Abstract
Oleaginous yeasts have received significant attention due to their substantial lipid storage capability. The accumulated lipids can be utilized directly or processed into various bioproducts and biofuels. Lipomyces starkeyi is an oleaginous yeast capable of using multiple plant-based sugars, such as glucose, xylose, and cellobiose. It is, however, a relatively unexplored yeast due to limited knowledge about its physiology. In this study, we have evaluated the growth of L. starkeyi on different sugars and performed transcriptomic and metabolomic analyses to understand the underlying mechanisms of sugar metabolism. Principal component analysis showed clear differences resulting from growth on different sugars. We have further reported various metabolic pathways activated during growth on these sugars. We also observed non-specific regulation in L. starkeyi and have updated the gene annotations for the NRRL Y-11557 strain. This analysis provides a foundation for understanding the metabolism of these plant-based sugars and potentially valuable information to guide the metabolic engineering of L. starkeyi to produce bioproducts and biofuels.
Key points
• L. starkeyi metabolism reprograms for consumption of different plant-based sugars.
• Non-specific regulation was observed during growth on cellobiose.
• L. starkeyi secretes β-glucosidases for extracellular hydrolysis of cellobiose.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oleaginous yeasts have been studied extensively due to their substantial lipid storage capability (Ageitos et al. 2011; Shi and Zhao 2017). The accumulated lipids provide alternatives to plant oils for biodiesel production (Spagnuolo et al. 2019). Lipomyces starkeyi is a promising oleaginous yeast, which was isolated from soil by Starkey (1946) and later described by Lodder and Kreger-Van Rij (1952). It can grow on various plant-based sugars and is an excellent lipid producer (McNeil and Stuart 2018a). Because of its ability to grow on a wide variety of sugars, many enzymes like dextranases, amylase, levoglucosan kinase, and other hydrolases have been studied and characterized from L. starkeyi (Bao et al. 2019; Kang et al. 2004, 2009; Nishimura et al. 2006; Rother et al. 2018; Ryu et al. 2000).
Nutrient limitation in L. starkeyi has been reported as a mechanism to induce lipid production (Calvey et al. 2016; Juanssilfero et al. 2018). Recent work has also explored the pathways involved in lipid metabolism (Kamineni and Shaw 2020; Takaku et al. 2020a). In addition to lipids, L. starkeyi has been engineered for fatty alcohol production (McNeil and Stuart 2018b; Wang et al. 2016). Lipid production in L starkeyi has been optimized using two-stage fermentation for industrial use (Lin et al. 2011; Zhang et al. 2021, 2022). In addition to accumulating large amounts of lipids, L. starkeyi can naturally utilize the sugars present in lignocellulosic hydrolysates and is tolerant to the inhibitors present in these hydrolysates (Monteiro de Oliveira et al. 2021; Pomraning et al. 2019). However, there is a considerable lack in our understanding of L. starkeyi, primarily due to insufficient knowledge of its physiology and the lack of efficient genetic tools.
The genome sequence of L. starkeyi NRRL Y-11557 was reported in 2016, along with 15 other biotechnologically relevant yeasts (Riley et al. 2016). Since then, a few studies have used the gene models of L. starkeyi NRRL Y-11557 in different bioinformatics analyses. Theoretical lipid yields on various sugars were reported using a small-scale metabolic model of L. starkeyi NRRL Y-11557 (Zhou et al. 2021). A recent study reported the transcriptional changes in L. starkeyi NRRL Y-11558 resulting from growth on hydrolysate (Pomraning et al. 2019). In addition, two studies have used bioinformatics analysis to identify and characterize the sugar transporters present in L. starkeyi: LST1_120451 as a cellobiose transporter (de Ruijter et al. 2020) and LST1_205437 as a co-transporter for glucose and xylose (Kuanyshev et al. 2021). Another study constructed the regulatory network of triacylglycerol (TAG) biosynthesis in L. starkeyi using metabolic profiling under nitrogen limiting conditions (Aburatani et al. 2020). Multiple transformation protocols have been established for the genetic engineering of L. starkeyi (Calvey et al. 2014; Dai et al. 2017; Lin et al. 2017; Takaku et al. 2020b), and heterologous gene expression has been realized (Oguro et al. 2017; Wang et al. 2016), enabling L. starkeyi to serve as a promising oleaginous yeast for chemical and fuel production.
In this work, we have systematically evaluated the growth of L. starkeyi NRRL Y-11557 on the plant-based sugars: glucose, xylose, and cellobiose. We performed transcriptomic and metabolomic analyses to better understand the underlying mechanisms of sugar uptake by this yeast. This work reports the different pathways activated during growth on these sugars based on the genome-wide gene expression differences. In addition, it also refines the gene annotation information of L. starkeyi NRRL Y-11557.
Materials and methods
Strains, media, and culture conditions
L. starkeyi NRRL Y-11557 (DSM 70,295; sourced from DSMZ—German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany) was grown on YPD medium (10 g/L yeast extract, 20 g/L peptone, and 20 g/L glucose) at 30 °C for routine culture. YPX medium (10 g/L yeast extract, 20 g/L peptone, and 20 g/L xylose), YPC medium (10 g/L yeast extract, 20 g/L peptone, and 20 g/L cellobiose), and YPD medium were used for transcriptomic and metabolomic analysis.
HPLC analysis of extracellular metabolites
Yeast growth (OD600) was measured using a spectrophotometer (Biomate 5, Thermo Fischer, Waltham, MA, USA). Extracellular metabolites such as glucose, xylose, cellobiose, glycerol, acetate, and ethanol were measured by HPLC (Agilent Technologies 1200 Series, Santa Clara, CA, USA) with a RezexTMROA-Organic Acid H + (8%) column (Phenomenex Inc., Torrance, CA, USA) and a refractive index detector (RID). The column was eluted with 0.005 N H2SO4 at a 0.6 mL/min flow rate at 50 °C.
