Abstract
Food supplement authentication is an important concern worldwide due to the ascending consumption related to health benefits and its lack of effective regulation in underdeveloped countries, making it a target of fraudulent activities. In this context, this study evaluated fish oil supplements by comprehensive two-dimensional gas chromatography coupled to mass spectrometry (GC×GC–MS) to obtain fingerprints, which were used to build predictive models for automated authentication of the most popular products sold in Brazil. The authentication process relied on a one-class classifier model using data-driven soft independent modeling of class analogy (DD-SIMCA). The output of the model was a binary classifier: certified IFOS fish oils and non-certified ones — regardless of the source of adulteration. The compositional analysis showed a significant variation in the samples, which validated the need for reliable statistical models. The DD-SIMCA algorithm is still incipient in GC×GC studies, but it proved to be an excellent tool for authenticity purposes, achieving a chemometric model with a sensitivity of 100%, specificity of 98.6%, and accuracy of 99.0% for fish oil authentication. Finally, orthogonalized partial least square discriminant analysis (OPLS-DA) was used to identify the features that distinguished the groups, which ascertained the results of the DD-SIMCA model that IFOS-certified oils are positively correlated to omega-3 fatty acids, including eicosapentaenoic acid (EPA, C20:5 n-3) and docosahexaenoic acid (DHA, C22:6 n-3).
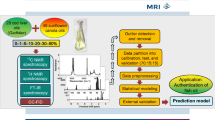
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Food supplements are an important source of bioactive substances, which can be sold as capsules or liquid dosage forms to complement the regular diet [1,2,3,4]. In 2016, the food supplement industry contributed with over 120 billion dollars to the US economy [5].
Many food supplements are commercially available, and fish oil is the most popular product worldwide [6, 7], probably due to several reports showcasing important health benefits [8,9,10]. Such benefits are related to the occurrence of polyunsaturated fatty acids, wherein the most important are the omega-3 (n-3) constituents [11, 12]. Fish oil is rich in eicosapentaenoic acid (EPA, C20:5 n-3) and docosahexaenoic acid (DHA, C22:6 n-3), which are different omega-3 fatty acids exclusively found in marine products [11,12,13]. Other fatty acids are found in seeds and plant oils such as alpha-linolenic acid (C18:3 n-3), linoleic acid (C18:2 n-6), arachidonic acid (C20:3 n-6), oleic acid (C18:1 n-9), and nervonic acid (C24:1 n-9) [11, 12].
The increasing demand for fish oil and the lack of effective regulation in some countries, such as underdeveloped countries, are a potential scenario for fraudulent practices, like mislabeling and adulteration with lower quality ingredients [1, 3, 13, 14]. For this reason, the assessment of fish oil quality has raised concerns among consumers and regulatory agencies worldwide [13, 14]. This awareness has led to the implementation of the international fish oil standards (IFOS), which is the oldest and most popular certification by Nutrasource, a Canadian company [15]. The IFOS-certified products meet the levels of omega-3 fatty acid described on the labels and are tested to check for possible oxidation and contaminants found in fish, including dioxins, furans, mercury, and lead [15]. Consequently, the IFOS-certified products are typically more expensive than the non-certified analogs, which raises concerns about the safety and quality of the latter products. Hence, quality assurance protocols are desperately needed to monitor fish oil supplements.
Multiple analytical platforms are needed to assess the inorganic and organic constituents of fish oils. However, this study focuses on the analysis of the fatty acids found in such products, which are the main ingredients of this supplement. Several studies were reported evaluating fish oil quality using different analytical techniques [14], such as vibrational spectroscopy [16,17,18,19,20], nuclear magnetic resonance [19, 21, 22], liquid chromatography (LC) [23,24,25], and gas chromatography (GC) [13, 26,27,28]. More specifically, an interesting technique for the analysis of the individual fatty acid isomers is comprehensive two-dimensional gas chromatography (GC×GC) [29].
Comprehensive two-dimensional gas chromatography is a multidimensional separation technique that explores two sequential GC stages to improve the overall peak capacity of the composite system [30,31,32,33], being considered an ideal platform for untargeted analysis in foodomic investigations [34]. Important reports of fish oil analysis using GC×GC include analysis of dioxins in fish oil [35], profiling of fatty acid methyl esters (FAME) in menhaden fish oil [36], and occurrence of persistent organic pollutants in fish oils [37]. However, to the best of our knowledge, there are no studies using GC×GC to study fish oil certification.
Fundamental studies with product certification and adulteration are challenging because the quality assurance and nature of potential adulterants are highly dependent on the country. Usually, underdeveloped countries are more susceptible to adulterations, which challenges multiclass classification. For instance, discriminant analysis using partial least squares (PLS-DA) may be unsuccessful if many impurities and adulterants are used, as the intra-class variation may become too large, jeopardizing the performance of the chemometric model. Moreover, if an adulterant was not used in the training dataset but is detected in test data, then the sample might be misclassified by the PLS-DA model. In this context, one-class chemometric models are particularly useful, as the certification model relies solely on the characteristics of the desired product, regardless of the impurities or adulterants [38, 39].
More specifically, data-driven soft independent modeling of class analogy (DD-SIMCA) may be explored for one-class classifiers, being an ideal solution for food supplement (e.g., fish oil) authentication resulting in a binary decision. In other words, the model output informs the analyst if the sample is authentic or not — regardless of the reason why the adulterated exhibited a different chromatographic profile. If discriminant analysis was used, the number of classes being modeled likely would increase with the number of adulterants being addressed by the investigation [40, 41]. To accomplish this goal, authentic samples are graphically assigned within a statistically relevant acceptance area, while the model rejects non-authentic samples, as described in the “DD-SIMCA” section.
