4) [8]. The follow-up data were obtained through electronic medical records and telephone interviews. Progression-free survival (PFS) and overall survival (OS) were chosen as endpoints for evaluating the prognosis of DLBCL patients. PFS was calculated as the time between diagnosis and the first relapse, progression, death from any cause, or last follow-up. OS was calculated as the time between diagnosis and death from any cause or last follow-up.
PET/CT Scanning Protocol
All patients underwent PET/CT scans with one of the following systems: Gemini GXL (Philips Corp, Netherlands) for the training cohort and Biograph 16 PET/CT scanner (Siemens Health care, Erlangen, Germany) for the external validation cohort, based on standard clinical scanning protocols. All patients fasted for at least 6 hours before scans, resulting in blood glucose levels under 8.7 mmol/L. After 6 hours of fasting (no oral or intravenous fluids containing sugar or dextrose), 185–370 MBq of [18F]-FDG (5.18 MBq/kg) was administered intravenously. Each patient was weighed to determine the standardized uptake value (SUV) prior to each scan. Whole-body PET/CT scans (from the base of the skull to the upper thigh) were performed 60 minutes after the radiopharmaceutical injection. Emission data were acquired for 2 minutes in each bed position. CT acquisition data were used for attenuation correction, and corrected PET images were reconstructed using ordered-subset expectation maximization (OSEM). Images were reconstructed with standard 4 × 4 × 4 mm3 voxels using 3 iterations of 33 subsets.
VOI drawing and radiomics processing
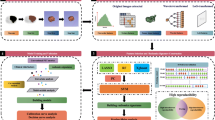
PET images were uploaded to 3D Slicer software (version 4.8.0; http://www.slicer.org). The Grow-Cut algorithm in 3D Slicer was used to semiautomatically generate the volume of interest (VOI). Two physicians manually adjusted the VOI to ensure that the measurement was reliable. If there was a discrepancy, the VOIs were reviewed and determined by a senior nuclear medical scientist. The 3 largest lesions were chosen as the targets for radiomic feature extraction. For the three largest VOIs, we summed the radiomic size and shape features, while all other features were averaged for each VOI [26]. Feature extraction was performed based on PET images by the built-in open-source PyRadiomics package (http://www.radiomics.io/pyradiomics.html) in 3D Slicer. Before feature extraction, we preprocessed the PET images, including resampling all images to a voxel size of 1×1×1 mm3 using bilinear interpolation and discretization with a fixed bin count of 128, as recommended by [27]. To obtain the comprehensive radiomics features, we converted the original PET images into wavelet and Laplacian of Gaussian (LoG) images with wavelet transforms and LoG transforms, respectively. Then, three types of radiomics features were extracted from the original, wavelet and LoG images: (I) first-order statistics, (II) shape features and (III) texture features. The full list of characteristics is provided in Supplementary Table 1. The workflow of our radiomic analysis is shown in Fig. 2.
In addition, the conventional metabolic parameters, including the SUVmax, MTV, and TLG, were calculated by volume viewer software on a dedicated workstation (CompassView 5.0, Philips Corp., Amsterdam, the Netherlands). Regions of interest (ROIs) were manually placed to cover the lesion, and the maximal SUV (SUVmax) value was recorded for each lesion in the PET images of the patients. For each PET dataset, the SUVmax was defined as the highest SUV among all hypermetabolic tumour foci. The MTV was determined by drawing a circular ROI that included all involved lesions in the axial, coronal, and sagittal PET images. The voxel boundaries were produced automatically with the 41% SUVmax threshold method recommended by the European Association of Nuclear Medicine [28]. The total metabolic tumour volume (TMTV) was obtained by summing the MTV of all lesions. The TLG was calculated as the sum of the product of the SUVmean and MTV of all lesions (TLG = sum of [SUVmean × MTV]).
Radiomics feature selection and signature construction
First, all extracted radiomics features were harmonized using z scores. The interobserver repeatability of each radiomic feature was evaluated in 50 randomly chosen patients using the intraclass correlation coefficient (ICC) method, and features with an ICC greater than 0.75 were selected.
Then, we calculated the effective and robust R-signature based on the cross-combination method of seven machine learning models. Specifically, we choose seven classical machine learning models: support vector machines (SVMs), gradient boosting decision trees (GBDTs), extreme gradient boosting (XGBoost), random forest (RF), light gradient boosting machines (LGB), the least absolute shrinkage and selection operator (LASSO), and logistic regression (Logistic). The SVM [29] maps the input samples to points in high-dimensional space to maximize the gap between two categories. The GBDT [30] is a robust prediction model that consists of a series of weak decision trees. The XGBoost method extends GBDTs [31] in terms of structure and training strategy for more efficient predictions. Furthermore, the LGB [32] outperforms XGBoost in terms of accuracy and operation speed due to its tree structure and optimization algorithm. The RF method [33] consists of a large number of individual decision trees that operate as an ensemble. Each individual tree in the RF produces a class prediction, and the class with the most votes becomes the model’s prediction. LASSO [34] is a classical regression analysis method that performs both feature selection and regularization, with features with nonzero coefficients viewed as useful features. Logistic regression [35] combines linear regression and activation functions to estimate the probability of a certain class.
