Abstract
NK cells of metastatic melanoma (MM) patients display impaired function, making them incapable to mount an effective antitumor response. In this study, we evaluated immunophenotypic characteristics and functional capacity of CD3−CD16+ NK cells of MM patients in an in vitro model based on NK cell contact with an NK sensitive, K562, and a tumor-specific, melanoma FemX tumor cell line. Although our results indicate similar NK cell antitumor cytotoxic potential of MM patients in contact with both cell lines based on the expression of CD107a degranulation marker, there is a discrepancy in NK cell IFNγ production, as it is not significantly induced by FemX tumor cells, found to be, contrary to K562, HLA class I positive. Furthermore, we show NKG2D receptor downregulation by K562 tumor cell line, only. This may result from the obtained higher gene expression of TGFβ and VEGFA growth factors in K562 tumor cells that can negatively regulate NKG2D expression. Additionally, aside from postcontact downmodulation of activating CD16 receptor, there are no significant changes in the expression of CD161, CD158a, and CD158b NK cell receptors. Therefore, the applied in vitro model shows that, compared to the full NK cell functional capacity of MM patients displayed in a tumor-sensitive setting represented by contact with K562 cells, tumor-specific melanoma setting provided by FemX tumor cells leads to reduced NK functional potential. The obtained insight into NK cell capacity may be of use for evaluation of the state of disease and can help in selecting effective immunotherapeutic agents for MM patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Natural killer cells (NK) as the main constituent of innate antitumor immunity are phenotypically defined as CD3−CD16+ NK cells and are composed of two functionally distinct subsets: cytotoxic CD16bright and regulatory CD16dim subset based on CD16 receptor expression (Nagler et al. 1989). Additionally, expression of functionally opposing, activating, and inhibitory NK cell receptors regulates NK cell cytotoxicity against malignant cells. Ligation of activating receptors such as NKG2D to stress-induced MICA/B ligands stimulates, while binding of inhibitory killer immunoglobulin-like (KIR) receptors to major histocompatibility complex (MHC) class I molecules on tumor cells inhibits NK cell cytotoxicity. In this sense, it has been well established that the balance of activating and inhibitory signals mediated by these receptors affects functional profiles of NK cells (Lanier 2003, 2005; Farag and Caligiuri 2006).
Malignant melanoma (MM) is a potentially fatal form of skin cancer characterized by a rapid progression, metastasis to regional lymph nodes and distant organs, as well as by limited efficiency of currently applied standard and novel therapy, including immunotherapy (Monzon and Dancey 2012; de Coana et al. 2015). Although MM is an immunogenic tumor, a complex tumor-induced immunosuppressive network evolves with disease progression that includes a number of distinct effector cells and their secreted factors that induce NK cell anergy (Umansky and Sevko 2012; Gabrilovich and Nagaraj 2009). Previous immunological investigations have shown that NK cells of MM patients have multiple defects, including impaired cytotoxicity and subset distribution, displayed in CD3−CD16bright cytotoxic subset depletion, as well as in downregulation of activating NK cell receptors and poor cytokine production (Sibbitt et al. 1984; Konjevic et al. 2007, Konjevic et al. 2009a, b). Considering that NK cells are an important component of antitumor immunity, the extent of tumor-induced NK cell anergy in MM patients determines their ability to mount efficient antitumor response.
In light of this, in this study we used an in vitro model of NK cell-tumor cell contact to evaluate functional and immunophenotypic characteristics of NK cells of MM patients. In this model, two human tumor cell lines were used, K562 an erythromyleoid cell line (Klein et al. 1976) and FemX a tissue-specific melanoma-derived tumor cell line (Wong et al. 2011). K562 tumor cell line is an NK sensitive cell line as it does not express MHC class I molecules, a characteristic that distinguishes susceptible from less NK cell susceptible tumor cells (Lisovsky et al. 2015). Furthermore, K562 tumor cell line has a high expression of stress-induced MICA/B ligands for activating NKG2D receptor that further augments its NK cell susceptibility (Bae et al. 2012). Contrary to this, FemX melanoma cell line has been shown to be less NK cell sensitive, although it has not been well characterized especially with respect to the expression of MHC class I molecules and NK cell activating ligands (Ito et al. 2007). In this sense, additional characterization of these tumor cell lines would be helpful in NK cell functional evaluation after NK cell-tumor cell contact. Therefore, the aim of this study was to investigate the functional and immunophenotypic characteristics of NK cells of MM patients by using an in vitro model of NK cell-tumor cell contact with two tumor cell lines that differ in NK cell sensitivity. The obtained data should give insight into NK cell functionality relevant for disease evaluation and also show how a tumor-specific setting affects NK cell antitumor capacity of MM patients that may be of help in the selection of immunotherapeutic agents that can enhance NK cell activity.
