Abstract
Bacillus thuringiensis bacteria show insecticidal activities that rely upon the production of insecticidal crystal proteins, which are encoded by cry or cyt genes and can target a variety of insect pests. It has been shown that cry1Ac is the only cry gene in B. thuringiensis subsp. kurstaki HD73 (B. thuringiensis HD73) and its expression is controlled by both σE and σK. Here, we report a novel σE-dependent strong promoter of a non-cry gene (HD73_5014), which can direct strong cry1Ac gene expression in B. thuringiensis HD73. We constructed an E. coli-B. thuringiensis shuttle vector (pHT315-P 5014 -1Ac) for cry1Ac gene expression, using the HD73_5014 gene promoter. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis and western blot analysis showed that expression of the cry1Ac gene directed by the HD73_5014 gene promoter was at the same level as that directed by the previously known strongest cry promoter, P cry8E . However, this strain did not form typical bipyramidal crystals in mother cells, as observed by transmission electron microscopy and atomic force microscope. The strain with Cry1Ac protein expression under the control of the HD73_5014 gene promoter (P 5014 -cry1Ac) showed insecticidal activity against Plutella xylostella similar to that under the control of the orf1cry8E gene promoter (P cry8E -cry1Ac). Collectively, these results suggest that the HD73_5014 gene promoter, as a non-cry gene promoter, would be an efficient transcriptional element for cry gene expression. These data also show the possibility for improving Cry production by searching for transcriptional elements in not only cry genes, but also non-cry genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacillus thuringiensis is a gram-positive, spore-forming bacterium that belongs to the Bacillus cereus (Bc) group (Skerlova et al. 2014). It is characterized by forming parasporal crystal proteins and spores during the stationary phase of its growth cycle (Bravo et al. 2011). These crystal proteins, encoded by cry or cyt genes, possess highly specialized insecticidal activities against numerous insect species, including Lepidoptera, Coleoptera, and Diptera (Schnepf et al. 1998). The insecticidal specificity of B. thuringiensis strains toward different insects is determined by these cry and cyt genes. Due to its insecticidal and environmentally friendly properties, B. thuringiensis has been used commercially as a biocontrol agent worldwide, accounting for approximately 50% of the ever-growing biopesticide market (Gerwick and Sparks 2014; Lacey et al. 2015).
To date, approximately 825 cry and cyt genes have been discovered (http://www.lifesci.sussex.ac.uk/home/Neil_Crickmore/Bt/).The cry genes are expressed during the stationary phase, and their products generally accumulate in the mother cell compartment to form a crystal inclusion that can account for 20 to 30% of the dry weight of the sporulating cells (Schnepf et al. 1998). The cry genes expressed during the stationary phase are classified as sporulation-dependent and sporulation-independent, based on how their expression is regulated (Agaisse and Lereclus 1995; Schnepf et al. 1998; Kroos et al. 1999). A typical example of a sporulation-independent cry gene is the cry3Aa gene, which is weakly but constitutively expressed during vegetative growth under the control of the sigma factor, σA (Desouza et al. 1993; Agaisse and Lereclus 1994). However, most cry genes (e.g., cry1Aa, cry1Ba, cry1Ac, cry2Aa, cry4Aa, cry4Ba, cry11Aa, and cry15Aa) are sporulation-dependent, which are active during the sporulation stages (Brizzard et al. 1991; Brown 1993; Yoshisue et al. 1993; Dervyn et al. 1995; Bravo et al. 1996; Zhang et al. 1998; Kroos et al. 1999). These sporulation-dependent cry genes are expressed in the mother cell and the crystals accumulate there. This progress is dependent on highly ordered cell programs and is regulated by a series of sigma factors (e.g., σE and σk) in signaling cascades (Agaisse and Lereclus 1995; Schnepf et al. 1998; Kroos et al. 1999). For example, transcription of the double promoter-containing gene cry1Aa is controlled by both σE and σK, which bind to the BtI and BtII promoters during the early-to-middle and middle-to-late sporulation phases, respectively (Wong et al. 1983; Brown and Whiteley 1988, 1990; Bravo et al. 1996; Saxild et al. 1996). Another sporulation-dependent cry gene, cry8Ea1 (also named cry8E), is transcribed from two promoters, namely P orf1 (located upstream of the orf1 gene) and P cry8E (located in the intergenic region between orf1 and cry8Ea1), which are regulated by σE and σH, respectively (Du et al. 2012).
Despite the fact that B. thuringiensis has been used as an insecticidal agent worldwide, it has only captured approximately 2% of the total insecticides market owing to the relatively low yield and instability of the crystal protein in the field (Bradley et al. 1995; Huang et al. 2007; Bravo et al. 2011). Some methods have been developed to increase the yield, titer, and stability of insecticidal crystal proteins. For example, it has been reported that Cry1Ac production was enhanced when the gene was controlled by the heterologous cry8E gene promoter P cry8E , compared with that in wild-type strain (HD73) (Li et al. 2013). Interestingly, it has been reported that a non-cry gene promoter can also successfully drive cry1Ac expression. ExsY is a basal layer protein in the exosporium of B. anthracis spores (Boydston et al. 2006). The promoter of the exsY gene (P exsY ) can direct cry1Ac gene expression during the late sporulation stage although it leads to a lower Cry protein yield (Zheng et al. 2014). Those studies provide us alternative strategies of improving Cry production and efficacy of Bt strains as insecticidal agents. Nevertheless, the pick of non-cry gene promoters and the efficiency of using those promoters to direct cry gene expression and Cry protein production remain to be further investigated.
In this study, we identified a novel strong promoter from a gene encoding a putative DeoR family transcriptional regulator (P 5014 , as it is encoded by HD73_5014). We showed that this non-cry promoter can efficiently drive cry1Ac gene expression in B. thuringiensis to a level comparable to that from the strong cry gene promoter P cry8E . A crystal protein-producing strain, HD−P 5014 -1Ac, in which cry1Ac is expressed under the HD73_5014 gene promoter, was constructed. We found that HD−P 5014 -1Ac was similar to the wild-type strain (HD73), both in terms of Cry1Ac production and insecticidal activity against P. xylostella larvae. More importantly, when introduced into a sigK mutant, we obtained a higher yield of encapsulated crystals since the non-cry gene promoter P 5014 is sigK-independent. Encapsulated crystals showed better resistance to environmental factors such as UV light inactivation and better efficacy when applied in the field in previous studies (He et al. 2017). Our work suggested that the non-cry gene promoter P 5014 showed good prospects in constructing engineered B. thuringiensis strains for application in agriculture.