These seven models acted as feature selection and classification models for predicting the PFS and OS. The feature selection models were used to select the most significant radiomics features, and the classification models were used to construct an R-signature based on the selected features. In previous works, LASSO was generally used, as it produced a simpler and more interpretable format than the other models. However, the effectiveness of LASSO has not been thoroughly considered. In this paper, we generated a set of feature selection-classification candidates with cross-combinations of the seven machine learning models, such as LASSO-SVM, LGB-XGBoost and Logistic-RF. Then, the R-signature was built by the optimal candidate. The optimal candidate was determined with tenfold cross-validation in four steps (see Supplementary Fig. 1): (I) the samples were split into 10 groups of approximately equal size. One group was used for validation, and the remaining 9 groups were used for training. This process was repeated 10 times, with different groups used for validation each time. Considering that our samples came from two medical centres, in the split procedure, we ensured that the cohorts retained the same proportion of sample sources as the original dataset. (II) Given a training cohort, we trained a feature selection model with all PET radiomics features for predicting the PFS/OS to obtain the corresponding feature weights. Based on the feature weights, we trained a feature classification model by recursively considering the subsets of the radiomics features. (III) We identified the feature subset with the largest AUC value as containing the most important features, and the corresponding feature selection-classification model was identified as the optimal candidate. The R-signature was the prediction result of the optimal classification model. (IV) Based on the optimal candidate, we calculated the AUC of each validation cohort and outputted the average AUC of all 10 validation cohorts.
Development and validation of the models
Univariate and multivariate Cox regression analyses were used to select the potential independent prognostic factors of PFS and OS in the training cohort. All the significant clinical factors (age, LDH, B symptoms, ECOG, Ann Arbor stage, extranodal involvement, bulky disease) in the univariate analysis were included in the multivariate analysis. Independent predictors (age, Ann Abor stage, bulky disease) were used to build the clinical model. Then, all statistically significant PET factors (SUVmax, TMTV, TLG, R-signature) in the univariate analysis were included in the multivariate analysis. The independent predictors (SUVmax, TMTV, R-signature) were used to build a combined PET-based model. After that, we assembled the models by combining all the independent clinical and PET predictors to build the combined model. Calibration curves were used to assess the models. To quantify the discrimination performance of the models, the Harrell concordance index (C-index) and the time-dependent area under the ROC curve (tdROC) were calculated. The performance of the models was also externally tested in the independent validation cohort with the formula and cut-off values derived from the training cohort.
Clinical benefit analysis based on the models
To estimate the clinical utility of the models, decision curve analysis (DCA) was performed by calculating the net benefits for a range of threshold probabilities in the training cohort and validation cohort.
Statistical analysis
All statistical tests were performed using SPSS 22.0 (IBM Corp., Armonk, NY, USA) and R statistical software (version 4.0.2). A P value less than 0.05 was considered statistically significant. The difference in the related clinical information between the training and validation cohorts was assessed using χ2 tests or Mann–Whitney U tests, as appropriate. We determined the optimal threshold of the R-signature based on the receiver operating characteristic (ROC) curve. Cox regression analysis was used to analyse the potential independent predictors and build the models. The survival conditions were evaluated by the Kaplan–Meier method and compared by log-rank tests. The calibration curves, C-index, tdAUC and DCA were calculated for the models in the training and validation cohorts.
Results
Patient characteristics
The baseline characteristics of the included patients in the training and validation cohorts are summarized in Table 1. No statistically significant (P < 0.05) difference was observed between the two cohorts. A total of 14 participants had SUVmean values in the liver<1.3 (training cohort: n=3; validation cohort: n=2) or ≥3.0 (training cohort: n=5; validation cohort: n=4). The median follow-up times for the training and validation cohorts were 35.0 and 23.5 months, respectively. Of the participants who experienced disease relapse or progression in the training cohort, 103 and 76 died, respectively, and the 1-year, 3-year, and 5-year PFS and 1-year, 3-year, and 5-year OS rates were 77.3%, 66.1%, and 54.6% and 85.7%, 76.3%, and 66.8%, respectively. Of the participants who experienced disease relapse or progression in the validation cohort, 33 and 25 died, respectively, and the 1-year, 3-year, and 5-year PFS and 1-year, 3-year, and 5-year OS rates were 83.5%, 67.9%, and 63.4% and 88.9%, 74.1%, and 71.9%, respectively.