Materials and Methods
Patients
In this study 14 patients with histologically proven MM in Stage IV, according to modified AJCC/UICC staging system, with median age of 55 years and no evidence of any other disease or infection were included. Before inclusion in the study, informed consent was signed by each patient and approved by the Ethical committee of Institute of Oncology and Radiology of Serbia.
Peripheral Blood Mononuclear Cell (PBMC) Isolation
PBMC were isolated from heparinized blood obtained from MM patients using Lymphoprep (Nypacon, Oslo, Norway) density gradient, centrifuged at 500×g for 40 min, and washed three times in RPMI 1640 culture medium (CM) (Sigma, St. Louis, USA) supplemented with 10% fetal calf serum (FCS) (Sigma).
Co-Culture of PBMC with Tumor Cell Lines
Freshly isolated PBMC of MM patients were adjusted to 3 × 106 cells/mL. One hundred µL of the PBMC suspension were mixed with tumor target cells, 100 µL of K562 cell line (2 × 106/mL) and 100 µL of human melanoma cell line FemX (2 × 106/mL) in six-well plates. Cell mixture was centrifuged at 100×g for 3 min and samples were incubated for 4 h at 37 °C in a humidified atmosphere in a CO2 incubator. PBMC were precipitated, washed and used for further immunophenotypical analysis before and after in vitro contact with tumor cell lines.
Flow Cytometric Analysis
Surface immuno-phenotype of peripheral blood lymphocytes (PBL) was identified using the following combinations of directly labeled monoclonal antibodies (mAbs): CD16FITC/CD3PE, CD161FITC/CD16PE/CD3PerCP, CD158aFITC/CD16PE/CD3PerCP, CD16FITC/CD158bPE/CD3PerCP (Becton–Dickinson, San Jose, USA), and CD16FITC/NKG2DPE/CD3PerCP (R&D, USA). The samples were prepared using 1 × 105 freshly isolated PBMC in 100 μL RPMI 1640 supplemented with 10% FCS, incubated for 30 min at 4 °C with 20 μL of appropriate mAb combination, washed twice with ice-cold phosphate-buffered saline (PBS), and fixed with 1% paraformaldehyde prior to FACS analyses (Jackson and Warner 1986). Surface marker expression was quantified on FACSCalibur flow cytometer (Becton–Dickinson, San Jose, USA). A total of 10,000–50,000 gated events verified as PBL, according to their physical characteristics, forward scatter characteristics (FSC), and side scatter characteristics (SSC), were collected per sample and analyzed using CellQUEST software. NK cells were gated within the lymphocyte subpopulation and according to their expression of CD3 and CD16 receptors were defined as CD3–CD16+. The two NK cell subsets were defined based on the density of CD16 antigen expression as CD3–CD16dim+ or CD3–CD16bright+ NK cells. NK cell receptors, NKG2D, CD161, CD158a, and CD158b on CD3–CD16+ NK cells were expressed as the percentage in PBL.
Flow Cytometric Analysis of Staining for Intracellular Molecules
Intracellular perforin, IFN-γ, pSTAT-4, pSTAT-5, pNFκB, and pZAP-70 were estimated in 1 × 106 PBMC. For the analysis of IFN-γ in the last 3 h of PBMC in vitro contact with tumor cell lines, Brefeldin A (10 µg/mL) was added. Cells were first stained for surface antigens with CD3PerCP and CD16 FITC or CD16 PE antibodies, fixed and after permeabilization with BD FACS permeabilizing solution 2 (BD Biosciences) stained with anti-perforin PE (R&D), anti-IFN-γ FITC, anti-pSTAT-4 PE, anti-pSTAT-5 PE, anti-pNFκB PE, and anti-pZAP-70 PE (Becton–Dickinson).
CD107 Expression Assay
As previously shown, CD107a (Becton–Dickinson) expression was estimated on CD3−CD16+ NK cells before and after 4 h of PBMC stimulation with target K562 and FemX tumor cells (Rubio et al. 2003).
Trypan Blue viability assay
Before and after contact with PBMC, viability of K562 and FemX tumor cells was evaluated with 0.4% of Trypan blue in PBS, and cell viability was determined as the number of viable cells divided by the total number of cells within the grids on the hemacytometer (Strober 2001).
