Abstract
The objective of this study was to determine which major grape varieties are present in a given wine using both high-performance thin-layer chromatography (HPTLC) fingerprinting and multivariate analysis. For this purpose, 40 mono- and multi-varietal commercial wine samples from four vintages between 2003 and 2012 were collected and analyzed for their polyphenolic composition using HPTLC peak profiles. Polyphenolic compounds such as gallic acid, caffeic acid, resveratrol and rutin (each belonging to one of the four common classes of wine polyphenolic antioxidants) were identified. Unsupervised chemometric method, principal component analysis was used to analyze variance in HPTLC patterns as a function of wine grape variety. An artificial neutral network, as the efficient supervised chemometric tool, was used to develop a predictive model for classification of wine samples and discrimination between them.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Wine characteristics and quality are generally described by aroma, taste, color and flavor and are the result of complex interactions between chemical composition, wine grape variety [1], geographical origin [2], i.e., climate and soil characteristics [3], and vinification conditions such as temperature used during grape maceration, maceration time, frequency, and intensity of maceration. [4]. Wine contains a wide variety of organic and inorganic compounds, with variation in chemical composition occurring due to the natural variability of grape composition and variability in the vinification process [5].
Wine grape variety is an important factor influencing wine flavor, with particular wine grape varieties considered to be an important factor in producing premium wines. Adulteration of these wines using cheaper (and inferior) grape varieties in order to produce a cheaper wine means that the ability to accurately authenticate and differentiate wines is very important in order to protect consumers [6]. Note that under food legislation the inclusion of other wine grape varieties in small, legally defined percentages is permitted (the permitted amount varying between countries). In authenticating a wine, the use of sensory evaluation alone cannot guarantee correct results; hence, it is important to develop more objective methods for characterization of wines based on reproducible physicochemical parameters and standardized control procedures.
A wine can be characterized by its phytochemical fingerprint which can be determined using various qualitative and quantitative analytical methods, especially hyphenated techniques which are usually a combination of chromatographic and spectroscopic methods. High-performance liquid chromatography (HPLC) is often a common method of choice. However, with the development of high-performance adsorbent layers and sophisticated instrumentation for sample application, chromatogram development, derivatization and evaluation, high-performance thin-layer chromatography (HPTLC) has become a powerful analytical method for analyzing complex inorganic and organic matrixes such as those found in wine samples [7]. The main advantages of HPTLC over both gas chromatography (GC) and HPLC are high sample throughput and rapid low-cost analysis. Many samples can be analyzed in parallel on the same plate with better precision and accuracy as a result of simultaneous analysis of both samples and standards under the same conditions with the added benefit of shorter analysis times [8]. Also, careful selection and use of derivatizing agents can enhance selectivity in visualization of target chromatographic zones. Choosing an appropriate scanning wavelength or storing color information resulting from splitting a plate image through red, green and blue channel filters can further enhance selectivity of the method [9]. There is an increasing interest in the application of multivariate statistical methods to differentiate wines according to grape variety, geographical origin and vinification method [10]. In this work, unsupervised pattern recognition technique, principal component analysis (PCA), was used to analyze variance in chromatographic fingerprints as a function of wine grape variety followed by ANN as a supervised technique to develop a predictive chemometric model. The main advantage in using ANNs is the ability to implicitly detect complex nonlinear relationships among variables, which makes it suitable for prediction and pattern recognition applications. ANNs were used to develop a model that can predict grape variety from the wine HPTLC fingerprint. To our knowledge, this is the first report which combines a HPTLC fingerprint with supervised and unsupervised statistical methods.