Radiomics feature selection and R-signature construction
In the first step of reproducible feature selection, we excluded features with ICCs less than 0.75, and 274 features with high reproducibility were selected for further analysis. Based on the results of the feature extraction-classification models, we determined 12 radiomic features for constructing the R-signaturePFS for the SVM-SVM model (AUC=0.757) and 31 radiomic features for constructing the R-signatureOS for the XGBoost-XGBoost model (AUC=0.762) (Fig. 3 and Supplementary Tables 2 and 3). The optimal cut-off values of the R-signatures were 0.386 for PFS and 0.268 for OS. Using these cut-offs, the risk groups identified by the R-signatures showed distinct OS and PFS outcomes in both the training and validation cohorts (Fig. 3).
Heatmaps indicate the AUC performance of the cross-combinations of the feature selection methods (columns) and classification models (rows) in predicting PFS (a) and OS (c). The spectra demonstrating the selected features to build R-signaturePFS (b) and R-signatureOS (d). Kaplan–Meier plots for patients stratified by risk classification according to R-signatures for patients in the training (e, f) and validation (g, h) cohorts
Univariate and multivariate analysis results
The differences in the clinical characteristics between the dichotomized R-signaturePFS and R-signatureOS groups are shown in Table 2.
For the clinical variables, we performed univariate and multivariate Cox regression analyses to identify the independent factors associated with survival. We identified age (HR=2.460, P<0.001; HR=2.810, P<0.001), Ann Abor stage (HR=3.139, P<0.001; HR=3.105, P<0.001) and bulky disease (HR=1.518, P=0.043; HR=1.631, P=0.041) as independent risk factors for PFS and OS (Table 3).
For the PET variables, univariate and multivariate Cox regression analyses showed that the R-signature (HR=5.024, P<0.001; HR=4.054, P<0.001), SUVmax (HR=2.027, P=0.001; HR=2.005, P=0.004) and TMTV (HR=2.265, P<0.001; HR=1.789, P=0.036) were independent risk factors for PFS and OS (Table 3).
Assessment and validation of models built for predicting PFS and OS
The independent clinical (age, Ann Abor stage, and bulky disease) and PET (R-signature, TMTV and SUVmax) predictors that were significant in the multivariate analysis were used to build the combined models for predicting the PFS and OS rates (Fig. 4 and Table 4). Individual clinical models and PET-based models were also constructed (Table 4).
a The combined model for predicting PFS. b, d Calibration curves of the model for predicting PFS in the training and validation cohorts at the 1-year, 3-year, and 5-year time points. c, e Time-dependent area under the ROC curve of the models for predicting PFS in the training and validation cohorts. f The combined model for predicting OS. g, i Calibration curves of the models for predicting OS in the training and validation cohorts at the 1-year, 3-year, and 5-year time points. h, j Time-dependent area under the ROC curve of the models for predicting OS in the training and validation cohorts
For PFS, the C-indices of the combined model in the training and validation cohorts were 0.801 (95% CI: 0.745-0.848) and 0.758 (95% CI: 0.618-0.795), respectively; these values were superior to those of the clinical model (C-index: 0.732, 95% CI: 0.679-0.785 in the training cohort, and C-index: 0.621, 95% CI: 0.512-0.731 in the validation cohort), PET-based model (C-index: 0.785, 95% CI: 0.734-0.835 in the training cohort, and C-index: 0.732, 95% CI: 0.642-0.823 in the validation cohort) and NCCN-IPI model (C-index: 0.720, 95% CI: 0.668-0.772 in the training cohort, and C-index: 0.673, 95% CI: 0.584-0.763 in the validation cohort) (Table 5).
For OS, the C-indices of the combined model in the training and validation cohorts were 0.807 (95% CI: 0.736-0.854) and 0.794 (95% CI: 0.668-0.881); these values were superior to those of the clinical model (C-index: 0.740, 95% CI: 0.680-0.800 in the training cohort, and C-index: 0.696, 95% CI: 0.584-0.807 in the validation cohort), PET-based model (C-index: 0.773, 95% CI: 0.713-0.833 in the training cohort, and C-index: 0.781, 95% CI: 0.697-0.902 in the validation cohort) and NCCN-IPI model (C-index: 0.726, 95% CI: 0.665-0.786 in the training cohort, and C-index: 0.708, 95% CI: 0.610-0.805 in the validation cohort) (Table 5).
The calibration curves for predicting the probability of PFS and OS at 1, 3, or 5 years for each model after 1000 bootstrap replicates are shown in Fig. 4. The calibration curves showed satisfactory agreement between the estimations and actual observations in the training and validation cohorts for the combined models.
