Abstract
Hepatic cirrhosis is a severe chronic disease of the liver accompanied by massive changes in the physiology of the cells constituting the hepatic tissue. Success or failing of a therapeutic effort is difficult to recognize because of its late manifestation in the tissue morphology. In this study, the complex course of hepatic cirrhosis and its regression is followed in a rodent carbon tetrachloride model. Raman spectroscopy, which senses molecular vibrations and reflects the molecular composition of a sample, was applied to gain label- and destruction-free insights into the process of cirrhosis and to conclude on the hepatic disease state on the cellular level. Hematoxylin and eosin staining and immunofluorescence labeling were used to obtain complementary information. False color images derived from maps of Raman spectra by spectral unmixing revealed individual nucleus positions giving structural information. Spectral data unraveled chemical changes associated with liver damage on the cellular level. Upon carbon tetrachloride treatment, a higher lipid content and the presence of catabolic products indicated cirrhosis in tissue samples. Furthermore, 77 % of the Raman spectra recorded from treated rats were classified as diseased, whereas 96 % of the Raman spectra recorded from untreated rats were classified as healthy. Importantly, samples from rats that experienced a recovery period revealed a chemical composition highly similar to the ones from healthy rats while morphologically clear signs of tissue damage were still obvious. Eighty-three percent of their Raman spectra were classified as healthy. The vibrational fingerprint of tissue provides characteristic information that might serve as prognostic biomarker.

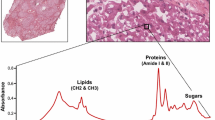
Images of hematoxylin and eosin stained tissue slices together with an average Raman spectrum belonging to healthy and cirrhotic rat liver, respectively, visualize morphological and spectral differences between the two states of the liver
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fibrotic diseases of the liver are a serious health concern [1]. If the liver is insulted, the cells within the liver may undergo apoptosis or become necrotic. Insults may be caused by chemicals and drugs such as alcohol or by viruses, especially hepatitis viruses, or autoimmune diseases. A dying cell releases a number of cytokines as alarm signals. These attract immune cells and induce a response in neighboring cells. If a lot of cell death occurs or if this happens chronically, an excess of extra cellular matrix (ECM) molecules is deposited at the sites of injury. It is known that ECM molecules originate mainly from activated fibroblasts. Yet, there is some controversy to what extend activated hematopoietic stem cells (HSCs) and immigrated fibroblasts are involved in hepatic fibrosis [2–4]. Macrophages seem to play an important role in the regulation of the fibrotic progression and regression [5]. The deposition of an excess of ECM molecules leads to scarring and clinically manifests in hepatic fibrosis. In an advanced stage, especially if vessels are interconnected by scars, the disease pattern is called cirrhosis. Different pathology scoring systems exist to classify hepatic tissues in healthy, fibrotic, and cirrhotic tissue based on a histologic assessment [6]. In the long term, hepatic cirrhosis also raises the risk for the development of hepatocellular carcinoma [7]. Several options for the treatment of hepatic fibrotic diseases exist. However, their success or failure is extremely individual. Thus, a strategy for therapeutic monitoring would be highly desirable [1, 8].
In the research field of digital pathology, aiming at the assistance of the pathologist in the daily routine, some effort is made to automate tissue classification in healthy and diseased liver. Therefore, images are taken from stained tissue samples. Virtual slides are constructed. The approach then mostly relies on image analysis. Thus, the work of the pathologist is imitated by a camera and algorithms. They are needed to segment the image, e.g., to identify the nuclei, and to classify the samples according to the disease state [9, 10]. The sample preparation requirements preceding the digital pathology assessment remain the same as in the classic approach as does the information accessible from the sample. Especially, the sample definitely has to be removed from the organism.
Via Raman spectroscopy chemical information from a tissue sample is collected in a different way. Laser light is focused onto the sample to excite molecular vibrations. A portion of the excitation light is inelastically scattered by the molecules in the sample. This Raman scattered light is detected and informs about the chemical composition of the sample. Raman data can be acquired from unstained samples. This minimizes the preparation effort [11, 12]. Often, the approach is referred to as spectral histopathology [11, 13]. It is possible to generate images from the Raman information [14, 15]. In addition, several studies report on the discrimination, especially of healthy and cancerous tissue based on Raman data [13, 16] or the disease-related Raman spectroscopic analysis of blood samples [17, 18]. Raman spectroscopy has also been applied in vivo [19].
The present study was based on the hypothesis that Raman spectra would as well reflect the hepatic health state with respect to cirrhosis and regression. In order to test this, Raman-based imaging and statistical analysis of the Raman data were performed. The Raman-based images were combined and compared with immunofluorescence images from the same sample site. The Raman data were found to distinguish health/recovery and ongoing cirrhosis.
Material and methods
Animal experiments and sample acquisition
Male Wistar rats (Wistar RccHan:WIST; Harlan Laboratories Inc., Indianapolis, IN, USA) were used in all animal experiments. Rats were kept under an artificial day-night cycle (12 h light-dark cycles; 23 °C room temperature; 30–60 % relative humidity) and received standard diet with water ad libitum. They were allowed to adapt to laboratory conditions for at least 3 d. All procedures involving animals and their care were performed with permission of the regional animal welfare committee of Thuringia (permit number: TVA Reg. No. 02-062/11) as well as in accordance with German legislation on protection of animals, with UK Home Office guidelines under the 1986 Scientific Procedures Act and with US National Institutes of Health guidelines.
