Abstract
Erhuangquzhi granules (EQG) have been clinically proven to be effective in nonalcoholic steatohepatitis (NASH) treatment. However, the active components and molecular mechanisms remain unknown. This study aimed to screen active components targeting tumor necrosis factor α (TNF-α) in EQG for the treatment of NASH by a surface plasmon resonance (SPR) biosensor–based active ingredient recognition system (SPR–AIRS). The amine-coupling method was used to immobilize recombinant TNF-α protein on an SPR chip, the specificity of the TNF-α-immobilized chip was validated, and nine medicinal herbs in EQG were prescreened. Nuciferine (NF), lirinidine (ID), and O-nornuciferine (NNF) from lotus leaves were found and identified as TNF-α ligands by UPLC‒MS/MS, and the affinity constants of NF, ID, and NNF to TNF-α were determined by SPR experiments (Kd = 61.19, 31.02, and 20.71 µM, respectively). NF, ID, and NNF inhibited TNF-α-induced apoptosis in L929 cells, the levels of secreted IL-6 and IL-1β were reduced, and the phosphorylation of IKKβ and IκB was inhibited in lipopolysaccharide (LPS)-stimulated RAW264.7 cells. In conclusion, a class of new active small-molecule TNF-α inhibitors was discovered, which also provides a valuable reference for the material basis and mechanism of EQG action in NASH treatment.
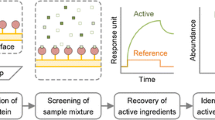
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nonalcoholic steatohepatitis (NASH) is the accumulation of fat accompanied by hepatic steatosis, inflammatory infiltrate, and cellular injury, with or without hepatic fibrosis [1]. The prevalence of NASH in the general population could be as high as 12% [2]. Many liver-related deaths are related to NASH, as the ultimate progression of NASH is cirrhosis and liver cancer, constituting the third leading cause of cancer-related death worldwide. There is no definitive regimen for NASH treatment; although liver transplantation is effective, there is a shortage of appropriate donors [3]. Therefore, novel pharmacotherapy for the treatment of NASH is urgently needed. Tumor necrosis factor α (TNF-α) is considered a potential drug therapeutic target for NASH therapy because NASH pathogenesis is associated with many proinflammatory cytokines. TNF-α, a secreted protein, plays an essential role among the proinflammatory cytokines in NASH progression [4, 5]. Anti-TNF-α monoclonal antibodies are widely used to treat inflammatory diseases in the clinic and have been repurposed for preclinical studies in the treatment of NASH. For example, Schramm et al. conducted a study showing that adalimumab, a humanized TNF-α antibody, could improve liver function and ameliorate NASH symptoms [6]. Anti-TNF-α antibodies have also shown therapeutic effects. However, TNF-α antibodies have limitations, such as serious side effects, high cost, and poor stability, which have prompted the need for small-molecule inhibitors that target TNF-α [7]. The small-molecule anti-TNF-α agent pentoxifylline is considered an effective drug for the treatment of NASH [8], and patients with NASH treated with pentoxifylline have reduced mortality and improved hepatorenal syndrome [9]. However, pentoxifylline can cause symptoms in the digestive system [10]. Thus, lower toxicity and effective TNF-α inhibitors are urgently needed for the treatment of NASH.
Traditional herbal medicines are not only used as complementary and alternative therapies for diseases [11] but also considered a component library for lead compound discovery. Erhuangquzhi granules (EQG) derived from Shengjiang Powder (SJP) have a history of more than 230 years and have lipid-lowering and anti-inflammatory functions. Thus, EQG is clinically used to treat patients with NASH [12]. Although EQG has been clinically proven to be effective in NASH treatment, its pharmacologically active components and molecular mechanisms remain unknown. Only a portion of the components have pharmacological activity in herbal extracts, but screening the active components from the EQG is a time-consuming and complex process. The commonly used strategy for active component identification begins with the separation of all components from traditional herbal medicine, and active ingredient analysis is performed individually. However, inactive components will waste many reagents and time, which is a drawback of this strategy.
There are many methods to screen the active components of traditional herbal medicines. Some approaches are based on the specific binding of the target protein to the active components, such as ultrafiltration (UF) LC/MS [13, 14] and microscale thermophoresis (MST) [15] approaches. UF LC/MS is used to determine the small molecules that bind to the target protein by incubating the target protein together with the traditional herbal medicine extract, intercepting the small molecules that bind to the protein through an ultrafiltration tube, and comparing the small molecules intercepted by denatured histone proteins versus normal proteins. MST technology requires fluorescent labeling of target proteins and changes in the fluorescence of labeled proteins caused by thermal surges caused by protein interactions with small molecules to analyze the affinities of small molecules of drugs with target proteins according to the changes in fluorescence [16, 17]. However, the above methods have limitations. For example, UF consumes a large amount of protein, and it is impossible to determine the affinity information of the compound and the target protein to sort the compound according to the affinity value. MST requires the fluorescent labeling of target proteins, which makes MST experiments more expensive and longer-term. Hence, a more effective method than traditional screening methods should be introduced.
