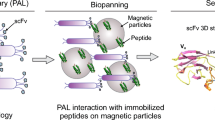
Abstract
Infection with Helicobacter pylori may result in the emergence of gastric adenocarcinoma. Among various toxins assisting pathogenesis of H. pylori, the vacuolating cytotoxin A (VacA) is one of the most potent toxins known as the major cause of the peptic ulcer and gastric adenocarcinoma. To isolate single-chain variable fragments (scFvs) against two conserved regions of VacA, we capitalized on the phage display technology and a solution-phase biopanning (SPB). Characterization of scFvs was carried out by enzyme-linked immunosorbent assay (ELISA), immunoblotting, and surface plasmon resonance (SPR). Bioinformatics analyses were also performed in order to characterize the structural and functional properties of the isolated scFvs and the interaction(s) between the isolated antibodies (Ab)-antigen (Ag). After four rounds of biopanning, the positive colonies detected by scFv ELISA were harvested to extract the plasmids and perform sequencing. Of several colonies, three colonies showed high affinity to the VacA1 and two colonies for the VacA2. Further complementary examinations (e.g., sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), western blot, SPR, and flow cytometry) displayed the high affinity and specificity of the isolated scFvs to the VacA. Docking results revealed the interaction of the complementarity-determining regions (CDRs) with the VacA peptide. In conclusion, for the first time, we report on the isolation of several scFvs against conserved residues of VacA toxin with high affinity and specificity, which may be used as novel diagnostic/therapeutic tool in the H. pylori infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Helicobacter pylori is a gram-negative bacterium, which is colonized in the human stomach. It has infected almost half of the world’s population, with the highest prevalence in the developing countries due to poor health and economic backgrounds (Adler et al. 2014; Lim et al. 2013; Oleastro and Menard 2013). In patients with H. pylori, in the absence of appropriate antibiotic treatment, long-term persistence of H. pylori could contribute to possible gastritis, peptic or duodenal ulcers, gastric adenocarcinoma, or mucosa-associated lymphoid tissue (MALT) lymphoma (Chey et al. 2007; Kuo et al. 2014; Pritchard and Crabtree 2006). A number of investigations have confirmed that the strong colonization of H. pylori might give rises to stomach-related malignancies, resulting in the classification of H. pylori as one of the main carcinogenic agents (Hagymasi and Tulassay 2014).
While the eradication of H. pylori infection is much of a concern, attempts for accurate detection of the infection appears as an ongoing challenge (Garza-Gonzalez et al. 2014). The invasive diagnosis methods are mostly on the basis of histology and biopsy of the stomach and further detection by molecular methods such as polymerase chain reaction (PCR) and fluorescent in situ hybridization (FISH) or physiological methods like rapid urease (RU) test. Despite being the most common method for the detection of H. pylori, the endoscopy and biopsy are assumed to be labor intensive for patients especially children and elderly patients (Carvalho et al. 2012). The noninvasive serology tests proceeding based on the presence of immunoglobulin G are the most prevalently used procedures. However, there exist some drawbacks, including misinterpretation of the data, and the prolonged existence of the Igs after eradication limits the application of immunodiagnostic method (Patel et al. 2014). The most prevalent procedure in post-treatment patients is known as the urea breath test (UBT) which may provide false-negative results after treatment with concomitant drug combinations (Capurso et al. 2006). As a result, in order to provide an effective therapy for eradication of the infection with H. pylori, a new method of detection for the infection is required.
Following the production of monoclonal antibodies (mAbs) by Köhler and Milstein in 1986, the biopharmaceutical market of commercialized mAbs has exponentially expanded (Kohler and Milstein 1975; Siddiqui 2010). Even though production of the mAbs has commenced a new era in the diagnosis methods, some shortcomings have urged rationalized strategies for development of mAbs, including low yielding antibodies and time-consuming and exerting immunological side effects in patients such as human anti-mouse antibody (HAMA) (Liu 2014; Reichert 2012). Among various antibody (Ab) development approaches, phage Ab-display system is a powerful technique for production of fully mAbs with high affinity and avidity towards extensive types of antigens (Ags) (Tohidkia et al. 2012; Zhao et al. 2016). Among several types of characterized Ab formats incorporated onto the phages, scFv particles are the most common types of Abs exploited for the establishment of phage display libraries. The main principle of this technology is the production of a large and diverse set of phages (the so-called library) that encompasses variable light and heavy chains (VL and VH, respectively) repertoires through randomized complementarity-determining regions (CDRs) and successive rounds of selection against any desired Ags. Despite the great variety of the expressive platforms, the E. coli is considered as one of the best candidates for the amplification and expression of the phage-scFv insert particles in vitro (Georgieva and Konthur 2011; Haque and Tonks 2012; Sidhu 2001; Verma et al. 1998). Recursive convenient isolation cycles and ability to simultaneously isolate several fragments against various epitopes of a given Ag has made it an advantageous method of the isolation of Abs (Ayriss et al. 2007).
Besides the presence of urease as the most potential virulence factor of H. pylori, the vacuolating toxin (VacA) is one of the most crucial factors for the bacterial colonization (Bessede et al. 2015; Cover et al. 2016). This toxin, which is present in almost 90% of the strains, is a 140-kDa weight toxin and produces an 88-kDa mature toxin containing two domains known as p33 and p55 after a two-step proteolytic cleavage (Papini et al. 2001; Salama et al. 2013). The p55 domain is responsible for the toxin binding to various cell surface receptors (e.g., receptor protein tyrosin phosphatases (RPTP) α and β, epidermal growth factor receptor (EGFR), and lymphocyte function-associated antigen 1 (LFA-1) in T cells), while the functional p33 domain contributes to vacuolation and pore formation by oligomerization on the cellular membrane (Rieder et al. 2005). The different cytotoxic activity of the VacA arises from the high polymorphism of the toxin among various strains of H. pylori. One region inside the p55 domain (the so-called M region) is known as one of the most diverse sequences of the toxin and based on the residues is classified in two groups, including m1 and m2 (Figura et al. 2015). Recent investigations determined two highly conserved regions between m1 and m2, containing residues 668–678 and 730–734 at C-terminal and is hypothesized to be the surface-exposed area responsible for the cell receptor binding (Gangwer et al. 2007).
Former diagnostic humoral Ab-based lab tests utilized for the detection of H. pylori were included polyclonal Abs harvested from patient’s serum (Hensel et al. 1999). The VacA, as a membrane-associated protein on the H. pylori, is one of the major toxins responsible for peptic ulcers (Fahimi et al. 2017). Thus, development of scFvs against conserved regions of this toxin provides a great possibility for the accurate diagnosis at the early stages of H. pylori-mediated infection and peptic ulcers and the precise treatment of such formidable disease. In this study, for the first time, we report on the isolation of scFvs from a semisynthetic phage antibody library (PAL) against C-ter conserved domains of VacA toxin, which were further characterized using both wet-lab and bioinformatic techniques to attain scFvs with high binding affinity.