RNA sequencing analysis
A single colony from a YPD agar plate was inoculated into YPD liquid medium to obtain L. starkeyi seed cultures. Seed cultures were then used to inoculate 50 mL YPD, YPX, and YPC medium in 250 mL baffled shake flasks, with a starting OD600 of 1. The cells were then grown at 250 rpm, 30 °C. The growth was monitored by measuring OD600 and sugar concentration. The samples for transcriptomics analysis were withdrawn at 30 h for YPD, YPX, and 52 h for YPC, as shown in Fig. 1. The samples were collected and washed using cold ddH2O. Total RNA was extracted using RNeasy Mini Kit (Qiagen, Hilden, Germany) and then treated with DNA-free DNase using the TURBO DNA-free kit (Ambion, Austin, TX, USA) to remove genomic DNA. The quality of RNA was confirmed by agarose gel electrophoresis and bioanalyser, DNA gel and Nanodrop. The stranded RNAseq libraries were prepared with Illumina’s TruSeq Stranded RNA Sample Prep kit. The libraries were pooled in equimolar concentration and sequenced for 101 cycles from each single end of the fragments on a HiSeq2500 (Illumina, San Diego, CA, USA). Fastq files were generated and demultiplexed with the bcl2fastq v1.8.4 Conversion Software (Illumina, San Diego, CA, USA). Raw sequencing reads are available at NCBI (BioProject ID: PRJNA808049).
RNA sequencing resulted in 9 samples with 13–19 million reads. The sequencing analysis was conducted using an in-house pipeline, as previously described (Chong et al. 2019; Jagtap et al. 2021). Adaptor sequences and low-quality reads were trimmed and analyzed using Trimmomatic (Bolger et al. 2014) and FastQC (Andrews et al. 2012). Reads were mapped to the L. starkeyi NRRL Y-11557 reference genome (NCBI GenBank assembly accession: GCA_001661325.1) (Riley et al. 2016) with STAR version 2.5.3a (Dobin et al. 2013). Read counts were calculated using featureCounts from the Subread package, version 1.5.2 (Liao et al. 2014). Differential expression analysis was performed on the reads counts in R v4.0.5 (R Core Team, 2021) using edgeR v3.32.1 and limma v3.46.0 (Ritchie et al. 2015; Robinson et al. 2010). Graphical representation of expression data was constructed using R packages: PCAtools v2.2.0 (Blighe and Lun 2021), gplots v3.1.1 (Warnes et al. 2011), and Glimma v2.0.0 (Su et al. 2017). Functional annotation of L. starkeyi was obtained from DOE Joint Genome Institute website Mycocosm (Grigoriev et al. 2014). Data analysis scripts, along with the results, can be downloaded from the following link: https://github.com/raogroupuiuc/lipo11557_growth.
Pairwise enrichment analysis was performed using GSEA (Subramanian et al. 2005). Briefly, the KEGG and GO annotation files downloaded from JGI Mycocosm (Grigoriev et al. 2014) were reformatted to the GSEA guidelines using Python. All parameters were set to default in GSEA, except “Permutation Type” was set to “gene_set.” Enrichment analysis was performed separately for different annotation classes (GO: biological process, molecular function, and cellular component and KEGG: pathways and pathway class) and visualized using Cytoscape v3.8.2 (Shannon et al. 2003). DeepLoc-1.0 was used to predict the subcellular localization of proteins, reported in Supplementary Dataset S1 (Almagro Armenteros et al. 2017).
Metabolomics analysis
Yeast cells grown in YPD, YPX, and YPC were collected during exponential phase and vacuum filtered using Vac-Man Laboratory Vacuum Manifold (Promega, Madison, WI, USA) assembled with a nylon membrane filter (pore size, 0.45 μm; diameter, 13 mm; Whatman, Piscataway, NJ, USA) and a filter holder (Millipore, Billerica, MA, USA). The filtered cell culture was washed with 2.5 mL of prechilled distilled water. The entire process of fast filtration was completed within 1 min as previously described (Jagtap et al. 2021; Yun et al. 2018). The filter membrane containing the washed cells was quickly mixed with 1 mL prechilled acetonitrile–water mixture (1:1, v/v) and 100 µL of glass beads. The mixture was vortexed for 3 min to disrupt cell membranes, allowing extraction of intracellular metabolites. The extraction mixture was then centrifuged at 16,100 × g for 3 min at 4 °C, and 0.8 mL of the supernatant containing the intracellular metabolites was dried in a speed vacuum concentrator for 6 h.
Before GC/MS analysis, the samples were derivatized by methoxyamination and trimethylsilylation, as previously described (Jagtap et al. 2019; Liu et al. 2021). For GC/MS, the derivatized metabolite samples were applied to an Agilent 7890A GC/5975C MSD system (Agilent Technologies, Santa Clara, CA, USA) equipped with an RTX-5Sil MS capillary column (30 m × 0.25 mm, 0.25 µm film thickness: Restek, Bellefonte, PA, USA) and an additional 10-m-long integrated guard column. One microliter of the derivatized sample was injected into the GC inlet in splitless mode. The oven temperature was initially set to 150 °C for 1 min, after which the temperature was increased to 330 °C at 20 °C/min, where it was held for 5 min. The mass spectra were recorded in a scan range 85–500 m/z at an electron impact of 70 eV, and the temperatures of the ion source and transfer line were 230 and 280 °C, respectively.
The raw data obtained from the GC–MS analysis were processed using an automated mass spectral deconvolution and identification system (AMDIS) software for peak detection and deconvolution of mass spectra. The processed data were uploaded to SpectConnect (http://spectconnect.mit.edu) for peak alignment and generation of the data matrix with the Golm Metabolome Database mass spectral reference library (Kopka et al. 2005). The normalized abundance values for each metabolite were obtained by dividing peak intensity with dry cell weight. For statistical analysis, such as principal component analysis (PCA) and clustering analysis (represented as a heatmap), Statistica (version 7.1; StatSoft, Tulsa, OK, USA), MetaboAnalyst, and MultiExperiment Viewer software were used (Chong et al. 2019; Howe et al. 2011). In addition, GraphPad Prism 6 (GraphPad, San Diego, CA, USA) was used for plotting bar graphs.