Surface plasmon resonance (SPR) is a label-free method for detecting biomolecular interactions in real time [18]. SPR has been widely used in biomolecular interactions, including enzyme–substrate interactions [19], protein‒protein interactions [20], and drug–target interactions [21]. The significant advantages of SPR in detecting proteins and small-molecule interactions have been recognized [22, 23]. In terms of active compound identification from medicinal herbal extracts, the advantage of SPR in the screening of active ingredients is that active components can be selected directly from medicinal herbal extracts [24], which reduces the purification and separation steps. Researchers have used SPR and ultra-performance liquid chromatography–tandem mass spectrometry in combination (SPR-UPLC‒MS/MS) to screen active components from medicinal herbal extracts. Chen et al. applied this method to screen signal transducer and activator of transcription 3 (STAT3) ligands, and features of the method were investigated, including the specificity, linearity, limit of detection, saturability, and robustness. Five compounds could bind to the STAT3 SH2 domain and inhibit STAT3-driven transcriptional activity. Among them, polydatin and neobaicalein were first reported to be STAT3 ligands [25]. Fu et al. demonstrated that avenanthramides (AVNs) significantly reduced DDX3 expression and used the SPR-UPLC‒MS/MS method to identify avenanthramide A as specifically binding to the oncogenic DDX3 protein. Avenanthramide A is an active component in AVNs and selectively kills tumor cells that highly express DDX3 [26]. Hence, the purpose of the study was to screen active components targeting TNF-α in EQG by SPR–AIRS.
In this study, SPR–AIRS was applied to screen direct small-molecule TNF-α inhibitors from EQG. The experimental workflow is illustrated schematically in Fig. 1. Recombinant TNF-α protein was immobilized on an SPR chip surface by the amine-coupling method, and the activity and specificity of the TNF-α-immobilized chip were validated by the direct small-molecule TNF-α inhibitor SPD304. Nine medicinal herbal extracts were used for prescreening, and lotus leaf extract had a high response to the TNF-α-immobilized chip. Lotus leaf extract was injected into the TNF-α-immobilized chip surface for ligand fishing, and recovery samples were analyzed by a UPLC-Q Exactive HF-X mass spectrometer (UPLC-QE HF-X MS) to confirm their structures. Nuciferine (NF), lirinidine (ID), and O-nornuciferine (NNF) from lotus leaves were found and identified, and their affinities were validated by SPR assay and molecular docking, which could be used as TNF-α ligands. NF and NNF significantly neutralized TNF-α-induced cytotoxicity in L929 cells. Due to its cytotoxicity, ID did not exert a good neutralizing TNF-α-induced cytotoxicity effect. Enzyme-linked immunosorbent assay (ELISA) experiments demonstrated that NF, ID, and NNF could significantly induce IL-6 and IL-1β protein levels in LPS-induced RAW264.7 macrophages. Western blotting (WB) revealed that NF, ID, and NNF could inhibit the phosphorylation of IκBα and IKKβ. These results validated that NF, ID, and NNF can be used as potential inhibitors of TNF-α. Among them, NF and NNF have low toxicity in RAW264.7 cells and good pharmacological effects. ID also has good pharmacological effects but is highly toxic. These results demonstrate that EQG contains inhibitors of TNF-α, while NF, ID, and NNF may be partially active components in the treatment of NASH by EQG.
Workflow for identifying potential TNF-α inhibitors from 9 medicinal herbs of EQG by SPR-UPLC‒MS/MS. A Nine medicinal herbs of EQG were extracted. B TNF-α protein was immobilized to the surface of the chip, and TNF-α ligands were screened from 9 medicinal herbs of EQG. C The results were validated by SPR and molecular docking. D The pharmacological activities of NF, ID, and NNF were validated by TNF-α-induced cytotoxicity in L929 cells and anti-inflammatory in LPS-induced RAW264.7 cells
Materials and methods
Chemicals and reagents
CM5 chips, an amine coupling kit, and PBS were purchased from Cytiva (Shanghai, China). TNF-α protein was from Sino Biological (Beijing, China). The purities of NF, ID, and NNF were over 98%, and these standard compounds were purchased from ChemFaces (Wuhan, China). SPD304 and actinomycin D were from MCE (Medchem Express, USA). ELISA kits were from Westang Biotechnology (Shanghai, China). Phospho-IKKα/β (Ser176/180) Antibody II, IKKβ (D30C6) Rabbit mAb, IκBα (L35A5) Mouse mAb, and Phospho-IκBα (Ser32) (14D4) Rabbit mAb were from Cell Signal Technology Inc. (Beverly, MA, USA). The L929 cell lines were purchased from Boster Bioengineering Company (Wuhan, China). The RAW264.7 cell lines were obtained from the cell bank of Shanghai Institutes for Biological Sciences. Fetal bovine serum (FBS) was produced by Biological Industries (Grand Island, NY, USA).
Nine medicinal herbs of EQG include Turmeric (Curcuma longa L.); Rhubarb (Rheum palmatum L.); Herba Gynostemmatis Pentaphylli (Gynostemma pentaphyllum (Thunb.) Mak.); Rhizoma Atractylodis Macrocephalae (Atractylodes macrocephala Koidz.); Lotus Leaf (Nelumbo nucifera Gaertn.); Oriental Waterplantain Rhizome (Alisma plantago-aquatica Linnaeus); Giant Knotweed Rhizome (Polygoni Cuspidati Rhizoma et Radix.); Radix Glycyrrhizae (Glycyrrhiza uralensis Fisch.); and Lobed Kudzuvine Root (Pueraria montana (Willd.) Ohwi) purchased from the First Affiliated Hospital of Shihezi University School of Medicine in Xinjiang Uygur Autonomous Region of China in December 2021. They were identified by Professor Bo Han. Voucher specimens (No. 20211220) were preserved in the Key Laboratory of Xinjiang Phytomedicine Resource and Utilization (School of Pharmacy, Shihezi University). Nine medicinal herbs of EQG were used for this work.