Materials and methods
Standard solutions and wine samples
Antioxidant standards, caffeic acid (98 %), gallic acid (97 %), resveratrol (99 %) were purchased from Sigma Chemicals (Balcatta WA, Australia), while rutin (97 %) was purchased from Alfa Aesar (Ward Hill, Massachusetts). A gradient elution method was set up using dichloromethane (Merck, Victoria, Australia), methanol (Merck), formic acid (Ajax Chemicals, Sydney, Australia) and sodium dodecyl sulfate (Sigma), butanol (Chem Supply, Gillman, South Australia), Milli-Q water (Millipore, Bayswater, Australia) and heptane (BDH, Tingalpa, Australia). Standard solutions of resveratrol (1 mg/mL) and 0.1 mg/mL standard solutions of caffeic acid, gallic acid and rutin were made in absolute ethanol. A methanolic solution (1 % w/v) of 2-aminoethyl diphenylborinate (Alfa Aesar) was used for plate derivatization. A total of 40 wine samples (38 red wines and 2 white wines) were collected from different regions of Australia (32 wine samples) and overseas (8 wine samples) (Table 1). All samples and standards were refrigerated at 4 °C to prevent degradation [11].
HPTLC analysis of caffeic acid, gallic acid, resveratrol and rutin
HPTLC was performed on HPTLC Silica gel 60 F254 glass plates 20 cm × 10 cm (Merck, Germany). Plates were pre-washed before use with a blank run of ethanol, then dried and activated by heating in an oven at 110 °C for 10 min. Samples were applied to the plates as 10 mm narrow bands using a 100-µL syringe with a semi-automatic sample applicator [Linomat 5 (CAMAG, Muttenz, Switzerland)], 8 mm from the lower edge plate, with 10 mm distance from each side and a track distance of 7 mm. This resulted in 15 sample applications per plate consisting of 10 μL of 11 wine samples, together with 2 μL of caffeic acid, 10 μL of gallic acid, 5 μL of resveratrol and 10 μL of rutin standard solutions. Samples of wine were used without any pre-treatment. Standard solutions were applied in duplicates on either side of the wine samples in order to prevent parallax error through uneven solvent fronts. HPTLC plates were developed in an automated multiple development chamber (AMD2, CAMAG) using a two-step (9:5) gradient elution method. Dichloromethane/methanol/formic acid (73:20:7) was the first mobile phase, and a water-in-oil micro-emulsion consisting of sodium dodecyl sulfate/n-butanol/water/heptane (8 g:25 mL:8 mL:160 mL) was the second mobile phase.
The specific optimized gradient elution separation used in this work was previously reported [12]. The window diagram technique was used to optimize the three components of eluent in the first step for complete separation of caffeic acid and resveratrol (Fig. 1).
Further separation of flavonoids and phenolic acids in wine samples was then achieved by using a micro-emulsion as an eluent in the second step. Micelle eluents containing surfactant as one of the components provide unique separation selectivity, by adding a secondary partitioning equilibrium from the silica gel stationary phase into the micro-emulsion droplets.
Images of plates for both standards and samples were captured using a TLC-Visualizer (CAMAG) with a 12-bit camera (CAMAG) under UV light at 366 nm before and after spraying each plate with 1 % w/v 2-aminoethyl diphenylborinate derivatizing agent solution. The derivatizing reagent reacts with polyphenolic bands on the plate turning them deep blue. Capture parameters (focal length, focus and aperture) were fixed to ensure the quality of images and reproducibility of results between plates. Images were stored in Joint Photographic Experts Group (JPEG) file format [13].