To investigate hepatic cirrhosis, the rodent carbon tetrachloride (CCl4) model, which reflects a typical course of chronic liver damage, was used.
In the CCl4 treatment group (four rats), animals received subcutaneous CCl4 injections (0.5 mL per kilo body weight, dissolved in olive oil 2:3 v/m, this is equivalent to 0.75 mL of a 7 M mixture of CCl4 in olive oil per kilo body weight) into the nuchal fold twice a week for 9 wk. Two animals were sacrificed immediately after the CCl4 treatment period, while the two remaining animals were allowed to recover from CCl4 treatment for 2 wk For comparison with the healthy state, livers from two untreated control animals were included.
All rats were sacrificed during isoflurane inhalation. Afterwards, the livers were sampled; parts of the liver lobes were sectioned, transferred into cryo-vials, and immediately frozen in liquid nitrogen. Samples were stored at –80 °C.
Fabrication of cryo-slices
Cryo-slices of the liver samples were cut in a cryostat (Leica CM3050 S; Leica Biosystems GmbH, Nussloch, Germany) at a temperature range between –19 °C and –23 °C. For slicing, the samples were stuck onto a plate with Tissue Tek (Sakura Finetek Germany GmbH, Staufen, Germany). Tissue slices for Raman measurements and subsequent immunofluorescence staining were cut from tissue parts free of Tissue Tek at a thickness of 10 μm and transferred onto CaF2 slides (Crystal GmbH, Berlin, Germany). Three slices were cut per liver for Raman measurements. Tissue slices for hematoxylin and eosin (H&E) staining and immunofluorescence labeling of the combination CD68 and CD19 were cut at a thickness of 5 μm and transferred onto standard glass slides (SuperFrost; Gerhard Menzel GmbH, Braunschweig, Germany).
Acquisition of Raman data
Raman spectra were acquired by a micro-Raman spectrometer (alpha 300; Witec, GmbH, Ulm, Germany) operating in the continuous mode for acquisition of the spectra. Excitation light was generated by a 532 nm frequency-doubled Nd:YAG laser. The laser light was coupled into an upright microscope (Carl Zeiss, AG, Oberkochen, Germany) by a fiber and focused on the sample by an EC Epiplan-Neofluar 100x LD objective with NA 0.75 (Carl Zeiss AG, Oberkochen, Germany). Laser power before the objective was set to 15 mW. The same objective and a 50 μm fiber served for the collection and transport of backscattered light from the sample. The light was dispersed on a 300 lines per mm grating before hitting on a back-illuminated CCD (DV401A-BV-352; Andor Technology Ltd., Belfast, United Kingdom) cooled to –60 °C. The spectrometer was operated using Witec software (Witec Control 1.5, Witec, GmbH, Ulm, Germany).
Spectra for classification and Raman imaging were collected in mapping mode with a step size of 0.5 μm and an acquisition time of 1 s per spectrum. One map had a dimension of 15 μm × 15 μm. Three maps were acquired per tissue slice. Three tissue slices were included per liver. Thus, nine maps were acquired per liver, yielding 8100 spectra per animal. Mapping positions were chosen randomly on the slices, however, avoiding severe scars, holes or large vessels. Before acquisition of the Raman maps, a mark for later image correlation was introduced into the tissue using maximum laser power (35 mW before the objective).
Raman imaging and statistical analysis of Raman data
In order to derive false color Raman images, the spectra maps were analyzed with the N-FINDR algorithm [20] in MatLab (The MathWorks GmbH, Ismaning, Germany). Prior, the spectra received a pretreatment including: truncation of the wavenumber axis keeping the wavenumbers from 600 to 1800 cm–1, spike removal by MatLab’s cosmic ray removal function and PCA-based noise reduction [20] keeping the first 10 components. The N-FINDR algorithm included a normalization based on extended multiplicative signal correction [21]. As input for the N-FINDR algorithm a number of three endmembers was used. The average spectra for each endmember were calculated from the raw spectra. A spectrum was included for further calculation if the relevant endmember contributed to it by at least 75 %.
For spectral characterization of the tissue samples, a principal component analysis (PCA) with subsequent linear discriminant analysis (LDA) was chosen. The PCA is a common, unsupervised chemometric method employed as a preprocessing step to reduce data dimension and separate high variance information in the data set from general noise variation in the data set. The subsequently carried out LDA is a supervised algorithm that searches for a separating hyperplane that maximizes the inter-class distance while keeping the intra-class distances minimal. Thus, the data are finally arranged according to predefined classes, revealing similarity or difference [22].
In order to perform PCA-LDA, spectra maps were loaded into GNU R [23]. All work in R was done in the hyperSpec [24] environment. The spectra received a pretreatment including: truncation of the wavenumber axis to 500–1800 cm–1 and 2800–3050 cm–1, polynomial baseline subtraction using a second order polynomial, vector normalization, spike removal, and removal of spectra from burned sites that only contain carbon signals. The burning events occurred probably because of the occasional presence of erythrocytes in the tissue. Erythrocytes are easily destroyed from 532 nm laser light as this wavelength is resonant with the Q-band of hemoglobin. The liver is well supplied with blood and erythrocytes visible in the H&E images (Fig. 1a–c).