Recombinant TNF-α protein immobilized on the chip surface
A Biacore T200 instrument (GE Healthcare, Sweden) was used to perform SPR experiments. Flow cell 1 was the reference channel, and other cells were the detection channels. The CM5 chip with a carboxymethylated dextran–coated surface was activated by a 1:1 mix of N-hydroxysuccinimide (NHS) and 1-ethyl-3-[3-dimethylaminopropyl] carbodiimide HCl (EDC). Recombinant TNF-α protein was diluted in 10 mM sodium acetate (pH 5.0, 50 µg/mL), the ligand was attracted and covalently coupled to the dextran matrix, and the contact time was 900 s with a flow rate of 10 μL/min. Ethanolamine was used to deactivate and further wash away loosely associated ligand.
TNF-α-immobilized chip specificity detection
SPD304 (10 µM) and INT747 (10 µM) sample solutions were injected into the TNF-α-immobilized chip surface to determine their response values to TNF-α protein, while running buffer was used as a blank control. The contact and dissociation times were 60 s and 180 s, respectively, and the flow rate was 30 µL/min. Subsequently, the affinity with TNF-α protein was tested over a concentration range of 0.5 to 32 µM. The steady-state affinity model (1:1) was used to perform affinity fitting to obtain the affinity constant (Kd) by Biacore T200 evaluation software.
Preparation of herbal extracts and prescreening
The nine kinds of medicinal herbs were crushed, and the powder was passed through a 50-mesh sieve and retained. The powdered medicinal herbs (1.00 g each) were accurately weighed and subjected to ultrasound extraction with 10 mL 80% methanol/water (v/v) for 40 min. Subsequently, the herbal extracts were centrifuged at 8000 × g for 5 min and were stored at 4 °C before analysis.
The nine herbal extracts were each diluted in PBS (1 µg/mL) and filtered through a 0.22-µm filter. The samples were flowed over the TNF-α-immobilized chip, and the contact and dissociation times were 60 s and 180 s, respectively. PBS was injected between different samples for the SPR system cleaning.
Recovery of TNF-α-bound active components
The TNF-α-bound active components were recovered through the Biacore T200 system. The process of the recovery experiment was divided into four parts: (1) Herbal extracts were flowed over the surface of the TNF-α-immobilized chip, and active components were bound to TNF-α protein. (2) The system washed away the residual sample solution. (3) The recovery solution was incubated on the surface of the TNF-α-immobilized chip, and the active ingredients dissociated from the TNF-α protein. Finally, (4) the flow direction of the TNF-α-immobilized chip surface was reversed, and the recovery solution containing active components was mixed with the deposition solution. Herbal extracts were flowed over the TNF-α-immobilized chip surface for 180 s, and the flow rate was maintained at 5 µL/min. The chip surface was cleaned with distilled water. Formic acid (0.5%) in water was used as the recovery solution and incubated for 20 s. A recovery solution containing active components was deposited in 10 µL ammonium bicarbonate (50 mM). This process was repeated 5 times as a cycle, and lotus leaf extract was recovered for 15 cycles. Recovered samples were dried with nitrogen gas and dissolved in 100 µL methanol. The samples were centrifuged at 8000 × g for 5 min and filtered through a 0.22-µm filter. The supernatants were taken and transferred into glass vials with micro inserts.
UPLC-QE HF-X MS analyses of active components
SPR-recovered samples and lotus leaf extract were analyzed on a Thermo Vanquish UPLC system (Thermo Fisher Scientific, USA). Chromatographic separation was carried out on a Waters ACQUITY UPLC®HSS T3 (Ф 2.1 × 100 mm, 1.8 μm) at 40 °C. Each sample was measured with different gradients of mobile phases of 0.1% aqueous formic acid (v/v) (A) and acetonitrile (B), and the injection volume was 5 µL. The elution procedure was 0% B at 1 min, 0 ~ 5% B at 1 ~ 5 min, 5 ~ 100% B at 5 ~ 20 min, 100% B at 20 ~ 25 min, and 100 ~ 0% B at 25 ~ 30 min. The flow rate was a constant speed of 0.3 mL/min. Samples were selected for MS/MS using 25/35/45% normalized collision energies (NCE), data-dependent MS/MS Top 20. The MS scans used positive ion mode, and the m/z scan range was 100 to 1000. All the data were processed by Thermo Scientific FreeStyle™ 1.7.
Affinity validation
The active component standards were dissolved in PBS with 5% DMSO and diluted from a 256-µM gradient to 1 µM. The samples were injected through a TNF-α-immobilized chip surface at a flow rate of 30 µL/min. The contact and dissociation times were 60 s and 120 s, respectively. The steady-state affinity model (1:1) was used to perform the affinity curve fitting to obtain the Kd by Biacore T200 evaluation software.
Molecular docking
Docking analysis was performed between standard compounds and TNF-α by AutoDock Vina. The crystal structure of TNF-α was retrieved from the Protein Data Bank (PDB: 2AZ5) [27]. To prepare for docking, hydrogen atoms were added, while water molecules and co-crystallized ligand were removed from the TNF-α crystal structure. Standard compound structures were retrieved from PubChem. The semiempirical MM method was applied to optimize the structures of small molecules in Chem Office Ultra 14.0 (Cambridge software). The docking site of the original ligand (x = − 8.33388, y = 68.21730, z = 19.96260) was selected as the center of the binding pocket with a 10-Å radius. The optimal posture is evaluated by calculating the binding energy. The docking results were visualized in PyMol.
Inhibition of TNF-α-induced cytotoxicity
The inhibition rate of TNF-α-induced cytotoxicity was determined in the L929 cell line after standard compound therapy. L929 cells were resuspended in RPMI-1640 complete medium containing 10% FBS at a density of 104 cells/mL and subsequently seeded in a 96-well plate for 24 h. After 24 h of incubation, the complete medium was discarded and replaced with 100 µL of RPMI-1640 incomplete medium containing different concentrations of standard compounds with 1 µg/mL actinomycin D and 2 ng/mL recombinant human TNF-α. The total volume of the mixture was 100 µL, and it was incubated with the L929 cells for 24 h. Cell morphological changes were observed under an inverted microscope, and images were taken, while the cell survival rate was measured using the CCK8 method.
