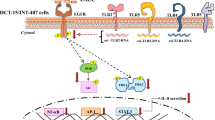
Abstract
Emerging evidence have suggested that aberrant sialylation on cell-surface carbohydrate architecture may influence host–pathogen interactions. The α2,6-sialyltransferase (ST) enzymes were found to alter the glycosylation pattern of the pathogen-infected host cell-surface proteins, which could facilitate its invasion. In this study, we assessed the role of specific α2,6-ST enzymes in the regulation of enteroaggregative E. coli (EAEC)–induced cell signaling pathways in human intestinal epithelial cells. EAEC-induced expression of α2,6-ST family genes in HCT-15 and INT-407 cell lines was assessed at mRNA level by qRT-PCR. Specific esi-RNA was used to silence the target ST-gene in each of the EAEC-infected cell type. Subsequently, the role of these enzymes in regulation of EAEC-induced cell signaling pathways was unraveled by analyzing the expression of MAPkinases (ERK1/2, p38, JNK) and transcription factors (NFκB, cJun, cFos, STAT) at mRNA and protein levels by qRT-PCR and western immunoblotting, respectively, expression of selected sialoglycoproteins by western immunoblotting along with the secretory IL-8 response using sandwich ELISA. ST6GAL-1 and ST6GAL-2 were efficiently silenced in EAEC-infected HCT-15 and INT-407 cells, respectively. Significant reduction in EAEC-induced activation of MAPKs, transcription factors, sialoglycoproteins, and IL-8 secretion was noted in ST-silenced cells in comparison to the respective control cells. We propose that ST6GAL-1 and ST6GAL-2 are quintessential for EAEC-induced stimulation of MAPK-mediated pathways, resulting in activation of transcription factors, leading to an inflammatory response in the human intestinal epithelial cells. Our study may be helpful to design better therapeutic strategies to control EAEC- infection.
Key points
• EAEC induces α2,6-sialyltransferase (ST) upregulation in intestinal epithelial cells
• Target STs (ST6GAL-1 & ST6GAL-2) were efficiently silenced using specific esiRNAs
• Expression of MAPKs, transcription factors & IL-8 was reduced in ST silenced cells
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Altered glycosylation, especially aberrant sialylation of the host cell-surface glycans was found to be associated with bacterial infections (Varki 2017). Such modulation of the glycosylation pattern was shown to mediate bacterial entry and influence the host immune response, hence essential in facilitating bacterial pathogenesis (Day et al. 2015; Lin et al. 2020; Poole et al. 2018). The potential role of pathogen-induced modulation of sialylation has been well established in the case of H. pylori infection, in which an increased expression of sialyl-Lewisa and sialyl-Lewisx was noted on the gastric mucosa. These sialylated glycans were shown as the binding sites for the sialic acid binding adhesin (SabA) of this organism, leading to successful bacterial colonization (Sun et al. 2022). An enhanced sialylation of mucins, found in cystic fibrosis patients, could make them more prone to lung infection by P. aeruginosa (Caballero et al. 2021). Recently, the attachment of spike protein of the SARS-COV-2 to the host’s sialic acids has been found to be essential for the entry of the virus followed by its fusion to the membrane of the host cells (Nguyen et al. 2022).
It was reported that hemagglutinin of human influenza virus A could utilize ST6GAL-1 (ST6β- galactoside α-2,6-sialyltransferase-1)–transferred α2,6-linked sialic acids as the binding partners for its attachment to the human respiratory tract. Further, downregulation of ST6GAL-1 by siRNAs specific for the target gene led to a reduction in virus internalization, thereby generating an effective antiviral response (Wu et al. 2014). Also, an increased level of β3GlcNAcT5, a GlcNAc transferase, has been observed in H. pylori–infected gastric carcinoma cell lines (Marcos et al. 2008). Cumulatively, these evidences suggested that overexpression of ST enzymes could result in hypersialylation, leading to disease progression and thus making ST inhibition as a promising strategy to control the infection. Among various ST inhibitors, P-3Fax-Neu5Ac (peracetylated analog of sialic acid) and cytidine-5′-yl sialyl ethylphosphonate (bisubstrate CMP-sialic acid analog) have been used to inhibit the activity of α2,3- as well as α2,6-STs effectively (Rillahan et al. 2012; Izumi et al. 2005).
Studies have suggested that pathogens could manipulate a myriad of signaling pathways associated with pathogen recognition and invasion, leading to inflammatory responses (Alto and Orth 2012; Bahia et al. 2018). It was shown that Salmonella typhimurium infection could trigger the activation of various MAPKs (mitogen-activated protein kinases), which could play an important role in the stimulation of pro-inflammatory cytokine responses (Hobbie et al. 1997). In this context, it can be mentioned that earlier studies from our laboratory have also shown that IL-8 response induced by EAEC in INT-407 cells might be due to activation of NF-κB and AP-1, mediated by the stimulation of MAPKs (Khan et al. 2010). Further, it was reported that a galactose specific adhesin of EAEC might be a major contributor to IL-8 secretion by these cells through STAT-3 activation in concert with the activation of ERK1/2 (Goyal et al. 2010). A recent report unraveled the role of ST6GAL-1 in promoting sustained signaling through NF-κB and JAK/STAT pathways, the prominent inflammatory pathways in human monocytic cell line (Holdbrooks et al. 2020). Also, in cystic fibrosis patients, an inflammation-induced upregulated expression of IL-8 and IL-6 was shown to be associated with an enhanced level of ST3GAL-6 and GlcNAc-6-O-sulfotransferases, resulting in an accumulation of sialyl-Lewisx and 6-sulfo-sialyl-Lewisx determinants on human airway mucins, which might act as the ligands for P. aeruginosa infection (Groux-Degroote et al. 2008).
However, EAEC-induced modulation of the expression of relevant glycosyltransferases leading to a change in glycosylation pattern in the human intestinal epithelium and their effect on the disease process have not been established till date. Intriguingly, previous reports from our laboratory have shown that EAEC-T8 (an invasive clinical isolate) could induce sialylation (α2,6 linked) of the membrane proteins including VDAC-2 and Prohibitin-2 of HCT-15 and INT-407 cells (Chandel et al. 2022). Therefore, in the present study, we focused on assessing the EAEC-induced expression of all the members of α2,6-ST family in human colonic as well as small intestinal epithelial cell lines. We also investigated the role of specific ST enzymes on the expression of VDAC-2 and Prohibitin-2, various signal transduction pathways, and their involvement in the secretion of IL-8 by these cells.
Materials and methods
Chemicals
The Roswell Park Memorial Institute Medium (RPMI) 1640, Dulbecco’s modified Eagle’s medium (DMEM, high glucose), and fetal bovine serum (FBS) used in the present study were obtained from GIBCO (Thermo Fisher Scientific, Waltham, MA, USA). All the primary and secondary antibodies were procured from Santa Cruz Biotechnology (St. Louis, USA). Specific primers for the various target genes and 3Fax-Peracetyl Neu5Ac were obtained from Sigma Aldrich (USA). Lipofectamine 2000 and TRIzol® were procured from Invitrogen (CA, USA). Luria–Bertani (LB) broth, Trypsin, and EDTA were procured from Hi-media (India).
Cell culture
HCT-15 (human colon carcinoma cell line) and INT-407 (human embryonic small intestinal epithelial cell line), obtained from NCCS (Pune, India), were cultured in RPMI-1640 and high glucose-containing DMEM, respectively, in a CO2 incubator (New Brunswick Eppendorf, Germany) at 37 °C. The media were supplemented with 10% fetal bovine serum (heat inactivated for 30 min at 56 °C), antibiotics (penicillin, 50units/ml; streptomycin, 50 μg/ml), 10-mM HEPES (pH 7.4), sodium bicarbonate (1.0 g/l), glucose (4.5 g/l), and sodium pyruvate (1 mM).
