Abstract
Owing to the functional versatility and potential applications in industry, interest in lipolytic enzymes tolerant to organic solvents is increasing. In this study, functional screening of a compost soil metagenome resulted in identification of two lipolytic genes, est1 and est2, encoding 270 and 389 amino acids, respectively. The two genes were heterologously expressed and characterized. Est1 and Est2 are thermostable enzymes with optimal enzyme activities at 80 and 70 °C, respectively. A second-order rotatable design, which allows establishing the relationship between multiple variables with the obtained responses, was used to explore the combined effects of temperature and pH on esterase stability. The response curve indicated that Est1, and particularly Est2, retained high stability within a broad range of temperature and pH values. Furthermore, the effects of organic solvents on Est1 and Est2 activities and stabilities were assessed. Notably, Est2 activity was significantly enhanced (two- to tenfold) in the presence of ethanol, methanol, isopropanol, and 1-propanol over a concentration range between 6 and 30% (v/v). For the short-term stability (2 h of incubation), Est2 exhibited high tolerance against 60% (v/v) of ethanol, methanol, isopropanol, DMSO, and acetone, while Est1 activity resisted these solvents only at lower concentrations (below 30%, v/v). Est2 also displayed high stability towards some water-immiscible organic solvents, such as ethyl acetate, diethyl ether, and toluene. With respect to long-term stability, Est2 retained most of its activity after 26 days of incubation in the presence of 30% (v/v) ethanol, methanol, isopropanol, DMSO, or acetone. All of these features indicate that Est1 and Est2 possess application potential.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Extreme environments exhibiting elevated temperatures, extreme pH values, and exposure to organic solvents or high salinity are used to recover novel robust bioactive molecules that can be applied under industrial conditions (Antranikian and Egorova 2007). The targeted environments such as hot springs, compost, oil fields, and deep-sea marine sediments are reservoirs for extremophilic microorganisms that could produce potentially relevant industrial enzymes (Auernik et al. 2008). Culture-independent metagenomic approaches are alternatives to conventional culture-based screening methods. Recently, some extremozymes, such as amylases, amidases, proteases, cellulases, and esterases, have been successfully identified through metagenomic approaches (Daniel 2005; Simon and Daniel 2011; González-González et al. 2017; Jayanath et al. 2018; Martínez-Martínez et al. 2018).
Lipolytic enzymes, which catalyze the hydrolysis and synthesis of acylglycerols, are considered as one of the most important groups of biocatalysts. Lipolytic enzymes include esterases (EC 3.1.1.1, carboxylesterases) and true lipases (EC 3.1.1.3, triacylglycerol acyl hydrolases) and are widespread in bacteria, archaea, and eukaryotes (Hasan et al. 2006). Due to their broad substrate, pH, and temperature spectra combined with high regio- and enantioselectivity, lipolytic enzymes are of interest for food, paper, medical, detergent, and pharmaceutical industries (Hita et al. 2009; Romdhane et al. 2010; Ferrer et al. 2015; Sarmah et al. 2018). In particular, lipolytic enzymes that function in non-aqueous solvents have attracted considerable attention, as they offer new possibilities for bioprocesses, such as shifting of thermodynamic equilibrium in favor of synthesis (esterification and transesterification), controlling substrate specificity and solubility by solvent engineering, and suppressing water-dependent side reactions (Secundo and Carrea 2002; Hun et al. 2003; Ahmed et al. 2010). However, the inhibition or inactivation of enzyme activity resulting from organic solvents has restricted the use of many lipolytic enzymes (Klibanov 2001; Jin et al. 2012). To overcome this limitation, some organic solvent–tolerant (OST) lipolytic enzymes have been isolated, including enzymes from Bacillus licheniformis S-86 (Torres et al. 2009), Streptomyces coelicolor A3(2) (Brault et al. 2012), Psychrobacter celer 3Pb1 (Wu et al. 2013), Alcanivorax dieselolei B-5(T) (Zhang et al. 2014), and Acetomicrobium hydrogeniformans (Kumagai et al. 2018), as well as from metagenomes of seawater (Chu et al. 2008), compost (Kang et al. 2011), lipid-contaminated soil (Glogauer et al. 2011), mountain soil (Jin et al. 2012), swamp sediment (Seo et al. 2014), and deep-sea hydrothermal vents (Yang et al. 2018). However, these enzymes only show tolerance towards specific organic solvents.
Industrially required versatile lipolytic enzymes that exhibit satisfactory activity and stability in both water-miscible and water-immiscible organic solvents are rare (Doukyu and Ogino 2010). It has been shown that the thermostability of an enzyme in water is correlated to its tolerance against organic solvents (Kumar et al. 2016). Thus, it is straightforward to screen naturally evolved OST enzymes from thermostable ones (Lotti and Alberghina 2007; Ahmed et al. 2010). Most industrial processes utilizing lipolytic enzymes are carried out at higher temperatures (above 45 °C); it is required that the enzymes exhibit activity and stability optima around 50 °C (Sharma et al. 2002). Thus, thermostable lipolytic enzymes exhibiting organic solvent tolerance are of high importance with respect to industrial applications.
Composting is the process of biological, aerobic decomposition of organic waste by microorganisms (Ryckeboer et al. 2003). During the thermophilic phase of composting, heat generated by microbial succession can raise temperatures to above 50 °C (Dougherty et al. 2012). Correspondingly, compost is a potential source for recovery of thermostable enzymes. Recently, lipolytic enzymes have been isolated from compost (Lämmle et al. 2007; Tirawongsaroj et al. 2008; Kim et al. 2010; Ohlhoff et al. 2015; Woo Lee et al. 2016), but only one (EstCS2) of the isolated enzymes was moderately thermostable (optimum 55 °C) and showed resistance to certain water-miscible organic solvents (Kang et al. 2011).
In this study, two genes encoding lipolytic enzymes (est1 and est2) were identified from a thermophilic compost metagenome. The corresponding enzymes were purified and characterized. Enzyme characterizations are usually conducted as one-factor-at-a-time for comparison with reported enzyme features from other studies. However, this methodology ignores interacting effects between factors, which may result in misleading conclusions, especially when at least two requirements must be fulfilled simultaneously. An alternative is to employ design of experiments (DOE) methodologies, i.e., second-order rotatable design approach, which use statistical and mathematical approaches to evaluate the combined effect of factors. DOE has been successfully applied in different aspects related to lipolytic enzymes such as the growth condition optimization and enzyme activity or stability measurements (Kamimura et al. 2001; Shieh et al. 2003; Benaiges et al. 2010). In this study, the combined effect of pH and temperature on the stability of Est1and Est2 was evaluated by the second-order rotatable design approach. Analysis of the recovered two metagenome-derived enzymes showed that they are thermophilic and tolerant towards organic solvents. In addition, Est2 was remarkably resistant to both water-miscible and immiscible organic solvents.
Materials and methods
Strains and plasmids
For the construction of metagenomic plasmid libraries, Escherichia coli TOP10 and pFLD (Invitrogen GmbH, Karlsruhe, Germany) were used as host and vector, respectively. Escherichia coli BL21 Star (DE3) and the pET101/D-TOPO® vector (Invitrogen GmbH) were used for heterologous expression of the recovered lipolytic genes.
Sample collection and DNA extraction
Compost samples were collected at a composting company (Göttingen GmbH, Göttingen, Germany, 51° 34′ 25.1″ N 9° 54′ 33.0″ E). The sampling pile was the fermentation product of household waste and fresh tree branches. To ensure using mainly thermophilic microorganisms as a source for metagenomic library construction, compost at the core zone of compost pile was collected. The temperature at the sampling spot was 55 °C. The compost soil sample (50 g) was collected in sterile plastic bags and stored at − 20 °C until required.
Metagenomic DNA of the compost sample was extracted following a phenol-chloroform method according to Zhou et al. (1996). In addition, DNA was also isolated with MoBio Power Soil DNA extraction kit following the protocol of the manufacturer (MoBio Laboratories, Carlsbad, CA, USA). DNA obtained from these two methods was pooled and stored at − 20 °C until required.
Metagenomic library construction and screening for lipolytic activity
To construct a metagenomic plasmid library, DNA was sheared, and fragments from 6 to 12 kb were recovered by gel extraction with the peqGold gel extraction kit (Peqlab Biotechnologie GmbH, Erlangen, Germany). End-repaired DNA fragments and PmlI-digested pFLD vector were ligated by employing T4 DNA ligase at 16 °C, overnight as recommended by the manufacturer (Thermo Scientific, Bremen, Germany). To screen for lipolytic activity, metagenomic library-bearing cells were plated onto LB agar plates containing 100 μg/ml ampicillin and 1% (v/v) emulsified tributyrin (Sigma, Germany) and subsequently incubated at 30 °C 1 to 7 days. Lipolytic-positive clones were identified by the formation of clear zones (halos) around individual colonies. The phenotype of positive clones was confirmed by the isolation of recombinant plasmids from positive strains, transformation of the isolated plasmids into the host, and rescreening on indicator agar plates.
Sequence analysis and homology modeling
Lipolytic genes (est1 and est2) were initially predicted by using the ORF Finder program (http://www.ncbi.nlm.nih.gov/gorf/gorf.html), and verified by Clone Manager and FramePlot analysis (Ishikawa and Hotta 1999). Similarity searches of the deduced amino acid sequences were performed by BLASTP program against the public GenBank database (Ye et al. 2006). Signal peptides were detected by using the SignalP 4.0 server (Bendtsen et al. 2004). The deduced amino acid sequences of est1 and est2 and reference sequences retrieved from GenBank were used to construct a phylogenetic tree with the neighbor-joining method by using MEGA version 6 (Tamura et al. 2013). Bootstrapping of 1000 replicates was used to estimate the confidence level.
Based on the deduced amino acid sequence, secondary structure and tertiary structure predictions were performed with I-TASSER (Zhang 2008). The identified structural analogs were used for multiple-sequence alignment using the Expresso webserver (Notredame et al. 2000). Figures showing secondary structure alignments were exported by ESPript3 (Robert and Gouet 2014). The analog with the highest TM score was also selected for structural superimposition.
Expression and purification of recombinant proteins Est1 and Est2
The primer pairs 5′-CACCATGCCCCTGGCCCGAGTGGA-3′ and 5′-GGCGCCCACCGGC ACCTGAGTC-3′ and 5′-CACCATGACCGAGCTGCCGGTGGGAG-3′ and 5′-GCG TCTTAGCGCGCGGTACAC-3 were used to amplify est1 and est2, respectively. The resulting PCR products were purified and ligated into expression vector pET101/D-TOPO® according to the protocol of the manufacturer (Invitrogen). To produce His6-tagged Est1 and Est2, recombinant plasmid DNA was transformed into E. coli BL21 (DE3) cells and plated on LB agar plates with 100 μg/ml ampicillin. A single colony was picked and grown overnight at 30 °C in 60-ml LB medium containing 100 μg/ml ampicillin. Subsequently, this pre-culture was added to 600-ml LB medium with 100 μg/ml ampicillin and grown with shaking at 30 °C. At an optical density (OD600) of 0.6, isopropyl-beta-D thiogalactopyranoside (IPTG) was added to a final concentration of 0.5 mM. After a 6-h induction at 30 °C, cells were harvested by centrifugation (7000×g, 4 °C, 10 min). Cell pellets were washed with 100 ml LEW buffer (50 mM NaH2PO4, 300 mM NaCl, pH 8) and stored at − 20 °C until required.