Cell viability detection
RAW264.7 cells were cultured in DMEM containing 10% FBS at 37 °C with 5% CO2. RAW264.7 cells were trypsinized and resuspended at a density of 1.5 × 105 cells/mL and subsequently seeded in a 96-well plate for 24 h. At the end of the incubation time, the medium was replaced with DMEM containing different concentrations of standard compounds (10, 20, 40, 60, 80, and 100 µM). The viability of RAW264.7 cells was measured after 24 h by adding CCK8 solution and incubating for 2 h. The absorbance was determined at 450 nm with a Model 680 Microplate Reader (Multiskan FC, Thermo).
Detection of cell supernatant cytokine levels
RAW264.7 cells (106 cells/mL) were treated with NF, ID, and NNF (10, 20, 40 µM) for 3 h in a 12-well plate and then incubated with LPS (1 μg/mL) for another 12 h. Next, the cell supernatant was collected and centrifuged at 1000 × g for 20 min. ELISA was used to detect the levels of IL-6 and IL-1β in the medium, and the absorbance was determined at 450 nm with a microplate reader.
Western blot
RAW264.7 cells (5 × 105 cells/mL) were cultured overnight. Then, the medium was discarded and replaced with DMEM containing different concentrations of standard compounds (10, 20, and 40 µM). LPS (1 µg/mL) was added after 12 h and incubated for another 2 h. The cellular proteins were extracted with RIPA lysis buffer (Solarbio, China) and centrifuged to remove insoluble matter. Protein concentrations were measured using bicinchoninic acid (BCA) experiments (Solarbio, China). Equal amounts of protein were separated by 10% SDS‒PAGE and transferred to PVDF membranes. After blocking with 2% BSA powder-TBST for 90 min, the membranes were incubated with primary antibodies against IκB-α (1:1000), p-IκB-α (1:1000), IKKβ (1:1000), p-IKKβ (1:1000), and GAPDH (1:1000). The protein band was incubated for 1 h with a goat anti-rabbit or anti-mouse IgG secondary antibody (1:10,000). Enhanced chemiluminescence (ECL) reagents were used to detect protein bands.
Statistical analysis
The results are presented as the means ± standard deviations (SD) and were analyzed using SPSS software version 16.0. One-way analysis of variance (ANOVA) was used to determine significant differences between groups, and P < 0.05 was considered a statistically significant difference.
Results and discussion
Protein precoupling and immobilization
To determine the optimal equivalent point of the protein, more recombinant TNF-α proteins can be immobilized. Sodium acetate at different pH values (4.0, 4.5, 5.0, and 5.5) was selected as the dilution buffer for TNF-α protein in the precoupling assay. Recombinant TNF-α protein was diluted in sodium acetate buffers with different pH values to a concentration of 50 µg/mL in each sample and analyzed by the Biacore T200 system. The results showed that a high immobilization level was achieved at pH 5.0 (Fig. 2A). Therefore, sodium acetate at pH 5.0 was used for protein immobilization. Recombinant TNF-α protein was immobilized on three detection channels of the CM5 chip with immobilization levels of 21,500.3, 23,127.8, and 22,841.7 response units (RU), respectively (Fig. 2B).
Immobilization of TNF-α proteins and screening of active ingredients. A Response of TNF-α protein diluted in sodium acetate buffers with different pH values. B Sensorgrams showing the immobilization of TNF-α proteins via amine coupling at pH 5.0 in flow cell 2. C The specificity of the TNF-α-immobilized chip was validated. D Sensorgrams of SPD304 and fitting curves are shown. E The prescreening response signals of 4 medicinal herbs. F Schematic diagram of the active ingredient recovery process
Specificity of the TNF-α-immobilized chip
A direct inhibitor and a negative control compound were used to verify the specificity of the TNF-α-immobilized chip. SPD304 was selected a direct TNF-α inhibitor as a positive control, and INT-747, a direct ligand for farnesoid X receptor [28], was selected as a negative control. Standard solutions of different concentrations of SPD304 and INT-747 were flowed through the surface of the TNF-α-immobilized chip. The blank with the addition of PBS was used as a control. SPD304 produced a high response signal with TNF-α protein, while INT-747 responded to a signal of approximately 0 RU, which is close to the signal produced by PBS (Fig. 2C). This result demonstrates the specificity of the TNF-α-immobilized chip. In addition, SPD304 could bind to the surface of the TNF-α-immobilized sensor chip in a concentration-dependent manner, and the Kd of SPD304 was 11.09 µM (Fig. 2D).
Screening of active components from herbal extracts
The SPR system was used to screen TNF-α ligands from EQG. Nine medicinal herbs of EQG were prescreened. The herb extraction samples (10 µg/mL) were injected into the surface of the TNF-α-immobilized chip. The prescreening results are shown in Fig. 2E. Among the 4 herb extraction samples, the response signal of lotus leaf was the highest, at 55.20 RU. The prescreening results for 5 other herbal extracts are shown in Electronic supplementary material Fig. S1.
According to the prescreening results, the lotus leaf was selected for further screening by combining SPR and UPLC-QE HF-X MS. An injection and recovery program was applied to recover the TNF-α-bound components in lotus leaf; the recovery process is shown (Fig. 2F). First, due to the specificity of the TNF-α-immobilized chip, components that could interact with TNF-α bound to TNF-α remained on the chip, while other components that did not bind to TNF-α eluted from the TNF-α-immobilized chip surface. Second, the wash solution was flowed over the TNF-α-immobilized chip surface to wash off components not bound to TNF-α protein. Third, the recovery solution was used for incubation and dissociation of the active ingredients. Finally, the flow direction of the recovery solution changed and mixed with the deposition solution to neutralize excess formic acid.