Images from hematoxylin and eosin stained tissue slices, from healthy control rats (a) and (b), CCl4-treated rats without recovery (c) and (d), and CCl4-treated rats with 2-wk recovery (e) and (f). For each rat included in the study, one image is depicted. The arrow heads in (a), (b), and (c) point at erythrocytes. Note also the prominent vesicles in the hepatocytes, especially in (c) and (d), which indicate steatosis. Nuclei are hematoxylin-stained and are visible in blue, cytoplasm is eosin-stained and is visible in red. The scale bar marks 20 μm and is valid for all images
In the first step of PCA-LDA, models were established from a number of training data according to the classification tasks (“healthy” versus “diseased” and “healthy” versus “recovered”). Afterwards, these models were employed to assign the class of independent test data without any additional information from the test data sets.
PCA-LDA was run independently for each classification task in a leave-one-animal-out manner to assure biological independence of the test data set. For the training data, nearly equal numbers of spectra regarding both classes were applied. The first five components from PCA were kept for LDA. They described at least 98 % of the variances of the respective datasets. The values given in Fig. 4 are averages of the respective classification task using repeated leave-one-out calculations. One representative loading for each of the two classification tasks is shown in Fig. 5.
H&E staining and image acquisition
The tissue slices were fixed to the glass slides by subsequent incubations with methanol (Carl Roth GmbH & Co. KG, Karlsruhe, Germany) and acetone (Carl Roth GmbH & Co. KG, Karlsruhe, Germany). Afterwards the slices were incubated in hemalum solution (Mayer’s recipe) and blued in tab water. Thereafter, they were incubated in Eosin Y solution (Sigma-Aldrich Chemie GmbH, Munich, Germany). The dehydration was done in an ascending ethanol (Nordbrand Nordhausen GmbH, Nordhausen, Germany) line. Subsequently, the slices were incubated in xylol (Carl Roth GmbH & Co. KG, Karlsruhe, Germany). Finally, they were embedded in Pertex (Medite GmbH, Burgdorf, Germany). Images were acquired with a Zeiss Axio Vert.A1 microscope (Carl Zeiss AG Oberkochen, Germany) equipped with a LED illumination light source and a LD A-Plan 20x/0.35 Ph1 objective (Carl Zeiss AG Oberkochen, Germany).
Immunofluorescence labeling and image acquisition
The tissue slices were fixed to the CaF2 slides or glass slides by acetone (Carl Roth GmbH & Co. KG, Karlsruhe, Germany) and permeabilized with Tween-20 in PBS (Carl Roth GmbH & Co. KG, Karlsruhe, Germany). The labeling procedure started with a blocking step with blocking reagent (Life Technologies GmbH, Darmstadt, Germany). For the double- and triple-staining, consecutive two-step labeling was performed. Each two-step procedure relied on a primary antibody recognizing the antigen of interest and a fluorophore-coupled secondary antibody recognizing the constant domain of the primary antibody. The following pairs were employed: rabbit-anti-glial fibrillary acidic protein (GFAP) 1:200 (Abcam plc, Cambridge, UK) together with goat-anti-rabbit-fluorescinisothiocyanat 1:400 (Dianova GmbH, Hamburg, Germany), mouse-anti- cluster of differentiation 68 (CD68) 1:100 (Bio-Rad AbD Serotec GmbH, Puchheim, Germany) together with donkey-anti-mouse-Alexa647 1:400 (Abcam plc, Cambridge, UK), chicken-anti-E-cadherin 1:50 (Abcam plc, Cambridge, UK) together with donkey-anti-chicken-Cy3 (Dianova GmbH, Hamburg, Germany) 1:400, rabbit-anti-cluster of differentiation 19 (CD19) 1:100 (Cell Signaling Technology Inc., Danvers, MA, USA) together with goat-anti-rabbit-fluorescinisothiocyanat 1:400 (Dianova GmbH, Hamburg, Germany). Antibody dilutions were prepared in blocking reagent (Life Technologies GmbH, Darmstadt, Germany). Between the incubation steps, the samples were rinsed in PBS (Biochrom AG, Berlin, Germany) containing 0.1 % Tween-20. Finally, the samples were embedded in anti-fading medium with 4',6-Diamidino-2-Phenylindol (DAPI) (Vectashield, Linaris GmbH, Dossenheim, Germany).
Fluorescence images were acquired using a laser scanning microscope (LSM 510 META, Carl Zeiss AG, Oberkochen, Germany). The following lasers, beam splitters, and filters were used for fluorescence excitation and emission detection: DAPI: laser diode 405 nm, HFT 405/514, BP 420–480, FITC: Argon laser 488 nm, HFT 488, NFT 490, BP 505–550, Cy3: HeNe laser 543 nm, HFT 405/488/543, NFT 490, BP 560–615, Alexa647: HeNe laser 633 nm, HFT UV/488/543/633.
Results
Histology
The liver samples involved in the study were characterized by visual inspection of H&E-stained tissue slices. In Fig. 1 corresponding images are depicted. In the healthy samples (Fig. 1a, b) the orderly arranged liver parenchyma is visible, vessels with smooth borders and clear separation from the surrounding parenchyma are depicted. In livers from animals treated with CCl4 (both with and without recovery) the well-arranged hepatocyte assemblage is destroyed (Fig. 1c–f). Cells are separated by long and thick scars and circular gaps. The scars often lead from one vessel to another (not shown). A number of infiltrates of immune cells is visible in the samples from animals that were treated with CCl4 without a recovery period. Accordingly, increased numbers of CD19+-B-lymphocytes and CD68+-macrophages are visible in Fig. 2c–f compared with Fig. 2a, b. Vesicles in the cytoplasm of the hepatocytes indicating steatosis are striking (Fig. 1c, d). In all samples originating from CCl4-treated rats (Fig. 1c–f) signs of cirrhosis are evident. However, steatosis seems to be regressive in animals with a recovery period (Fig. 1e, f).
