Abstract
Substrate preparation (i.e., composting) for Agaricus bisporus cultivation is the most critical point of mushroom production. Among many factors involved in the composting process, the microbial ecology of the system is the underlying drive of composting and can be influenced by composting management techniques. Pasteurization temperature at the beginning of phase II, in theory, may influence the bacterial community and subsequently the “selectivity” and nutrition of the final substrate. Therefore, this hypothesis was tested by simulation in bioreactors under different pasteurization conditions (57 °C/6 h, 60 °C/2 h, and 68 °C/2 h), simulating conditions adopted by many producers. Bacterial diversity, based on 16S ribosomal RNA obtained by high-throughput sequencing and classified in operational taxonomic units (OTUs), was greater than previously reported using culture-dependent methods. Alpha diversity estimators show a lower diversity of OTUs under a high-temperature pasteurization condition. Bacillales order shows a relatively higher OTU abundance under a high-pasteurization temperature, which also was related to high ammonia emission measurements. On the other hand, beta diversity analysis showed no significantly changes in the bacterial community structure under different conditions. Agaricus bisporus mycelium growth during a standard spawn run period was significantly slower in the compost pasteurized at high temperature. Since the bacterial community structure was not greatly affected by different pasteurization conditions but by-products left (e.g., ammonia) at the end of compost conditioning varied, further studies need to be conducted to determine the functional role of the microbial communities found during substrate preparation for Agaricus bisporus cultivation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Agaricus bisporus (both button and portabella mushroom) cultivation contributes greatly to the agriculture economy in the USA, which in 2015 produced approximately 418,000 t valued at 1.19 billion USD [1]. Among many factors involved in A. bisporus production, substrate preparation (i.e., composting) is considered the most critical step. Mushroom substrate preparation is a complex biological process to manage and represents a major cost for mushroom producers.
The meticulous substrate preparation process has improved over recent years by utilizing specific composting materials such as straw-bedded horse manure, wheat straw, mulch hay, corn stover, poultry manure, grain distillers waste, oilseed meal, and gypsum [2, 3]. However, it is still based on a simple understanding of phase I and II composting concepts developed during the last century [4,5,6]. Phases I and II composting is a continuous process but varies in the nature and function of each phase. Phase I can be carried out in a rick (outdoor windrow) or an aerated bunker (indoor) over a 5–14-day period, depending on the raw materials and composting system. The intense microbial activity releases heat (temperatures can exceed 80 °C within 24–36 h) and breaks down complex carbohydrates such as cellulose and hemicellulose, resulting in a lignin-humus complex [2, 7]. Phase II has two main objectives, pasteurization and conditioning. Both stages are executed in a controlled environment, either in aerated tunnels (rectangular-shaped buildings containing bulk compost) or in a growing room (shelf beds or trays) under specific temperature regimes. Pasteurization temperatures are reached in tunnels by self-heating and by utilizing steam if done in growing rooms. Pasteurization set points typically range between 57 and 62 °C for 2–8 h to kill insects, nematodes, competitor species, and pathogens. Conditioning temperature ranges are typically between 47 and 48 °C for 5–8 days and is necessary to remove the ammonia formed during phase I and to establish a beneficial microbiota which can feed and stimulate A. bisporus growth [7,8,9,10].
The microbial ecology of the substrate is the underlying drive of the composting processes and has been the impetus of several studies in the past. This effort to isolate microorganisms in culture and subsequently identify by morphology, and more recently by molecular analyses, has been investigated to look for a relationship with A. bisporus growth and/or yield [11,12,13,14,15,16,17]. The majority of these reports [11,12,13,14,15] focused on thermophilic bacteria and fungi such as Scytalidium, Torula, Chaetomium, Humicola, Mucor, Penicillium, Aspergillus, Monilia, Fusarium, Epicoccum, Trichoderma, Nocardia, Pseudonocardia, Streptomyces, Thermoactinomyces, Thermomonospora, Talaromyces, and Stibella, which are typically cultured in the laboratory (culture-dependent methods). More recently [16, 17], a fingerprint and library clones sequencing methods was used to reveal the presence of certain bacterial orders during composting for A. bisporus cultivation such as Bacillales, Xanthomonadales, Clostridiales, Pseudomonadales, Themales, Halanaerobiales, Thermoanaerobacteriales, Actinomycetales, and Acidimicrobiales. The bacterial succession previously identified during composting should be reinvestigated, since in recent years, improvements in molecular biology tools have expanded the microbiology frontiers, revealing that a vast majority of bacteria found in environmental samples were not able to be cultured in the laboratory [18,19,20]. In addition, recent studies [21, 22] using high-throughput sequencing revealed the presence of bacterial 16S rRNA sequences for several phyla during composting processes (Verrucomicrobia, Spirochaetes, Proteobacteria, Planctomycetes, Gemmatimonadetes, Firmicutes, Chloroflexi, Chlorobi, Bacteroidetes, Actinobacteria, and Acidobacteria). These observations were done based on waste composting management (over time and compost recipes) with samples taken over long periods of time (months).
Different from a conventional manure or waste composting, with the goal of making a stable soil organic amendment, substrate preparation for A. bisporus is an extremely controlled, high-temperature system that occurs for a much shorter period of time. The microbial ecology of this system is not well understood and can be influenced by temperature management during phase II [23,24,25]. Since the conditioning step is recognized as fundamental to compost preparation, the adjustment of temperature at the beginning of phase II (pasteurization), in theory, can modify the bacterial community for the subsequent conditioning step and ultimately affect the compost selectivity for A. bisporus cultivation. Therefore, the objective of this work was to use a culture-independent method (direct recover amplicon) was used to evaluate the bacterial community changes under different phase II temperature regimes during pasteurization in an experimental composting system.
