Abstract
Cut190 from Saccharomonospora viridis AHK190 (Cut190) is the only cutinase that exhibits inactive (Ca2+-free) and active (Ca2+-bound) states, although other homologous cutinases always maintain the active states (Ca2+-free and bound). The X-ray crystallography of the S176A mutant of Cut190* (Cut190_S226P/R228S) showed that three Ca2+ ions were bound at sites 1–3 of the mutant. We analyzed the roles of three Ca2+ ions by mutation and concluded that they play different roles in Cut190* for activation (sites 1 and 3) and structural and thermal stabilization (sites 2 and 3). Based on these analyses, we elucidated the mechanism for the conformational change from the Ca2+-free inactive state to the Ca2+-bound active state, proposing the novel Ca2+ effect on structural dynamics of protein. The introduction of a disulfide bond at Asp250 and Glu296 in site 2 remarkably increased the melting temperatures of the mutant enzymes by more than 20–30 °C (while Ca2+-bound) and 4–14 °C (while Ca2+-free), indicating that a disulfide bond mimics the Ca2+ effect. Replacement of surface asparagine and glutamine with aspartic acid, glutamic acid, or histidine increased the melting temperatures. Engineered mutant enzymes were evaluated by an increase in melting temperatures and kinetic values, based on the hydrolysis of poly(butylene succinate-co-adipate) and microfiber polyethylene terephthalate (PET). A combined mutation, Q138A/D250C-E296C/Q123H/N202H, resulted in the highest thermostability, leading to the maximum degradation of PET film (more than 30%; approximately threefold at 70 °C, compared with that of Cut190* at 63 °C).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Aromatic polyesters are recalcitrant for enzymatic hydrolysis, owing to their high glass transition temperatures (Tg). Aromatic polyesters are primarily polyethylene terephthalate (PET) and have a high Tg, which is approximately 75–80 °C in air, but decreases to 60–65 °C in aqueous solution (Kawai et al. 2014). At temperatures above Tg, even the brittle amorphous regions of PET become flexible and more accessible to an enzymatic attack. In fact, the enzymatic hydrolysis of PET film by cutinases has been verified at temperatures higher than 60 °C, preferably ≥ 65 °C (Kawai et al. 2014; Herrero Acero et al. 2011; Sulaiman et al. 2014). The effect of Ca2+ on the thermal stability has been shown with regard to several thermostable cutinases from actinomycetes (Kawai et al. 2013, 2014; Sulaiman et al. 2014; Thumarat et al. 2012; Miyakawa et al. 2015; Kitadokoro et al. 2012). We have analyzed the crystal structures of a mutant of Cut190 from Saccharomonospora viridis AHK 190 (Cut190_S226P) in the Ca2+-free (inactive form; PDB ID: 4WFI) and Ca2+-bound (active form; PDB IDs: 4WFJ and 4WFK) states, which were accompanied by dynamic conformational changes in the protein structure (Miyakawa et al. 2015). Homologous cutinases such as Est119 (PDB IDs: 3VIS and 3WYN), Tf_Cut2 (PDB IDs: 4CG1-3), LC-cutinase (PDB ID: 4EB0) and Thc_Cut1 and Cut2 (PDB IDs: LUI and LUJ) maintained constant overall structures, either in the presence or absence of Ca2+. Therefore, only Cut190 displayed active (Ca2+-bound) (PDB IDs: 4WFJ and 4WFK) and inactive (Ca2+-free) (PDB ID: 4WFI) states, although the active state overlapped with Est119 and the others (PDB IDs: 4EB0, 3VIS, 3WYN, 4CG1-3, and 5LUI-L) (Sulaiman et al. 2014; Kitadokoro et al. 2012; Kawai et al. 2013; Roth et al. 2015; Ribitsch et al. 2017). Their activities and thermal stabilities increased in the presence of Ca2+ (Kawai et al. 2013, 2014; Sulaiman et al. 2014; Thumarat et al. 2012; Then et al. 2015; Miyakawa et al. 2015). Est119 maintained the same overall structure with or without Ca2+ (PDB IDs: 3WYN, 3VIS). Cutinase 1 or 2 from Thermobifida cellulosilytica (Thc_Cut1 and Thc_Cut2) and Thc_Cut2 mutants, either bound or not bound to Mg2+ or Ca2+ (PDB IDs: 5LUI-5LUL), also maintained the same overall structures. The Ca2+-binding site for thermal stabilization in 3WYN was different from the Ca2+-binding site necessary for the activation of Cut190 (Miyakawa et al. 2015). Very recently, we analyzed three Ca2+-binding sites of the mutant S176A of Cut190_S226P/R228S (Cut190*) (PDB ID: 5ZNO), based on molecular dynamics simulations to show that the Ca2+-bound structure fluctuated less than the Ca2+-free structure (Numoto et al. 2018).
In this paper, we analyzed the roles of the Ca2+ ions bound to sites 1–3 of Cut190* and elucidated the conformational change from the Ca2+-free inactive state to the Ca2+-bound active state. We also replaced the surface glutamine and asparagine of Cut190*, which are prone to deamidation, with glutamic acid, aspartic acid, alanine, or histidine to improve the activity and thermostability and finally constructed a robust mutant, which achieved the best degradation rate of PET film at 70 °C.