Images from immunofluorescently stained tissue slices from healthy control rats (a) and (b), CCl4-treated rats without recovery (c) and (d), and CCl4-treated rats with 2 wk recovery (e) and (f). For each rat included in the study one image is depicted. Nuclei are stained with DAPI and are visible in blue, CD68+-cells (macrophages) are visible in red, CD19+-cells (B-Lymphocytes) are visible in green. First, the immune cells are identified in liver samples of rats treated with CCl4. However, a number of liver-dwelling macrophages is present in the samples of healthy rats as well. The scale bar marks 50 μm and is valid for all images
Immunofluorescence
Immunofluorescence assessments allowed for the visualization of several cell types of the liver depending on the antibodies employed, i.e., hepatocytes (E-cadherin positive), macrophages (CD68 positive), and HSCs (GFAP positive) could be identified within the same sample (Fig. 3b).
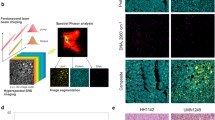
Combination of immunofluorescence and Raman imaging. (a) Superposition of immunofluorescence and false color N-FINDR Raman image (smaller inset) of a liver section from a healthy animal. White lines indicate the laser label used to overlay both images. (b) Side-by-side comparison of immunofluorescence (left) and false color N-FINDR Raman image (right) of a liver section from a rat treated with CCl4 and allowed to recover for 2 wk. The white square indicates the region of Raman measurement. Both figures (a) and (b) demonstrate that information on the cellular distribution as visualized by cell nuclei can be obtained from immunofluorescence images as well as from false color Raman images; (c) and (d) the Raman spectra contain information on the chemical composition of the tissue. In order to avoid the overlap of the spectra and to facilitate the recognizability of all features, the spectra are shifted along the intensity axis. The scale bar marks 50 CCD counts. The average Raman spectrum in (c) belongs to the N-FINDR image in (a) and the average Raman spectrum in (d) belongs to the N-FINDR image in (b). The colors chosen for the spectra correspond to the colors in the respective N-FINDR image. Cell type assignments are possible from immunofluorescence labeling (a) and (b): GFAP labels hepatic stellate cells (green), E-cadherin labels hepatocytes (red), CD68 labels macrophages (violet). Cell nuclei are stained with DAPI (blue), the scale bar marks 10 μm
Label-free imaging using Raman spectroscopy and N-FINDR
False color images could be obtained from the liver tissue without the need of any label using Raman spectroscopic mapping and a subsequent unmixing algorithm. The N-FINDR algorithm defines each Raman spectrum to be a mixture of a number of pure spectra (endmembers). Because each of the endmembers is represented by a specific color, each spectrum in the Raman false color image is represented by a mixed color according to the endmember contributions in the spectrum [20]. Thus, gradual changes in the chemical composition of a sample can be visualized in the Raman false color images.
Three endmembers constitute the Raman false color images in Fig. 3a (healthy sample) and b (CCl4-treated sample with 2 wk recovery). Especially, cell nuclei can be distinguished in the Raman false color images. In the corresponding average spectra (Fig. 3c, d) features of nucleic acid [25] appear at 788 cm–1 (PO2 –) and 1092 cm–1 (PO2 –). The remaining average spectra corresponding to the two other endmembers show mainly contributions from proteins and lipids [25, 26]. All spectra are dominated by cytochrome contributions [27] at 750, 1130, and 1586 cm–1.
Correspondence of immunofluorescence and Raman-based images
The recording of the Raman data preceded the immunofluorescence staining and image acquisition procedures. With the help of a laser induced mark (Fig. 4), the positions of the maps of Raman spectra could be assigned in the immunofluorescence preparations. Nuclei positions are accessible from both methods and show very good agreement when comparing the images (Fig. 3a, b). Importantly, in general, the immunofluorescence preparations did not show any signs of damage to the tissue that could have been introduced by the excitation of the Raman process (Figs. 3b and 4).
Immunofluorescence images taken from a site where Raman measurements were done previously. The nuclei are stained with DAPI (blue), CD68 labels macrophages (violet), and GFAP labels hepatic stellate cells (green); the scale bar marks 20 μm. The arrow points at the laser-induced mark, which was introduced for re-identification of the site. The white squares indicate the Raman mapping positions
Spectral tissue characterization using Raman data and PCA-LDA
In order to assess how the morphologic information accessible from H&E staining is reflected in the chemical information contained in the Raman spectra, slices of the livers characterized in Fig. 1 were subjected to Raman spectroscopic measurements. The excitation of the Raman scattering with 532 nm was in resonance with an absorption of cytochromes. Within the Raman spectra, features of cytochromes are obvious (Figs. 3 and 5b, Electronic Supplementary Material (ESM) Fig. S1). They vanished during prolonged excitation at the same spot (ESM Fig. S1). The Raman spectra were further analyzed by PCA-LDA. The analysis revealed two obviously distinct groups of spectra. On the one hand, there were those originating from the untreated animals [Fig. 5b (1)], on the other hand, those of the CCl4-treated animals without a recovery period [Fig. 5b (2)]. With an automated assignment of biological independent repeat measurements (different animals), 96 % of the spectra originating from the untreated animals ended up in the class “healthy,” whereas 77 % of the spectra from CCl4-treated animals without a recovery period were grouped into the class “diseased” (Fig. 5a).
