Abstract
Helicobacter pylori bacteria are involved in gastroduodenal disorders, including gastric adenocarcinoma. Since the current therapies encounter with some significant shortcomings, much attention has been paid to the development of new alternative diagnostic and treatment modalities such as immunomedicines to target H. pylori. Having used phage display technology, we isolated fully humane small antibody (Ab) fragment (VL) against the Flap region of urease enzyme of H. pylori to suppress its enzymatic activity. Solution biopanning (SPB) and screening process against a customized biotinylated peptide corresponding to the enzyme Flap region resulted in the selection of VL single domain Abs confirmed by the enzyme-linked immunosorbent assay (ELISA), sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and Western blotting. The selected Ab fragments showed a high affinity with a KD value of 97.8 × 10−9 and specificity to the enzyme with high inhibitory impact. For the first time, a VL single domain Ab was isolated by SPB process against a critical segment of H. pylori urease using a diverse semi-synthetic library. Based on our findings, the selected VL Ab fragments can be used for the diagnosis, imaging, targeting, and/or immunotherapy of H. pylori. Further, Flap region shows great potential for vaccine therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Helicobacter pylori, a gram-negative spiral bacterium, is recognized as a major health threat worldwide. It is estimated that H. pylori bacteria are colonized in more than half of the world’s population (Maleki Kakelar et al. 2018). The World Health Organization (WHO) has classified H. pylori as a group 1 carcinogen because its infection initiates several inflammatory states in the gastrointestinal tract (e.g., superficial gastritis, chronic atrophic gastritis, peptic ulceration), leading to the mucosa-associated lymphoma and gastric cancer (Blaser and Berg 2001). In addition, recent studies have revealed the association of H. pylori infection with other irrelevant disorders, including cardiovascular, neurologic, ocular, and dermatological diseases (Ardekani et al. 2013; Izzotti et al. 2009; Lee et al. 2001).
The currently used diagnostic methods and antibiotic combination therapies often fail to be fully efficient in controlling the H. pylori infection, in large part because of high cost, increasing antibiotic resistance, and high rate of infection recurrence (Michetti 1997; Telford and Ghiara 1996). Therefore, the development of efficient prophylactic vaccines or immunotherapies might provide promising alternative therapies (Nakayama and Graham 2004). Further, monoclonal antibody (mAb)-based immunotherapy has been introduced against the H. pylori infection. In this respect, several studies have been conducted to produce specific Abs against both the whole cell and the well-known virulence factors of H. pylori, including the cag pathogenicity island (cag PAI), vacuolating toxin (VacA), motility, and the urease enzyme (Reiche et al. 2002; Schmausser et al. 2002). Of note, the mucosal sIgA Abs, specific to the urease, were found in 50% of the patients with H. pylori gastritis (Schmausser et al. 2002). Furthermore, due to the importance of urease in neutralizing highly acidic stomach environment and also the colonization of urease enzyme, it can be considered as a specific molecular marker to be targeted. So far, several research groups have selected different formats of neutralizing Abs (i.e., full-length Abs, nanobodies, and scFvs) to target the whole urease enzyme with neutralizing capability (Ardekani et al. 2013; Houimel et al. 2001; Ko et al. 1997; Li et al. 2008; Nagata et al. 1992; Reiche et al. 2002).
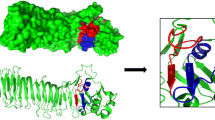
H. pylori urease enzyme consists of two subunits of β (61.2 kDa) and α (26.5 kDa). The whole active enzyme is a dodecameric macromolecule that is assembled from three units of the α- and β-subunits forming a heterotrimeric (αβ)3 molecular structure. Then, four of the (αβ)3 units form a spherical and dodecameric are assembled (1100 kDa) forming a huge internal hollow. The supramolecular enzyme has 12 separated active sites on the UreB subunit (i.e., the large subunit of H. pylori urease), which could protect each other from the acidic inactivation by spreading a neutral layer with ammonia produced from the urea hydrolysis over the molecular and bacterial surfaces. Superposition of the uninhibited and acetohydroxamic acid (AHA)-inhibited H. pylori urease structure have revealed a mobile region consisting of the 313–346 amino acid (aa) residues forming a helix-turn-helix motif at the entrance of the active site cavity—so-called Flap (Ha et al. 2001). Several researchers have reported that the Flap motion possibly contributes to the establishment of the catalytic site, whose active site disappears in the open status of the Flap (Ha et al. 2001; Hirota et al. 2001b). Thus, the Flap region can be a promising candidate for the targeted therapy using mAbs/Ab fragments.
We have previously developed scFv Ab fragments and successfully targeted the VacA toxin of H. pylori (Fahimi et al. 2018). In the present study, having considered the proven importance of Flap region in the activity of the urease enzyme, we aimed to isolate and characterize Ab fragments against the most critical segment of urease enzyme by means of a human semi-synthetic phage antibody library (PAL), Tomlinson human library I+J. To this end, human single-fold scFv library (Tomlinson I) was panned against the customized biotinylated peptide corresponding to the Flap region of the enzyme through a solution-phase biopanning (SPB) process, as reported previously (Abdolalizadeh et al. 2013; Tohidkia et al. 2013; Tohidkia et al. 2017). The selected Ab fragments were then characterized for binding affinity to the whole urease enzyme by means of Western blotting and bioinformatic analysis to analyze the potential of the VL Ab fragment in neutralizing H. pylori urease enzyme.
Methods and materials
Peptide design
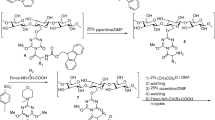
The Flap region of urease enzyme (i.e., from 313 to 346 amino acid residues) was chosen as the target for the selection of specific Ab fragments from the semi-synthetic Tomlinson library I using SPB process. The structural screening of the urease enzyme was performed using PDB viewer software to ensure the accessibility of the Flap segment. Then, the desired peptide sequence was customized by BIOMATIK (Cambridge, Ontario, Canada) with the purity of 98% while the amino acid residues were confirmed using high-performance liquid chromatography (HPLC). The amino acid sequences of the target peptide (EHMDMLMVCHHLDKSIKEDVQFADSRIRPQTIAA-COOH) were modified with biotin and mini-polyethylene glycol (PEG) at the N-terminal of the peptide.
Bacterial strains and library preparation
Tomlinson human scFv library I, TG1 Tr and HB2151 Escherichia coli strains used to produce respectively the scFv phage and the soluble scFv fragments, and KM13 helper phage were purchased from Source Bioscience (Nottingham, UK). The diversity of the library was estimated to be approximately 1.4 × 108 transformants. This phage library was constructed by the side chain diversifications of the complementarity-determining regions (CDRs) (i.e., CDR2 and CDR3, while CDR1 was kept constant) on a single framework from VH (i.e., V3–23/DP-47 and JH4b) and VL (i.e., O12/O2/DPK9 and Jj1). The Ab fragments diversified were fused to g3p in phagemid vector, pIT2, with histidine (His) and Myc tags.