Bacterial strains
EAEC-T8 (an invasive clinical isolate) and EAEC- O42 (non-invasive prototype strain) strains, procured from the NICED (Kolkata, India), and a plasmid-cured EAEC-T8 strain (EAEC-pT8) were used in this study(Konar et al. 2012). Bacteria were cultivated overnight at 37 °C, 150 rpm in LB broth, washed, and suspended (O.D.600 nm = 1 for 6 × 108 bacteria/ml) in appropriate antibiotic-free media for infecting the cell lines.
Infection of cell lines
For EAEC infection, 5 × 106 cells (HCT-15 and INT-407) were seeded in 6-well tissue culture plates and grown to ~ 85% confluency. EAEC strains diluted in serum and antibiotic-free media were used to infect the cell lines at a multiplicity of infection (MOI) of 100 bacteria per epithelial cell for 3 h at 37 °C. The monolayers were washed with PBS (phosphate buffer saline, pH 7.2) to remove extracellular bacteria. The infected cells were detached using Trypsin (0.25%)/1 M EDTA solution and used for subsequent experiments.
RNA isolation
Total RNA was extracted from each cell type, infected with each EAEC strain (EAEC-T8, EAEC-pT8, and EAEC-O42) separately as well as from the uninfected cells (controls) using TRIzol® as per manufacturers’ instructions (Chomczynski and Sacchi 2006). Briefly, TRIzol™ (1 ml) was used to lyse the cells (3 × 106) following incubation (30 min, 4 °C). Subsequently, RNA was extracted using a mixture of chloroform (20%) and isopropanol (50%). The sample was centrifuged (12,000 × g, 10 min, 4 °C), and total RNA was precipitated as a white gel-like pellet. The precipitated RNA pellet was subjected to ethanol (75%) washing by centrifugation (7500 rpm, 5 min) and resuspended in RNase-free water. The RNA concentration was measured spectrophotometrically (Infinite® 200 PRO Microplate reader, TECAN, Switzerland). The absorbance of each sample was taken at 260 nm and 280 nm. The RNA concentration was calculated in μg/ml, where 1O.D. at 260 nm corresponds to 40 μg/ml of RNA. Only samples with A260/A280 ratios of 1.8–2.0 were considered pure and used for further experiments.
cDNA synthesis and qRT-PCR
Briefly, 1 μg of total RNA isolated from both types of cultured cells infected with and without EAEC was reverse transcribed into cDNA, using First Strand cDNA synthesis kit (Thermo Scientific, USA) according to the manufacturers’ instructions. Using cDNA as the template, the expression of each gene of α2,6- ST family in the cells was assessed at mRNA level by qRT-PCR (quantitative reverse transcriptase polymerase chain reaction) on a LightCycler® 96 Instrument (Roche Life Sciences, Germany). For qRT-PCR, the reaction mixture (10 µl) contained 1 µl of template cDNA, 0.75 µl of 0.2 µM specific primer pair (Table 1; KiCqStart primers, Sigma Aldrich, USA), and 5 μl of SYBR green master mix [DyNAmo ColorFlash SYBR Green qPCR Kit (F416L, Thermo Fisher Scientific, USA)]. The thermal profile of the qRT-PCR reaction is given in Table 2. For negative controls, the addition of template cDNA was excluded. The threshold cycle (Ct value) for each mRNA was calculated using the Lightcycler® 96 software. The Ct value of each mRNA was normalized to that of 18S rRNA (internal control) to obtain ΔCt value (ΔCt = Ct target gene – Ct internal control). Subsequently, the fold change of mRNA expression of each gene in the infected cells relative to that in the control cells (uninfected cells) was calculated by using the formula 2–ΔΔCt, where ΔΔCt = ΔCt infected cells – ΔCt control cells (Livak and Schmittgen 2001). All the experiments were conducted thrice in triplicates.
Silencing of specific sialyltransferases
For gene silencing, both the human intestinal cell lines were seeded at a density of 3 × 105 per well in 6-well plates (Greiner, USA) and incubated overnight in a CO2 incubator at 37 °C until they reach 60–70% confluency. The cells were incubated for 2 h in antibiotic- and serum-free media, after which they were transfected for 8 h with 50-pmol esi-RNA against the mRNA of each target sialyltransferase (Sigma Aldrich, USA) separately using lipofectamine 2000. Cells transfected with esi-RNA targeting FLUC (firefly luciferase)-mRNA were used as a non-target negative control, whereas cells treated with only lipofectamine 2000 were taken as experimental control. The transfection media was removed, and the cells were incubated in complete media for an additional 36 h at 37 °C following which infection with EAEC-T8 (at 1:100 MOI) was given for 3 h. Cells were trypsinized, harvested, and washed stringently to remove extracellular bacteria. The transfection efficiency was assessed in view of mRNA expression of the target sialyltransferase by qRT-PCR. 18S rRNA was used as an internal control. Each experiment was done thrice in triplicates.
Effect of silencing of specific sialyltransferase on the activation of MAPKs, transcription factors, and IL-8 response in EAEC-T8–infected cell types
Evaluation of the expression of MAPKs and transcription factors by qRT-PCR
The EAEC-T8–induced expression of 18S rRNA, MAPKs [ERK-1 (MAPK3), ERK-2 (MAPK1), JNK (MAPK8), and p38 (MAPK14)], and transcription factors [NFκB, AP-1 (cJun and cFos), and STAT-3] at mRNA level in both cell lines, pre-transfected with esi-specific sialyltransferase-RNA and esi-FLUC-RNA separately as well as treated with only lipofectamine, was assessed by qRT-PCR. Specific primer sequences (KiCqStart primers, Sigma Aldrich, USA) for each molecule are given in Table 3. Relative fold change of the expression of mRNA of each gene was evaluated as 2–ΔΔCt in the target sialyltransferase-silenced cells in comparison to that of the negative control (esi-FLUC-RNA–transfected cells) and experimental control (lipofectamine-treated cells). All the experiments were performed thrice in triplicates.
Western immunoblotting
The expression of the MAPKs (pERK-1/2, ERK1/2, pp38, p38, pJNK, and JNK) was assessed at the protein level by western immunoblotting of the cell lysates, obtained from each of the esi-specific-sialyltransferase-RNA–transfected, esi-FLUC-RNA–transfected (negative control), and lipofectamine-treated (experimental control) EAEC-T8–infected cell type. In addition, their basal protein expression was also assessed in the uninfected as well as in non-transfected EAEC-T8–infected HCT-15 and INT-407 cells. For this, following incubation (30 min, 4 °C) in lysis buffer [RIPA (50-mM Tris/HCl (pH 8.0) containing 150-mM NaCl, 1.0% NP-40, 0.5% sodium deoxycholate, 0.1% SDS, 10-mM NaVO4, and cocktail protease inhibitors)], the cells were sonicated (30 s on and 30 s off, 5 min). The lysates were centrifuged (10,000 rpm, 10 min, 4 °C), and the supernatants were subjected to 10% SDS-PAGE (Laemmli 1970). After protein transfer (1 h, 350 mA, 20 V) to nitrocellulose membranes (NCM, GE Healthcare, USA) (Towbin et al. 1979), the membranes were treated with 5% BSA in TBST0.1% (TBS containing 0.1% Tween-20) at 37 °C for 2 h. For non-phosphorylated antibodies, 5% skim milk (SM) in TBST0.1% was used instead of BSA. Subsequently, the membranes were incubated (4 °C, overnight) with specific antibody against pERK, pp38, and pJNK (Santa Cruz Biotechnology, USA) separately. The membranes were washed with TBST0.1%, treated (37 °C, 1 h) with the respective HRP-conjugated secondary antibody, finally developed using ECL Western Blotting Detection Reagent (GE Healthcare, USA), and photographed in the FluorChem M system (ProteinSimple, USA). Likewise, to assess the expression of total ERK1/2, p38, and JNK, the bands corresponding to pERK, pp38, and pJNK on the blots were stripped off by incubating the membranes in warm stripping buffer [1.5% glycine, 0.1% SDS, and 1% (v/v) Tween-20 in distilled water (pH 2.2)] for 20 min at 55 °C. This was followed by washing with TBST0.1%, blocking; incubation with specific antibody against total ERK-1/2, p38, and JNK separately as the primary probes; and developed as mentioned before. The dilution used for each antibody is shown in Table 4. The band intensity of each MAPK and the respective phosphorylated MAPK in western immunoblot was quantified by scanning densitometry using ImageJ software. The densitometric value for pERK, ERK1/2, pp38, p38, JNK, and pJNK in each of the EAEC-infected cell type pre-transfected with esi-RNA against specific sialyltransferase and esi-FLUC-RNA separately as well as treated with only lipofectamine was normalized against that of β-actin. The level of expression of pERK, pp38, and pJNK was assessed by normalization of each value against the value of total ERK1/2, p38, and JN,K respectively. Each experiment was done in triplicates.