Material and Methods
Composting Phases I and II
Phase I composting was prepared at the Mushroom Research Center (MRC) at The Pennsylvania State University, University Park campus, PA, USA, in an aerated bunker, following standard methods, followed by phase II composting simulated in bioreactors. The standard formula consisted of wheat straw-bedded horse manure, dried poultry manure, dried distiller’s grain, and gypsum (Table S1). The raw materials were mixed in a Jaylor® feed mixer to reach approximately a 60% starting moisture. The mixed ingredients were filled into a bunker with a forced aeration system supplying fresh air to prevent anaerobic conditions. At day 3, the compost was removed from the bunker, remixed, watered, and returned to the bunker for an additional 3 days, totaling 6 days of phase I. In phase II, bioreactors were adapted to simulate three different pasteurization conditions, 57 °C/6 h, 60, °C/2 h, and 68 °C/2 h (Fig. 1). Four-liter reactors were made from 15.2-cm-diameter polyvinyl chloride (PVC) pipe, compost was supported on a rigid, and stainless steel grid was fixed 5 cm from the bottom of the reactor. Fresh humidified air was supplied to the bottom of the bioreactors through plastic tubing, providing an aerobic condition for the compost (Fig. S1). Each bioreactor was filled with 1.5 kg of fresh compost collected immediately at the end of phase I, and three replicates were applied for each time/temperature combination during the different pasteurization conditions. After pasteurization, the transition from pasteurization to conditioning was managed quickly and adjusted to a constant conditioning temperature at 47 °C for 6 days, similar to commercial production practices.
Experimental design and temperature settings during composting phases I and II. Vertical line in the middle of graph separates phases I and II. Diagram above shows sampled points. Phase I (self-heating) two samples: D0 and D6. Phase II (bioreactors) 6 samples: after pasteurization P57, P60, and P68 and after conditioning C57, C60, and C68
Sampling and DNA Extraction
Samples were obtained from eight different points during the composting period: mixed raw materials at the beginning of phase I (D0), at the end of phase I (D6), after pasteurization (P57, P60, and P68), and after conditioning (C57, C60, and C68) (Fig. 1). For each sampling point/period, three samples were collected and immediately frozen in liquid nitrogen and kept at −80 °C until DNA extraction. Briefly, 10 g of frozen compost were transferred to a 50-ml falcon tube with 30 ml of saline solution (NaCl–0.9%, w/w), shook for 6 h on an orbital table at 150 rpm, and 12 ml of the supernatant was transferred to a 15-ml falcon tube. The supernatant was centrifuged at 4000 rpm for 20 min at 4 °C, and 300 mg of the pelletized cells was used for DNA extraction using PowerLyser™ PowerSoil DNA Isolation Kit (Mo Bio, Carlsbad, CA, USA) following manufacturer’s instructions. Subsequently, extracted genomic DNA was visualized in an agarose gel (1%, 100 v for 45 min) and quantified using a spectrophotometer (NanoDrop, 2000c model, Thermo Scientific). To improve the genomic DNA quality, a DNA cleanup kit was used (PowerClean Pro DNA kit—Carlsbad, CA, USA) to remove humic acids and other PCR inhibitory substances with a subsequent visualization in agarose gel and quantification in a spectrophotometer following the cleanup step.
Library Preparation and Sequencing of Bacterial Ribosomal Markers 16S rRNA
The V4 variable region of the 16S ribosomal RNA gene was amplified in triplicate (pooled after amplification) using primers F515 5′-GTGCCAGCMGCCGCGGTAA-3′ and R806 5′-GGACTACHVGGGTWTCTAAT-3′ [26], with Illumina’s barcodes on the forward and reverse primers. For each PCR reaction, 40 ng of extracted genomic DNA was used. The reaction was carried out with 5.0 μl of buffer 5X (Q5 reaction buffer), 0.5 μl of dNTPs (10 μM each), 1.25 μl each primer (forward and reverse, 10 μM each), 0.25 μl Q5 high-fidelity DNA polymerase and adjusted with nuclease-free water for a final volume of 25 μl. PCR conditions were an initial denaturation at 98 °C for 30 s, followed by 30 cycles at 98 °C for 10 s, 50 °C for 30 s, 72 °C for 30 s and ended with a final extension step at 72 °C for 5 min. After amplification, PCR products were verified in a 1% agarose gel (45 min/100 v) and quantified by spectrophotometer NanoDrop 2000c (Thermo Scientific). Subsequent steps included a cleaning step using Agencourt AMPure magnetic beads (Beckman Coulter, Brea, CA, USA); visualization in the Bioanalyzer; and quantification using an Invitrogen Qubit HS double-stranded DNA (dsDNA) kit (Invitrogen Carlsbad, CA, USA). Amplicon samples were pooled in equimolar concentrations (10 nM) following Illumina’s protocol. Sequencing was carried out using Illumina MiSeq platform (paired 300-bp read lengths) at Huck Institutes of the Life Sciences at The Pennsylvania State University, University Park, PA, USA.
Sequences Processing
Raw Illumina bacterial sequence reads were processed using mothur package v.1.38.1 [27] following the pipeline analysis from Schloss laboratory (SOP–Standard Operating Procedures) [28]. Forward and reverse reads were paired based on their quality score. In subsequent screenings, no ambiguous base codes were allowed, and primers as well as reads shorter than 250 and longer than 260 bp were removed, generating a read average length of 253 bp. The dataset was dereplicated and unique sequences were aligned to a 50,000-column wide SILVA-based reference alignment using the Needleman-Wunsch pairwise alignment method. Sequence classifications were made against the SILVA database (release v123) containing 152,308 bacterial small subunit (SSU) rRNA sequences as well as Archaea and Eukarya sequences [29]. After alignment, sequences with homopolymers exceeding 8 bp were removed. To reduce the sequencing error and number of unique sequences, the dataset was further denoised through a pre-clustering algorithm initially developed by Huse et al. [30] and modified and implemented in mothur [31]. Using this method, sequences were sorted by decreasing abundance and then sequentially compared to each of the rarer sequences. If a rare sequence is less than a specified number of bases (1-nt difference per 100-nt) different from the more abundant sequence, the rare sequence is removed from the dataset and its abundance is added to the more abundant sequence. Chimera sequences were treated separately from sequencing error using the UCHIME algorithm [32] by comparing all sequences to the most abundant sequences in the dataset, which is implemented in mothur. Unspecific amplification products (mitochondria, chloroplasts, Archaea, Eukarya, and unknown domain) were removed by a mothur command. Denoised and high-quality sequences were classified by a naïve Bayesian classifier [33] implemented in mothur. Finally, sequences were split into groups corresponding to their taxonomy at the level of order and then assigned to operational taxonomic units (OTUs) at a 3% dissimilarity level [34]. To normalize the sequencing effort in all 24 samples without compromising the estimated amplicon diversity, all samples were randomly subsampled 1000 times to the number of reads in the sample with the fewest reads, n = 22,000 and n = 25,000 excluding and including OTU singletons, respectively. Downstream alpha- and beta-diversity measurements were performed excluding OTU singletons’ and results and discussion focused on datasets excluding OTU singletons but a consideration was made to both strategy analyses (see, e.g., [35]).
