Abstract
Vancomycin is an important glycopeptide antibiotic which is used to treat serious infections caused by Gram-positive bacteria. However, during the last years, a tremendous rise in vancomycin resistances, especially among Enterococci, was reported, making fast diagnostic methods inevitable. In this contribution, we apply Raman spectroscopy to systematically characterize vancomycin-enterococci interactions over a time span of 90 min using a sensitive Enterococcus faecalis strain and two different vancomycin concentrations above the minimal inhibitory concentration (MIC). Successful action of the drug on the pathogen could be observed already after 30 min of interaction time. Characteristic spectral changes are visualized with the help of multivariate statistical analysis (linear discriminant analysis and partial least squares regressions). Those changes were employed to train a statistical model to predict vancomycin treatment based on the Raman spectra. The robustness of the model was tested using data recorded by an independent operator. Classification accuracies of >90 % were obtained for vancomycin concentrations in the lower range of a typical trough serum concentration recommended for most patients during appropriate vancomycin therapy. Characterization of drug–pathogen interactions by means of label-free spectroscopic methods, such as Raman spectroscopy, can provide the knowledge base for innovative and fast susceptibility tests which could speed up microbiological analysis as well as finding applications in novel antibiotic screenings assays.

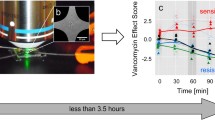
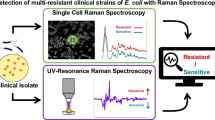
E. faecalis is incubated with vancomycin and characterized by means of Raman spectroscopy after different time points. Characteristic spectral changes reveal efficient vancomycin-enterococci-interaction
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Enterococcus faecalis are Gram-positive paired or chained cocci. They are part of the normal human gastrointestinal tract. However, in some cases, they can cause urinary tract infections, endocarditis and seldom septicaemia. Because enterococci have several natural resistances against a wide range of antibiotics, vancomycin is often used in enterococcal infection as a so-called “antibiotic of last resort”. Vancomycin is a glycopeptide antibiotic and active against aerobe and anaerobe Gram-positive bacteria. It entered clinical practice in the 1950s, but was soon substituted with alternatives that were considered to be less toxic or more efficacious [1]. Increasing resistances of pathogens against those alternative drugs resulted in a return of vancomycin [1–3].
Vancomycin inhibits the last stage, stage III, of the bacterial cell wall synthesis by binding to the d-alanyl-d-alanine terminus of the peptidoglycan precursor UDP-MurNAc-pentapeptide or nascent glycan chains. During the interaction, the peptide moiety of vancomycin forms five hydrogen bonds with the d-Ala-d-Ala dipeptide from the cell wall precursor. This bulky complex prevents glycan polymerization (transglycosylation) as well as cross-linking of the individual peptidoglycan chains (transpeptidation). This leads to an instable bacterial cell wall resulting in lysis of the cell [4].
In the 1980s, the first vancomycin-resistant enterococci (VRE) emerged which can be associated with the continuous use of vancomycin and broad spectrum antibiotics [5]. Nowadays, VRE have spread extensively worldwide and cause increasing problems in clinical practice [6–10]. However, not only the VRE themselves, but also the risk of transferring vancomycin-resistance genes to other pathogens, for example methicillin-resistant Staphylococcus aureus (MRSA), marks a major reason of concern [2, 11–13]. In order to quickly identify, isolate and efficiently treat patients suffering from VRE, fast diagnostic methods are needed. Current standard microbiological methods are very time-consuming as they require bacterial cultivation for detection of bacterial growth or growth behaviour [14, 15]. Therefore, research efforts are undertaken to speed up analysis time. Such alternative approaches comprise polymerase chain reaction (PCR)-based methods that identify the resistance genes [16–18], mass spectrometry [19–21], flow cytometry [22, 23] as well as combinations of phenotypic and genotypic approaches [19, 24, 25]. Among the spectroscopic methods, vibrational spectroscopy, such as IR absorption and Raman scattering, gained a lot of importance as they require only minimal sample amounts down to a single bacterial cell. This enables insights into cellular heterogeneity and the possibility to study difficult-to-culture pathogens with easy sample preparation [14, 26–28]. Moreover, vibrational spectroscopic techniques are label-free and non-destructive, allowing further analysis after spectroscopic characterization [29, 30]. The “vibrational spectroscopic fingerprint” is characteristic for each bacterial species due to the unique cellular composition made up of various nucleic acids, lipids, proteins, polysaccharides and carbohydrates. During recent years, Raman spectroscopy was already successfully applied to characterize bacteria, discriminate Gram-positive and Gram-negative bacteria as well as identify and classify various pathogens [26, 28, 31–37] and also pathogens under antibiotic influence [27]. An advance offers the combination of Raman spectroscopy with microfluidic devices with further functionalities, such as dielectrophoresis for the enrichment of bacteria directly in suspension, making sample preparation faster and very easy [30, 38, 39]. Furthermore, Raman spectroscopy can be successfully applied to investigate bacteria–drug interactions with antibiotics as was shown for the interaction of fluoroquinolones with Gram-positive bacteria [40, 41], the antibiotic effects on Pseudomonas aeruginosa biofilms [42] as well as the interaction of clinical relevant antibiotics with Escherichia coli [43–45]. Vancomycin–bacteria interactions have been employed to capture bacteria on silver nanoparticles for further spectroscopic characterization [46] as well as to differentiate resistant and sensitive enterococci [38].