Identification of TNF-α-bound components
The recovered samples were analyzed by UPLC-QE HF-X MS. The total ion chromatogram (TIC) of the lotus leaf extract is shown in Fig. 3A. Ion signals were identified in the lotus leaf extract sample (Rt = 12.00 min, m/z = 296.1641 [M + H]+) and the lotus leaf recovery sample (Rt = 12.19 min, m/z = 296.1642 [M + H]+) (Fig. 3B, C). The structure was identified by matching precise mass numbers and secondary mass spectrometry. The ion m/z 296.1641 of the lotus leaf extract sample produced fragment ions at m/z 265.1219, 250.0984, and 234.1036 (Fig. 3E), while the ion m/z 296.1642 of the lotus leaf recovery sample produced the same fragment ions (Fig. 3F). According to fragment characteristics (Fig. 3G), this compound was speculated to be NF. Subsequently, the NF standard solution was injected into the UPLC-QE HF-X MS system under the same conditions. The Rt of NF was 12.33 min, and the m/z was 296.1641 ([M + H]+) (Fig. 3D), which was the same as the ion signal in the lotus leaf extract and recovery samples. Therefore, NF was identified as a candidate TNF-α ligand from lotus leaf extract. The secondary mass spectrometry of NF is shown in Electronic supplementary material Fig. S2. In addition, ID and NNF were identified as candidate compounds from lotus leaf extract in the same way. The Rt and m/z of NF, ID, and NNF in lotus leaf extract, recovery samples, and standard samples are shown in Table 1. The ion chromatogram and secondary mass spectrometry of ID and NNF are shown in Electronic supplementary material Fig. S3.
Identification of active ingredients. A TIC of lotus leaf extract. EIC of NF in lotus leaf extract samples (B), lotus leaf recovery samples (C), and NF standard samples (D). The secondary mass spectrum of NF in lotus leaf extract samples (E) and lotus leaf recovery samples (F). Probable fragmentation pathways of NF (G)
Validation of NF, ID, and NNF binding to TNF-α
To validate the interactions between TNF-α and NF, ID, and NNF, an SPR affinity assay was performed. The sensorgrams and fitting curve of NF are shown in Fig. 4A, B. The results of ID and NNF are shown in Fig. 4C, D and Fig. 4E, F, respectively. According to the SPR assay results, NF, ID, and NNF had good affinities with TNF-α protein (Kd = 61.19, 31.02, and 20.71 µM, respectively).
To further validate that NF, ID, and NNF can bind to TNF-α, molecular docking was applied to evaluate the interactions. The X-ray cocrystal structure of a TNF-α dimer with SPD304 (PDB code: 2AZ5) was applied as the model to validate NF, ID, and NNF, which bind to TNF-α. The conformation with the most favorable orientation was the most stable for further structural analysis. The structures of the complex between SPD304, NF, ID, or NNF and the TNF-α dimer are shown in Fig. 5. According to the results, the binding energies of NF, ID, and NNF were negative, which suggested that NF, ID, and NNF can bind to TNF-α. Thus, NF, ID, and NNF were submitted to further biological evaluations.
Inhibition of TNF-α-induced cytotoxicity in L929 cells
Cell apoptosis plays an important role in chronic inflammation, is associated with inflammatory destruction, and affects tissue repair. TNF-α has been shown to induce L929 cell apoptosis [29]. The inhibitory effects of NF, ID, and NNF on TNF-α-induced L929 cell cytotoxicity were detected by the CCK-8 method. Actinomycin D is an autophagic activator [30] that can increase TNF-α-induced L929 cell cytotoxicity. As shown in Fig. 6B, C, actinomycin D and TNF-α inhibited the viability of L929 cells; cell viability was only 2.99 ± 0.18%, but with treatment with NF and NNF, the cell survival rate improved significantly. Due to its cytotoxicity, ID did not exert a good neutralizing TNF-α-induced cytotoxicity effect at 40 µM. These results indicate that NF, ID, and NNF can neutralize the cytotoxicity of TNF-induced L929 cells and suggest that the SPR–AIRS method specifically recognizes NF, ID, and NNF as TNF-α-binding ingredients.
NF, ID, and NNF can inhibit TNF-α-induced cytotoxicity in L929 cells. A Chemical structures of NF, ID, and NNF. B Inhibition of TNF-α-induced cytotoxicity in L929 cells. The inhibition rate was determined after treatment with different concentrations of NF, ID, and NNF (10, 20, and 40 µM). Survival rates were determined as the average ± SD. C Morphology of L929 cells after treatment with actinomycin D, actinomycin D with TNF-α, actinomycin D, and TNF-α with 10, 20, or 40 µM NF, ID, and NNF
Effects of NF, ID, and NNF on cell viability
To determine whether NF, ID, and NNF had toxic effects on RAW264.7 macrophages, CCK-8 experiments were applied to cell viability assays. After treatment with different concentrations (0, 10, 20, 40, 60, 80, and 100 μM) of NF, ID, and NNF, the results showed that treatment with 100 μM NF (Fig. 7A) and 100 μM NNF (Fig. 7C) had no obvious effect on cell viability. However, after treatment with ID, cell viability was significantly reduced (Fig. 7B). The cell survival rate was approximately 80% after treatment with ID at 40 μM. Thus, 10, 20, and 40 μM NF, ID, and NNF were applied in the next experiments.