Materials and methods
Peptide design
Two consensus surface-exposed residues, mediating host receptors and toxin interaction, were selected as the target for the isolation of scFvs (Fig. 1). The two mentioned residues were separately customized by BIOMATIK (Cambridge, Ontario, Canada) with a purity value of 98% verified by a high-performance liquid chromatography (HPLC). The VacA1 and VacA2 peptides (668–678 and 730–734 at C-terminal, respectively) were tagged with mini polyethylene glycol (PEG) and biotin. As a result, mini-PEG and biotin-tagged VacA1 (at C-terminal) and mini-PEG- and biotin-tagged VacA2 (at N-terminal) were used for the affinity capture during the biopanning process.
The p55 domain of vacuolating toxin A (VacA) with conserved domains. The image was produced based on the crystallography of p55 domain of toxin. Two consensus residues at the C-ter of p55 domain considered as receptor binding domains of the toxin were chosen as separate peptides for the isolation of scFvs. VacA1 and VacA2 sequences are shown as red and blue, respectively
Bacterial strains and library preparation
The TG1 Tr E. coli used for the amplification of scFv-phage, HB2151 E. coli used for the production of soluble scFvs, the semisynthetic Tomlinson library I+J, and KM13 helper phage were purchased from Source Bioscience (Nottingham, UK). The PAL was constructed by side chain diversification of CDRs encompassing CDR2 and CDR3 of a single framework from VH (V3-23/DP-47 and JH4b), VL (O12/O2/DPK9 and Jκ1), and a CDR1 that was kept constant. The diversifications were incorporated at 18 residues that are highly diverse in the primary repertoire and function as Ag-binding sites. Sequences encoding scFvs were cloned next to gene 3 protein in phagemid vector, pIT2, with histidine (His) and myc tags.
Selection of VacA-specific scFvs by a solution-phase biopanning
According to the Tomlinson protocol, 106 colony forming unit (cfu) phage colonies were rescued by helper phages from the library I and subjected to four rounds of panning with descending concentrations of peptide for VacA1 and VacA2. Briefly, after blockage of Strep Dynabead Myone-T1 (Thermo Fisher, Waltham, MA, USA) and phages with bovine serum albumin (BSA) as 0.3% in phosphate-buffered saline (PBS), pre-absorption was performed for each round in order to eliminate the strep-binder phages. Biotinylated peptides were exposed to the blocked phages for 120 min at room temperature. Then, peptide-bound phages were captured by blocked strep Dynabeads. The Dynabeads-captured phage-peptide particles were separated by a magnetic apparatus, DynaMag-2 (Invitrogen, Karlsruhe, Germany). Stringent washing steps (6×) were performed for all rounds of panning in order to isolate any cross-reacting phage particles. Phages were eluted by 500 μL pancreas trypsin (1 mg/mL) (Sigma-Aldrich Co., Taufkirchen, Germany) for 20 min. Half of the eluates (250 μL) were exploited for the next round of biopanning process and transferred to TG1 E. coli at the optical density (OD) of 0.4, then plated on trypton yeast extract (TYE) media containing 100 mg ampicillin/mL and 2% (w/v) glucose (TYE-Amp-Glc) as amplified phages for the next rounds of panning.
Polyclonal phage enzyme-linked immunosorbent assay (ELISA)
ELISA was performed on polystyrene 96-well plates to monitor the rate of enrichment of phages bound to peptides following the panning cycles. ELISA plates coated with 2 μg/mL BSA (Sigma-Aldrich Co., Taufkirchen, Germany) and 1 mg/mL streptavidin (S888, Invitrogen, Karlsruhe, Germany) were added with 100 μL/well peptide and mini-PEG respectively as positive and negative wells and then incubated for 1.5 h at room temperature. Following blocking, 100 μL/well of the eluted phages of each round were incubated for 1 h at room temperature. The selected phages with target scFvs obtained from the panning process were exposed to the primary anti-M13 mAb (GE Healthcare, Amersham, UK) and the secondary antibody goat anti-mouse immunoglobulin (IgG) conjugated with horse radish peroxidase (HRP) [(ab6789), Abcam, Cambridge, MA, USA] respectively at dilutions of 1:3000 and 1:5000. After performing optical reaction by adding the HRP substrate tetramethylbenzidine (TMB), the intensity of color was quantified by an ELISA reader at 450–630, ELx808™ (Biotek, Winooski, VT, USA). All the steps were followed by three rounds of washing with phosphate-buffered saline and tween (PBST) 0.1%.
Soluble antibody fragment ELISA
For the production of soluble scFv fragments, 200 μL of the non-suppressor strain of E. coli HB2151 at the exponential phase (OD 0.4) was combined with 10 μL of eluted phages of round 4 of either VacA1 or VacA2 and incubated on water bath 37 °C for 45 min. Single colonies resulted from the culture of eluates in HB2151 E. coli were transferred to master plates containing 2× TY with 100 μg/mL ampicillin and 2% (w/v) glucose (2× YT/Amp/Glc) and incubated overnight at 37 °C. A small quantity (5 μL) of each well was transferred to 200 μL 2× TY/Amp/Glc 0.1% and incubated on shaker plate at 37 °C for 3 h to reach to OD 0.9. The production of soluble scFvs was stimulated by addition of 25 μL induction media, containing 9 mM isopropyl β-d-1-thiogalactopyranoside (IPTG), 0.4 M sucrose, and 100 μg/mL ampicillin. Then, it was incubated on shaker plate overnight at 30 °C. After centrifugation, the plates were resuspended in 50 μL periplasmic extraction (PE) buffer (3× PE) for the extraction of periplasmic scFvs. The extracted soluble scFvs were exploited for monoclonal scFv ELISA as same as polyclonal phage ELISA as mentioned previously, except that the detection of scFvs was carried out in one step with the addition of HRP-Protein A at dilution 1:5000.
Sequence analysis
ScFv-encoding phagemids of each positive colony were extracted by QIAprep Spin Miniprep Kit (Qiagen, Hilden, Germany). The PCR analysis together with gel electrophoresis confirmed the quality of two-strand plasmids and presence of scFv genes inside plasmids using pHEN and LMB3 primers as forward and reverse primers respectively (data not shown). Having used LMB3 primer, ABI 3730XL DNA sequencer (Applied Biosystems, Darmstadt, Germany) was exploited for determination of sequences. The sequencing data were analyzed by Chromas 2.4.4 (Technelysium Pty Ltd., South Brisbane, Australia), and the alignment of CDR regions was accomplished by the VBASE2 database. Furthermore, unique sequence alignments with the human germline genes were performed using the IMGT/V-QUEST (http://www.imgt.org).