Solution-phase biopanning
About 3 × 1010 colony-forming units (CFUs) of the recombinant phages displaying scFvs were rescued according to the supplier’s protocol as described previously (Hawkins et al. 1992; Henderikx et al. 1998). Briefly, blocking of the recombinant phages and Dynabeads MyOne Streptavidin T1 (Invitrogen, Oslo, Norway) process were separately fulfilled using phosphate-buffered saline (PBS) containing 0.05% Tween 20 (Sigma-Aldrich, Steinheim, Germany) and 3% bovine serum albumin (BSA) (Sigma-Aldrich, Taufkirchen, Germany) at room temperature (RT) for 30 min. The biotinylated peptide (at the designated regressive concentrations of 200 to 50 nM) was respectively used for the selection rounds 1 to 5 by incubation with the blocked phages at RT for 120 min. To capture the peptide-bound phage scFvs, the blocked Dynabeads was added to the phage-peptide mixture and incubated at RT for 20 min. A magnetic particle concentrator, DynaMag-2 (Invitrogen, Karlsruhe, Germany), was used to separate the peptide-bound phage scFvs from the non-bound phage scFvs (non-specific binder) by washing with PBST (PBS-0.1% Tween 20). The peptide-bound phages were eluted with 0.5 mL of bovine pancreas trypsin (1 mg/mL) (Sigma-Aldrich Co., Taufkirchen, Germany) at RT for 10 min. The eluted phages were incubated with the exponential-grown E. coli TG1 at 37 °C for 30 min to amplify specific phage binders for the next round of the selection. To deplete streptavidin phage binders, before each panning process, the blocked phages were incubated with the blocked Dynabeads streptavidin at RT for 1 h.
Polyclonal phage enzyme-linked immunosorbent assay
Coating of the enzyme-linked immunosorbent assay (ELISA) plate with the biotinylated peptide was indirectly performed through precoated biotinylated BSA and streptavidin on polystyrene 96-well plates as described previously (de Haard et al. 1999). Both the positive and the negative plates were coated with 100 μL/well of ImmunoPure biotinylated BSA (Sigma-Aldrich Co., Taufkirchen, Germany) at a final concentration of 2 μg/mL in PBS at 4 °C overnight. After washing with PBST, 100 μL/well of 1 mg/mL streptavidin (S888, Invitrogen, Karlsruhe, Germany) was dissolved in PBS containing 0.5% gelatin and added to the plates and kept at RT for 1 h. All incubation steps were followed by three times washing for 5 min with PBST. The positive and negative plates were incubated, respectively, with 100 μL/well of the biotinylated peptide and mini-PEG (200 nM) at RT for 1.5 h, and then blocked with 200 μL/well of 2% skim milk-PBS (M-PBS) at RT for 2 h. The plates were incubated with the polyclonal phages (10 μL) resulting from each round of the selection combined with 2% M-PBS (100 μL) at RT for 1 h.
The specific phage binders to the peptide were detected by incubating the plates at RT for 90 min with the primary anti-M13 mAb (GE Healthcare, Amersham, UK) and the secondary goat anti-mouse IgG conjugated with HRP (ab6789, Abcam, San Francisco, CA, USA) at dilutions of 1:3000 and 1:5000 in 2% M-PBS, respectively. After adding tetramethylbenzidine (TMB), peroxidase substrate solution, the enzyme reaction was stopped by the addition of 5% H2SO4. Finally, the optical density (OD) was read at OD450–OD650 (Emadi et al. 2007) by ELx808 Absorbance Microplate Reader (BioTek, Winooski, VT, USA).
Soluble antibody fragment ELISA
To screen specific Ab clones, soluble Ab fragments were produced in 96-well microplates as described previously by Marks et al. (1991) with slight modifications. Briefly, 200 μL of the non-suppressor strain HB2151 in the exponential growth phase (OD600 = 0.4) was infected with the phages eluted from the round 4 and incubated at 37 °C for 45 min. Serial dilutions of the infected HB2151 were cultured on TYE/ampicillin/glucose media (TYE containing 100 μg/mL ampicillin and 2% glucose) and incubated at 37 °C overnight. The single colonies were picked and cultured in 100 μL of 2xTY/Amp/Glc 2% and incubated at 37 °C overnight under gentle shaking. Then, 5 μL of the overnight culture was inoculated to a new plate containing 200 μL 2xTY/Amp/Glc 0.1% and incubated at 37 °C under shaking until the OD600 reached 0.9 (approximately 180 min). The expression of Ab fragment was induced by adding 25 μL/well of 2xTY containing 9 mM isopropyl β-D-thiogalactopyranoside (IPTG) and 0.9 M sucrose to a final concentration of 1 mM and 100 mM, respectively. Then, the culture plates were incubated at 30 °C overnight under shaking. To facilitate the release of the periplasmic Ab fragments into the culture supernatant, the culture plates were incubated with periplasmic extraction buffer consisting of 60% (w/v) sucrose, 150 mM Tris, and 3 mM ethylenediaminetetraacetic acid (EDTA), pH 8.0 at 4 °C for 30 min. Following centrifugation of the microplate at 2000×g at 4 °C for 15 min, the culture supernatant containing scFvs in M-PBS was incubated with the peptide-coated ELISA plates at RT for 90 min. The peptide coating and detection processes were performed similarly to the aforementioned method for the polyclonal phage ELISA, in which the HRP protein A (Invitrogen, Karlsruhe, Germany) was used at a dilution of 1:5000 instead of anti-M13 mAb.
Sequence analysis
Double-stranded plasmids of each positive colony were extracted by QIAprep Spin Miniprep Kit. Polymerase chain reaction (PCR) and gel electrophoresis were also exploited to confirm the presence of scFv genes within the plasmid using LMB3 (5′-CTA TGC GGC CCC ATT CA-3′) and pHEN (5′-CAG GAA ACA GCT ATG AC-3′) primers as forward and reverse primers, respectively. Sequencing was performed using ABI3730XL DNA sequencer (Applied Biosystems, Darmstadt, Germany) using LMB3 primer. The nucleotide sequence of the VL antibody gene was submitted to the DNA Data Bank of Japan (accession number LC375193). The sequencing data were analyzed by Chromas 2.4.4, and the alignment of CDRs was done by VBASE2 database to distinguish the different Ab clones.
Large-scale periplasmic expression of VL
Antibody clones with different CDRs were picked for the expression in 400-mL growth medium. Next, 4 mL of the overnight culture was inoculated into 400 mL 2xTY/Amp/Glc 2% and incubated for 180 min until reaching OD600 of 0.9. After centrifugation at 1800×g for 10 min, the culture medium was changed to 400-mL induction media [2xTY/amp supplemented with 1 mM IPTG] and incubated at 37 °C for 4 h under shaking. To obtain the periplasmic-expressed scFvs, the culture medium was centrifuged at 2800×g for 20 min, and then, the bacterial pellets were re-suspended in 4% culture volume of ice-cold TES buffer (Tris 50 mM-sucrose 20% and 1 mM EDTA, pH 8) while incubating on the ice for 1 h. Subsequent to the centrifugation at 20,000×g for 30 min, the supernatant containing the extracted scFv was retrieved for the next steps (Tohidkia et al. 2012).