In this contribution, we present a systematic characterization of the interaction of vancomycin with enterococci by means of Raman spectroscopy. The spectral changes upon efficient action of the drug is analyzed under different experimental conditions. The aim is to achieve a robust and fast differentiation of vancomycin treated and untreated E. faecalis. Inter-personal operator variation was taken into account by training a classification model with samples prepared and measured by one operator and testing its prediction accuracy with biological replicates prepared and measured later on by another operator.
Such drug–bacteria interaction studies provide the knowledge base for the interpretation of experiments with resistant bacteria, such as various VRE strains. Furthermore, they are also valuable for the assignment of the mode of action when studying the effect of different antibiotics on bacteria.
Materials and methods
Cultivation and treatment of microorganisms
E. faecalis ATCC® 29212™ was cultivated in CASO broth (ROTH GmbH) overnight at 37 °C while shaking at 90 rpm. The overnight culture was used to inoculate a starter culture with an optical density (OD) of 0.1 (d = 1 cm) at λ = 600 nm. To monitor bacterial growth, the OD was measured over time at 600 nm (with pathlength 1 cm) using a Cary 60 UV-Vis spectrophotometer (Agilent Technologies Cary 60). At an OD of 0.6–0.8, vancomycin hydrochloride hydrate (SIGMA-ALDRICH, Steinheim, Germany) was added to the bacteria culture giving a final vancomycin concentration of either 100 μg/ml (batches 1–4) or 10 μg/ml (batches 5–11). In the following, this time point is called time point 0 min. Each batch is an independent biological replicate (new overnight culture) that comprises a vancomycin-treated sample and a control where no vancomycin was added, but which was otherwise treated identically.
Raman sample preparation
Bacteria sampling for Raman spectroscopic investigations was carried out according to previously published protocols [40, 47]: Samples of the bacterial culture were taken directly (0 min) and 30, 60 and 90 min after addition of vancomycin. At each time point, 1 ml bacterial culture was harvested. The medium was removed by centrifugation (Eppendorf centrifuge 5415, 13,000 rpm, 3 min, room temperature) and the cells were washed twice: first, with 0.9 % sodium chloride solution and, secondly, with deionized water. The pellet was resuspended in 50 μl deionized water and dried at 55 °C for 20 min on a calcium fluoride (CaF2, Crystal GmbH, Berlin, Germany) slide. Raman spectra were taken immediately after sample preparation.
Vancomycin hydrochloride hydrate and peptidoglycan (from S. aureus cell wall component, SIGMA-ALDRICH, Steinheim, Germany) were measured as solids under the same experimental conditions as the bacteria samples.