Materials and methods
Materials
The poly(butylene succinate-co-adipate) (PBSA) suspension (BIONOLLE™ EM-301; weight-average Mw = 1.0 × 105) is a product of Showa Denko K. K. (Tokyo, Japan). A 0.25-mm-thick film of amorphous PET (PET-GF; crystallinity of 6.3%) was obtained from Goodfellows Cambridge, Ltd. (Tokyo, Japan). The PET package (approximately 0.6 mm thick; PET-S; crystallinity of 8.4%) for PC peripherals (Sanwa Supply Inc., Okayama, Japan) was also used for degradation tests, as the same material used in the previous report (Sulaiman et al. 2014). APEXA 4027™ film was prepared by heat-cast (approximately 40-μm thick) from APEXA 4027™ resin (Dupont, Co., Ltd., Tokyo, Japan) (Kawai et al. 2017). Microfiber PET was prepared as follows: PET pellets (Teijin Ltd., Osaka, Japan) were solved in 1,1,1,3,3,3-hexafluore-2-propanol at 10 wt%, which was extruded from a metallic nozzle (0.2 mm inner diameter) at a constant rate of 1.0 μl/min using a micro-feeder pump (Furue Science, Kawaguchi, Saitama, Japan). The electospinning conditions were set at the applied voltage of 10 kV using a high-voltage supply device Model-600F (Pulse Electronics Engineering Co., Ltd., Noda, Chiba, Japan) and at the distance of 20 cm between the nozzle and the metal target. The diameter of obtained microfiber was approximately 1–3 μm. The density of microfiber was approximately 1.3 mg/cm2. The microfiber PET is considered as approximately 100% amorphous state. Bis(2-hydroxyethyl) terephthalate (BHET) is a product of Tokyo Chemical Industry Co., Ltd. (Tokyo, Japan). Mono(2-hydroxyethyl) terephthalate (MHET; CAS No. 1137-99-1) was synthesized as described in the Supporting Information (Scheme S1). All other chemicals were the highest grade available through commercial routes.
Site-directed mutagenesis and expression and purification of mutant enzymes
Site-directed mutagenesis was performed to introduce a specific mutation using a KOD plus a mutagenesis kit (TOYOBO, Osaka, Japan) according to the manufacturer’s instructions. Cut190* was used as the template enzyme in this paper. The PCR products were obtained using pQE80L-Cut190* as the DNA template and specific primers for the site-directed mutagenesis (Kawabata et al. 2017). The parental pQE80L-Cut190* plasmid was removed via digestion with DpnI, and the PCR products were transformed directly into Escherichia coli Rosetta-gami B (DE3) cells. Mutagenesis results were verified by sequencing. Mutant enzymes were expressed and purified under the same conditions as described previously (Kawai et al. 2014). The SDS-PAGE analysis revealed that the purities of the enzymes were greater than 90%. The protein concentration was determined using a Bio-Rad protein assay kit (Bio-Rad Laboratories, Hercules, CA, USA), with bovine serum albumin as a standard.
Enzymatic activity measurements
As described previously, Cut190* can act on both PBSA and PET (Kawai et al. 2014) and results of 3D modeling of Cut190* in complexes with PET and PBSA models were almost identical (Kawabata et al. 2017), indicating the usefulness of PBSA for a conventional assay of Cut190*. Actually the enzyme reaction with PBSA followed general enzymatic reaction curves, which were converted to kinetic values following Michaelis-Menten equation. The enzymatic activity was quantitatively measured by monitoring the decrease in the turbidity of a PBSA suspension at 600 nm (Kawai et al. 2014). For the PBSA assay, the reaction mixture included PBSA, 50 mM Tris-HCl (pH 8.2), 2.5 mM Ca2+, and an appropriate quantity of Cut190* or its mutants (0.9–1.3 μM in general), in a total volume of 2 ml. The reaction mixture was incubated at 37 °C for 30 min, with gentle shaking in a water bath. An absorbance of 1 at 600 nm corresponded to 0.25 mg dry weight PBSA/ml and a decrease of 1 OD600 of the PBSA suspension was calculated to correspond to 1.5 nkat (Kawai et al. 2014). Kinetic values were calculated from Michaelis-Menten plots. The value of an enzyme reaction at each PBSA concentration was subtracted from the control value that did not include the enzyme, and the net decrease in the OD600 was subjected to the calculation described above. The measurements were performed in duplicate or triplicate.