Materials and methods
Bacterial strains, plasmids, and growth conditions
B. thuringiensis subsp. kurstaki HD73 (BGSC strain number BGSC 4D4, hereafter designated as HD73), the acrystalliferous mutant strain HD73− (Lereclus et al. 1989) and their derivatives were grown at 30 °C in Luria–Bertani (LB) medium (1% tryptone, 0.5% yeast extract, and 0.5% NaCl) or on solid LB medium supplemented with 1.5% agar. Schaeffer’s sporulation medium (Schaeffer et al. 1965) (SSM; 0.8% nutrient broth, 1 mM MgSO4, 13.4 mM KCl, 0.5 mM NaOH, 1 mM Ca(NO3)2, 0.01 μM MnCl2, and 1 μM FeSO4) was used to measure promoter activities. Escherichia coli JM109 was used for molecular cloning experiments, and E. coli ET1 was used for producing non-methylated plasmid DNA for B. thuringiensis transformations (Macaluso and Mettus 1991; Wang et al. 2006); both strains were grown at 37 °C in LB medium. When required, antibiotics were added at the following concentrations for growth of B. thuringiensis: 5 μg/ml erythromycin and 50 μg/ml kanamycin. For growth of E. coli, 100 μg/ml ampicillin was added when needed. The HD73 complete genomic sequence had been submitted to NCBI database (GenBank Accession No. NC_020238.1), all gene sequences could be found in this database. The bacterial strains and plasmids used in this study are summarized in Table 1.
DNA manipulation and transformation
Plasmid DNA was extracted from E. coli cells with a Plasmid Miniprep Kit (Axygen, Beijing, China). Restriction enzymes and T4 DNA ligase (Takara Biotechnology Corporation, Dalian, China) were used according to the manufacturer’s instructions. PCR was performed with the high-fidelity PrimeSTAR HS DNA polymerase (Takara Biotechnology Corporation, Beijing, China) or Taq DNA polymerase (BioMed, Beijing, China). DNA fragments were purified from 1% agarose gels, using a AxyPrep DNA Gel Extraction Kit (Axygen, Beijing, China). Standard procedures were followed for E. coli transformation (Sambrook and Russell 2015). B. thuringiensis cells were transformed by electroporation as previously described (Lereclus et al. 1989).
RNA-Seq analysis
Total RNA was extracted from HD73 cells grown in SSM medium until T 7 stage (7 h after the end of the exponential phase) following the previously published method (Du et al. 2012). Briefly, 1 ml of Bt cells were harvested by centrifugation (14,000×g, 1 min at 4 °C), and the pellets were resuspended in 1 ml of cold TRI-Reagent (Invitrogen, San Diego, CA). The RNA was extracted with the Qiagen Easy RNA kit according to the manufacturer’s instructions. The residual DNA was removed using RNase-free DNase I (New England BioLabs). The pure total RNA was used for RNA-Seq analysis. Briefly, rRNA (including 16S and 23S) was removed from 4 μg of total RNA by Microbexpress™ (Ambion), and the left RNA was chemically fragmented. The sequence library construction is carried out according to ScriptSeq: trademark: mRNA-Seq Library Preparation Kit (Illumina). In brief, the fragmented RNA is reverse-transcribed into cDNA using the SuperScript double-stranded cDNA synthesis kit (Invitrogen) with the addition of SuperScript III reverse transcriptase (Invitrogen) and random primers containing a tagging sequence at their 3′ ends. This was followed by RNase A (Roche, Germany) treatment, phenol-chloroform extraction, and ethanol precipitation. The resulting cDNAs were ligated to a 5′ DNA/DNA adaptor and the di-tagged cDNA was purified by PAGE gel. The size of the inserted fragment is ~ 150–250 bp. The purified products were PCR amplified to in 18 cycles using a high-fidelity DNA polymerase. PCR products were purified using the PAGE gel. cDNAs were sequenced using a single flow cell of the Illumina Hiseq 2000. FPKM = cDNA fragments/(mapped reads (millions)×transcript length (kb)).
Construction of the gene promoter with the lacZ reporter gene
To compare the transcriptional activity of the target genes, the promoter sequences of the chosen genes were amplified from HD73 genomic DNA (GenBank Accession No. NC_020238.1) using different primers (Table 2). The length of the amplified fragment (P 5014 ) is 710 base pairs upstream of the HD73_5014 translational start codon. The amplified fragments were digested with PstI and BamHI, followed by ligation into the linearized pHT304-18Z plasmid, which harbors a promoterless lacZ gene (Agaisse and Lereclus 1994). The recombinant plasmids were introduced into HD73 cells by electroporation using a Gene Pulser II apparatus (Bio-Rad, USA). The resulting strains were selected on the agar plates supplemented with erythromycin (validation is done by sequencing).
Construction of the P 5014 -Cry1Ac fusion
Overlapping PCR was performed to generate the P 5014 -cry1Ac fusion. The primer pairs, pHT5014-5, pHT5014-3 and pHT1Ac-5, pHT1Ac-3, were used to amplify P 5014 and the cry1Ac ORF (GenBank Access Number: AAB46989.1) (Wong et al. 1983), respectively. Long-fragment PCR was conducted with the pHT5014-5/pHT1Ac-3 primer set, using the PCR products as templates. The generated P 5014 -cry1Ac fusion product was then digested with SalI and SphI and integrated into the pHT315 vector, which was also digested with SalI and SphI, resulting in pHT315-P 5014 -1Ac. The plasmid was verified by Sanger sequencing, followed by introduction into HD73− (acrystalliferous mutant strain by curing of the plasmid carrying the cry1Ac gene) (Lereclus et al. 1989) and HDsigK− strain (acrystalliferous sigK mutant strain) (Zhou et al. 2014). Transformants were selected on erythromycin agar plates at 30 °C. In addition, the pHT315-8E-1Ac plasmid, HD8E-1Ac, and ∆sigK strains were used as controls (Table 1) (Zhou et al. 2014).