Data acquisition and statistical analysis
Images of the HPTLC chromatograms were exported from winCATS to ImageJ (1.48c version, Wayne Rasband), an image processing program developed at the National Institute of Health, USA (http://rsb.info.nih.gov/ij). ImageJ is a freely available Java-based program for digital picture manipulation that can be used for advanced picture transformations such as filtering, smoothing, background subtraction, auto-balance or grayscale conversion and other signal transformations [14, 15]. It is able to plot the intensity associated with each pixel, as well as export raw data which is particularly useful for further chemometric data handling. ImageJ was designed with open architecture that provides extensibility via Java plugins and recordable macros. The subtracks were cut out (594 pixel length and 60 pixel width for each subtrack) from every track on the plate. The stack of all subtracks was separated into three independent channels for red, green and blue primary color components. The color value of a given point from the chromatographic plate depends on the channel at which it was observed. Images were processed using ImageJ software with raw data exported for further chemometric data handling. The blue channel was chosen because almost all bands are blue and chemometric models obtained for blue channel showed best results. Once the data have been acquired, the usual data pretreatment procedures are denoising, normalization, followed by warping/registering of the chromatogram are analyzed [16, 17]. Normalization of the images was performed by scaling each sample to the sum of intensity. Denoising of the images was done using the 3 pixels median filter. Peak alignment was employed to correct the inter- and intra-plate peak shift due to variations in mobile phase composition, humidity, temperature, operator handling and instrumental instability. The HPTLC chromatograms were warped to the reference by deleting or adding baseline segments near the selected signals using Correlation Optimized Warping (COW), so that the peak R F values were equalized [18]. The data were additionally preprocessed by using mean centering, which is the preferred option when the classification of samples is based on variables that are all measured using the same unit.
Multivariate analysis and data acquisition
Principal component analysis was performed with PLS ToolBox, v.6.2.1, for MATLAB 7.12.0 (R2011a). PCA was carried out as an exploratory data analysis by using a singular value decomposition algorithm (SVD) and a 0.95 confidence level for Q and T 2 Hotelling limits for outliers. Hierarchical cluster analysis (HCA) was obtained using the Ward method to calculate cluster distances and by applying Euclidean distance as a measure of distance between the samples. Artificial neural networks Microsoft Windows™ based simulator software Statistica™ 2009 (StatSoft®, USA) was used to develop a predictive model. Analyzed chromatographic fingerprints provided 594 inputs (peak intensities) for each wine sample. The number of inputs was reduced to 244 by averaging data of consecutive peaks. If a large number of inputs are used to build an ANN, the number of interconnection weights will increase and the training of the neural networks will be extremely slow. Obtained chromatograms were smoothed to reduce the noise and improve signal-to-noise ratio [19] and therefore improve accuracy and precision of a model. Different grape blends provided 19 corresponding categorical outputs (i.e., Cabernet, Cabernet Merlot, Cabernet Sauvignon, Chambourcin Shiraz, Chardonnay, Cienna, Dolcetto, Durif, Merlot, Cabernet Sauvignon, Muscat, Petit Verdot, Pinot Noir, Primitivo, Prunesta Malvasia Nera, Shiraz, Shiraz Cabernet, Shiraz Merlot and Zinfandel). From these data, training (70 % of data set), testing (15 % of data set) and validation sets (15 % of data set) were randomly selected before each training epoch to develop an ANN model.
Results and discussion
The HPTLC profile of a wine sample according to image analysis
HPTLC analysis is generally affected by noise, which is related to the grainy nature of the HPTLC plates, non-uniformity of the plate spraying during derivatization, progressive degradation of spot color, etc. Therefore, an increase in the signal-to-noise ratio could improve analytical results. Investigated wine samples contained characteristic phenolic compounds at R F values of: 0.13 ± 0.02 (rutin), 0.26, 0.48, 0.55 ± 0.0.03 (gallic acid), 0.78 ± 0.02 (caffeic acid) and 0.87 ± 0.02 (resveratrol) (Fig. 2). In this study, it is observed in the chromatographic HPTLC profiles of wine that bands are observed over a large range, with R F values from 0 to 0.9.