The expression of transcription factors [NF-κB, AP-1(cFos and cJun), and STAT-3] as well as IκB was assessed at the protein level in each of the target sialyltransferase-silenced EAEC-T8–infected cell type along with the negative control as well as experimental control by western immunoblotting. As mentioned earlier, their basal protein expression was also assessed in the uninfected as well as non-transfected EAEC-T8–infected HCT-15 and INT-407 cells. For this, the cytoplasmic and nuclear extracts from the cells were prepared and fractionated separately by using NE-PER™ Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific™, USA) kit as per manufacturers’ protocol. The cytoplasmic and nuclear proteins were then subjected to western immunoblotting. After blocking, membranes containing the cytoplasmic proteins were incubated (overnight, 4 °C) separately with the specific antibody (diluted in 2.5% SM-TBST0.1%) against NF-κB and total STAT-3 (Santa Cruz Biotechnology), while the membranes containing the nuclear proteins were incubated (overnight, 4 °C) separately with the specific antibody against NF-κB, pSTAT-3, cFos, and cJun (Santa Cruz Biotechnology, USA; Table 4). After washing, the membranes were treated with the respective secondary antibody conjugated with HRP (diluted as before) for 1 h at 37 °C. Bands were developed as described earlier. Also, for the assessment of the expression of cytoplasmic IκB and total nuclear STAT-3, the bands corresponding to cytoplasmic NF-κB and nuclear pSTAT-3 on the blots were stripped off. The blots were reprobed with the specific antibody against IκB and total STAT-3, respectively, and developed. The expression of each protein in each of the sialyltransferase–transfected EAEC-T8–infected cell line was evaluated by scanning densitometry (ImageJ software), as discussed earlier. Further, the level of expression of pSTAT-3 and nuclear NF-κB was obtained by normalization of each value (normalized against β-actin value) against that of STAT-3 (normalized against cytoplasmic total STAT-3-value) and cytoplasmic NF-κB, respectively. Each experiment was performed in triplicates.
Further, to assess the basal level of expression of the MAPKs and transcription factors, their band intensity was also quantified in both uninfected and non-transfected EAEC-T8–infected cell lines. The densitometric values obtained for various cell signaling molecules in both uninfected and EAEC-T8–infected cell lines were normalized against that of β-actin. Each experiment was done in duplicates.
Assessment of IL-8 response
EAEC-T8–induced expression of IL-8 was assessed in each of the esi-FLUC-RNA–transfected (negative control), lipofectamine-treated (experimental control), and specific sialyltransferase-silenced cell type at the mRNA level. Briefly, the RNA was extracted from the cells, and the cDNA was synthesized as mentioned earlier. Subsequently, qRT-PCR was done using specific primers for IL-8 [(forward: 5′-TCCAAACCTTTCCACCCCAA-3′; reverse: 5′-TGCTTGA- AGT TTCACT- GGCATC-3′) KiCqStart primers; Sigma Aldrich, USA], as described earlier. The relative fold change of the expression of IL-8-mRNA was evaluated in view of 2–ΔΔCt in the specific sialyltransferase-silenced cells with respect to negative control and experimental control. 18S rRNA was used as an internal control. All the experiments were done thrice in triplicates.
Further, the level of secretory IL-8 was assessed in the culture supernatants, obtained from each of the specific sialyltransferase-silenced EAEC-T8–infected cell line by IL-8 sandwich ELISA [BD OptEIA ELISA kit (BD PharMingen, CA, USA)], as per manufacturers’ instructions. This assay utilizes anti-human IL-8 as the capture antibody and biotinylated anti-human IL-8 along with HRP-conjugated streptavidin as the detection probe. To determine the unknown IL-8 concentration, a standard curve was made by plotting the known concentrations of IL-8 versus their absorbance values. The level of IL-8 was expressed as pg/ml. Appropriate controls were run in parallel. Each set of the experiment was done thrice in triplicates.
Effect of sialyltransferase inhibitor (3Fax-Peracetyl Neu5Ac) on IL-8 level of EAEC-T8–infected cells
In another set of experiments, the effect of sialyltransferase inhibitor (3Fax-Peracetyl Neu5Ac) on IL-8 response in each cell type (uninfected and EAEC-T8–infected cells) was assessed. Briefly, HCT-15 and INT-407 cells (5 × 105 cells/well in a 6-well plate) were incubated in the presence and absence of P-3Fax-Neu5Ac (64 µmol/l) (Rillahan et al. 2012). The cells were cultured for 72 h followed by infection with EAEC-T8 for 3 h. The cell pellets and the culture supernatants were collected. RNA was extracted from both types of cells, and cDNA was synthesized as mentioned earlier. The expression of IL-8 was assessed at mRNA level by qRT-PCR. Further, the level of secretory IL-8 in the culture supernatants was estimated by sandwich ELISA. All the experiments were performed thrice in triplicates.
Effect of silencing of specific sialyltransferase on the expression of EAEC-induced sialoglycoproteins in each cell line
Western immunoblotting was performed to assess the level of expression of sialoglycoproteins (VDAC-2 and Prohibitin-2) in each of the EAEC-T8–infected cell line, pre-transfected with esi-RNA against each specific sialyltransferase and esi-FLUC-RNA (negative control) separately as well as in the cells pre-treated with lipofectamine (experimental control). For this, membrane proteins were isolated from each cell type and subjected to western immunoblotting (Chandel et al. 2022). The membranes were incubated separately with specific antibodies against each sialoglycoprotein and developed using the above mentioned protocol. The glycoprotein expression was assessed in each of the EAEC-infected specific sialyltransferase-silenced cell line in comparison to the respective control cells using scanning densitometry (ImageJ software), as described earlier. Each experiment was done in triplicates.
Effect of silencing of specific sialyltransferase on adherence of EAEC to each cell type
The effect of silencing of specific sialyltransferase on the adherence of EAEC to each cell line was assessed. Briefly, cells were grown to 50% confluent monolayers, and the media were replaced with antibiotic-free D-mannose (0.5%) containing fresh media. Silencing of each specific sialyltransferase in HCT-15 and INT-407 cell lines was carried out as mentioned earlier. This was followed by EAEC-T8 infection (3 h at 37 °C) of the cells and washing to remove the non-adherent bacteria. The cells were fixed in methanol (70%, 5 min), stained with Giemsa stain (10%, Hi-Media, India), and observed under light microscope for the assessment of EAEC adherence. Also, in parallel, cells under the same conditions were washed with sterile warm PBS to remove non-adherent bacteria followed by lysis in PBS containing 0.5% (v/v) Triton X-100 for 5 min at 37 °C. The suspension containing bacterial cells was serially diluted in PBS and plated on LB agar to quantify the number of colony-forming units (CFU) per ml (Feeney et al. 2017). Cells transfected with esi-FLUC-RNA and treated only with lipofectamine 2000 were run as controls.