Results
Growth and utilization of different substrates
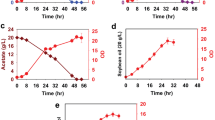
The hydrolysis of lignocellulosic biomass releases monosaccharide and disaccharide sugars, principally in the form of glucose, xylose, and cellobiose (Xu and Li 2017). L. starkeyi is an oleaginous yeast capable of utilizing these lignocellulosic sugars. In this work, we test the growth of L. starkeyi NRRL Y-11557 on glucose, xylose, and cellobiose. L. starkeyi was grown on yeast extract and peptone (YP) medium containing one of the following sugars: glucose (YPD), xylose (YPX), and cellobiose (YPC). Growth in YP medium leads to higher growth and cell mass. It also, however, adds addition proteins and complex sugars to the media. The results presented here represent co-utilization of proteins from YP media and the supplemented sugars (Nancib et al. 1991). Figure 1 shows the growth and substrate utilization of L. starkeyi on the different sugars. Growth on the three sugars led to similar final cell densities. However, we observed differences in sugar utilization. The cells ultimately utilized 20 g/L glucose and xylose in 48 h, whereas consumption of 20 g/L cellobiose was slower and took 84 h of growth. L. starkeyi consumed glucose marginally faster than xylose, which is evident from the time point for 50% sugar utilization. The cells consumed 10 g/L of glucose, xylose, and cellobiose in 28 h, 32 h, and 56 h, respectively.
Identification of genes associated with substrate utilization
We used whole-genome RNA sequencing to analyze the growth of L. starkeyi on the three sugars. RNA was extracted approximately at the time point for 50% sugar utilization, with 3 biological replicates for each condition (30 h for glucose and xylose, and 52 h for cellobiose). We chose the mid-point of sugar utilization in each case to capture the gene expression profiles for sugar uptake and metabolism. The sequencing resulted in an average of 16.3 million reads across 9 samples. More than 85% of reads were mapped to a unique location in the reference genome (NCBI GenBank assembly accession: GCA_001661325.1) (Supplementary Fig. S1) (Riley et al. 2016). The different clusters on the principal component analysis (PCA) plot show the distinct gene expression profiles of the three sugars (Fig. 2a). PC1 axis, representing 70% variation, highlights the difference between growth on glucose and cellobiose, whereas PC2 axis, which represents 22% variation, distinguished the growth on xylose from the other two sugars.
Overview of the transcriptomics analysis. a Principal component analysis plot generated from gene expression profiles of L. starkeyi grown on glucose, xylose, and cellobiose. RNAseq data were collected in triplicates for each condition. KEGG pathway enrichment analysis for growth on cellobiose vs. glucose (b) and xylose vs. glucose (c). The vertical axis represents the pathway category, and the horizontal axis represents the pathway’s enrichment score [− log(p value)]. Significantly enriched KEGG pathways (p < 0.05 and FDR < 0.25) are plotted. The data were analyzed using the GSEA tool and plotted in Origin (OriginLab Corporation, Northampton, MA, USA)
We chose glucose as a control to compare the differences in gene expression profiles of different sugars for further analysis. During the growth on cellobiose compared to glucose, expression of 851 genes was significantly increased, and expression of another 374 genes was significantly decreased (Supplementary Fig. S2a). During growth on xylose compared to glucose, expression of 392 genes was significantly increased and 120 genes was significantly decreased (Supplementary Fig. S2b). Fold change > 2 and adjusted p value < 0.05 were considered significant for comparative gene expression analysis. These numbers are in line with the clustering trend on the PCA plot. A complete list of differential gene expression of all genes in L. starkeyi is provided in Supplementary Dataset S1.
To further investigate the differences in global gene expression profiles of these sugars, we performed gene set enrichment analysis (GSEA) using KEGG annotations. Differential gene expression during growth on cellobiose is limited to sugar uptake and metabolism, like starch metabolism, pentose and glucuronate interconversion pathway, and galactose metabolism (Fig. 2b). On the other hand, growth on xylose resulted in broader changes in the central carbon metabolism, as noted in Fig. 2c. We note that while cellobiose has a more extensive set of differentially expressed genes than xylose (Supplementary Fig. S2), the list of enriched pathways is smaller than xylose (Fig. 2b, c). This is potentially because we have limited information about the gene annotation of those gene sets. Genes with limited annotation information but significant differential expression are listed in the “Other genes of interest” sheet in Supplementary Dataset S1.
Comparative analysis of growth on cellobiose
We first explored cellobiose metabolism in L. starkeyi by comparing the gene expression on YP medium containing cellobiose (YPC) versus YP medium containing glucose (YPD). Figure 3a highlights the cellobiose uptake pathways. Cellobiose can be transported into and utilized by the cell in two ways. The first is the import of cellobiose with the aid of a cellobiose transporter and further hydrolysis of intracellular cellobiose into glucose by β-glucosidase. A recent study identified LST1_120451 as a cellobiose transporter (CDT1) (de Ruijter et al. 2020). Expression of the CDT1 gene was increased 11-fold during growth on cellobiose versus glucose. L. starkeyi has multiple copies of genes encoding for β-glucosidase (BGL1-6), with the increased expression of five BGLs during growth on cellobiose as compared to glucose. Expression of glucokinase (GLK1), the enzyme that facilities the phosphorylation of glucose, was increased 2.3-fold. We also used DeepLoc (Almagro Armenteros et al. 2017), a localization prediction tool for eukaryotic genes, to predict the localization of BGL1-6. We note that BGL1 and BLG3 are extracellular, BGL5 and BGL6 are peroxisomal, whereas BGL2 and BGL4 are cytosolic. The increased expression of BGL3 was the highest amongst the six BGLs (1252-fold), which alludes to the second mechanism of cellobiose utilization. Cellobiose is hydrolyzed into glucose extracellularly, aided by secreted β-glucosidases, and the resultant glucose is transported into the cell using glucose transporters. Multiple membrane-localized genes, annotated as predicated transporters belonging to the major facilitator superfamily, exhibited increased expression on cellobiose. However, we lack experimental validation for these predicted transporters. We previously tested a few predicted transporters for glucose uptake and found that LST1_205437 facilitates both glucose and xylose uptake (Kuanyshev et al. 2021). Details on gene annotation, differential gene expression, and gene localization are provided in Supplementary Dataset S1.