Micro-Raman spectroscopy
Raman spectra of the bacteria on CaF2 slides were recorded with an upright micro-Raman setup (CRM 300, WITec GmbH, Germany) equipped with a 600 grooves/mm grating and a back-illuminated CCD camera (DV401 BV, ANDOR, 1024 × 127 pixels) cooled down to −60 °C. A frequency doubled Nd:YAG solid-state laser (532 nm, Excelsior 532-60) was used to excite the Raman signal. The laser light was focused onto the sample using a 50x objective (Zeiss EC EpiPlan, NA 0.7) for the batches 1–3, and a 100x objective (Carl Zeiss LD EC EpiPlan-Neofluar, NA 0.75) for batches 4–11. For the 50x objective, chromatic aberrations were encountered when studying thin bacterial films which were not observed with the 100x objective. Laser power was 15 mW at the sample. The 180° backscattered light was collected and delivered to the spectrograph by a multimode optical fibre with 50 μm core diameter. Performance and alignment of the system were checked each day by recording Raman spectra of silicon as well as of 4-acetamidophenol (ACP, paracetamol, SIGMA-ALDRICH, Steinheim, Germany). ACP spectra were used to correct possible shifts in the laser wavelength. Reference band positions were taken from the literature [48].
Two different sampling modes were applied. For batches 1–7, an automated scanning mode was used to collect 49 spectra evenly distributed over an area of 20 μm × 20 μm integrating 20 s per spectrum. However, sample roughness led to varying signal-to-noise ratio within each area. Furthermore, spectra with cosmic spikes in the fingerprint region were discarded prior to analysis. For batches 8–11, 20 single spectra (without cosmic spikes) were recorded at different locations on the sample with an integration time of 30 s. The total numbers of spectra per time point and treatment are summarized in Table 1.
Data analysis
Statistical analysis was carried out in the statistical language GNU R, version 3.0.3 [49] using the “hyperSpec” [50] environment for handling the spectra. The package “ggplot2” [51] was employed for graphical display (Fig. 4).
Spectra pre-processing
Spectra were initially truncated to the spectral ranges 350–2000 and 2500–3300 cm−1. Cosmic rays in the Raman maps were removed with an interactive, in-house written spike filter. Baseline correction in the fingerprint region (350–2000 cm−1) was performed with an automated linear baseline fit with 50 support points. For the C–H stretching region, a quadratic baseline was used. The fitting ranges were from 2500 to 2775 cm−1 and from 3150 to 3300 cm−1 where no Raman signal occurs, and the CH region between 2825 and 3050 cm−1 was extrapolated with the fit. Next, “loess” smoothing interpolation [52] was used to downsample the spectra onto an evenly spaced wavenumber axis from 600 to 1750 and 2825 to 3050 cm−1 with data point spacing of 5 cm−1. This interpolation took the laser wavelength correction calculated from the ACP spectra into account and, at the same time, provided an evenly spaced wavenumber axis common to all measurements. The smoothing parameter was set accordingly corresponding to 10 cm−1 spectral resolution. Interpolation was necessary due to slightly varying excitation wavelengths over several months (as measured by ACP spectra). This operation also trades some spectral resolution for a better signal-to-noise ratio. Fig. S1 in the Electronic Supplementary Material (ESM) visualizes the pre-treatment procedure for an exemplarily Raman spectrum of E. faecalis.
For the 50x objective (NA 0.7) employed initially to cover a large area of the bacteria film, significant chromatic aberrations were observed in the spectral region between 600 and 3000 cm−1 (corresponding to a wavelength range of 549 to 633 nm). Those chromatic aberrations were observed in the experimental data as changes in the relative intensities of corresponding Raman bands, such as, e.g. C–H deformation and C–H stretching, with different foci (z-positions) in the thin bacterial film. Similar findings have been described measuring microbial spores [53]. In a principal component analysis (PCA), this aberration initially yielded a component which was essentially the average spectrum with a change in sign between the fingerprint and the C–H stretching regions. This component carried ca. 90 % of the total variance and was uncorrelated with batch, time and treatment. The chromatic aberrations were approximately corrected by normalizing the fingerprint and C–H stretching regions separately to their mean intensity. In addition, from batch 4, another objective with improved colour correction (100x EC Epiplan Neofluar) was used.
Descriptive LDA
Descriptive linear discriminant analysis (LDA) was carried out to reveal spectral differences due to vancomycin treatment with 100 μg/ml vancomycin (four batches). These models were regularized by partial least squares regression (PLS). PLS is a latent variable approach which models the covariance structures of a matrix of predictors X (in our case the Raman spectra) and a matrix of response variables Y (which are the treatment groups “treated” and “untreated” in our example). LDA maximizes the variance between given groups (the treatment) with respect to the variance within these groups. However, in contrast to PLS, LDA needs large sample sizes relative to the number of variates. Therefore, appropriate regularization to few input variables, such as the first (few) PLS scores, is important [54, 55]. These combined PLS-LDA models are provided by the R package “cbmodels” [56] relying on packages “pls” [57] and “MASS” [52]. Separate descriptive PLS-LDA models were calculated for each of the four treatment time points (0, 30, 60 and 90 min) by taking just one PLS latent variable. The coefficients extracted from these models were used to analyse relevant changes in spectra over time.