Spectral characterization of liver samples (a) schematic representation of the results obtained by PCA-LDA, and (b) average spectra from 200 single Raman spectra of each class [(1) – untreated, (2) – CCl4-treated without recovery, (3) – CCl4-treated with 2-wk recovery]. In order to compare Raman data reflecting different hepatic health states, as a first step a PCA-LDA model was established discriminating “healthy” and “diseased.” It was applied on biologically and statistically independent data sets from differently treated animals. Percentages in (a) represent the relative number of Raman spectra of the liver tissue slices assigned to one of the LDA classes. The spectra in (b) are baseline-corrected and shifted along the intensity axis. The scale bar marks 50 CCD counts
In order to visualize the spectral features that were responsible for the discrimination between “healthy” and “diseased,” a representative LDA coefficient is displayed in Fig. 6a along with a scatter plot in Fig. 6c visualizing the projection of independent test spectra into its prediction model. The prominent bands in the coefficient point towards those molecular vibrations that influenced the discrimination most. Bands scoring negative in the coefficient contributed more in the untreated samples (“healthy”). Bands scoring positive in the coefficient contributed more in the CCl4-treated samples without a recovery period (“diseased”). For the discrimination of “healthy” and “diseased”, especially vibrational contributions around 1660 cm–1 (C=C), around 1450 cm–1 (CH2), and the band from 1200 to 1300 cm-1 (=CH, CH2) were of relevance and can be assigned to lipids [26]. As those vibrational contributions are positive in the coefficient, the lipid content is higher in the diseased livers compared with the untreated ones. In the scatter plot, each spectrum is represented by a dot. Spectra scoring negative were predicted as “healthy,” spectra scoring positive were predicted as “diseased.”
DA coefficients and scatter plots. (a) and (b) LDA coefficients visualizing features of the Raman spectra that are important for the discrimination between two data sets; (c) and (d) prediction of test data from corresponding LDA models, each test spectrum is reflected by a dot in the scatter plot (black – untreated, red – treated)
The spectra originating from CCl4-treated animals with 2 wk recovery were grouped into the class “healthy” in their majority (83 %) when forced into one of the classes, “healthy” or “diseased” (Fig. 5a). Characteristic features in the spectra originating from CCl4-treated animals with recovery, which would have allowed their discrimination from the spectra originating from the untreated animals, could not be identified. The average spectra resemble each other [Fig. 5b (1) and (3)]. Comparing “healthy” against “recovered”, 85 % of the spectra originating from the untreated animals and 71 % of the spectra originating from CCl4-treated animals with recovery were grouped into the class “healthy.”
The LDA coefficient corresponding to the discrimination of tissue from untreated rats and from rats with CCl4-treatment and a subsequent recovery period (Fig. 6b) contains a lot of noise. In the corresponding scatter plot (Fig. 6d) the spectra originating from CCl4-treated animals with recovery score highly variable.
The detailed confusion tables for the classification of the test data are depicted in Table S1 in the ESM.
Discussion
In the present study, it is shown that Raman data in combination with chemometric analysis are suitable to characterize the state of health of liver tissue with regard to cirrhotic alteration.
The Raman data were acquired after an excitation with 532 nm laser light. This is in resonance with an absorption of cytochromes. Especially hepatocytes are known to contain high amounts of cytochromes due to their important role in the detoxification of xenobiotics [28]. Pronounced contributions of cytochrome vibrations were recognizable in the Raman spectra regardless of the treatment of the rats (Fig. 3). However, cytochromes degrade quickly under the applied conditions of measurement (ESM Fig. S1). This needs to be taken into account for the experimental design. Herein, the time for the acquisition of a single spectrum was 1 s, the step size 1 μm, and the laser spot size can be calculated as 865 nm. Thus, the effect of photo-degradation seems tolerable regarding a stable appearance of the cytochrome band at about 1585 cm–1 for tens of seconds of continuous irradiation (ESM Fig. S1b).
A spectral unmixing algorithm (N-FINDR) was found to be suitable to gain imaging information from the Raman data. Especially, the location of cell nuclei in the tissue slices (N-FINDR endmember depicted in blue in Fig. 3a, b) could be revealed by the N-FINDR analysis. In addition, differences in the overall chemical composition of the samples were reflected by the occurrence of patterns of the “yellow” and the “green” N-FINDR endmembers (Fig. 3a, b). The corresponding Raman spectra (Fig. 3c, d) unraveled the presence of high amounts of cytochrome in hepatocytes. Generally, comprehensive information is gained via Raman microspectroscopy and it is unnecessary to choose a small number of molecules for the investigation. Nevertheless, the features that were found in the Raman false color images corresponded very well with the ones in the immunofluorescence images that were recorded from the same site (Fig. 3a, b).
Statistical analysis of the Raman data from the tissue samples by PCA-LDA allowed for the discrimination of healthy and diseased liver tissue. In order to assess how the morphologic information accessible from H&E staining was reflected in the chemical information contained in the Raman spectra, slices of the livers characterized in Fig. 1 were subjected to Raman spectroscopic measurements. The statistical approach was employed to gain access to that information hidden in the bulk Raman data, which indeed is meaningful for the state of health of the tissue. This particular information usually is not accessible in an intuitive way from a look at the Raman spectra because the spectra are as complex as the sample is chemically complex.