In this study, we evaluated an alternative method for fish oil authentication according to the IFOS certification. The thirty-two selected brands were the most popular fish oil supplements sold through e-commerce in Brazil. The experiments were carefully designed for the use of one-class classifiers using DD-SIMCA chemometric modeling [38, 39]. A pixel-based data analysis approach was employed using the GC×GC chromatograms as chemical fingerprints (i.e., untargeted analysis) for the classification model [42,43,44,45], followed by orthogonalized partial least squares discriminant analysis (OPLS-DA) with the sole purpose of generating a loading tensor to validate the findings of the DD-SIMCA. Finally, important figures of merit for the DD-SIMCA classification model were determined during method development, including sensitivity, specificity, and accuracy [46]. It is hoped that this report will highlight the potential application of one-class classifiers to authentication studies with chemometrics in the field of GC×GC.
Materials and methods
Materials and samples
The chemicals used during the derivatization process were sodium methoxide, formic acid, dichloromethane, and sodium sulfate (Sigma-Aldrich, St, Louis, MO, USA). The 2-mL tubes were purchased from Eppendorf (São Paulo, SP, Brazil). Methanol and methylene chloride were purchased from Synth (Diadema, São Paulo, Brazil), and sodium sulfate was purchased from CAQ (Diadema, São Paulo, Brazil). A set of 2-mL screw-top glass vials with PFTE/PDMS septum was purchased from Nova Analítica (São Paulo, São Paulo, Brazil). The samples consisted of 32 brands of commercial fish oil supplement capsules purchased from e-commerce (Table 1). The experiments were performed with true replicates (N = 3). Peak identification was confirmed using the Supelco 37 Component FAME Mix (Sigma-Aldrich), alongside confirmation with mass spectral library search (Wiley and NIST) and selected references [36, 47].
Derivatization method
The procedure was adapted from Bogusz et al. [48]. An aliquot of 200 mg of fish oil sample was weighted in a 2 mL tube. An aliquot of 200 μL of saturated sodium methoxide in methanol was added. The reaction proceeded at 70ºC for 5 min using a MULTI REAX vortex mixer (Heidolph Instruments GmbH & Co. KG, Schwabach, Germany). Next, an aliquot of 100 μL of formic acid solution (0.5 mol L−1) was added to neutralize the basic catalyst. To remove all remaining water, sodium sulfate was added to the tube. Then, 1 mL of methylene chloride was added and vortexed for 15 s. The tubes containing the samples were centrifuged (ROTANTA 460R, Hettich, Beverly, MA, USA) for 5 min at 9056 × units of gravity (g). Finally, 600 μL of the supernatant was collected and transferred to a 2-mL screw-top glass for GC×GC analysis.
Instrumentation
Comprehensive two-dimensional gas chromatography coupled with mass spectrometry (GC×GC–MS) was used to obtain the FAME profile of the 32 fish oil supplements. The system consisted of a TRACE 1300 GC coupled to an ISQ single transmission quadrupole mass spectrometer (ThermoFisher Scientific, Waltham, MA, USA). A TriPlus RSH autosampler (ThermoFisher Scientific) was used to inject 1 µL of the liquid sample with a 60:1 split ratio at 280 °C. The non-polar × mid-polar column configuration consisted of a first 20 m × 0.18 mm-ID (0.18 µm film thickness) SLB-1 ms column (100% poly(dimethylsiloxane)) (Supelco, Bellefonte, PA, USA) and a second 2.5 m × 0.25 mm-ID (0.25 µm film thickness) MEGA-17 column (poly(diphenyl-dimethylsiloxane) with 50% diphenylsiloxane monomer incorporation) — adapted from [49].
The flow modulation was performed using the reverse fill/flush configuration [50] employing three-port and four-port SilFlow GC splitters (Trajan Scientific and Medical, Melbourne, Australia) [51, 52]. A 50-µL sampling loop comprising a 23 cm × 0.53 mm-ID MXT deactivated guard column (Restek Corporation) and a bleed capillary of 2.5 m × 0.10 mm-ID were employed for flow modulation [53]. A three-way miniature diaphragm isolation valve (The Lee Company, Westbrook, CT, USA) was used to actuate the auxiliary gas during flow modulation. A modulation period of 4 s with a flush period of 250 ms was used. For flow splitting, an unpurged 3-port SilFlow GC splitter (Trajan Scientific) and two 5.0 m × 0.18 mm-ID and 5.0 m × 0.32 mm-ID fused silica capillaries (Restek Corporation) were used for passive division of the 2D flow. The 0.18 mm-ID and 0.32 mm-ID capillaries were used as the transfer line to the MS and FID, respectively.
The oven temperature was programmed from 160 °C to 290 °C at 3 °C min−1. The transfer line and ion source temperatures were set at 280 °C and 250 °C, respectively. Helium was used as carrier and auxiliary gas at 0.5 mL min−1 and 25 mL min−1, respectively. EI was performed at 70 eV. Full MS (centroid mode) was used for data acquisition (25.60 Hz) with a spectral range of 50 to 400 u at nominal mass resolution. Blank runs were performed every 5 injections for quality assurance.
Xcalibur (ThermoFisher Scientific) software was used for instrument control and data acquisition. ChromSpace (SepSolve Analytical, Waterloo, ON, Canada) was employed to control the flow-modulator and synchronize the GC run. GC Image (GC Image, LLC, Lincoln, NE, USA) was used for qualitative analysis using the template matching feature for batch processing. The “*.RAW” Xcalibur files were converted to “*.CDF” ANDI/netCDF format using the file converter plug-in. Chemometrics was performed on MATLAB R2021a (MathWorks, Natick, MA, USA) environment. The netCDF files were imported to MATLAB to generate three-way data tensors. DD-SIMCA was performed using a modified algorithm from Pomerantsev et al. [38, 39].