Expression of scFvs in large scale
Colonies scored the highest ODs in monoclonal scFv ELISA with different CDRs were selected for the expression of soluble scFvs in 400 mL broth culture. The overnight grown colonies in 400 mL 2× TY/Amp/Glc 2% were incubated for OD 0.9 to precipitate by centrifuge. Then, they were resuspended in 400 mL 2× TY/Amp/Sucrose 0.4 M supplemented with 1 mM IPTG and incubated for 4 h at 30 °C. Following centrifugation at 2800×g for 20 min, the pellets were resuspended in 4% culture volume of ice-cold periplasmic extraction TES buffer [Tris 0.1 mM, ethylenediaminetetraacetic acid (EDTA) and 100 mM sucrose 2 M]. Then, the extracted scFvs in the supernatant were collected as final samples and stored for further steps. The expression of scFvs in large scale was proved by SDS-PAGE 12% under reducing condition using a Mini-PROTEAN Tetra Cell system (Bio-Rad, Munich, Germany) and western blot. The primary antibody (c-myc) and the secondary antibody conjugated with HRP were added at dilutions of 1:3000 and 1:4000, respectively. Visualization was enhanced by the addition of electrochemiluminescence (ECL) solution of Western blotting kit (GE Healthcare, Amersham, UK), and developed on an x-ray film.
Purification of colonies by affinity chromatography
The dialyzed expressed scFvs by 12 kDa cut-off membrane (D9652, Sigma-Aldrich Co., Taufkirchen, Germany) undergone the purification process by immobilized metal affinity chromatography (IMAC) resin TALON™ (Clontech Laboratories, Inc. Mountain View, CA, USA) based on the manufacturer’s protocol. Briefly, the slurry-containing resin was applied to the resins and incubated under gentle shaking for 20 min at room temperature. Further, the sample-resin complex was centrifuged at 1500×g for 10 min, and the resin was collected followed by a couple of washing. The detachment of resins from the histidine-tag scFvs was accomplished by adding of the elution buffer and collecting as final 0.5 mL aliquots. The concentration of scFvs and the extinction coefficient was measured via the ProtParam software freely available at http://web.expasy.org/protparam/. SDS-PAGE and western blot analyses were also carried out to determine high-throughput affinity as mentioned previously.
Affinity determination of scFvs by western blot
To determine the specificity of scFvs against peptide, immunoblotting was performed by running 3 μL recombinant p55 domain of VacA toxin (AMSbio LLC, Cambridge, USA) on SDS-PAGE 10% and immunoblotted on nitrocellulose membrane on a Trans-blot® SD semidry (BIO-RAD, CA, USA) with voltage of 10 V for 25 min. After staining with purified scFvs (20 μg/mL), detection of scFvs bound to the recombinant protein was carried out by the primary polyclonal anti-VacA IgG (AMSbio LLC, Cambridge, USA) and the secondary anti-rabbit HRP-conjugate IgG at dilutions 1:1000 and 1:4000, respectively.
Surface plasmon resonance (SPR) analysis
A multi-parameter SPR instrument (MP-SPR Navi 210A, BioNavis Ltd., Tampere, Finland) and streptavidin (SA) sensor from BioNavis company were applied for the biomolecular interaction analysis and investigation of kinetic parameters in real-time. For the assay, the gold slide was exploited in both channels at 670 nm wavelength at temperature 28 °C using PBS as running buffer. About 10 μg/mL diluted biotinylated peptide was injected for 7 min at a flow rate of 30 μL/min in the flow cell 1 (Fc1), resulting in the immobilization of 100 response units (RU) for this capture method. The Fc2, with no peptide injection, was indicated as the reference channel. In order to determine binding affinities, each mAb was injected above the reference and peptide surfaces at five concentration ratios ranged from 100 to 1000 nM. Regeneration process was performed after Ab type alteration for three times with 1 min injection of 10 mM glycin-HCl at pH 2.5. Data studies of all measurements and calculation of kinetic parameters were carried out using the Data Viewer and Trace Drawer software (Bionavis).
Fluorescence-activated cell sorting (FACS) flow cytometry analysis
The functionality of the scFvs against VacA-expressed H. pylori was carried out using flow cytometry analysis using BD FACS Calibur ™ (BD Biosciences, San Jose, USA). The urease-positive biopsy samples from patients in Imam Reza Hospital (Tabriz, Iran) were cultured on Brucella blood agar base supplemented with 7% fresh defibrinated sheep blood, trimethoprim (6 μg/mL), amphotericin B (7 μg/mL), and vancomycin (10 μg/mL) under microaerophilic condition (i.e., 10% CO2, 5% O2, 85% N2) at 37 °C for 6 days. Grown agar plates with positive VacA on immunoblotting (data not shown) were scrapped and harvested for further steps. About 0.5 McFarland of harvested H. pylori was centrifuged at 2500×g for 10 min, washed twice with PBS-BSA 0.1%, and resuspended in the FACS buffer containing PBS-BSA 1% for 30 min for blocking. The scFvs were diluted 1:4 in PBS-BSA 0.1% and incubated with the prepared H. pylori samples at 4 °C for 1 h. After washing with cooled PBS-BSA, the scFv-bound H. pylori bacteria were exposed to 1 mg/mL (diluted in PBS-BSA) primary anti-c-myc antibody (Dallas, TX, USA) for 1 h at 4 °C. Detection of the bound antibody was accomplished by addition of the fluorescein isothiocyanate (FITC)-conjugated antibody at a dilution ratio of 1:64. A negative control containing no scFv was also tested.