Assays of β-galactosidase activities
To assay β-galactosidase activities, the B. thuringiensis strains were grown in SSM medium at 30 °C with shaking (220 rpm). Two milliliters of culture were collected at 1-h intervals from T 1 to T 12 (T 0 indicates the end of the exponential growth phase and T n indicates n hours after T 0 ). Cells were centrifuged (14,000×g, 1 min) and pellets were stored at – 20 °C ready for use. β-galactosidase activities were measured as previously described (Yang et al. 2012) and expressed as Miller units (Miller 1972). Values are reported as the mean and standard error of at least three independent assays.
SDS-PAGE analysis of Cry protein production
B. thuringiensis cells were grown to T 24 in 50 ml of fresh SSM at 30 °C with shaking (220 rpm), and the cells were centrifuged at 4 °C for 10 min at 8000 rpm, followed by freeze-drying for ~ 48 h until the pellets became lyophilized powders. An appropriate volume of double-distilled water was added to each sample to adjust them to equivalent bacterial biomass concentrations (mg/ml). The pellets were thoroughly vortexed for 20 s and resuspended. Then, the cells were disrupted with a Mini-BeadBeater (Biospec Products, Inc., Bartlesville, OK, USA) in a 2-ml centrifuge tube containing approximately 200 μg of glass beads (0.1 mm diameter). After a low-speed (1000×g) centrifugation, the total cell lysates were transferred to a new centrifuge tube. The mixture of 40 μl of cell lysates and 10 μl of 5× loading buffer was boiled for 10 min, followed by centrifugation at 12000×g for 1 min at 4 °C, then the cell lysates were analyzed to detect Cry protein production by SDS-PAGE and Coomassie Brilliant Blue staining. Protein band intensities were determined using ImageJ software (Version 1.6.0_24, National Institutes of Health) by comparison to bovine serum albumin (BSA).
Western blot analysis
Western blot experiments were performed after the samples were separated by SDS-PAGE (4% polyacrylamide stacking gel, 10% polyacrylamide separating gel) as previously described (Wang et al. 2006). The proteins were transferred to a polyvinylidene difluoride (PVDF) membrane. The protein-laden PVDF membranes were probed with a primary antibody against Cry1Ac (Beijing Protein Innovation Inc., Beijing, China) at a dilution of 1:5000. Antibody binding was detected with HRP-conjugated goat anti-mouse IgG (CWBiotech, Beijing, China). Visualization was performed as previously described (Ni et al. 2016).
Determination of the transcriptional start site
Total RNA was extracted from HD73 cells grown in SSM until T 7 stage, and reverse transcription PCR was conducted as previously described (Du et al. 2012). We used the SMARTer RACE (switching mechanism at the 5′ end of the RNA transcript-rapid amplification of cDNA ends) cDNA amplification kit (Clontech, Mountain View, CA) to determine the transcription start site, following the manufacturer’s instructions. P5014Race, located ~ 200 bp downstream of the HD73_5014 start codon (TTG), was designed as the specific reverse primer. NestRace was the forward primer provided in the kit (Clontech, Mountain View, CA). P5014Race and NestRace were used as specific primers for amplifying the 5′ end of P 5014 cDNA. The sequences of the primers used in this study are shown in Table 2.
Observation of cell shape and crystal structure
Transmission electron microscopy (TEM) and atomic force microscopy (AFM) were employed to analyze the B. thuringiensis cell shape and structure of the insecticidal crystals. Samples were prepared for TEM as follows. Briefly, the cells were grown until T 24 in SSM at 30 °C. Twenty-milliliter samples were harvested by centrifugation, followed by adding 1 ml of 3% glutaraldehyde solution. After performing a series of procedures (fixation, dehydration, replacement, soaking, embedding, sectioning, and staining), the cell shape was observed as previously described (Liu et al. 2008; Weigel and Glazebrook 2010). For AFM experiments, the strains were grown until T 24 or a later stage to release the crystals. The MultiMode 8 SPM instrument was operated according to the guidelines of the manufacturer (Bruker, Germany).
Bioassay of insecticidal activities
Biological assays were performed as described by (Zhou et al. 2014), using equivalent bacterial biomass concentrations for the HD−P 5014 -1Ac, HD8E-1Ac, HD73, and HD73− strains. Briefly, insecticidal activities were tested by exposing second instar larvae (diamondback moth, P. xylostella) to an artificial diet incorporating 1 of 7 dilutions (bacterial lyophilized powder concentrations of 1.25, 2.5, 5, 10, 20, 40, and 80 μg/ml) of each preparation in water (Xue et al. 2008). A 6-cm-diameter cabbage leaf disc was pretreated with a gradient bacterial concentration as described above and then transferred to a new plastic culture dish, after which 30 s instar larvae were placed in each dish. The numbers of surviving larvae were counted after 3 days, and the LC50 was calculated by probit analysis. The test for each concentration was performed in triplicate.