Data preprocessing
A pixel-based approach was selected for multivariate data processing [54, 55]. The interested reader is directed elsewhere for more information on handling non-integer acquisition rates [56]. The GC×GC chromatograms were imported to MATLAB as three order tensors (i.e., data cube) (1D by 2D by m/z). The augmented matrices were prepared by unfolding them to a matrix (1D by 2D × m/z) for one-class analysis using DD-SIMCA [38, 39]. The peaks from the blank runs were removed from the dataset prior to chemometrics. Next, intensities of all ions in the corresponding mass spectra (i.e., belonging to the same scan) were summed up to create the total ion chromatograms (TIC), which were organized into a matrix (samples by TIC). Finally, chromatograms were normalized, Pareto-scaled, and mean-centered before the modeling step. The interested reader is directed elsewhere for more information on the preprocessing steps [57]
DD-SIMCA
Classification is a statistical problem that aims to answer to which class an unknown object belongs, based on a model that contains objects whose categories are well-established [41]. A category or class is a group of objects that can be defined by common properties shared among all its members. Therefore, classification methods will differ from each other on how to statistically estimate the class limits in a multivariate space.
Pomerantsev et al. [38] proposed in 2008 an algorithm that uses distances based on leverage and residual variance from principal component analysis (PCA) to assort unknown objects. The method named DD-SIMCA is based on SIMCA, which is a multiclass well-established classification algorithm proposed by Wold [58]. In DD-SIMCA, score distance (SD) and orthogonal distance (OD) standardized by a chi-squared (χ2) distribution are used to describe a function that delimits a single-class acceptance area with a significance level (e.g., 95%). To calculate these limits, scaling factors (\({h}_{0}\) and \({v}_{0}\)) and the number of degrees of freedom (\({N}_{h}\) and \({N}_{v}\)) are needed. These parameters can be estimated from a training dataset (X). The first step consists in the decomposition of X (I × J), as in PCA:
where T (I × A) is the score matrix, with A being related to the used number of principal components (PC), P (J × A) is the loadings matrix, and E (I × J) is the residuals matrix. In the second step, the calculated scores (T) and loadings (P) enable the extraction of \({h}_{i}\) and \({v}_{i}\) for score and orthogonal distances, respectively:
where λa, a = 1, …, A are the diagonal elements of the matrix TtT:
When both OD and OS are calculated, the total distance c can be calculated for each sample:
where \({N}_{h}\), \({h}_{0}\), \({N}_{v}\), and \({v}_{0}\) are unknown at first but are estimated by a data-driven approach [38, 59]. Thus, based on a ccrit value, we can develop the acceptance area for a given value α [59]. So, from now on any, new sample will be considered authentic, or otherwise, it will be classified as unauthentic — regardless of the features responsible for such differentiation:
This binary decision rule is what delimits objects considered authentic and non-authentic. This characteristic is what differentiates DD-SIMCA from discriminant analysis, such as PLS-DA. In DD-SIMCA, only one well-known class of authentic objects (X, target class) is necessary to allocate news samples as being authentic (or not). Conversely, PLS-DA intrinsically needs predefined classes to do the classification task, which makes it difficult to classify an object that does not belong to any of the pre-established specific classes. In summary, a PLS-based discriminant model will classify a new sample only if this sample is a member of one of the predefined classes, which makes such methods not the most suitable for authentication investigations [41].
OPLS-DA
In this study, OPLS-DA modeling was performed only to obtain the loading array to evaluate the features and ascertain the results obtained with DD-SIMCA [60,61,62].
Results and discussion
Sample discussion
Fish oil supplement may be considered a complex sample due to the occurrence of many critical pairs of positional isomers of the polyunsaturated fatty acid methyl esters. The successful use of GC×GC was reported to improve the chromatographic resolution of such challenging peak clusters [36, 47]. In addition to the enhanced separation, qualitative analysis also benefited from the ordered and structured GC×GC chromatograms, which evidenced the elution patterns of the FAME peaks. For example, using a non-polar × mid-polar column configuration, it was possible to establish elution regions for the peaks of saturated, mono-, di-, tri-, tetra-, penta-, and hexa-unsaturated FAMEs. This pattern alongside the use of authentic standards enabled the identification of the peaks detected in the GC×GC chromatograms. Interestingly, the number of detected peaks in the fish oil supplements seemed to exhibit fewer peaks than those found in biological samples, like fresh herring oil [47]. Furthermore, the current method was capable of distinguishing between triglycerides and free fatty acids that composed the fish oil supplements — as the evaluated products did not specify the type of lipids used in the formulation. Since transesterification was used for analyte derivatization, it can be stated that the FAMEs originated from triglyceride-like structures due to the nature of the base-catalyzed reaction. However, such reaction does not methylate free fatty acids, so it was possible to detect the free fatty acids as well. While this condition allowed profiling the composition of triglyceride-based fatty acids and the actual free fatty acids, it was observed some tailing in the 2D for the hydrogen-bonding analytes. However, such tailing did not cause any peak overlap in the GC×GC chromatograms. For most samples, the relative abundance of free fatty acids was found between 0 and 2%. Conversely, the relative concentration was 23 to 69% for the samples A6, A7, B3, and B20.