Setup and validation of a predictive model
In order to train a model for the prediction of efficient bacteria–drug interaction, only those time points for which the PLS-LDA coefficients were steady (30, 60 and 90 min) were used. For this predictive PLS-LDA model, only two batches (batches 1 and 2) prepared and measured by operator 1 (SK) were used for training. Here, a high vancomycin concentration (100 μg/ml) was applied to observe strong drug-induced spectral variations. The first latent variable was sufficient to discriminate treated and untreated spectra. The second latent variable already carried much noise (even though some spectral features are still recognizable). While the first latent variable explains 89 % of the Y-variance in the training data using 37 % of the variance in the spectra, the second latent variable almost doubles the explained variance in the spectra, but adds only 3 % to the explained Y-variance. A closer inspection of the scores reveals that the second latent variable captures mostly variation between the biological replicates. Therefore, only one latent variable was considered for modelling. This also keeps the number of variates within the model as low as possible which is advisable for a dataset with, in total, 11 biological independent replicates [58]. The model performance was evaluated with nine independent biological replicates prepared and measured by operator 2 (CA), two replicates with 100 μg/ml and seven with only 10 μg/ml vancomycin.
Calculation of changes in the relative nucleic acid/protein content
To follow changes in the relative nucleic acid/protein ratio with time and vancomycin treatment, ratios of mean Raman intensities in the spectral range 775–800 cm−1, representing nucleic acid (785 cm−1), to the 995–1020 cm−1 region, representing protein (1005 cm−1), were calculated (Fig. 1b) by numerically integrating the spectrum in the respective spectral range.
a Growth curve of Enterococcus faecalis measured as optical density (OD) at 600 nm over time in the absence and presence of vancomycin. The arrow marks the time point of antibiotic application. b Nucleic acid/protein ratio during cultivation without (0 μg/ml) and with 10 or 100 μg/ml of vancomycin calculated using the integrated intensity of the Raman bands at 785 cm−1 (uracil, cytosine, thymine and PO2 −) and at 1005 cm−1 (phenylalanine). The mean and standard deviation were calculated from the 11 biological replicates
Results and discussion
Characterization of the growth of E. faecalis using optical density
The growth of E. faecalis ATCC® 29212™ was measured as optical density of the bacteria culture over a time of 3 h (1.5 h before to 1.5 h after vancomycin addition). Each biological replicate (batch) comprises an E. faecalis control culture without vancomycin and an E. faecalis culture incubated with vancomycin. Figure 1a shows a typical E. faecalis growth curve which can be divided in lag phase (first 30 min of growth), followed by the exponential growth phase and stationary phase (>90 min, not shown). From the slope of the growth curve in the exponential phase the specific growth rate μ of E. faecalis ATCC® 29212™ in CASO at 37 °C was determined to be μ = 1.315 ± 0.07 h−1, the generation time g = ln2/μ = 32 ± 2 min and the dividing rate n = 1/g = 1.9 ± 0.1 h−1. Mean and standard deviation were calculated from six independent biological replicates. These values are in good agreement with other data published in the literature [59–61].
Characterization of vancomycin–E. faecalis interaction using established microbiological methods
The minimum inhibitory concentration (MIC) of E. faecalis ATCC® 29212™ was determined by automated analysis using a VITEK® 2 system (bioMérieux, Nürtingen, Germany) to be 2 μg/ml. This is in good agreement with literature values reporting MICs for vancomycin between 1 and 4 μg/ml [62]. With a MIC of vancomycin ≤4 mg/l, E. faecalis ATCC® 29212™ is a vancomycin susceptible strain according to the EUCAST-approved susceptibility breakpoint [63].