To purify Est1 and Est2, Protino® Ni-TED 2000 packed column (Macherey-Nagel, Germany) was used following the manufacturer’s protocol, however, with the modified LEW buffer (50 mM NaH2PO4, 1 M NaCl, 10% v/v glycerol, 0.05% v/v Triton X-100, pH 8). Protein concentration was measured by the Bradford method (Bradford 1976). Purity and molecular mass of the purified proteins were determined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) using the procedure of Laemmli (1970). Fractions derived from affinity chromatography showing a single band with the estimated molecular mass of the targeted proteins were pooled, dialyzed with 50 mM sodium phosphate buffer (pH 8), and stored in 50% (v/v) glycerol at − 20 °C until use.
Standard enzyme assays
Esterase activity was measured by a spectrophotometric method (Jaeger et al. 1999) using p-nitrophenyl (p-NP) acyl esters (Sigma) as substrates. To minimize substrate auto-hydrolysis at high temperatures, p-NP caprylate (C8) was used as a standard substrate. Unless otherwise indicated, Est1 activity was measured at 80 °C in 1 ml containing 50 mM sodium phosphate assay buffer (pH 8),1 mM p-NP caprylate (C8), and 1% (v/v) isopropanol, while Est2 activity was measured at 70 °C in 1 ml containing 50 mM TAPS (3-(2, 4 dinitrostyrl)-(6R,7R-7-(2-thienylacetamido)-ceph-3-em-4-carboxylic acid) assay buffer (pH 9), 1 mM p-NP caprylate (C8), and 1% (v/v) isopropanol. The assay buffer was initially incubated in a screwed-cap test tube for 10 min at assay temperature. Then, the reaction was initiated by adding enzyme and substrate to the buffer. The amount of p-nitrophenol released by esterase-catalyzed hydrolysis was continuously monitored at a wavelength of 410 nm against an enzyme-free blank. One unit (U) of enzyme activity was defined as the amount of enzyme that released 1 μmol of p-nitrophenol per minute. All experiments were performed in at least triplicate. Results are shown as mean values ± standard deviation (SD).
Substrate specificities of Est1 and Est2 were checked towards the following p-NP acyl esters of different chain lengths: p-NP acetate (C2), p-NP butyrate (C4), p-NP valerate (C5), p-NP caproate (C6), p-NP caprylate (C8), p-NP caprate (C10), p-NP laurate (C12), p-NP myristate (C14), and p-NP palmitate (C16). Considering the instability of short-chain substrates, the assay temperature was decreased to 50 °C. Initial rates of reaction for p-NP butyrate and p-NP valerate were calculated by estimating Est1 and Est2 activities with different substrate concentrations ranging from 1 to 2000 μM. Values for Km and Vmax were determined by employing the Lineweaver-Burk plots (Lineweaver and Burk 1934). Lipolytic activity towards different triacylglycerides was also measured qualitatively by incubating Est1 and Est2 on agar plates emulsified with tributyrin (C4), tricaproin (C6), tricaprylin (C8), tricaprin (C10), trilaurin (C12), trimyristin (C14), or tripalmitin (C16). Formation of clearing zones (halos) on agar plates indicated lipolytic activity. Beta-lactamase activity of Est2 was tested spectrophotometrically at 486 nm, under standard assay conditions with 1 mM nitrocefin (E-isomer) as substrate.
Effect of temperature and pH
The effect of pH on Est1 and Est2 activities was measured at 348 nm (the pH-independent isosbestic wavelength) under standard assay conditions (Glogauer et al. 2011). The following overlapping buffer systems were used: 50 mM acetate buffer (pH 3.0 to 6.0), 50 mM sodium phosphate buffer (pH 6.0 to 8.0), 50 mM TAPS buffer (pH 8.0 to 9.0), and 50 mM CHES (N-cyclohexyl-2-aminoethanesulfonic acid) buffer (pH 9.0 to 10.0). Temperature optima for Est1 and Est2 activities were measured in a temperature range of 20 to 100 °C. Thermostability of enzyme activity was determined by incubating Est1 and Est2 in their optimal buffers at various temperatures (50 to 80 °C) for up to 6 days. Subsequently, Est1 and Est2 activities were determined under standard assay conditions.
Combined effect of pH and temperature on the stabilities of Est1 and Est2
Second-order rotatable design was applied to study the combined effect of pH and temperature on the stabilities of Est1and Est2. The design was based on five levels and two variables (Table S2). Experimental data were fitted to the empirical model using Eq. (1):
in which Z was residual relative activity, presented as the percentage of activity measured before incubation and under standard assay conditions; X and Y were code values of pH and temperature shown in Table S2; b0, b1, b2, b12, b11, and b22 were regression coefficients. Significance of regression coefficients was checked by Student’s t test (α = 0.05). Statistically non-significant coefficients were removed, and best-fit parameters were recalculated (Lazić 2004). The consistency of regression models was checked by Fisher’s test (α = 0.05). The ratios of the following mean squares were compared with the F-criterion tabular values. Based on the following mean square ratios (Box et al. 2005), models were accepted if:
Est1 and Est2 were incubated under conditions described in Table S1 for 2 h, and residual activity was subsequently measured under the respective standard assay conditions.
Effect of miscible and immiscible organic solvents
The following organic solvents with different log p values were used in this study: water-miscible organic solvents of DMSO (− 1.3), methanol (− 0.75), ethanol (− 0.24), acetone (− 0.24), isopropanol (0.074), and 1-propanol (0.28), as well as the water-immiscible organic solvents of ethyl acetate (0.68), diethyl ether (0.85), chloroform (2.0), and toluene (2.5). The effects of water-miscible organic solvents on Est1 and Est2 activities were measured by adding each organic solvent into the assay buffer to obtain a final concentration ranging from 6 to 30% (v/v) under standard assay conditions. Enzyme activity measured in organic solvent-free assay buffer was regarded as 100%. Appropriate controls were also set to eliminate changes in extinction coefficients due to the presence of solvents.
To evaluate short-term stability towards water-miscible and water-immiscible organic, Est1 and Est2 were incubated in 100-μl aliquots with different amounts of water-miscible organic solvents (0 to 75%, v/v for Est1; 0 to 95%, v/v for Est2) or water-immiscible organic solvents (15 and 30%, v/v) at 30 °C for 2 h with vigorous shaking (300 rpm). The long-term stability towards water-miscible solvents was only measured for Est2. In the presence of 30 or 60% (v/v) organic solvents, Est2 was incubated at 30 °C with constant shaking in a screwed-cap test tube for up to 26 or 13 days, respectively. Enzyme activities in either the aqueous phase (for water-immiscible solvents) or the mixture (for water-miscible solvents) were measured. Each water-miscible organic solvent was equalized to the same final concentration in the assay buffer and the residual activity was measured under standard assay conditions. A blank reference was prepared by using the same buffer solution without enzyme containing and the same amount and type of organic solvent (Shao et al. 2013). Residual activity was subsequently measured under the respective standard assay conditions. The activity measured at the start of the experiment was taken as 100%.
Effect of additives on Est1 and Est2 activities
The effects of metal ions on Est1 and Est2 activities were examined in 50 mM sodium phosphate buffer (pH 8) at 50 °C, in the presence of 1 mM and 10 mM KCl, CaCl2, MnCl2, MgCl2, ZnSO4, FeSO4, CuCl2, NiSO4, FeCl3, and AlCl3. The inhibitory effect on enzyme activity was measured under standard assay conditions with the following known esterase effectors (each 1 mM and 10 mM): phenylmethyl-sulfonyl fluoride (PMSF), dithiothreitol (DTT), and ethylenediaminetetraacetic acid (EDTA). In addition, the effect of the following detergents (each 0.1 and 1%, v/v), Triton X-100, Tween 20, Tween 80, and SDS, was determined. Furthermore, the effect of NaCl and KCl on enzyme activity in a range between 0.5 and 4 M was assessed. Activity measured in additive-free assay buffer was regarded as 100% activity, while reactions that included corresponding additive but no enzyme were used as blanks.
Accession number
The gene sequences are available at the GenBank database under accession numbers KR149567.1 (Est1) and KR149568.1 (Est2).
Results
Metagenomic library screening and analysis of two novel esterase-encoding genes
To isolate novel lipolytic enzymes, a compost sample at the thermophilic stage (55 °C) was used for constructing a metagenomic plasmid library. The library consisted of approximately 675,200 clones with an average insert size of 5.3 kb and comprised a total size of 3.58 GB. Among the 279 lipolytic-positive clones, two E. coli clones harboring the plasmids pFLD_Est1 and pFLD_Est2 showed strong lipolytic activity (large halos) on indicator plates and were selected for further characterization.
Sequence analyses of the plasmids pFLD_Est1 and pFLD_Est2 revealed that each contained one putative esterase-encoding gene, est1 (813 bp) and est2 (1170 bp), respectively. The deduced proteins comprised 270 (Est1) and 389 (Est2) amino acids. Putative signal peptides indicating extracellular localization were not detected in the deduced protein sequences. Sequence similarity searches showed that Est1 exhibited 49% identity to a hypothetical protein from Candidatus Entotheonella (GenBank: ETW96815) and 43% identity to Est28 from a grassland soil metagenomic library (Nacke et al. 2011). Est2 showed 53% sequence identity to a beta-lactamase from Streptomyces lavenduligriseus (GenBank: WP_030784121) and 52% sequence identity to a putative esterase from Streptomyces bottropensis (GenBank: EMF58012).
Sequence analysis and the subsequently constructed phylogenetic tree revealed that Est1 belonged to family V and Est2 to family VIII of lipolytic enzymes (Fig. S1). The tertiary structure predicted by I-TASSER obtained the C-scores of 1.03 for Est1 and 1.27 for Est2, which indicated a significant confidence of good quality.
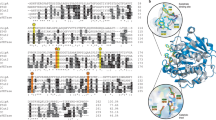
The two conserved family V motifs G-X-S-X-G-G and P-T-L were present in the Est1 protein sequence at amino acid positions 92 to 97 and 208 to 210, respectively (Fig. S2a). The tertiary structure of Est1 was composed of a cap domain with five α-helices (α4 to α8, Fig. S2a) and an α/β-hydrolase fold core domain (Fig. 1a). The core domain consists of six helices surrounded by eight β-strands that form parallel structures, in which Ser94 is located between β5 and α3, Asp217 after β7, and His268 between β8 and α10. The overall structure of Est1 superimposed on MGS-M2 (TM score 0.96; RMSD 0.94) (Alcaide et al. 2015), with a global amino acid sequence identity of 21.9% (Fig. 1c).