Gene Expression Analyses
For the purpose of quantitative PCR (qPCR), total RNA was extracted from cell lines using TRI Reagent® BD kit (Sigma) according to the manufacturer`s recommendations. RNA bands were visualized on UV transilluminator and RNA concentration was determined spectrophotometrically (BioSpec Nano, Shimadzu). To prepare primary complementary DNA (cDNA) with random primers by RT-PCR, 2 µg total RNA was used as template for MultiScribe™ Reverse Transcriptase in a High-Capacity cDNA Reverse Transcription kit (Thermo Fisher Scientific). The reverse transcription reaction was conducted according to the manufacturer`s instructions.
qPCR Amplification
All target transcripts were detected using quantitative Real-time PCR (qPCR) and Taqman assays. TaqMan® Gene Expression Assays (VEGFA- Hs00900055_m1, VEGFARc1 Hs01052961_m1, TGFB-Hs00998133_m1, MMP2- Hs01548727_m1, BCL-2- Hs00608023_m, BAX- Hs00180269_m1, MET- Hs01565584_m) consist of a 20X mix of unlabeled PCR primers and TaqMan® MGB probes (FAM™ dye-labeled). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) for which data was obtained using TaqMan control reagents (Applied Biosystems- Hs02758991_g1) served as an endogenous control. PCR reactions were performed using an ABI Prism 7500 Sequence Detection System (Applied Biosystems). The PCR reaction conditions were described previously (Zec et al. 2014).
Statistical Analysis
Significance of differences for obtained results was carried out by nonparametric Wilcoxon test.
Results
After 4 h of in vitro contact of freshly isolated PBL of MM patients with K562 and FemX tumor cell lines, there is a significant increase (p < 0.05, Wilcoxon signed rank test) in the expression of CD107a degranulation marker on CD3−CD16+ NK cells (Fig. 1a). There is a decrease in the postcontact expression of cytolitic molecule perforin in NK cells as shown in a representative Flow cytometry histogram (Fig. 1b). After contact with NK cells, a significant decrease (p < 0.05, Wilcoxon signed rank test) in tumor cell viability is obtained in K562, while the decrease in FemX cell line is not significant (p > 0.05, Wilcoxon signed rank test) (Fig. 1c). However, analyses of intracellular IFNγ expression shows a significant increase (p < 0.05, Wilcoxon signed rank test) in the entire NK cell subset after a 4 h contact with K562 tumor cell line, only (Fig. 1d). The evaluation of pSTAT4, pSTAT5, and pZAP70 expression in NK cells after a 4 h contact with K562 and FemX tumor cell lines shows a decrease, although without statistical significance (p > 0.05, Wilcoxon signed rank test). However, postcontact expression of pNFκB in NK cells shows a nonsignificant increase (p > 0.05, Wilcoxon signed rank test) (Fig. 1e).
Evaluation of NK cell functional capacity of MM patients after 4 h in vitro contact with K562 and FemX tumor target cells shows that a the expression of CD107a degranulation marker on CD3−CD16+ NK cells of MM patients is significantly (*p < 0.05, Wilcoxon signed rank test) increased, b the expression of cytolytic molecule perforin in NK cells is decreased (a representative Flow cytometry histogram is given), c the viability of K562 tumor cells is significantly decreased (*p < 0.05, Wilcoxon signed rank test), while FemX cell line viability is not significantly decreased (p > 0.05, Wilcoxon signed rank test), d intracellular expression of IFNγ is significantly increased (*p < 0.05, Wilcoxon signed rank test) in NK cells after contact with K562 tumor cell line (a representative Flow cytometry dot plot is given), while after contact with FemX cell line it is not significantly increased (p > 0.05, Wilcoxon signed rank test) and e the expression of pSTAT4, pSTAT5, and pZAP70 in NK cells shows a nonsignificant (p > 0.05, Wilcoxon signed rank test) decrease, while postcontact expression of pNFκB in NK cells shows a nonsignificant increase (p > 0.05, Wilcoxon signed rank test)
The analysis of the expression of CD16 on CD3−CD16+ NK cells after 4 h in vitro contact of MM patients’ PBL with K562 and FemX tumor cell lines shows a significant decrease (p < 0.05, Wilcoxon signed rank test) after contact with K562, only (Fig. 2a). The analysis of the change in the postcontact distribution of the two NK cell subsets, shows a significant increase (p < 0.05, Wilcoxon signed rank test) in the CD3-CD16dim subset and a significant decrease (p < 0.05, Wilcoxon signed rank test) in the CD3−CD16bright NK cell subset (Fig. 2b, c).