The average composition (N = 3) of the 10 most representative samples is shown using a heatmap plot (Fig. 1). Interestingly, there were no clear patterns among the samples, as the fish oil supplements seemed to be composed of triglycerides and free fatty acids arbitrarily with respect to the IFOS certification. In addition, it was seen that the intra-group compositional variation was significant, as highlighted by the hierarchical cluster analysis (HCA) (Fig. 1), which supports the use of a one-class classifier model like DD-SIMCA. The assigned classes 0 (red) and 1 (green) in Fig. 1 represent the samples with and without IFOS certification, respectively.
Hierarchical clustering heatmaps of the fatty acid composition of representative fish oil supplements. The peak areas were normalized and autoscaled. The distance measured was Euclidean. The clustering method used was Ward. The dendrogram illustrates the large intra-class variation among the two evaluated classes, namely, IFOS-certified supplements (class 0, red) and IFOS-non-certified (class 1, green). Abbreviations: free fatty acids (FA), fatty acid methyl esters (FAME)
Noteworthy, the most important fatty acids in fish oil supplements are C20:5 n-3 (EPA) and C22:6 n-3 (DHA). Correlation analysis of the composition of all the samples (Fig. 2) shows that the peak intensities of EPA and DHA are negatively correlated within the samples evaluated in this study. In other words, when the concentration of DHA is high in a particular brand of fish oil, there is less amount of EPA in the same product. Conversely, positive correlations were observed between DHA with shorter chain fatty acids, like C16 and C18, while the same was not be seen for EPA (Fig. 2).
Correlation heatmaps of the fatty acid composition of fish oil supplements. The data input was the peak areas of the free fatty acids (FA) and the fatty acid methyl esters (FAME). The distance measured was Pearson r. The cell color indicates the correlation coefficient between the two selected variables (or features). Caption: positive correlation values (red), negative correlation values (blue)
The chemometric approach selected herein was a pixel-based approach for one-class classification using DD-SIMCA. It must be stated that the DD-SIMCA algorithm is suitable for both peak table and pixel-based multivariate data analysis. Particularly, this pixel-based approach was interested in evaluating how the DD-SIMCA algorithm handled chromatograms (Fig. 3) with peak intensities that varied over a wide dynamic range, including column overloading — as highlighted in Fig. 4. This specific broad dynamic range problem is already known to be challenging for peak table processing, as overloaded peaks generally cause peak splitting during integration, resulting in a single feature being described with two peak apexes [63].
Expanded total ion chromatogram (TIC) obtained by GC×GC–MS of fish oil supplements. The peaks of the C18:1 isomers exhibited a broad range of intensities in the evaluated fish oil supplements yielding no column overloading (A), column overloading (B), and excessive column overloading (C). As a consequence, the automated peak integration resulted in split peaks (multiple apexes/features for the same compound). This finding motivated the evaluation of pixel-based multivariate data processing to handle the wide dynamic range problem
DD-SIMCA
The first and most important step in creating one-class models is to establish a well-describe class — named target class — in this case, samples with IFOS certificate.
Next, the model was tested against non-certified samples to estimate the figures of merit to evaluate this proof of concept.
All supplements were used for modeling and evaluation, totalizing 7 certified and 25 non-certified samples. The 31 and 72 chromatograms were used for calibration and external validation of the one-class model, respectively. Each chromatogram exhibited 48,123 variables (i.e., pixels). To build the model, the calibration data were normalized and Pareto-scaled. Chi-squared-type acceptance areas were determined, and the number of PCs of the model was set to five the leave-one-out cross-validation method. Outlier significance was set to 0.01, and type I error (α) was calculated as approximately 0.04. The classic estimation method [64] was chosen for the final model (Fig. 5).
The one-class model was mathematically successful as there were no extreme or outliers found in the training dataset (green markers, Fig. 6), corroborating the expected value of 4% type I error (reference value). In the external validation step (blue markers, Fig. 6), only one sample was misclassified, resulting in a type 2 error of 1.4%, which is three times less than the 4.3% calculated for this data set [38, 65]. Important figures of merit like sensitivity (Eq. 6), specificity (Eq. 7), and accuracy (Eq. 8) were calculated for this model by considering the occurrence of true positive (TP), true negative (TN), false positive (FP), and false negative (FN) [46]. This model achieved a sensitivity value of 100%, specificity of 98.6%, and accuracy of 99.0%, which suggests an excellent performance for fish oil authentication, due to a single misclassification. However, when assessing which sample was mismatched, only the second replicate of sample B1 was found. Accordingly, no further actions were necessary.
Remarkably, despite the significant intra-class compositional variation and occurrence of non-linear chromatography (i.e., overloaded peaks in some samples), the one-class DD-SIMCA model was performed adequately for fish oil authentication. This pixel-based chemometric protocol is an interesting alternative for routine studies as all steps involved in data processing can be readily automated and parameters are not subjective. For instance, this pixel-based approach successfully bypassed the need for peak integration, which may require analyst intervention to address split peaks (i.e., one feature and two apexes).
The output differences between DD-SIMCA and PLS-DA must be highlighted to showcase the benefits of one-class classifiers for authentication studies. It is clear from Fig. 6 that the non-certified samples are comprised by more than one class of external objects, above and below log (1 + \(\frac{v}{{v}_{0}}\)). This means that in terms of multiclass classification (e.g., PLS-DA), more than one class (at least 2) of non-certified samples was needed to allocate all possibilities of the accredited samples. Furthermore, even if it was beforehand possible to prepare such sub-classes, a discrimination model will likely provide a wrong answer to the authenticity question, once any new object must be attributed to one of the predefined classes. The reason for this limitation is that the analyst must know exactly which adulterants or ingredients differ in the formulations to estimate the number of classes. This procedure is important for multiclass models to minimize the intra-class variation.