The modeled three-dimensional structure of Est1 and Est2. a A 3D structure model of Est1. The overall structure is composed of two domains: cap domain and catalytic domain. b A 3D structure model of Est2. The overall structure harbors two domains: a large α/β domain and a small α-helix domain. α-helices and β-strands are colored in cyan and magenta, respectively. The catalytic triad of Est1 (residues Ser94, Asp215, and His245) and that of Est2 (residues Ser70, Lys73, and Try160) are indicated in stick representation. The cap domain for Est1 and the small α-helix domain for Est2 are presented as transparent surface. c Structural superposition of Est1 (pink) onto its structural homolog MGS-M2 (yellow; PDB: 4Q3L). d Structural superposition of Est2 (pink) onto its structural homolog CcEstA (yellow; PDB:5GKV)
Due to the high sequence similarity between family VIII lipolytic enzymes and class C beta-lactamases/penicillin-binding proteins, amino acid sequences in the two categories were aligned (Fig. S2b). All the aligned sequences shared the same catalytic triad of serine, lysin, and tyrosine. Conserved family VIII motifs including S70-X-X-K73, G154-X-X-X-X-H159, and H/W341-X-G343 were detected in Est2 sequences. However, classical β-lactamase motifs such as Y-A-N and L-S/T-G (KTG-box) were absent in the Est2 sequence (Fig. S2b). Similar to Est1, the tertiary structure of Est2 also consists of two domains: a large α/β domain and a small α-helical domain. The small α-helix domain contains four helices and two 310 helices. For the large α/β domain, a central eight-stranded antiparallel β-sheet is flanked by six helices on one face and two on the opposite face (Fig. 1c). The deduced catalytic residues of Ser70, Lys73, and Tyr159 are located in the large cavity between the two domains (Fig. 1b). The overall structure of Est2 superimposed well on CcEstA (TM score 0.95; RMSD 0.46) (Woo Lee et al. 2016), with a global amino acid sequence identity of 39.6% (Fig. 1d).
Purification of Est1 and Est2 and substrate specificity towards p-NP acyl esters
The esterase-encoding genes est1 and est2 were successfully cloned in expression vectors and expressed in E. coli BL21(DE3). The produced gene products Est1 and Est2 were subsequently purified by Ni-TED affinity chromatography, yielding 28.9-fold and 16.9-fold purification values and specific activities of 22.2 and 7.3 U/mg, respectively (Table S2). SDS-PAGE analysis revealed single bands with molecular masses of approximately 35 kDa (Est1) and 48 kDa (Est2) (Fig. S3), which were in accordance with the calculated masses including during cloning added V5 epitope and His6-tag.
Est1 and Est2 exhibited a substrate preference for esters with short-chain fatty acids. The maximal activities were detected with p-NP butyrate (C4) for Est1 and p-NP valerate (C5) for Est2 as substrates (Fig. 2). Correspondingly, p-NP butyrate and p-NP valerate were used for calculating the Km and Vmax values. The Km and Vmax values of Est1 were 3.0 μM and 31.2 U/mg, respectively, and that of Est2 2.7 μM and 19.0 U/mg, respectively. With respect to esters with long-chain fatty acids as substrates (C10 to C16), Est1 was active with p-NP caprate (C10) retaining 45% activity, but barely with other substrates. In comparison, Est2 hydrolyzes more acyl esters with long-chain fatty acids. It showed 80%, 45%, 12%, and 30% activity towards p-NP caprate (C10), laurate (C12), p-NP myristate (C14), and p-NP palmitate (C16), respectively. Chain length selectivity towards triacylglycerides was recorded for the substrates tributyrin (C4) and tricaproin (C6) for both enzymes. The preference for short substrates (< C10) indicates that Est1 and Est2 are esterases, rather than “true” lipases which are commonly found in family I of lipolytic enzymes (Arpigny and Jaeger 1999). Moreover, beta-lactamase activity of Est2 with nitrocefin as substrate was tested, but no significant beta-lactamase activity was detected (data not shown).
Effect of pH and temperature on Est1 and Est2 activities
Est1 and Est2 were active over the entire tested pH range (3 to 10), with the exception of Est1 at pH 3 (Fig. 3a, d). Est1 exhibited maximal activity at pH 7 and retained more than 95% activity between pH 6 and 8. Est2 activity increased with pH and peaked at pH 9. With respect to the temperature dependence of enzyme activity, Est1 and Est2 displayed a similar ascending trend along the temperature gradient, showing a maximal activity at 80 and 70 °C, respectively. At higher temperatures (90 and 100 °C), approximately 80% activity was retained (Fig. 3b, e).
Effect of pH and temperature on esterase activity. a Effect of pH on Est1 activity. Maximal activity at pH 7 (45.8 U/mg) was taken as 100%. b Effect of temperature on Est1 activity. Maximal activity at 80 °C (46.1 U/mg) was taken as 100%. c Thermostability of Est1 at 50 °C (closed circle), 60 °C (open circle), and 70 °C (closed diamond); activity measured before incubation (40.4 U/mg) was taken as 100%. d Effect of pH on Est2 activity. Maximal activity at pH 9 (14.4 U/mg) was taken as 100%. e Effect of temperature on Est2 activity. Maximal activity at 70 °C (15.2 U/mg) was taken as 100%. f Thermostability of Est2 at 50 °C (closed triangle), 60 °C (open triangle), 70 °C (closed square), and 80 °C (open square); activity measured before incubation (12.0 U/mg) was taken as 100%
Despite that the activity maximum of Est1 was at 80 °C, stability at 70 °C was low as a rapid drop of activity to 50% was detected after 15 min of incubation. Similar results were obtained for Est2 at 80 °C with 35% residual activity after 30 min. Strikingly, Est2 showed significant stability at 70 °C, with 50% residual activity after 12 h of incubation. At lower temperatures (50 or 60 °C), the activities of both enzymes were remarkably stable for extended incubation times; nonetheless, Est1 displayed a higher stability than Est2 (Fig. 3c). Est1 retained more than 80% activity at 50 °C over the entire incubation period (7 days), whereas 52% residual activity was observed for Est2 after 5 days. When incubated at 60 °C, Est1 exhibited a half-life of 2 days, which is nearly twice as that of Est2 (Fig. 3f).
Combined effect of pH and temperature on Est1 and Est2 stabilities
The combined effect of temperature and pH on the stability of Est1 and Est2 was visualized by response surfaces (Fig. 4). After removing the insignificant terms and refitting to the empirical model, the following equations for Est1 (4) and Est2 (5) stability under different conditions were obtained:
Response surface corresponding to the combined effect of pH and temperature on Est1 (a) and Est2 (b) stabilities. Residual activity was expressed as a percentage of the initial activity measured before incubation under standard assay conditions. The elliptical contour plot on the pH–T dimension demonstrates the change of residual activity (%)
Variance analysis implied that the models for Est1 and Est2 significantly fitted to the experimental data (\( {F}_1\ge {\mathrm{F}}_{\mathrm{den}}^{\mathrm{num}} \), \( {F}_2\le {\mathrm{F}}_{\mathrm{den}}^{\mathrm{num}} \), α = 0.05; Table S3). In equations (4) and (5), the negative coefficients for T, pH2 and T2 indicate the existence of an absolute maximum of residual activity. The calculated optimal conditions were located at pH 8.2 and 39.1 °C (Est1) and pH 8.7 and 48.9 °C (Est2). It was predicted that Est1 and Est2 retain 98.6 and 96.3% residual activity, respectively, under these conditions. Moreover, according to the elliptical contour plot on the pH–T (x–y) dimension, the pH and temperature ranges in which Est1 and Est2 retain more than 80% residual activity were pH 7.2 to 9.2 and 23.6 to 54.6 °C for Est1 and pH 7.0 to 10.4 and 37.4 to 60.4 °C for Est2 (Fig. 4). In conclusion, Est1 and Est2 are stable under thermophilic and alkaline conditions.
Effect of water-miscible organic solvents on Est1 and Est2 activities
Est1 activity increased to approximately 150% in the presence of low concentrations of 1-propanol (below 6%, v/v), and ethanol and isopropanol (< 12%, v/v). At higher concentration, Est1 activity decreased rapidly and was not detectable at 18% (v/v) 1-propanol, 30% (v/v) ethanol, and 24% (v/v) isopropanol (Fig. 5a). Addition of DMSO, methanol, and acetone inhibited Est1 activity over the tested concentration range, retaining 50% activity at 30% (v/v) DMSO, or less than 10% activity at 30% (v/v) methanol and acetone.
Effects of water-miscible organic solvents on of Est1 (a) and Est2 (b) activities. Catalytic activity was measured under standard assay conditions in the presence of different amounts of ethanol (closed square), methanol (open square), isopropanol (closed circle), 1-propanol (open circle), DMSO (closed triangle), and acetone (open triangle). Specific activities corresponding to 100% relative activity were 21.9 U/mg (Est1) and 5.1 U/mg (Est2)
Ethanol, methanol, isopropanol, and 1-propanol had a stimulatory effect on Est2 activity almost at all tested concentrations (Fig. 5b). In the presence of ethanol and methanol, Est2 activity increased continually with raising concentrations. At 30% (v/v) ethanol and methanol, Est2 activity increased 8.8-fold and 5.4-fold, respectively. For isopropanol and 1-propanol, optimal Est2 activity was recorded at concentrations of 18 and 12% (v/v), respectively (Fig. 5b). In the presence of DMSO, Est2 retained more than 70% activity at all tested concentrations, with the maximal activity at 6% (v/v). Acetone caused an activity decrease to 34.1% at 30% (v/v).
Effect of water-miscible organic solvents on Est1 and Est2 stabilities
Est1 retained almost unchanged residual activity after 2 h of incubation in the presence of 15 and 30% (v/v) of all tested organic solvents, with the exception of 1-propanol (Fig. 6a). Est1 residual activity rapidly dropped to 17.0% at 30% (v/v) 1-propanol. Decreased tolerance was also detected after exposure to 45% (v/v) isopropanol, methanol, acetone, and DMSO for 2 h (Fig. 6a). At 60% (v/v) solvent concentration, Est1 exhibited moderate tolerance to DMSO (60.5% residual activity), but lost most of its activity (less than 10% residual activity) in the presence of the other tested organic solvents. Est1 activity was almost inactivated by all the tested organic solvents at 75% (v/v).