NF, ID, and NNF inhibit the production of inflammatory mediators mediated by LPS. Cell viabilities of NF (A), ID (B), and NNF (C) on RAW264.7 cells at 24 h. The levels of IL-6 and IL-1β were assessed by ELISA (D, E). RAW264.7 cells were incubated with the indicated concentrations of NF (F, G), ID (H, I), and NNF (J, K), and the phosphorylation levels of IKKβ and IκB were detected by western blotting. Data are expressed as the means ± SDs, #P < 0.05, ##P < 0.01 compared with the normal group; *P < 0.05, **P < 0.01 compared with the model group. (n = 3 in each group)
NF, ID, and NNF block TNF-α-mediated signaling pathways
The TNF-α-mediated inflammatory process is regulated by several sequences in gene promoters [31]. The TNF-α-mediated classical nuclear factor kappa-B (NF-κB) pathway has received much attention. The effects of NF, ID, and NNF on the activation of the NF-κB pathway in RAW264.7 cells were investigated. The phosphorylation levels of IKKβ and IκB were enhanced by LPS compared to the control group. Encouragingly, IKKβ and IκB phosphorylation levels showed dose-dependent decreases after NF pretreatment (Fig. 7F, G). The same effect can be seen in the presence of ID (Fig. 7H, I) and NNF (Fig. 7J, K). NF, ID, and NNF exert an in vitro anti-inflammatory effect by inhibiting the TNF-α pathway.
Discussion
SPR–AIRS for the discovery of TNF-α ligand active components in EQG was established in this study. Compounds of the same parent nucleus (NF, ID, and NNF) were discovered as TNF-α ligands from complex extracts. NF, ID, and NNF as TNF-α ligands were verified by SPR affinity analysis and molecular docking, and cell assays demonstrated that they inhibited TNF-α-induced apoptosis in L929 cells and reduced LPS-induced RAW264.7 cellular inflammation levels. The experimental results demonstrate that SPR–AIRS could be an effective method to analyze the active components of traditional herbal medicines.
Interestingly, NF, ID, and NNF have similar chemical structures and all belong to the category of aporphine alkaloid [32, 33]. The chemical structures are shown in Fig. 6A. The structure of NF contains methyl groups at the positions of R1 and R2, that of ID contains a methyl at the R1 position and an H atom at the R2 position, and that of NNF contains an H atom at the R1 position and a methyl at the R2 position. NF and NNF did not exhibit obvious cytotoxicity at 100 μM, while ID showed obvious cytotoxicity from 40 to 100 μM. In terms of structure, NF and NNF are not cytotoxic to the cell, and the structures of NF and NNF contain methyl groups at the R2 position. ID showed cytotoxicity, and the structure of ID contains an H at the R2 position. In TNF-α-induced L929 cell cytotoxicity experiments, NNF exhibited superior activity over NF because, structurally, NF has a methyl at R1 and NNF has an H at R1. In simple terms, if NF is used as the structural parent nucleus, the methyl group at R1 is replaced with an H, and its activity is enhanced, while the methyl group at R2 is replaced by an H, and its cytotoxicity increases. In addition, NNF exhibited inhibition of TNF-α-induced L929 apoptosis activity better than NF and ID, as shown in the SPR assay, in which the TNF-α affinity of NNF (Kd = 20.71 µM) was better than those of NF and ID (Kd = 61.19 and 31.02), indicating that compounds with good specificity for TNF-α proteins will have a strong effect on neutralizing TNF-α-induced L929 cell cytotoxicity.
It has been reported that NF has anti-inflammatory activity [34]. Previous studies have indicated that NF reduces the level of IL-6 and the phosphorylation level of IκB to alleviate LPS-induced inflammation [35]. Other studies have demonstrated that NF has anti-inflammatory and antitumor properties through the regulation of NF-κB [36]. Tumor necrosis factor receptor type 1 (TNF-R1) is widely distributed on the surfaces of normal cell membranes [37], and crosslinking with TNF-α produces a classic proinflammatory response, the most classic of which is the activation of NF-κB [38]. The reason why NF regulates NF-κB can be explained in part because NF acts as a TNF-α ligand. Additionally, there have been many reports of NF having anti-obesity activity and improving obesity-related diseases [39]. NF is a major aporphine alkaloid constituent present in lotus leaves, and through its activity, the lipid-lowering, antioxidant and anti-inflammatory properties of the lotus leaves [40] are explained. Therefore, screening NF, ID, and NNF from lotus leaves not only identified new TNF-α inhibitors but also provided a valuable reference for explaining the anti-inflammatory activity of lotus leaves.
Conclusions
SPR–AIRS was used to screen active components from nine medicinal herbs of EQG. NF, ID, and NNF were screened out to make TNF-α ligands and validated by SPR binding experiments and molecular docking. Experiments with L929 cells revealed that the binding of NF, ID, and NNF to TNF-α resulted in inhibitory effects on TNF-α-induced cytotoxicity. NF and NNF had no cytotoxicity to RAW264.7 cells and could inhibit the phosphorylation of IKKβ and IκB in LPS-induced RAW264.7 cells. Although ID inhibited the phosphorylation of IKKβ and IκB, ID was cytotoxic to RAW264.7 cells. Taken together, these results demonstrate that NF, ID, and NNF are potential TNF-α inhibitors. Our study showed the feasibility of applying SPR–AIRS to screen active components targeting TNF-α and high-resolution MS identification for screening TNF-α ligands from lotus leaf extract. This illustrates that NF, ID, and NNF may be partially active components in the treatment of NASH by EQG. This also provides a valuable reference for the material basis and mechanism of action of EQG in the treatment of NASH.
References
Winkelmayer WC, Finkel KW. Prevention of acute kidney injury using vasoactive or antiplatelet treatment: three strikes and out? JAMA. 2014;312(21):2221–2.