Purification by affinity chromatography
Before conducting the purification process, the extracted Ab fragments were dialyzed against PBS buffer using a 12-kDa cutoff membrane (D9652, Sigma-Aldrich Co., Taufkirchen, Germany) at 4 °C for 24 h. Then, TALON® immobilized metal affinity chromatography (IMAC) resin (Clontech Laboratories, Inc., Mountain View, CA, USA) was used to purify the scFvs following the manufacturer’s protocol. Briefly, 1 mL of slurry TALON® resin was equilibrated with 5-bed volumes of PBS, pH 7.2, and then incubated with the dialyzed samples on a rotator at RT for 20 min. After centrifugation at 700×g for 2 min, the resin pellet was washed (×2) by incubating with 10-bed volumes of PBS for 10 min. After the centrifugation, the resin-bound VL was eluted by 5-bed volumes of imidazole buffer and collected separately in five tubes.
SDS-PAGE and Western blotting
The SDS-PAGE and Western blotting were performed to ensure the accuracy of expression and purification processes. The induced and un-induced periplasmic extracts of the desired scFv samples were run on two separate 12% SDS-polyacrylamide gels under reducing condition using a Mini-PROTEAN Tetra Cell system (Bio-Rad, Munich, Germany). Coomassie Brilliant Blue, R-250, was used to stain one of the gels, and the other one was blotted onto nitrocellulose membrane (10 V, 20 min) using semi-dry trans-blot system (Bio-Rad, Munich, Germany). After blocking with MPBS 5% at 4 °C overnight, the membrane was incubated with the primary anti-c-Myc tag antibody (Santa Cruz Biotechnology, Dallas, USA) and the secondary anti-HRP antibody by gentle shaking for 90 min at the dilutions of 1:1000 and 1:8000, respectively. After all the incubation steps, the membranes were washed (×3) with the PBST for 5 min. Finally, they were visualized by the enhanced chemiluminescence Western blotting kit (GE Healthcare, Amersham, UK) on the X-ray film.
Surface plasmon resonance analysis
To study the biomolecular interaction(s) and the kinetic parameters in a real-time manner, we capitalized on a multi-parameter surface plasmon resonance (SPR) instrument (MP-SPR Navi 210A, BioNavis Ltd., Tampere, Finland) and streptavidin (SA) sensor provided from BioNavis company. Briefly, the gold slide was applied in both channels at 670-nm wavelength and the temperature of 28 °C by means of PBS running buffer. Approximately 10 μg/mL diluted biotinylated peptide was injected in the flow cell 1 (Fc1) for 7 min (at a flow rate of 30 μL/min), which resulted in the immobilization of 100 response units (RU) for this capture method. The Fc2, with no peptide injection, was designated as the reference channel. To verify the binding affinities, each Ab fragment was injected above the reference and peptide surfaces at five concentration ratios ranged from 100 to 1000 nM. Data analysis of all the measurements and calculation of the kinetic parameters was carried out by means of Bionavis Data Viewer and Trace Drawer software.
Immunoblotting with soluble VL
A volume of 5 μL recombinant UreB (1 mg/mL), HPM-5030-5, and a concentrated fraction of bacterial cells disrupted by the sonication were run on 10% SDS-polyacrylamide gel under reducing condition. Once blotted onto a nitrocellulose membrane, the membrane was prepared to be incubated with the purified soluble VL. After blocking with MPBS 5%, the membrane was incubated with purified VL (20 μg/mL) for 2 h. The detection and visualization processes were done as mentioned above.
The cultivation of H. pylori and urease enzyme concentration
Donated fresh and positive urease biopsy samples of the gastric patients from Imam Reza Hospital (Tabriz, Iran) were cultured on Colombia blood agar base containing 7% fresh defibrinated sheep blood, amphotericin B (7 μg/mL), vancomycin (10 μg/mL), and trimethoprim (6 μg/mL) under microaerophilic conditions (i.e., 10% CO2, 5% O2, 85% N2) at 37 °C for 5–6 days. Afterward, these positive strains were subcultured using brain-heart infusion medium (BHI) supplemented with 10% fetal bovine serum (FBS) and antibiotic to be amplified (Blanchard and Nedrud 2012). The agar plates were then scraped, and also broth flasks were centrifuged at 2800×g to be prepared for the urease extraction using distilled water and sonication. Disrupted cells were centrifuged at 3300×g, and the supernatant containing urease enzyme was concentrated by Vivaspin®20 (50-kDa kit, GE Healthcare, Amersham, UK). The identity of H. pylori was confirmed by VacA and urease enzyme immunoblotting assay, fast urease, and gram staining test.
Analysis of urease inhibitory effect
The H. pylori urease inhibition test was carried out using a standard protocol with slight modifications. In brief, 25 μL of concentrated H. pylori urease was incubated with 50 μL of VL (equivalent to 0 to 80 μg/well) in 96 microwell at 4 °C overnight. Afterward, 75 μL of 50 mM phosphate buffer (pH 6.8, containing 500 mM urea, 0.02% phenol red) was added to each well. Then, 5 × 109 VL phages were added with the same number of helper phage into two separate wells as the negative controls. The color development was determined at 550 nm. The inhibition percentage was calculated by the following equation during 3-h incubation period at 23 °C: [(activity without Ab _ activity with Ab) / (activity without Ab)] × 100, as reported previously (Mobley et al. 1988).
Bioinformatic workflow
Structural modeling and refinement
The 3D homology modeling of VL was accomplished by the Iterative Threading ASSEmbly Refinement (I-TASSER) from Zhang server ranked as one of the top-ranked servers for the prediction of protein structure (Roy et al. 2010; Yang et al. 2015; Yang and Zhang 2015; Zhang 2008). Further, the secondary structure of the VL was predicted using PSSpred (Yan et al. 2013). The modeling process in the I-TASSER starts with the identification of templates by the LOMETS from the PDB library. The LOMETS is a meta-server threading approach containing multiple threading programs, in which each threading program can generate lots of template alignments (Wu and Zhang 2007). In this work, the 10 top-ranked templates were selected from the LOMETS threading programs. Usually, one template with the highest Z score is selected from each threading program, in which the threading programs are sorted by the average performance in large-scale benchmark test experiments. For each target, the I-TASSER simulation process leads to a generation of a large ensemble of structural conformations (so-called decoys). In the next step of selecting the final top-ranked model, the I-TASSER exploits the SPICKER program to cluster all the decoys based on the pair-wise structure similarity (Zhang and Skolnick 2004). After selecting the best model with consideration of the C score, the overall refinement process was performed using Princeton TIGRESS server (Khoury et al. 2014).