Heatmap displaying the effect of enzyme stability towards water-miscible organic solvents. Short-term stability of Est1 (a) and Est2 (b) towards different concentrations of organic solvents at 30 °C for 2 h. The specific activity expressed as percentages of Est1 reference reactions (100%) are 23.3, 17.5, 24.3, 6.6, 15.3, and 13.6 U/mg for reactions in the presence of ethanol, methanol, isopropanol, 1-propanol, DMSO, and acetone, respectively, and that of Est2 are 32.7,14.4, 17.7, 42.7, 4.34, and 3.1 U/mg, respectively. Long-term stability of Est2 towards 30% (v/v; c) and 60% (v/v; d) organic solvents for prolonged time periods at 30 °C. The specific activity values expressed as percentages of Est2 reference reactions (100%) are 14.3, 7.1, 9.2, 25.9, 5.0, and 4.1 U/mg for reactions in the presence of ethanol, methanol, isopropanol, 1-propanol, DMSO, and acetone, respectively
In comparison with Est1, Est2 displayed higher resistance towards water-miscible solvents (Fig. 6b). In the organic solvent concentration range of 0 and 60% (v/v), Est2 retained its full activity in the presence of isopropanol, methanol, and ethanol and had a slight activity loss by the addition of DMSO and acetone. On the contrary, Est2 stability decreased along the rising concentration of 1-propanol, retaining 32.4% residual activity at 60% (v/v). Incubating with concentrations above 60% (v/v) of isopropanol, 1-propanol ethanol, or methanol led to a rapid decline of Est2 residual activity (Fig. 6b). Exposure to 75% (v/v) DMSO and acetone led to inactivation of Est2 activity.
Est2 exhibited high stability in the presence of 30% (v/v) ethanol, methanol, isopropanol, DMSO, and acetone, as its residual activity was above 70% over the extended incubation period of up to 26 days (Fig. 6c). This residual activity was even higher than that Est2 exhibited during incubation without additions (Fig. S4). However, a continuous drop in Est2 residual activity was observed when exposed to 1-propanol. Est2 also displayed decreased tolerance towards 60% (v/v) organic solvents (Fig. 6d). Incubation with 1-propanol, ethanol, and methanol reduced Est2 activity rapidly (< 10% residual activity). Nonetheless, Est2 exhibited substantial tolerance against isopropanol, DMSO, and acetone, retaining approximately 40% residual activity after the 13-day incubation.
Effect of water-immiscible organic solvents on Est1 and Est2 stabilities
Incubation with water-immiscible organic solvents of diethyl ether, chloroform, and toluene resulted in deleterious effects on Est1 enzyme activity, which was not detectable at the tested concentrations (Table 1). However, Est1 activity displayed some resistance towards ethyl acetate, with 46.6 and 21.1% residual activities at the tested concentrations of 15 and 30% (v/v), respectively. In contrast, Est2 was tolerant to ethyl acetate, diethyl ether, and toluene. Est2 retained its activity after incubation with ethyl acetate and diethyl ether. In the presence of toluene, Est2 retained 87.0 and 68.8% activities at the tested concentrations of 15 and 30% (v/v), respectively. In addition, Est2 was as Est1 inactivated by chloroform.
Effect of other additives on Est1 and Est2 activities
Metal ions, inhibitors, detergents, and salts were also analyzed for their effects on Est1 and Est2 activities. Est1 and Est2 are generally resistant to various metal ions (Table S4). The addition of tested metal ions at 1 mM concentration had minor effects on Est1 and Est2 activities as approximately 90% of the activity was retained. Moreover, both enzymes exhibited substantial tolerance (above 50% activity) towards some metal ions such as Ca2+, Zn2+, Cu2+, Mn2+, Fe3+, and Al3+ at a concentration of 10 mM. Enzyme activity was slightly enhanced in the presence of 10 mM Mg2+ (Est1) and Zn2+ (Est2). The presence of the chelating agent EDTA did not affect Est1 activity but decreased Est2 activity to less than 70%. The latter results indicated that Est1 activity is independent of metal ions, whereas Est2 might be a metalloenzyme (Mohamed et al. 2013).
With respect to the detergents, Est1 activity was enhanced in the presence of 0.1% (v/v) Tween 20 and 0.1 and 1% (v/v) Tween 80. Est2 activity was less resistant than that of Est1 towards the tested detergents. Est2 retained more than 70% activity after addition of 1 mM Triton-100, Tween 20, and Tween 80. The anionic detergent SDS inhibited the activity of both enzymes entirely (Table S4). The activity of Est1 and Est2 was substantially decreased at 10 mM DTT and PMSF. DEPC displayed detrimental effects on Est1 activity, while Est2 activity was almost unaffected even at a concentration of 10 mM. The inhibition of enzyme activity by SDS and PMSF indicates that Est1 and Est2 belong to the serine hydrolases (Peng et al. 2011). In addition, Est1 and Est2 were active in the presence of 0 to 4 M NaCl and KCl. Both enzymes retained more than 50% of activity up to 2.5 M NaCl and KCl, which is indicative of halotolerance (Table S5).
Discussion
Extreme environments such as compost have been used for mining biocatalysts, which are likely to be adapted to harsh industrial reaction conditions (Ryckeboer et al. 2003). In this study, compost derived from the thermophilic core of the pile was used as DNA source to construct a metagenomic library, whereby two putative genes encoding lipolytic enzymes were identified. The low identities of the deduced amino acid sequences to known proteins (Est1 43% and Est2 53%) indicate that Est1and Est2 are novel lipolytic enzymes. Est1 is most related to the characterized esterase EstPS5, which was derived from a screening of a peat-swamp forest soil metagenome (Bunterngsook et al. 2010). Est2 shared the highest sequence identity with a beta-lactamase from Streptomyces achromogenes. In addition, most of the enzyme sequences similar to Est2 were derived from members of the Streptomyces genus. Streptomyces strains are predominantly found in soil and decaying vegetation and well-known as important natural sources for antibiotics (Raja and Prabakarana 2011).
Phylogenetic analyses indicate that Est1 and Est2 belong to family V and family VIII of lipolytic enzymes, respectively (Fig. S1). For most of the known lipolytic enzymes, the catalytic activity generally relies on the typical catalytic triad of Ser, Asp/Gly, and His (Ollis et al. 1992; Jaeger et al. 1999). This triad is also present in the Est1 amino acid sequence (Fig. S2a) and its tertiary structure (Fig. 1a). However, another catalytic triad consisting of Ser70, Lys73, and Tyr159 (Sakai et al. 1999) is responsible for Est2 catalytic activity (Fig. 1c, Fig. S2b). This triad is conserved among family VIII lipolytic enzymes, class C β-lactamases, and penicillin-binding proteins (Arpigny and Jaeger 1999; Hausmann and Jaeger 2010; Biver and Vandenbol 2013; Popovic et al. 2017). Despite the same catalytic triad and high amino acid sequence identity to β-lactamases, some family VIII esterases show promiscuous β-lactamase activity. Similar to Est2, the family VIII esterases EstB (Petersen et al. 2001), Lip8 (Ogino et al. 2004), Est2K (Kim et al. 2010), and Est7K (Woo Lee et al. 2016) showed negligible or no detectable β-lactamase activity, whereas EstC (Rashamuse et al. 2009), EstM-N1 (Yu et al. 2011), and PBS-2 (Boyineni et al. 2014) exhibited moderate or high β-lactamase activity. Yu et al. (2011) suggested that those promiscuous β-lactamase activities could be a result of family VIII esterases evolving from class C β-lactamases or vice versa.
According to the predicted tertiary structure (Fig. 1), the active sites of Est1 and Est2 are protected by an α-helix domain (cap domain). This domain acts as a shield for the catalytic site of many lipolytic enzymes and appears to play a key role in several functional aspects, such as activity, substrate specificity, and thermostability (Gall et al. 2014; Li et al. 2015; Kim 2017). Est2 was capable of utilizing acyl esters with long-chain fatty acids as substrate (C10, C12, and C16) (Fig. 2b). In addition, the Est2 structural homolog Est-Y29 (Ngo et al. 2013) was also reported to hydrolyze a wide variety of hydrophobic compounds, as a result of the deep hydrophobic patch between the large α/β domain in which the small α-helix domain defines a wide active site (Ngo et al. 2014).
Est1 and Est2 were generally thermoalkaline esterases (Fig. 3). In terms of thermophilicity, defined as increased enzymatic activity along a temperature gradient (Georis et al. 2000), Est1 and Est2 have great advantages in comparison with its homologs from other sources. Although esterases from Fervidobacterium nodosum (Yu et al. 2010), Anoxybacillus gonensis (Faiz et al. 2007), Sulfolobus tokodaii (Angkawidjaja et al. 2012), and a compost metagenome (Riedel et al. 2015) displayed optimal activities above 70 °C, Est1and Est2 exhibited higher activities (above 80%) at 90 and 100 °C. Moreover, Est1 and Est2 displayed unprecedented thermostability at high temperatures, with a half-life of more than 7 days at 50 °C and 2 days at 60 °C for Est1 (Fig. 4c) and 5 days at 50 °C, 1 day at 60 °C, and 12 h at 70 °C for Est2 (Fig. 4f). This remarkable feature distinguished the two enzymes from their thermophilic counterparts, such as esterases from the thermophilic microorganisms Archaeoglobus fulgidus (40% residual activity after 30 min at 60 °C; D’Auria et al. 2000), Thermogutta terrifontis (75% residual activity after 30 min at 70 °C; Sayer et al. 2015a), and Thermoanaerobacter tengcongensis (half-life of 2 h at 70 °C; Cook et al. 1996), as well as those from metagenomes of compost (approx. 90% residual activity after 1 h at 60 °C; Kang et al. 2011) and hot spring (half-life of 6 h at 60 °C; Zarafeta et al. 2016).
In general, thermostability is dependent on the structural rigidity, which is an accumulation of various features, including but not limited to amino acid composition, ion pairing, hydrogen bonds, hydrophobic interactions, and sulfide bridges (Sadeghi et al. 2006; Jochens et al. 2010; Ebrahimi et al. 2011; Pezzullo et al. 2013). Interestingly, the two characterized structural homologs of Est1, MGS-M2 (Alcaide et al. 2015) and TtEst (Sayer et al. 2015b), were reported as thermostable esterases (Alcaide et al. 2013; Sayer et al. 2015a). The thermostability of Est1 and Est2 is advantageous for industrial applications, as higher reactivity, stability, and process yields, as well as lower viscosity and fewer contamination problems, can be achieved at an elevated operation temperature (Lima et al. 2004; Panda and Gowrishankar 2005; Doukyu and Ogino 2010; Sood et al. 2016).
Temperature and pH play important roles in enzyme-catalyzed reactions and the two factors have to be considered together for optimization of enzyme reactions. Est1 and Est2 are most stable at conditions which are close to that of the original compost habitat (Fig. 4a, b). Generally, enzymes are stable at conditions similar to their original habitats (Elend et al. 2006; Kovacic et al. 2016), but show maximal activities at higher or lower temperatures (Hardeman and Sjoling 2007; Hu et al. 2010), which is also the case for Est1 and Est2 (Fig. 3). Est1 and Est2 were predicted to retain more than 80% residual activity over a broad temperature and pH range (Fig. 4). Thus, together with the feature of broad substrate specificity (Fig. 2), the two enzymes, particularly Est2, could be potentially utilized in detergents, in which high enzyme stability is required during washing conditions of pH 8 to 11 and temperatures of 30 to 60 °C, as well as versatile substrate specificity (Nerurkar et al. 2013; Bora 2014).