Statistical analysis
GraphPad Prism 5 Version 5.01 (GraphPad Software, San Diego, USA) was employed for statistical analysis. The data presented as the mean with standard deviation (mean ± S.D.) were analyzed using ANOVA, followed by Tukey’s test, where p < 0.05 was considered as significant.
Results
Expression of α2,6-ST family genes was altered in EAEC-infected intestinal epithelial cell lines at mRNA level
In the present study, qRT-PCR data revealed altered expression of α2,6-ST family genes (ST6GAL-1, ST6GAL-2, ST6GALNAc-1, ST6GALNAc-2, ST6GALNAc-3, ST6GALNAc-4, ST6GALNAc-5, ST6GALNAc-6) at mRNA level in both EAEC-T8, EAEC-pT8, and EAEC-O42–infected HCT-15 and INT-407 cells as compared to the respective uninfected cells (control). The ΔCt values of the α2,6-ST family genes in both cell lines are given in Table S1 (A and B), respectively. Figure 1 shows the graphical representation of the expression of all the eight genes of α2,6-ST family in each of the control and EAEC-infected cell line. Analysis of data revealed that EAEC-T8 could induce maximum expression of ST6GAL-1 in HCT-15 cells and ST6GAL-2 in INT-407 cells at mRNA level among all other α2,6-ST family genes, coding the enzymes, known to be responsible for the transfer of sialic acids in α2,6 linkage to the oligosaccharides of the glycoproteins. Thus, ST6GAL-1 and ST6GAL-2 were selected as the target ST genes to be silenced in the EAEC-T8–infected respective cell lines.
Relative expression profile of ST6-family genes at mRNA level in both cell lines infected with and without EAEC. One-way ANOVA, followed by Tukey’s multiple comparison test, was applied; ***p < 0.001, **p < 0.01, and *p < 0.05 [EAEC-T8, EAEC-pT8, and EAEC-O42 vs. uninfected (UI) cells]; ns, non-significant
The target ST genes were silenced efficiently in EAEC-T8–infected intestinal cells
The level of expression of EAEC-T8–induced ST6GAL-1 and ST6GAL-2 transcripts was found to be reduced significantly in the esi-RNA-ST6GAL-1 and esi-RNA-ST6GAL-2–silenced HCT-15 and INT-407 cells, respectively, as compared to the esi-FLUC-RNA–transfected (negative control) and lipofectamine-treated cells (experimental control), as assessed by qRT-PCR (Fig. 2). ∆Ct values of ST6GAL-1 and ST6GAL-2 transcripts in each of the esi-ST6GAL-1-RNA, esi-ST6GAL-2-RNA, and esi-RNA-FLUC–transfected and only lipofectamine-treated EAEC-T8–infected cell type are shown in Table S2. The silencing efficiencies of ST6GAL-1 and ST6GAL-2 were calculated as 70.8% and 95.7% in the respective cell line, respectively.
Relative expression profile of EAEC-T8–induced target sialyltransferases at mRNA level in both cell lines infected with and without EAEC. The bar diagrams show the expression fold of a ST6GAL-1 and b ST6GAL-2 at mRNA level in ST6GAL-1 and ST6GAL-2–silenced respective cell type as compared to the negative control (EAEC-T8–infected cells transfected with esi-FLUC RNA) and experimental control (EAEC-T8–infected cells treated with only lipofectamine); one-way ANOVA, followed by Tukey’s multiple comparison test, was used; ***p < 0.001 (ST6GAL-1 and ST6GAL-2–silenced cells vs. experimental control); ns, non-significant (negative control vs. experimental control)
Silencing of target sialyltransferases in EAEC-T8–infected intestinal epithelial cells attenuated the expression of MAPKs, transcription factors, and IL-8 secretion
A decreased expression of EAEC-T8–induced activated MAPKs [ERK-1/2 (MAPK3/1), p38 (MAPK14), and JNK (MAPK8)], transcription factors (NFκB, AP-1, and STAT-3), and IL-8 at both mRNA and protein levels was noted in ST6GAL-1 and ST6GAL-2–silenced HCT-15 and INT-407 cells, respectively. The ∆Ct values for the transcript of each of these genes in target sialyltransferase-silenced cells in comparison to the respective negative control and experimental control are shown in Table S3 (A and B). Expression analysis of the EAEC-T8–induced transcripts in esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA–transfected respective cell type revealed downregulation of ERK1, ERK2, JNK, p38, NFκB, cJun, cFos, STAT-3, and IL-8 as compared to the respective controls (Fig. 3A). The fold change for each of these genes in target sialyltransferase-silenced cells in comparison to the respective controls is shown in Table S3 (C).
A Relative expression profile of cell signaling molecules at mRNA level in EAEC-T8–infected target ST-silenced respective cell types. The bar diagrams indicate EAEC-T8–induced expression of ERK1, ERK2, JNK, p38, NFκB, cJun, cFos, STAT3, and IL-8 in ST6GAL-1 and ST6GAL-2–silenced HCT-15 and INT-407 cells, respectively, as compared to respective negative control and experimental control. One-way ANOVA, followed by Tukey’s multiple comparison test, was used; ***p < 0.001, **p < 0.01, and *p < 0.05 indicate the level of significance; ns, non-significant. B Expression profile of cell signaling molecules at protein level in cells. Western blots showing the expression of a MAPKs [pERK1/2, ERK1, ERK2, pJNK, JNK, pp38, and p38] and b transcription factors [IκB, NF-κB (cytoplasmic), NF-κB (nuclear), cFos (nuclear), cJun (nuclear), pSTAT-3 (nuclear) and STAT-3 (nuclear)] in ST6GAL-1 and ST6GAL-2–silenced respective HCT-15 and INT-407 cell types as compared to the respective negative control and experimental control. In the case of MAPKs, the blots were probed with anti-pERK, anti-pJNK, and anti-pp38 as the primary antibodies followed by incubation with HRP-conjugated secondary antibody. The bands on the blots were then stripped off and reprobed with the antibody to total ERK2 and ERK1, JNK, and p38. For transcription factors, the blots were probed with NF-κB (cytoplasmic), NF-κB (nuclear), cFos, cJun, and pSTAT-3 (nuclear) as the primary antibodies followed by incubation with HRP-conjugated secondary antibody. The NF-κB (cytoplasmic) and pSTAT-3 (nuclear) bands on the blots were then stripped off and reprobed with the antibody to IκB and STAT-3. Normalization was done by using β-actin (internal control). C1, lipofectamine-treated; C2, esi-FLUC-RNA–transfected; T, esi-ST6GAL-1-RNA/esi-ST6GAL-2-RNA–transfected EAEC-T8–infected HCT-15 and INT-407 cells; Mr, protein markers. C Bar diagrams revealing the expression of a MAPKs and b transcription factors in EAEC-T8–infected ST6GAL-1 and ST6GAL-2–transfected respective cell line as evaluated by Scion analysis with ImageJ software. The intensity of the band of each parameter was normalized to that of β-actin as the internal control. Further, the level of expression of each activated MAPK was obtained by normalization of each value against the value of respective total MAPK. The expression of each molecule in lipofectamine-treated control cells (C1) was set to 1, with respect to which the expression of the respective molecule in esi-FLUC RNA–transfected cells (C2) and esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA–transfected cells (T) was calculated. D ELISA-based estimation of IL-8 secretion by EAEC-T8–infected target ST-silenced respective cell types as compared to that of response in control cells (lipofectamine-treated cells and esi-FLUC RNA–transfected cells). One-way ANOVA, followed by Tukey’s multiple comparison test, was applied which revealed a significant reduction in secretory IL-8 concentration in each of the ST-silenced cell line [ST6GAL-1–silenced HCT-15 cells, 1451.7 pg/ml; ST6GAL-2–silenced INT-407 cells, 1389.1 pg/ml] in comparison to the respective vehicle control [HCT-15, 2269.583 pg/ml; INT-407, 2230 pg/ml); ***p < 0.001 (esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA vs. the lipofectamine-treated cells); ns, non-significant (esi-FLUC-RNA vs. the lipofectamine-treated cells)
The results obtained by qRT-PCR were further corroborated at the protein level by western immunoblotting. The intensity of the bands corresponding to the MAPKs (pERK1/2, ERK1, ERK2, pJNK, JNK pp38, and p38) and transcription factors (cytoplasmic NFκB, nuclear NFκB, nuclear cFos, nuclear cJun, nuclear pSTAT3, and nuclear STAT3) was found to be reduced in contrast to cytoplasmic IκB which was found to show an increased expression in each of the target ST-silenced cell type in comparison to the respective controls as shown in Fig. 3B (a and b). Replica blots have been shown in Fig. S1. Figure 3C shows the representative bar diagrams demonstrating the reduced expression of pERK, pJNK, pp38, nuclear NF-κB, cJun, cFos, and nuclear pSTAT-3 along with an increased expression of IκB in the case of esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA–transfected respective cell type as compared to the respective controls. The percentage change in the expression for each of these genes in esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA–silenced cells as compared to the respective controls is shown in Table S3 (D). Also, the representative western immunoblots displaying the basal level of protein expression of various MAPKs and transcription factors along with the respective bar diagrams in the case of both uninfected and EAEC-T8–infected cell lines have been shown in Fig. S2. The bar diagrams reflecting the expression fold (as assessed by densitometric analysis) in ST6GAL1 and ST6GAL2–silenced EAEC-T8–infected HCT-15 and INT-407 cells in comparison to the uninfected, esi-RNA-FLUC–transfected, and only lipofectamine-treated EAEC-T8–infected cells are shown in Fig. S3.