Results
The HD73_5014 gene promoter showed high activities during sporulation
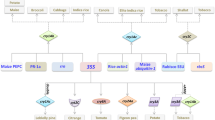
cry1Ac, the only cry gene in HD73, is sporulation-dependent since it is regulated by the sporulation sigma factor K (SigK) (Schnepf et al. 1998). The cry1Ac gene was expressed after entry into sporulation phase and the transcriptional activity was maintained at a high level from T 6 (Zhou et al. 2014). We aimed to decouple high expression of cry1Ac at the late sporulation stage from its dependence on SigK. We assumed that some promoters of the genes with high transcriptional activity at the late sporulation phase might be applied to direct cry1Ac gene expression efficiently in a SigK-independent fashion. To obtain a potential strong promoter with this feature, we analyzed gene-expression levels from microarray data (NCBI Gene Expression Omnibus Accession No. GSE 48410) at the late sporulation phase (here, we chose T 7 ) of the B. thuringiensis HD73 strain (Peng et al. 2015) and selected 100 candidate genes with top-level transcriptional activity (Table S1). Then, RNA-Seq data (SRA accession: SRP127175; Table 3) were utilized to check the relative transcriptional levels of those candidate genes at the late sporulation phase (T 7 ). We obtained seven candidate genes with high transcriptional activity both in the microarray data (Table S1) and RNA-Seq data (value > 20,000). The top seven genes with relatively high transcriptional activity, from high to low, are HD73_0588, HD73_6004 (cry1Ac), HD73_4734, HD73_1453, HD73_4944, HD73_0992, and HD73_5014 (Table 3). To confirm the high transcriptional activity of these candidate genes (except for cry1Ac), we fused each of the corresponding promoter regions to the lacZ gene to create the lacZ reporter fusion and constructed individual strains bearing each of those reporter fusions (see Materials and Methods, Fig. 1a, b). The assays of the β-galactosidase activity demonstrated that the transcriptional activity of P 5014 was the highest (Fig. 1c). In addition, we also found that HD73_5014 was transcribed at a relatively high level even at T −1 (relative expression level is 1881.97, data not shown). Finally, the HD73_5014 gene, a putative DeoR family transcriptional regulator, was selected since it showed high expression levels at the sporulation stage (Fig. 1c).
The promoter of the HD73_5014 gene showed high transcriptional activity in the sporulation stages. a The arrangement of six putative highly transcriptional genes and their promoter regions in HD73 genome. The white open arrows indicate the candidate genes and black open arrows show adjacent genes. b Construction of a transcriptional reporter by fusing the HD73_5014 gene promoter with lacZ. c Comparison of the transcriptional activities of the 6 putative strong promoters. d Comparison of the transcriptional activities between the HD73_5014 and cry8E gene promoters. The β-galactosidase activities of three clones were determined at the indicated times after growing the cells in SSM at 30 °C. Each value represents the mean and standard error of at least three independent replicates
It has been reported that the cry8E gene promoter (P cry8E ) is a strong cry promoter that can direct high cry1Ac gene expression during the sporulation stages (Zhou et al. 2014). To compare the high transcriptional activity of HD73_5014 gene promoter (P 5014 ) and the transcriptional levels with P cry8E , we performed β-galactosidase assays using two strains harboring these lacZ fusions (see Materials and Methods, Fig. 1b). β-Galactosidase assays indicated that the transcriptional activities of both P 5014 and P cry8E were high during the sporulation stages, especially from T 6 to T 12 (Fig. 1d). The transcriptional activities of both promoters were similar, and both increased until they peaked at T 8 and then decreased (Fig. 1d). In general, the transcriptional activity of P 5014 was comparable to that of P cry8E , demonstrating that P 5014 is also a strong promoter during the sporulation stages.
The HD73_5014 gene promoter functions in a σE-dependent manner
We wanted to investigate the transcriptional regulation of H73_5014 in sporulation stages. We started by analyzing the HD73_5014 gene promoter sequence in detail. The transcription start site was confirmed to be a single 5′-end nucleotide residue “A,” located 55 nucleotides upstream of the HD73_5014 translational start codon, based on the sequences of random clones obtained by 5′-RACE (Fig. 2a).
The activity of the HD73_5014 gene promoter relies on σE. a Analysis of the promoter of the HD73_5014 gene in B. thuringiensis HD73. The − 35 and − 10 motifs and the transcription start site (+ 1) are annotated. The underlined TTG is the start codon of HD73_5014 gene. b Analyses of HD73_5014 gene promoter activities in the wild-type HD73, the sigK mutant, and the sigE mutant strains. Promoter-directed β-galactosidase expression in three clones was determined at the indicated times after growing the cells in SSM at 30 °C. The values shown represent the mean and standard error of at least three independent replicates
Our analysis of the HD73_5014 promoter region revealed that − 35 (GCATGA) and − 10 (AGTACAAT) regions (Fig. 2a) were similar to the consensus sequences of σE-dependent promoters (KHATANHT…MATANNHT) (Zhang et al. 1998). These findings indicated that the HD73_5014 promoter might function in a σE-depended manner. The cry1Ac gene is a sporulation-dependent gene that has two promoters, which are recognized by σE and σK, respectively (Wong et al. 1983). We aimed to use the HD73_5014 gene promoter to drive cry1Ac gene expression; thus, we were interested in determining whether the HD73_5014 promoter is controlled by σE and σK. To that end, we introduced the previously constructed pHT304-P 5014 plasmid containing the P 5014 -lacZ fusion into the wild-type strain (HD73), the sigE mutant (HDΔsigE), and the sigK mutant (HDΔsigK) and then compared the β-galactosidase activities of these three strains. In the sigE mutant, the P 5014 -lacZ fusion was expressed at a much lower level, confirming that the HD73_5014 promoter is controlled by σE (Fig. 2b). The transcriptional activity of P 5014 increased rapidly during T 1 to T 8 in both the wild-type and sigK mutant strains (Fig. 2b). However, the transcriptional activity of P 5014 in the wild-type strain was lower than that in the sigK mutant from T 9 to T 12 (Fig. 2b), consistent with the transcriptional activity of P cry8E observed previously (Du et al. 2012). Our results confirmed that the HD73_5014 gene promoter functions specifically in a σE-, but not σK-, dependent manner during sporulation. Thus, using the HD73_5014 gene promoter to direct expression of cry1Ac would allow us to achieve high expression and presumably high yield of crystal proteins independent on SigK.