Water-miscible organic solvents are generally detrimental to enzymes. In contrast, Est2 activity was significantly enhanced by the addition of ethanol, isopropanol, methanol, and 1-propanol (Fig. 5b). The stimulated catalytic activity of esterase from Burkholderia cepacia was also observed in the presence of DMSO, DMF, methanol, ethanol, 2-propanol, and acetone (Takeda et al. 2006). Similar results were obtained for family VIII esterases Est2K, lpc53E1, and Est7K in the presence of isopropanol and methanol (Kim et al. 2010; Selvin et al. 2012; Woo Lee et al. 2016). The significant activating effect could be attributed to the uniform water phase formed by water-miscible solvents (Ogino and Ishikawa 2001) or the high diffusion rate of substrate in the presence of water-miscible solvents (Metin et al. 2006), which enables substrates quick and easy access to the active site. Moreover, Est2 activity generally showed a bell-shaped dependence on water-miscible organic solvent concentration (Fig. 5b), which indicated an optimal water activity for its hydrolytic activity (Léonard-Nevers et al. 2009; Adlercreutz 2013). Other esterases, including Est1 (Fig. 5a), commonly showed slightly increased (Hotta et al. 2002; Schütte and Fetzner 2007; Faulds et al. 2011; Kang et al. 2017), or decreased (Li and Yu 2013; Monsef Shokri et al. 2014; Dukunde et al. 2017) activity towards water-miscible organic solvents. For these enzymes, water-miscible solvents strip off the crucial water monolayer around the enzyme surface and compete for hydrogen bonds, which at higher solvent concentrations finally leads to denaturation (Ó’Fágáin 2003; Doukyu and Ogino 2010; Monsef Shokri et al. 2014).
In addition, Est1 was stable in the presence of 30% (v/v) water-miscible organic solvents (Fig. 6a). Similar or even lower solvent tolerance was found for most of the reported OST esterases (Doukyu and Ogino 2010; Kang et al. 2011; Brault et al. 2012; Xing et al. 2012; Zhang et al. 2014; Kang et al. 2017; Kumagai et al. 2018). Thus, in comparison, Est2 exhibited superior stability against higher concentrations of the analyzed water-miscible organic solvents (Fig. 6b). Among the rare counterparts, a hyper-thermophilic archaeal esterase (Hotta et al. 2002) and EstB from Alcanivorax dieselolei B-5(T) (Zhang et al. 2014) displayed substantial stability towards certain water-miscible organic solvents at high concentrations. An increase of hydrophobic interactions and hydrogen bonds are essential in enhancing esterase tolerance against water-miscible organic solvents (Song and Rhee 2001; Kawata and Ogino 2009; Park et al. 2012). Est2 also showed considerable tolerance towards 30% (v/v) ethanol, isopropanol, DMSO, methanol, and acetone for up to 26 days (Fig. 6c) and towards 60% (v/v) isopropanol, DMSO, and acetone up to 13 days (Fig. 6d). The preservation of high esterase activity over an extended period has been rarely described previously (Sana et al. 2007; Jin et al. 2012; Li and Yu 2013). This feature could allow to apply Est2 as an immobilized biocatalyst in non-aqueous-based continuous bioprocesses (Sana et al. 2007; Yang et al. 2011).
Overall, both enhanced activity and adequate stability towards water-miscible organic solvents are pre-requisite properties for applications of esterases in organic synthesis (Panda and Gowrishankar 2005; Lopez-Lopez et al. 2014). In non-aqueous media, such enzymes could be exploited for ester synthesis and transesterification reactions and a variety of other chemical reactions (Salihu and Alam 2015; Sood et al. 2016; Sarmah et al. 2018). Thus, Est2 could be advantageous for use in biodiesel production, as the acyl acceptors methanol or ethanol are added in esterase/lipase-catalyzed transesterification reactions (Srimhan et al. 2011; Nasaruddin et al. 2014; Wang et al. 2017). In addition, Est1 and Est2 activities responded differently to the presence of polar protic solvents (methanol, ethanol, isopropanol, and 1-propanol) and polar aprotic solvents (acetone and DMSO), which further illustrates that the nature of the solvent influences esterase activity (Torres and Castro 2004). However, the relationship between the corresponding organic solvent logPow value and esterase activity/stability remains uncertain.
Comparing with water-miscible organic solvents, water-immiscible organic solvents are less deleterious for enzyme-solvent interactions (Rahman et al. 2005). However, most of the reported esterases were similar to Est1 (Table 1), showing a significant decrease of activity or inactivation after the incubation with water-immiscible organic solvents (Schütte and Fetzner 2007; Berlemont et al. 2013; Jin et al. 2012). In contrast, lipolytic enzymes of family I, also known as true lipases, are commonly resistant to water-immiscible organic solvents. In the presence of a water-solvent (hydrophobic) interface, the hydrophobic amino acid residues in the lid/flap region stabilize lipases in a flexible, open conformation (Dandavate et al. 2009; Yang et al. 2011; Kamal et al. 2013). However, the cap domain for most esterases is not intrinsically flexible and does not provide open and closed conformations (Bornscheuer 2002; Gall et al. 2014; Kim 2017). This could be the reason that water-immiscible tolerant esterases are rare. To the best of our knowledge, only Est2 and the esterases EstC23 (Jin et al. 2012), LipBL (Pérez et al. 2012), Lpc53E1 (Selvin et al. 2012), RBest1 (Berlemont et al. 2013), Pf_Est (Mandelli et al. 2016), EST4 (Gao et al. 2016), and LipA9 (Park et al. 2018) show substantial resistance towards certain water-immiscible organic solvents. This feature further expands the application potential of Est2 to synthetic reactions in the presence of water-immiscible solvents (Gao et al. 2016; Sarmah et al. 2018).
Est1 and Est2 are to some extent resistant to metal ions (Table S4), which is an important feature in the bioremediation of environmental waste (Brault et al. 2012). Est1 and Est2 were also active at a salinity range of up to 4 M (Fig. S5), suggesting halotolerance (Jeon et al. 2012). Halotolerant enzymes are desirable in processes in which water activity is low (Delgado-García et al. 2012). In combination with the tolerance of Est1 and Est2 to organic solvents, it can be further confirmed that halotolerance is somehow positively correlated with organic solvent tolerance (Berlemont et al. 2013).
In conclusion, the characterization of Est1 and Est2 revealed that both, especially Est2, exhibit several application-relevant features, such as a broad substrate range, thermostability, halotolerance, and resistance to various organic solvents. In addition, as revealed by second-order rotatable design, Est1 and Est2 were predicted to be stable at a broad crossed range of temperature and pH. To the best of our knowledge, this is the first time to report an esterase, Est2, which simultaneously exhibits a significantly enhanced activity and unprecedented high stability towards water-miscible organic solvents and substantial tolerance towards water-immiscible organic solvents.
References
Adlercreutz P (2013) Immobilisation and application of lipases in organic media. Chem Soc Rev 42:6406–6436. https://doi.org/10.1039/c3cs35446f
Ahmed EH, Raghavendra T, Madamwar D (2010) An alkaline lipase from organic solvent–tolerant Acinetobacter sp. EH28: application for ethyl caprylate synthesis. Bioresour Technol 101:3628–3634. https://doi.org/10.1016/J.BIORTECH.2009.12.107
Alcaide M, Tornés J, Stogios PJ, Xu X, Gertler C, Di Leo R, Bargiela R, Lafraya Á, Guazzaroni M-E, López-Cortés N, Chernikova TN, Golyshina OV, Nechitaylo TY, Plumeier I, Pieper DH, Yakimov MM, Savchenko A, Golyshin PN, Ferrer M (2013) Single residues dictate the co-evolution of dual esterases: MCP hydrolases from the α/β hydrolase family. Biochem J 454:157–166. https://doi.org/10.1042/BJ20130552
Alcaide M, Stogios PJ, Lafraya Á, Tchigvintsev A, Flick R, Bargiela R, Chernikova TN, Reva ON, Hai T, Leggewie CC, Katzke N, La Cono V, Matesanz R, Jebbar M, Jaeger K-E, Yakimov MM, Yakunin AF, Golyshin PN, Golyshina OV, Savchenko A, Ferrer M, MAMBA Consortium (2015) Pressure adaptation is linked to thermal adaptation in salt-saturated marine habitats. Environ Microbiol 17:332–345. https://doi.org/10.1111/1462-2920.12660
Angkawidjaja C, Koga Y, Takano K, Kanaya S (2012) Structure and stability of a thermostable carboxylesterase from the thermoacidophilic archaeon Sulfolobus tokodaii. FEBS J 279:3071–3084. https://doi.org/10.1111/j.1742-4658.2012.08687.x
Antranikian G, Egorova K (2007) Extremophiles, a unique resource of biocatalysts for industrial biotechnology. In: Physiology and biochemistry of extremophiles. American Society of Microbiology, Washington DC, pp 361–406
Arpigny JL, Jaeger K-E (1999) Bacterial lipolytic enzymes: classification and properties. Biochem J 343:177. https://doi.org/10.1042/0264-6021:3430177
Auernik KS, Cooper CR, Kelly RM (2008) Life in hot acid: pathway analyses in extremely thermoacidophilic archaea. Curr Opin Biotechnol 19:445–453. https://doi.org/10.1016/j.copbio.2008.08.001
Benaiges MD, Alarcón M, Fuciños P, Ferrer P, Rua M, Valero F (2010) Recombinant Candida rugosa lipase 2 from Pichia pastoris: immobilization and use as biocatalyst in a stereoselective reaction. Biotechnol Prog 26:1252–1258. https://doi.org/10.1002/btpr.444
Bendtsen JD, Nielsen H, Von Heijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795. https://doi.org/10.1016/j.jmb.2004.05.028
Berlemont R, Spee O, Delsaute M, Lara Y, Schuldes J, Simon C, Power P, Daniel R, Galleni M (2013) Novel organic solvent–tolerant esterase isolated by metagenomics: insights into the lipase/esterase classification. Rev Argent Microbiol 45:3–12