Further, the EAEC-T8–induced secretory IL-8 concentration in each of the ST-silenced cell line was reduced significantly in comparison to the respective vehicle control as shown in Fig. 3D.
P-3Fax-Neu5Ac–mediated inhibition of sialylation in intestinal epithelial cell lines hindered EAEC-T8–induced IL-8 secretion
A significant reduction in the expression of EAEC-T8–induced IL-8 at the mRNA level was noted in each intestinal epithelial cell type in the presence of a pan-sialyltransferase inhibitor [3Fax-Peracetyl Neu5Ac (P-3Fax-Neu5Ac)] as compared to the respective control cells in the absence of the inhibitor (Fig. 4A). The ∆Ct values for the IL-8 transcript in both the uninfected and EAEC-T8–infected cell lines pre-treated with P-3Fax-Neu5Ac as well as control cells [DMSO (vehicle)] are shown in Table S4.
A Expression profile of IL-8 at mRNA level and B ELISA-based estimation of IL-8 secretion in the case of both cell lines infected with and without EAEC-T8 in the presence of sialyltransferase inhibitor (P-3Fax-Neu5Ac). Reduced levels of secretory IL-8 in both inhibitor-treated EAEC-T8–infected cell lines (HCT-15, 2041.7 pg/ml; INT-407, 1177.7 pg/ml) in comparison to the respective infected cells (HCT-15, 2501.4 pg/ml; INT-407, 2063.7 pg/ml) in the absence of inhibitor. One-way ANOVA, followed by Tukey’s multiple comparison test, was used; ***p < 0.001 indicates the level of significance
This observation was supported by the reduced level of secretory IL-8 in both inhibitor-treated EAEC-T8–infected cell lines in comparison to the respective infected cells in the absence of inhibitor (Fig. 4B).
Silencing of target sialyltransferases downregulated the expression of Prohibitin-2 and VDAC-2 sialoglycoproteins
The expression of the identified sialoglycoproteins (VDAC-2 and Prohibitin-2) was found to be reduced in the membrane fractions of each of the target ST-silenced cell type as compared to the respective control cells as observed in Fig. 5. The corresponding bar diagrams reflecting their expression fold (as assessed by densitometric analysis) in HCT-15 in comparison to the respective controls are shown in Fig. 5.
A Expression profile of Prohibitin-2 and VDAC-2 at protein level in EAEC-T8–infected ST6GAL-1 and ST6GAL-2–silenced respective cell types. The blots were probed with the antibodies against Prohibitin-2 and VDAC-2 separately followed by incubation with HRP-conjugated secondary antibody. Normalization was done by using β-actin as the internal control. C1, lipofectamine-treated; C2, esi-FLUC RNA–transfected; T, esi-ST6GAL-1-RNA/esi-ST6- GAL-2-RNA–transfected EAEC-T8–infected HCT-15 and INT-407 cells; Mr, protein markers. Corresponding bar diagrams indicate the percentage decrease in the expression of Prohibitin-2 and VDAC-2 in ST6GAL-1 and ST6GAL-2–silenced respective cell lines [(HCT-15: PHB2, 56.36%; VDAC2, 82.30%); (INT-407 cells: PHB2, 29.33%; VDAC2, 81.47%)] as obtained by ImageJ analysis of western blots in comparison to control cells (lipofectamine-treated cells and esi-FLUC RNA–transfected cells). One-way ANOVA, followed by Tukey’s multiple comparison test, was applied; ***p < 0.001 (esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA vs. the lipofectamine-treated cells); ns, non-significant (esi-FLUC-RNA vs. the lipofectamine-treated cells)
Silencing of sialyltransferases diminishes EAEC adherence to the cells
The aggregative adherence of EAEC-T8 to ST6GAL-1 and ST6GAL-2–silenced cells was found to be reduced appreciably as compared to the respective control cells, as depicted in Fig. 6A and B. Figure 6C and D shows a 45.2% and 38.2% reduction in view of CFU counts (as reflected in bar diagrams) in ST6GAL-1 and ST6GAL-2–silenced HCT-15 and INT-407 cells, respectively, in comparison to vehicle control.
Aggregative adherence of EAEC-T8 to A HCT-15 and B INT-407 cells transfected with esi-ST6GAL-1-RNA and esi-ST6-GAL-2-RNA as assessed by Giemsa staining. C and D depict the aggregative adherence of EAEC-T8 to silenced HCT-15 and INT-407 cells as shown by bar diagrams in view of CFU. **p < 0.01 (iii vs. i in the case of HCT-15); ***p < 0.001 (iii vs. i in the case of INT-407) and non-significant (ii vs. i). One-way ANOVA followed by Tukey’s multiple comparison test in comparison to lipofectamine-treated cells, was applied. Each bar represents the mean ± S.D. of three independent experiments. (iii) Cells transfected with esi-ST6GAL-1-RNA and esi-ST6GAL-2-RNA, (ii) cells transfected with esi-FLUC-RNA, and (i) cells treated with lipofectamine
Discussion
Host–pathogen crosstalk is crucial in understanding the underlying mechanism of any successful microbial infection. Pathogens are known to manipulate the glycosylation patterns of the host cell-surface proteins, which are mediated by specific glycosyltransferases (Lin et al. 2020). Our previous report regarding identification of several differentially expressed α2,6-linked sialic acid–containing glycoproteins on EAEC-T8 (an invasive clinical isolate of EAEC)–infected human colonic (HCT-15) and small intestinal (INT-407) epithelial cell lines as well as the importance of pathogen-induced α2,6-linked sialylation in infection led us to evaluate the expression level of all the genes of α2,6 sialyltransferase family in these cells infected with EAEC-T8, its plasmid-cured counterpart, and EAEC-O42 (a prototype non-invasive strain of EAEC) separately (Chandel et al. 2022). Among various upregulated transcripts of α2,6-ST family members, known to catalyze the transfer of Neu5AC from donor CMP-Neu5Ac to the terminal sugar residue of the oligosaccharides on acceptor glycoproteins in α2,6 linkage, ST6GAL-1 and ST6GAL-2 were selected as the targets to be silenced using specific esi-RNA in EAEC-T8–infected HCT-15 and INT-407 cells, respectively. Although an increased expression of ST6GALNAc-3, ST6GALNAc-5, and ST6GALNAc-6 in HCT-15 cell line and ST6GALNAc-5 in INT-407 cell line was observed following EAEC infection, these sialyltransferases are responsible for catalyzing the transfer of sialic acid to the Gal residue of glycolipids via an α2,6 linkage (Hugonnet et al. 2021).