The enzyme activity was also measured with microfiber amorphous PET. The reaction mixture contained Tris buffer (pH 8.5), 100 μmol; CaCl2, 2.5 μmol; glycerol, 24%; enzyme, 2 nmol; and microfiber PET, approximately 2 mg (10.4 μmol as MHET unit) in a total volume of 1 ml, which was incubated at 65 °C for 15 h with shaking. An aliquot of the supernatant was withdrawn, diluted with 50 mM Tris buffer (pH 7.0), and subjected to terephthalic acid (TPA) measurements by absorbance at 240 nm (ε = 1.38 × 104). Degradation products such as BHET, MHET, and TPA were confirmed by high-performance liquid chromatography (HPLC). The aliquots of the reaction mixtures were withdrawn, filtered with 0.2 μm pore size and subjected to a HPLC system LC-20AB with a system controller SCL-10A (Shimadzu, Kyoto, Japan): Analytical conditions were as follows: detection, 240 nm with a Shimadzu UV-VIS detector SPD-10AV; column, TOSOH TSK-GEL ODS-100S (4.6 × 150 mm, TOSOH, Tokyo, Japan); eluent, a mixture of 0.3% formic acid/20% acetonitrile/Milli-Q water; flow rate, 1.0 ml/min; temperature, 25 °C; injection volume, 200 μl. MHET, BHET, and TPA were detectable at 1 μM.
Circular dichroism (CD) analysis and measurement of melting temperature
CD measurements were obtained using a model J-725 Jasco spectropolarimeter (JASCO Corp., Tokyo, Japan). The far-UV CD spectra were obtained at 20 °C for protein concentrations of 0.04 mg/ml in 10 mM Tris-HCl buffer (pH 8.0) in quartz cells with path length of 10 mm. No serious change was found with the spectra of all the mutants, suggesting that the secondary structures of the enzymes were not changed by mutation. Thermal denaturation curves were recorded in temperature mode at 222 nm from 20 to 90 °C with a heating rate of 1.0 °C/min. The analysis of the transition curves obtained was performed on the basis of two-state transition model, as described previously (Inaba et al. 2013) to determine melting temperature (Tm).
Degradation tests of PET films
PET films were cut into a circle of approximately 6 mm ϕ (PET-GF and PET-S) or 1-cm2 (APEXA 4027™) and incubated in a 1-ml reaction mixture including 100 mM Tris-HCl buffer (pH 8.2) or HEPES buffer (pH 8.5), 2.5 mM CaCl2, 24% glycerol, and approximately 2 μM Cut190* or its mutants at different temperatures under shaking. Prior to the incubation, PET films were incubated at indicated temperatures for 1 h in a reaction mixture (750 μ) that the enzyme was omitted and cooled down to room temperature. Then the enzyme and the films were mixed at room temperature and kept for 30 min and the incubation was started at indicated temperatures. An aliquot of the reaction mixture was withdrawn at intervals and diluted with 25 mM Tris-HCl buffer (pH 7.0); the absorbance of which was measured at 240 nm, as described above with microfiber PET. The contribution of the enzyme to the absorbance at 240 nm was negligible. After the incubation, the films were withdrawn and washed with 0.2% sodium carbonate and 0.1% nonionic surfactant (Brij 35) to remove proteins before a final wash with 50% ethanol following the method previously described (Silva et al. 2011). The films were dried below 60 °C overnight prior to weight loss and scanning electron microscope (SEM) measurements using an S-300 N scanning electron microscope (Hitachi Ltd., Tokyo, Japan) (Kawai et al. 2014).
Amino acid sequence accession number
The amino acid sequence of Cut190 (304 amino acids including the signal peptide sequence of 44 amino acids) was deposited to the GenBank under the accession number of BA042836.1.
Results
Structural and functional analyses of Cut190* at three Ca2+-binding sites
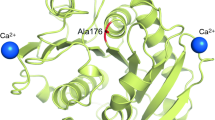
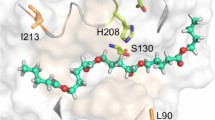
As found in PDB ID: 5ZNO, the amino acids involved in the three Ca2+-binding sites of Cut190* were identified at site 1 (Ser76, Ala78, and Phe81), site 2 (Glu220, Asp250, and Glu296), and site 3 (Asp204 and Thr206), as shown in Fig. 1a. We analyzed the role of Ca2+ at each site by mutating amino acids involved in Ca2+ binding.
The crystal structure of Cut190*S176A in complex with Ca2+ (PDB ID; 5ZNO). The orange spheres indicate Ca2+ molecules. The side chains of residues indicated are shown by stick models. The figure was generated by PyMOL. a Overall structure. b The structure around the second Ca2+-binding site. c The structure around the third Ca2+-binding site
Site 1 (Ser76, Ala78, and Phe81)
As previously described (Miyakawa et al. 2015), Ca2+ is indispensable for the activation of Cut190. On the other hand, CD measurements of Cut190* displayed the increased Tm values with increased Ca2+ concentrations (Fig. 2). Therefore, it should be concluded that Ca2+ plays different roles in Cut190, that is, activation and thermal stabilization. As described previously (Miyakawa et al. 2015), the Ser76-Phe81 region of the β1-β2 loop is relevant for the activation of Cut190 by Ca2+. Ser76-Phe81 and Phe106 (oxyanion hole-forming amino acid) are positioned differently in the Ca2+-free (PDB ID: 4WFI) and the Ca2+-bound (PDB ID: 4WFK) states of Cut190, although Arg70-Phe76 (corresponding to Ser76-Phe81) and Tyr99 (corresponding to Phe106) are kept in the same positions in the Ca2+-free (PDB ID: 3VIS) and the Ca2+-bound (PDB ID: 3WYN) states of Est119 (Fig. S1). The significant difference between Cut190 and the other cutinases is the deficiency of one amino acid in the β1-β2 loop of Cut190, which corresponds to Gly72 and Gly38 of Est119 and Tc_Cut2 (serine is at the same positions in Tf_Cut2 and LCC), respectively (Fig. S2). We introduced a serine between Phe77 and Ala78 of Cut190* (78S) to increase the number of amino acids in the β1-β2 loop, which showed sufficient activity even in the absence of Ca2+ (Table 1), although the activity was still remarkably enhanced by Ca2+. The expression of 78S reached a maximum 2 h after addition of isopropyl β-D-1-thiogalactopyranoside (IPTG), and the yield was less than 10% of that of Cut190* incubated for 12–15 h after IPTG addition. Longer incubation after IPTG addition resulted in a decrease in the soluble expression and an increase in insoluble expression, suggesting that the mutant (78S) was expressed as an inappropriate protein structure.