Biver S, Vandenbol M (2013) Characterization of three new carboxylic ester hydrolases isolated by functional screening of a forest soil metagenomic library. J Ind Microbiol Biotechnol 40:191–200. https://doi.org/10.1007/s10295-012-1217-7
Bora L (2014) Purification and characterization of highly alkaline lipase from Bacillus licheniformis MTCC 2465: and study of its detergent compatibility and applicability. J Surfactant Deterg 17:889–898. https://doi.org/10.1007/s11743-013-1517-6
Bornscheuer UT (2002) Microbial carboxyl esterases: classification, properties and application in biocatalysis. FEMS Microbiol Rev 26:73–81. https://doi.org/10.1111/j.1574-6976.2002.tb00599.x
Box GEP, Hunter JS, Hunter WG (2005) Statistics for experimenters: design, innovation and discovery, 2nd edn. Wiley-Interscience, Hoboken
Boyineni J, Kim J, Kang BS, Lee C, Jang SH (2014) Enhanced catalytic site thermal stability of cold-adapted esterase EstK by a W208Y mutation. Biochim Biophys Acta, Proteins Proteomics 1844:1076–1082. https://doi.org/10.1016/j.bbapap.2014.03.009
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brault G, Shareck F, Hurtubise Y, Lépine F, Doucet N (2012) Isolation and characterization of EstC, a new cold-active esterase from Streptomyces coelicolor A3(2). PLoS One 7:e32041. https://doi.org/10.1371/journal.pone.0032041
Bunterngsook B, Kanokratana P, Thongaram T, Tanapongpipat S, Uengwetwanit T, Rachdawong S, Vichitsoonthonkul T, Eurwilaichitr L (2010) Identification and characterization of lipolytic enzymes from a peat-swamp forest soil metagenome. Biosci Biotechnol Biochem 74:1848–1854. https://doi.org/10.1271/bbb.100249
Chu X, He H, Guo C, Sun B (2008) Identification of two novel esterases from a marine metagenomic library derived from South China Sea. Appl Microbiol Biotechnol 80:615–625. https://doi.org/10.1007/s00253-008-1566-3
Cook GM, Rainey FA, Patel BKC, Morgan HW (1996) Characterization of a new obligately anaerobic thermophile, Thermoanaerobacter wiegelii sp. nov. Int J Syst Bacteriol 46:123–127. https://doi.org/10.1099/00207713-46-1-123
D’Auria S, Herman P, Lakowicz J, Bertoli E, Tanfani F, Rossi M, Manco G (2000) The thermophilic esterase from Archaeoglobus fulgidus: structure and conformational dynamics at high temperature. Proteins Struct Funct Genet 38:351–360
Dandavate V, Jinjala J, Keharia H, Madamwar D (2009) Production, partial purification and characterization of organic solvent–tolerant lipase from Burkholderia multivorans V2 and its application for ester synthesis. Bioresour Technol 100:3374–3381. https://doi.org/10.1016/j.biortech.2009.02.011
Daniel R (2005) The metagenomics of soil. Nat Rev Microbiol 3:470–478. https://doi.org/10.1038/nrmicro1160
Delgado-García M, Valdivia-Urdiales B, Aguilar-González CN, Contreras-Esquivel JC, Rodríguez-Herrera R (2012) Halophilic hydrolases as a new tool for the biotechnological industries. J Sci Food Agric 92:2575–2580. https://doi.org/10.1002/jsfa.5860
Dougherty MJ, D’haeseleer P, Hazen TC, Simmons BA, Adams PD, Hadi MZ (2012) Glycoside hydrolases from a targeted compost metagenome, activity screening and functional characterization. BMC Biotechnol 12:38. https://doi.org/10.1186/1472-6750-12-38
Doukyu N, Ogino H (2010) Organic solvent-tolerant enzymes. Biochem Eng J 48:270–282. https://doi.org/10.1016/J.BEJ.2009.09.009
Dukunde A, Schneider D, Lu M, Brady S, Daniel R (2017) A novel, versatile family IV carboxylesterase exhibits high stability and activity in a broad pH spectrum. Biotechnol Lett 39:577–587. https://doi.org/10.1007/s10529-016-2282-1
Ebrahimi M, Lakizadeh A, Agha-Golzadeh P, Ebrahimie E, Ebrahimi M (2011) Prediction of thermostability from amino acid attributes by combination of clustering with attribute weighting: a new vista in engineering enzymes. PLoS One 6:e23146. https://doi.org/10.1371/journal.pone.0023146
Elend C, Schmeisser C, Leggewie C, Babiak P, Steele HL, Reymond J, Jaeger K, Streit R, Carballeira JD, Streit WR (2006) Isolation and biochemical characterization of two novel metagenome-derived esterases. Appl Environ Microbiol 72:3637–3645. https://doi.org/10.1128/AEM.72.5.3637
Faiz O, Colak A, Saglam N, Canakçi S, Beldüz AO (2007) Determination and characterization of thermostable esterolytic activity from a novel thermophilic bacterium Anoxybacillus gonensis A4. J Biochem Mol Biol 40:588–594. https://doi.org/10.5483/BMBRep.2007.40.4.588
Faulds CB, Pérez-Boada M, Martínez ÁT (2011) Influence of organic co-solvents on the activity and substrate specificity of feruloyl esterases. Bioresour Technol 102:4962–4967. https://doi.org/10.1016/j.biortech.2011.01.088
Ferrer M, Bargiela R, Martínez-Martínez M, Mir J, Koch R, Golyshina OV, Golyshin PN (2015) Biodiversity for biocatalysis: a review of the α/β-hydrolase fold superfamily of esterases-lipases discovered in metagenomes. Biocatal Biotransformation 33:235–249. https://doi.org/10.3109/10242422.2016.1151416
Gall MG, Nobili A, Pavlidis IV, Bornscheuer UT (2014) Improved thermostability of a Bacillus subtilis esterase by domain exchange. Appl Microbiol Biotechnol 98:1719–1726. https://doi.org/10.1007/s00253-013-5053-0
Gao W, Wu K, Chen L, Fan H, Zhao Z, Gao B, Wang H, Wei D (2016) A novel esterase from a marine mud metagenomic library for biocatalytic synthesis of short-chain flavor esters. Microb Cell Factories 15:41. https://doi.org/10.1186/s12934-016-0435-5
Georis J, Esteves FDL, Lamotte-Brasseur J, Bougnet V, Giannotta F, Frère J-M, Devreese B, Granier B (2000) An additional aromatic interaction improves the thermostability and thermophilicity of a mesophilic family 11 xylanase: structural basis and molecular study. Protein Sci 9:466–475. https://doi.org/10.1110/ps.9.3.466
Glogauer A, Martini VP, Faoro H, Couto GH, Müller-Santos M, Monteiro RA, Mitchell DA, de Souza EM, Pedrosa FO, Krieger N (2011) Identification and characterization of a new true lipase isolated through metagenomic approach. Microb Cell Factories 10:54. https://doi.org/10.1186/1475-2859-10-54
González-González R, Fuciños P, Rúa ML (2017) An overview on extremophilic esterases. In: Extremophilic enzymatic processing of lignocellulosic feedstocks to bioenergy. Springer International Publishing, Cham, pp 181–204
Hardeman F, Sjoling S (2007) Metagenomic approach for the isolation of a novel low-temperature-active lipase from uncultured bacteria of marine sediment. FEMS Microbiol Ecol 59:524–534. https://doi.org/10.1111/j.1574-6941.2006.00206.x
Hasan F, Shah AA, Hameed A (2006) Industrial applications of microbial lipases. Enzym Microb Technol 39:235–251. https://doi.org/10.1016/J.ENZMICTEC.2005.10.016
Hausmann S, Jaeger K-E (2010) Lipolytic enzymes from bacteria. In: Handbook of Hydrocarbon and Lipid Microbiology. Springer Berlin Heidelberg, Berlin, pp 1099–1126
Hita E, Robles A, Camacho B, González PA, Esteban L, Jiménez MJ, Muñío MM, Molina E (2009) Production of structured triacylglycerols by acidolysis catalyzed by lipases immobilized in a packed bed reactor. Biochem Eng J 46:257–264. https://doi.org/10.1016/J.BEJ.2009.05.015
Hotta Y, Ezaki S, Atomi H, Imanaka T (2002) Extremely stable and versatile carboxylesterase from a hyperthermophilic archaeon. Appl Environ Microbiol 68:3925–3931. https://doi.org/10.1128/AEM.68.8.3925-3931.2002
Hu X, Thumarat U, Zhang X, Tang M, Kawai F (2010) Diversity of polyester-degrading bacteria in compost and molecular analysis of a thermoactive esterase from Thermobifida alba AHK119. Appl Microbiol Biotechnol 87:771–779. https://doi.org/10.1007/s00253-010-2555-x
Hun CJ, Rahman RNZA, Salleh AB, Basri M (2003) A newly isolated organic solvent–tolerant Bacillus sphaericus 205y producing organic solvent-stable lipase. Biochem Eng J 15:147–151. https://doi.org/10.1016/S1369-703X(02)00185-7
Ishikawa J, Hotta K (1999) FramePlot: a new implementation of the frame analysis for predicting protein-coding regions in bacterial DNA with a high G + C content. FEMS Microbiol Lett 174:251–263
Jaeger K-E, Dijkstra BW, Reetz MT (1999) Bacterial biocatalysts: molecular biology, three-dimensional structures, and biotechnological applications of lipases. Annu Rev Microbiol 53:315–351. https://doi.org/10.1146/annurev.micro.53.1.315
Jayanath G, Mohandas SP, Kachiprath B, Solomon S, Sajeevan TP, Bright Singh IS, Philip R (2018) A novel solvent–tolerant esterase of GDSGG motif subfamily from solar saltern through metagenomic approach: recombinant expression and characterization. Int J Biol Macromol 119:393–401. https://doi.org/10.1016/j.ijbiomac.2018.06.057
Jeon JH, Lee HS, Kim JT, Kim SJ, Choi SH, Kang SG, Lee JH (2012) Identification of a new subfamily of salt-tolerant esterases from a metagenomic library of tidal flat sediment. Appl Microbiol Biotechnol 93:623–631. https://doi.org/10.1007/s00253-011-3433-x
Jin P, Pei X, Du P, Yin X, Xiong X, Wu H, Zhou X, Wang Q (2012) Overexpression and characterization of a new organic solvent-tolerant esterase derived from soil metagenomic DNA. Bioresour Technol 116:234–240. https://doi.org/10.1016/j.biortech.2011.10.087
Jochens H, Aerts D, Bornscheuer UT (2010) Thermostabilization of an esterase by alignment-guided focussed directed evolution. Protein Eng Des Sel 23:903–909. https://doi.org/10.1093/protein/gzq071
Kamal Z, Yedavalli P, Deshmukh MV, Rao NM (2013) Lipase in aqueous-polar organic solvents: activity, structure, and stability. Protein Sci 22:904–915. https://doi.org/10.1002/pro.2271
Kamimura ES, Medieta O, Rodrigues MI, Maugeri F (2001) Studies on lipase-affinity adsorption using response surface analysis. Biotechnol Appl Biochem 33:153–159
Kang CH, Oh KH, Lee MH, Oh TK, Kim BH, Yoon JH (2011) A novel family VII esterase with industrial potential from compost metagenomic library. Microb Cell Factories 10:41. https://doi.org/10.1186/1475-2859-10-41