ST6GAL-1 is known to catalyze the transfer of Neu5Ac to the terminal Gal residue of Galβ1,4- GlcNAc on N-glycans of glycoproteins (Hugonnet et al. 2021). It is known as an acute phase reactant and is released during generation of an inflammatory state in humans, induced by trauma, bacterial infections, and stress (Irons et al. 2020). Studies have reported a link between virus replication and modifications in the cell-surface N-glycans via an increased level of ST6GAL-1 in HBV-infected hepatoma cell line (HepAD38). Further, knockdown of ST6GAL-1 was found to reduce the virus multiplication, as revealed by 90% decrease in the HBV cDNA (Priyambada et al. 2018).
ST6GAL-2 has also been found to catalyze the transfer of sialic acid to an oligosaccharide; however, unlike ST6GAL-1, it preferentially utilizes GalNAcβ1,4GlcNAc as the acceptor substrate (Laporte et al. 2009). Although studies have illustrated the involvement of ST6GAL-2 in follicular thyroid carcinoma, melanoma, and breast carcinoma, its importance in bacterial infections is still unexplored. In this study, we focused on elucidating the role of ST6GAL-1 and ST6GAL-2 on EAEC-induced signal transduction pathways in colonic and small intestinal epithelial cells, respectively.
The MAPK signaling cascade is known to respond to external stimuli including growth factors and stress signals, resulting in induction of cellular growth, differentiation, and proliferation (Cargnello and Roux 2011). Microbes may manipulate these signaling pathways to subvert the host immune responses, subsequently facilitating the pathogenesis (Krachler et al. 2011). Prominent alterations in the expression of various MAPKs and IL-8 secretion were reported during infection of intestinal epithelial cell lines by enteropathogenic, enterotoxigenic, and enterohemorrhagic E. coli (de Grado et al. 2001; Dahan et al. 2002; Ren et al. 2014).
The activated NF-κB (a transcription factor) has been found to play a central role in the regulation of infection-associated inflammation. High level of activated NF-κB was found to be of significance in the pathogenesis of organisms like Salmonella typhimurium and Helicobacter pylori (Hobbie et al. 1997; Kim et al. 2006). Additionally, the activity of AP-1 (activator protein-1; composed of cJun and cFos proteins) along with the JAK/STAT signaling pathway might also be altered in response to bacterial infections (Mohammed et al. 2007). Both AP-1 and STAT pathways have been implicated in cellular events of apoptosis, differentiation, cell survival, and inflammation. Studies have highlighted an elevated level of activated AP-1 and STAT-3 in respiratory tract infections due to Chlamydia pneumoniae and gastrointestinal infections due to Helicobacter pylori (Lu et al. 2017; Wang et al. 2013). The concerted action of these three transcription factors resulted in an increased secretion of IL-8 (Liu et al. 2018). These observations are in consensus with our previous findings, where EAEC and its galactose specific adhesin–mediated secretion of IL-8 by human small intestinal epithelial cells was found to be orchestrated via activation of AP-1 and STAT-3–mediated signaling pathways (Khan et al., 2010; Goyal et al. 2010).
Although the role of sialyltransferases has not been thoroughly investigated in pathogen-induced cell signaling pathways, several reports have indicated their involvement in cancers of different origin. An elevated expression of ST6GAL-1 was found to be associated with colon carcinoma (Swindall et al. 2013). Further, ST6GAL-1 mediated activation of PI3K/AKT signaling in hepatocellular carcinoma cells and prostate cancer cells suggesting its role in mediating proliferation of these cells (Wei et al. 2016; Zhao et al. 2014). In the present study, the potential role of ST6GAL-1 and ST6GAL-2 in modulation of EAEC-induced signal transduction pathways was highlighted, as a reduced level of MAPKs and transcription factors was noted in ST6GAL-1 and ST6GAL-2–silenced HCT-15 and INT-407 cells following EAEC infection.
A correlation between altered glycosylation and inflammation was shown in several reports (Kreisman and Cobb 2012; Reily et al. 2019). Matsuzwa et al. (2003) demonstrated an upregulated expression of α1,4GlcNAc- transferase (α4GnT) in H. pylori–induced gastritis and a positive correlation between the increased level of α4GnT and the degree of inflammation in gastric biopsy samples. Our findings regarding the decrease in the level of EAEC-induced secretory IL-8 in the ST6GAL-1 and ST6GAL-2–silenced respective cell type revealed their involvement in the regulation of IL-8 response in these cells. Further, a synonymous reduction in IL-8 levels was also observed in each of the EAEC-infected cell type following treatment with 3Fax-Peracetyl Neu5Ac (a pan-sialyltransferase inhibitor), which is in good agreement with our earlier observation. Although not much information exists regarding the link between IL-8 secretion and sialyltransferases in bacterial infections, an increased expression of ST6GAL-1 has been implicated in augmenting the levels of both IL-8 and IL-6 in the case of pancreatic cancer and ovarian cancer (Britain et al. 2017). In this context, it is to be mentioned that the anti-inflammatory role IgG, a pro-inflammatory mediator of the humoral immune response, was reported due to the addition of terminal sialic acid to the glycan of its Fc fragment, resulting in the conformational alteration and acquisition of novel binding activity of this protein (Anthony et al., 2008). This report is contradictory to our finding regarding the role of α2,6-linked sialic acid in EAEC-induced inflammatory response in the intestinal epithelial cells. However, Ramos et al. (2011) have shown that avian influenza viruses could bind to α2,3-linked sialic acids on human host cell surface and aggravate pro-inflammatory immune response and cytokine release, which corroborates our observation.
In a previous study, we have shown that EAEC infection could induce the expression of α-2,6 sialylated Prohibitin-2 and VDAC2 on the host cell surface, which might aid in the aggregative adherence of EAEC to the intestinal epithelial cells. In the present study, the reduced expression of both these sialylated glycoproteins in each of the target sialyltransferase-silenced EAEC-infected cell type coupled with attenuated aggregative adherence of this organism to these cells suggested the possible role of ST6GAL-1 and ST6GAL-2 in the addition of sialic acid to these proteins in a specific linkage thus influencing bacterial adherence. In consensus with our findings, Lin et al. (2002) have demonstrated that silencing of ST6GAL-1 in breast carcinoma cell lines resulted in reduced expression of membrane-bound α2 → 6-linked sialic acid glycoconjugates. Further, Sakarya et al. (2010) reported that adherence of Salmonella typhi to colonic cells could be mediated via sialic acid moieties and removal of these sialic acid residues could reduce S. typhi binding by 41%.