Williams CD, Stengel J, Asike MI, Torres DM, Shaw J, Contreras M, Landt CL, Harrison SA. Prevalence of nonalcoholic fatty liver disease and nonalcoholic steatohepatitis among a largely middle-aged population utilizing ultrasound and liver biopsy: a prospective study. Gastroenterology. 2011;140(1):124–31.
Wong RJ, Aguilar M, Cheung R, Perumpail RB, Harrison SA, Younossi ZM, Ahmed A. Nonalcoholic steatohepatitis is the second leading etiology of liver disease among adults awaiting liver transplantation in the United States. Gastroenterology. 2015;148(3):547–55.
Lu S, Wang Y, Liu J. Tumor necrosis factor-α signaling in nonalcoholic steatohepatitis and targeted therapies. J Genet Genomics. 2022;49(4):269–78.
Schuster S, Cabrera D, Arrese M, Feldstein AE. Triggering and resolution of inflammation in NASH. Nat Rev Gastroenterol Hepatol. 2018;15(6):349–64.
Schramm C, Schneider A, Marx A, Lohse AW. Adalimumab could suppress the activity of non-alcoholic steatohepatitis (NASH). Z Gastroenterol. 2008;46(12):1369–71.
Chaparro M, Guerra I, Muñoz-Linares P, Gisbert JP. Systematic review: antibodies and anti-TNF-α levels in inflammatory bowel disease. Aliment Pharmacol Ther. 2012;35(9):971–86.
Mahjoubin-Tehran M, De Vincentis A, Mikhailidis DP, Atkin SL, Mantzoros CS, Jamialahmadi T, Sahebkar A. Non-alcoholic fatty liver disease and steatohepatitis: state of the art on effective therapeutics based on the gold standard method for diagnosis. Mol Metab. 2021;50: 101049.
Akriviadis E, Botla R, Briggs W, Han S, Reynolds T, Shakil O. Pentoxifylline improves short-term survival in severe acute alcoholic hepatitis: a double-blind, placebo-controlled trial. Gastroenterology. 2000;119(6):1637–48.
Acedo SC, Caria CR, Gotardo ÉM, Pereira JA, Pedrazzoli J, Ribeiro ML, Gambero A. Role of pentoxifylline in non-alcoholic fatty liver disease in high-fat diet-induced obesity in mice. World J Hepatol. 2015;7(24):2551–8.
Yan T, Yan N, Wang P, Xia Y, Hao H, Wang G, Gonzalez FJ. Herbal drug discovery for the treatment of nonalcoholic fatty liver disease. Acta Pharm Sin B. 2020;10(1):3–18.
Wu X, Yuan M, He NS, Zou N, Qin DM, Zhang XM, Gan X, Yang JY, Wang XL, Yang BJ, Yuan JQ. Effect of Erhuang quzhi granules on liver function and blood lipid in patients with nonalcoholic fatty liver disease. J Tradit Chin Med. 2021;62(14):1243–6.
Xiao S, Yu R, Ai N, Fan X. Rapid screening natural-origin lipase inhibitors from hypolipidemic decoctions by ultrafiltration combined with liquid chromatography-mass spectrometry. J Pharm Biomed Anal. 2015;104:67–74.
Choi Y, Jung Y, Kim SN. Identification of Eupatilin from Artemisia argyi as a selective PPARα agonist using affinity selection ultrafiltration LC-MS. Molecules. 2015;20(8):13753–63.
Jerabek-Willemsen M, Wienken CJ, Braun D, Baaske P, Duhr S. Molecular interaction studies using microscale thermophoresis. Assay Drug Dev Technol. 2011;9(4):342–53.
Al Hamoui Dit Banni G, Nasreddine R, Fayad S, Colas C, Marchal A, Nehmé R. Investigation of lipase-ligand interactions in porcine pancreatic extracts by microscale thermophoresis. Anal Bioanal Chem. 2021;413(14):3667–81.
Seidel SA, Wienken CJ, Geissler S, Jerabek-Willemsen M, Duhr S, Reiter A, Trauner D, Braun D, Baaske P. Label-free microscale thermophoresis discriminates sites and affinity of protein-ligand binding. Angew Chem Int Ed Engl. 2012;51(42):10656–9.
Chavanieu A, Pugnière M. Developments in SPR fragment screening. Expert Opin Drug Discov. 2016;11(5):489–99.
Mulla T, Patil S, Jadhav J. Exploration of surface plasmon resonance for yam tyrosinase characterization. Int J Biol Macromol. 2018;109:399–406.
Moscetti I, Cannistraro S, Bizzarri AR. Surface plasmon resonance sensing of biorecognition interactions within the tumor suppressor p53 network. Sensors. 2017;17(11):2680–97.
Liu L, Hua Y, Wang D, Shan L, Zhang Y, Zhu J, Jin H, Li H, Hu Z, Zhang W. A sesquiterpene lactone from a medicinal herb inhibits proinflammatory activity of TNF-α by inhibiting ubiquitin-conjugating enzyme UbcH5. Chem Biol. 2014;21(10):1341–50.
Navratilova I, Besnard J, Hopkins AL. Screening for GPCR ligands using surface plasmon resonance. ACS Med Chem Lett. 2011;2(7):549–54.
Zeng KW, Wang JK, Wang LC, Guo Q, Liu TT, Wang FJ, Feng N, Zhang XW, Liao LX, Zhao MM, Liu D, Jiang Y, Tu P. Small molecule induces mitochondrial fusion for neuroprotection via targeting CK2 without affecting its conventional kinase activity. Signal Transduct Target Ther. 2021;6(1):71–83.