Merlot wine contains significantly high peaks at R F values of 0.15, 0.20, 0.26, 0.55 (gallic acid), and 0.78 (caffeic acid). Note that the high gallic acid and caffeic acid content observed in Merlot in this work (Fig. 2) has previously been reported in Merlot Chilean and South American wine [20]. Furthermore, almost all Shiraz wines excluding Shiraz Cabernet wine contain three characteristic peaks at R F = 0.15, 0.55, and 0.78 corresponding to rutin, gallic acid and caffeic acid and a compound with R F value 0.26, with a few samples also showing a resveratrol peak at R F = 0.87 (resveratrol). Durif wine samples had almost a similar pattern as Shiraz wine with four characteristic peaks at R F values of 0.26, 0.55 and 0.78. These phenolic compounds play an important role in wine quality as they significantly contribute to the bitterness, astringency and color of wine [21]. Pinot Noir and Petit Verdot are both observed to have an almost identical chromatographic profile to Shiraz Cabernet wines. Furthermore, one Cabernet wine sample had the same characteristic peaks as Shiraz Cabernet with R F = 0.15, 0.40 and 0.66 and showed a different HPTLC pattern from other Cabernet wines (Fig. 2). The peak profiles for Cabernet, Cabernet Merlot and Cabernet Sauvignon wines show they all contain peaks at 0.15, 0.26, 0.66 and 0.87. When compared to the other wines, Muscat and chardonnay showed a different pattern with only a few other small peaks observed between R F = 0.2 to 0.9 and no resveratrol peak observed. Wine samples for Cienna, the Italian wines (Prunesta, Primitivo) and American wine (Zinfandel) showed similar patterns and were also observed to have significant amounts of rutin and caffeic acid.
Multivariate analysis
While classical HPTLC fingerprinting is done by visual inspection, based on an analyst’s perception, a chemometric approach, based on different statistical models, is able to provide better objectivity. The chromatographic system is optimized to separate and identify all compounds from different varieties present in a wine sample. The HPTLC fingerprint treats the chromatogram as a unique signal, without a need to identify and interpret the peaks. Moreover, it contains a vast amount of information that is able to be used for qualitative and quantitative analysis of analytes present in wine. Both unsupervised and supervised techniques were applied on the data set in order to analyze variance in HPTLC patterns according to grape variety and to develop a model that could predict a grape variety from chromatographic fingerprint.
Principal component analysis
Principal component analysis (PCA), a commonly used multivariate technique, is a variable reduction procedure able to be used to account for most of the variation of total variability in a HPTLC fingerprint. PCA visualizes the structure of data, grouping wine samples according to variety, and identifying important variables (different phenolic compounds/bands) for discrimination between samples [22]. Principal component analysis as an initial multivariate technique was applied on the data matrix (40 samples × 594 variables) obtained from HPTLC plate images.
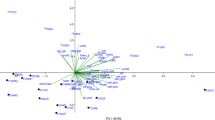
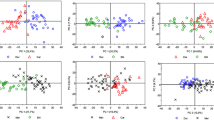
Variables represent the intensities of pixels along the 594 length lines. The Merlot Alpine Valley sample lied outside the Hotelling T 2 ellipse, suggesting that sample was recognized as an outlier and was therefore removed from data set. The first two rotated factors [principal components (PCs)] have the highest eigenvalues and accounted for 45.02 and 13.23 % of the total variability, respectively. The first four principal components describe 73.87 % of total variability. From the mutual projections of PC factor scores (Fig. 3a), all wine samples were clustered into one of three groups; (a) Cabernet, Cabernet Sauvignon and Cabernet Merlot formed one group (upper left hand side), while one Cabernet sample is misclassified and positioned on right side of PCs score close to Shiraz Cabernet samples, (b) Shiraz and Merlot as monovarietal wines formed two subgroups (lower left hand side), and (c) other multi-variety Shiraz wines formed a third group together with Italian and French wines (middle right hand side). One Cabernet sample positioned on right side of PCs score had a similar HPTLC profile as Shiraz Cabernet (Fig. 3). Further, Primitivo (Italian wine), Cienna, and two white wines (Chardonnay and Muscat) formed one subgroup in the central region of the PC plot. Our previous study has shown that Cabernet wine produced in Heathcote, Victoria, has a slightly different HPTLC fingerprint and chemical composition when compared to the other Cabernet wines tested (Table 1) [23].