Assessment of the model quality and specifications
Frustratometer was used to get insight into the landscape of distribution of energy in a refined model that helps to an understanding about the molecule sites that have the higher level of energy and serves important roles such as the interaction with other molecules and active sites (Parra et al. 2016). The simulated refined structure was evaluated by the most known benchmarking tools, including Verify3D (Luthy et al. 1992), ProsA (Wiederstein and Sippl 2007), and ERRAT2 (Colovos and Yeates 1993). The CASTp was used to specify the packets and cavities on the surface of the antibody (Dundas et al. 2006). Furthermore, Ramachandran plot analysis was performed by RAPPER server to deepen our understanding of the structural details of the modeled VL (http://mordred.bioc.cam.ac.uk/~rapper/rampage.php).
Ligands and binding site prediction and in silico docking
To characterize the potential ligands along with defining the biological annotations of the target protein and ligand binding site residues of ligand peptides on the target VL, COACH as a meta-server approach to predict ligand binding site was applied. In the next step, the UreB large subunit of H. pylori (PDB accession number 1e9z) was retrieved from the PDB. Then, the docking of VL and UreB was done using the ClusPro server that is a protein-protein docking server (Kozakov et al. 2017; Porter et al. 2017). The visualization of PDB files and screening for finding any interactions between CDR regions of VL and UreB were carried out using YASARA and PyMol (Krieger and Vriend 2014; Schrodinger 2015).
Results
Biopanning
The polyclonal phage ELSA was conducted to monitor the successful enrichment of specific phage binders during the biopanning process. The PEG-precipitated phage scFvs amplified from each round of the selections were incubated with both the biotinylated peptide and mini-PEG-coated wells, respectively, as the positive and negative controls to eliminate the signals related to BSA, streptavidin, or mini-PEG phage binders. As shown in Fig. 1a, the peptide-specific phages were enriched after the five consecutive rounds of selection. Unlike rounds 1 and 2, the enrichment of peptide-specific phages in the rounds 3 and 4 was significantly promoted without any notable non-specific signals. In contrary to the round 1, increasing 20-fold signals in round 4 indicated successful accomplishment of the biopanning process.
The biopanning and screening processes. a For the biopanning process, polyclonal phages amplified after each round of panning were incubated with positive (the biotinylated peptide coated) and negative (mini-PEG coated) wells, detected with HRP protein A, and the absorbance was measured at 450 nm subtracted from 630 nm as the reference absorbance. b The specific antibody clone screening. Individual bacterial clones producing soluble antibody fragments were screened for the specific binding to the biotinylated peptide using ELISA assay. The culture supernatants containing antibody fragments were incubated with the positive (i.e., the biotinylated peptide-coated) and the negative (mini-PEG-coated) wells. The clones showing positive to negative ratio more than 2-fold were considered as the positive hit rates
Screening for specific antibody clones
To identify the specific peptide-binding antibody clones, the polyclonal phages obtained from round 4 of the selection were used to infect the non-suppressor HB2151 E. coli producing soluble antibody fragments without g3p. The supernatants containing soluble Ab fragments from each cultured clone in a 96-well microplate were incubated with both the positive- (i.e., the coated peptide) and negative-background (i.e., the coated mini-PEG) ELISA plates. As shown in Fig. 1b, having considered the positive/negative signal ratios over 10-fold, screening results revealed three clones (3%), designated as H2, C3, and A5, which were able to recognize the peptide.
Antibody sequence analysis
To determine full-length insert and sequence diversity, PCR and sequencing were applied to the positive hit clones. The PCR and gel electrophoresis results showed that all the positive clones harbor a partial insert of 635 base-pair nucleotides (DNA Data Bank of Japan, accession number LC375193, Supplementary Material). Nucleotide data obtained from the sequencing were aligned in the VBASE2 database. The CDR comparison revealed that three selected antibody clones were single domain antibody, VL, having identical amino acid sequences.
Expression and purification analysis
The VL antibody C3 was subjected to express in a large scale to produce adequate amounts of reagents for the post panning processes. The expression and purification process of the antibody was confirmed by the SDS-PAGE and Western blotting. The expression results showed a clear protein band at approximately 18 kDa in the induced sample, equivalent to the expected molecular weight of the VL domain antibody (Fig. 2). Further, the SDS-PAGE and Western blotting analyses were performed to verify the accuracy of purification. As shown in Fig. 2, an intense protein band corresponding to the predicted molecular weight of the VL domain antibody was purified with high purity (over 90%), in two elution fractions. Overall, the expression yield of VL antibody C3 was found to be about 1 mg per 1 l of the culture.
The authenticity of expression and purification procedures was confirmed by 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and Western blotting assay by using of anti-C-myc antibody. Expression analysis of C3 by a Coomassi Brilliant Blue staining and b Western blotting assay was certified by clear protein band at approximately 18 kDa. Purification analysis of VL by c SDS-PAGE and d Western blotting. Lane UN, uninduced preplasmic fraction lane IN indicate enhancement of VL expression after addition of 1 mM IPTG, lane F; final fraction, sample of periplasmic fractions after 4 h of induction, lane Flow indicate flow-through fraction lane W washed fraction lane E elution fraction
Surface plasmon resonance analysis
The multi-parameter SPR instrument together with streptavidin (SA) sensor was conducted to study the binding kinetics of the selected Ab fragments. The SPR analysis resulted in binding affinity at a nanomolar range with a KD value of 97.8 × 10−9 M for C3 clone as shown in Table 1 and Fig. S1 (Supplementary Material).
Immunoblotting analysis by purified VL antibody
Determining the specificity of purified VL antibody to urease large subunit (UreB) was performed by running 5 μg recombinant UreB and concentrated cell lysate fraction UreB on 10% acrylamide gel and immunoblotting. As shown in Fig. 3, the immunoblotting results indicated that our VL antibody recognized both recombinant and cell lysate fraction with the molecular weight of approximately 62 kDa even greater than commercial full-length antibody as the positive controls.