Site 2 (Glu220, Asp250, and Glu296)
The amino acids involved in the second Ca2+-binding site (Glu220, Asp250, and Glu296; Fig. 1b) are conserved among homologous cutinases (Fig. S2) and must be related to the thermostability (Thumarat et al. 2012). Based on previous reports demonstrating that an engineered disulfide bridge mimics the effect of calcium on proteases and a cutinase (Duerschmidt et al. 2005; Then et al. 2016), we replaced Asp250 and Glu296 with cysteines to introduce a disulfide bond (D250C-E296C). D250C-E296C showed an increased Tm value in the Ca2+-free state (Table 1), suggesting that an engineered disulfide bridge at D250C-E296C also contributes to the thermal stabilization, instead of Ca2+ binding. As previously reported (Kawabata et al. 2017), we generated two mutants of Cut190* with significantly increased activities, Q138A and I224A. To intensify these mutants by the introduction of the disulfide bond, we compared the abilities of both mutants to hydrolyze PET, using microfiber PET, as shown in Table 2. Microfiber PET was completely degraded to TPA by Q138A (Fig. S3). Q138A showed a higher degradation rate than Cut190*, but the degradation rate of I224A was lower than that of Cut190*. These results could be explained by the fact that the Q138A mutant showed lower binding energies and larger binding surface areas by modeling with PET, but the I224A mutant showed similar energies and slightly larger surface areas, compared with Cut190* (Kawabata et al. 2017). Based on these results, we constructed Q138A/D250C-E296C, which showed approximately the same Tm value as D250C-E296C and a better kcat/Km than that of D250C-E296C, as shown in Table 1. Further, we replaced Glu220 of Q138A/D250C-E296C with arginine (Q138A/D250C-E296C/E220R) and then Gly251 of Q138A/D250C-E296C/E220R with aspartic acid to construct a salt bridge between Arg220 and Asp251 (Q138A/D250C-E296C/E220R-G251D). The three mutants with D250C-E296C had no activity in the absence of Ca2+, that is, site 2 is unrelated to activation. On the other hand, all of the mutants displayed Tm values that increased by 20–23 °C (compared with Cut190* in the absence of Ca2+) and 4–10 °C (compared with Cut190* in the presence of Ca2+). Except Q138A/D250C-E296C/E220R, other mutants with D250C-E296C showed rather close Tm values with and without Ca2+. As Ca2+ is requisite to activation, decrease of Tm values in the presence of Ca2+ is unfavorable. Taken together, Q138A/D250C-E296C was considered the best mutant with regard to its activity and thermal stability.
Site 3 (Asp204, Thr206, and Gln209)
Ca2+ was bound directly to Asp204 and Thr206 and indirectly to Gln209 via water (Fig. 1c); the binding of Ca2+ at the third site has not been reported to date. Glu184 is located close to Asp204 (Fig. 1c). We constructed E184R, which was considered to form a salt bridge with Asp204 and might be efficient for stabilization. E184R showed an increased kcat/Km, but a reduced Tm value, suggesting the inhibition of Ca2+ binding. On the other hand, E184R unexpectedly showed sufficient activity in the absence of Ca2+ as well as S78 (Fig. 3).
Modification of the surface asparagine and glutamine
Glutamine and asparagine are known to be prone to deamidation, leading to the instability of proteins (Kaneko et al. 2005; Wright 1991). For the purpose of improving the thermostability, we replaced the surface glutamines and asparagines of Cut190* with glutamates and asparatates, respectively, or alanine. The kinetic properties of the mutants were compared with those of Cut190*, as shown in Table 1. Q110A resulted in no expression of the protein, although Q110E was expressed normally and showed a slightly increased Tm value. N168D was expressed normally, but its kinetic values and Tm were approximately equal to those of Cut190*. On the other hand, N168A exhibited expression that was approximately half of the normal expression of Cut190*, whereas approximately half of the protein was expressed in insoluble forms. Therefore, the replacement of Asn and Gln with Ala seemed to be inappropriate. Q141E resulted in an increased Tm value, but Q138A/Q141E had a reduced Tm value and kcat/Km. Q110E, N133D, and Q209E had Tm values that increased by 3–5 °C. Both Gln123 and Asn202 in Cut190 were replaced with histidine in other cutinases (Fig. S2). Both Q123H and N202H showed increased Tm values by 2.7 and 6.5 °C, respectively. Taken together with the results of the Ca2+ binding, we finally constructed Q138A/D250C-E296C/Q123H/N202H, which showed reasonable kinetic values and the highest Tm values either with or without Ca2+.