Performance analysis of the combined models in clinical use
The decision curve analyses for the combined models are presented in Fig. 5. The decision curve analysis showed that the combined models had a higher overall net benefit than the clinical models, PET-based models, and NCCN-IPI across most of the risk thresholds in both the training and validation cohorts for PFS and OS.
Outcome according to R-signature in the subgroups of the NCCN-IPI model
To confirm the added prognostic value of radiomics features to the NCCN-IPI, we evaluated the prognostic value of R-signatures in low/low-intermediate risk (NCCN-IPI of <4) and high-intermediate/high (NCCN-IPI of ≥4) groups.
For the training cohort, in the low/low-intermediate risk group (n=138), patients with a low R-signature had a better survival rate (92.3% for PFS and 89.5% for OS) than those with a high R-signature (51.1% for PFS and 35.7% for OS), and the P values were both <0.001. In the high-intermediate/high risk group (n=135), patients with a low R-signature had a better survival rate (75.7% for PFS and 77.0% for OS) than those with a high R-signature (34.7% for PFS and 39.3% for OS), and the P values were both 0.001 (Fig. 6 and Table 6).
Kaplan–Meier plots for PFS and OS in patients with different NCCN-IPI scores in relation to their R-signatures. a-d Low/low-intermediate NCCN-IPI (score: 0,1/2,3) and high-intermediate/high NCCN-IPI (score: 4,5/6-8) in relation to R-signaturePFS for PFS stratification in the training (a, b) and validation (c, d) cohorts. e-h. Low/low-intermediate NCCN-IPI (score: 0,1/2,3) and high-intermediate/high NCCN-IPI (score: 4,5/6-8) in relation to R-signatureOS for OS stratification in the training (e, f) and validation (g, h) cohorts
For the validation cohort, in the low/low-intermediate risk group (n=68), patients with a low R-signature had a better survival rate (97.4% for PFS and 96.2% for OS) than those with a high R-signature (55.2% for PFS and 56.3% for OS), and the P values were both statistically significant (<0.001). In the high-intermediate/high risk group (n=42), patients with a low R-signature had a better survival rate (75.0% for PFS and 83.3% for OS) than those with a high R-signature (36.4% for PFS and 33.3% for OS), and the P values were 0.031 and 0.007 (Fig. 6 and Table 6).
Discussion
Our study analysed PET radiomics features to predict survival outcomes in DLBCL patients. The results of this study indicated that combined models that integrated R-signatures calculated by radiomics features extracted from pretreatment [18F]-FDG PET using machine learning methods could be used as a preoperative tool for individualized prognosis prediction treatment guidance in patients with DLBCL.
Accurately predicting the prognosis of patients is of great importance for optimizing DLBCL treatment strategies. Several studies have attempted to assess the predictive value of radiomics information from PET images in DLBCL [21,22,23]. Aide et el. recruited 132 patients with DLBCL to evaluate the ability of pretreatment PET radiomics features to predict prognosis and found that LZLGE was the only independent predictor of 2-year event-free survival (HR=2.84, P=0.01) with ROC analysis accuracy (area under curve=0.76) [21]. Lue KH et el. extracted 80 PET-based radiomics features from 171 patients with DLBCL and found that RLNGLRLM was independently associated with PFS (HR=15.7, P=0.007) and OS (HR=8.64, P=0.040) [22]. Eertink et al. found that PET radiomics features extracted from the largest lesion of DLBCL patients were able to predict the 2-year time to progression (area under curve=0.67) [23]. Consistent with previous studies, the R-signatures consisting of PET radiomics features identified with machine learning in the present study were independently associated with PFS (HR=4.15, P<0.001) and OS (HR=4.029, P<0.001). Both our study and previous studies support the viewpoint that pivotal radiomics feature data extracted from PET images could help predict clinical outcomes in patients with DLBCL. The underlying cause might be explained by the hypothesis that radiomics features are strongly correlated with tumour heterogeneity [36,37,38], which is a prognostic determinant of patient survival [39,40,41].
Recently, machine learning models have attracted great attention in both research and industry because they can automatically detect inherent patterns in data distribution [42]. With advancements in machine learning, different machine learning models, such as random forest (RF) and support vector machines (SVMs), have been utilized to extract data features. These methods differ from each other in terms of structure, optimization algorithm, scale, and performance in different datasets. For example, SVMs are more suitable than logistic regression for large datasets [43]. However, most existing works have only focused on a single method for feature selection and R-signature construction, and few comparisons have been performed with other machine learning methods to assess robustness and efficiency. The cross-combination of feature selection-classification models has shown great efficiency and robustness in clinical practice [24, 25]. Thus, we developed a cross-combination approach for determining the optimal feature selection-classification pair based on AUC results. As shown in Fig. 3, the AUC used by the LASSO method for predicting OS was poor (0.696). Therefore, we chose the most powerful selection-classification pair from 49 possible combinations. The SVM-SVM model had an AUC of 0.757 for predicting PFS, and the XGBoost-XGBoost model had an AUC of 0.762 for predicting OS. In addition, we found that the selected features (see Fig. 3) were mainly texture features with the wavelet and LoG filtering algorithms, indicating that these features have better discrimination than shape and first-order features.