Loadings assessment using OPLS-DA
Lastly, the features were evaluated to ascertain the chemical validity of the DD-SIMCA model. Considering that DD-SIMCA does not report loadings arrays, OPLS-DA was used to generate results as similar as possible to the DD-SIMCA model. The interested reader should keep in mind that the classification model used a linear combination of the original pixels to perform the one-class predictions. In other words, the entire chromatogram is used as an input for chemometric modeling, instead of only a handful of “markers”.
OPLS-DA was applied to determine which variables are responsible for distinguishing between certified and uncertified samples. Training and validation sets consisted of both IFOS certified and uncertified samples, which were randomly divided to contain 75 and 25% of the samples of each class in those sets respectively, i.e., 23 certified and 54 uncertified chromatograms in the training set and 8 certified and 18 uncertified chromatograms in the validation set. Three latent variables (LV) were selected based on the lowest values of venetian-blind cross-validation average errors. It was found that the first latent variable (LV) was responsible for the discrimination between IFOS-certified and uncertified samples. The certified samples were found in the negative region and the latter in the positive region of the scores plot. Accordingly, the same is valid for the loadings (Fig. 7), since negative loadings are related to the certified samples, whereas the positive loadings are related to the uncertified ones. The peaks in the negative region are related to peaks of C20:5 n-3 (EPA) and C22:6 n-3 (DHA). Conversely, peaks in the positive region are related to other fatty acids (C16 and C18), which was consistent with the compositional analysis reported in Fig. 2. Interestingly, compositional analysis can lead to misunderstandings in the interpretation of the loadings, as EPA and DHA are not correlated to each other, but might be correlated to C16 and C18 fatty acids (Fig. 2). However, it is important to keep in mind that loadings are coefficients of the linear combination of the variables from which the principal components were constructed. Therefore, higher loading values do not necessarily mean higher peak intensities (or areas for the peak table approach), but rather more importance to the chemometric model.
Conclusion
The proposed method combining GC×GC–MS and one-class classifiers was shown to be interesting alternative for automated authentication of fish oil supplements. The chromatographic method was able to distinguish between the types of lipids that that composed the fish oil supplements, like free fatty acids and triglycerides. Also, it was found that EPA and DHA were negatively correlated within the evaluated samples. In other words, it was found that when DHA is present with higher peak areas, less amount of EPA was found in the same product. In terms of authentication, the chemometric model was considered successful with only one replicate misclassified. Therefore, the final DD-SIMCA model presented a type 2 error of 1.4%, 100% of sensitivity, 98.6% of specificity, and 99.0% of accuracy. Lastly, loadings obtained by OPLS-DA confirmed the findings of DD-SIMCA, wherein the peaks of C20:5 n-3 (EPA) and C22:6 n-3 (DHA) were important to build the chemometric model used for fish oil authentication. However, this task is not to be taken lightly as there was significant intra-class variation, as shown by the HCA dendrogram. Finally, we hope to illustrate the interesting opportunities of one-class classifier models for GC×GC-based chemometrics, as this approach is still incipient in the separation community.
References
Fibigr J, Šatínský D, Solich P. Current trends in the analysis and quality control of food supplements based on plant extracts. Anal Chim Acta. 2018;1036:1–15. https://doi.org/10.1016/j.aca.2018.08.017.
Costa JG, Vidovic B, Saraiva N, Costa do Ceu M, Del Favero G, Marko D, Oliveira NG, Fernandes AS. Contaminants: a dark side of food supplements? Free Radic Res. 2019;53:1113–35. https://doi.org/10.1080/10715762.2019.1636045.
Kowalska A, Bieniek M, Manning L. Food supplements’ non-conformity in Europe – Poland: a case study. Trends Food Sci Technol. 2019;93:262–70. https://doi.org/10.1016/j.tifs.2019.09.022.
Czepielewska E, Makarewicz-Wujec M, Różewski F, Wojtasik E, Kozłowska-Wojciechowska M. Drug adulteration of food supplements: a threat to public health in the European Union? Regul Toxicol Pharmacol. 2018;97:98–102. https://doi.org/10.1016/j.yrtph.2018.06.014.
Economic impact of the dietary supplement industry. https://www.crnusa.org/resources/economic-impact-dietary-supplement-industry accessed 20 Aug 2022.
Mishra S, Stieman B, JGahche J, Potischman N. Data brief 399: dietary supplement use among adults: United States, 2017–2018. Hyattsville. 2021.
Kamiński M, Kręgielska-Narożna M, Bogdański P. Determination of the popularity of dietary supplements using Google Search rankings. Nutrients. 2020;12:908. https://doi.org/10.3390/nu12040908.
Ellulu MS, Khaza’ai H, Abed Y, Rahmat A, Ismail P, Ranneh Y. Role of fish oil in human health and possible mechanism to reduce the inflammation. Inflammopharmacology. 2015;23:79–89. https://doi.org/10.1007/s10787-015-0228-1.
Siscovick DS, Barringer TA, Fretts AM, Wu JHY, Lichtenstein AH, Costello RB, Kris-Etherton PM, Jacobson TA, Engler MB, Alger HM, Appel LJ, Mozaffarian D. Omega-3 polyunsaturated fatty acid (fish oil) supplementation and the prevention of clinical cardiovascular disease. Circulation. 2017;135: https://doi.org/10.1161/CIR.0000000000000482
Gao H, Geng T, Huang T, Zhao Q. Fish oil supplementation and insulin sensitivity: a systematic review and meta-analysis. Lipids Health Dis. 2017;16:131. https://doi.org/10.1186/s12944-017-0528-0.