Data Analysis
An average (%) of assigned OTUs per sample was plotted in a bar plot at phylum and order levels. Sampling effort was verified by Good’s coverage measure [36] and rarefaction curves in mothur. Rarefaction curves were calculated using 1000-fold resampling without replacement. Alpha diversity estimators (Shannon-Wiener [H], the inverse Simpson [1/D], richness estimator Chao 1, and Sobs–observed OTUs) were applied to compare the total diversity among the communities. The reciprocal Simpson index (Inverse Simpson) was selected for the results and discussion because it represents the number of uniformly distributed OTUs that were required to have the same diversity as the actual community; thus giving it an easier biological interpretation compared to others indices (e.g., Shannon) [31]. Alpha diversity estimations generated in mothur were compared by ANOVA (p < 0.05) followed by t-test pairwise comparisons with Bonferroni correction in R program version 3.3.1 (R Development Core Team; http://www.R-project.org [37]). All graphic visualization was generated by ggplot2 R package [38], except Venn diagram and rarefaction curves, which were plotted in Excel. Beta diversity was calculated applying Jaccard and Theta Yue and Clanton [39] dissimilarity coefficients implemented in mothur. Beta diversity measures were supported by non-metric multidimensional scaling ordination plots [40], heatmap, and a dendrogram [27]. Statistical analyses of beta diversity measures were computed using parsimony, analysis of molecular variance (AMOVA), and homogeneity of molecular variance (HOMOVA) [41,42,43] by pairwise comparisons with Bonferroni correction in mothur. Good’s coverage [36], rarefaction curves, shared OTUs (Venn diagram), and alpha- and beta-diversity measures were all calculated in mothur to a normalized number of OTUs (observed richness).
Ammonia Emission during Phase II
Ammonia emission concentration (NH3) of exhaust gas from the bioreactors (phase II) was evaluated over two periods, during pasteurization and during conditioning. Boric acid flasks were kept in a cold water bath in a 4 °C refrigerator (Fig. S1). The boric acid flask was filled with a solution of 42 g l−1 boric acid plus a bromocresol green-methyl red indicator. The boric acid flasks were replaced after pasteurization and again after conditioning. To determine the amount of ammonia released from the compost and captured in the flasks, the boric acid solution was titrated with 0.7 M hydrochloric acid (HCL) with the end point defined as the color change from yellow to pink (original color) [44] . Each milliliter of HCL consumed during the titration represents 11.9 mg of NH3. Ammonia emission was expressed in mg of NH3/ton of substrate/h. In addition, the total nitrogen (Kjeldahl method) and total carbon (by loss ignition method) were analyzed for the compost samples.
Agaricus bisporus Mycelium Growth Ratio during Phase III
To evaluate A. bisporus mycelium growth rate in the compost generated under different pasteurization conditions, an experimental spawn run was modeled in “race tubes” (glass tube, 4 cm diameter×20 cm height). Conditioned compost obtained from bioreactors (100 g after conditioning) was inoculated with 4% commercial millet spawn (w/w, strain 901 Lambert) and incubated at 22 °C. The growth measurement was initiated 3 days after inoculation with subsequent 2-day intervals until complete compost colonization. Growth rate comparison was made against a commercial standard spawn run period (~12 days) by ANOVA and posterior Tukey test (p < 0.05).
Results
As a basic indicator of the composting progress, compost temperatures of 70 and 80 °C were recorded on days 2 and 4 of the phase I process, respectively (Fig. 1). These are temperatures typically observed in commercial phase I mushroom composting systems. Bacterial communities recovered by direct amplicon sequencing generated approximately 1.8 million raw paired Illumina reads for eight sampled points (24 samples). After processing the quality of the reads in mothur, ~1.2 million reads remained, comprised of 152,617 unique sequences which were clustered into 96,227 OTUs, including OTU singletons (Table S2). Sampling effort verified to a normalized number of sequences still yielded sufficient resolution of bacterial communities (as suggested by a Good’s coverage measures averaged across all sampled points) of 90.5 and 98% including and excluding OTU singletons, respectively (Table S3). Rarefaction curves based on the number of observed OTUs revealed a better sampling intensity excluding OTU singletons (Fig. S2).
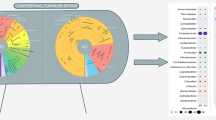
Relative abundances of OTUs per sample, at both the phylum and order levels, are plotted in Fig. 2. Firmicutes were dominant at all sample points followed by Proteobacteria, Actinobacteria, and Bacteroidetes, whether including OTU singletons or not. These four phyla had an average of ~91% assigned OTUs across all sampled points and ~9% for the other phyla, whether including OTU singletons or not (Table S4). Removing OTU singletons changed the total number of phyla from 19 to 16, excluding Spirochates, Saccharibacteria, and Synergistetes (Table S4). During the pasteurization stage (P57, P60, and P68), Firmicutes showed an average of ~81% abundance, excluding OTU singletons (Fig. 2a). Bacillales was the most abundant order (Firmicutes) among all sampled points, particularly during the pasteurization stage comprising 50.2, 54.1, and 66.5% of OTUs of the P57, P60, and P68 sample points, respectively (Fig. 2b and Table S4). During conditioning, the relative abundance of Bacillales was still predominant, but decreased to 11.4, 14.5, and 43.4% at C57, C60, and C68, respectively. Clostridiales, Lactobacillales, and Micrococcales, also Firmicutes members, were the most abundant orders besides Bacillales at all sample times (Fig. 2b), excluding or including OTU singletons. The total number of classified orders dropped to 13.2%, removing OTU singletons (82 to 72 orders, data not shown).