Comparison of PET hydrolysis by Cut190* mutants
We have already reported that Cut190* can degrade PET films at 63 °C for 3 days; the weight losses calculated by the TPA measurements were 10.9 ± 1.5% for PET-GF and 26.2 ± 0.6% for PET-S (Kawai et al. 2014). Here, we tested the degradation rates of two PET films and one modified PET film (APEXA 4027™) with Cut190* at different temperatures (Fig. 4). PET-GF degradation increased with higher reaction temperatures (50–65 °C), but PET-S degradation was maximal at 60 °C and decreased at 65 °C. APEXA 4027™ was degraded at temperatures less than 55 °C.
Q138A/D250C-E296C displayed a Tm value of 77.9 °C without Ca2+ (Table 1), suggesting an increase in the degradation rate of PET-GF at temperatures higher than 63 °C, which has been previously shown (Kawai et al. 2014). As expected, the mutant showed a higher degradation rate of PET-GF at 70 °C (Table 3). Apparent surface erosion was observed by SEM (Fig. 5a). As weight loss of the enzyme-treated film was approximately in accordance with the degradation rate based on the absorbance at 240 nm, as found in a previous report (Kawai et al. 2014), the degradation rate was measured by the absorbance thereafter. Q138A/D250C-E296C/E220R-G251D showed the decreased degradation of PET-GF at 70 °C, although its kcat/Km on PBSA at 37 °C was almost identical to that of Q138A/D250C-E296C. Introduction of Q123H/N202H improved the degradation rates of PET-GF by Q138A/D250C-E296C and Q138A/D250C-E296C/E220R-G251D (Table 3 and Fig. 5b). According to the increase of the degradation rate from 25% at 70 °C by Cut190*(Q138A/D250C-E296C) to 34% at 70 °C by Cut190*(Q138A/D250C-E296C/Q123H/N202H), the appearance of the film surface was dramatically changed with a far uneven structure. To prevent product inhibition, the reaction volumes were changed to 1.0–3.0 ml with the same enzyme concentration (2 μM), but the total amount of the degradation products showed no significant differences. Therefore, no product inhibition, up to approximately 15 mM (PET-GF; 70 °C), was suggested.
Discussion
X-ray crystallography of a mutant S176A of Cut190* displayed binding of three Ca2+ ions at different sites (PDB ID: 5ZNO), as shown in Fig. 1. The first Ca2+-binding site is the same as previously reported (Miyakawa et al. 2015). The main reason for the inactive state (Ca2+-free) of Cut190* is considered due to the deficiency of one amino acid in the β1-β2 loop, compared with other homologous cutinases maintaining the constant active state. Introduction of serine between Phe77 and Ala78 of Cut190* led to sufficient activity even in the absence of Ca2+, the activity of which was enhanced by the presence of Ca2+. However, the expression level of the mutant was only less than 10% of that of Cut190*, indicating the inappropriate protein structure, as found in some mutants relevant to substrate binding (Kawabata et al. 2017). This has to be caused by replacement of Arg76, which is conserved in other cutinases with serine in Cut190*. Arginine must prevent Ca2+ binding at this site and at the same time stabilize the β1-β2 loop by a cation-π interaction (stronger than a salt bridge) with Phe77, in which the cationic side chain of arginine typically interacts with the center of the electron-rich aromatic ring (Dougherty 1996, 2007) (Fig. S4). Instead, Cut190 stabilizes the β1-β2 loop by Ca2+ binding, which fixes the big α3 helix and pulls β2 and β4 sheets up, resulting in the open conformation of Phe106 in the β3-α2 loop (Miyakawa et al. 2015).
The amino acids involved in the second Ca2+-binding site (Glu220, Asp250, and Glu296) are conserved among homologous cutinases (Fig. S2) and must be related to thermostability (Thumarat et al. 2012). The mediation of thermostability by Ca2+ has been reported with regard to thermophilic protease (Smith et al. 1999; Teplyakov et al. 1990). As Ca2+ preferentially binds carboxylate and other oxygen ligands (most likely to be located on the protein surface), Ca2+ is considered to play a significant stabilizing role in proteins (Vieille and Zeikus 2001). Q138A degraded microfiber PET better than Cut190* (Table 2), suggesting that Q138A might be better as a template for further mutation. Actually Q138A/D250C-E296C showed approximately the same Tm value as D250C-E296C and a better kcat/Km than that of D250C-E296C. Thermal stabilization effect by introduction of a disulfide bridge between Asp250 and Glu296 is most probably due to the fixing of flexible β7-α6 and α8-β9 loops. There are many reports on the increased thermal stability by introduction of a disulfide bond, which is reasonable from the viewpoint of less entropy in a protein unfolded state. The Tm values of Cut190* increased with Ca2+ (0~25 mM), as shown in Fig. 2, but those of the D250C-E296C mutants were almost constant with and without Ca2+, suggesting that the introduced disulfide bond mimic the Ca2+ effect for thermostabilization.