Kang LJ, Meng ZT, Hu C, Zhang Y, Guo HL, Li Q, Li M (2017) Screening, purification, and characterization of a novel organic solvent-tolerant esterase, Lip2, from Monascus purpureus strain M7. Extremophiles 21:345–355. https://doi.org/10.1007/s00792-016-0907-x
Kawata T, Ogino H (2009) Enhancement of the organic solvent-stability of the LST-03 lipase by directed evolution. Biotechnol Prog 25:1605–1611. https://doi.org/10.1002/btpr.264
Kim TD (2017) Bacterial hormone-sensitive lipases (bHSLs): Emerging enzymes for biotechnological applications. J Microbiol Biotechnol 27:1907–1915. https://doi.org/10.4014/jmb.1708.08004
Kim YH, Kwon EJ, Kim SK, Jeong YS, Kim J, Yun HD, Kim H (2010) Molecular cloning and characterization of a novel family VIII alkaline esterase from a compost metagenomic library. Biochem Biophys Res Commun 393:45–49. https://doi.org/10.1016/j.bbrc.2010.01.070
Klibanov AM (2001) Improving enzymes by using them in organic solvents. Nature 409:241–246. https://doi.org/10.1038/35051719
Kovacic F, Mandrysch A, Poojari C, Strodel B, Jaeger K-E (2016) Structural features determining thermal adaptation of esterases. Protein Eng Des Sel 29:65–76. https://doi.org/10.1093/protein/gzv061
Kumagai PS, Gutierrez RF, Lopes JLS, Martins JM, Jameson DM, Castro AM, Martins LF, DeMarco R, Bossolan NRS, Wallace BA, Araujo APU (2018) Characterization of esterase activity from an Acetomicrobium hydrogeniformans enzyme with high structural stability in extreme conditions. Extremophiles 22:781–793. https://doi.org/10.1007/s00792-018-1038-3
Kumar A, Dhar K, Singh Kanwar S, Arora PK (2016) Lipase catalysis in organic solvents: advantages and applications. Biol Proced Online 18:2. https://doi.org/10.1186/s12575-016-0033-2
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Lämmle K, Zipper H, Breuer M, Hauer B, Buta C, Brunner H, Rupp S (2007) Identification of novel enzymes with different hydrolytic activities by metagenome expression cloning. J Biotechnol 127:575–592. https://doi.org/10.1016/j.jbiotec.2006.07.036
Lazić ŽR (2004) Design of experiments in chemical engineering. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim
Léonard-Nevers V, Marton Z, Lamare S, Hult K, Graber M (2009) Understanding water effect on Candida antarctica lipase B activity and enantioselectivity towards secondary alcohols. J Mol Catal B Enzym 59:90–95. https://doi.org/10.1016/J.MOLCATB.2009.01.008
Li X, Yu H (2013) Purification and characterization of an extracellular esterase with organic solvent tolerance from a halotolerant isolate, Salimicrobium sp. LY19. BMC Biotechnol 13:108
Li P, Chen X, Ji P, Li C, Wang P, Zhang Y, Xie B, Qin Q, Su H, Zhou B, Zhang Y, Zhang X (2015) Interdomain hydrophobic interactions modulate the thermostability of microbial esterases from the hormone-sensitive lipase family. J Biol Chem 290:11188–11,198. https://doi.org/10.1074/jbc.M115.646182
Lima VMG, Krieger N, Mitchell DA, Baratti JC, de FI, Fontana JD (2004) Evaluation of the potential for use in biocatalysis of a lipase from a wild strain of Bacillus megaterium. J Mol Catal B Enzym 31:53–61. https://doi.org/10.1016/J.MOLCATB.2004.07.005
Lineweaver H, Burk D (1934) The determination of enzyme dissociation constants. J Am Chem Soc 56:658–666. https://doi.org/10.1021/ja01318a036
Lopez-Lopez O, Cerdan M, Siso M (2014) New extremophilic lipases and esterases from metagenomics. Curr Protein Pept Sci 15:445–455. https://doi.org/10.2174/1389203715666140228153801
Lotti M, Alberghina L (2007) Lipases: molecular structure and function. In: Industrial Enzymes. Springer Netherlands, Dordrecht, pp 263–281
Mandelli F, Goncalves TA, Gandin CA, Oliveira ACP, Oliveira Neto M, Squina FM (2016) Characterization and low-resolution structure of an extremely thermostable esterase of potential biotechnological interest from Pyrococcus furiosus. Mol Biotechnol 58:757–766. https://doi.org/10.1007/s12033-016-9975-5
Martínez-Martínez M, Coscolín C, Santiago G, Chow J, Stogios PJ, Bargiela R, Gertler C, Navarro-Fernández J, Bollinger A, Thies S, Méndez-García C, Popovic A, Brown G, Chernikova TN, García-Moyano A, Bjerga GEK, Pérez-García P, Hai T, Del Pozo M V., Stokke R, Steen IH, Cui H, Xu X, Nocek BP, Alcaide M, Distaso M, Mesa V, Peláez AI, Sánchez J, Buchholz PCF, Pleiss J, Fernández-Guerra A, Glöckner FO, Golyshina O V., Yakimov MM, Savchenko A, Jaeger K-E, Yakunin AF, Streit WR, Golyshin PN, Guallar V, Ferrer M, The INMARE Consortium TI (2018) Determinants and prediction of esterase substrate promiscuity patterns. ACS Chem Biol 13:225–234. doi: https://doi.org/10.1021/acschembio.7b00996
Metin K, Burcu Bakir Ateslier Z, Basbulbul G, Halil Biyik H (2006) Characterization of esterase activity in Geobacillus sp. HBB-4. J Basic Microbiol 46:400–409. https://doi.org/10.1002/jobm.200510121
Mohamed YM, Ghazy MA, Sayed A, Ouf A, El-Dorry H, Siam R (2013) Isolation and characterization of a heavy metal-resistant, thermophilic esterase from a Red Sea Brine Pool. Sci Rep 3:3358. https://doi.org/10.1038/srep03358
Monsef Shokri M, Ahmadian S, Akbari N, Khajeh K (2014) Hydrophobic substitution of surface residues affects lipase stability in organic solvents. Mol Biotechnol 56:360–368. https://doi.org/10.1007/s12033-013-9716-y
Nacke H, Will C, Herzog S, Nowka B, Engelhaupt M, Daniel R (2011) Identification of novel lipolytic genes and gene families by screening of metagenomic libraries derived from soil samples of the German Biodiversity Exploratories. FEMS Microbiol Ecol 78:188–201. https://doi.org/10.1111/j.1574-6941.2011.01088.x
Nasaruddin RR, Alam MZ, Jami MS (2014) Evaluation of solvent system for the enzymatic synthesis of ethanol-based biodiesel from sludge palm oil (SPO). Bioresour Technol 154:155–161. https://doi.org/10.1016/j.biortech.2013.11.095
Nerurkar M, Joshi M, Pariti S, Adivarekar R (2013) Application of lipase from marine bacteria Bacillus sonorensis as an additive in detergent formulation. J Surfactant Deterg 16:435–443. https://doi.org/10.1007/s11743-012-1434-0
Ngo TD, Ryu BH, Ju H, Jang E, Park K, Kim KK, Kim TD (2013) Structural and functional analyses of a bacterial homologue of hormone-sensitive lipase from a metagenomic library. Acta Crystallogr Sect D Biol Crystallogr 69:1726–1737. https://doi.org/10.1107/S0907444913013425
Ngo TD, Ryu BH, Ju H, Jang EJ, Kim KK, Kim TD (2014) Crystallographic analysis and biochemical applications of a novel penicillin-binding protein/β-lactamase homologue from a metagenomic library. Acta Crystallogr Sect D Biol Crystallogr 70:2455–2466. https://doi.org/10.1107/S1399004714015272
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217. https://doi.org/10.1006/JMBI.2000.4042
Ó’Fágáin C (2003) Enzyme stabilization—recent experimental progress. Enzym Microb Technol 33:137–149. https://doi.org/10.1016/S0141-0229(03)00160-1
Ogino H, Ishikawa H (2001) Enzymes which are stable in the presence of organic solvents. J Biosci Bioeng 91:109–116
Ogino H, Mimitsuka T, Muto T, Matsumura M, Yasuda M, Ishimi K, Ishikawa H (2004) Cloning, expression, and characterization of a lipolytic enzyme gene (lip8) from Pseudomonas aeruginosa LST-03. J Mol Microbiol Biotechnol 7:212–223. https://doi.org/10.1159/000079830
Ohlhoff CW, Kirby BM, Van Zyl L, Mutepfa DLR, Casanueva A, Huddy RJ, Bauer R, Cowan DA, Tuffin M (2015) An unusual feruloyl esterase belonging to family VIII esterases and displaying a broad substrate range. J Mol Catal B Enzym 118:79–88. https://doi.org/10.1016/J.MOLCATB.2015.04.010
Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J, Sussman JL, Verschueren KHG, Goldman A (1992) The α/β hydrolase fold. Protein Eng Des Sel 5:197–211. https://doi.org/10.1093/protein/5.3.197
Panda T, Gowrishankar BS (2005) Production and applications of esterases. Appl Microbiol Biotechnol 67:160–169. https://doi.org/10.1007/s00253-004-1840-y
Park HJ, Joo JC, Park K, Yoo YJ (2012) Stabilization of Candida antarctica lipase B in hydrophilic organic solvent by rational design of hydrogen bond. Biotechnol Bioprocess Eng 17:722–728. https://doi.org/10.1007/s12257-012-0092-4
Park SH, Kim S, Park S, Kim HK (2018) Characterization of organic solvent-tolerant lipolytic enzyme from Marinobacter lipolyticus isolated from the Antarctic Ocean. Appl Biochem Biotechnol. https://doi.org/10.1007/s12010-018-2865-5
Peng Q, Zhang X, Shang M, Wang X, Wang G, Li B, Guan G, Li Y, Wang Y (2011) A novel esterase gene cloned from a metagenomic library from neritic sediments of the South China Sea. Microb Cell Factories 10:95. https://doi.org/10.1186/1475-2859-10-95
Pérez D, Kovačic F, Wilhelm S, Jaeger K-E, García MT, Ventosa A, Mellado EN (2012) Identification of amino acids involved in the hydrolytic activity of lipase LipBL from Marinobacter lipolyticus. Microbiology 158:2192–2203. doi: https://doi.org/10.1099/mic.0.058792-0
Petersen EI, Valinger G, Sölkner B, Stubenrauch G, Schwab H (2001) A novel esterase from Burkholderia gladioli which shows high deacetylation activity on cephalosporins is related to beta-lactamases and DD-peptidases. J Biotechnol 89:11–25
Pezzullo M, Del Vecchio P, Mandrich L, Nucci R, Rossi M, Manco G (2013) Comprehensive analysis of surface charged residues involved in thermal stability in Alicyclobacillus acidocaldarius esterase 2. Protein Eng Des Sel 26:47–58. https://doi.org/10.1093/protein/gzs066
Popovic A, Hai T, Tchigvintsev A, Hajighasemi M, Nocek B, Khusnutdinova AN, Brown G, Glinos J, Flick R, Skarina T, Chernikova TN, Yim V, Brüls T, Le PD, Yakimov MM, Joachimiak A, Ferrer M, Golyshina OV, Savchenko A, Golyshin PN, Yakunin AF (2017) Activity screening of environmental metagenomic libraries reveals novel carboxylesterase families. Sci Rep 7:44103. https://doi.org/10.1038/srep44103
Rahman RNZRA, Baharum SN, Basri M, Salleh AB (2005) High-yield purification of an organic solvent-tolerant lipase from Pseudomonas sp. strain S5. Anal Biochem 341:267–274. https://doi.org/10.1016/j.ab.2005.03.006