The loading plots (Fig. 3b, c) demonstrate the significant contribution of polyphenolic compounds to the total variability. The most influential polyphenolic compounds discriminating between Cabernet, multi-varietal Cabernet wines, and Merlot and Shiraz wine, from other wine samples are rutin, compounds with R F values at 0.26, 0.48, 0.66, 0.73, and resveratrol. In contrast to other types of wines and one Cabernet sample, Cabernet, Cabernet Merlot and Cabernet Sauvignon wines contain significant amounts of phenolic compounds with R F values at 0.13 (rutin), 0.26, 0.66 and 0.88 (resveratrol). These phenolic compounds may be identified as characteristic markers for a wine grape variety. Further, wine samples positioned on the left side of PCs score such as Merlot and Shiraz showed a similar pattern and contain polar compounds with R F values 0.15, 0.26, 0.55, 0.78 and may be characteristic markers for these grape varieties. Also, wines which formed a third cluster (Prunesta, Primitivo, Muscat, Chardonnay) contain polar phenolic compounds, with R F at values of 0.09 and 0.13. These compounds may be important markers for wines containing these grape varieties. According to Fig. 3c, bands with R F values of 0.09, 0.27 and 0.73 have a positive correlation with PC2, while rutin (R F = 0.13) and resveratrol (R F = 0.88) and a compound with a R F value of 0.66 showed the most influence in distinguishing between separated Cabernet and multi-varietal Cabernet wines from other wine samples. Wines belonging to the third cluster do not contain compounds with R F value 0.88, while wines which formed a third cluster contain bands with R F values of 0.09, 0.13 and 0.27. Gallic acid and caffeic acid are polyphenolic antioxidants most commonly found in the investigated wines, while rutin and resveratrol were able to be quantified in only selected wine varieties which means they may be useful as characteristics markers of grape variety, which is in agreement with PCA.
An artificial neural network (ANN) is an artificial intelligence tool that mimics the way that human brain works, processes information and learns from data. ANNs are mathematical models composed of a large number of highly inter-connected processing elements (artificial neurons) operating in parallel (layers). Neurons are organized into layers, an input layer, one or more hidden layers, and an output layer (targets). There are different ways in which information can be processed by a neuron and different ways of connecting each neuron to one another. Different neural network structures can be constructed by using different processing elements and by the specific way in which they are connected. Thus, connection weights and the number of hidden neurons in an ANN are adjustable parameters that are optimized during the learning phase.
The ANN learns by adjusting the strength of connection weights and by optimizing the number of hidden layers and hidden neurons during the learning phase. Commonly, ANNs are trained through a specific learning rule so that a particular input leads to a specific target output. Supervised learning is a recursive learning process where inputs fed in the ANN are mapped in the output. The output is then compared with the target, and network weights are adjusted to match the target. The key strength of an ANN is its ability to model both, linear and nonlinear relationships [24], and to estimate any mathematical functions using fewer parameters than other methods [25]. After running 30 tests, each evaluating 250 different ANN topologies, the multilayer perceptron (MLP) with limited memory Broyden–Fletcher–Goldfarb–Shanno (BFGS) optimization algorithm was selected, based on its predictive performance (Table 2) in the validation data set. Broyden–Fletcher–Goldfarb–Shanno, or Quasi-Newton, is a powerful second-order training algorithm with very fast convergence but high memory requirements. More details about the ANNs and learning algorithms can be found in the literature [26].
The best ANN model trained by the Broyden–Fletcher–Goldfarb–Shanno (BFGS) optimization algorithm was a MLP with a 244-26-21 topology. It correctly classified 96 % (25 out of 26) of the wine grape varieties in the training set, 100 % (8 out of 8) in the testing set and 67 % (4 out of 6) in the external validation set. The mistake in the training set was in predicting the Shiraz variety instead of Durif for one of the wine samples. This can be explained by the fact that there was only one Durif wine sample so the ANN did not have enough data to memorize the fingerprint pattern for the Durif variety. Durif, also known as a Petit Shiraz, is named after its discoverer French nurseryman Dr. François Durif. Petit Syrah (Durif) is a black-skinned grape variety, which is the result of a cross between Syrah and a less well-known Rhône variety, Peloursin. It was created to improve the grape’s ability to resist mildew. However, it could not grow in the humid Rhône region and was brought to California [27]. Although Shiraz and Petit Shiraz are two completely different grape varieties, they both make big, rich red wines and both are considered as Rhône varietals [28].