As expected, the livers sampled from CCl4-treated rats without a recovery period were very different from those livers sampled from untreated rats according to the PCA-LDA analysis. In contrast, the livers sampled from CCl4-treated rats with a 2 wk recovery period were very similar to those livers sampled from untreated rats and different from those of CCl4-treated rats without a recovery period in the PCA-LDA (Fig. 5).
The CCl4-intoxication causes massive cell damage and cell death. The response towards this includes the release of plenty of cytokines and growth factors [6]. The cells, also those cells that are situated away from the fibrotic scars, are transferred in an alarm condition. Thus, their transcription rate and metabolism changes dramatically. Steatosis develops, as is also visible from the H&E preparations (Fig. 1c, d). Such changes are detected by Raman spectroscopy and they can be the reason for the differences between the tissue experiencing ongoing chronic CCl4-intoxication and healthy tissue in the PCA-LDA.
Accordingly, the LDA coefficient in Fig. 6a points first of all towards a higher lipid content in the livers of CCl4-treated rats without a recovery period in comparison with the livers of untreated rats. This is in good agreement with the steatosis visible in the H&E images (Fig. 1c, d). In addition, the coefficient shows a band at 1598 cm–1 that might correspond to a catabolic or an oxidation product of retinol. Hepatic stellate cells store retinyl esters in high amounts. A hallmark of chronic liver injury is hepatic stellate cell proliferation and activation [29]. Hepatic stellate cell proliferation might be the reason for a gradually higher content of retinoids in the diseased samples. However, hepatic stellate cells are known to lose retinol in the course of activation [30]. Thus, the presence and detection of retinol catabolites is more likely than the presence and detection of retinyl esters in the diseased samples.
Of course, not all the cells in a tissue are affected in the same magnitude by a toxin. A considerable number of the Raman spectra recorded from the CCl4-treated rats without a recovery period were classified as “healthy.” This fact might be explained by the occurrence of less severely affected cells.
The suspicion of physiologically differing cellular states being reflected in the Raman spectra is supported by the scatter plots in Fig. 6c, d. In general, the spectra originating from CCl4-treated rats score a lot more variable in the scatter plots than the spectra originating from untreated animals. This is especially true for a few spectra derived from rats experiencing a recovery period. As soon as the cause of damage (herein: CCl4) is omitted, the liver starts to recover. Its unrivaled potential of self-renewal allows for a compensation of two-thirds of its weight. If the fibrotic state was not passed, the regression process leads to the disappearance of the scars and the parenchyma can recover completely. If the cirrhotic state was already achieved, the scars stay but the remaining parenchyma recovers its function [6]. During the time of recovery, the cells are metabolically and mitotically active. Thus, their composition varies and, along with it, the spectral fingerprint.
The majority of the spectra of CCl4-treated rats with a 2 wk recovery period were classified as “healthy” in the PCA-LDA. This means their cell physiology must have changed back from the pathologic to a physiological state. A considerable recovery can as well be concluded from the H&E images (Fig. 1e, f compared with Fig. 1c, d). However, there are still scars visible in those images and conclusions on the state of health of the cells between the scars are difficult to make from the H&E stains.
Obviously, the recovery had proceeded so far that a clearly separate class “recovered” next to the class “healthy” could not be established by PCA-LDA for the dataset analyzed herein. In accordance, the LDA coefficient only showed a high noise level for the classification task “healthy” versus “recovered” (Fig. 6b). Owing to the low signal-to-noise-ratio, the spectral features cannot be interpreted, but might result from variations between different animals.
The present study provides evidence that the statistical analysis of the Raman spectra reveals the (patho-)physiological state of the cells constituting the hepatic tissue. Regarding the difference of the healthy state from the chronic pathologic state maintained by ongoing intoxication, the Raman results agree with the conclusions of a visual inspection of H&E stained tissue slices. Concerning the advanced phase of recovery, the Raman data, which were recorded from parts of the tissue away from scars, reflect the healthy cell physiology rather than the overall still pathologic tissue morphology obvious from the H&E stained samples. First of all, this highlights the complementarity of vibrational spectroscopy and histochemistry in the assessment of tissue. Furthermore, a shift in the cell physiology towards health necessarily precedes the manifestation of recovery in the morphology of the tissue. Thus, the early recognition of the recovery of the liver from fibrotic diseases is an important advantage of the Raman-based approach for liver assessment applied herein.
Yet, the approach was only tested regarding the CCl4 model of hepatic cirrhosis so far, meaning for a toxin-driven variant of cirrhosis. However, there are other relevant inducers of hepatic cirrhosis (e.g., viruses and alterations of the bile duct [1, 8]). The differences in the etiology might influence the results of the Raman spectroscopy-based assessment. Another limitation of the present study is the low number of biological replicates included. A higher number of independent replicates might have allowed to reveal the rather subtle chemical differences and, thereby, to distinguish between healthy and recovered tissue as well [31]. This possibility is likely since even the assignment of specific cell types in bulk tissue can be achieved by Raman spectroscopy if characteristic chemical differences exist and a sufficient amount of reference data (like a fingerprint database) is available, as described in a previous study for the identification of HSCs in fresh tissue [32]. However, such an analysis was beyond the scope of the present study.