Immunoblotting analysis was performed to determine specificity of our VL antibody on urease large subunit (UreB). Lane A, recombinant UreB was detected by commercial antiurease monoclonal antibody (positive control); lane B, recombinant UreB was detected by our C3 VL antibody; lanes C and D, detection of extracted UreB from wild-type H. pylori by commercial mAb (positive control) and C3 VL antibody, respectively. Lane NC represents the negative control. E. coli was used as NC since bacteria do not contain the urease enzyme
Inhibitory effect of VL antibody
Serial dilutions of VL purified antibody were incubated in duplicate with concentrated urease extraction of fresh H. pylori culture. After the addition of substrate solution buffer containing urea and appropriate a pH indicator, the spectrophotometry analysis indicated that VL antibody is able to inhibit the activity of urease up to 40% at 80 μg/well (Fig. 4). The VL-phage complex inhibitory effect was estimated to be over 60%, while the negative controls of each experiment carried out by the soluble VL and helper phage did not show a notable response.
a Dose-dependent response curves of H. pylori urease activity inhibition by isolated single domain VL antibody (C3). The pre-extracted and concentrated urease fraction from wild-type H. pylori was treated overnight with various concentrations of C3 (0 to 80 μg/well) at 4 °C on microplates. After addition of appropriate substrate mixture, the color development was measured at 550 nm. b Inhibitory effect of VL-phage complex in comparison with helper phage, as negative control, is high significantly. Points indicate averages for duplicate determinations
Alignment and prediction of the secondary and 3D structure
In the first step of the modeling and evaluation of bioinformatics pipeline, LOMETS analysis resulted in ten top rank templates selected from the LOMETS threading programs (Fig. 5a). In the next step, PSSpred predicted the secondary structure of VL (Fig. 5b). As previously mentioned in “Materials and methods” section, the I-TASSER through the SPICKER program clusters decoys based on the structural similarity and reports five models that have in concordance with the five argest structure clusters. The quality of models was quantitatively analyzed by the C score which is a statistical value calculated based on the significance of threading template alignments and the convergence parameters of the structure assembly simulations. The C score range is [− 5, 2], and the higher value is in concordance with the higher quality of the model. Thus, the C score of the model from the biggest cluster was about − 0.74 (Fig. 5c, d).
Alignment and structural visualization analyses. a Ten templates was aligned by I-TASSER, showing Norm.Z score > 1. b The predicted secondary structure of VL by PSSpred. The higher score, the more confident of predicted secondary structure. c The 3D structure of refined VL, in which the yellow region represents His tag; also, the blue, green, red, and magenta regions represent Mys tag, VL, CDRs, and linker regions, respectively. d The surface view of the protein and CDR regions. Pymol was used for the visualization
Refinement and assessment of the quality of the model
After selecting the best model, the Princeton TIGRESS server was used to refine the model. Comparison of the models before and after the refinement processes was done using verify3D and ERRAT2. The ERRAT2 overall quality of the model was respectively about 90.98 and 97.25 before and after refinement of the model. Further, the Verify3D score showed 70.27% for the model before the refinement and 87.84% after the refinement process, which indicates a significant improvement in the 3D structure and limitation of errors and clashes (Table 2). The torsion angle, as an important local structure parameter that has a pivotal role in the folding of proteins, was calculated using the Ramachandran plot analysis. The results indicated an acceptable enhancement in the model quality after the refinement. The statistical results of the Ramachandran plot are presented in Fig. S2A (Supplementary Material). Further, three residues were found in the outlier region (i.e., Ser 47, His132 in His Tag region, and Gly135 in Mys tag region), while the Verify3D and ERRAT2 analyses resulted in an acceptable 3D structural model.
In the last section of the quality assessment of the model, the PDB format of the model was analyzed via ProSA. Based on the ProSA analysis, the Z score was about − 4.55, indicating an overall quality of the model, which is within the range of the values typically found for the native proteins of similar sizes as presented in Fig. S2B and C (Supplementary Material) and Table 2.
Structural and physicochemical specifications
The use of frustratometer (http://www.frustratometer.tk/) for defining the local energy landscape in a protein molecule can result in important information in terms of the biological behavior of a protein. In fact, it is necessary to address how the energy is distributed in a protein structure and how the conformational changes lead to changes in the energy distributions, in which the sites with the high local frustration often specify the biologically important regions (e.g., binding or allosteric sites). Minimally frustrated linkages constitute a stable folding core of the molecule. The highly and minimally frustrated regions in the refined model are shown in Fig. S3 (Supplementary Material).
The CASTp server was used to find the pockets and cavities of VL. The CASTp analysis revealed several areas as ligand-binding pockets or cavities, in which the largest pocket (VOL = 1141.2) in this protein was composed of 32 residues that mostly located in the C-terminal of the VL (Fig. 6).
As a part of the evaluation of the VL, the prediction of ligand binding sites of this protein was accomplished by means of COACH. Figure S4 (Supplementary Material) represents the predicted results for the ligand binding sites that ranked and ordered based on the C score values.
Docking results
Antibody mode of the ClusPro was used along with the non-CDR regions masking strategy to find the most important regions involved in the interactions between VL and UreB. The computational showed a good correlation with the experimental data. The second-ranked docking model was found to possess the lowest energy of − 352.0 comparable to the first ranked model that showed the lowest energy of − 354.9. Multiple interactions between the UreB and the VL resulted in a tight binding of the VL to UreB, through hydrogen bonding, salt bridges, hydrophobic interactions, cation–pi, and pi–pi interactions (Fig. 7). Screening in CDR regions showed multiple contacts by defining 5A as a cutoff for distance (Fig. 7). Selecting less strict cutoff (8A) for the distance, to find interactions between the CDR regions and the Flap regions in UreB, indicated an interaction between GLN 106 from CDR3 and HIS 323 from the UreB Flap. Further, with 5A cutoff for the distance, we did not find any interactions between CDRs and the Flap region.
Docking analysis. a, b Two view modes from the best docking model between VL and UreB, respectively. c–e The schematic views of interactions between the CDR regions of VL (CDR1, 2, and 3, respectively) and UreB with distance < 5.0 as cutoff. The blue molecule represents the UreB, and the magenta molecule represents the VL. The CDR regions of VL represented in green. f The interaction between the CDR3 (GLN 106) and the UreB Flap (HIS 323) with 8A as cutoff
Discussion
Considering the increased rate of the H. pylori-related diseases, in particular, gastric cancer as the second most frequent cause of the cancer-related deaths worldwide (Arkenau 2009; Yamashita et al. 2011), much more effective therapeutic and diagnostic methods are indeed required. The immunological approaches can be counted as one of the most prominent current issue studied, including the discovery of efficient prophylactic epitopes, and the production of antibodies against crucial virulence factors for passive immunization and detection. In recent decades, many immunologists have focused on the control of H. pylori infections. For instance, Cao et al. succeeded to isolate a murine anti-H. pylori scFv, recognizing a 30-kDa surface protein with profound inhibitory effects on the H. pylori in vitro and its colonization in the mouse stomach (Cao et al. 2000). Further, several phage display/hybridoma-based antibody fragments and epitopes have been identified against H. pylori. The passive immunotherapy has been considered as a new strategy for controlling the H. pylori infection, in particular in the countries with the high incidence rates of infection. Although a couple of H. pylori antigens have successfully been utilized for the vaccination against H. pylori in animal models, these antigens do not show high immunogenicity to elicit a protective immune response in human clinical trials. Presumably, the lack of efficiency is because of the ineffectiveness of IgA and IgM monoclonal antibodies (Abs) in comparison with the secretory IgA (sIgA). Further, the paucity of our understanding in terms of the immune responses to the H. pylori infection together with lack of information on epitope(s) and appropriate adjuvant(s) might be the main reason for the failure of the vaccine therapies against the H. pylori infection (Berdoz and Corthesy 2004). Among H. pylori virulence factors, the urease enzyme is considered a crucial factor in all pathogenic strains, which helps the bacterium to tolerate and pass through the harsh acidic environment of the stomach and colonize beneath the mucous lining. The UreB, as a subunit, catalyzes the urea to ammonia and provides a safe microenvironment for such process. This subunit is antigenic and can stimulate the immune system response. Therefore, neutralizing antibody directed to this subunit might potentially be a promising therapeutic modality.