Taken together, our findings have unraveled that EAEC induced upregulation of ST6GAL-1 and ST6GAL-2 in human colonic and small intestinal epithelial cells. The sialyltransferase-mediated α2,6 sialylation of the cell surface of VDAC-2 and Prohibitin-2 might be quintessential for the activation of various cell signaling molecules leading to the inflammatory response in these cells. The silencing of these sialyltransferases revealed reduced expression of the sialoglycoproteins, MAPKs, transcription factors, and secretory IL-8. Further, a decrease in the IL-8 level in each intestinal cell type following treatment with a pan-sialyltransferase inhibitor (P-3Fax-Neu5Ac) suggested that the infection process may be modulated by using specific inhibitors of these sialyltransferases. Based on the existing literature, this is the first report highlighting the EAEC-induced alteration in the expression of specific sialyltransferase enzymes and their subsequent effect on signal transduction pathways, leading to IL-8 secretion.
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Alto NM, Orth K (2012) Subversion of cell signaling by pathogens. Cold Spring Harb Perspect Biol 4(9):a006114. https://doi.org/10.1101/cshperspect.a006114
Anthony RM, Nimmerjahn F, Ashline DJ, Reinhold VN, Paulson JC, Ravetch JV (2008) A recombinant IgG Fc that recapitulates the anti-inflammatory activity of IVIG. Science 320(5874):373. https://doi.org/10.1126/science.1154315
Bahia D, Satoskar AR, Dussurget O (2018) Editorial: cell signaling in host-pathogen interactions: the host point of view. Front Immunol 9:221. https://doi.org/10.3389/fimmu.2018.00221
Britain CM, Dorsett KA, Bellis SL (2017) The glycosyltransferase ST6Gal-I protects tumor cells against serum growth factor withdrawal by enhancing survival signaling and proliferative potential. J Biol Chem 292(11):4663–4673. https://doi.org/10.1074/jbc.M116.763862
Caballero I, Ringot-Destrez B, Si-Tahar M, Barbry P, Guillon A, Lantier I, Berri M, Chevaleyre C, Fleurot I, Barc C, Ramphal R, Pons N, Paquet A, Lebrigand K, Baron C, Bähr A, Klymiuk N, Léonard R, Robbe-Masselot C (2021) Evidence of early increased sialylation of airway mucins and defective mucociliary clearance in CFTR-deficient piglets. J Cyst Fibros 20(1):173–182. https://doi.org/10.1016/j.jcf.2020.09.009
Cargnello M, Roux PP (2011) Activation and function of the MAPKs and their substrates, the MAPK-activated protein kinases. Microbiol Mol Biol Rev 75(1):50–83. https://doi.org/10.1128/mmbr.00031-10
Chandel S, Joon A, Ghosh S (2022) Enteroaggregative Escherichia coli induces altered glycosylation in membrane proteins of cultured human intestinal epithelial cells. Biochimie 199:68–80. https://doi.org/10.1016/j.biochi.2022.04.002
Chomczynski P, Sacchi N (2006) The single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction: twenty-something years on. Nat Protoc 1(2):581–585. https://doi.org/10.1038/nprot.2006.83
Dahan S, Busuttil V, Imbert V, Peyron JF, Rampal P, Czerucka D (2002) Enterohemorrhagic Escherichia coli infection induces interleukin-8 production via activation of mitogen-activated protein kinases and the transcription factors NF-kappaB and AP-1 in T84 cells. Infect Immun 70(5):2304–2310. https://doi.org/10.1128/IAI.70.5.2304-2310.2002
Day CJ, Tran EN, Semchenko EA, Tram G, Hartley-Tassell LE, Ng PS, King RM, Ulanovsky R, McAtamney S, Apicella MA, Tiralongo J, Morona R, Korolik V, Jennings MP (2015) Glycan:glycan interactions: high affinity biomolecular interactions that can mediate binding of pathogenic bacteria to host cells. Proc Natl Acad Sci USA 112(52):E7266–E7275. https://doi.org/10.1073/pnas.1421082112
de Grado M, Rosenberger CM, Gauthier A, Vallance BA, Finlay BB (2001) Enteropathogenic Escherichia coli infection induces expression of the early growth response factor by activating mitogen-activated protein kinase cascades in epithelial cells. Infect Immun 69(10):6217–6224. https://doi.org/10.1128/IAI.69.10.6217-6224.2001
Feeney S, Ryan JT, Kilcoyne M, Joshi L, Hickey R (2017) Glycomacropeptide reduces intestinal epithelial cell barrier dysfunction and adhesion of entero-hemorrhagic and entero-pathogenic Escherichia coli in vitro. Foods 6(11):93
Goyal A, Konar M, Setia A, Narang A (1800) Ghosh S (2010) Galactose specific adhesin of enteroaggregative E. coli induces IL-8 secretion via activation of MAPK and STAT-3 in INT-407 cells. Biochim Biophys Acta 6:574–9. https://doi.org/10.1016/j.bbagen.2010.03.009
Groux-Degroote S, Krzewinski-Recchi MA, Cazet A, Vincent A, Lehoux S, Lafitte JJ, Van Seuningen I, Delannoy P (2008) IL-6 and IL-8 increase the expression of glycosyltransferases and sulfotransferases involved in the biosynthesis of sialylated and/or sulfated Lewisx epitopes in the human bronchial mucosa. Biochem J 410(1):213–223. https://doi.org/10.1042/bj20070958
Hobbie S, Chen LM, Davis RJ, Galán JE (1997) Involvement of mitogen-activated protein kinase pathways in the nuclear responses and cytokine production induced by Salmonella typhimurium in cultured intestinal epithelial cells. J Immunol 159(11):5550–5559
Holdbrooks AT, Ankenbauer KE, Hwang J, Bellis SL (2020) Regulation of inflammatory signaling by the ST6Gal-I sialyltransferase. PLoS One 15(11):e0241850. https://doi.org/10.1371/journal.pone.0241850
Hugonnet M, Singh P, Haas Q, von Gunten S (2021) The distinct roles of sialyltransferases in cancer biology and onco-immunology. Front Immunol 12:799861. https://doi.org/10.3389/fimmu.2021.799861
Irons EE, Punch PR, Lau JTY (2020) Blood-borne ST6GAL1 regulates immunoglobulin production in B cells. Front Immunol 11:617. https://doi.org/10.3389/fimmu.2020.00617
Izumi M, Wada K, Yuasa H, Hashimoto H (2005) Synthesis of bisubstrate and donor analogues of sialyltransferase and their inhibitory activities. J Org Chem 70(22):8817–8824. https://doi.org/10.1021/jo0512608
Khan K, Konar M, Goyal A, Ghosh S (2010) Enteroaggregative Escherichia coli infection induces IL-8 production via activation of mitogen-activated protein kinases and the transcription factors NF-kappaB and AP-1 in INT-407 cells. Mol Cell Biochem 337(1–2):17–24. https://doi.org/10.1007/s11010-009-0282-3
Kim SY, Lee YC, Kim HK, Blaser MJ (2006) Helicobacter pylori CagA transfection of gastric epithelial cells induces interleukin-8. Cell Microbiol 8(1):97–106. https://doi.org/10.1111/j.1462-5822.2005.00603.x
Konar M, Sachin O, Priya A, Ghosh S (2012) Identification of key proteins of cultured human intestinal cells involved in interaction with enteroaggregative Escherichia coli. FEMS Immunol Med Microbiol 66(2):177–190. https://doi.org/10.1111/j.1574-695X.2012.00998.x
Krachler AM, Woolery AR, Orth K (2011) Manipulation of kinase signaling by bacterial pathogens. J Cell Biol 195(7):1083–1092. https://doi.org/10.1083/jcb.201107132