Detailed information about the underlying molecular biological cause of differences between well distinguishable health states of the tissue, especially about the presence of specific proteins, is hardly accessible by Raman spectroscopy. It requires other techniques, such as immunolabeling. In immunofluorescence images specific cell types can be distinguished easily (Fig. 3b), and even protein subtypes or splicing variants can be recognized [33].
Therefore, ideally, Raman microspectroscopy and immunolabeling are employed in combination benefiting from advantages of both. Since Raman spectroscopy generally works in a nondestructive way, the sample is suited for immunofluorescence application after Raman analysis. The introduction of a mark into the sample by the Raman excitation laser allows for the recognition of the same site in the fluorescent sample later on as illustrated herein (Figs. 3a and 4). While the Raman false color images presented in this contribution cover only a rather small area, in several recent publications Raman-based images of more extended tissue parts have been shown [13, 34].
In addition, Raman data have been the basis for health-oriented classification tasks in several studies. First of all, these studies concentrated on cancerous tissue [35] or the identification of bacteria and their resistance against antibiotics [36, 37]. Conversely, the present study addresses a clinical field, fibrotic diseases of the liver, in which differences between classes (healthy versus diseased) are not difficult to distinguish by the naked eye. Yet, the liver is an inner organ and, thus, generally invisible for the naked eye. Furthermore, considerable time is required from the onset of recovery from hepatic injury until its manifestation in hepatic morphology. An early prognostic marker for this recovery would be of high relevance to govern hepatic anti-fibrotic therapy [1, 8]. Raman spectroscopy has been applied to study freshly sampled, unprocessed liver lobes ex vivo [38]. In addition, it is in principle applicable in vivo [39]. Hence, it might even be a future option to evaluate the effect of anti-fibrotic treatment on the course of disease by minimally invasive examination, thereby omitting the inherent risks of taking biopsies of liver tissue and decisively meeting the need for the surveillance of anti-fibrotic therapy.
References
Lee YA, Wallace MC, Friedman SL. Pathobiology of liver fibrosis: a translational success story. Gut. 2015;64(5):830–41.
Desmouliere A. Hepatic stellate cells: the only cells involved in liver fibrogenesis? A dogma challenged. Gastroenterology. 2007;132(5):2059–62.
Iwaisako K, Jiang C, Zhang M, Cong M, Moore-Morris TJ, Park TJ, et al. Origin of myofibroblasts in the fibrotic liver in mice. Proc Natl Acad Sci U S A. 2014;111(32):E3297–305.
Mederacke I, Hsu CC, Troeger JS, Huebener P, Mu X, Dapito DH, et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat Commun. 2013;4:2823.
Wynn TA, Barron L. Macrophages: master regulators of inflammation and fibrosis. Semin Liver Dis. 2010;30(3):245–57.
Bedossa P, Paradis V. Liver extracellular matrix in health and disease. J Pathol. 2003;200(4):504–15.
Zhang DY, Friedman SL. Fibrosis-dependent mechanisms of hepatocarcinogenesis. Hepatology. 2012;56(2):769–75.
Trautwein C, Friedman SL, Schuppan D, Pinzani M. Hepatic fibrosis: concept to treatment. J Hepatol. 2015;62(1 Suppl):S15–24.
Kayser K, Gortler J, Bogovac M, Bogovac A, Goldmann T, Vollmer E, et al. AI (artificial intelligence) in histopathology—from image analysis to automated diagnosis. Folia Histochem Cytobiol. 2009;47(3):355–61.
Schochlin M, Weissinger SE, Brandes AR, Herrmann M, Moller P, Lennerz JK. A nuclear circularity-based classifier for diagnostic distinction of desmoplastic from spindle cell melanoma in digitized histological images. J Pathol Inform. 2014;5:40.
Diem M, Mazur A, Lenau K, Schubert J, Bird B, Miljkovic M, et al. Molecular pathology via IR and Raman spectral imaging. J Biophotonics. 2013;6(11/12):855–86.
Galler K, Brautigam K, Grosse C, Popp J, Neugebauer U. Making a big thing of a small cell—recent advances in single cell analysis. Analyst. 2014;139(6):1237–73.
Mavarani L, Petersen D, El-Mashtoly SF, Mosig A, Tannapfel A, Kotting C. Spectral histopathology of colon cancer tissue sections by Raman imaging with 532 nm excitation provides label-free annotation of lymphocytes, erythrocytes, and proliferating nuclei of cancer cells. Analyst. 2013;138(14):4035–9.
Lattermann A, Matthaus C, Bergner N, Beleites C, Romeike BF, Krafft C, et al. Characterization of atherosclerotic plaque depositions by Raman and FTIR imaging. J Biophotonics. 2013;6(1):110–21.
Okada M, Smith NI, Palonpon AF, Endo H, Kawata S, Sodeoka M, et al. Label-free Raman observation of cytochrome c dynamics during apoptosis. Proc Natl Acad Sci U S A. 2012;09(1):28–32.
Wang W, Zhao J, Short M, Zeng H. Real-time in vivo cancer diagnosis using Raman spectroscopy. J Biophotonics. 2015;8(7):527–45.
Neugebauer U, Trenkmann S, Bocklitz T, Schmerler D, Kiehntopf M, Popp J. Fast differentiation of SIRS and sepsis from blood plasma of ICU patients using Raman spectroscopy. J Biophotonics. 2014;7(3/4):232–40.