Bioinformatic analysis
Iterative Threading ASSEmbly Refinement (I-TASSER) from the Zhang server was used for the homology modeling of the selected scFvs. I-TASSER has been ranked as one of the most applicable servers for the prediction of protein structure in CASP7-CASP11 experiments, which benefits from cutting-edge algorithms to predict the 3D structure and function of proteins (Roy et al. 2010; Yang et al. 2015; Yang and Zhang 2015; Zhang 2008). Further, the secondary structure of the scFvs was predicted using PSSpred, which is a neural network based algorithm for the prediction of protein secondary structure (Yan et al. 2013). The first step of I-TASSER pipeline was the identification of templates by the Local Meta-Threading-Server (LOMETS) from the Protein Data Bank (PDB) library. The LOMETS is a meta-server threading approach containing multiple threading programs, where each threading program can generate tens of thousands of template alignments (Wu and Zhang 2007). Each threading programs are sorted by the average performance in the large-scale benchmark test experiments and then one template with the highest Z score is selected from each threading program. In this work, 10 top-ranked templates were selected from the LOMETS threading programs. For each target, I-TASSER simulation process generates a large ensemble of structural conformations, called decoys. In the next step, the SPICKER program clustered all the decoys based on the pairwise structure similarity, then the best model was selected with consideration of C score (Zhang and Skolnick 2004). The models with the highest C scores were refined by Princeton the TIGRESS server (atlas.princeton.edu/refinement/first) (Khoury et al. 2014). Furthermore, the Frustratometer server was applied to assess the landscape of energy distribution in the refined model, which helped to understand the molecule sites that have a higher level of energy serving important roles such as the interaction with other molecules and active sites (Parra et al. 2016). Evaluation of refined models have done by Verify3D (Eisenberg et al. 1997), Protein Structure Analysis (ProsA) (Wiederstein and Sippl 2007), and ERRAT (Colovos and Yeates 1993). To evaluate the 3D structure of refined scFv models, Ramachandran plots were created by the RAPPER server (Fig. S4 and Table S1) (http://mordred.bioc.cam.ac.uk/~rapper/rampage.php). The Computed Atlas of Surface Topography of proteins (CASTp) was used to specify the packets and cavities on the surface of the scFvs (Dundas et al. 2006). To predict ligand binding sites, a consensus approach (COACH), which is a meta-server approach to protein-ligand binding site prediction, was applied (Brylinski and Skolnick 2008; Capra et al. 2009; Roy et al. 2012; Yang et al. 2013). The pepATTRACT server was used to predict the best state of docking between scFvs and intended peptides (Schindler et al. 2015). The best-predicted docking model was selected based on the lowest energy and experimental results. The visualization of PDB files and screening to find the interactions between CDR regions of scFvs and VacA was performed using the “yet another scientific artificial reality application” (YASARA) (Krieger et al. 2002; Krieger and Vriend 2014) and PyMOL (Schrodinger 2015).
Results
Determining binding specificities of cross-reacting polyclonal phage particles by ELISA
For affinity selection of recombinant phages, biopanning steps were performed in four successive rounds accompanied by a subtractive approach for the non-specific binder omission. Amplified phage-scFvs related to each round of selections were coated with biotinylated peptide and mini-PEG in positive and negative wells respectively. The panning of the library directed against VacA1 and VacA2 was assessed through the count of eluted phages, which determined a dramatic increase up to 100-fold in the last round corresponding to the enrichment of VacA-specific phages (data not shown). Polyclonal phage ELISA was performed for each round in order to assess the reactivity of scFv-phages with peptides. As shown in Fig. 2, peptides determined an exponential increase in the absorbance after the first round of panning.
Polyclonal phage enzyme-linked immunosorbent assay for the VacA1 (a) and VacA2 (b) specified phage particles. PEG-precipitated combinatorial phage eluates were utilized to recognize peptide-specific phage-scFv particles. Negative and positive wells contain mini-PEG and peptide, respectively. VacA, vacuolating toxin A; PEG, polyethylene glycol
Isolating soluble scFv fragments and scFv ELISA
The binding specificity of the selected phage clones was assessed by the production of soluble scFvs via infection of the E. coli HB2151 with the phage-scFvs. Unlike TG-1, the amber stop codon located upstream of the gene III is not suppressed in this strain, thus producing free scFv fragments. Soluble scFvs were produced from the TYE plates of the last enriched steps of biopanning for both VacA1 and VacA2. The binding activity of soluble scFvs was confirmed by scFv ELISA with the criterion of optical density higher than 0.1- and 2-fold greater absorption ratio for positive binders. As illustrated in Fig. 3, about 18 and 21 positive colonies showed specificity to the VacA1 and VacA2 peptides, respectively.
Binding efficiency of the single colonies to the VacA1 (a) and VacA2 (b) peptides. Individual clones of the last round of selection were expressed in the non-suppressor strain of E. coli, HB2151 in order to yield soluble antibody fragments. Harvested soluble scFvs were added to an enzyme-linked immunosorbent assay plate coated with biotinylated peptides and the effectiveness of the Ag-Ab interaction was measured on 450–630 wavelength. VacA, vacuolating toxin A
Analysis of scFv diversity by DNA sequencing
Of the panning process, 22 individually derived scFvs from the selection round #4 were subjected to phagemid DNA extraction and sequencing. The panning process yielded intact antibodies in all the colonies of VacA1 and two of the VacA2 (data not shown). Homology alignment of colonies determined a broad diversity among VacA1-specific scFvs and two diverse colonies for VacA2. The DNA segments encoding the scFv gene were also analyzed by the IMGT/V-QUEST tool to identify any similarity between the framework of scFvs and the human germline V, D, and J segments. As illustrated in Table 1, for the heavy chain, clones of VacA1 determined no diversity in the V and J segments representing IGHV3-23*01 F and IGHJ4*02 F, respectively. The highest difference observed in the D segment with four immunoglobulin heavy constant delta (IGHD) genes (i.e., IGHD3, IGHD1, IGHD4, and IGHD5) so as the IGHD3 gene determined the most occurrence. For the light chain, all the V and J segments were completely conserved, including immunoglobulin kappa variable (IGKV)1-39*01 F and IGKJ1*01 F, respectively. Furthermore, as shown in Table 2, the alignment of VacA2 colonies confirmed the identical V and J segments of IGHV3-23*01 F and IGHJ4*02 F and also the high rated diversity in the D segment in IGHD2 gene, including IGHD2-8*02 F, IGHD2-21*01 F, and IGHD3 gene containing IGHD3-3*01 F, IGHD3-10*01 F and IGHD3-16*01 F fragments. Same as the VacA1, the light chain was consistent with two IGKV1-39*01 F and IGKJ1*01 F genes for the V and J segments.
Expression and purification of soluble scFvs
Intact positive colonies with high reactivity in scFv ELISA from VacA1 (i.e., F10, D1, A5, and G12 colonies) and VacA2 (i.e., H1 and F4 colonies) were produced, and the supernatant was examined using SDS-PAGE and immunoblotting assays for further confirmation. The SDS-PAGE results of expressed scFvs harboring PelB leader for directing to periplasmic environment confirmed the presence of scFvs with the molecular mass of 28 kDa as well as immunoblotting (Fig. 4a, b). The expressed scFvs were also purified by the affinity chromatography exploiting their polyhistidine-tag. The quality of purified scFvs by TALON metal affinity resins following expression was observed through SDS-PAGE and western blot as well (Fig. 5a, b).