As one of the unique properties of H. pylori urease, the low Km (18 mM) has made this enzyme stronger than other ureolytic strains encountering the harsh acidic stomach condition. The cavity site of the most important ureolytic bacteria is conserved. This is an extremely important trait that is involved in different motions and flexibility of the Flap segment located exactly at the entrance of the catalytic site cavity (Ha et al. 2001). By screening a panel of overlapping synthetic peptides covering the entire sequence of the two subunits (i.e., UreA and UreB), two stretches of UreB-derived 19 and 15 aa residues, (UB-33; aa 321 to 339, CHHLDKSIKEDVQFADSRI) and (U211–225, IEAGAIGFKIHEDWG), were identified (Hirota et al. 2001a; Li et al. 2008). They were found to be localized in the neighborhood of the active site of urease specifically recognized by the isolated L2 and 6E6 hybridoma-based antibody. Some mAbs have been generated against the conformational epitopes. Given that mAbs against these two linear antigenic epitopes show marked neutralizing impacts with high inhibitory effect, these linear epitopes are deemed to play crucial roles. Further, a minimal epitope as 8 amino acid residues (F8; SIKEDVQF) for L2 reactivity is a part of the Flap segment, which might be a key segment in terms of targeting of the bacterium. Based on previous reports, this segment plays a critical role in the formation of the catalytic cavity site pocket. In the current study, having recruited the phage display technology, we aimed to isolate and characterize fully human antibody fragments against the purified and customized Flap segment of H. pylori urease that also presents as an efficient epitope for the vaccination. So far, against the whole two subunits of urease enzyme, some mAbs have been isolated with somewhat success. In this current study, for the first time, we capitalized on peptide-based and solution-phase biopanning to isolate antibody fragments against UreB.
Once the Flap accessibility was confirmed by bioinformatic analysis, the peptide was customized with some necessary modifications and the solution-phase biopanning process confirmed the enrichments of high-affinity scFvs (Figs. 1, 2, 3, and 4). Of these, only three identical VL antibody fragments were identified based on the sequencing. Further, stringent strategies were applied, including (a) decreasing peptide concentration through five successive rounds and (b) an increasing number of wash steps (i.e., 5 to 20 times for rounds 1–5) leading to decrease signals in the rounds 4 and 5. This method maximized the selection of highly specific antibodies against native conformation of targets while minimizing the isolation of an antibody with an inability of recognizing the whole native protein in the physiological context. Based on the concentrations used in the biopanning process, we estimated that the approximate affinity of our target to be in the nanomolar range as validated by the SPR analysis (Table 1 and Fig. S1). In fact, the affinity of the antibody fragments below 100 nM is considered as high affinity, upon which the isolated VL fragment could be counted as an Ab fragment with high affinity. The low affinity of the C3 in comparison with other isolated antibody fragments may be due to use of semi-synthetic library instead of immune or other kinds of libraries. In fact, semi-synthetic libraries contain numerous types of antibody fragments with low affinity while antibodies constructed through immune libraries show high affinity and low diversity antibodies. It is worth mentioning that coating plates indirectly for the ELISA makes the peptide much more accessible. In other words, the conformation of the peptide is more native leading to isolating high affinity and accurate scFvs. In short, the indirect coating seems to be superior to the direct one in terms of protecting peptide conformation. The results of the inhibitory experiments (Fig. 4) showed that the isolated Ab fragment can neutralize the activity of urease up to 40%, where the percentage of the phage-VL complex was calculated to be about 60% as compared to controls. The greater inhibitory effect of the phage-VL complex may be referred to the physical elements. First, the space occupancy of the massive body of M13 capsid may show some interference with the urease active site. Subsequently, the entry of urea as the substrate to the cavity pocket can be limited. Second, different conformations of VL when fused to Pro3 (one of the major capsid protein) could be another important issue.
We also applied several computational techniques to further validate the elected Ab fragments (Figs. 5, 6, 7, S2, S3, and S4). Based on the docking results (Fig. 7), the CDRs of VL displayed interactions with the UreB. Such impacts could potentially change the structural conformation of the target UreB near the Flap region and disturb the function of it. This in silico finding may help to explain the mechanism of the VL impacts on the UreB. It seems that a 30 aa (200–230) motif (Fig. 7b, the blue region of the UreB) can provide a potential deduced from the epitope mapping screening. Having compared this region to the Flap region (313–346) in terms of the interactions with and distance to the CDRs, as shown in Fig. 7b (the red region of UreB), it was found that the VL interact with 200–230 region more than 313–346 region. As this 30-residue motif is located exactly at the vicinity of the Flap segment, interaction of the isolated VL with this region probably cause to disturb flexibility and function of the Flap. We speculate that the inhibitory effect of the isolated VL and some other reported Abs could be due to their interaction(s) with different sites of the UreB resulting in the different degrees of inhibition of the UreB. We also ponder that such different outcomes may be attributed to (a) the different conformation of VL-pro3 fusion and (b) the space prevention made by the phage capsid at the entrance of enzyme cavity. Further, owing to the small size and shape of the isolated VL fragment, it might be more resistant to the temperature and denaturing agents in comparison with the full-length Ab and even Ab fragments such as Fab. Besides, the penetration of the isolated VL fragment into the human tissue and especially stomach mucosa, where H. pylori are colonized, can be substantially high. It has been shown that the human polymeric IgA is superior to other antibody fragments of the same monoclonal specificity to neutralize the activity of H. pylori urease enzyme (Berdoz and Corthesy 2004). Furthermore, the existence of mucosal secretory IgA (sIgA) antibody specific to the urease in more than 50% of H. pylori-induced gastritis patients confirms that the urease enzyme is a perfect candidate for the targeted therapy of this disease. Based on the bold role of the sIgA in H. pylori-infected patients, the selected Ab fragments could be reformatted into various full-length human antibody isotypes for efficient targeted therapy. Owing to the high avidity of sIgA, it shows superior blocking efficacy in comparison with the IgG counterparts or even another format of Ab fragments such as scFv.