For the validation data set, mistakes were made by predicting Pinot Noir wine grape variety instead of Cabernet, and Shiraz Cabernet instead of Prunesta Malvasia Nera (Black Malvasia). Malvasia Nera is the darker-skinned red wine grape variety that is grown throughout Italy and the Mediterranean and is primarily used as a blending grape in Italy, being valued for the dark color and aromatic qualities it can add to a wine. Cabernet Sauvignon offers bolder and more pronounced expressions of similar characteristics and has largely replaced Malvasia Nera in Tuscan blends [29].
Conclusion
The development of analytical methods for the assessment of authenticity and for detection of adulteration of food and beverages has become very important, especially for expensive products such as premium wine. The HPTLC fingerprint evaluation method that has been developed, combined with digital image processing and consequent chemometric techniques offers a simple and effective way to differentiate wines according to grape varieties. Given that the model is based on limited number of wine samples, model represents a preliminary attempt to distinguish wine categories based on HPTLC fingerprints. Gallic and caffeic acid are phenolic antioxidants most commonly found in wine, while rutin and resveratrol were quantified in only selected wine varieties and therefore are recognized by PCA as potential phenolic markers of wine grape variety. This type of methodology can be used to create a large digital database of fingerprints from different wine varieties and develop a predictive model to characterize wines and wine grape variety from a wine’s HPTLC fingerprint.
References
Armanino C, Casolino MC, Casale M, Forina M (2008) Modelling aroma of three Italian red wines by headspace-mass spectrometry and potential functions. Anal Chim Acta 614(2):134–142
Koundouras S, Marinos V, Gkoulioti A, Kotseridis Y, van Leeuwen C (2006) Influence of vineyard location and vine water status on fruit maturation of nonirrigated cv. Agiorgitiko (Vitis vinifera L.). Effects on wine phenolic and aroma components. J Agric Food Chem 54(14):5077–5086
Sabon I, de Revel G, Kotseridis Y, Bertrand A (2002) Determination of volatile compounds in Grenache wines in relation with different terroirs in the Rhone Valley. J Agric Food Chem 50(22):6341–6345
Esti M, Tamborra P (2006) Influence of winemaking techniques on aroma precursors. Anal Chim Acta 563(1–2):173–179
Hosu AD, Cimpoiu C, Pop N, Miclaus V, Bolboaca SD, Jantschi L (2011) The analysis of different factors affecting the red wines antioxidant content. Not Bot Hort Agrobot Cluj 39(1):159–164
Faria M, Nunes E, Oliveira MP (2008) Relative quantification of Vitis vinifera L. varieties in musts by microsatellite DNA analysis. Eur Food Res Technol 227(3):845–850
Ogegbo OL, Eyob S, Parmar S, Wang Z-T, Annie Bligh SW (2012) Metabolomics of four TCM herbal products: application of HPTLC analysis. Anal Methods 4(8):2522–2527
Morlock G, Schwack W (2008) Planar chromatography-back to the future? LC GC Eur 21(7):366–371
Milojković-Opsenica D, Ristivojević P, Andrić F, Trifković J (2013) Planar chromatographic systems in pattern recognition and fingerprint analysis. Chromatographia 76(19–20):1239–1247
Gonzalez-San Jose ML, Santa Maria G, Diez C (1990) Anthocyanins as parameters for differentiating wines by grape variety, wine-growing region, and wine-making methods. J Food Compos Anal 3:54–66
Spangenberg B, Poole CF, Weins C (2011) Quantitative thin-layer chromatography: a practical survey. Springer, Berlin