Relative abundance of operational taxonomy units (OTUs) at the phylum (a) and order level (b) across composting phases I and II. OTUs were classified using the Silva database (release 123) > 97% of identity. Samples: phase I–D0 and D6; phase II (pasteurization)–P57, P60, and P68; phase II (conditioning)–C57, C60, and C68
Alpha diversity is a general measure of species based on OTUs’ richness and is summarized in Fig. 3. All diversity indices applied (Inverse Simpson, Chao1 and Shannon) as well as observed number of OTUs (Sobs) were significantly higher in the low temperature points (mixed raw materials–D0 and conditioning stage C57, C60, and C68) (Fig. 3 and Table S6). Each index has a different approach behind the calculation but similar diversity tendencies were observed with each method; however, different methods used to calculate the indices and the exclusion of OTU singletons can change the diversity among samples [35]. Removing OTU singletons changed the magnitude of total diversity (Table S3), but not the statistical significantly among sampled points (data not show).
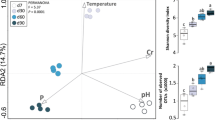
Among several beta measures, Jaccard and ThetaYC were chosen and applied to examine the extent to which different composting stages (mainly pasteurization temperatures) can influence the microbial community structure during composting. Community structure dissimilarity among sampled points during composting were compared using ThetaYC index and visualized by non-metric multidimensional scaling plot (nMDS) and by heatmap (Fig. 4). At the end of phase II, composting samples C57 and C60, previously pasteurized at low temperature (P57 and P60), showed higher similarity as observed in heatmap (Fig. 4a). Phase I samples (D0 and D6) grouped separately from each other as well as from phase II samples (nMDS) (Fig. 4b). The high pasteurization temperature sample (P68) grouped separately from samples submitted at low pasteurization temperatures (P57, P60), which grouped together. After conditioning the samples, previously submitted at low pasteurization temperature (C57 and C60) also grouped separately from the sample pasteurized at high-temperature C68. AMOVA applied to the community structure showed no statistical difference, whether including OTU singletons or not (Table S7). HOMOVA also showed no statistical difference among sampled points (Table S7). Community relationships during composting were calculated using Jaccard and ThetaYC dissimilarity coefficients and verified by dendrogram and analyzed by parsimony. Four distinct clusters were observed in the dendrogram (Fig. S3): samples D0 and D6, samples P57, P60, P68, and C68 as well as samples C57 and C60, in both coefficients. Parsimony analysis showed no statistical difference between samples points (Table S8).
Heatmap (a) and non-metric multidimensional scaling plot ((b), two dimensions) comparing the structure similarity among the communities sampled across composting phases I and II. Both visualization methods were applied by ThetaYC (Yue and Clayton) coefficient as measure of similarity. Heatmap–0 (blue) more similarity than 1 (red). Samples: phase I–D0 and D6; phase II (pasteurization)–P57, P60, and P68; phase II (conditioning)–C57, C60, and C68
OTUs’ distribution across sample points during composting is summarized in Fig. 5. During phase I, a total of 1703 OTUs were assigned. At D0 (mixed materials) a total of 989 OTUs were assigned, and at D6 (pasteurization peak heating) 427 OTUs were assigned; between these samples, 287 OTUs were shared. Immediately following pasteurization, the 3-way Venn diagram (P57, P60, and P68) showed a total of 4161 OTUs of which 275 were shared among the three communities. The samples taken after pasteurization at 57 °C (P57) had the highest number of OTUs assigned, 1700 (including shared OTUs). After the conditioning stage (C57, C60, and C68) a total of 3902 OTUs were assigned and the highest number of OTUs was again associated with the lowest pasteurization temperature (C57), 1847 (including shared OTUs). It should be noted that the number of OTUs, after excluding singletons, dropped approximately 40% but still demonstrated a similar distribution across sampled points (Table S2).
As an important nutrient, the nitrogen loss of the compost was monitored by quantifying ammonia emissions during phase II composting under different pasteurization conditions (temperatures). Ammonia emission during pasteurization and conditioning was recorded for each treatment during phase II composting. Ammonia emissions were higher during the pasteurization stage for the high-temperature treatment (P68) compared to the emissions at the other two pasteurization temperatures, which were 35 and 34% lower (P57 and P60 respectively) (Fig. 6b). After pasteurization, during conditioning, both compost samples pasteurized at the lower temperatures (P57 and P60) showed lower ammonia emissions (81 and 79% less, respectively) than compost pasteurized at 68 °C (Fig. 6b). The total nitrogen at the end of conditioning was slightly lower for the compost pasteurized at 68 °C (Fig. 6c). However, the C/N ratio of compost pasteurized at 68 °C was higher at the end of conditioning (Fig. 6d).
Mycelial growth in experimental spawn run (a), ammonia (NH3) emission (b) during pasteurization and conditioning (phase II), total nitrogen % (c) and C/N ratio % (d). Significant differences between groups (C57, C60, and C68) were evaluated by ANOVA, and t test pairwise comparisons are indicated by bars at the top of graph (a), mycelial growth (p-value legend: “null” = ns, * = <0.05, ** = <0.01, *** = <0.001)
High-temperature pasteurization compost (68 °C/2 h) demonstrated significantly slower mycelial growth in the “race tubes” after spawning compared to growth rates observed in the 57 and 60 °C pasteurized composts. Mycelial growth rates between the 57 and 60 °C pasteurization temperature compost showed no statistical difference from each other (Fig. 6a).