Substrate utilization in L. starkeyi (a and b). Substrate uptake and metabolism for cellobiose (a) and xylose (b). Differential gene expression is reported in comparison to glucose. Upregulated genes (fold change > 2 and p value < 0.05) are highlighted in red and downregulated genes in blue. Heatmap of gene expression data in KEGG pathways: starch and sucrose metabolism (c) and pentose and glucuronate metabolism (d). RNAseq data were collected in triplicate for each condition. Color key represents the z-score for each gene (normalized for all growth conditions). e Intensity levels of intracellular metabolites profiled using GC/MS during growth on different sugars. The data represents the average of six independent measurements, and bars indicate standard deviation
We also observed non-specific regulation in L. starkeyi during growth on cellobiose. This is likely due to the cells utilizing sugars present in the yeast extract. The effect of yeast extract on gene expression may be most noticeable on cellobiose because it is a less preferred carbon source than the sugar monomers. Preferential sugar uptake has also been reported in the yeast Spathaspora passalidarum during fermentation of hydrolysate containing glucose, xylose, and cellobiose (Long et al. 2012). Expression of hydrolase genes, involved in the breakdown of other oligosaccharides, was also increased (Fig. 3a). Nine α-glucosidase genes (MAL), which are involved in the breakdown of maltose, and three α-amylase genes (AMY), which are responsible for the breakdown of maltodextrin, and five β-fructofuranosidase genes (SUC) exhibited increased expression. Significant increases in expression were observed for the genes belonging to the starch and sucrose metabolism pathway of the KEGG pathway database (Fig. 3c) (Kanehisa and Goto 2000).
We next focused on the genes involved in central carbon metabolism (Fig. 4). Cellobiose breaks down into glucose and enters glycolysis. Genes in glycolysis did not exhibit significant differences in expression, except for enolase (ENO1), whose expression decreased 2.2-fold, and fructose bisphosphatase (FBP1), whose expression increased 2.4-fold. We also noticed decreased expression of both the pyruvate decarboxylases (PDC1: 2.6-fold, PDC2: 3.4-fold) and a 4.9-fold increase in expression of pyruvate carboxylase (PYC2). However, the small change in the expression of these genes indicates the absence of any global shifts in regulation induced from growth on cellobiose. One interesting result was the increased expression of multiple aldehyde dehydrogenases (ALD1: 20-fold, ALD4: threefold, and ALD5: tenfold). We also observed decreased expression of acetate kinase (ackA) and acetyl-CoA synthetase (ACS1). It is plausible that L. starkeyi accumulates a smaller pool of acetyl-CoA during growth on cellobiose compared to glucose. This observation is in line with the decreased concentration of hexadecenoic acid during growth on cellobiose (discussed in detail in the last subsection of results).
Differential gene expression of the central metabolic pathway during growth on cellobiose, reported in comparison to glucose. Upregulated genes are highlighted in red, and downregulated genes in blue. Metabolites with significantly higher abundance are highlighted in green, and metabolites in lower abundance in purple. Gene and metabolite names and supporting quantitative data are included in Supplementary Datasets S1 and S2
In the tricarboxylic acid (TCA) cycle, we observed increased expression of one of the three citrate synthases (CIT3) by 7.9-fold and fumarate reductase (FRD1) by 14.3-fold. In addition, expression of both genes in the glyoxylate bypass, malate synthase (MLS1) and mitochondrial isocitrate lyase (ICL1), was increased 3.4-fold and sevenfold, respectively. Supplementary Dataset S1 lists the differential expression for all genes in central carbon metabolism.
Differential analysis of growth on xylose
We next analyzed xylose metabolism in L. starkeyi. Gene expression during growth on YP medium containing xylose (YPX) is compared to YPD. Overall, 392 genes were significantly upregulated, and 120 genes were significantly downregulated during growth on xylose compared to glucose. The growth on xylose had a closer gene expression profile to glucose, with key differences in carbohydrate metabolism (Fig. 2c).
A prominent pathway for xylose utilization in eukaryotes is the oxidoreductive pathway, also known as the XR-XDH pathway (Fig. 3b) (Jagtap and Rao 2018a; Lee et al. 2021). Xylose reductase (XR) reduces intracellular xylose to xylitol, which is further oxidized to xylulose by xylitol dehydrogenase (XDH) or sorbitol dehydrogenase (SOR1-6). The last step of the XR-XDH pathway is the phosphorylation of xylulose by xylulokinase (XKS), which enters the pentose phosphate pathway. In L. starkeyi, we found that expression of all genes in the XR-XDH pathway was increased. For instance, we noted significant increases in expression for XR and SOR2, which increased 30- and 302-fold, respectively. In addition, expression of XKS was 27-fold increased as well. Increased expression was also observed in the genes belonging to the pentose and glucuronate interconversion pathway of the KEGG pathway database (Kanehisa and Goto 2000) (Fig. 3d). Some oleaginous yeasts have an arabitol bypass (Jagtap and Rao 2018b), where the carbon flux from xylose is directed through arabinitol dehydrogenase (ARD1) and ribulokinase (RK) instead of the one-step XKS. Expression of ARD1 and RK was increased 2.4- and 5.4-fold, respectively. The extent of upregulation of these pathways in L. starkeyi indicates that the one-step XKS phosphorylation draws a higher carbon flux than the arabitol bypass.
Xylose uptake pathway converts xylose to xylulose-5P and ribulose-5P, which enter the pentose phosphate pathway (Fig. 5). Few genes in pentose phosphate pathways exhibited increased expression in L. starkeyi, which is consistent with the mechanism of xylose metabolism. Most notably, expression of transketolase (TKL2), transaldolase (TAL1,2), and ribokinase (RBK1) was increased. We also observed increased expression of a few genes in upper glycolysis, namely, glucose-6-phosphate isomerase (PGI1) and fructose bisphosphate (FBP1). Genes involved in glyoxylate bypass also exhibited increased expression during growth on xylose, which was also the case with cellobiose growth. We observed a few more similarities in the gene expression on xylose and cellobiose. For instance, expression of both fumarate reductase and citrate synthase was increased.
Differential gene expression of the central metabolic pathway during growth on xylose, reported in comparison to glucose. Upregulated genes are highlighted in red, and downregulated genes in blue. Metabolites with significantly higher abundance are highlighted in green, and metabolites with lower abundance in purple. Gene and metabolite names and supporting quantitative data are included in Supplementary Datasets S1 and S2
Measurement of intracellular metabolites
To supplement the RNA sequencing data, we used gas chromatography-mass spectrometry (GCMS) and measured changes in the concentration of intracellular metabolites. Of the 55 metabolites measured, significant differences were observed in the concentrations of 37 metabolites during growth on glucose, xylose, and cellobiose (Fig. 6). Significance was evaluated using two-sample t-tests, and the cut-off was relative metabolite concentration > 2 and adjusted p value < 0.05. Compared to glucose, the concentration of 9 and 3 metabolites was higher on cellobiose and xylose, respectively. Whereas another 16 and 4 metabolites had a lower concentration in cellobiose and xylose, respectively, the changes in the metabolite concentrations align with those of differential gene expression. Metabolite concentrations are reported in Supplementary Dataset S2.