In conclusion, in the current study, we isolated a fully human VL Ab fragment with high affinity specific to the most critical segment of the urease enzyme of H. pylori (UreB). The isolated VL showed good inhibitory impacts on H. pylori. As a result, it is proposed as a therapeutic agent for the clinical and/or preclinical applications. The engineered VL Abs can be as efficient as the other formats of small fragment antibodies (e.g., scFvs, Fabs, and nanobodies). Further, VL Abs can easily be matured and reformatted to an appropriate subtype of antibodies with much more desirable characteristics like L2 and 6E6 in order to exploit as a targeting agent in the development of the targeted nanomedicines and immunosensors. In addition, we confirmed previous reports reporting the importance of Flap segment that gives unique characteristic to urease of H. pylori. In fact, this segment has a crucial role in the urease activity and can potentially be chosen as a valuable candidate for the passive and active immunization. Taken all, the flexible localization of this important segment (Flap) directly or indirectly may lead to the reduction of the urease enzyme activity. Thus, theoretically and potentially, the Flap segment can be proposed as a target for the immunization if the epitope designing and all related issues are taken into consideration.
References
Abdolalizadeh J, Nouri M, Zolbanin JM, Barzegari A, Baradaran B, Barar J, Coukos G, Omidi Y (2013) Targeting cytokines: production and characterization of anti-TNF-alpha scFvs by phage display technology. Curr Pharm Des 19(15):2839–2847. https://doi.org/10.2174/1381612811319150019
Ardekani LS, Gargari SL, Rasooli I, Bazl MR, Mohammadi M, Ebrahimizadeh W, Bakherad H, Zare H (2013) A novel nanobody against urease activity of Helicobacter pylori. International journal of infectious diseases : IJID : official publication of the International Society for Infectious Diseases 17(9):e723–e728. https://doi.org/10.1016/j.ijid.2013.02.015
Arkenau HT (2009) Gastric cancer in the era of molecularly targeted agents: current drug development strategies. J Cancer Res Clin Oncol 135(7):855–866. https://doi.org/10.1007/s00432-009-0583-7
Berdoz J, Corthesy B (2004) Human polymeric IgA is superior to IgG and single-chain Fv of the same monoclonal specificity to inhibit urease activity associated with Helicobacter pylori. Mol Immunol 41(10):1013–1022. https://doi.org/10.1016/j.molimm.2004.05.006
Blanchard TG, Nedrud JG (2012) Laboratory maintenance of Helicobacter species. Current protocols in microbiology Chapter 8:Unit8B.1 https://doi.org/10.1002/9780471729259.mc08b01s24
Blaser MJ, Berg DE (2001) Helicobacter pylori genetic diversity and risk of human disease. J Clin Invest 107(7):767–773. https://doi.org/10.1172/jci12672
Cao J, Sun Y, Berglindh T, Mellgard B, Li Z, Mardh B, Mardh S (2000) Helicobacter pylori-antigen-binding fragments expressed on the filamentous M13 phage prevent bacterial growth. Biochim Biophys Acta 1474(1):107–113. https://doi.org/10.1016/S0304-4165(00)00005-2
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519. https://doi.org/10.1002/pro.5560020916
de Haard HJ, van Neer N, Reurs A, Hufton SE, Roovers RC, Henderikx P, de Bruine AP, Arends JW, Hoogenboom HR (1999) A large non-immunized human Fab fragment phage library that permits rapid isolation and kinetic analysis of high affinity antibodies. J Biol Chem 274(26):18218–18230. https://doi.org/10.1074/jbc.274.26.18218
Dundas J, Ouyang Z, Tseng J, Binkowski A, Turpaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34(Web Server):W116–W118. https://doi.org/10.1093/nar/gkl282
Emadi S, Barkhordarian H, Wang MS, Schulz P, Sierks MR (2007) Isolation of a human single chain antibody fragment against oligomeric alpha-synuclein that inhibits aggregation and prevents alpha-synuclein-induced toxicity. J Mol Biol 368(4):1132–1144. https://doi.org/10.1016/j.jmb.2007.02.089
Fahimi F, Sarhaddi S, Fouladi M, Samadi N, Sadeghi J, Golchin A, Tohidkia MR, Barar J, Omidi Y (2018) Phage display-derived antibody fragments against conserved regions of VacA toxin of Helicobacter pylori. Appl Microbiol Biotechnol 102(16):6899–6913. https://doi.org/10.1007/s00253-018-9068-4
Ha NC, Oh ST, Sung JY, Cha KA, Lee MH, Oh BH (2001) Supramolecular assembly and acid resistance of Helicobacter pylori urease. Nat Struct Biol 8(6):505–509. https://doi.org/10.1038/88563
Hawkins RE, Russell SJ, Winter G (1992) Selection of phage antibodies by binding affinity. Mimicking affinity maturation J Mol Biol 226(3):889–896. https://doi.org/10.1016/0022-2836(92)90639-2
Henderikx P, Kandilogiannaki M, Petrarca C, von Mensdorff-Pouilly S, Hilgers JH, Krambovitis E, Arends JW, Hoogenboom HR (1998) Human single-chain Fv antibodies to MUC1 core peptide selected from phage display libraries recognize unique epitopes and predominantly bind adenocarcinoma. Cancer Res 58(19):4324–4332
Hirota K, Nagata K, Norose Y, Futagami S, Nakagawa Y, Senpuku H, Kobayashi M, Takahashi H (2001a) Identification of an antigenic epitope in Helicobacter pylori urease that induces neutralizing antibody production. Infect Immun 69(11):6597–6603. https://doi.org/10.1128/IAI.69.11.6597-6603.2001
Hirota K, Nagata K, Norose Y, Futagami S, Nakagawa Y, Senpuku H, Kobayashi M, Takahashi H (2001b) Identification of an antigenic epitope in Helicobacter pylori urease that induces neutralizing antibody production. Infect Immun 69(11):6597–6603. https://doi.org/10.1128/iai.69.11.6597-6603.2001
Houimel M, Corthesy-Theulaz I, Fisch I, Wong C, Corthesy B, Mach J, Finnern R (2001) Selection of human single chain Fv antibody fragments binding and inhibiting Helicobacter pylori urease. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine 22(1):36–44. https://doi.org/10.1159/000030153
Izzotti A, Durando P, Ansaldi F, Gianiorio F, Pulliero A (2009) Interaction between Helicobacter pylori, diet, and genetic polymorphisms as related to non-cancer diseases. Mutat Res 667(1–2):142–157. https://doi.org/10.1016/j.mrfmmm.2009.02.002
Khoury GA, Tamamis P, Pinnaduwage N, Smadbeck J, Kieslich CA, Floudas CA (2014) Princeton_TIGRESS: protein geometry refinement using simulations and support vector machines. Proteins 82(5):794–814. https://doi.org/10.1002/prot.24459
Ko GH, Park HB, Shin MK, Park CK, Lee JH, Youn HS, Cho MJ, Lee WK, Rhee KH (1997) Monoclonal antibodies against Helicobacter pylori cross-react with human tissue. Helicobacter 2(4):210–215. https://doi.org/10.1111/j.1523-5378.1997.tb00090.x