Discussion
This was the first report applying direct recovered amplicons to explore the bacterial community profiles during phase II substrate preparation for A. bisporus cultivation under different pasteurization conditions (temperatures). The number of bacterial taxa represented by 16 phyla (Fig. 2a) and approximately 80 orders (only the top 20 were plotted in Fig. 2b) which was higher than previously reported using fingerprint and clone library sequencing methods during substrate preparation (phases I and II) and during A. bisporus cultivation [16, 17, 45, 46]. Some of the previous reports [16, 17] were able to detect, by fingerprint and library clones sequencing, phyla such as Firmicutes, Proteobacteria, Deinococcus-Thermus and Actinobacteria, including 9 orders (Bacillales, Xanthomonadales, Clostridiales, Pseudomonadales, Thermales, Halanaerobiales, Thermoanaerobacteriales, Actinomycetales, and Acidimicrobiales. During mushroom cropping (casing layer samples), Siyoum et al. [45] using fingerprint (DGGE, denaturing gradient gel electrophoresis) and sequencing (purified sequence fragments from gel) were able to identify two phyla, Proteobacteria and Bacteroidetes. Wang et al. [46] using DGGE to compare the bacterial communities of two different substrate formulations (based on rice- and wheat-straw) during composting for A. bisporus cultivation, reported differences on gel bands intensity, without any taxa information. These bacterial groups previously reported, especially Firmicutes, Proteobacteria, and Actinobacteria were three most abundant taxa detected by high-throughput sequencing in the current study, which can possibly explain the detection by less robust methods such as fingerprint and clones library sequencing. Most classified OTUs (~90%, Fig. 2a) were distributed among Firmicutes, Proteobacteria, Actinobacteria, and Bacteroidetes. However, Firmicutes was the most abundant phylum in the current survey, differing from previous reports, which used direct recovered amplicons with differences in primers (target region), sequences analysis (analysis strategies), and composting methods (waste composting) [21, 22]. Bacteroidetes was reported as the most abundant phylum during waste composting [21]. Proteobacteria was the most abundant phylum found during food and animal waste composting [22]. On the other hand, Ntougias et al. [47] reported that Firmicutes was the most abundant phylum in spent mushroom compost (compost remaining after cropping) by a clone library sequencing method. These results support the initial hypothesis that drove us to explore the bacterial communities present during substrate preparation under different pasteurization conditions using a robust amplicon sequencing methodology. It should be noted that results comparison between different studies should be carefully examined since the differences in experimental designs are considerably broad (samples type, primers, bioinformatics analysis, etc.) [48,49,50]. In addition, variability in both chemical as well as physical differences is difficult to account for as is collecting compost samples representing specified temperatures in a large system [51]. Collecting a representative sample from a commercial composting system is very difficult due to the heterogeneity of the compost matrix [52, 53]. The experimental design applied here, including bioreactors and high-throughput sequencing, was used to improve on the accuracy of bacterial community exploration during substrate preparation for A. bisporus cultivation. The bioreactor system and temperature settings were simulated based on commercial phase II compost observations. One of the advantages in an experimental scale (i.e., bioreactors) is the possibility to uniformly mix the raw materials as well as control the airflow (oxygen level) and temperatures, which are critical to solid fermentation performance [54, 55].
Another expected finding was the bacterial diversity based on total number of assigned OTUs, which was significantly lower from high-temperature compost, i.e., peak heating (D6), and after the pasteurization stage (P57, P60, and P68) (Fig. 3 and Table S6). The temperature within a composting matrix determines the rate at which many of the biological processes take place [56,57,58], which controls microbial population succession during composting [59,60,61]. Therefore, the temperature represents a significant factor in determining the relative advantage of some populations over others [62, 63]. Previous [64, 65] studies that isolated microorganisms during phase I (at the beginning and during the turns) and phase II (after pasteurization and conditioning) reported that some species present at the beginning of the process (mixed raw materials) disappeared during composting and other species, as well as the number of CFU’s (colony-forming unit), decreased at the high-temperature stages (peak heating and after pasteurization stage). It should be noted that analysis based on assigned OTUs is not a quantitative method. However, the compost temperature during pasteurization affected the bacterial diversity at the end of the conditioning stage (C57, C60, and C68). After conditioning, compost pasteurized at 68 °C demonstrated significantly lower diversity than the compost treatments pasteurized at lower temperatures (C57 and C60).
Although the alpha diversity was significantly different, pasteurization temperature settings had only limited or no statistical difference on the bacterial community structure among composting stages (Fig. 4 and Tables S7 and S8). It is worth remembering that the interval between sample points D6 (end of phase I) and P57, P60, and P68 (at the end of pasteurization) was separated by only a few hours (Fig. 1), which could contribute to the similarity observed among these communities. On the other hand, comparison among sample points after pasteurization and at the end of conditioning also showed no statistical differences in the communities’ structure (after 6 days of composting). Overlap or shifting of OTUs within the three composting stages (D0-D6; P57-P60-P68; C57-C60-C68) revealed that less than 50% of OTUs were shared between the three different composting stages (Fig. 5). Compost chemical changes based on carbon, nitrogen, and ammonia emission (Fig. 6), during composting possibly favored specialized bacterial groups across these different composting stages (Fig. 5) without affect the bacterial community structure.