SDS-PAGE and immunoblot analysis of the individual unique clones. Expression analysis of scFvs by Coomassie brilliant blue (a) and western blot (b). UN T IPTG-uninduced cell lysates incubated for 4 h, IN T IPTG-induced cell lysates incubated for 4 h after without periplasmic buffer, UN P.P uninduced sample incubated for 4 h and periplasmic extraction. G12, A5, D1, F10, F4, and H1 IPTG-induced samples for 4 h and periplasmic extraction assembly determined with the name of clones. IPTG, isopropyl β-d-1-thiogalactopyranoside
Binding characterization of scFvs to the recombinant VacA
Binding profile of the soluble scFvs was characterized by immunoblotting against the recombinant p55 domain of the VacA toxin under reducing condition. As shown in Fig. 6, results of the western blot analyses confirmed retained binding affinity of the F10, G12, and A5 colonies to the VacA1 and F4 and H1 to the VacA2. The D1, specified clone with the significant binding capacity to the VacA1, determined no positive signal after visualization.
Peptide-scFv binding kinetic measurement by SPR analysis
For investigating the affinities of selected phage clones to VacA1 peptide, SPR analysis was performed. In the adsorption step, the RU test upon the clones D1, F10, and G12 confirmed the high affinity of the clones against the VacA1 peptide (Fig. S1). However, the clone A5 determined the same RU signal of the PBS, suggesting no affinity to the VacA1 peptide. The binding kinetics of each clone is summarized in Table 3.
Binding specificity analysis by flow cytometry
Immunofluorescent analysis was performed to investigate the binding of the scFvs to the membrane-associated VacA on H. pylori. ScFvs bound to the FITC-conjugated antibody demonstrated a fluorescent signal and were compared to the negative control sample containing no scFv. As shown in Fig. 7, all the isolated scFvs showed a fair binding capacity to H. pylori.
FACS analysis of isolated scFvs against membrane-associated VacA toxin of H. pylori bacteria. a–f The isolated scFvs named G12, D1, A5, F10, F4, and H1, respectively. The red curve in all histograms shows untreated bacteria and the black curve indicates the scFv-treated bacteria. VacA, vacuolating toxin A
Docking results evaluated by bioinformatic analysis
While the ligand binding analyses validated the specific binding of the isolated scFvs (Fig. S7 and Table S2), the docking between scFvs and VacA epitopes as ligands was carried out using the pepATTRACT server that performed a novel docking protocol in a fully blind manner for protein-peptide docking (Fig. 8). A summary of the CDR regions of the light and heavy chains of scFvs interacting with VacA ligand is shown in Table 4. Furthermore, as presented in Table 4, the sequences of scFVs were submitted to the DNA Data Bank of Japan (DDBJ). The structural and physiochemical details of each colony are presented in Figs. S2 to S8. The 3D structures and sequences of the isolated scFvs are shown in Fig. 9.
The 3D structures and sequences of the isolated scFvs. a, c, e, g, i, k The G12, D1, A5, F10, F4, and H1 clones, respectively. Sphere residues indicate the biggest pocket in each scFv. Red, blue, cyan, white, and dirty pink residues represent CDRs, residues consisting the pocket, residues of CDRs from scFvs that have overlap with pockets, residues from the myc-tag that involved in pocket formation, and residues from linker that have role in forming the packet, respectively. b, d, f, h, l Define sequences of the G12, D1, A5, F10, F4, and H1 scFvs, respectively. Highlighted regions indicate residues consisting pockets
Discussion
The VacA toxin is known as one of the most potent toxins of H. pylori, which is involved in the pathogenesis of the bacteria in the digestion system of hosts (Fig. 1). It is an essential part of the Ag-based diagnosis for patients with the peptic ulcer (Palframan et al. 2012). Despite the high importance of the VacA in the development of diseases such as peptic ulcer and gastric cancer, to the best of our knowledge, no VacA-based monoclonal antibody against all subtypes of VacA has been recognized. Perhaps, the high occurrence of polymorphisms in this overwhelming toxin has made it difficult to generate mAbs against the VacA species regardless of their diversity (Figura et al. 2015). Notwithstanding, almost most of the diagnostic methods of H. pylori infection are based on the histologic samples obtained from the infected patients. Such sampling is an invasive procedure and often associated with some errors, in large part due to an uneven distribution of H. pylori in different sections of the organ. Further, most of the methods used for the diagnosis of the infections seem to be somewhat imprecise, providing possible false outcomes. The Ab-based diagnosis procedures are deemed to minimize possible inaccuracies of the currently used detection methods in patients infected with H. pylori (Pourakbari et al. 2013).
In this study, we aimed to produce fully human scFvs by capitalizing on a semisynthetic PAL containing diverse scFvs and SPB procedure. Considering possible polymorphism(s) in VacA gene, two surfaces of conserved domains involved in the binding of the toxin to the host cell should be taken into account, which can result in possible isolation of scFvs against all types of the VacA toxin (Gangwer et al. 2007). Having considered such fact, we hypothesized that exploiting these regions would produce efficient scFvs for diagnosis of the H. pylori infection by the VacA toxin.
For the isolation of human scFvs against two conserved domains of VacA, the Tomlinson library I was applied using the SPB procedure. The SPB, which enables the reaction between target and phage particles on an affinity matrix, provides some advantages over the traditionally used immobilized selection methods. Among them, an enhanced accessibility of the target to scFv-containing phage particles and conformational endurance of the target due to lack of extreme immobilizing conditions causing target denaturation are counted as the advantages of SPB (Aghebati-Maleki et al. 2016). Further, as washing steps separate immobilized targets on a solid surface, the evaluation of the exact concentration of the target is often impossible/hard to be accomplished. Given that the SPB is supplemented with mobile Ags in a solution, the calculation of precise Ag amount based on the kinetic properties and capacity of the phage infection appears to be plausible (Ahmad et al. 2012; Tikunova and Morozova 2009). On the other hand, exploiting low volumes of the peptide with decreasing concentration on each round may result in isolation of high-affinity scFv fragments (Fellouse and Pal 2015).
The IMGT/V-QUEST analysis was used to detect any human germline gene similarity to the isolated scFvs (Tables 1 and 2). Further steps were applied for the validation of the scFvs isolated by biopanning steps (Fig. 2) using several techniques such as western blot and ELISA. Among various positive colonies of VacA1 and VacA2 resulted from the screening step, four colonies of VacA1 and two colonies of VacA2 indicated high optical density in the monoclonal scFv ELISA (Figs. 3, 4, and 5), even though the expression of the desired colonies in the suppressor strain, TG1, showed some difference of the OD when phage particles were loaded scFvs on their surface. This phenomenon seems to be a prevalent complication pertinent to the antibody phage display. The most frequent reason seems to be the presence of various stop codons inside the gene responsible for the expression of the scFvs; hence, the protein translated from the gene could be defective/destructed (Barderas et al. 2006; Tohidkia et al. 2012). The second cause may attribute to the nature of the ELISA in each method. In the soluble scFv ELISA, the produced signal is based on the interaction of the tags bound to scFv and reporting antibody, while the pvɪɪɪ coat protein of phage bound to scFvs is the substrate of the primary antibody, which is approximately 2700 times more frequent than the pɪɪɪ on the surface of the phage. As a result, the OD of the phage ELISA is much significant than the scFv ELISA (Jensen et al. 2002; Wang et al. 2004). It should be noted that the production of scFvs in the non-suppressor E. coli strain, HB2151, could provide high yielding antibody fragments in comparison with phage-scFv particles. Therefore, it may compensate the low absorbance of scFv clones in the scFv ELISA.