Overview of the metabolomics analysis. Heatmap of intracellular metabolites from xylose and cellobiose uptake pathways (a), glycolysis (b), pentose phosphate pathway (c), TCA cycle (d), and lipid biosynthesis pathway (e). All experiments were performed with six replicates. Color key represents the z-score for each metabolite (normalized across all 18 samples)
During growth on cellobiose, we observed a 345-fold increase in the concentration of intracellular cellobiose, which is consistent with the upregulation of a cellodextrin transporter (CDT1). Again, in alignment with the gene regulation, metabolite concentrations in central carbon metabolism exhibit mixed changes, with valine and allantoin present in higher concentrations during growth on cellobiose and others like fumarate, malate, and α-ketoglutarate are present in lower concentrations. An interesting result is decreased concentration of hexadecenoic acid, indicating a potentially lower intracellular pool of acetyl-CoA. We previously highlighted that the gene expression results also point to reduced production of acetyl-CoA. While this study did not focus on lipid production in L. starkeyi, it has been previously reported that L. starkeyi produces slightly lower lipid titers in flask cultures when grown on cellobiose, compared to glucose (Gong et al. 2012).
The metabolite concentrations during growth on xylose are in line with the gene expression results. Metabolites in the xylose utilization and pentose phosphate pathways are in higher concentration (Fig. 6). We notice a 29-fold higher concentration of intracellular xylose. Only a few metabolites are present in lower concentrations in glycolysis and TCA cycle, indicating little difference in the regulation of central carbon metabolism between growth on xylose and glucose.
Discussion
L. starkeyi is a promising oleaginous yeast that can utilize a variety of plant-based substrates. It can grow on various hydrolysates derived from wheat straw and corn stover and exhibits tolerance to the aromatic inhibitors present in hydrolysates (Brandenburg et al. 2021; Pomraning et al. 2019). It is also an excellent lipid producer and can store triacylglycerides up to 70% of its dry cell weight. While various studies have optimized lipid accumulation using different hydrolysates, a few focus on governing metabolic pathways and physiology.
In this work, we analyze the transcriptome and metabolome of L. starkeyi during growth on three plant-based sugars in nutrient-rich YP medium: glucose, xylose, and cellobiose. Cell culture samples for RNA and metabolite extraction were collected at the mid-point of sugar utilization (Fig. 1). It has been previously reported that L. starkeyi does not enter the oleaginous phase in nitrogen-rich conditions (Pomraning et al. 2019). We observed significant changes in the gene expression and metabolite concentration resulting from the utilization of different sugars (Figs. 2 and 6) and mapped their gene regulation to relevant metabolic pathways (Figs. 3, 4, and 5).
The bulk of the differential gene expression analysis focuses on central carbon metabolism. Of the 8192 genes in L. starkeyi, 10% are upregulated and 5% are downregulated during growth on cellobiose compared to glucose. On the other hand, growth on xylose resulted in comparatively lesser regulation: 5% of genes are upregulated, and another 1.5% downregulated (Supplementary Fig. S2). However, we observed a reverse trend from the enrichment analysis (Fig. 2b, c), where more pathways were enriched during growth on xylose, despite lesser differential genes, compared to cellobiose. We suspect this is because the gene expression changes in cellobiose are global. Since the gene annotation information is limited, our analysis has elaborated on the differences mainly limited to the central carbon metabolism.
We also note that some genes in the L. starkeyi genome were incorrectly annotated. For instance, we found 86 genes annotated incorrectly as L-arabinose isomerase in the KEGG annotations. Growth of L. starkeyi on L-arabinose was tested (Supplementary Fig. S3). L. starkeyi can utilize 5 g/L L-arabinose in 5 days of growth in YP media, which is much slower than glucose and xylose utilization (20 g/L in 2 days). Using bioinformatics tools such as BLAST and DeepLoc, we note that all 86 genes are predicted membrane proteins, some bound to the cell membrane (Supplementary Dataset S1 Sheet 2). A recent study found that the gene LST1_120451, marked as L-arabinose isomerase in the annotation, is a functional cellobiose transporter (de Ruijter et al. 2020), and another gene, LST1_205437 is a co-transporter for glucose and xylose (Kuanyshev et al. 2021). The prediction models have annotated most of these 86 genes as transporter proteins.
In an attempt to improve the gene annotations of the L. starkeyi genome, we have manually verified the annotations of 190 genes, belonging to the sugar uptake pathways and central carbon metabolism (Supplementary Dataset S1 Sheet 1 – Key genes). Sheets 2 and 3 have gene function and localization information of genes that were annotated incorrectly by the annotation pipelines. Differential expression and expression values for all genes in L. starkeyi genes are present in sheet 6 of Supplementary Dataset S1. Supplementary Dataset S3 contains a list of orthologous genes between S. cerevisiae and L. starkeyi, generated using the orthology tool, InParanoid (O'Brien et al. 2005).
During growth on cellobiose, we saw upregulation of both cytosolic and secreted β-glucosidases, along with upregulation of a cellobiose transporter, CDT1 (Fig. 3a). This is in line with cellobiose utilization in other fungi like Neurospora crassa (Ha et al. 2011). We also note non-specific regulation during growth on cellobiose. Genes involved in the hydrolysis of other oligosaccharides and polysaccharides, like maltose, sucrose, maltodextrin, and β-glucan, are also upregulated. Uptake of xylose in L. starkeyi is also in line with other xylose assimilating yeasts like Scheffersomyces stipitis (Kwak et al. 2019). Xylose is converted to xylulose through the oxidoreductase pathway (XR, XDH) and channeled into the pentose phosphate pathway primarily through xylulokinase (XKS). All three genes, XR, XDH, and XKS, exhibited increased expression during growth on xylose. Pentose phosphate pathway and upper glycolysis are also partially upregulated. We also observed increased expression of the genes in the glyoxylate bypass (MLS1 and ICL1), phosphoenolpyruvate carboxylase (PCK1), and fructose bisphosphate (FBP1) during growth on both xylose and cellobiose, indicating the upregulation of gluconeogenesis in these growth conditions. In line with these results, we also report the increased expression of genes in the biosynthesis of the secondary metabolites like glycogen synthase (GSY1), glycogenin glucosyltransferase (GLG1), and myo-inositol dehydrogenase (idhA).