The binding of Ca2+ at the third site has not been reported so far. As Glu184 is located close to Asp204, E184R was considered to form a salt bridge with Asp204 and might be efficient for stabilization. However, E184R showed a reduced Tm value, suggesting the inhibition of Ca2+ binding. Ca2+ binding at the third site must fix the long flexible β6-β7 loop, resulting in the fixing of the α5 helix, which reinforces the Ca2+ effect at site 1 to fix the α3 helix. On the other hand, E184R unexpectedly showed sufficient activity in the absence of Ca2+ as well as S78, as shown in Fig. 3. Glu184 was somewhat closer to Gly140 than to Asp204 (Fig. 1c). In other homologous cutinases (Est119, Tc_Cut2, Tf_Cut2, and LCC), Arg exists at the same position as Glu184, and Glu exists at the same position as Gly140, making a stronger salt bridge (Fig. S2). A certain interaction between E184R and Glu140 must fix the α5 (E184R) and α3 (Gly140) helices, which should serve as an alternative to Ca2+ binding at site 1. In conclusion, the differences in the first and third Ca2+-binding sites of Cut190 compared to other homologous cutinases caused the inactive state of Cut190, which were compensated by Ca2+ binding that lead to the active state, whereas other homologous cutinases maintained constant active states.
Glutamine and asparagine are known to be prone to deamidation, leading to instability of proteins (Kaneko et al. 2005; Wright 1991). Q141E, N133D, Q110E, and Q209E had Tm values that increased by 3–5 °C, but N168D had approximately the same Tm value as Cut190*. Asn133 is involved in the first Ca2+ binding in Cut190_S26P (Miyakawa et al. 2015), but N133D showed no inhibition on the activity and the slightly increased Tm value. This is not surprising, as the carbonyl group of the main chain of Asn133 is involved in Ca2+ binding in site 1. Gln110 (NH2) was suggested to interact with Asp131 (COOH) in Ca2+-bound state (Miyakawa et al. 2015), but Q110E showed approximately the same kinetic values and the rather increased Tm value. In a mutant S176A of Cut190*, Gln110 (CO in CONH2) seemed to interact with Asp131 (NH in a peptide bond), thereby intensifying Q110E-Asp131 interaction (PDB ID: 5ZNO). Both Gln123 and Asn202 in Cut190 were replaced with histidine in other cutinases (Fig. S2). Both Q123H and N202H showed the increased Tm values by 2.7 °C and 6.5 °C, respectively. Q138A/D250C-E296C/Q123H/N202H showed the highest Tm values either with or without Ca2+.
Degradation of PET-GF, PET-S, and Apexa 4027 films was compared with Cut190* at different temperatures (Fig. 4). PET-GF degradation increased with higher reaction temperatures (50–65 °C), but PET-S degradation was maximal at 60 °C and decreased at 65 °C. APEXA 4027™ was degraded at temperatures less than 55 °C. Introduced hydrolyzable residues in APEXA 4027™ and some minor components in PET-S (as PET packages are often molded in the presence of additives) must have caused aging of the film at higher temperatures, resulting in the decreased degradation. This result contradicts the report of Sulaiman et al. (2014) that suggested a higher degradation rate of PET-S at 70 °C than at 60 °C. On the other hand, the higher degradation of PET-S at 60 °C than that of PET-GF at 65 °C may be because of the orientation of the polymer chains. It is noteworthy that degradation rate of PET films is highly dependent on the purity of PET and orientation of the polymer chains besides its crystallinity. Therefore, the comparison of the degradation rates has to be based on the standard film such as PET-GF that is commercially available. The degradation of PET-GF (pure and amorphous PET) at 50 °C is low (approximately 5%), and the degradation increased with increased temperatures (55, 60, and 65 °C) to more than 30%.
PET hydrolases in general need to exhibit thermal stability properties at ≥ 65 °C (Wei and Zimmermann 2017). Recently, the crystal structures of a cutinase-like enzyme, referred to as PETase from Ideonella sakaiensis (IsPETase) while working at 30 °C (Yoshida et al. 2016) were solved (PDB IDs: 5G0 and 5XJH) (Han et al. 2017; Joo et al. 2018). However, the enzyme is apparently heat-labile with Tm values of lower than 50 °C, suggesting that the enzyme might not significantly degrade the building blocks of PET, and actually, the degradation rate of PET was less than 1% at 30 °C (only the surface modification level) (Yoshida et al. 2016). Surface modification of the PET film at temperatures lower than Tg has been reported by various hydrolases (Zimmermann and Billig 2011), as the ends of the polymer chains protrude on the surface of the PET film, as imaged in Fig. S5, which leads to the release of hydrolytic products, such as BHET, MHET, and TPA at low levels, such as less than 1%. Whether IsPETase could reach the degradation rate corresponding to those of thermostable PET hydrolases (Wei and Zimmermann 2017) or not awaits further study.