The bacterial community present in compost has a direct effect on the mycelium nutrition mainly during the initial growth (spawn run) and represents an easily degradable source of carbon and nitrogen [66, 67]. Nonetheless, some compounds left by the microbial communities at the end of conditioning can inhibit or stimulate A. bisporus mycelium growth. One of the most notable compounds that can affect mycelial growth is ammonia. Ammonia concentration has a strong inhibition effect on A. bisporus mycelium growth and thus on mushroom production [68]. Free ammonia is required during phase I composting because it stimulates the bioconversion of mixed raw materials and releases polysaccharides from the lignin matrix making the straw carbohydrates more accessible to A. bisporus mycelium. However, free ammonia should be avoided at the end of conditioning to not adversely affect A. bisporus mycelium growth. As observed during phase II, ammonia emission was higher during pasteurization at 68 °C (Fig. 6b) with a significant effect on A. bisporus growth under an experimental spawn run condition (Fig. 6a). In addition, total nitrogen content was slightly lower at the end of conditioning for the compost pasteurized at 68 °C (Fig. 6c). These observations agree with previous studies [69, 70], which, using different bio-wastes and temperature profiles, demonstrated that increased ammonia emissions during composting have a correlation with high-temperature regimes. Carbon content also is an important nutrient and serves primarily as an energy source for the microorganisms, while a fraction of the carbon is incorporated into the microbial cell and the other fraction is lost to the atmosphere through CO2 emissions. Carbon loss during the decomposition process was more pronounced than nitrogen loss partially because of their relative abundance in the compost matrix (C > N), and both impacted the C/N ratio [71]. The carbon content at the beginning of the process (D0–mixed raw material) was 41.82% and dropped to 39.24, 38.22, and 39.66% at the end of conditioning for the compost pasteurized at 57, 60, and 68 °C, respectively (Fig. 6d). Eklind et al. [72] reported that waste composting in a laboratorial scale under different temperatures (40, 55, and 67 °C) showed highest CO2 emissions at a composting temperature of 55 °C (~15 days). Other variables can affect the carbon dynamics during composting such as oxygen level [73, 74] and initial compost composition [75]. In addition, the compost temperature, C/N ratio, and oxygen levels (phase I and phase II) used in the current study are quite different from waste composting.
Firmicutes members, in particular bacteria belonging to the Bacillales and Clostridiales orders (Fig. 2b) which are spore-formers and have the capability to grow in environments with high ammonia levels and high temperatures [76, 77], were the most abundant orders observed during high ammonia emission stages, pasteurization at 68 °C (Fig. 6b). These results do not provide an understanding regarding the metabolic role of Firmicutes members during composting but rather support the need for further explorations into the functionality of this particular bacteria group during A. bisporus substrate preparation with the goal focusing on better management of phase I and phase II composting and reducing free ammonia as efficiently as possible at the end of conditioning. Future studies should include analysis of RNA (metatranscriptomics) as well as proteomics to explore the functionality of these bacterial taxa for this particular composting process.
Conclusions
Bacterial dynamics documented during these experimental composting conditions revealed a broad bacterial diversity not reported before by less robust methods. Furthermore, limited changes in the community structure during composting should be examined because by-products left by the communities could vary at different pasteurization conditions and affect A. bisporus growth. As observed in ammonia emissions and mycelium growth, the high-temperature pasteurization setting had a negative impact during initial A. bisporus growth (spawn run). Bacillales members showed potential to be a key group of bacteria important during phase I composting, related to the breakdown and raw materials and creating a selective substrate for A. bisporus growth. Looking forward, further studies should include fungal community analyses by direct recover amplicons as well as investigating the functionality that these communities play in making a selective substrate for mushroom production. A better understanding of the biology of substrate preparation will lead to better decision-making for growers as they manage the composting process. Further experimentation is also needed to determine what role the microbial communities play in fruit body formation as well as disease development during mushroom cropping.
References
USDA, United States Department of Agriculture (2016) Farm resources, income, and expenses: mushrooms. Agricultural statistics: 2016 Washington, DC: US Government Printing Office; National Agricultural Statistics Service, Agricultural Statistics Board
Gerrits JPG (1988) Nutrition and compost. In: Van Griensven LJLD (ed) The cultivation of mushrooms, 1st ed. Rustington - Darlington Mushroom Laboratories, Sussex, United Kingdom, pp 29–72
Beyer DM (2003) Basic procedures for Agaricus mushroom growing. The Pennsylvania State University, University Park, PA, USA
Lambert EB (1931) Studies on the preparation of mushroom compost J. Agric. Res. 62(7):415–422
Sinden JW, Hauser E (1950) The short method of composting. In: Atkins FC (ed) Science and cultivation of edible fungi. Proceedings of the 1st international congress on the science and cultivation of edible fungi. Peterborough Mush, CA. pp 52–59. http://www.isms.biz/proceedings/
Sinden JW, Hauser E (1954) Mushrooms: cultivation, nutritional value, medicinal and its relation to short composting. In: Edwards RL (ed) Science and cultivation of edible fungi. Proceedings of the 1st international congress on the science and cultivation of edible fungi. Gembloux, BE, pp 123–131. http://www.isms.biz/proceedings/
Chang ST, Miles PG (2004) Mushrooms: cultivation, nutritional value, medicinal effect, and environmental impact. CRC, Boca Raton,