Harris WS. Fish oil supplementation: evidence for health benefits. Cleve Clin J Med. 2004;71:208–10. https://doi.org/10.3949/ccjm.71.3.208.
Johnson M. Omega-3, omega-6 and omega-9 fatty acids: implications for cardiovascular and other diseases. J Glycomics Lipidomics. 2014;04: https://doi.org/10.4172/2153-0637.1000123
Galuch M, Carbonera F, Magon T, da Silveira R, dos Santos P, Pizzo J, Santos O, Visentainer J. Quality assessment of omega-3 supplements available in the Brazilian market. J Braz Chem Soc. 2017: https://doi.org/10.21577/0103-5053.20170177
Rohman A, Putri AR, Irnawati WA, Nisa K, Lestari LA. The employment of analytical techniques and chemometrics for authentication of fish oils: a review. Food Control. 2021;124:107864. https://doi.org/10.1016/j.foodcont.2021.107864.
IFOS. https://certifications.nutrasource.ca/about/how-certifications-work/ifos. Accessed 6 May 2022.
Cascant MM, Breil C, Fabiano-Tixier AS, Chemat F, Garrigues S, de la Guardia M. Determination of fatty acids and lipid classes in salmon oil by near infrared spectroscopy. Food Chem. 2018;239:865–71. https://doi.org/10.1016/j.foodchem.2017.06.158.
Hall DW, Marshall SN, Gordon KC, Killeen DP. Rapid quantitative determination of squalene in shark liver oils by Raman and IR spectroscopy. Lipids. 2016;51:139–47. https://doi.org/10.1007/s11745-015-4097-6.
Rohman A, Che Man YB. Analysis of chicken fat as adulterant in cod liver oil using Fourier transform infrared (FTIR) spectroscopy and chemometrics Análisis de grasa de pollo como adulterante en aceite de hígado de bacalao usando espectroscopía infrarroja con transformada de Fo. CyTA - J Food. 2011;9:187–91. https://doi.org/10.1080/19476337.2010.510211.
Giese E, Winkelmann O, Rohn S, Fritsche J. Determining quality parameters of fish oils by means of 1H nuclear magnetic resonance, mid-infrared, and near-infrared spectroscopy in combination with multivariate statistics. Food Res Int. 2018;106:116–28. https://doi.org/10.1016/j.foodres.2017.12.041.
Killeen DP, Card A, Gordon KC, Perry NB. First use of handheld Raman spectroscopy to analyze omega-3 fatty acids in intact fish oil capsules. Appl Spectrosc. 2020;74:365–71. https://doi.org/10.1177/0003702819877415.
Giese E, Rohn S, Fritsche J. Chemometric tools for the authentication of cod liver oil based on nuclear magnetic resonance and infrared spectroscopy data. Anal Bioanal Chem. 2019;411:6931–42. https://doi.org/10.1007/s00216-019-02063-y.
Standal IB, Axelson DE, Aursand M. 13C NMR as a tool for authentication of different gadoid fish species with emphasis on phospholipid profiles. Food Chem. 2010;121:608–15. https://doi.org/10.1016/j.foodchem.2009.12.074.
Araujo P, Tilahun E, Zeng Y. A novel strategy for discriminating marine oils by using the positional distribution (sn-1, sn-2, sn-3) of omega-3 polyunsaturated fatty acids in triacylglycerols. Talanta. 2018;182:32–7. https://doi.org/10.1016/j.talanta.2018.01.030.
Suh JH, Ho C-T, Wang Y. Evaluation of carbonyl species in fish oil: an improved LC–MS/MS method. Food Control. 2017;78:463–8. https://doi.org/10.1016/j.foodcont.2017.03.014.
Chatterjee NS, Singh A, Vishnu KV, Ajeeshkumar KK, Anandan R, Ashok Kumar K, Mathew S. Authentication of two bio-active fish oils by qualitative lipid profiling using semi-targeted approach: an exploratory study. J AOAC Int. 2020;103:78–82. https://doi.org/10.5740/jaoacint.19-0208.
Gómez-Cortés P, Camiña JM. Oxidomics on the omega-3 volatile degradation pattern to determine differences between vegetable and marine oils. Food Res Int. 2019;122:10–5. https://doi.org/10.1016/j.foodres.2019.03.064.
Araujo P, Zeng Y, Du Z-Y, Nguyen T-T, Frøyland L, Grung B. Discrimination of n-3 rich oils by gas chromatography. Lipids. 2010;45:1147–58. https://doi.org/10.1007/s11745-010-3483-3.
Khoomrung S, Raber G, Laoteng K, Francesconi KA. Identification and characterization of fish oil supplements based on fatty acid analysis combined with a hierarchical clustering algorithm. Eur J Lipid Sci Technol. 2014;116:795–804. https://doi.org/10.1002/ejlt.201300369.
Arena A, Zoccali M, Trozzi A, Tranchida PQ, Mondello L. Occurrence of mineral oil hydrocarbons in omega-3 fatty acid dietary supplements. Foods. 2021;10:2424. https://doi.org/10.3390/foods10102424.
Bahaghighat HD, Freye CE, Synovec RE. Recent advances in modulator technology for comprehensive two dimensional gas chromatography. TrAC Trends Anal Chem. 2019;113:379–91. https://doi.org/10.1016/j.trac.2018.04.016.
Tranchida PQ. Comprehensive two-dimensional gas chromatography: a perspective on processes of modulation. J Chromatogr A. 2018;1536:2–5. https://doi.org/10.1016/j.chroma.2017.04.039.