Q138A/D250C-E296C showed a higher degradation rate of PET-GF at 70 °C than at 63 °C, as previously reported (Table 3). Q138A/D250C-E296C/E220R-G251D showed the decreased degradation of PET-GF at 70 °C, although its kcat/Km on PBSA at 37 °C was almost identical to that of Q138A/D250C-E296C. This result is most probably due to the substrate recognition or Tm values of the two mutants. Introduction of Q123H/N202H improved the degradation rates of PET-GF by Q138A/D250C-E296C and Q138A/D250C-E296C/E220R-G251D (Table 3). SEM observation of enzyme-treated PET-GF films supported these results (Fig. 5). The degradation rates of the Cut190* mutants are ranked next to those of HiC from Thermomyces insolens and TfCut2 mutants (Wei and Zimmermann 2017). No product inhibition by Cut 190* is beneficial to its application and would be due to Ala108, as glycine at this position is related to product inhibition, but the replacement of glycine with alanine releases the product inhibition by MHET (Wei et al. 2016).
In conclusion, a PET hydrolase, Cut190*, is an exception among homologous cutinases as Cut190* can be converted from a Ca2+-free inactive state to a Ca2+-bound active state, in which three Ca2+ ions can bind at three binding sites 1–3, whereas other cutinases maintain constant active states, although they showed increased activity with Ca2+. We analyzed the functional roles of the three binding sites by mutation: Site 1 is related to activation, site 2 is related to thermostabilization, and site 3 is supplementally related to activation and thermostabilization. A mutant, Q138A/D250C-E296C/Q123H/N202H, showed the highest Tm value and highest degradation of PET-GF by more than 30% at 70 °C with apparent surface erosion of the film.
References
Dougherty DA (1996) Cation-π interactions in chemistry and biology. A new view of benzene, Phe, Tyr, and Trip. Science 271:163–168
Dougherty DA (2007) Cation-π interactions involving aromatic amino acids. J Nutr 137:1504S–1508S
Duerschmidt P, Mansfeld J, Ulbrich-Hofmann R (2005) An engineered disulfide bridge mimics the effect of calcium to protect neutral protease against local unfolding. FEBS J 272:1523–1534
Han X, Liu W, Huang J-W, Ma J, Zheng Y, Ko T-P, Xu L, Cheng Y-S, Chen C-C, Guo R-T (2017) Structural insight into catalytic mechanism of PET hydrolase. Nat Commun 8:2106
Herrero Acero E, Ribitsch D, Steinkelner GT, Gruber K, Greimel K, Eiteljoerg I, Trotscha E, Wei R, Zimmermann W, Zinn M, Cavaco-Paulo A, Freddi G, Schwab H, Guebitz GM (2011) Enzymatic surface hydrolysis of PET: effect of structural diversity on kinetic properties of cutinases from Thermobifida. Macromolecules 44:4632–4640
Inaba S, Fukada H, Ikegami T, Oda M (2013) Thermodynamic effects of multiple protein conformations on stability and DNA binding. Arch Biochem Biophys 537:225–232
Joo S, Cho IJ, Seo H, Son HF, Sagong HY, Shin TJ, Choi SY, Lee SY, Kim KJ (2018) Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation. Nat Commun 9:382
Kaneko H, Minagawa H, Shimada J (2005) Rational design of thermostable lactate oxidase by analyzing quaternary structure and prevention of deamidation. J Biotechnol Lett 27:1777–1784
Kawabata T, Oda M, Kawai F (2017) Mutational analysis of cutinase-like enzyme, Cut190, based on the 3D docking structure with model compounds of polyethylene terephthalate. J Biosci Bioeng 124:28–35
Kawai F, Thumarat U, Kitadokoro K, Waku T, Tada T, Tanaka N, Kawabata T (2013) Comparison of polyester-degrading cutinases from genus Thermobifida. In: Chen et al (eds) Green polymer chemistry: biocatalysis and materials II. American Chemical Society, Washington, DC, pp 111–120
Kawai F, Oda M, TamashiroT WT, Tanaka N, Yamamoto M, Mizushima H, Miyakawa T, Tanokura M (2014) A novel Ca2+-activated, thermostabilized polyesterase capable of hydrolyzing polyethylene terephthalate from Saccharomonospora viridis AHK 190. Appl Microbiol Biotechnol 98:10053–10064
Kawai F, Kawase T, ShionoT UH, Sukigara S, Tu C, Yamamoto M (2017) Enzymatic hydrophilization of polyester fabrics using a recombinant cutinase Cut190 and their surface characterization. J Fiber Sci Technol 73:8–18
Kitadokoro K, Thumarat U, Nakamura R, Nishimura K, Karatani H, Suzuki H, Kawai F (2012) Crystal structure of cutinase Est119 from Thermobifida alba AHK119 that can degrade modified polyethylene terephthalate at 1.76 Å resolution. Polym Degrad Stab 97:771–775