The expression of CD16 on CD3−CD16+ NK cells, CD3−CD16bright and CD3−CD16dim NK cell subsets of MM patients after 4 h of in vitro contact with K562 and FemX tumor cell lines shows a a significant (*p < 0.05, Wilcoxon signed rank test) decrease in the expression of CD16 after contact with K562, only, b a significant increase in the CD3−CD16dim subset and a significant decrease in the CD3−CD16bright NK cell subset (**p < 0.01 and *p < 0.05, Wilcoxon signed rank test, respectively). c Representative flow cytometry histograms showing the change of CD16 antigen expression on CD3−CD16+ NK subsets
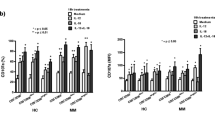
The analysis of receptor expression shows a significant decrease (p < 0.05, Wilcoxon signed rank test) in the expression of NKG2D on CD3−CD16+ NK cells in MM patients after 4 h contact with K562, only. With respect to CD161, there is no significant (p > 0.05, Wilcoxon signed rank test) change in its expression on CD3−CD16+ NK cells after 4 h contact either with K562 or FemX cell line (Fig. 3a). Analysis of the expression of two inhibitory receptors shows that after a 4 h contact with K562 or FemX cell lines there is no significant (p > 0.05, Wilcoxon signed rank test) change in the expression of CD158a or CD158b in MM patients on CD3−CD16+ NK cells (Fig. 3b).
The expression of NK cell activating and inhibitory receptors on CD3−CD16+ NK cells in MM patients after 4 h of in vitro contact with K562 and FemX tumor cell lines shows a a significant decrease (*p < 0.05, Wilcoxon signed rank test) in NKG2D after contact with K562 cells, only, and no significant (p > 0.05, Wilcoxon signed rank test) change in CD161 receptor, b representative flow cytometry dot plots for NKG2D are given, c no significant (p > 0.05, Wilcoxon signed rank test) change in the expression of inhibitory receptors CD158a or CD158b on NK cells
Evaluation of HLA class I molecule expression shows that K562 tumor cell line lacks, while FemX tumor cell line strongly expressesses HLA class I molecules (Fig. 4a). The evaluation of several gene expression using Quantitative Real-Time PCR (qPCR) shows differences between the two investigated tumor cell lines, K562 and FemX. The expression of VEGFA, its receptor VEGFRc1 and TGF-β1 gene is greater in K562 compared to FemX tumor cell line. On the contrary, matrix metalloproteinase-2 (MMP-2) gene expression is higher in FemX compared to K562 tumor cell line. Evaluation of proapoptotic BAX and antiapoptotic BCL-2 gene expression shows that K562 has a much greater proapoptotic potential compared to FemX cell line. However, the expression of MET gene that defines tumor invasiveness is similar in K562 and FemX tumor cells (Fig. 4b).
Molecular characterization of K562 and FemX tumor cell lines shows a high MHC class I molecule expression in FemX and a lack of expression in K562 tumor cell line given as flow cytometric histograms and b higher VEGFA and its receptor VEGFARc1, TGF-β1 and BAX gene expression, as opposed to lower BCL-2 and MMP-2 gene expression in K562 compared to FemX tumor cells, while MET gene expression is similar in these cell lines based on relative quantification of gene products using quantitative real-time RT-PCR
Discussion
Extensive former evaluation has shown that NK cells of MM patients in advanced clinical stage of disease display numerous functional profile impairments due to tumor-induced exhaustion (Konjevic et al. 2007, 2009b). In this sense, in this study we evaluated tumor-induced NK cell degranulation as an indicator of NK cell cytotoxic activity. Marker of cytotoxic degranulation, CD107a, is a lysosomal-associated membrane protein-1 (LAMP-1) expressed on the preformed lytic granules of NK cells that contain cytolitic molecules such as perforin and granzymes. NK cell-tumor cell contact strongly upregulatd expression of CD107a on NK cells that we found to be associated with intracellular loss of perforin owing to granule exocytoses, a finding that is in concordance with previous studies (Betts et al. 2003) as shown in Fig. 1a, b. In this study, we found that after contact with NK cell-sensitive K562 and tissue-specific melanoma-derived FemX tumor cell lines NK cells show similar and substantial expression of CD107a that suggests sufficient level of activation by both cell lines and a satisfactory NK antitumor cytotoxic mechanisms of MM patients.
Inspite of the similarly induced degree of NK cell cytotoxicity, we show that the decrease in cell viability is more pronounced in K562 than in FemX tumor cell line. This finding is supported by the difference in apoptotic potential of these two tumor cell lines, as shown by the ratio of evaluated proapoptotic Bax to antiapoptotic Bcl-2 genes. Based on this, compared to FemX tumor cell line, K562 tumor cells are expected to be more readily killed by NK cells as they have a greater apoptotic potential (Kornblau et al. 2000). However, inspite of the difference in the susceptibility to apoptosis of the two tumor cell lines, we show that they have similar invasive potential obtained by the evaluation of the expression of Met oncogene.