These results improve our understanding of glucose, xylose, and cellobiose assimilation by L. starkeyi NRRL Y-11557 and provide a global overview of gene expression during growth on these plant-based sugars. The associated data sets from the transcriptomics and bioinformatics analysis present potentially valuable information to help guide the metabolic engineering of L. starkeyi and assist in developing regulatory networks and genome-scale models.
Data availability
RNA sequencing reads are uploaded to a NCBI bioproject (Accession number: PRJNA808049). Additional datasets are uploaded at the research group GitHub page: https://github.com/raogroupuiuc/lipo11557_growth. Other datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Aburatani S, Ishiya K, Itoh T, Hayashi T, Taniguchi T, Takaku H (2020) Inference of regulatory system for TAG biosynthesis in Lipomyces starkeyi. Bioeng (Basel) 7:4. https://doi.org/10.3390/bioengineering7040148
Ageitos JM, Vallejo JA, Veiga-Crespo P, Villa TG (2011) Oily yeasts as oleaginous cell factories. Appl Microbiol Biotechnol 90(4):1219–1227. https://doi.org/10.1007/s00253-011-3200-z
Almagro Armenteros JJ, Sonderby CK, Sonderby SK, Nielsen H, Winther O (2017) DeepLoc: prediction of protein subcellular localization using deep learning. Bioinformatics 33(21):3387–3395. https://doi.org/10.1093/bioinformatics/btx431
Andrews S, Krueger F, Segonds-Pichon A, Biggins L, Krueger C, Wingett S (2012) FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics, Babraham Institute, Cambridge, United Kingdom. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Blighe K, Lun A (2021) PCAtools: everything principal components analysis. R package version 2.6.0. https://github.com/kevinblighe/PCAtools
Bao M, Niu C, Xu X, Zheng F, Liu C, Wang J, Li Q (2019) Identification, soluble expression, and characterization of a novel endo-inulinase from Lipomyces starkeyi NRRL Y-11557. Int J Biol Macromol 137:537–544. https://doi.org/10.1016/j.ijbiomac.2019.06.096
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Brandenburg J, Blomqvist J, Shapaval V, Kohler A, Sampels S, Sandgren M, Passoth V (2021) Oleaginous yeasts respond differently to carbon sources present in lignocellulose hydrolysate. Biotechnol Biofuels 14(1):124. https://doi.org/10.1186/s13068-021-01974-2
Calvey CH, Willis LB, Jeffries TW (2014) An optimized transformation protocol for Lipomyces starkeyi. Curr Genet 60(3):223–230. https://doi.org/10.1007/s00294-014-0427-0
Calvey CH, Su YK, Willis LB, McGee M, Jeffries TW (2016) Nitrogen limitation, oxygen limitation, and lipid accumulation in Lipomyces starkeyi. Bioresour Technol 200:780–788. https://doi.org/10.1016/j.biortech.2015.10.104
Chong J, Wishart DS, Xia J (2019) Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr Protoc Bioinformatics 68(1):e86. https://doi.org/10.1002/cpbi.86
Dai Z, Deng S, Culley DE, Bruno KS, Magnuson JK (2017) Agrobacterium tumefaciens-mediated transformation of oleaginous yeast Lipomyces species. Appl Microbiol Biotechnol 101(15):6099–6110. https://doi.org/10.1007/s00253-017-8357-7
de Ruijter JC, Igarashi K, Penttila M (2020) The Lipomyces starkeyi gene Ls120451 encodes a cellobiose transporter that enables cellobiose fermentation in Saccharomyces cerevisiae. FEMS Yeast Res 20:3. https://doi.org/10.1093/femsyr/foaa019
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21. https://doi.org/10.1093/bioinformatics/bts635
Gong Z, Wang Q, Shen H, Hu C, Jin G, Zhao ZK (2012) Co-fermentation of cellobiose and xylose by Lipomyces starkeyi for lipid production. Bioresour Technol 117:20–24. https://doi.org/10.1016/j.biortech.2012.04.063
Grigoriev IV, Nikitin R, Haridas S, Kuo A, Ohm R, Otillar R, Riley R, Salamov A, Zhao X, Korzeniewski F, Smirnova T, Nordberg H, Dubchak I, Shabalov I (2014) MycoCosm portal: gearing up for 1000 fungal genomes. Nucleic Acids Res 42(Database issue):699–704. https://doi.org/10.1093/nar/gkt1183
Ha SJ, Galazka JM, Kim SR, Choi JH, Yang X, Seo JH, Glass NL, Cate JH, Jin YS (2011) Engineered Saccharomyces cerevisiae capable of simultaneous cellobiose and xylose fermentation. Proc Natl Acad Sci U S A 108(2):504–509. https://doi.org/10.1073/pnas.1010456108
Howe E, Sinha R, Schlauch D, Quackenbush J (2011) RNA-Seq analysis in MeV. Bioinformatics 27(22):3209–3210. https://doi.org/10.1093/bioinformatics/btr490
Jagtap SS, Rao CV (2018a) Microbial conversion of xylose into useful bioproducts. Appl Microbiol Biotechnol 102(21):9015–9036. https://doi.org/10.1007/s00253-018-9294-9
Jagtap SS, Rao CV (2018b) Production of D-arabitol from D-xylose by the oleaginous yeast Rhodosporidium toruloides IFO0880. Appl Microbiol Biotechnol 102(1):143–151. https://doi.org/10.1007/s00253-017-8581-1
Jagtap SS, Bedekar AA, Liu JJ, Jin YS, Rao CV (2019) Production of galactitol from galactose by the oleaginous yeast Rhodosporidium toruloides IFO0880. Biotechnol Biofuels 12:250. https://doi.org/10.1186/s13068-019-1586-5
Jagtap SS, Deewan A, Liu JJ, Walukiewicz HE, Yun EJ, Jin YS, Rao CV (2021) Integrating transcriptomic and metabolomic analysis of the oleaginous yeast Rhodosporidium toruloides IFO0880 during growth under different carbon sources. Appl Microbiol Biotechnol 105(19):7411–7425. https://doi.org/10.1007/s00253-021-11549-8