Wuest PJ, Bengtson GD (1982) Penn State handbook for commercial mushroom growers. The Pennsylvania State University, University Park, PA, USA
Van Griensven LJLD (1988) The cultivation of mushrooms. Rustington - Darlington Mushroom Laboratories, Sussex, United Kingdom
Straatsma G, Gerrits JPG, Thissen JTNM, Amsing JGM, Loeffen H, Van Griensven LJLD (2000) Adjustment of the composting process for mushroom cultivation based on initial substrate composition Bioresour. Technol. 72(1):67–74
Fergus CL (1964) Thermophilic and thermotolerant molds and actinomycetes of mushroom compost during peak heating Mycologia 56(2):267–284
Fordyce Jr C (1970) Relative number of certain microbial groups present in compost used for mushroom (Agaricus bisporus) propagation Appl. Microbiol. 20(2):196–199
Straatsma GJPG, Gerrits MPAM, Augustijn HJM, Op den Camp DV, Van Griensven LJLD (1989) Population dynamics of Scytalidium thermophilum in mushroom compost and stimulatory effects on growth rate and yield of Agaricus bisporus J. Gen. Microbiol. 135(4):751–759
Op den Camp HJM, Stumm CK, Straatsma GPJL, Derikx PJL, Van Griensven LJLD (1990) Hyphal and mycelial interactions between Agaricus bisporus and Scytalidium thermophilum on agar medium Microb. Ecol. 19(3):303–309
Straatsma GPJL, Samson RA, Olijnsma TW, Op Den Camp HJM, Gerrits JPG, Van Griensven LJLD (1994) Ecology of thermophilic fungi in mushroom compost, with emphasis on Scytalidium thermophilum and growth stimulation of Agaricus bisporus Appl Envir Microbiol 60(2):454–458
Székely AJ, Sipos R, Berta B, Vajna B, Hajdú C, Márialigeti K (2009) DGGE and T-RFLP analysis of bacterial succession during mushroom compost production and sequence-aided T-RFLP profile of mature compost Microb. Ecol. 57(3):522–533
Zhang X, Zhong Y, Yang S, Zhang W, Xu M, Ma A, Zhuang G, Chen G, Liu W (2014) Diversity and dynamics of the microbial community on decomposing wheat straw during mushroom compost production Bioresour. Technol. 170:183–195
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities Annu. Rev. Genet. 38:325–552
Rappé MS, Giovannoni SJ (2003) The uncultured microbial majority Annu. Rev. Microbiol. 57:369–394
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms Microbiol. Mol. Biol. Rev. 68(4):669–685
Neher DA, Weicht TR, Bates ST, Leff JW, Fierer N (2013) Changes in bacterial and fungal communities across compost recipes, preparation methods and composting times PLoS One 8(11):e79512. doi:10.1371/journal.pone.0079512
Martins FL, Antunes LP, Pascon RC, Oliveira JCF, Digiampietri LA, Barbosa D, Peixoto BM, Vallim MA, Viana-Niero C, Ostroski EH, Telles GP, Dias Z, Cruz BJ, Juliano L, Verjovski-Amleida S, Silva AM, Setubal JC (2013) Metagenomic analysis of a tropical composting operation at the São Paulo zoo park reveals diversity of biomass degradation functions and organisms PLoS One 8(4):e61928. doi:10.1371/journal.pone.0061928
Ross RC, Harris PJ (1983) An investigation into the selective nature of mushroom compost Sci. Hortic. 19(1–2):55–64
Ross RC, Harris PJ (1982) Some factors involved in phase II of mushroom compost preparation Sci. Hortic. 17(3):223–229
Savoie J-M, Olivier J-M, Laborde J (1996) Changes in nitrogen resources with increases in temperature during production of mushroom compost Would J Microbiol Biotechnol 12(4):379–384
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample Proc. Natl. Acad. Sci. U. S. A. 108(1):4516–4522
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities Appl. Environ. Microbiol. 75(23):7537–7541
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform Appl. Environ. Microbiol. 79(17):5112–5120
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools Nucleic Acids Res. 41:590–596
Huse SM, Welch DM, Morrison HG, Sogin ML (2010) Ironing out the wrinkles in the rare biosphere through improved OTU clustering Environ. Microbiol. 12(7):1889–1898
Schloss PD, Gevers D, Westcott LS (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies PLoS One 6(12):e27310. doi:10.1371/journal.pone.0027310
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection Bioinformatics 27(16):2194–2200
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy Appl. Environ. Microbiol. 73(16):5261–5267
Schloss PD, Westcott SL (2011) Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rRNA gene sequence analysis Appl. Environ. Microbiol. 77(10):3219–3226
Zhan A, Xiong W, He S, Maclsaac HJ (2014) Influence of artifact removal on rare species recovery in natural complex communities using high-throughput sequencing PLoS One 9(5):e96928. doi:10.1371/journal.pone.0096928
Good IJ (1953) The population frequencies of species and the estimation of population parameters Biometrika 40(3–4):237–264
R Development Core Team (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, http://www.R-project.org
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer Science & Business Media, New York, NY,
Yue JC, Clayton MK (2005) A similarity measure based on species proportion Commun Stat 34(11):2123–2131
Borg I, Groenen PJF (1997) Modern multidimensional scaling. Springer, New York, NY,
Martin AP (2002) Phylogenetic approaches for describing and comparing the diversity of microbial communities Appl. Environ. Microbiol. 68(8):3673–3682
Schloss PD (2008) Evaluating different approaches that test whether microbial communities have the same structure ISME J 2(3):265–275
Stewart Jr CN, Excoffier L (1996) Assessing population genetic structure and variability with RAPD data: application to Vaccinium macrocarpon (American cranberry) J. Evol. Biol. 9(2):153–171
Hong JH, Keener HM, Elwell DL (1998) Preliminary study of the effect of continuous and intermittent aeration on composting hog manure amended with sawdust Compost Sci Util 6(3):74–88
Siyoum NA, Surridge K, van der Linde EJ, Korsten L (2016) Microbial succession in white button mushroom production system from compost and casing to a marketable packed product Ann Microbiol 66(1):151–164
Wang L, Mao J, Zhao H, Li M, Wei Q, Zhou Y, Shao H (2016) Comparison of characterization and microbial communities in rice straw- and wheat straw-based compost for Agaricus bisporus production J. Ind. Microbiol. Biotechnol. 43(9):1248–1260
Ntougias S, Zervakis GI, Kavroulakis N, Ehaliotis C, Papadopoulou KK (2004) Bacterial diversity in spent mushroom compost assessed by amplified rDNA restriction analysis and sequencing of cultivated isolates Syst. Appl. Microbiol. 27(6):746–754