Furthermore, in this model we also evaluated NK cell immunoregulatory function by measuring intracellular IFNγ expression. We found that, unlike degranulation, significant IFNγ production by NK cells of MM patients is obtained only after contact with K562 tumor cells. Considering that the production of IFNγ by NK cells is a process that requires a much higher stimulation threshold (Romee et al. 2013) compared to NK cell cytotoxic function, the obtained result confirms that K562 tumor cells provide higher activation potential compared to FemX tumor cell line. This may follow from the difference in the expression of HLA class I molecules, as our finding shows that, unlike HLA-null K562, FemX tumor cell line is HLA class I positive and able to engage HLA-specific NK cell inhibitory KIR receptors that by interfering with activating signaling pathways lead to inadequate IFNγ production (Bryceson et al. 2011; Fauriat et al. 2010). Aside from this, high expression of stress-induced MICA/B ligands (Lopez-Larrea et al. 2008) specific for NK cell activating NKG2D receptor may contribute to higher activation potential of K562 tumor cells. Moreover, inadequate NK cell IFNγ production by FemX cell line may also result from matrix metalloproteinase (MMP)-induced shedding of NK cell ligands from FemX cells and shedding of receptors from NK cells that reduces receptor-ligand interaction and decreases the input of activating signals (Tekle et al. 2012; Gonzalez-Gugel et al. 2016). Therefore, our finding of greater production of MMP2 by FemX tumor cell line may support this process.
In order to gain insight into the engagement of signaling pathways relevant for evaluated NK cell functions, we analyzed the level of active phosphorylated form of signal transducers and activators of transcription 4 and 5 (STAT4 and 5), ZAP70, and NFκB molecules in NK cells. In this sense, we show that the level of active phosphorylated STAT4, as a major transcription factor for IFNγ synthesis, after a 4 h contact of NK cells with K562, as well as FemX tumor cell lines shows a tendency of decrease reflecting postactivational dephosphorylation and degradation (Konjevic et al. 2013). Similarly, it was found after contact with both tumor cell lines that the level of active phosphorylated form of STAT5, a transcription factor important in NK cell cytotoxicity as it is linked to the regulation of perforin synthesis (Yu et al. 1999), as well as NKG2D signaling (Sutherland et al. 2002), also shows a tendency of postcontact decrease. Furthermore, we show in our experimental model a tendency of increase of phosphorylated NFκB transcription factor as it is engaged in different NK cell responses during target cell recognition (Zhou et al. 2002; Tato et al. 2006; Kwon et al. 2016).
In this study we also show that a 4 h in vitro contact of NK cells of MM patients with K562 and FemX tumor cell lines leads to a decrease in the percent of CD3−CD16+ NK cells. It has been established that CD16 mediates direct NK cell cytotoxicity of some virus infected and tumor cells and that post contact CD16 loss can be used as a sensitive and specific measure of NK cell cytotoxicity toward target cells (Jewett and Tseng 2011; Warren 2011; Warren et al. 2013). Several mechanisms, aside from the standard postactivational receptor internalization (Jewett and Tseng 2011; Romme et al. 2013), are involved in the observed depletion of CD16 surface receptor, that include shedding through the activation of a transmembrane proteolytic enzyme, metalloproteinase ADAM-17 (Romme et al. 2013). Furthermore, the expression of phosphorylated ZAP-70, a nonreceptor protein tyrosine kinase associated with CD16 also shows a tendency of decrease after NK cell-tumor cell contact reflecting postactivational dephosphorylation and ubiquitin-dependent proteasome-mediated degradation (Paolini et al. 2001). CD16 loss in this model resulted in a decrease in the otherwise prevailing cytotoxic CD3−CD16bright NK cell subset and consequently in an increase in the regulatory CD3−CD16dim subset, although postactivational CD16 depletion following NK cell conjugate formation with tumor cells was previously found even after 30 min (Konjevic et al. 2009b). Moreover, the effect of contact of NK cells with tumor cells in vivo is also reflected in the decrease in cytotoxic CD3−CD16bright NK cell subset in freshly isolated NK cells of MM patients (Konjevic et al. 2009a).
We further analyzed the effect of the two tumor cell lines on the expression of several activating and inhibitory NK cell receptors. We found that NKG2D, one of the main activating NK cell receptors, decreased significantly only after contact with K562 tumor cell line, shown in Fig. 3a, b, as K562 tumor cells highly express MICA/B ligands specific for NKG2D receptors (Ito et al. 2007). This postactivational NKG2D receptor downregulation appears to be an early event evident after 4 h NK cell-tumor cell contact that is in accordance with data of NKG2D decrease reported upon contact with several ovarian tumor cell lines (Jimenez-Perez et al. 2012). In this sense, it has been shown that postcontact NKG2D downregulation is associated with degradation of its intracellular signal transducing adapter, DAP-10 (Gonzalez-Gugel et al. 2016).