Juanssilfero AB, Kahar P, Amza RL, Miyamoto N, Otsuka H, Matsumoto H, Kihira C, Thontowi A, Yopi OC, Prasetya B, Kondo A (2018) Effect of inoculum size on single-cell oil production from glucose and xylose using oleaginous yeast Lipomyces starkeyi. J Biosci Bioeng 125(6):695–702. https://doi.org/10.1016/j.jbiosc.2017.12.020
Kamineni A, Shaw J (2020) Engineering triacylglycerol production from sugars in oleaginous yeasts. Curr Opin Biotechnol 62:239–247. https://doi.org/10.1016/j.copbio.2019.12.022
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Kang HK, Lee JH, Kim D, Day DF, Robyt JF, Park KH, Moon TW (2004) Cloning and expression of Lipomyces starkeyi alpha-amylase in Escherichia coli and determination of some of its properties. FEMS Microbiol Lett 233(1):53–64. https://doi.org/10.1016/j.femsle.2004.01.036
Kang HK, Park JY, Ahn JS, Kim SH, Kim D (2009) Cloning of a gene encoding dextranase from Lipomyces starkeyi and its expression in Pichia pastoris. J Microbiol Biotechnol 19(2):172–177. https://doi.org/10.4014/jmb.0802.100
Kopka J, Schauer N, Krueger S, Birkemeyer C, Usadel B, Bergmüller E, Dormann P, Weckwerth W, Gibon Y, Stitt M, Willmitzer L, Fernie AR, Steinhauser D (2005) GMD@CSB.DB the Golm metabolome database. Bioinformatics 21(8):1635–8. https://doi.org/10.1093/bioinformatics/bti236
Kuanyshev N, Deewan A, Jagtap SS, Liu J, Selvam B, Chen LQ, Shukla D, Rao CV, Jin YS (2021) Identification and analysis of sugar transporters capable of co-transporting glucose and xylose simultaneously. Biotechnol J 16(11):e2100238. https://doi.org/10.1002/biot.202100238
Kwak S, Jo JH, Yun EJ, Jin YS, Seo JH (2019) Production of biofuels and chemicals from xylose using native and engineered yeast strains. Biotechnol Adv 37(2):271–283. https://doi.org/10.1016/j.biotechadv.2018.12.003
Lee JW, Yook S, Koh H, Rao CV, Jin YS (2021) Engineering xylose metabolism in yeasts to produce biofuels and chemicals. Curr Opin Biotechnol 67:15–25. https://doi.org/10.1016/j.copbio.2020.10.012
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Lin J, Shen H, Tan H, Zhao X, Wu S, Hu C, Zhao ZK (2011) Lipid production by Lipomyces starkeyi cells in glucose solution without auxiliary nutrients. J Biotechnol 152(4):184–188. https://doi.org/10.1016/j.jbiotec.2011.02.010
Lin X, Liu S, Bao R, Gao N, Zhang S, Zhu R, Zhao ZK (2017) Development of an Agrobacterium-mediated transformation method and evaluation of two exogenous constitutive promoters in oleaginous yeast Lipomyces starkeyi. Appl Biochem Biotechnol 183(3):867–875. https://doi.org/10.1007/s12010-017-2469-5
Liu JJ, Woodruff W, Deewan A, Jagtap SS, Yun EJ, Walukiewicz HE, Jin YS, Rao CV (2021) Investigating the role of the transcriptional regulator Ure2 on the metabolism of Saccharomyces cerevisiae: a multi-omics approach. Appl Microbiol Biotechnol 105(12):5103–5112. https://doi.org/10.1007/s00253-021-11394-9
Lodder J, Kreger-van Rij NJW (1952) Genus Lipomyces. In: Lodder J, Kreger-van Rij NJW (eds) The yeasts, a taxonomic study. North-Holland Publishing Company (Elsevier), Amsterdam, pp 669–700
Long TM, Su YK, Headman J, Higbee A, Willis LB, Jeffries TW (2012) Cofermentation of glucose, xylose, and cellobiose by the beetle-associated yeast Spathaspora passalidarum. Appl Environ Microbiol 78(16):5492–5500. https://doi.org/10.1128/AEM.00374-12
McNeil BA, Stuart DT (2018a) Lipomyces starkeyi: an emerging cell factory for production of lipids, oleochemicals and biotechnology applications. World J Microbiol Biotechnol 34(10):147. https://doi.org/10.1007/s11274-018-2532-6
McNeil BA, Stuart DT (2018b) Optimization of C16 and C18 fatty alcohol production by an engineered strain of Lipomyces starkeyi. J Ind Microbiol Biotechnol 45(1):1–14. https://doi.org/10.1007/s10295-017-1985-1
Monteiro de Oliveira P, Aborneva D, Bonturi N, Lahtvee PJ (2021) Screening and growth characterization of non-conventional yeasts in a hemicellulosic hydrolysate. Front Bioeng Biotechnol 9:659472. https://doi.org/10.3389/fbioe.2021.659472
Nancib N, Branlant C, Boudrant J (1991) Metabolic roles of peptone and yeast extract for the culture of a recombinant strain of Escherichia coli. J Ind Microbiol 8(3):165–169. https://doi.org/10.1007/BF01575849
Nishimura K, Harada T, Arita Y, Watanabe H, Iwabuki H, Terada A, Naganuma T, Uzuka Y (2006) Identification of enzyme responsible for erythritol utilization and reaction product in yeast Lipomyces starkeyi. J Biosci Bioeng 101(4):303–308. https://doi.org/10.1263/jbb.101.303
O’Brien KP, Remm M, Sonnhammer EL (2005) Inparanoid: a comprehensive database of eukaryotic orthologs. Nucleic Acids Res 33(Database issue):D476-80. https://doi.org/10.1093/nar/gki107
Oguro Y, Yamazaki H, Ara S, Shida Y, Ogasawara W, Takagi M, Takaku H (2017) Efficient gene targeting in non-homologous end-joining-deficient Lipomyces starkeyi strains. Curr Genet 63(4):751–763. https://doi.org/10.1007/s00294-017-0679-6
Pomraning KR, Collett JR, Kim J, Panisko EA, Culley DE, Dai Z, Deng S, Hofstad BA, Butcher MG, Magnuson JK (2019) Transcriptomic analysis of the oleaginous yeast Lipomyces starkeyi during lipid accumulation on enzymatically treated corn stover hydrolysate. Biotechnol Biofuels 12:162. https://doi.org/10.1186/s13068-019-1510-z