[18F]-FDG PET results can be used as a measure of cellular glucose metabolism to provide biological information about tumours. Our study demonstrated that the metabolic parameter TMTV was an independent predictor of PFS (HR=1.90, P=0.003), and SUVmax was found to be an independent predictor of both PFS (HR=2.03, P=0.001) and OS (HR=1.77, P=0.015). These results were consistent with previous studies [44,45,46,47]. To apply these results in clinical practice, we developed user-friendly predictive combined models that incorporated the R-signature, metabolic metrics (SUVmax and TMTV), and clinical risk factors (age, stage and bulky disease), and we compared the predictive performance of the models with clinical, PET-based, and NCCN-IPI prognostic models. The study found that the combined models had satisfactory agreement with the calibration curve and exhibited significant prognostic superiority over other models in terms of C-indices and tdAUCs.
To confirm the combined model’s risk discrimination, we compared the clinical utility of the combined model with that of other models (such as the clinical model, PET-based model and NCCN-IPI model). The DCA demonstrated that the combined models were superior to the other models across most of the range of reasonable threshold probabilities, thus suggesting that they are more suitable for estimating individual survival. Consistent with previous studies [6,7,8], our results showed that the NCCN-IPI might be improved for identifying high-risk groups for which novel treatment approaches are most needed (Supplementary Fig. 2 and Supplementary Table 4). The possible additional predictive value of the combined R-signature and the NCCN-IPI model was explored in this study. The results showed that the addition of the R-signature to the NCCN-IPI model yielded a better stratification for DLBCL patients in both high-intermediate/high risk (NCCN-IPI of ≥4) and low/low-intermediate risk (NCCN-IPI of <4) groups, in both the training and validation cohorts. Interestingly, several studies have shown that the TMTV also had additive prognostic value in patients in the NCCN-IPI subgroup [48, 49]. In agreement with the work of Eertink et al. [23], our results support further exploration of combining radiomics features with clinical predictors to more accurately identify high-risk patients. To uncover the potential additional prognostic value of these radiomics features, these results may contribute to better understanding of the clinically beneficial effects of these parameters with machine learning methods.
One of the limitations of our study was its retrospective nature. Additionally, the biological features of DLBCL, including the double expressor, double hit, and genomic prognostic factors, were not investigated in this study. Furthermore, the SUVmean of the liver was outside the normal range in a small portion of our patients (n=14). According to the quality control (QC) criteria described by EANM guidelines, the SUVmean of the liver should be between 1.3 and 3.0. Moreover, given the low number of progression or death events in the NCCN-IPI categories, conclusions about the additional predictive value of the R-signature to the NCCN-IPI model should be interpreted with caution, and larger cohorts are needed in the future. In addition, we used only the 3 largest lesions as targets for radiomic feature calculation in the present study. To date, there is no consensus on the tumour segmentation method for radiomic feature calculation in patients with DLBCL. Due to the high distribution variability of nodal and/or extranodal lesions with heterogeneous volumes and variable metabolic activity, lymphoma segmentation is more challenging than that of primary tumour lesions. Therefore, we proposed a concise tumour segmentation approach for the assessment of radiomics features that may be easier to apply in clinical settings. Finally, the 3D slicer and the growth cut algorithm applied in the present study have not been clinically approved. Although the development of PyRadiomics is involved in the standardization effort of the IBSI (The Image Biomarker Standardization Initiative) team, there are still some differences between PyRadiomics-based feature extraction and IBSI-recommended feature extraction. The main differences are due to grey value discretization, resampling (grid alignment, grey value rounding, and mask resampling), resegmentation, and kurtosis calculation. All of these differences may alter the extrapolation of our results to other clinical practices. The main strengths of our study are the homogeneity of the included patients (de novo DLBCL histology and R-CHOP-like regimens as standard treatment) and the use of an external validation method that supports the general applicability of the models.
Conclusion
The R-signature obtained with the cross-combination method could be used as a survival predictor for patients with DLBCL. Furthermore, the combined models that incorporate R-signature metabolic metrics and clinical risk factors may allow for accurate risk stratification and individualized management of DLBCL patients.