Kreisman LS, Cobb BA (2012) Infection, inflammation and host carbohydrates: a glyco-evasion hypothesis. Glycobiology 22(8):1019–1030. https://doi.org/10.1093/glycob/cws070
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685. https://doi.org/10.1038/227680a0
Laporte B, Gonzalez-Hilarion S, Maftah A, Petit JM (2009) The second bovine beta-galactoside-alpha2,6-sialyltransferase (ST6Gal II): genomic organization and stimulation of its in vitro expression by IL-6 in bovine mammary epithelial cells. Glycobiology 19(10):1082–1093. https://doi.org/10.1093/glycob/cwp094
Lin B, Qing X, Liao J, Zhuo K (2020) Role of protein glycosylation in host-pathogen interaction. Cells 9(4):1022 https://doi.org/10.3390/cells9041022
Lin S, Kemmner W, Grigull S, Schlag PM (2002) Cell surface alpha 2,6 sialylation affects adhesion of breast carcinoma cells. Exp Cell Res 276(1):101–110. https://doi.org/10.1006/excr.2002.5521
Liu X, Yin S, Chen Y, Wu Y, Zheng W, Dong H, Bai Y, Qin Y, Li J, Feng S, Zhao P (2018) LPS-induced proinflammatory cytokine expression in human airway epithelial cells and macrophages via NF-κB, STAT3 or AP-1 activation. Mol Med Rep 17(4):5484–5491. https://doi.org/10.3892/mmr.2018.8542
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta C(T)) Method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Lu R, Zhang YG, Sun J (2017) STAT3 activation in infection and infection-associated cancer. Mol Cell Endocrinol 451:80–87. https://doi.org/10.1016/j.mce.2017.02.023
Marcos NT, Magalhães A, Ferreira B, Oliveira MJ, Carvalho AS, Mendes N, Gilmartin T, Head SR, Figueiredo C, David L, Santos-Silva F, Reis CA (2008) Helicobacter pylori induces beta3GnT5 in human gastric cell lines, modulating expression of the SabA ligand sialyl-Lewis x. J Clin Invest 118(6):2325–2336. https://doi.org/10.1172/jci34324
Matsuzwa M, Ota H, Hayama M, Zhang MX, Sano K, Honda T, Ueno I, Akamatsu T, Nakayama J (2003) Helicobacter pylori infection up-regulates gland mucous cell-type mucins in gastric pyloric mucosa. Helicobacter 8(6):594–600. https://doi.org/10.1111/j.1523-5378.2003.00185.x
Mohammed KA, Nasreen N, Antony VB (2007) Bacterial induction of early response genes and activation of proapoptotic factors in pleural mesothelial cells. Lung 185(6):355–365. https://doi.org/10.1007/s00408-007-9046-6
Nguyen L, McCord KA, Bui DT, Bouwman KM, Kitova EN, Elaish M, Kumawat D, Daskhan GC, Tomris I, Han L, Chopra P (2022) Sialic acid-containing glycolipids mediate binding and viral entry of SARS-CoV-2. Nat Chem Biol 18(1):81–90. https://doi.org/10.1038/s41589-021-00924-1
Poole J, Day CJ, von Itzstein M, Paton JC, Jennings MP (2018) Glycointeractions in bacterial pathogenesis. Nat Rev Microbiol 16(7):440–452. https://doi.org/10.1038/s41579-018-0007-2
Priyambada SA, Misaki R, Okamoto T, Ohashi T, Ueda K, Matsuura Y, Fujiyama K (2018) St6gal1 knockdown alters HBV life cycle in HepAD38 cells. Biochem Biophys Res Commun 503(3):1841–1847. https://doi.org/10.1016/j.bbrc.2018.07.124
Ramos I, Bernal-Rubio D, Durham N, Belicha-Villanueva A, Lowen AC, Steel J, Fernandez-Sesma A (2011) Effects of receptor binding specificity of avian influenza virus on the human innate immune response. J Virol 85(9):4421–4431. https://doi.org/10.1128/JVI.02356-10
Reily C, Stewart TJ, Renfrow MB, Novak J (2019) Glycosylation in health and disease. Nat Rev Nephrol 15(6):346–366. https://doi.org/10.1038/s41581-019-0129-4
Ren W, Yin J, Duan J, Liu G, Zhu X, Chen S, Li T, Wang S, Tang Y, Hardwidge PR (2014) Mouse intestinal innate immune responses altered by enterotoxigenic Escherichia coli (ETEC) infection. Microbes Infect 16(11):954–961. https://doi.org/10.1016/j.micinf.2014.09.005
Rillahan CD, Antonopoulos A, Lefort CT, Sonon R, Azadi P, Ley K, Dell A, Haslam SM, Paulson JC (2012) Global metabolic inhibitors of sialyl- and fucosyltransferases remodel the glycome. Nat Chem Biol 8(7):661–668. https://doi.org/10.1038/nchembio.999
Sakarya S, Göktürk C, Oztürk T, Ertugrul MB (2010) Sialic acid is required for nonspecific adherence of Salmonella enterica ssp. enterica serovar Typhi on Caco-2 cells. FEMS Immunol Med Microbiol 58(3):330–5. https://doi.org/10.1111/j.1574-695X.2010.00650.x
Sun X, Zhang S, Ren J, Udenigwe CC (2022) Sialic acid-based strategies for the prevention and treatment of Helicobacter pylori infection: emerging trends in food industry. Crit Rev Food Sci Nutr 62(7):1713–1724. https://doi.org/10.1080/10408398.2020.1846157
Swindall AF, Londoño-Joshi AI, Schultz MJ, Fineberg N, Buchsbaum DJ, Bellis SL (2013) ST6Gal-I protein expression is upregulated in human epithelial tumors and correlates with stem cell markers in normal tissues and colon cancer cell lines. Cancer Res 73(7):2368–2378. https://doi.org/10.1158/0008-5472.Can-12-3424
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76(9):4350–4354. https://doi.org/10.1073/pnas.76.9.4350
Varki A (2017) Biological roles of glycans. Glycobiology 27(1):3–49. https://doi.org/10.1093/glycob/cww086
Wang A, Al-Kuhlani M, Johnston SC, Ojcius DM, Chou J, Dean D (2013) Transcription factor complex AP-1 mediates inflammation initiated by Chlamydia pneumoniae infection. Cell Microbiol 15(5):779–794. https://doi.org/10.1111/cmi.12071
Wei A, Fan B, Zhao Y, Zhang H, Wang L, Yu X, Yuan Q, Yang D, Wang S (2016) ST6Gal-I overexpression facilitates prostate cancer progression via the PI3K/Akt/GSK-3beta/beta-catenin signaling pathway. Oncotarget 7(40):65374–65388. https://doi.org/10.18632/oncotarget.11699
Wu D, Huang W, Wang Y, Guan W, Li R, Yang Z, Zhong N (2014) Gene silencing of β-galactosamide α-2,6-sialyltransferase 1 inhibits human influenza virus infection of airway epithelial cells. BMC Microbiol 14:78. https://doi.org/10.1186/1471-2180-14-78
Zhao Y, Li Y, Ma H, Dong W, Zhou H, Song X, Zhang J, Jia L (2014) Modification of sialylation mediates the invasive properties and chemosensitivity of human hepatocellular carcinoma. Mol Cell Proteomics 13(2):520–536. https://doi.org/10.1074/mcp.M113.034025
Funding
Student research fellowship was sanctioned by the Indian Council of Medical Research, India. This work was financially supported by the Department of Biotechnology (India BT/PR21686/MED/29/1142/ 2016).
Author information
Authors and Affiliations
Contributions
Dr. SG contributed to designing the research work and organizing the final draft of the manuscript. Dr. SC conducted experiments, analyzed, and interpreted the data as well as wrote the manuscript. AJ assisted in carrying out some experiments. SK helped significantly in carrying out qRT-PCR experiments. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chandel, S., Joon, A., Kaur, S. et al. Role of ST6GAL1 and ST6GAL2 in subversion of cellular signaling during enteroaggregative Escherichia coli infection of human intestinal epithelial cell lines. Appl Microbiol Biotechnol 107, 1405–1420 (2023). https://doi.org/10.1007/s00253-022-12321-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12321-2