The HD73_5014 gene promoter directed cry1Ac expression and crystal accumulation
To test whether HD73_5014 promoter could effectively direct Cry1Ac expression in B. thuringiensis, we first constructed the pHT315-P 5014 -1Ac plasmid (Fig. 3a), which contains the P 5014 -cry1Ac fusion. The reaction product Cry1Ac on pHT315-P5014-1Ac is fused protein of N-terminal seven amino acids (1–7) for HD73_5014 and Cry1Ac (Fig. 3a). In parallel, the pHT315-8E-1Ac (Zhou et al. 2014) plasmid was used as a control. Subsequently, these two plasmids were introduced into the acrystalliferous HD73− strain, which could not produce the Cry protein (Lereclus et al. 1989), resulting in the HD−P 5014 -1Ac and HD8E-1Ac strains (Zhou et al. 2014), respectively. The pHT315-P 5014 -1Ac plasmid was also introduced into HDsigK− strain (Zhou et al. 2014), resulting in the ∆sigK−P 5014 -1Ac strain. The formation of crystal proteins was examined by TEM and high-resolution AFM (Fig. 4). In contrast to the HD73− strain (Fig. 4d), HD−P 5014 -1Ac (Fig. 4a), HD8E-1Ac (Fig. 4b), and HD73 (Fig. 4c) all could produce clear crystalline inclusions. Interestingly, HD−P 5014 -1Ac (Fig. 4a) and HD8E-1Ac (Fig. 4b) could not form typical bipyramidal crystals during the sporulation stage, while the typical bipyramidal crystalline inclusion was clear in HD73 (Fig. 4c). For further confirmation, we examined the crystal formation by AFM. Compared with the perfect bipyramidal crystals produced by HD73 (Fig. 4g), we saw imperfect bipyramidal crystals in both the HD−P 5014 -1Ac (Fig. 4e) and HD8E-1Ac strains (Fig. 4f), some of which had a rounded, blunt shape. These results suggested that the regulation of Cry1Ac expression significantly affected bipyramidal crystal formation.
The HD73_5014 promoter drives Cry1Ac expression with high efficiency in B. thuringiensis. a Physical map and expression region sequence of pHT315-P 5014 -1Ac. b SDS-PAGE analysis of Cry1Ac expression in the HD−P 5014 -1Ac, HD8E-1Ac, HD73, and HD73− strains. The numbers below indicate the Cry1Ac yield in the four strains after normalization to 0.1 μg/μl BSA. c Comparison of Cry1Ac levels detected by western blotting using a specific antibody against Cry1Ac. Shown is a comparison of Cry1Ac levels from HD73− with those of HD−P 5014 -1Ac, HD8E-1Ac, and HD73 in liquid shaking SSM during stationary phase. The triangle points to the Cry1Ac-specific band. d SDS-PAGE analysis of Cry1Ac expression in strains sigK−, HD∆sigK, and sigK−P 5014 -1Ac. The relative protein yield of Cry1Ac was calculated with Image J software
Cry1Ac expression under different promoters in B. thuringiensis strains. a–d TEM images of different B. thuringiensis cells grown for T 24 in SSM medium at 30 °C. a HD−P 5014 -1Ac strain. b HD8E-1Ac strain. c HD73, a wild-type strain. d HD73−, a negative control strain. e–g AFM pictures of Cry1Ac crystals in different B. thuringiensis strains. e HD−P 5014 -1Ac strain. f HD8E-1Ac strain. g HD73, a wild-type strain
The Cry protein was detected by performing SDS-PAGE (Fig. 3b). The results showed that HD−P 5014 -1Ac, HD8E-1Ac, and HD73 all produced the ~ 130-kDa protein Cry1Ac with approximately the same yield, and no ~ 130-kDa proteins were expressed in the control strain HD73−. In the ∆sigK strain, the production of Cry1Ac was dramatically decreased as that the native promoter of cry1Ac gene was partially σK dependent (Zhou et al. 2014), whereas the sigK−P 5014 -1Ac produced ~ 130-kDa Cry1Ac in a large amount (Fig. 3d), indicating that P 5014 could efficiently direct Cry1Ac expression. This result suggested that P 5014 may be applied to achieve high yield of the crystals in the sigK gene deletion background.
We also performed immunoblot analysis to investigate the specific Cry1Ac proteins produced in different strains using antibodies against Cry1Ac. Protein was extracted from stationary phase cells grown in SSM and tested for Cry1Ac. As expected, ~ 130-kDa Cry1Ac proteins were found produced by the HD−P 5014 -1Ac, HD8E-1Ac, and HD73 strains, but not by HD73− (Fig. 3c). Thus, all results showed that the promoter of the HD73_5014 gene could direct cry1Ac gene expression correctly and efficiently.
The HD73_5014 promoter drove high expression of the insecticidal Cry1Ac protein in B. thuringiensis
HD−P 5014 -1Ac did not produce typical bipyramidal crystals that the wild-type strain (HD73) did, although we saw obvious Cry1Ac protein accumulation in HD−P 5014 -1Ac compared with HD73− (Figs. 3 and 4). Thus, we determined the toxicity of Cry1Ac protein driven by HD73_5014 promoter to P. xylostella in comparison to that of HD8E-1Ac (Zhou et al. 2014). The second instar P. xylostella larvae were fed cabbage leaves pretreated with HD−P 5014 -1Ac, HD8E-1Ac, HD73, or HD73−. The toxicity assay was carried out accordingly (see Materials and Methods). Our results showed that the LC50 value of HD−P 5014 -1Ac against P. xylostella was close to those of HD8E-1Ac and HD73 (Table 4), indicating that crystals from HD−P 5014 -1Ac were capable of killing P. xylostella effectively.
Discussion
A principal contribution of this investigation to the literature is the identification of a new strong non-cry gene promoter, P 5014 , which could effectively drive cry1Ac expression. This could be important since decoupling high expression of the cry genes from their native promoters is particularly useful in engineering Bt-based biological pesticides as we discuss next. The promoter of HD73_5014 among six candidate genes showed the highest transcriptional activity at T 7 stage by using the lacZ reporter fusions (Fig. 1c), which differs from the RNA-Seq and microarray data (Table 3 and S1). One possibility is that RNA-Seq and microarray are the general methods useful for identification of novel transcripts. Differential expression of the genes of interest based on RNA-Seq or microarray has to be further confirmed by other methods such as qRCR or the lacZ reporter fusion (Chen et al. 2014). The promoter of the HD73_5014 gene is a good candidate for directing the Cry1Ac expression since P 5014 maintains a very high transcriptional activity from exponential phase to stationary phase (Fig. 1c, d).