Riley R, Haridas S, Wolfe KH, Lopes MR, Hittinger CT, Goker M, Salamov AA, Wisecaver JH, Long TM, Calvey CH, Aerts AL, Barry KW, Choi C, Clum A, Coughlan AY, Deshpande S, Douglass AP, Hanson SJ, Klenk HP, LaButti KM, Lapidus A, Lindquist EA, Lipzen AM, Meier-Kolthoff JP, Ohm RA, Otillar RP, Pangilinan JL, Peng Y, Rokas A, Rosa CA, Scheuner C, Sibirny AA, Slot JC, Stielow JB, Sun H, Kurtzman CP, Blackwell M, Grigoriev IV, Jeffries TW (2016) Comparative genomics of biotechnologically important yeasts. Proc Natl Acad Sci U S A 113(35):9882–9887. https://doi.org/10.1073/pnas.1603941113
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47. https://doi.org/10.1093/nar/gkv007
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Rother C, Gutmann A, Gudiminchi R, Weber H, Lepak A, Nidetzky B (2018) Biochemical characterization and mechanistic analysis of the levoglucosan kinase from Lipomyces starkeyi. ChemBioChem 19(6):596–603. https://doi.org/10.1002/cbic.201700587
Ryu SJ, Kim D, Ryu HJ, Chiba S, Kimura A, Day DF (2000) Purification and partial characterization of a novel glucanhydrolase from Lipomyces starkeyi KSM 22 and its use for inhibition of insoluble glucan formation. Biosci Biotechnol Biochem 64(2):223–228. https://doi.org/10.1271/bbb.64.223
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Shi S, Zhao H (2017) Metabolic engineering of oleaginous yeasts for production of fuels and chemicals. Front Microbiol 8:2185. https://doi.org/10.3389/fmicb.2017.02185
Spagnuolo M, Yaguchi A, Blenner M (2019) Oleaginous yeast for biofuel and oleochemical production. Curr Opin Biotechnol 57:73–81. https://doi.org/10.1016/j.copbio.2019.02.011
Starkey RL (1946) Lipid production by a soil yeast. J Bacteriol 51:33–50. https://doi.org/10.1128/JB.51.1.33-50.1946
Su S, Law CW, Ah-Cann C, Asselin-Labat ML, Blewitt ME, Ritchie ME (2017) Glimma: interactive graphics for gene expression analysis. Bioinformatics 33(13):2050–2052. https://doi.org/10.1093/bioinformatics/btx094
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Takaku H, Matsuzawa T, Yaoi K, Yamazaki H (2020a) Lipid metabolism of the oleaginous yeast Lipomyces starkeyi. Appl Microbiol Biotechnol 104(14):6141–6148. https://doi.org/10.1007/s00253-020-10695-9
Takaku H, Miyajima A, Kazama H, Sato R, Ara S, Matsuzawa T, Yaoi K, Araki H, Shida Y, Ogasawara W, Yamazaki H (2020b) A novel electroporation procedure for highly efficient transformation of Lipomyces starkeyi. J Microbiol Methods 169:105816. https://doi.org/10.1016/j.mimet.2019.105816
Wang W, Wei H, Knoshaug E, Van Wychen S, Xu Q, Himmel ME, Zhang M (2016) Fatty alcohol production in Lipomyces starkeyi and Yarrowia lipolytica. Biotechnol Biofuels 9:227. https://doi.org/10.1186/s13068-016-0647-2
Warnes GR, Bolker B, Bonebakker L, Gentleman R, Huber W, Liaw A, Lumley T, Maechler M, Magnusson A, Moeller S, Schwartz M, Venables B (2011) Gplots: various R programming tools for plotting data. R package version 2.6.0
Xu F, Li Y (2017) Biomass digestion. In: Abrams MA (ed) Encyclopedia of sustainable technologies. Elsevier, Amsterdam, pp 197–204
Yun EJ, Oh EJ, Liu JJ, Yu S, Kim DH, Kwak S, Kim KH, Jin YS (2018) Promiscuous activities of heterologous enzymes lead to unintended metabolic rerouting in Saccharomyces cerevisiae engineered to assimilate various sugars from renewable biomass. Biotechnol Biofuels 11:140. https://doi.org/10.1186/s13068-018-1135-7
Zhang L, Lim EY, Loh KC, Dai Y, Tong YW (2021) Two-stage fermentation of Lipomyces starkeyi for production of microbial lipids and biodiesel. Microorganisms 9:8. https://doi.org/10.3390/microorganisms9081724
Zhang L, Lee JTE, Ok YS, Dai Y, Tong YW (2022) Enhancing microbial lipids yield for biodiesel production by oleaginous yeast Lipomyces starkeyi fermentation: a review. Bioresour Technol 344(Pt B):126294. https://doi.org/10.1016/j.biortech.2021.126294
Zhou W, Wang Y, Zhang J, Zhao M, Tang M, Zhou W, Gong Z (2021) A metabolic model of Lipomyces starkeyi for predicting lipogenesis potential from diverse low-cost substrates. Biotechnol Biofuels 14(1):148. https://doi.org/10.1186/s13068-021-01997-9
Acknowledgements
We thank Dr. Christopher J. Fields and Dr. Jenny Drnevich, affiliated with High-performance Computing in Biology (HPCBio) at the University of Illinois at Urbana-Champaign, for their help with the initial RNAseq data analysis.
Funding
This study was supported by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research under Award Numbers DE-SC0018420 and DE-SC0018260.
Author information
Authors and Affiliations
Contributions
JL, YJ, and CR conceived the project. JL, SJ, AD, EY, and HW designed and performed experiments. AD, JL, SJ, EY, and CR analyzed the data. AD, JL, and CR wrote the manuscript and designed the figures. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deewan, A., Liu, JJ., Jagtap, S.S. et al. System analysis of Lipomyces starkeyi during growth on various plant-based sugars. Appl Microbiol Biotechnol 106, 5629–5642 (2022). https://doi.org/10.1007/s00253-022-12084-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12084-w