Further, the immunoblot analyses confirmed that three colonies of the VacA1 (i.e., A5-G12-F10) and two colonies of the VacA2 (F4 and H1) as strong VacA binders (Fig. 6). Given that the immunoblotting assay is on the basis of denaturing the 3D structure of the peptide, the D1 could possibly interact with the conformational epitope of the peptide, but not the linear structure (Forsstrom et al. 2015; Trott et al. 2014).
The functionality of the scFvs analyzed by the flow cytometry method confirmed that all the isolated scFvs show high cell-associated fluorescence signal when they are linked to the surface of the VacA toxin on the surface of H. pylori (Fig. 7). Furthermore, the binding affinity calculation performed by SPR analysis showed an overall dissociation equilibrium constants (KD) of 100 nM for all the scFvs (Table 3).
Despite the overall similarity between the scFv sequences, bioinformatic results showed that this small diversity in sequence might lead to the significant difference in conformational structures and consequently functional characterization of each scFv (Figs. S2, S6, and S8). Such sequence diversity causes the formation of the biggest pocket in different sites and also possible changes in the density of frustration index in scFv structures (Figs. S5 and S6). The aforementioned structural and consequently physicochemical diversity might eventuate difference in the tendency of CDR regions to target peptides (Table 4). For example, in silico docking result indicates only CDR3H of H1 scFv is involved in the interaction with the VacA2 peptide, while all CDRs from F10 were found to interact with the VacA1 (Fig. S8). Bioinformatic analyses confirmed almost all the experimental results; nonetheless, the D1 and A5 colonies showed somewhat deviation from the in silico results in western blot analysis for D1 and VacA1 and SPR analysis for A5 and VacA1. Despite that pocket finder analysis showed overlap between major scFv regions and pockets as cavities that define potential interacting capacity (Table S3), docking results proved that affinity for interacting antigen with CDR is more than the off-sequential regions such as myc-tag and linker taking part in cavity formation (Figs. 8 and S8). Taken all, the 3D structures and sequences of the isolated scFvs were fully determined (Fig. 9).
Two scenarios can be assumed by the comparison of bioinformatic results with the Western blot and SPR analyses. First, there is a false-negative result in western blot analysis of D1 and VacA1 SPR analysis for A5 and VacA1, even though in silico docking results confirmed desirable affinity of D1 and A5 to VacA2 and VacA1, respectively. Second, there is a false-positive response in SPR analysis of D1 and VacA2 and also a false-positive response in the western blotting analysis of A5 and VacA1.
It should be noted that a high in vitro blocking capacity of the IgAs has already been reported, in which IgA was shown to display high stability in the gastric environment with good avidity and to the membrane-associated proteins of H. pylori (Berdoz and Corthesy 2004). Further, new drug delivery agents resistant to mucus barrier of the stomach have brought new insights to the efficient delivery of antimicrobial agents (Jain et al. 2012; Jain and Jain 2013).
Altogether, it seems that the conventional treatments of the H. pylori-mediated gastric diseases are shifting towards immunopharmaceuticals. In brief, in this present study, we isolated scFvs against VacA toxin of H. pylori for the first time by means of a solution-phase biopanning. The isolated scFvs displayed high binding affinity confirmed by experimental and bioinformatic approaches. On the basis that a portion of the VacA is attached to the bacterial membrane and the high affinity of the isolated scFvs to the surface-exposed areas of the VacA toxin, we propose the selected scFvs as new Ab fragments that can be further developed towards both research and clinical applications in the H. pylori-infected patients.
References
Adler I, Muino A, Aguas S, Harada L, Diaz M, Lence A, Labbrozzi M, Muino JM, Elsner B, Avagnina A, Denninghoff V (2014) Helicobacter pylori and oral pathology: relationship with the gastric infection. World J Gastroenterol 20(29):9922–9935. https://doi.org/10.3748/wjg.v20.i29.9922
Aghebati-Maleki L, Bakhshinejad B, Baradaran B, Motallebnezhad M, Aghebati-Maleki A, Nickho H, Yousefi M, Majidi J (2016) Phage display as a promising approach for vaccine development. J Biomed Sci 23(1):66. https://doi.org/10.1186/s12929-016-0285-9
Ahmad ZA, Yeap SK, Ali AM, Ho WY, Alitheen NB, Hamid M (2012) scFv antibody: principles and clinical application. Clin Dev Immunol 2012:980250. https://doi.org/10.1155/2012/980250
Ayriss J, Woods T, Bradbury A, Pavlik P (2007) High-throughput screening of single-chain antibodies using multiplexed flow cytometry. J Proteome Res 6(3):1072–1082. https://doi.org/10.1021/pr0604108
Barderas R, Shochat S, Martinez-Torrecuadrada J, Altschuh D, Meloen R, Ignacio Casal J (2006) A fast mutagenesis procedure to recover soluble and functional scFvs containing amber stop codons from synthetic and semisynthetic antibody libraries. J Immunol Methods 312(1–2):182–189. https://doi.org/10.1016/j.jim.2006.03.005
Berdoz J, Corthesy B (2004) Human polymeric IgA is superior to IgG and single-chain Fv of the same monoclonal specificity to inhibit urease activity associated with Helicobacter pylori. Mol Immunol 41(10):1013–1022. https://doi.org/10.1016/j.molimm.2004.05.006
Bessede E, Dubus P, Megraud F, Varon C (2015) Helicobacter pylori infection and stem cells at the origin of gastric cancer. Oncogene 34(20):2547–2555. https://doi.org/10.1038/onc.2014.187
Brylinski M, Skolnick J (2008) A threading-based method (FINDSITE) for ligand-binding site prediction and functional annotation. Proc Natl Acad Sci U S A 105(1):129–134. https://doi.org/10.1073/pnas.0707684105