Tremblay J, Singh K, Fern A, Kirton ES, He S, Woyke T, Lee J, Chen F, Dangl JL, Tringe SG (2015) Primer and platform effects on 16S rRNA tag sequencing Front Microbial 6:771
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies Nucleic Acids Res. 41(1):e1
Rosselli R, Romoli O, Vitulo N, Vezzi A, Campanaro S, Pascale F, Schiavon R, Tiarca M, Poletto F, Concheri G, Valle G, Squartini A (2016) Direct 16S rRNA-seq from bacterial communities: a PCR-independent approach to simultaneously assess microbial diversity and functional activity potential of each taxon Sci Rep 6:32165
Jurak E, Punt AM, Arts W, Kabel MA, Gruppen H (2015) Fate of carbohydrates and lignin during composting and mycelium growth of Agaricus bisporus on wheat straw based compost PLoS One 10(10):e013909
Van Lier JJC, Van Ginkel JT, Straatsma G, Gerrits JPG, Van Griensven LJLD (1994) Composting of mushroom substrate in a fermentation tunnel: compost parameters and a mathematical model Netherlands J Agr Sci 42(4):271–292
Mason IG (2006) Mathematical modeling o the composting process: a review Waste Manag. 26:3–21
Mohee R, Mudhoo A (2005) Analysis of the physical properties of an in-vessel composting matrix Powder Technol. 155:92–99
Agnew JM, Leonard JJ (2003) Literature review–the physical properties of compost Compost Sci Util 11(3):238–264
Barrena R, Canovas C, Sánchez A (2006) Prediction of temperature and thermal inertia effect in the maturation stage and stockpiling of a large composting mass Waste Manag. 26:953–959
Liang C, Das KC, McClendon RW (2003) The influence of temperature and moisture contents regimes on the aerobic microbial activity of a biosolids-composting blend Bioresour. Technol. 86(2):131–137
Smith SR, Lang NL, Cheung KHM, Spanoudaki K (2005) Factors controlling pathogen destruction during anaerobic digestion of biowastes Waste Manag. 25(4):417–425
Tiquia SM, Wan HC, Tam NFY (2002) Microbial population dynamics and enzyme activities during composting Compost Sci Util 10(2):150–161
Yu H, Zeng G, Huang H, Xi X, Wang R, Huang D, Huang G, Li J (2007) Microbial community succession and lignocellulose degradation during agricultural waste composting Biodegradation 18(6):793–802
Gajalakshmi S, Abbasi S (2008) Solid waste management by composting: state of the art Crit Rev Env Sci Tec 38(5):311–400
Komilis DP, Ham RK (2003) The effect of lignin and sugars to the aerobic decomposition of solid wastes Waste Manag. 23(5):419–423
Ishii K, Fukui M, Takii S (2000) Microbial succession during a composting process as evaluated by denaturing gradient gel electrophoresis analysis J. Appl. Microbiol. 89:768–777
Adhikary RK, Barua P, Bordoloi DN (1992) Influence of microbial pretreatment on degradation of mushroom nutrient substrate Int Biodeterior Biodegradation 30:233–241
Evered CE, Noble R, Atkey PT (1995) Microbial populations and straw degradation in mushroom composts prepared in controlled environments. In: Elliot, TJ (ed) Science and cultivation of edible fungi. Proceedings of the 14th International Congress on the Science and Cultivation of Edible Fungi. Oxford, UK. pp 245–250 http://www.isms.biz/proceedings/
Fermor TR, Wood DA (1981) Degradation of bacteria by Agaricus bisporus and other fungi J. Gen. Microbiol. 126:377–387
Fermor TR, Grant WD (1985) Degradation of fungal and actinomycete mycelia by Agaricus bisporus J. Gen. Microbiol. 131:1729–1734
González-Matute R, Rinker DL (2006) Compatibility of ammonia suppressants used in poultry litter with mushroom compost preparation and production Bioresour. Technol. 97:1679–1686
Pagans E, Barrena R, Font X, Sánchez A (2006) Ammonia emissions from the composting of different organic wastes. Dependency on process temperature Chemosphere 62:1534–1542
Fukumoto Y, Osada T, Hanajima D, Haga K (2003) Patterns and quantities of NH3, N2O and CH4 emissions during manure composting without forced aeration-effect of compost pile scale Bioresour. Technol. 89:109–114
Ryckeboer J, Mergaert J, Coosemans J, Deprins K, Swings J (2003) Microbiological aspects of biowaste during composting in a monitored compost bin J. Appl. Microbiol. 94:127–137
Eklind Y, Sundberg C, Smars S, Steger K, Sundh I, Kirchmann H, Jönsson H (2007) Carbon turnover and ammonia emissions during composting of biowaste at different temperatures J. Environ. Qual. 36:1512–1520
Tiquia SM, Richard TL, Honeyman MS (2002) Carbon, nutrient, and mass loss during composting Nutr Cycl Agroecosys 62:15–24
Richard TL, Walker LP, Gossett JM (2006) The effects of oxygen on solid-state biodegradation kinetics Biotechnol. Prog. 22(1):60–69
Eklind Y, Kirchmann H (2000) Composting and storage of organic household waste with different litter amendments. I: carbon turnover Bioresour. Technol. 74(2):115–124
Kelly LC, Cockell CS, Summers S (2012) Diverse microbial species survive high ammonia concentrations Int J Astrobiology 11(2):125–131
Partanen P, Hultman J, Paulin L, Auvinen P, Romantschuk M (2010) Bacterial diversity at different stages of the composting process BMC Microbiol. 10:94
Acknowledgments
This project was funded by the Coordination for the Improvement of Higher Education Personnel (CAPES–No. 005707/2012-05) and Penn State Mushroom Industry Endowment (Penn State University, USA). Thanks to Carl Schlagnhaufer for help during DNA extraction and amplification as well the Department of Plant Pathology and Environmental Microbiology at Penn State University for providing lab space and instruments. Sequence data for this study is deposited in NCBI SRA (Sequence Read Archive) under BioProject accession PRJNA377263 and will be available upon publication.
Authors’ Contribution
J.A.P. and F.R.V. conceived and designed the experiments. F.R.V and J.A.P. performed the experiments. F.R.V. analyzed the data. J.A.P. contributed reagents/materials/analysis tools. F.R.V. and J.A.P. wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 291 kb).
Rights and permissions
About this article
Cite this article
Vieira, F.R., Pecchia, J.A. An Exploration into the Bacterial Community under Different Pasteurization Conditions during Substrate Preparation (Composting–Phase II) for Agaricus bisporus Cultivation. Microb Ecol 75, 318–330 (2018). https://doi.org/10.1007/s00248-017-1026-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1026-7