Data availability
The datasets generated and analysed during the current study are available from the Nanjing Drum Tower Hospital, the Affiliated Hospital of Nanjing University Medical School, and the Jiangsu Province Hospital, the First Affiliated Hospital of Nanjing Medical University.
References
Liu Y, Barta SK. Diffuse large B-cell lymphoma: 2019 update on diagnosis, risk stratification, and treatment. Am J Hematol. 2019;94(5):604–16.
Bakhshi TJ, Georgel PT. Genetic and epigenetic determinants of diffuse large B-cell lymphoma. Blood Cancer J. 2020;10(12):123.
Kwak JY. Treatment of diffuse large B cell lymphoma. Korean J Intern Med. 2012;27(4):369–77.
Coiffier B, Thieblemont C, Van Den Neste E, Lepeu G, Plantier I, Castaigne S, Lefort S, Marit G, Macro M, Sebban C, et al. Long-term outcome of patients in the LNH-98.5 trial, the first randomized study comparing rituximab-CHOP to standard CHOP chemotherapy in DLBCL patients: a study by the Groupe d'Etudes des Lymphomes de l'Adulte. Blood. 2010;116(12):2040–2045.
Coiffier B, Sarkozy C. Diffuse large B-cell lymphoma: R-CHOP failure-what to do? Hematol Am Soc Hematol Educ Program. 2016;2016(1):366–78.
Sehn LH, Berry B, Chhanabhai M, Fitzgerald C, Gill K, Hoskins P, Klasa R, Savage KJ, Shenkier T, Sutherland J, et al. The revised International Prognostic Index (R-IPI) is a better predictor of outcome than the standard IPI for patients with diffuse large B-cell lymphoma treated with R-CHOP. Blood. 2007;109(5):1857–61.
International Non-Hodgkin's Lymphoma Prognostic Factors P. A predictive model for aggressive non-Hodgkin's lymphoma. N Engl J Med. 1993;329(14):987–994.
Zhou Z, Sehn LH, Rademaker AW, Gordon LI, Lacasce AS, Crosby-Thompson A, et al. An enhanced International Prognostic Index (NCCN-IPI) for patients with diffuse large B-cell lymphoma treated in the rituximab era[J]. Blood. 2014;123(6):837–42.
Gallicchio R, Mansueto G, Simeon V, Nardelli A, Guariglia R, Capacchione D, Soscia E, Pedicini P, Gattozzi D, Musto P, et al. F-18 FDG PET/CT quantization parameters as predictors of outcome in patients with diffuse large B-cell lymphoma. Eur J Haematol. 2014;92(5):382–9.
Jiang C, Teng Y, Chen J, Wang Z, Zhou Z, Ding C, Xu J. Value of (18)F-FDG PET/CT for prognostic stratification in patients with primary intestinal diffuse large B cell lymphoma treated with an R-CHOP-like regimen. Ann Nucl Med. 2020;34(12):911–9.
Xie M, Zhai W, Cheng S, Zhang H, Xie Y, He W. Predictive value of F-18 FDG PET/CT quantization parameters for progression-free survival in patients with diffuse large B-cell lymphoma. Hematology. 2016;21(2):99–105.
Xie M, Wu K, Liu Y, Jiang Q, Xie Y. Predictive value of F-18 FDG PET/CT quantization parameters in diffuse large B cell lymphoma: a meta-analysis with 702 participants. Med Oncol. 2015;32(1):446.
Vercellino L, Cottereau AS, Casasnovas O, Tilly H, Feugier P, Chartier L, Fruchart C, Roulin L, Oberic L, Pica GM, et al. High total metabolic tumor volume at baseline predicts survival independent of response to therapy. Blood. 2020;135(16):1396–405.
Stanta G, Bonin S. Overview on Clinical Relevance of Intra-Tumor Heterogeneity. Front Med (Lausanne). 2018;5:85.
McGranahan N, Swanton C. Clonal Heterogeneity and Tumor Evolution: Past, Present, and the Future. Cell. 2017;168(4):613–28.
Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017;14(12):749–62.
Lambin P, Rios-Velazquez E, Leijenaar R, Carvalho S, van Stiphout RG, Granton P, Zegers CM, Gillies R, Boellard R, Dekker A, et al. Radiomics: extracting more information from medical images using advanced feature analysis. Eur J Cancer. 2012;48(4):441–6.
Lue KH, Wu YF, Liu SH, Hsieh TC, Chuang KS, Lin HH, Chen YH. Prognostic Value of Pretreatment Radiomic Features of 18F-FDG PET in Patients With Hodgkin Lymphoma. Clin Nucl Med. 2019;44(10):e559–65.