Vancomycin inhibits cell wall synthesis during bacterial growth and division [4]; thus, it is most efficient on metabolically active bacteria. Therefore, in our experiments, vancomycin was always added during the exponential growth phase (90 min after inoculation, time point 0 min). To ensure a strong and fast effect of the antibiotic, a vancomycin concentration of 50 times the determined MIC, i.e. 100 μg/ml, was used to analyse typical spectral changes induced by the action of the drug and also to train the classification model to identify treated bacteria. The model was tested using vancomycin concentrations of 5 times the MIC (10 μg/ml vancomycin) which is the lower range of a typical trough serum concentration recommended for most patients during appropriate vancomycin therapy [1, 64].
The growth curve depicted in Fig. 1a demonstrates that both applied vancomycin concentrations (100 and 10 μg/ml) inhibit further growth of E. faecalis. For the high vancomycin concentration (100 μg/ml), this effect is visible already after 30 min for the lower concentration at 60 min after vancomycin addition.
Raman spectroscopic characterization of vancomycin–E. faecalis interaction
The antibiotic influence of vancomycin on E. faecalis was monitored by recording Raman spectra of the bacteria immediately after adding the drug (0 min) and every 30 min after vancomycin addition over a time span of 1.5 h, i.e. 30, 60 and 90 min of vancomycin–E. faecalis interaction time. The experiment was carried out using two different drug concentrations: 100 and 10 μg/ml vancomycin respectively. E. faecalis cultures without vancomycin treatment served as controls.
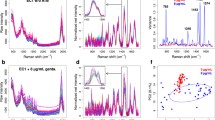
Figure 2 summarizes the pre-processed average Raman spectra of E. faecalis at the four different time points after addition of vancomycin, as well as Raman spectra of the untreated controls. Mean spectra including the 5th and 95th percentile are shown in Fig. S2 in the ESM. The Raman spectra show all characteristic features of typical bacteria spectra and are in good agreement with published data [26, 27, 30, 31]. The vibrational bands can be assigned according to contributions from the overall biochemistry content of the bacteria cells. Nucleic acid contributions show up at 785, 1335 and 1575 cm−1. Proteins are responsible for the amide I around 1660 cm−1 and the amide III band around 1250 cm−1 as well as the sharp ring breathing mode of phenylalanine at 1005 cm−1. The band around 1450 cm−1 can be assigned to C–H deformation vibrations and the prominent bands around 2935 cm−1 to C–H stretching vibrations. The Raman spectra of the different treatment methods and the different time points appear very similar at first glance. However, they contain significant differences that reflect metabolic changes. These differences can be extracted and visualized by multivariate statistical analysis methods.
Calculation of the nucleic acid/protein ratio to follow bacterial growth
The ratio of nucleic acid and protein content is known to be characteristic for the different metabolic stages during bacterial growth [41, 43, 65]. This ratio is highest in the exponential growth phase when bacteria have high transcription of genes and decreases in the stationary phase when the bacteria accumulate storing material and show reduced replication activities. Figure 1b shows the calculated nucleic acid/protein band ratios for the three different vancomycin concentrations. As expected, a slight decrease of the ratio is observed for the controls (0 μg/ml vancomycin). During the observation period (0–90 min), these cultures develop from the exponential growth phase towards the stationary phase. For bacteria treated with vancomycin, the nucleic acid/protein ratio stays constant within experimental error (Fig. 1b). This indicates that under the influence of the drug, bacteria cannot progress into another metabolic state and remain with a similar overall chemical composition as when hit by the insult. This observation is in good agreement with the action of the antibiotic: Vancomycin inhibits the cell wall biosynthesis, prevents bacteria to further divide and, ultimately, leads to cell death.