Capra JA, Laskowski RA, Thornton JM, Singh M, Funkhouser TA (2009) Predicting protein ligand binding sites by combining evolutionary sequence conservation and 3D structure. PLoS Comput Biol 5(12):e1000585. https://doi.org/10.1371/journal.pcbi.1000585
Capurso G, Carnuccio A, Lahner E, Panzuto F, Baccini F, Delle Fave G, Annibale B (2006) Corpus-predominant gastritis as a risk factor for false-negative 13C-urea breath test results. Aliment Pharmacol Ther 24(10):1453–1460. https://doi.org/10.1111/j.1365-2036.2006.03143.x
Carvalho MA, Machado NC, Ortolan EV, Rodrigues MA (2012) Upper gastrointestinal histopathological findings in children and adolescents with nonulcer dyspepsia with Helicobacter pylori infection. J Pediatr Gastroenterol Nutr 55(5):523–529. https://doi.org/10.1097/MPG.0b013e3182618136
Chey WD, Wong BC, Practice Parameters Committee of the American College of G (2007) American College of Gastroenterology guideline on the management of Helicobacter pylori infection. Am J Gastroenterol 102(8):1808–1825. https://doi.org/10.1111/j.1572-0241.2007.01393.x
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519. https://doi.org/10.1002/pro.5560020916
Cover TL, Holland RL, Blanke SR (2016) Helicobacter pylori vacuolating toxin. In: Backert S, Yamaoka Y (eds) Helicobacter pylori research: from bench to bedside. Springer, Tokyo, pp 113–141
Dundas J, Ouyang Z, Tseng J, Binkowski A, Turpaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34(Web Server):W116–W118. https://doi.org/10.1093/nar/gkl282
Eisenberg D, Luthy R, Bowie JU (1997) VERIFY3D: assessment of protein models with three-dimensional profiles. Methods Enzymol 277:396–404
Fahimi F, Tohidkia MR, Fouladi M, Aghabeygi R, Samadi N, Omidi Y (2017) Pleiotropic cytotoxicity of VacA toxin in host cells and its impact on immunotherapy. Bioimpacts 7(1):59–71. https://doi.org/10.15171/bi.2017.08
Fellouse FA, Pal G (2015) Methods for the construction of phage-displayed libraries In: Sidhu SS, Geyer CR (eds) Phage display in biotechnology and drug discovery. CRC Press
Figura N, Valassina M, Moretti E, Vindigni C, Collodel G, Iacoponi F, Giordano N, Roviello F, Marrelli D (2015) Histological variety of gastric carcinoma and Helicobacter pylori cagA and vacA polymorphism. Eur J Gastroenterol Hepatol 27(9):1017–1021. https://doi.org/10.1097/MEG.0000000000000414
Forsstrom B, Axnas BB, Rockberg J, Danielsson H, Bohlin A, Uhlen M (2015) Dissecting antibodies with regards to linear and conformational epitopes. PLoS One 10(3):e0121673. https://doi.org/10.1371/journal.pone.0121673
Gangwer KA, Mushrush DJ, Stauff DL, Spiller B, McClain MS, Cover TL, Lacy DB (2007) Crystal structure of the Helicobacter pylori vacuolating toxin p55 domain. Proc Natl Acad Sci U S A 104(41):16293–16298. https://doi.org/10.1073/pnas.0707447104
Garza-Gonzalez E, Perez-Perez GI, Maldonado-Garza HJ, Bosques-Padilla FJ (2014) A review of Helicobacter pylori diagnosis, treatment, and methods to detect eradication. World J Gastroenterol 20(6):1438–1449. https://doi.org/10.3748/wjg.v20.i6.1438
Georgieva Y, Konthur Z (2011) Design and screening of M13 phage display cDNA libraries. Molecules 16(2):1667–1681. https://doi.org/10.3390/molecules16021667
Hagymasi K, Tulassay Z (2014) Helicobacter pylori infection: new pathogenetic and clinical aspects. World J Gastroenterol 20(21):6386–6399. https://doi.org/10.3748/wjg.v20.i21.6386
Haque A, Tonks NK (2012) The use of phage display to generate conformation-sensor recombinant antibodies. Nat Protoc 7(12):2127–2143
Hensel F, Knorr C, Hermann R, Krenn V, Muller-Hermelink HK, Vollmers HP (1999) Mitogenic autoantibodies in Helicobacter pylori-associated stomach cancerogenesis. Int J Cancer 81(2):229–235
Jain AK, Agarwal A, Agrawal H, Agrawal GP (2012) Double-liposome-based dual-drug delivery system as vectors for effective management of peptic ulcer. J Liposome Res 22(3):205–214. https://doi.org/10.3109/08982104.2012.655284
Jain AK, Jain SK (2013) Development and characterization of nanolipobeads-based dual drug delivery system for H. pylori targeting. J Drug Target 21(6):593–603. https://doi.org/10.3109/1061186x.2013.784978
Jensen KB, Larsen M, Pedersen JS, Christensen PA, Alvarez-Vallina L, Goletz S, Clark BF, Kristensen P (2002) Functional improvement of antibody fragments using a novel phage coat protein III fusion system. Biochem Biophys Res Commun 298(4):566–573
Khoury GA, Tamamis P, Pinnaduwage N, Smadbeck J, Kieslich CA, Floudas CA (2014) Princeton_TIGRESS: protein geometry refinement using simulations and support vector machines. Proteins 82(5):794–814. https://doi.org/10.1002/prot.24459
Kohler G, Milstein C (1975) Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 256(5517):495–497
Krieger E, Koraimann G, Vriend G (2002) Increasing the precision of comparative models with YASARA NOVA—a self-parameterizing force field. Proteins 47(3):393–402
Krieger E, Vriend G (2014) YASARA view—molecular graphics for all devices—from smartphones to workstations. Bioinformatics 30(20):2981–2982. https://doi.org/10.1093/bioinformatics/btu426
Kuo SH, Yeh KH, Chen LT, Lin CW, Hsu PN, Hsu C, Wu MS, Tzeng YS, Tsai HJ, Wang HP, Cheng AL (2014) Helicobacter pylori-related diffuse large B-cell lymphoma of the stomach: a distinct entity with lower aggressiveness and higher chemosensitivity. Blood Cancer J 4:e220. https://doi.org/10.1038/bcj.2014.40
Lim SL, Canavarro C, Zaw MH, Zhu F, Loke WC, Chan YH, Yeoh KG (2013) Irregular meal timing is associated with Helicobacter pylori infection and gastritis. ISRN Nutr 2013:714970. https://doi.org/10.5402/2013/714970
Liu JK (2014) The history of monoclonal antibody development—progress, remaining challenges and future innovations. Ann Med Surg (Lond) 3(4):113–116. https://doi.org/10.1016/j.amsu.2014.09.001
Oleastro M, Menard A (2013) The role of Helicobacter pylori outer membrane proteins in adherence and pathogenesis. Biology (Basel) 2(3):1110–1134. https://doi.org/10.3390/biology2031110
Palframan SL, Kwok T, Gabriel K (2012) Vacuolating cytotoxin A (VacA), a key toxin for Helicobacter pylori pathogenesis. Front Cell Infect Microbiol 2:92. https://doi.org/10.3389/fcimb.2012.00092