Higgins Keppler EA, Jenkins CL, Davis TJ, Bean HD. Advances in the application of comprehensive two-dimensional gas chromatography in metabolomics. TrAC Trends Anal Chem. 2018;109:275–86. https://doi.org/10.1016/j.trac.2018.10.015.
Kulsing C, Nolvachai Y, Marriott PJ. Concepts, selectivity options and experimental design approaches in multidimensional and comprehensive two-dimensional gas chromatography. TrAC Trends Anal Chem. 2020;130:115995. https://doi.org/10.1016/j.trac.2020.115995.
Stilo F, Bicchi C, Reichenbach SE, Cordero C. Comprehensive two-dimensional gas chromatography as a boosting technology in food-omic investigations. J Sep Sci. 2021;44:1592–611. https://doi.org/10.1002/jssc.202100017.
Hoh E, Lehotay SJ, Mastovska K, Huwe JK. Evaluation of automated direct sample introduction with comprehensive two-dimensional gas chromatography/time-of-flight mass spectrometry for the screening analysis of dioxins in fish oil. J Chromatogr A. 2008;1201:69–77. https://doi.org/10.1016/j.chroma.2008.05.089.
Tranchida PQ, Franchina FA, Dugo P, Mondello L. Flow-modulation low-pressure comprehensive two-dimensional gas chromatography. J Chromatogr A. 2014;1372:236–44. https://doi.org/10.1016/j.chroma.2014.10.097.
Hoh E, Lehotay SJ, Mastovska K, Ngo HL, Vetter W, Pangallo KC, Reddy CM. Capabilities of direct sample introduction−comprehensive two-dimensional gas chromatography−time-of-flight mass spectrometry to analyze organic chemicals of interest in fish oils. Environ Sci Technol. 2009;43:3240–7. https://doi.org/10.1021/es803486x.
Pomerantsev AL. Acceptance areas for multivariate classification derived by projection methods. J Chemom. 2008;22:601–9. https://doi.org/10.1002/cem.1147.
Ferreira VHC, Hantao LW, Poppi RJ. Use of color based chromatographic images obtained from comprehensive two-dimensional gas chromatography in authentication analyses. Talanta. 2021;234:122616. https://doi.org/10.1016/j.talanta.2021.122616.
Toledo BR, Hantao LW, Ho TD, Augusto F, Anderson JL. A chemometric approach toward the detection and quantification of coffee adulteration by solid-phase microextraction using polymeric ionic liquid sorbent coatings. J Chromatogr A. 2014;1346:1–7. https://doi.org/10.1016/j.chroma.2014.04.035.
Rodionova OY, Titova AV, Pomerantsev AL. Discriminant analysis is an inappropriate method of authentication. TrAC Trends Anal Chem. 2016;78:17–22. https://doi.org/10.1016/j.trac.2016.01.010.
Paiva AC, Simões Oliveira D, Hantao LW. A bottom-up approach for data mining in bioaromatization of beers using flow-modulated comprehensive two-dimensional gas chromatography/mass spectrometry. Separations. 2019;6:46. https://doi.org/10.3390/separations6040046.
Sinha AE, Prazen BJ, Synovec RE. Trends in chemometric analysis of comprehensive two-dimensional separations. Anal Bioanal Chem. 2004;378:1948–51. https://doi.org/10.1007/s00216-004-2503-7.
Feizi N, Hashemi-Nasab FS, Golpelichi F, Saburouh N, Parastar H. Recent trends in application of chemometric methods for GC-MS and GC×GC-MS-based metabolomic studies. TrAC Trends Anal Chem. 2021;138:116239. https://doi.org/10.1016/j.trac.2021.116239.
Stefanuto P-H, Smolinska A, Focant J-F. Advanced chemometric and data handling tools for GC×GC-TOF-MS. TrAC Trends Anal Chem. 2021;139:116251. https://doi.org/10.1016/j.trac.2021.116251.
Ballabio D, Grisoni F, Todeschini R. Multivariate comparison of classification performance measures. Chemom Intell Lab Syst. 2018;174:33–44. https://doi.org/10.1016/j.chemolab.2017.12.004.
de Geus H-J, Aidos I, de Boer J, Luten JB, Brinkman UAT. Characterisation of fatty acids in biological oil samples using comprehensive multidimensional gas chromatography. J Chromatogr A. 2001;910:95–103. https://doi.org/10.1016/S0021-9673(00)01183-3.
Bogusz S, Hantao LW, Braga SCGN, de Matos França V de CR, da Costa MF, Hamer RD, Ventura DF, Augusto F. Solid-phase microextraction combined with comprehensive two-dimensional gas chromatography for fatty acid profiling of cell wall phospholipids. J Sep Sci. 2012;35:2438–2444. https://doi.org/10.1002/jssc.201200256
Mogollon NGS, Ribeiro FA de L, Lopez MM, Hantao LW, Poppi RJ, Augusto F. Quantitative analysis of biodiesel in blends of biodiesel and conventional diesel by comprehensive two-dimensional gas chromatography and multivariate curve resolution. Anal Chim Acta. 2013;796:130–136. https://doi.org/10.1016/j.aca.2013.07.071
Griffith JF, Winniford WL, Sun K, Edam R, Luong JC. A reversed-flow differential flow modulator for comprehensive two-dimensional gas chromatography. J Chromatogr A. 2012;1226:116–23. https://doi.org/10.1016/j.chroma.2011.11.036.
Crucello J, Pierone DV, Hantao LW. Simple and cost-effective determination of polychlorinated biphenyls in insulating oils using an ionic liquid-based stationary phase and flow modulated comprehensive two-dimensional gas chromatography with electron capture detection. J Chromatogr A. 2020;1610:460530. https://doi.org/10.1016/j.chroma.2019.460530.