Raja A, Prabakarana P (2011) Actinomycetes and drug-an overview. Am J Drug Discov Dev 1:75–84. https://doi.org/10.3923/ajdd.2011.75.84
Rashamuse K, Magomani V, Ronneburg T, Brady D (2009) A novel family VIII carboxylesterase derived from a leachate metagenome library exhibits promiscuous β-lactamase activity on nitrocefin. Appl Microbiol Biotechnol 83:491–500. https://doi.org/10.1007/s00253-009-1895-x
Riedel K, Sutherland R, Donato JJ, Liebl W, Leis B, Angelov A, Mientus M, Li H, Pham VTT, Lauinger B, Bongen P, Pietruszka J, Gonçalves LG, Santos H (2015) Identification of novel esterase-active enzymes from hot environments by use of the host bacterium Thermus thermophilus. Front Microbiol 6:27. https://doi.org/10.3389/fmicb.2015.00275
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:320–324. https://doi.org/10.1093/nar/gku316
Romdhane IB-B, Fendri A, Gargouri Y, Gargouri A, Belghith H (2010) A novel thermoactive and alkaline lipase from Talaromyces thermophilus fungus for use in laundry detergents. Biochem Eng J 53:112–120. https://doi.org/10.1016/J.BEJ.2010.10.002
Ryckeboer J, Mergaert J, Vaes K, Klammer S, Clercq D, Coosemans J, Insam H, Swings J (2003) A survey of bacteria and fungi occurring during composting and self-heating processes. Ann Microbiol 53:349–410
Sadeghi M, Naderi-Manesh H, Zarrabi M, Ranjbar B (2006) Effective factors in thermostability of thermophilic proteins. Biophys Chem 119:256–270. https://doi.org/10.1016/j.bpc.2005.09.018
Sakai Y, Ishikawa J, Fukasaka S, Yurimoto H, Mitsui R, Yanase H, Kato N (1999) A new carboxylesterase from Brevibacterium linens IFO 12171 responsible for the conversion of 1,4-butanediol diacrylate to 4-hydroxybutyl acrylate: purification, characterization, gene cloning, and gene expression in Escherichia coli. Biosci Biotechnol Biochem 63:688–697. https://doi.org/10.1271/bbb.63.688
Salihu A, Alam MZ (2015) Solvent tolerant lipases: a review. Process Biochem 50:86–96. https://doi.org/10.1016/j.procbio.2014.10.019
Sana B, Ghosh D, Saha M, Mukherjee J (2007) Purification and characterization of an extremely dimethylsulfoxide tolerant esterase from a salt-tolerant Bacillus species isolated from the marine environment of the Sundarbans. Process Biochem 42:1571–1578. https://doi.org/10.1016/j.procbio.2007.05.026
Sarmah N, Revathi D, Sheelu G, Yamuna Rani K, Sridhar S, Mehtab V, Sumana C (2018) Recent advances on sources and industrial applications of lipases. Biotechnol Prog 34:5–28. https://doi.org/10.1002/btpr.2581
Sayer C, Isupov MN, Bonch-Osmolovskaya E, Littlechild JA (2015a) Structural studies of a thermophilic esterase from a new Planctomycetes species, Thermogutta terrifontis. FEBS J 282:2846–2857. https://doi.org/10.1111/febs.13326
Sayer C, Szabo Z, Isupov MN, Ingham C, Littlechild JA (2015b) The structure of a novel thermophilic esterase from the Planctomycetes species, Thermogutta terrifontis reveals an open active site due to a minimal “Cap” domain. Front Microbiol 6:1294. https://doi.org/10.3389/fmicb.2015.01294
Schütte M, Fetzner S (2007) EstA from Arthrobacter nitroguajacolicus Ru61a, a thermo- and solvent-tolerant carboxylesterase related to class C b-Lactamases. Curr Microbiol 54:230–236. https://doi.org/10.1007/s00284-006-0438-2
Secundo F, Carrea G (2002) Lipase activity and conformation in neat organic solvents. J Mol Catal B Enzym 19:93–102. https://doi.org/10.1016/S1381-1177(02)00155-8
Selvin J, Kennedy J, Lejon DPH, Kiran GS, Dobson ADW (2012) Isolation identification and biochemical characterization of a novel halo-tolerant lipase from the metagenome of the marine sponge Haliclona simulans. Microb Cell Factories 11:72. https://doi.org/10.1186/1475-2859-11-72
Seo S, Lee YS, Yoon SH, Kim SJ, Cho JY, Hahn BS, Koo BS, Lee CM (2014) Characterization of a novel cold-active esterase isolated from swamp sediment metagenome. World J Microbiol Biotechnol 30:879–886. https://doi.org/10.1007/s11274-013-1496-9
Shao H, Xu L, Yan Y (2013) Isolation and characterization of a thermostable esterase from a metagenomic library. J Ind Microbiol Biotechnol 40:1211–1222. https://doi.org/10.1007/s10295-013-1317-z
Sharma R, Soni S, Vohra R, Gupta L, Gupta J (2002) Purification and characterisation of a thermostable alkaline lipase from a new thermophilic Bacillus sp. RSJ-1. Process Biochem 37:1075–1084. https://doi.org/10.1016/S0032-9592(01)00316-8
Shieh C-J, Liao H-F, Lee C-C (2003) Optimization of lipase-catalyzed biodiesel by response surface methodology. Bioresour Technol 88:103–116
Simon C, Daniel R (2011) Metagenomic analyses: past and future trends. Appl Environ Microbiol 77:1153–1161. https://doi.org/10.1128/AEM.02345-10
Song JK, Rhee JS (2001) Enhancement of stability and activity of phospholipase A1 in organic solvents by directed evolution. Biochim Biophys Acta Protein Struct Mol Enzymol 1547:370–378. https://doi.org/10.1016/S0167-4838(01)00204-7
Sood S, Sharma A, Sharma N, Kanwar SS (2016) Carboxylesterases: sources, characterization and broader applications. Insights Enzym Res 1:1. https://doi.org/10.21767/2573-4466.100002
Srimhan P, Kongnum K, Taweerodjanakarn S, Hongpattarakere T (2011) Selection of lipase producing yeasts for methanol-tolerant biocatalyst as whole cell application for palm-oil transesterification. Enzym Microb Technol 48:293–298. https://doi.org/10.1016/j.enzmictec.2010.12.004
Takeda Y, Aono R, Doukyu N (2006) Purification, characterization, and molecular cloning of organic-solvent-tolerant cholesterol esterase from cyclohexane-tolerant Burkholderia cepacia strain ST-200. Extremophiles 10:269–277. https://doi.org/10.1007/s00792-005-0494-8
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tirawongsaroj P, Sriprang R, Harnpicharnchai P, Thongaram T, Champreda V, Tanapongpipat S, Pootanakit K, Eurwilaichitr L (2008) Novel thermophilic and thermostable lipolytic enzymes from a Thailand hot spring metagenomic library. J Biotechnol 133:42–49. https://doi.org/10.1016/J.JBIOTEC.2007.08.046
Torres S, Castro GR (2004) Non-aqueous biocatalysis in homogeneous solvent systems. Crit Rev Biotechnol 42:271–277
Torres S, Martínez MA, Pandey A, Castro GR (2009) An organic-solvent-tolerant esterase from thermophilic Bacillus licheniformis S-86. Bioresour Technol 100:896–902. https://doi.org/10.1016/j.biortech.2008.07.009
Wang X, Qin X, Li D, Yang B, Wang Y (2017) One-step synthesis of high-yield biodiesel from waste cooking oils by a novel and highly methanol-tolerant immobilized lipase. Bioresour Technol 235:18–24. https://doi.org/10.1016/J.BIORTECH.2017.03.086
Woo Lee H, Kyeong Jung W, Ho Kim Y, Han Ryu B, Doohun Kim T, Kim J, Kim H (2016) Characterization of a novel alkaline family VIII esterase with s-enantiomer preference from a compost metagenomic library. J Microbiol Biotechnol 26:315–325. https://doi.org/10.4014/jmb.1509.09081
Wu G, Zhang S, Zhang H, Zhang S, Liu Z (2013) A novel esterase from a psychrotrophic bacterium Psychrobacter celer 3Pb1 showed cold-adaptation and salt tolerance. J Mol Catal B Enzym 98:119–126. https://doi.org/10.1016/j.molcatb.2013.10.012
Xing MN, Zhang XZ, Huang H (2012) Application of metagenomic techniques in mining enzymes from microbial communities for biofuel synthesis. Biotechnol Adv 30:920–929. https://doi.org/10.1016/j.biotechadv.2012.01.021
Yang Y, Yu Y, Zhang Y, Liu C, Shi W, Li Q (2011) Lipase/esterase-catalyzed ring-opening polymerization: a green polyester synthesis technique. Process Biochem 46:1900–1908. https://doi.org/10.1016/j.procbio.2011.07.016
Yang X, Wu L, Xu Y, Ke C, Hu F, Xiao X, Huang J (2018) Identification and characterization of a novel alkalistable and salt-tolerant esterase from the deep-sea hydrothermal vent of the East Pacific Rise. Microbiology 7:e00601. https://doi.org/10.1002/mbo3.601
Ye J, McGinnis S, Madden TL (2006) BLAST: improvements for better sequence analysis. Nucleic Acids Res 34:W6–W9. https://doi.org/10.1093/nar/gkl164
Yu S, Zheng B, Zhao X, Feng Y (2010) Gene cloning and characterization of a novel thermophilic esterase from Fervidobacterium nodosum Rt17-B1. Acta Biochim Biophys Sin Shanghai 42:288–295. https://doi.org/10.1093/abbs/gmq020
Yu EY, Kwon MA, Lee M, Oh JY, Choi JE, Lee JY, Song BK, Hahm DH, Song JK (2011) Isolation and characterization of cold-active family VIII esterases from an arctic soil metagenome. Appl Microbiol Biotechnol 90:573–581. https://doi.org/10.1007/s00253-011-3132-7
Zarafeta D, Moschidi D, Ladoukakis E, Gavrilov S, Chrysina ED, Chatziioannou A, Kublanov I, Skretas G, Kolisis FN (2016) Metagenomic mining for thermostable esterolytic enzymes uncovers a new family of bacterial esterases. Sci Rep 6:38886. https://doi.org/10.1038/srep38886
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 23:40. https://doi.org/10.1186/1471-2105-9-40
Zhang S, Wu G, Liu Z, Shao Z, Liu Z (2014) Characterization of EstB, a novel cold-active and organic solvent-tolerant esterase from marine microorganism Alcanivorax dieselolei B-5(T). Extremophiles 18:251–259. https://doi.org/10.1007/s00792-013-0612-y
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Acknowledgments
We thank Dr Silja Brady and Sarah Zachmann for providing assistance. We acknowledge the support of Mingji Lu by “Erasmus Mundus Action 2 – Lotus I Project.”
Author information
Authors and Affiliations
Contributions
The manuscript was written through the contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lu, M., Dukunde, A. & Daniel, R. Biochemical profiles of two thermostable and organic solvent–tolerant esterases derived from a compost metagenome. Appl Microbiol Biotechnol 103, 3421–3437 (2019). https://doi.org/10.1007/s00253-019-09695-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09695-1