We showed that the Cry1Ac protein directed by P 5014 in the engineered strain is functionally similar to that directed by the strong cry gene promoter P cry8E . This conclusion is supported by the following evidence. Firstly, SDS-PAGE and western blot analysis showed that the Cry1Ac proteins expressed under the HD73_5014 promoter, the cry8E promoter, and the native cry1Ac promoter were the same in molecular weight and similar in quantity (Fig. 3). Also, like the wild-type strain, the P 5014 -driven Cry1Ac-producing strain strongly accumulated crystal proteins in the mother cell (Fig. 4). Secondly, the P 5014 -controlled Cry1Ac protein showed similar insecticidal activity against P. xylostella as did the P cry8E -controlled Cry1Ac and Cry1Ac from the wild-type strain (Table 4).
In B. subtilis, DeoR (deoxyribonucleoside regulator) is homologous to the sorbitol operon regulator family of metabolic regulators, and contains a C-terminal effector-binding domain and an N-terminal DNA-binding domain (Skerlova et al. 2014). DeoR negatively regulates expression of enzymes involved in the catabolism of deoxyribonucleosides and deoxyribose (Zeng et al. 2000). It is located immediately upstream of the dra-nupC-pdp operon, which encodes three enzymes required for deoxyribonucleoside and deoxyribose utilization (Saxild et al. 1996). By sequence analysis, we found that a similar dra-nupC-pdp gene cluster exists in B. thuringiensis HD73 genome (Accession No. NC_020238.1). The gene located immediately upstream of this operon is HD73_2060, sharing 78% identity with deoR of B. subtilis 168, indicating that HD73_2060 is the ortholog of B. subtilis deoR. However, HD73_5014 is quite different from deoR, even though its product was annotated as the DeoR family transcriptional regulator. The HD73_5014 encoded protein is predicted to be 73 amino acids in length and comprises primarily the HTH domain (from amino acid 11 to 49). It shares a very low sequence identity with DeoR (only about 6%) and may not be related with deoxyribonucleoside regulation. HD73_5014 did not attract much attention in previous studies and its function remains unclear. We plan to determine the function of the HD73_5014 gene in our follow-up studies.
Many previous studies have focused on improving the yield and stability of insecticidal crystal proteins even after exposure to multiple environmental stresses (Sanchis et al. 1999; Yang et al. 2013; Zhou et al. 2014). One approach taken is to use strong non-cry gene promoters to direct expression of the cry gene. But only the gene promoters with high transcriptional activity are useful to be selected for directing cry gene expression. P cry8E is a relatively strong promoter of the cry8Ea1 gene and is regulated by both σE and σH (Du et al. 2012). This promoter was capable of increasing Cry1Ac protein yield in B. thuringiensis, especially in the sigK mutant (Li et al. 2013; Zhou et al. 2014). This engineered strain is of clear advantage since the P cry8E -driven Cry1Ac expression is at high levels even in the sigK mutant while crystals are encapsulated because of lack of mother cell lysis at the end of sporulation due to the sigK deletion mutation. Previous work has shown that encapsulation allows better protections from environmental stresses and more sustained insecticidal activities of the crystals when applied in the field (He et al. 2017).
Here, we report that P 5014 is another candidate of non-cry gene promoters with high transcriptional activity and could effectively direct high Cry1Ac expression, similar to the known so far the strongest cry gene promoter P cry8E (Fig. 1d and 3). Our results indicated that the cry gene expression driven by P 5014 was higher than that driven by most cry-type gene promoters, supporting the idea that non-cry-type gene promoter could be an alternative for directing crystal protein expression in the engineered Bt strains. This is particularly useful in the sigK mutant background. The deletion of sigK gene leads to the encapsulation of Cry1Ac inside the mother cell; however, the Cry1Ac production was dramatically decreased due to the dependence of the cry1Ac gene expression on SigK (Fig. 3d). In this study, we showed that the σK-independent promoter P 5014 could direct Cry1Ac expression efficiently in the sigK− strain, providing an alternative solution of producing high quantity of Cry1Ac in the sigK deletion background.
Interestingly, the Cry1Ac protein produced under the control of the P 5014 and P cry8E promoters could not assemble perfectly to form typical bipyramidal crystals (Fig. 4e–g). This is the first report describing the abnormal 3-dimensional crystal shape of Cry1Ac. Exactly how expression from the HD73_5014 gene promoter (or P cry8E ) led to abnormal assembly of the Cry1Ac protein remains unknown. We speculated one reason was that the Cry1Ac protein expressed under P 5014 was fused additional seven amino acids of HD73_5014 at N-terminal portion (Fig. 3a). However, Cry1Ac proteins produced from the three strains were very similar in terms of the quantity and insecticidal activity against P. xylostella (Table 4). Our data appeared to indicate that the abnormal crystal shape of Cry1Ac did not affect its biological activity. However, since we only tested the insecticidal activity against P. xylostella under laboratory conditions, it remains to be determined whether different promoter-driven crystals have the same or different performance in the field.
Taken together, our data provide a good example of Cry1Ac expression using a strong non-cry gene promoter in an engineered B. thuringiensis strain. The HD73_5014 gene promoter might also be useful for expressing other cry genes. In addition, non-cry gene promoters maybe capable of increasing Cry protein yield and insecticidal activity in engineered B. thuringiensis strains. The data generated in this study enable us to propose new strategies for improving other biopesticides in future studies and for applications in agriculture.