Descriptive LDA to analyze spectral changes due to vancomycin interaction
In order to visualize the spectral differences caused by the action of the drug, the Raman spectra of the untreated controls and the bacteria treated with the high vancomycin concentration (100 μg/ml) were compared for each time point by means of partial least square regression combined with linear discriminant analysis (PLS-LDA). The high vancomycin concentration was chosen for this analysis as the effect is expected to be strong. The analysis is carried out by individual comparison at each time point to unravel at which time point spectral changes happen. The coefficients of the PLS-LDA models are shown in Fig. 3a. Large coefficients (both, positive and negative) indicate wavenumbers that contribute to the differentiation of treated and untreated E. faecalis. Positive scores indicate vibrational bands that are more prominent in vancomycin-treated bacteria while negative scores originate from vibrational bands that are more intense in the untreated controls. The coefficient at time point 0 min shows no significant features, but rather spectral noise. This indicates that the pure presence of the antibiotic vancomycin in the bacterial culture does not have an effect on the Raman spectra of the bacteria. After 30 min of vancomycin interaction, time in the bacterial culture and characteristic spectral features emerge. These features stay the same or become even more prominent as the time elapses. All in all, the coefficients of the three interaction time points 30, 60 and 90 min look very similar. Most prominent positive signals are observed around 790, 1225 and 1490 cm−1. The signals around 1225 and 1490 cm−1 could be assigned protein contributions (amide III and amide II, respectively). Especially the band around 1225 cm−1 has been described to reflect intermolecular hydrogen bonds in bacterial cell wall peptides [66]. Such hydrogen bonds would be in good agreement with vancomycin binding to d-alanyl-d-alanine precursors of the cell wall. Strong negative signals are found around 1445 and 1655 cm−1. These signals can be assigned to CH2 deformation vibrations [31, 67] and to contributions from the amide I vibration from proteins [31] respectively. We attribute these spectral features to the increase in relative protein content with time, as discussed previously for the nucleic acid/protein ratio. However, the majority of the signals are a superposition of bands from several biochemical compounds of the complex bacterial cells allowing only wavenumber estimation instead of a well-defined band assignment. Raman spectra of pure substances are shown in Fig. 3b for comparison. The spectrum of solid vancomycin hydrochloride which is a glycopeptide consisting of amino acids and carbohydrates shows prominent Raman bands around 885, 993, 1232, 1320, 1610 and 2945 cm−1. These bands can be assigned to several vibrations corresponding to peptides/proteins [31, 67]. None of those bands contributes to the PLS-LDA coefficient separating treated and untreated bacteria. Thus again, not the pure presence of vancomycin but rather the action of the drug on the enterococci is detected. Vancomycin inhibits the bacterial cell wall biosynthesis by binding to cell wall precursors and hence, inhibiting peptidoglycan synthesis. The Raman spectrum of peptidoglycan, a polymer made of sugars (N-acetylglucosamine and N-acetylmuramic acid) and amino acids, shows prominent vibrational bands at 1670 and 1448 cm−1 originating from the peptide bonds and C–H deformation vibrations, respectively (Fig. 3b). Both of those vibrations appear as negative coefficients in Fig. 3a, a more pronounced contribution of peptidoglycans to the spectra of untreated bacteria. This is in agreement with an impaired peptidoglycan synthesis in the vancomycin-treated bacteria. However, it is impossible to assign the vibrational bands in the coefficient plot explicitly to changes in the bacterial cell wall. Similar bands could be observed due to changes in other proteins.
Classification model for the fast identification of vancomycin interaction
In order to rapidly detect efficient vancomycin-enterococci interaction, a classification model to distinguish treated and untreated bacteria has been trained. Based on the observation from the descriptive LDA that all interaction times >30 min show similar coefficients, all time points after at least 30 min of vancomycin treatment were included into one classification model.
Two different operators carried out the experiments. To verify the model including possible inter-personal variation, data from operator 1 were used to set up the classification model and data recorded by operator 2 were used to evaluate the model's predictive abilities. The preferred validation scheme in literature for those numbers of replicates would be a resampling method such as repeated cross validation or .632+ bootstrap on the level of biological replicates [68, 69]. In the present study, however, we were able to make use of the rare fact that two independent persons were involved in the experiments. This allowed checking the generalization ability for measurements acquired up to 1 year after the acquisition of the training replicates was finished. The prediction of the data (biological replicates in batches 3–11) recorded by operator 2 with this treatment-based classification model is visualized in the score plot of the PLS-LDA model (Fig. 4). Spectra were classified as either control or vancomycin-treated samples. For easier assessment, data are grouped according to true treatment, i.e. control (0 μg/ml vancomycin) and treated with 10 and 100 μg/ml vancomycin respectively. Table 2 summarizes the correct classifications for each time point and treatment. Already after 30 min of vancomycin incubation, the prediction accuracy is above 90 %. Longer incubation times (60 min) lead to some further increase in correct classifications (≥95 %) and stay so also for 90 min incubation time.