Miyakawa T, Mizushima H, Ohtsuka J, Oda M, Kawai F, Tanokura M (2015) Structural basis for the Ca2+-enhanced thermostability and activity of PET-degrading cutinase from Saccharomonospora viridis AHK190. Appl Microbiol Biotechnol 99:2486–2498
Numoto N, Kamiya N, Bekker G-J, Yamagami Y, Inaba S, Ishii K, Uchiyama S, Kawai F, Ito N, Oda M (2018) Structural dynamics of the PET-degrading cutinase-like enzyme from Saccharomonospora viridis AHK190 in substrate-bound states elucidates the Ca2+-driven catalytic cycle. Biochemistry 57: 5289–5300
Ribitsch D, Hromic A, Zitzenbacher S, Zartl B, Gamerith C, Pellis A, Jungbauer A, Lyskowski A, Steinkellner G, Gruber K, Tscheliessnig R, Acero EH, Guebitz GM (2017) Small cause, large effect: structural characterization of cutinases from Thermobifida cellulosilytica. Biotechnol Bioeng 114:2481–2488
Roth C, Wei R, Oeser T, Then J, Føllner C, Zimmermann W, Straeter N (2015) Structural and functional studies on a thermostable polyethylene terephthalate degrading hydrolase from Thermobifida fusca. Appl Microbiol Biotechnol 98:7815–7823
Silva C, Da S, Silva N, Matama T, Araujo R, Martins M, Chen S, Chen J, Wu J, Casal M, Cavaco-Paulo A (2011) Engineered Thermobifida fusca cutinase with increased activity on polyester substrates. Biotechnol J 6:1230–1239
Smith CA, Toogood HS, Baker HM, Daniel RM, Baker EN (1999) Calcium-mediated thermostability in the subtilisin superfamily: the crystal structure of Bacillus Ak 1 protease at 1.8 Å resolution. J Mol Biol 294:1027–1040
Sulaiman S, You D-J, Kanaya E, Koga Y, Kanaya S (2014) Crystal structure and thermodynamic and kinetic stability of metagenome-derived LC-compost. Biochemistry 53:1858–1869
Teplyakov AV, Kuranova IP, Harutyunyan EH, Vainshtein BK, Frommel C, Hohne WE, Wilson KS (1990) Crystal structure of thermitase at 1.4 Å resolution. J Mol Biol 214:261–279
Then J, Wei R, Oeser T, Barth M, Belisario-Ferrari MR, Schmidt J, Zimmermann W (2015) Ca2+ and Mg2+ binding site engineering increases the degradation of polyethylene terephtharate films by polyester hydrolase from Thermobifida fusca. Biotechnol J 10:592–598
Then J, Wei R, Oeser T, Gerdts A, Schmidt J, Barth M, Zimmermann W (2016) A disulfide bridge in the calcium binding site of a polyester hydrolase its thermal stability and activity against polyethylene terephthalate. FEBS Open Bio 6:425–432
Thumarat U, Nakamura R, Kawabata T, Suzuki H, Kawai F (2012) Biochemical and genetic analysis of a cutinase-type polyesterase from a thermophilic Thermobifida alba AHK 119. Appl Microbiol Biotechnol 95:419–430
Vieille C, Zeikus GJ (2001) Hyperthermophilic enzymes: sources, uses, and molecular mechanisms for thermostability. Microbiol Mol Biol Rev 65:1–43
Wei R, Zimmermann W (2017) Biocatalysis as a green route for recycling the recalcitrant plastic polyethylene terephthalate. Microb Biotechnol 10:1302–1307
Wei R, Oeser T, Schmidt J, Meier R, Barth M, Then J, Zimmermann W (2016) Engineered bacterial polyester hydrolases efficiently degrade polyethylene terephthalate due to relieved product inhibition. Biotechnol Bioeng 113:1658–1665
Wright HT (1991) Nonenzymatic deamidation of asparaginyl and glutamyl residues in proteins. Crit Rev Biochem Mol Biol 26:1–52
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Oda K (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351:1196–1199
Zimmermann W, Billig S (2011) Enzymes for the biofunctionalization of poly(ethylene terephthalate). Adv Biochem Eng Biotechnol 125:97–120
Acknowledgements
The authors thank Dr. Nobutaka Numoto of Tokyo Medical and Dental University for the helpful discussion APEXA 4027™ film was kindly provided by Dr. M. Kaku, Dupont Co., Ltd. Japan.
Funding
This study was funded by the Institute for Fermentation, Osaka (Japan) (2017). This research was partially supported by Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from Japan Agency for Medical Research and Development under Grant Number JP17am0101066. This research is partially linked with the Asia Core Program supported by JSPS (Japan) and NRCT (Thailand).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interests.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 615 kb)
Rights and permissions
About this article
Cite this article
Oda, M., Yamagami, Y., Inaba, S. et al. Enzymatic hydrolysis of PET: functional roles of three Ca2+ ions bound to a cutinase-like enzyme, Cut190*, and its engineering for improved activity. Appl Microbiol Biotechnol 102, 10067–10077 (2018). https://doi.org/10.1007/s00253-018-9374-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9374-x