Papini E, Zoratti M, Cover TL (2001) In search of the Helicobacter pylori VacA mechanism of action. Toxicon 39(11):1757–1767
Parra RG, Schafer NP, Radusky LG, Tsai MY, Guzovsky AB, Wolynes PG, Ferreiro DU (2016) Protein Frustratometer 2: a tool to localize energetic frustration in protein molecules, now with electrostatics. Nucleic Acids Res 44(W1):W356–W360. https://doi.org/10.1093/nar/gkw304
Patel SK, Pratap CB, Jain AK, Gulati AK, Nath G (2014) Diagnosis of Helicobacter pylori: what should be the gold standard? World J Gastroenterol 20(36):12847–12859. https://doi.org/10.3748/wjg.v20.i36.12847
Pourakbari B, Ghazi M, Mahmoudi S, Mamishi S, Azhdarkosh H, Najafi M, Kazemi B, Salavati A, Mirsalehian A (2013) Diagnosis of Helicobacter pylori infection by invasive and noninvasive tests. Braz J Microbiol 44(3):795–798. https://doi.org/10.1590/S1517-83822013005000052
Pritchard DM, Crabtree JE (2006) Helicobacter pylori and gastric cancer. Curr Opin Gastroenterol 22(6):620–625. https://doi.org/10.1097/01.mog.0000245539.50765.f6
Reichert JM (2012) Marketed therapeutic antibodies compendium. MAbs 4(3):413–415. https://doi.org/10.4161/mabs.19931
Rieder G, Fischer W, Haas R (2005) Interaction of Helicobacter pylori with host cells: function of secreted and translocated molecules. Curr Opin Microbiol 8(1):67–73. https://doi.org/10.1016/j.mib.2004.12.004
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5(4):725–738. https://doi.org/10.1038/nprot.2010.5
Roy A, Yang J, Zhang Y (2012) COFACTOR: an accurate comparative algorithm for structure-based protein function annotation. Nucleic Acids Res 40(Web Server issue):W471–W477. https://doi.org/10.1093/nar/gks372
Salama NR, Hartung ML, Muller A (2013) Life in the human stomach: persistence strategies of the bacterial pathogen Helicobacter pylori. Nat Rev Microbiol 11(6):385–399. https://doi.org/10.1038/nrmicro3016
Schindler CE, de Vries SJ, Zacharias M (2015) Fully blind peptide-protein docking with pepATTRACT. Structure 23(8):1507–1515. https://doi.org/10.1016/j.str.2015.05.021
Schrodinger, LLC (2015) The PyMOL Molecular Graphics System, version 1.8
Siddiqui MZ (2010) Monoclonal antibodies as diagnostics; an appraisal. Indian J Pharm Sci 72(1):12–17. https://doi.org/10.4103/0250-474X.62229
Sidhu SS (2001) Engineering M13 for phage display. Biomol Eng 18(2):57–63
Tikunova NV, Morozova VV (2009) Phage display on the base of filamentous bacteriophages: application for recombinant antibodies selection. Acta Nat 1(3):20–28
Tohidkia MR, Barar J, Asadi F, Omidi Y (2012) Molecular considerations for development of phage antibody libraries. J Drug Target 20(3):195–208. https://doi.org/10.3109/1061186X.2011.611517
Trott M, Weibeta S, Antoni S, Koch J, von Briesen H, Hust M, Dietrich U (2014) Functional characterization of two scFv-Fc antibodies from an HIV controller selected on soluble HIV-1 Env complexes: a neutralizing V3- and a trimer-specific gp41 antibody. PLoS One 9(5):e97478. https://doi.org/10.1371/journal.pone.0097478
Verma R, Boleti E, George AJ (1998) Antibody engineering: comparison of bacterial, yeast, insect and mammalian expression systems. J Immunol Methods 216(1–2):165–181
Wang X, Campoli M, Ko E, Luo W, Ferrone S (2004) Enhancement of scFv fragment reactivity with target antigens in binding assays following mixing with anti-tag monoclonal antibodies. J Immunol Methods 294(1–2):23–35. https://doi.org/10.1016/j.jim.2004.08.005
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(Web Server):W407–W410. https://doi.org/10.1093/nar/gkm290
Wu S, Zhang Y (2007) LOMETS: a local meta-threading-server for protein structure prediction. Nucleic Acids Res 35(10):3375–3382. https://doi.org/10.1093/nar/gkm251
Yan R, Xu D, Yang J, Walker S, Zhang Y (2013) A comparative assessment and analysis of 20 representative sequence alignment methods for protein structure prediction. Sci Rep 3:2619. https://doi.org/10.1038/srep02619
Yang J, Roy A, Zhang Y (2013) Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics 29(20):2588–2595. https://doi.org/10.1093/bioinformatics/btt447
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43(W1):W174–W181. https://doi.org/10.1093/nar/gkv342
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9:40. https://doi.org/10.1186/1471-2105-9-40
Zhang Y, Skolnick J (2004) SPICKER: a clustering approach to identify near-native protein folds. J Comput Chem 25(6):865–871. https://doi.org/10.1002/jcc.20011
Zhao A, Tohidkia MR, Siegel DL, Coukos G, Omidi Y (2016) Phage antibody display libraries: a powerful antibody discovery platform for immunotherapy. Crit Rev Biotechnol 36(2):276–289. https://doi.org/10.3109/07388551.2014.958978
Acknowledgments
The authors wish to thank all the members of the cell culture facility at the Research Center for Pharmaceutical Nanotechnology at Tabriz University of Medical Sciences for their help. The authors also thanks Dr. Morteza Eskandani, Mr. Abolfazl Barzegari, Ms. Farzaneh Fathi, Dr. Morteza Milani, Dr. Mohammadreza Sadeghi and Dr. Ailar Nakhlband for their assistance during this research.
Funding
The study was funded by postgraduate grant of the Research Center for Pharmaceutical Nanotechnology (RCPN) at Tabriz University of Medical Sciences (RCPN-93005; recipient Dr. M.R. Tohidkia). This work was a joint project for a postgraduate thesis between the RCPN and the School of Advanced Biomedical Sciences at Tabriz University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
The present study does not contain any experiments in relation to either human participants or animal models by any of the authors.
Electronic supplementary material
ESM 1
(DOCX 24.5 MB)
Rights and permissions
About this article
Cite this article
Fahimi, F., Sarhaddi, S., Fouladi, M. et al. Phage display-derived antibody fragments against conserved regions of VacA toxin of Helicobacter pylori. Appl Microbiol Biotechnol 102, 6899–6913 (2018). https://doi.org/10.1007/s00253-018-9068-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9068-4