It has also been established that tumor cells can display adverse effects on immune cells through the secretion of immunosuppressive soluble mediators, including growth factors TGFβ, VEGF, and GM-CSF (Umansky and Sevko 2012). As TGFβ has been reported to decrease the expression of NKG2D receptors on NK cells, we investigated TGFβ gene expression in the two tumor cell lines. We found this growth factor to be more highly expressed in K562, compared to FemX tumor cells, that may contribute to the shown downmodulation of NKG2D receptor after contact with this tumor cell line. Furthermore, we investigated gene expression of vascular endothelial growth factor A (VEGFA) and its receptor, VEGFRc1 in these tumor cell lines, as VEGFA, aside from angiogenic mediates several immunosuppressive effects, including TGFβ gene transcription. Our findings show that VEGFA and VEGFRc1 genes are more abundantly expressed in K562, and as VEGFA-VEGFRc axis may induce higher TGFβ production in the K562 setting, this can have an adverse effect on NKG2D receptor expression. Moreover, our novel finding showing VEGFRc1 gene expression in lymphocytes of MM patients (unpublished data) supports the recently shown direct suppressive effect of VEGFA on immune cells (Voron et al. 2015).
The results in this study regarding CD161, an NK cells receptor with controversial function (Azzoni et al. 1998; Lanier 1998; Rosen et al. 2005; Aldemir et al. 2005), show unchanged expression after contact with the two tumor cell lines that may result from the shown lack of expression of its ligands on K562 cells (Braud et al. 2007), while, there is no data for CD161 ligand expression on FemX tumor cells. Similarly, that lack of change in the expression of CD158a and CD158b inhibitory KIR receptors after contact with either tumor cell line may follow from the inability of these cell lines to engage these receptors, as K562 cell line is HLA-negative, whereas, FemX tumor cell line, shown in this study to be HLA class I positive, apparently does not express appropriate HLA-C ligands (Trowsdale and Parham 2004). In this sense, the finding in this study of unaltered postcontact expression of CD161 and KIR receptors may exclude their contribution to the evaluated NK cell functions, namely, NK cell degranulation and intracellular IFNγ production.
In this study we performed an evaluation of NK cell functional capacity of investigated MM patients by applying an in vitro model based on NK cell contact with two tumor cell lines K562 and FemX that differ in NK cell sensitivity. We show that contrary to the preserved and similar NK cell antitumor cytotoxicity induced by the both cell lines, NK cell IFNγ production as a function with higher activation requirements is not adequately induced by tumor-specific FemX, compared to NK cell sensitive K562 tumor cell line.
This model gives data of satisfactory NK antitumor activity of MM patients that is unable to be expressed in a tumor-specific setting and that may be achieved by application of appropriate agents that facilitate NK cell-target cell recognition and eliminate tumor-derived immunosuppressive factors. Therefore, the data obtained in this study may be of use for evaluation of the state of disease and for selection of immunotherapy to augment NK cell function of MM patients.
References
Aldemir H, Prod’homme V, Dumaurier MJ, Retiere C, Poupon G, Cazareth J, Bihl F, Braud Vm (2005) Cutting edge: lectin-like transcript 1 is a ligand for the CD161 receptor. J Immunol 175:7791–7795
Azzoni L, Zatsepina O, Abebe B, Bennett IM, Kanakaraj P, Perussia B (1998) Differential transcriptional regulation of CD161 and a novel gene, 197/15a, by IL-2, IL-15, and IL-12 in NK and T cells. J Immunol 161:3493–3500
Bae DS, Hwang YK, Lee JK (2012) Importance of NKG2D-NKG2D ligands interaction for cytolytic activity of natural killer cell. Cell Immunol 276:122–127
Betts MR, Brenchley JM, Price DA, De Rosa SC, Douek DC, Roederer M, Koup RA (2003) Sensitive and viable identification of antigen-specific CD8+ T cells by a flow cytometric assay for degranulation. J Immunol Methods 281:65–78
Braud VM, Josso-Aldemir H, Prod HommeV (2007) Use of LLT1 and/or CD161 for modulating the activity of cells of the immune system EP 1621551 A1 REEL/FRAME:019301/0826 https://www.google.ch/patents/US20090074756