References
Agaisse H, Lereclus D (1994) Structural and functional analysis of the promoter region involved in full expression of the cryIIIA toxin gene of Bacillus thuringiensis. Mol Microbiol 13:97–107. https://doi.org/10.1111/j.1365-2958.1994.tb00405.x
Agaisse H, Lereclus D (1995) How does Bacillus thuringiensis produce so much insecticidal crystal protein? J Bacteriol 177:6027–6032. https://doi.org/10.1128/jb.177.21.6027-6032.1995
Arantes O, Lereclus D (1991) Construction of cloning vectors for Bacillus thuringiensis. Gene 108:115–119. https://doi.org/10.1016/0378-1119(91)90495-W
Boydston JA, Yue L, Kearney JF, Turnbough CLJ (2006) The ExsY protein is required for complete formation of the exosporium of Bacillus anthracis. J Bacteriol 188:7440–7448. https://doi.org/10.1128/JB.00639-06
Bradley D, Harkey MA, Kim MK, Biever KD, Bauer LS (1995) The insecticidal CryIB crystal protein of Bacillus thuringiensis spp. thuringiensis has dual specificity to Coleopteran and Lepidopteran larvae. J Invertebr Pathol 65:162–173. https://doi.org/10.1006/jipa.1995.1024
Bravo A, Agaisse H, Salamitou S, Lereclus D (1996) Analysis of cryIAa expression in sigE and sigK mutants of Bacillus thuringiensis. Mol Gen Genet 250:734–741. https://doi.org/10.1007/BF02172985
Bravo A, Likivivatanavong S, Gill SS, Soberón M (2011) Bacillus thuringiensis: a story of a successful bioinsecticide. Insect Biochem Mol Biol 41:423–431. https://doi.org/10.1016/j.ibmb.2011.02.006
Brizzard BL, Schnepf HE, Kronstad JW (1991) Expression of the cryIB crystal protein gene of Bacillus thuringiensis. Mol Gen Genet 231:59–64. https://doi.org/10.1007/bf00293822
Brown KL (1993) Transcriptional regulation of the Bacillus thuringiensis subsp. thompsoni crystal protein gene operon. J Bacteriol 175:7951–7957. https://doi.org/10.1128/jb.175.24.7951-7957.1993
Brown KL, Whiteley HR (1988) Isolation of a Bacillus thuringiensis RNA polymerase capable of transcribing crystal protein genes. Proc Natl Acad Sci U S A 85:4166–4170. https://doi.org/10.1073/pnas.85.12.4166
Brown KL, Whiteley HR (1990) Isolation of the second Bacillus thuringiensis RNA polymerase that transcribes from a crystal protein gene promoter. J Bacteriol 172:6682–6688. https://doi.org/10.1128/jb.172.12.6682-6688.1990
Chen J, Hou K, Qin P, Liu H, Yi B, Yang W, Wu W (2014) RNA-Seq for gene identification and transcript profiling of three Stevia rebaudiana genotypes. BMC Genomics 15:571. https://doi.org/10.1186/1471-2164-15-571
Dervyn E, Poncet S, Klier A, Rapoport G (1995) Transcriptional regulation of the cryIVD gene operon from Bacillus thuringiensis subsp. israelensis. J Bacteriol 177:2283–2291. https://doi.org/10.1128/jb.177.9.2283-2291.1995
Desouza MT, Lecadet MM, Lereclus D (1993) Full expression of the cryIIIA toxin gene of Bacillus thuringiensis requires a distant upstream DNA sequence affecting transcription. J Bacteriol 175:2952–2960. https://doi.org/10.1128/jb.175.10.2952-2960.1993
Du C, Nickerson KW (1996) Bacillus thuringiensis HD-73 spores have surface-localized Cry1Ac toxin: physiological and pathogenic consequences. Appl Environ Microbiol 62:3722–3726
Du L, Wei J, Han L, Chen Z, Zhang J, Song F, Huang D (2011) Characterization of Bacillus thuringiensis sigK disruption mutant and its influence on activation of cry3A promoter. Wei Sheng Wu Xue Bao 51:1177–1184
Du L, Qiu L, Peng Q, Lereclus D, Zhang J, Song F, Huang D (2012) Identification of the promoter in the intergenic region between orf1 and cry8Ea1 controlled by sigma H factor. Appl Environ Microbiol 78:4164–4168. https://doi.org/10.1128/AEM.00622-12
Gerwick BC, Sparks TC (2014) Natural products for pest control: an analysis of their role, value and future. Pest Manag Sci 70:1169–1185. https://doi.org/10.1002/ps.3744
He X, Sun Z, He K, Guo S (2017) Biopolymer microencapsulations of Bacillus thuringiensis crystal preparations for increased stability and resistance to environmental stress. App Microbiol Biotechnol 101:2779–2789. https://doi.org/10.1007/s00253-016-8070-y
Huang D, Zhang J, Song F, Lang Z (2007) Microbial control and biotechnology research on Bacillus thuringiensis in China. J Invertebr Pathol 95:175–180. https://doi.org/10.1016/j.jip.2007.02.016
Kroos L, Zhang B, Ichikawa H, Yu YT (1999) Control of sigma factor activity during Bacillus subtilis sporulation. Mol Microbiol 31:1285–1294. https://doi.org/10.1046/j.1365-2958.1999.01214.x
Lacey LA, Grzywacz D, Shapiro-Ilan DI, Frutos R, Brownbridge M, Goettel MS (2015) Insect pathogens as biological control agents: back to the future. J Invertebr Pathol 132:1–41. https://doi.org/10.1016/j.jip.2015.07.009
Lereclus D, Arantes O, Chaufaux J, Lecadet M (1989) Transformation and expression of a cloned delta-endotoxin gene in Bacillus thuringiensis. FEMS Microbiol Lett 60:211–217. https://doi.org/10.1111/j.1574-6968.1989.tb03448.x
Li CR, Du LX, Peng Q (2013) Construction of high-level expression vector for Bacillus thuringiensis. Microbiol China 40:350–361
Liu R, Yu G, Zou W, Du T (2008) Improvements in technique of madding ultrathin section for transmission electron microscope. Jiangxi For Sci Technol 41–43
Macaluso A, Mettus AM (1991) Efficient transformation of Bacillus thuringiensis requires nonmethylated plasmid DNA. J Bacteriol 173:1353–1356. https://doi.org/10.1128/jb.173.3.1353-1356.1991
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Ni D, Xu P, Gallagher S (2016) Immunoblotting and immunodetection. In: Coligan JE (ed) Current protocols in immunology, Wiley, Hoboken, pp 8.10.11–18.10.36