Mayerhoefer ME, Riedl CC, Kumar A, Gibbs P, Weber M, Tal I, Schilksy J, Schöder H. Radiomic features of glucose metabolism enable prediction of outcome in mantle cell lymphoma. Eur J Nucl Med Mol Imaging. 2019;46(13):2760–9.
Wang H, Zhao S, Li L, Tian R. Development and validation of an (18)F-FDG PET radiomic model for prognosis prediction in patients with nasal-type extranodal natural killer/T cell lymphoma. Eur Radiol. 2020;30(10):5578–87.
Aide N, Fruchart C, Nganoa C, Gac AC, Lasnon C. Baseline (18)F-FDG PET radiomic features as predictors of 2-year event-free survival in diffuse large B cell lymphomas treated with immunochemotherapy. Eur Radiol. 2020;30(8):4623–32.
Lue KH, Wu YF, Lin HH, Hsieh TC, Liu SH, Chan SC, Chen YH. Prognostic Value of Baseline Radiomic Features of 18F-FDG PET in Patients with Diffuse Large B-Cell Lymphoma. Diagnostics (Basel). 2020;11(1):36.
Eertink JJ, van de Brug T, Wiegers SE, Zwezerijnen GJC, Pfaehler EAG, Lugtenburg PJ, van der Holt B, de Vet HCW, Hoekstra OS, Boellaard R, et al. 18F-FDG PET baseline radiomics features improve the prediction of treatment outcome in diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging. 2022;49(3):932–42.
Zhou Y, Ma XL, Zhang T, Wang J, Zhang T, Tian R. Use of radiomics based on (18)F-FDG PET/CT and machine learning methods to aid clinical decision-making in the classification of solitary pulmonary lesions: an innovative approach. Eur J Nucl Med Mol Imaging. 2021;48(9):2904–13.
Sun P, Wang D, Mok V C, Shi L. Comparison of feature selection methods and machine learning classifiers for radiomics analysis in glioma grading. IEEE Access. 2019;102010-102020.
Chang E, Joel MZ, Chang HY, Du J, Khanna O, Omuro A, et al. Comparison of radiomic feature aggregation methods for patients with multiple tumors. Sci Rep. 2021;11(1):9758.
Tixier F, Cheze-Le Rest C, Hatt M, Albarghach NM, Pradier O, Metges J-P, et al. Intratumor Heterogeneity Characterized by Textural Features on Baseline 18F-FDG PET Images Predicts Response to Concomitant Radiochemotherapy in Esophageal Cancer. J Nucl Med. 2011;52:369–78.
Boellaard R, Delgado-Bolton R, Oyen WJ, Giammarile F, Tatsch K, Eschner W. FDG PET/CT: EANM procedure guidelines for tumour imaging: version 2.0. Eur J Nucl Med Mol Imaging. 2015;42(2):328–354.
Huang S, Cai N, Pacheco PP, Narrandes S, Wang Y, Xu W. Applications of Support Vector Machine (SVM) Learning in Cancer Genomics. Cancer Genomics Proteomics. 2018;15(1):41–51.
Zhang Z, Jung C. GBDT-MO: Gradient-Boosted Decision Trees for Multiple Outputs. IEEE Trans Neural Netw Learn Syst. 2021;32(7):3156–67.
Hou N, Li M, He L, Xie B, Wang L, Zhang R, Yu Y, Sun X, Pan Z, Wang K. Predicting 30-days mortality for MIMIC-III patients with sepsis-3: a machine learning approach using XGboost. J Transl Med. 2020;18(1):462.
Zhang L, Liu M, Qin X, Liu G. Succinylation Site Prediction Based on Protein Sequences Using the IFS-LightGBM (BO) Model. Comput Math Methods Med. 2020;2020:8858489.
Sylvester EVA, Bentzen P, Bradbury IR, Clément M, Pearce J, Horne J, Beiko RG. Applications of random forest feature selection for fine-scale genetic population assignment. Evol Appl. 2018;11(2):153–65.
McEligot AJ, Poynor V, Sharma R, Panangadan A. Logistic LASSO Regression for Dietary Intakes and Breast Cancer. Nutrients. 2020;12(9):2652.
Nick TG, Campbell KM. Logistic regression. Methods Mol Biol. 2007;404:273–301.
Moon SH, Kim J, Joung JG, Cha H, Park WY, Ahn JS, Ahn MJ, Park K, Choi JY, Lee KH, et al. Correlations between metabolic texture features, genetic heterogeneity, and mutation burden in patients with lung cancer. Eur J Nucl Med Mol Imaging. 2019;46(2):446–54.
Choi ER, Lee HY, Jeong JY, Choi YL, Kim J, Bae J, Lee KS, Shim YM. Quantitative image variables reflect the intratumoral pathologic heterogeneity of lung adenocarcinoma. Oncotarget. 2016;7(41):67302–13.