Bryceson YT, Chiang SC, Darmanin S, Fauriat C, Schlums H, Theorell J, Wood SM (2011) Molecular mechanisms of natural killer cell activation. J Innate Immun 3:216–226
de Coaña YP, Choudhury A, Kiessling R (2015) Checkpoint blockade for cancer therapy: revitalizing a suppressed immune system. Trends Mol Med 21:482–491
Farag SS, Caligiuri MA (2006) Human natural killer cell development and biology. Blood Rev 20:123–137
Fauriat C, Long EO, Ljunggren HG, Bryceson YT (2010) Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood 115:2167–2176
Gabrilovich DI, Nagaraj S (2009) Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol 9:162–174
Gonzalez-Gugel E, Saxena M, Bhardwaj N (2016) Modulation of innate immunity in the tumor microenvironment. Cancer Immunol Immunother 65:1261–1268
Ito N, DeMarco RA, Mailliard RB, Han J, Rabinowich H, Kalinski P, Stolz DB, Zeh HJ 3rd, Lotze MT (2007) Cytolytic cells induce HMGB1 release from melanoma cell lines. J Leukoc Biol 81:75–83
Jackson A, Warner N (1986) Preparation, staining and analysis by flow cytometry of peripheral blood leukocytes. In: Rose N, Friedman H, Fahey J (eds) Manual of clinical laboratory immunology, 3rd edn. American Society for Microbiology, Washington, DC, pp 226–235
Jewett A, Tseng HC (2011) Tumor induced inactivation of natural killer cell cytotoxic function; implication in growth, expansion and differentiation of cancer stem cells. J Cancer 2:443–457
Jimenez-Perez MI, Jave-Suarez LF, Ortiz-Lazareno PC, Bravo-Cuellar A, Gonzalez-Ramella O, Aguilar-Lemarroy A, Hernandez-Flores G, Pereira-Suarez AL, Daneri-Navarro A, del Toro-Arreola S (2012) Cervical cancer cell lines expressing NKG2D-ligands are able to down-modulate the NKG2D receptor on NKL cells with functional implications. BMC Immunol 13:7
Klein E, Ben-Bassat H, Neumann H, Ralph P, Zeuthen J, Polliack A, Vánky F (1976) Properties of the K562 cell line, derived from a patient with chronic myeloid leukemia. Int J Cancer 18:421–431
Konjevic G, Radenkovic S, Vuletic A, Mirjacic Martinovic K, Jurisic V, Srdic T (2013) STAT Transcription Factors in Tumor Development and Targeted Therapy of malignancies, in “Oncogene” Editor: Yahawardiah Siregar, Open Access Book, Publisher In Tech, ISBN, 980-953-307-364-9
Konjević G, Mirjacić Martinović K, Vuletić A, Jović V, Jurisić V, Babović N, Spuzić I (2007) Low expression of CD161 and NKG2D activating NK receptor is associated with impaired NK cell cytotoxicity in metastatic melanoma patients. Clin Exp Metastasis 24:1–11
Konjević G, Mirjacić Martinović K, Jurisić V, Babović N, Spuzić I (2009a) Biomarkers of suppressed natural killer (NK) cell function in metastatic melanoma: decreased NKG2D and increased CD158a receptors on CD3−CD16+ NK cells. Biomarkers 14:258–270
Konjević G, Mirjacić Martinović K, Vuletić A, Jurisić V, Spuzić I (2009b) Distribution of several activating and inhibitory receptors on CD3−CD16+ NK cells and their correlation with NK cell function in healthy individuals. J Membr Biol 230:113–123
Kornblau SM, Vu HT, Ruvolo P, Estrov Z, O’Brien S, Cortes J, Kantarjian H, Andreeff M, May WS (2000) BAX and PKCalpha modulate the prognostic impact of BCL2 expression in acute myelogenous leukemia. Clin Cancer Res 6:1401–1409
Kwon HJ, Choi GE, Ryu S, Kwon SJ, Kim SC, Booth C, Nichols KE, Kim HS (2016) Stepwise phosphorylation of p65 promotes NF-κB activation and NK cell responses during target cell recognition. Nat Commun 7:11686. doi:10.1038/ncomms11686
Lanier LL (1998) NK cell receptors. Annu Rev Immunol 16:359
Lanier LL (2003) Natural killer cell receptor signaling. Curr Opin Immunol 15:308–314
Lanier LL (2005) NK cell recognition. Annu Rev Immunol 23:225–274
Lisovsky I, Isitman G, Bruneau J, Bernard NF (2015) Functional analysis of NK cell subsets activated by 721.221 and K562 HLA-null cells. J Leukoc Biol 97:761–767
López-Larrea C, Suárez-Alvarez B, López-Soto A, López-Vázquez A, Gonzalez S (2008) The NKG2D receptor: sensing stressed cells. Trends Mol Med 14:179–189
Monzon JG, Dancey J (2012) Targeted agents for the treatment of metastatic melanoma. Onco Targets Ther 5:31–46