Peng Q, Wang G, Liu G, Zhang J, Song F (2015) Identification of metabolism pathways directly regulated by sigma54 factor in Bacillus thuringiensis. Front Microbiol 6:407. https://doi.org/10.3389/fmicb.2015.00407
Sambrook BJ, Russell DW (2015) Molecular cloning. In: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sanchis V, Gohar M, Chaufaux J, Arantes O, Meier A, Agaisse H, Cayley J, Lereclus D (1999) Development and field performance of a broad-spectrum nonviable asporogenic recombinant strain of Bacillus thuringiensis with greater potency and UV resistance. Appl Environ Microbiol 65:4032–4039
Saxild HH, Andersen L, Hammer K (1996) Dra-nupC-pdp operon of Bacillus subtilis: nucleotide sequence, induction by deoxyribonucleosides, and transcriptional regulation by the deoR-encoded DeoR repressor protein. J Bacteriol 178:424–434. https://doi.org/10.1128/jb.178.2.424-434.1996
Schaeffer P, Millet J, Aubert JP (1965) Catabolic repression of bacterial sporulation. Proc Natl Acad Sci U S A 54:704–711. https://doi.org/10.1073/pnas.54.3.704
Schnepf E, Crickmore N, Van Rie J, Lereclus D, Baum J, Feitelson J, Zeigler DR, Dean DH (1998) Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol Mol Biol Rev 62:775–806
Shu C, Yu H, Wang R, Fen S, Su X, Huang D, Zhang J, Song F (2009) Characterization of two novel cry8 genes from Bacilllus thuringiensis strain BT185. Curr Microbiol 58:389–392. https://doi.org/10.1007/s00284-008-9338-y
Skerlova J, Fabry M, Hubalek M, Otwinowski Z, Rezacova P (2014) Structure of the effector-binding domain of deoxyribonucleoside regulator DeoR from Bacillus subtilis. FEBS J 281:4280–4292. https://doi.org/10.1111/febs.12856
Wang G, Zhang J, Song F, Wu J, Feng S, Huang D (2006) Engineered Bacillus thuringiensis GO33A with broad insecticidal activity against lepidopteran and coleopteran pests. Appl Microbiol Biotechnol 72:924–930. https://doi.org/10.1007/s00253-006-0390-x
Weigel D, Glazebrook J (2010) Transmission electron microscopy (TEM) freeze substitution of plant tissues. Cold Spring Harb Protoc. https://doi.org/10.1101/pdb.prot4959
Wong HC, Schnepf HE, Whiteley HR (1983) Transcriptional and translational start sites for the Bacillus thuringiensis crystal protein gene. J Biol Chem 258:1960–1967
Xue J, Liang G, Crickmore N, Li H, He K, Song F, Feng X, Huang D, Zhang J (2008) Cloning and characterization of a novel Cry1A toxin from Bacillus thuringiensis with high toxicity to the Asian corn borer and other lepidopteran insects. FEMS Microbiol Lett 280:95–101. https://doi.org/10.1111/j.1574-6968.2007.01053.x
Yang H, Wang P, Peng Q, Rong R, Liu C, Lereclus D, Zhang J, Song F, Huang D (2012) Weak transcription of the cry1Ac gene in nonsporulating Bacillus thuringiensis cells. Appl Environ Microbiol 78:6466–6474. https://doi.org/10.1128/aem.01229-12
Yang J, Peng Q, Chen Z, Deng C, Shu C, Zhang J, Huang D, Song F (2013) Transcriptional regulation and characteristics of a novel N-acetylmuramoyl-L-alanine amidase gene involved in Bacillus thuringiensis mother cell lysis. J Bacteriol 195:2887–2897. https://doi.org/10.1128/JB.00112-13
Yoshisue H, Fukada T, Yoshida K, Sen K, Kurosawa S, Sakai H, Komano T (1993) Transcriptional regulation of Bacillus thuringiensis subsp. israelensis mosquito larvicidal crystal protein gene cryIVA. J Bacteriol 175:2750–2753. https://doi.org/10.1128/jb.175.9.2750-2753.1993
Zeng XM, Saxild HH, Switzer RL (2000) Purification and characterization of the DeoR repressor of Bacillus subtilis. J Bacteriol 182:1916–1922. https://doi.org/10.1128/JB.182.7.1916-1922.2000
Zhang JB, Schairer HU, Schnetter W, Lereclus D, Agaisse H (1998) Bacillus popilliae cry18Aa operon is transcribed by sigma(E) and sigma(K) forms of RNA polymerase from a single initiation site. Nucleic Acids Res 26:1288–1293. https://doi.org/10.1093/nar/26.5.1288
Zheng Q, Wang G, Zhang Z, Qu N, Zhang Q, Peng Q, Zhang J, Gao J, Song F (2014) Expression of cry1Ac gene directed by PexsY promoter of the exsY gene encoding component protein of exosporium basal layer in Bacillus thuringiensis. Acta Microbiol Sin 54:1138–1145
Zhou C, Zheng Q, Peng Q, Du L, Shu C, Zhang J, Song F (2014) Screening of cry-type promoters with strong activity and application in Cry protein encapsulation in a sigK mutant. Appl Microbiol Biotechnol 98:7901–7909. https://doi.org/10.1007/s00253-014-5874-5
Acknowledgements
We thank F He (CAAS) and X Tian (CAAS) for participating in some of the work, S Huang (CAAS) for providing technical support in preparing the AFM pictures, and X Chen (CAAS) and Y Xiao (CAAS) for some experimental performances.
Funding
This study was funded by the National Natural Science Foundation of China (No. 31530095 and No. 31300085).
Author information
Authors and Affiliations
Contributions
XZ, JZ, DS, and FS designed the experiments. XZ performed the experiments. TG, QP, and FS analyzed the results. LS analyzed the RNA-Seq data. TG, YC, and FS wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors
Electronic supplementary materials
Supplementary Table S1
(PDF 266 kb)
Rights and permissions
About this article
Cite this article
Zhang, X., Gao, T., Peng, Q. et al. A strong promoter of a non-cry gene directs expression of the cry1Ac gene in Bacillus thuringiensis. Appl Microbiol Biotechnol 102, 3687–3699 (2018). https://doi.org/10.1007/s00253-018-8836-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8836-5