Sala E, Mema E, Himoto Y, Veeraraghavan H, Brenton JD, Snyder A, Weigelt B, Vargas HA. Unravelling tumour heterogeneity using next-generation imaging: radiomics, radiogenomics, and habitat imaging. Clin Radiol. 2017;72(1):3–10.
Burrell RA, McGranahan N, Bartek J, Swanton C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature. 2013;501(7467):338–45.
Liu J, Dang H, Wang XW. The significance of intertumor and intratumor heterogeneity in liver cancer. Exp Mol Med. 2018;50(1):e416.
Morris LG, Riaz N, Desrichard A, Şenbabaoğlu Y, Hakimi AA, Makarov V, Reis-Filho JS, Chan TA. Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget. 2016;7(9):10051–63.
Sharma N, Gautam S K, Henry AA, Kumar A. Application of Big Data and Machine Learning. Machine Learning and Big Data: Concepts, Algorithms, Tools and Applications. 2020;305–333.
Gondaliyaa C P, Patel A M, Parikh S M. A Comparative Study on Machine Learning Based Algorithms. Proceedings of 3rd International Conference on Internet of Things and Connected Technologies (ICIoTCT). 2018;26–27.
Sasanelli M, Meignan M, Haioun C, Berriolo-Riedinger A, Casasnovas RO, Biggi A, Gallamini A, Siegel BA, Cashen AF, Véra P, et al. Pretherapy metabolic tumour volume is an independent predictor of outcome in patients with diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging. 2014;41(11):2017–22.
Mikhaeel NG, Smith D, Dunn JT, Phillips M, Møller H, Fields PA, Wrench D, Barrington SF. Combination of baseline metabolic tumour volume and early response on PET/CT improves progression-free survival prediction in DLBCL. Eur J Nucl Med Mol Imaging. 2016;43(7):1209–19.
Zucca E, Cascione L, Ruberto T, Facchinelli D, Schär S, Hayoz S, Dirnhofer S, Giovanella L, Bargetzi M, Mamot C, et al. Prognostic models integrating quantitative parameters from baseline and interim positron emission computed tomography in patients with diffuse large B-cell lymphoma: post-hoc analysis from the SAKK38/07 clinical trial. Hematol Oncol. 2020;38(5):715–25.
Miyazaki Y, Nawa Y, Miyagawa M, Kohashi S, Nakase K, Yasukawa M, Hara M. Maximum standard uptake value of 18F-fluorodeoxyglucose positron emission tomography is a prognostic factor for progression-free survival of newly diagnosed patients with diffuse large B cell lymphoma. Ann Hematol. 2013;92(2):239–44.
Shagera QA, Cheon GJ, Koh Y, Yoo MY, Kang KW, Lee DS, Kim EE, Yoon SS, Chung JK. Prognostic value of metabolic tumour volume on baseline (18)F-FDG PET/CT in addition to NCCN-IPI in patients with diffuse large B-cell lymphoma: further stratification of the group with a high-risk NCCN-IPI. Eur J Nucl Med Mol Imaging. 2019;46(7):1417–27.
Zhao P, Yu T, Pan Z. Prognostic value of the baseline 18F-FDG PET/CT metabolic tumour volume (MTV) and further stratification in low-intermediate (L-I) and high-intermediate (H-I) risk NCCNIPI subgroup by MTV in DLBCL MTV predict prognosis in DLBCL. Ann Nucl Med. 2021;35(1):24–30.
Funding
This work was partially supported by fundings for Clinical Trials from the Affiliated Drum Tower Hospital, Medical School of Nanjing University under Grant No. 2021-LCYJ-MS-04.
This work was also partially supported by fundings for the Key Project of Medical Science and Technology of Nanjing under Grant No.ZKX21011.
Author information
Authors and Affiliations
Contributions
Chong Jiang, Ang Li, Yue Teng, Xiangjun Huang, and Chongyang Ding collected data and analysed data; Jingyan Xu, Jianxin Chen, and Zhengyang Zhou participated in the research design; Chong Jiang and Ang Li contributed to the writing of the manuscript, discussed data, and supervised the study, and all authors performed data analysis and interpretation and read and approved the final version of the article.
Corresponding authors
Ethics declarations
Conflicts of interest/Competing interests
The authors declare that they have no conflicts of interest
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Chong Jiang and Ang Li are co-first authors. They contributed equally to the work.
This article is part of the Topical Collection on Hematology
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, C., Li, A., Teng, Y. et al. Optimal PET-based radiomic signature construction based on the cross-combination method for predicting the survival of patients with diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging 49, 2902–2916 (2022). https://doi.org/10.1007/s00259-022-05717-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-022-05717-9