Kozakov D, Hall DR, Xia B, Porter KA, Padhorny D, Yueh C, Beglov D, Vajda S (2017) The ClusPro web server for protein-protein docking. Nat Protoc 12(2):255–278. https://doi.org/10.1038/nprot.2016.169
Krieger E, Vriend G (2014) YASARA View—molecular graphics for all devices—from smartphones to workstations. Bioinformatics 30(20):2981–2982. https://doi.org/10.1093/bioinformatics/btu426
Lee MH, Roussel Y, Wilks M, Tabaqchali S (2001) Expression of Helicobacter pylori urease subunit B gene in Lactococcus lactis MG1363 and its use as a vaccine delivery system against H. pylori infection in mice. Vaccine 19(28–29):3927–3935. https://doi.org/10.1016/S0264-410X(01)00119-0
Li HX, Mao XH, Shi Y, Ma Y, Wu YN, Zhang WJ, Luo P, Yu S, Zhou WY, Guo Y, Wu C, Guo G, Zou QM (2008) Screening and identification of a novel B-cell neutralizing epitope from Helicobacter pylori UreB. Vaccine 26(52):6945–6949. https://doi.org/10.1016/j.vaccine.2008.09.089
Luthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three-dimensional profiles. Nature 356(6364):83–85. https://doi.org/10.1038/356083a0
Maleki Kakelar H, Barzegari A, Dehghani J, Hanifian S, Saeedi N, Barar J, Omidi Y (2018) Pathogenicity of Helicobacter pylori in cancer development and impacts of vaccination. Gastric Cancer 22:23–36. https://doi.org/10.1007/s10120-018-0867-1
Marks JD, Hoogenboom HR, Bonnert TP, McCafferty J, Griffiths AD, Winter G (1991) By-passing immunization. Human antibodies from V-gene libraries displayed on phage J Mol Biol 222(3):581–597. https://doi.org/10.1016/0022-2836(91)90498-U
Michetti P (1997) Vaccine against Helicobacter pylori: fact or fiction? Gut 41(6):728–730
Mobley HL, Cortesia MJ, Rosenthal LE, Jones BD (1988) Characterization of urease from Campylobacter pylori. J Clin Microbiol 26(5):831–836
Nagata K, Mizuta T, Tonokatu Y, Fukuda Y, Okamura H, Hayashi T, Shimoyama T, Tamura T (1992) Monoclonal antibodies against the native urease of Helicobacter pylori: synergistic inhibition of urease activity by monoclonal antibody combinations. Infect Immun 60(11):4826–4831
Nakayama Y, Graham DY (2004) Helicobacter pylori infection: diagnosis and treatment. Expert Rev Anti-Infect Ther 2(4):599–610. https://doi.org/10.1586/14787210.2.4.599
Parra RG, Schafer NP, Radusky LG, Tsai MY, Guzovsky AB, Wolynes PG, Ferreiro DU (2016) Protein Frustratometer 2: a tool to localize energetic frustration in protein molecules, now with electrostatics. Nucleic Acids Res 44(W1):W356–W360. https://doi.org/10.1093/nar/gkw304
Porter KA, Xia B, Beglov D, Bohnuud T, Alam N, Schueler-Furman O, Kozakov D (2017) ClusPro PeptiDock: efficient global docking of peptide recognition motifs using FFT. Bioinformatics 33(20):3299–3301. https://doi.org/10.1093/bioinformatics/btx216
Reiche N, Jung A, Brabletz T, Vater T, Kirchner T, Faller G (2002) Generation and characterization of human monoclonal scFv antibodies against Helicobacter pylori antigens. Infect Immun 70(8):4158–4164. https://doi.org/10.1128/IAI.70.8.4158-4164.2002
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5(4):725–738. https://doi.org/10.1038/nprot.2010.5
Schmausser B, Eck M, Greiner A, Luhrs H, Vollmers HP, Muller-Hermelink HK (2002) Disparity between mucosal and serum IgA and IgG in Helicobacter pylori infection. Virchows Arch 441(2):143–147. https://doi.org/10.1007/s00428-002-0621-1
Schrodinger LLC (2015) The PyMOL molecular graphics system. Version 1:8
Telford JL, Ghiara P (1996) Prospects for the development of a vaccine against Helicobacter pylori. Drugs 52(6):799–804
Tohidkia MR, Barar J, Asadi F, Omidi Y (2012) Molecular considerations for development of phage antibody libraries. J Drug Target 20(3):195–208. https://doi.org/10.3109/1061186X.2011.611517
Tohidkia MR, Asadi F, Barar J, Omidi Y (2013) Selection of potential therapeutic human single-chain Fv antibodies against cholecystokinin-B/gastrin receptor by phage display technology. BioDrugs 27(1):55–67. https://doi.org/10.1007/s40259-012-0007-0
Tohidkia MR, Sepehri M, Khajeh S, Barar J, Omidi Y (2017) Improved soluble ScFv ELISA screening approach for antibody discovery using phage display technology. SLAS Discov:2472555217701059. https://doi.org/10.1177/2472555217701059
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(Web Server):W407–W410. https://doi.org/10.1093/nar/gkm290
Wu S, Zhang Y (2007) LOMETS: a local meta-threading-server for protein structure prediction. Nucleic Acids Res 35(10):3375–3382. https://doi.org/10.1093/nar/gkm251
Yamashita K, Sakuramoto S, Watanabe M (2011) Genomic and epigenetic profiles of gastric cancer: potential diagnostic and therapeutic applications. Surg Today 41(1):24–38. https://doi.org/10.1007/s00595-010-4370-5
Yan R, Xu D, Yang J, Walker S, Zhang Y (2013) A comparative assessment and analysis of 20 representative sequence alignment methods for protein structure prediction. Sci Rep 3:2619. https://doi.org/10.1038/srep02619
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43(W1):W174–W181. https://doi.org/10.1093/nar/gkv342
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9:40. https://doi.org/10.1186/1471-2105-9-40
Zhang Y, Skolnick J (2004) SPICKER: a clustering approach to identify near-native protein folds. J Comput Chem 25(6):865–871. https://doi.org/10.1002/jcc.20011
Acknowledgments
The authors like to thank all the members of the Research Center for Pharmaceutical Nanotechnology, Biomedicine Institute at Tabriz University of Medical Sciences for the technical support. The authors also like to thank Dr. Hengameh Zandi and Mr. Reza Mousavi for the gift of reagents used in the preparation of H. pylori culture.
Funding
This work was financially supported by the Research Center for Pharmaceutical Nanotechnology at Tabriz University of Medical Sciences (grant no. RCPN-93006 and 93016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
The ethical approval for the use of patients’ samples was obtained from the Ethical Committee of Tabriz University of Medical Sciences. All the gastric patients admitted to Imam Reza Hospital (Tabriz, Iran), whose fresh and positive urease biopsy samples were used in this current study, were informed on the use of the samples for the study with written consents.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 1294 kb)
Rights and permissions
About this article
Cite this article
Fouladi, M., Sarhadi, S., Tohidkia, M. et al. Selection of a fully human single domain antibody specific to Helicobacter pylori urease. Appl Microbiol Biotechnol 103, 3407–3420 (2019). https://doi.org/10.1007/s00253-019-09674-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09674-6