Agatonovic-Kustrin S, Morton DW, Yusof AP (2016) Development and validation of a simple high performance thin layer chromatography method combined with direct 1,1-diphenyl-2-picrylhydrazyl assay to quantify free radical scavenging activity in wine. Food Chem 197(Part A):285–290
Hess AVI (2007) Digitally enhanced thin-layer chromatography: an inexpensive, new technique for qualitative and quantitative analysis. J Chem Educ 84(5):842
Zarzycki PK, Ślączka MM, Zarzycka MB, Włodarczyk E, Baran MJ (2012) Application of micro-thin-layer chromatography as a simple fractionation tool for fast screening of raw extracts derived from complex biological, pharmaceutical and environmental samples. Anal Chim Acta 716:54–60
Olech M, Komsta Ł, Nowak R, Cieśla Ł, Waksmundzka-Hajnos M (2012) Investigation of antiradical activity of plant material by thin-layer chromatography with image processing. Food Chem 132(1):549–553
Komsta L (2012) Chemometrics in fingerprinting by means of thin layer chromatography. Chromatogr Res Int 2012:5. doi:10.1155/2012/893246
Daszykowski M, Stanimirova I, Bodzon-Kulakowska A, Silberring J, Lubec G, Walczak B (2007) Start-to-end processing of two-dimensional gel electrophoretic images. J Chromatogr A 1158(1–2):306–317
Wong KH, Razmovski-Naumovski V, Li KM, Li GQ, Chan K (2014) Differentiating Puerariae Lobatae Radix and Puerariae Thomsonii Radix using HPTLC coupled with multivariate classification analyses. J Pharm Biomed Anal 95:11–19
Fuller WA (1996) Introduction to statistical time series. Wiley, New York
Mendoza L, Matsuhiro B, Aguirre MJ, Isaacs M, Sotés G, Cotoras M, Melo R (2011) Characterization of phenolic acids profile from chilean red wines by high-performance liquid chromatography. J Chil Chem Soc 56(2):688–691
Rossouw M, Marais J (2004) The phenolic composition of South African Pinotage, Shiraz and Cabernet Sauvignon wines. S Afr J Enol Vitic 25(2):94–104
Ristivojević P, Andrić FL, Trifković JĐ, Vovk I, Stanisavljević LŽ, Tešić ŽL, Milojković-Opsenica DM (2014) Pattern recognition methods and multivariate image analysis in HPTLC fingerprinting of propolis extracts. J Chemom 28(4):301–310
Agatonovic-Kustrin S, Hettiarachchi CG, Morton DW, Razic S (2015) Analysis of phenolics in wine by high performance thin-layer chromatography with gradient elution and high resolution plate imaging. J Pharm Biomed Anal 102:93–99
Wikel JH, Dow ER (1993) The use of neural networks for variable selection in QSAR. Bioorg Med Chem Lett 3(4):645–651
Duprat AF, Huynh T, Dreyfus G (1998) Toward a principled methodology for neural network design and performance evaluation in QSAR. Application to the prediction of LogP. J Chem Inf Comput Sci 38(4):586–594
Haykin S (1999) Neural networks: a comprehensive foundation. Prentice-Hall, Englewood
Prial FJ (1997) Wine talk. The New York Times, New York
Jackson RS (2014) Wine science: principles and applications, 4th edn. Elsevier, Burlington
Robinson J (2006) The Oxford companion to wine. Oxford Univeristy Press, Oxford
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare there is no conflict of interest.
Compliance with Ethics Requirements
This article does not contain any studies with human or animal subjects.
Rights and permissions
About this article
Cite this article
Agatonovic-Kustrin, S., Milojković-Opsenica, D., Morton, D.W. et al. Chemometric characterization of wines according to their HPTLC fingerprints. Eur Food Res Technol 243, 659–667 (2017). https://doi.org/10.1007/s00217-016-2779-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-016-2779-9