Nagler A, Lanier LL, Cwirla S (1989) Comparative studies of human FcRIII-positive and negative natural killer cells. J Immunol 143:3183–3319
Paolini R, Molfetta R, Piccoli M, Frati L, Santoni A (2001) Ubiquitination and degradation of Syk and ZAP-70 protein tyrosine kinases in human NK cells upon CD16 engagement. Proc Natl Acad Sci USA 98:9611–9616
Romee R, Foley B, Lenvik T, Wang Y, Zhang B, Ankarlo D, Luo X, Cooley S, Verneris M, Walcheck B, Miller J (2013) NK cell CD16 surface expression and function is regulated by a disintegrin and metalloprotease-17 (ADAM17). Blood 121:3599–3608
Rosen DB, Bettadapura J, Alsharifi M, Warren Mathew PA, Lanier HS, Lanier LL (2005) Cutting edge: lectin-like transcript-1 is a ligand for the inhibitory human NKR-P1A receptor. J Immunol 175:7796–7799
Rubio V, Stuge TB, Singh N, Betts MR, Weber JS, Roederer M, Lee PP (2003) Ex vivo identification, isolation and analysis of tumorcytolytic T cells. Nat Med 9:1377–1382
Sibbitt WL Jr, Bankhurst AD, Jumonville AJ, Saiki JH, Saiers JH, Doberneck RC (1984) Defects in natural killer cell activity and interferon response in human lung carcinoma and malignant melanoma. Cancer Res 44:852–856
Strober W (2001) Trypan blue exclusion test of cell viability. Curr Protoc Immunol May; Appendix 3: Appendix 3B. doi: 10.1002/0471142735.ima03bs21
Sutherland CL, Chalupny NJ, Schooley K, VandenBos T, Kubin M, Cosman D (2002) UL16-binding proteins, novel MHC class I-related proteins, bind to NKG2D and activate multiple signaling pathways in primary NK cells. J Immunol 168:671–679
Tato CM, Mason N, Artis D, Shapira S, Caamano JC, Bream JH, Liou HC, Hunter CA (2006) Opposing roles of NF-kappaB family members in the regulation of NK cell proliferation and production of IFN-gamma. Int Immunol 18:505–513
Tekle C, Nygren MK, Chen YW, Dybsjord I, Nesland JM, Maelandsmo GM, Fodstad O (2012) B7-H3 contributes to the metastatic capacity of melanoma cells by modulation of known metastasis-associated genes. Int J Cancer 130:2282–2290
Trowsdale J, Parham P (2004) Mini-review: defense strategies and immunity-related genes. Eur J Immunol 34:7–17
Umansky V, Sevko A (2012) Melanoma-induced immunosuppression and its neutralization. Semin Cancer Biol 22(4):319–326
Voron T, Colussi O, Marcheteau E, Pernot S, Nizard M, Pointet AL, Latreche S, Bergaya S, Benhamouda N, Tanchot C, Stockmann C, Combe P, Berger A, Zinzindohoue F, Yagita H, Tartour E, Taieb J, Terme M (2015) VEGF-A modulates expression of inhibitory checkpoints on CD8+ T cells in tumors. J Exp Med 212:139–148
Warren HS (2011) Target-induced natural killer cell loss as a measure of NK cell responses. J Immunol Methods 370:86–92
Warren HS, Wu F, Horn PL, Pyne DB, West NP, Cripps AW (2013) Peripheral blood natural killer (NK) cell function in healthy adults assessed using the target-induced NK loss (TINKL) assay. J Immunol Methods 392:68–70
Wong JL, Mailliard RB, Moschos SJ, Edington H, Lotze MT, Kirkwood JM, Kalinski P (2011) Helper activity of natural killer cells during the dendritic cell-mediated induction of melanoma-specific cytotoxic T cells. J Immunother 34:270–278
Yu CR, Ortaldo JR, Curiel RE, Young HA, Anderson SK, Gosselin P (1999) Role of a STAT binding site in the regulation of the human perforin promoter. J Immunol 162:2785–2790
Zec M, Srdic-Rajic T, Krivokuca A, Jankovic R, Todorovic T, Andelkovic K, Radulovic S (2014) Novel selenosemicarbazone metal complexes exert anti-tumor effect via alternative, caspase-independent necroptotic cell death. Med Chem 10:759–771
Zhou J, Zhang J, Lichtenheld MG, Meadows GG (2002) A role for NF-kappa B activation in perforin expression of NK cells upon IL-2 receptor signaling. J Immunol 169:1319–1325
Acknowledgements
This study was supported by the grants of the Ministry of Education, Science and Technology of the Republic of Serbia: Grant Numbers 41031 and 175056.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Konjevic, G., Vuletic, A., Mirjacic Martinovic, K. et al. Evaluation of the Functional Capacity of NK Cells of Melanoma Patients in an In Vitro Model of NK Cell Contact with K562 and FemX Tumor Cell Lines. J Membrane Biol 250, 507–516 (2017). https://doi.org/10.1007/s00232-017-9977-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-017-9977-7