Kaiqi Su, Zou Q, Ning Hu, Wang P. High-sensitive and high-efficient biochemical analysis method using a bionic electronic eye in combination with a smartphone-based colorimetric reader system. Sens Actuators, B Chem. 2015;216(9):134–40.
Chen L, Lv D, Chen X, Liu M, Wang D, Liu Y, Hong Z, Zhu Z, Hu X, Cao Y, Yang J, Chai Y. Biosensor-based active ingredients recognition system for screening STAT3 ligands from medical herbs. Anal Chem. 2018;90(15):8936–45.
Fu R, Yang P, Li Z, Liu W, Amin S, Li Z. Avenanthramide A triggers potent ROS-mediated anti-tumor effects in colorectal cancer by directly targeting DDX3. Cell Death Dis. 2019;10(8):593–616.
He MM, Smith AS, Oslob JD, Flanagan WM, Braisted AC, Whitty A, Cancilla MT, Wang J, Lugovskoy AA, Yoburn JC, Fung AD, Farrington G, Eldredge JK, Day ES, Cruz LA, Cachero TG, Miller SK, Friedman JE, Choong IC, Cunningham BC. Small-molecule inhibition of TNF-alpha. Science. 2005;310(5750):1022–5.
Baghdasaryan A, Claudel T, Gumhold J, Silbert D, Adorini L, Roda A, Vecchiotti S, Gonzalez FJ, Schoonjans K, Strazzabosco M, Fickert P, Trauner M. Dual farnesoid X receptor/TGR5 agonist INT-767 reduces liver injury in the Mdr2-/- (Abcb4-/-) mouse cholangiopathy model by promoting biliary HCO-3 output. Hepatology. 2011;54 4:1303–12.
Ma L, Gong H, Zhu H, Ji Q, Su P, Liu P, Cao S, Yao J, Jiang L, Han M, Ma X, Xiong D, Luo HR, Wang F, Zhou J, Xu Y. A novel small-molecule tumor necrosis factor α inhibitor attenuates inflammation in a hepatitis mouse model. J Biol Chem. 2014;289(18):12457–66.
Merkel O, Wacht N, Sifft E, Melchardt T, Hamacher F, Kocher T, Denk U, Hofbauer JP, Egle A, Scheideler M, Schlederer M, Steurer M, Kenner L, Greil R. Actinomycin D induces p53-independent cell death and prolongs survival in high-risk chronic lymphocytic leukemia. Leukemia. 2012;26(12):2508–16.
Hayden MS, Ghosh S. Shared principles in NF-kappaB signaling. Cell. 2008;132(3):344–62.
Wang F, Cao J, Hou X, Li Z, Qu X. Pharmacokinetics, tissue distribution, bioavailability, and excretion of nuciferine, an alkaloid from lotus, in rats by LC/MS/MS. Drug Dev Ind Pharm. 2018;44(9):1557–62.
Wan Y, Xia J, Xu JF, Chen L, Yang Y, Wu JJ, Tang F, Ao H, Peng C. Nuciferine, an active ingredient derived from lotus leaf, lights up the way for the potential treatment of obesity and obesity-related diseases. Pharmacol Res. 2022;175: 106002.
Wu H, Yang Y, Guo S, Yang J, Jiang K, Zhao G, Qiu C, Deng G. Nuciferine ameliorates inflammatory responses by inhibiting the TLR4-mediated pathway in lipopolysaccharide-induced acute lung injury. Front Pharmacol. 2017;8:939.
Chen X, Zheng X, Zhang M, Yin H, Jiang K, Wu H, Dai A, Yang S. Nuciferine alleviates LPS-induced mastitis in mice via suppressing the TLR4-NF-κB signaling pathway. Inflamm Res. 2018;67(11–12):903–11.
Song C, Cao J, Lei Y, Chi H, Kong P, Chen G, Yu T, Li J, Kumar Prajapati R, Xia J, Yan J. Nuciferine prevents bone loss by disrupting multinucleated osteoclast formation and promoting type H vessel formation. FASEB J. 2020;34(3):4798–811.
Sedger LM, McDermott MF. TNF and TNF-receptors: from mediators of cell death and inflammation to therapeutic giants - past, present and future. Cytokine Growth Factor Rev. 2014;25(4):453–72.
Jaco I, Annibaldi A, Lalaoui N, Wilson R, Tenev T, Laurien L, Kim C, Jamal K, Wicky John S, Liccardi G, Chau D, Murphy JM, Brumatti G, Feltham R, Pasparakis M, Silke J, Meier P. MK2 phosphorylates RIPK1 to prevent TNF-induced cell death. Mol Cell. 2017;66(5):698–710.
Wang Y, Yao W, Li B, Qian S, Wei B, Gong S, Wang J, Liu M, Wei M. Nuciferine modulates the gut microbiota and prevents obesity in high-fat diet-fed rats. Exp Mol Med. 2020;52(12):1959–75.
Kim BM, Cho BO, Jang SI. Anti-obesity effects of Diospyros lotus leaf extract in mice with high-fat diet-induced obesity. Int J Mol Med. 2019;43(1):603–13.
Acknowledgements
This work was supported by the Financial Science and Technology Plan Project of Xinjiang (Grant Numbers: 2020AA005, 2020A03004-2), the Grant from the National Natural Science Foundation of China (82174075, U1903211), the Grant from Shihezi University (CGZH201806, CXBJ202006).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guan, Jw., Xu, Y., Yu, W. et al. Biosensor-based active ingredient recognition system for screening TNF-α inhibitors from lotus leaves. Anal Bioanal Chem 415, 1641–1655 (2023). https://doi.org/10.1007/s00216-023-04565-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-023-04565-2