Facanali R, Marques MOM, Hantao LW. Metabolic profiling of Varronia curassavica Jacq. terpenoids by flow modulated two-dimensional gas chromatography coupled to mass spectrometry. Separations. 2020;7:18. https://doi.org/10.3390/separations7010018.
Moreira de Oliveira A, Alberto Teixeira C, Wang Hantao L. Evaluation of the retention profile in flow-modulated comprehensive two-dimensional gas chromatography and independent component analysis of weathered heavy oils. Microchem J. 2022;172:106978. https://doi.org/10.1016/j.microc.2021.106978.
Paiva AC, Hantao LW. Exploring a public database to evaluate consumer preference and aroma profile of lager beers by comprehensive two-dimensional gas chromatography and partial least squares regression discriminant analysis. J Chromatogr A. 2020;1630:461529. https://doi.org/10.1016/j.chroma.2020.461529.
Crucello J, Miron LFO, Ferreira VHC, Nan H, Marques MOM, Ritschel PS, Zanus MC, Anderson JL, Poppi RJ, Hantao LW. Characterization of the aroma profile of novel Brazilian wines by solid-phase microextraction using polymeric ionic liquid sorbent coatings. Anal Bioanal Chem. 2018;410:4749–62. https://doi.org/10.1007/s00216-018-1134-3.
Replication data and code for: Bor et al “Police killings and their spillover effects on mental health among Black Americans in the US.” https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/VGYI0E Accessed 28 Aug 2022.
van den Berg RA, Hoefsloot HC, Westerhuis JA, Smilde AK, van der Werf MJ. Centering, scaling, and transformations: improving the biological information content of metabolomics data. BMC Genomics. 2006;7:142. https://doi.org/10.1186/1471-2164-7-142.
Wold S. Pattern recognition by means of disjoint principal components models. Pattern Recognit. 1976;8:127–39. https://doi.org/10.1016/0031-3203(76)90014-5.
Pomerantsev AL, Rodionova OY. Concept and role of extreme objects in PCA/SIMCA. J Chemom. 2014;28:429–38. https://doi.org/10.1002/cem.2506.
Brereton RG, Lloyd GR. Partial least squares discriminant analysis: taking the magic away. J Chemom. 2014;28:213–25. https://doi.org/10.1002/cem.2609.
Barker M, Rayens W. Partial least squares for discrimination. J Chemom. 2003;17:166–73. https://doi.org/10.1002/cem.785.
Hantao LW, Aleme HG, Passador MM, Furtado EL, Ribeiro FA de L, Poppi RJ, Augusto F. Determination of disease biomarkers in Eucalyptus by comprehensive two-dimensional gas chromatography and multivariate data analysis. J Chromatogr A. 2013;1279:86–91. https://doi.org/10.1016/j.chroma.2013.01.013
Bean HD, Hill JE, Dimandja J-MD. Improving the quality of biomarker candidates in untargeted metabolomics via peak table-based alignment of comprehensive two-dimensional gas chromatography–mass spectrometry data. J Chromatogr A. 2015;1394:111–7. https://doi.org/10.1016/j.chroma.2015.03.001.
Ogundare SA, van Zyl WE. A review of cellulose-based substrates for SERS: fundamentals, design principles, applications. Cellulose. 2019;26:6489–528. https://doi.org/10.1007/s10570-019-02580-0.
Zontov YV, Rodionova OY, Kucheryavskiy SV, Pomerantsev AL. DD-SIMCA – A MATLAB GUI tool for data driven SIMCA approach. Chemom Intell Lab Syst. 2017;167:23–8. https://doi.org/10.1016/j.chemolab.2017.05.010.
Funding
This study was financed by ANP/PETROBRAS (Grant 2019/00209–3 and 2019/00210–1, Brazil), the National Council for Scientific and Technological Development (CNPq) (Grant 316202/2021–5), the São Paulo Research Foundation (FAPESP) (Grant 14/50867–3 and 20/01064–6), and the Coordination for the Improvement of Higher Education Personnel (CAPES) — Finance Code 001.
Author information
Authors and Affiliations
Contributions
Conceptualization, Leandro Wang Hantao; methodology, Carlos Alberto Teixeira, Rássius Alexandre Medeiro Lima, Sofia Madruga Marcondes Ferraz, and Victor Gustavo Kelis Cardoso; formal analysis and investigation, Carlos Alberto Teixeira, Rássius Alexandre Medeiro Lima, Sofia Madruga Marcondes Ferraz, and Victor Gustavo Kelis Cardoso; writing — original draft preparation, Carlos Alberto Teixeira, Rássius Alexandre Medeiro Lima, Sofia Madruga Marcondes Ferraz, and Victor Gustavo Kelis Cardoso; writing — review and editing, Carlos Alberto Teixeira, Leandro Wang Hantao, Rássius Alexandre Medeiro Lima, Sofia Madruga Marcondes Ferraz, and Victor Gustavo Kelis Cardoso; funding acquisition, Leandro Wang Hantao; and supervision, Leandro Wang Hantao.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Comprehensive 2D Chromatography with guest editors Peter Q. Tranchida and Luigi Mondello.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lima, R.A.M., Ferraz, S.M.M., Cardoso, V.G.K. et al. Authentication of fish oil (omega-3) supplements using class-oriented chemometrics and comprehensive two-dimensional gas chromatography coupled to mass spectrometry. Anal Bioanal Chem 415, 2601–2611 (2023). https://doi.org/10.1007/s00216-022-04428-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04428-2