Ryzhikova E, Kazakov O, Halamkova L, Celmins D, Malone P, Molho E, et al. Raman spectroscopy of blood serum for Alzheimer's disease diagnostics: specificity relative to other types of dementia. J Biophotonics. 2014. doi:10.1002/jbio.201400060.
Bohndiek SE, Wagadarikar A, Zavaleta CL, Van de Sompel D, Garai E, Jokerst JV, et al. A small animal Raman instrument for rapid, wide-area, spectroscopic imaging. Proc Natl Acad Sci U S A. 2013;110(30):12408–13.
Hedegaard M, Matthaus C, Hassing S, Krafft C, Diem M, Popp J. Spectral unmixing and clustering algorithms for assessment of single cells by Raman microscopic imaging. Theor Chem Accounts. 2011;130(4/6):1249–360.
Afseth NK, Kohler A. Extended multiplicative signal correction in vibrational spectroscopy, a tutorial. Chemometr Intell Lab. 2012;117:92–9.
Krafft C, Steiner G, Beleites C, Salzer R. Disease recognition by infrared and Raman spectroscopy. J Biophotonics. 2009;2(1/2):13–28.
Core Team R. R: a language and environment for statistical computing. Austria: R Foundation for Statistical Computing; 2014.
Beleites C, Sergo V (2014) HyperSpec: a package to handle hyperspectral data sets in R.
Notingher I, Green C, Dyer C, Perkins E, Hopkins N, Lindsay C, et al. Discrimination between ricin and sulphur mustard toxicity in vitro using Raman spectroscopy. J R Soc Interface. 2004;1(1):79–90.
Chan JW, Motton D, Rutledge JC, Keim NL, Huser T. Raman spectroscopic analysis of biochemical changes in individual triglyceride-rich lipoproteins in the pre- and postprandial state. Anal Chem. 2005;77(18):5870–6.
Ogawa M, Harada Y, Yamaoka Y, Fujita K, Yaku H, Takamatsu T. Label-free biochemical imaging of heart tissue with high-speed spontaneous Raman microscopy. Biochem Biophys Res. 2009;382(2):370–4.
Pelkonen O, Turpeinen M, Hakkola J, Honkakoski P, Hukkanen J, Raunio H. Inhibition and induction of human cytochrome P450 enzymes: current status. Arch Toxicol. 2008;82(10):667–715.
Friedman SL. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev. 2008;88(1):125–72.
Blaner WS, O'Byrne SM, Wongsiriroj N, Kluwe J, D'Ambrosio DM, Jiang H, et al. Hepatic stellate cell lipid droplets: a specialized lipid droplet for retinoid storage. Biochim Biophys Acta. 2009;1791(6):467–73.
Beleites C, Neugebauer U, Bocklitz T, Krafft C, Popp J. Sample size planning for classification models. Anal Chim Acta. 2013;760:25–33.
Galler K, Schleser F, Frohlich E, Requardt RP, Kortgen A, Bauer M, et al. Exploitation of the hepatic stellate cell Raman signature for their detection in native tissue samples. Integr Biol. 2014;6(10):946–56.
Galler K, Junker K, Franz M, Hentschel J, Richter P, Gajda M, et al. Differential vascular expression and regulation of oncofetal tenascin-C and fibronectin variants in renal cell carcinoma (RCC): implications for an individualized angiogenesis-related targeted drug delivery. Histochem Cell Biol. 2012;137(2):195–204.
Yamazoe S, Naya M, Shiota M, Morikawa T, Kubo A, Tani T, et al. Large-area surface-enhanced Raman spectroscopy imaging of brain ischemia by gold nanoparticles grown on random nanoarrays of transparent boehmite. ACS Nano. 2014;8(6):5622–32.
Cals F, Bakker Schut TC, Hardillo JA, Baatenburg de Jong RJ, Koljenovic S, Puppels GJ. Investigation of the potential of Raman spectroscopy for oral cancer detection in surgical margins. Lab Investig. 2015;95(10):1186–96.
Assmann C, Kirchhoff J, Beleites C, Hey J, Kostudis S, Pfister W, et al. Identification of vancomycin interaction with Enterococcus faecalis within 30 min of interaction time using Raman spectroscopy. Anal Bioanal Chem. 2015;407(27):8343–52.
Kloss S, Rosch P, Pfister W, Kiehntopf M, Popp J. Toward culture-free Raman spectroscopic identification of pathogens in ascitic fluid. Anal Chem. 2015;87(2):937–43.
Galler K, Requardt RP, Glaser U, Markwart R, Bocklitz T, Bauer M, et al. Single cell analysis in native tissue: quantification of the retinoid content of hepatic stellate cells. Sci Rep. 2016;6:24155.
Matousek P, Stone N. Recent advances in the development of Raman spectroscopy for deep noninvasive medical diagnosis. J Biophotonics. 2013;6(1):7–19.
Acknowledgments
Financial support by the DFG via the research group FOR 1738 “Heme and heme degradation products” and the BMBF (FKZ 01EO1002 and FKZ 01EO1502, Integrated Research and Treatment Center “Center for Sepsis Control and Care”) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 474 kb)
Rights and permissions
About this article
Cite this article
Galler, K., Fröhlich, E., Kortgen, A. et al. Hepatic cirrhosis and recovery as reflected by Raman spectroscopy: information revealed by statistical analysis might lead to a prognostic biomarker. Anal Bioanal Chem 408, 8053–8063 (2016). https://doi.org/10.1007/s00216-016-9905-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9905-1