Classification model based on antibiotic treatment. Test results for nine independent biological replicates prepared and measured by an independent operator. Single Raman spectra are represented by symbols. Horizontal lines represent 5, 10, 25, 50, 75, 90 and 95 % predicted posterior probability of the spectrum coming from a vancomycin-treated sample
For test samples treated with 100 μg/ml vancomycin, already at time point 0 min, many of the treated samples are recognized as such. We attribute this mainly to the fact that due to sample handling, a short but existing duration of vancomycin treatment occurs: While the sample is taken immediately after vancomycin is added, some time is needed for centrifugation and washing during which the drug can affect the bacteria. With the lower vancomycin concentration, this effect is not as pronounced. At time points ≥30 min, the scores for both concentrations are practically the same. As PLS-LDA scores would reflect concentration changes in a linear manner, we again arrive at the conclusion that biochemical changes in the cells due to the interaction of the drug with the cell are observed rather than the pure presence of the drug. Furthermore, the drug-induced biochemical changes reflected in the Raman spectra are the same for vancomycin concentrations 5× and 50× of the MIC. This is in agreement with biological knowledge as the MIC denotes the lowest drug concentration that can inhibit further growth of the bacteria.
The slight trend of the PLS-LDA scores of the untreated control samples towards more negative coefficients as time progresses indicates that also growth-dependent biochemical changes as they occur when evolving from the mid to the late exponential growth phase might contribute to the PLS-LDA to recognize untreated cases.
Summary and conclusion
It was shown that micro-Raman spectroscopy can be used to describe biochemical changes within bacteria caused by the action of a drug as shown for the interaction of vancomycin with E. faecalis. Observed spectral changes can be explained using biological knowledge of the mode of action of the antibiotics. Furthermore, it could be shown that the observed spectral changes are not caused by the pure presence of the drug. Observing the effect rather than the presence of the antibiotic is a necessary (though not sufficient) condition for the future development of an analytical method to detect antibiotic susceptibility of unknown strains and maybe even species.
The spectral differences observed in the presence and absence of vancomycin were used to train a classification model which is able to discriminate between vancomycin-treated and untreated groups of E. faecalis. The quality of the model was assessed with an independent test data set recorded by an independent operator yielding prediction accuracies >90 %. This marks the high potential of the presented method for further diagnostic or pharmacological tests.
References
Levine DP (2006) Vancomycin: a history. Clin Infect Dis 42(Supplement 1):S5–S12. doi:10.1086/491709
Sood S, Malhotra M, Das BK, Kapil A (2008) Enterococcal infections & antimicrobial resistance. Indian J Med Res 128(2):111–121
Van Bambeke F, Van Laethem Y, Courvalin P, Tulkens PM (2004) Glycopeptide antibiotics from conventional molecules to new derivatives. Drugs 64(9):913–936. doi:10.2165/00003495-200464090-00001
Kahne D, Leimkuhler C, Wei L, Walsh C (2005) Glycopeptide and lipoglycopeptide antibiotics. Chem Rev 105(2):425–448. doi:10.1021/cr030103a
Arias CA, Murray BE (2012) The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol 10(4):266–278. doi:10.1038/nrmicro2761
Werner G, Strommenger B, Witte W (2008) Acquired vancomycin resistance in clinically relevant pathogens. Futur Microbiol 3(5):547–562. doi:10.2217/17460913.3.5.547
Kristich CJ, Rice LB, Arias CA (2014) Enterococcal infection—treatment and antibiotic resistance. Enterococci: from commensals to leading causes of drug resistant infection. Massachusetts Eye and Ear Infirmary, Boston
Kudo M, Nomura T, Yomoda S, Tanimoto K, Tomita H (2014) Nosocomial infection caused by vancomycin-susceptible multidrug-resistant Enterococcus faecalis over a long period in a university hospital in Japan. Microbiol Immunol 58(11):607–614. doi:10.1111/1348-0421.12190
Gastmeier P, Schröder C, Behnke M, Meyer E, Geffers C (2014) Dramatic increase in vancomycin-resistant enterococci in Germany. J Antimicrob Chemother 69(6):1660–1664. doi:10.1093/jac/dku035
Fernandes SC, Dhanashree B (2013) Drug resistance & virulence determinants in clinical isolates of Enterococcus species. Indian J Med Res 137(5):981–985
Noble WC, Virani Z, Cree RG (1992) Co-transfer of vancomycin and other resistance genes from Enterococcus faecalis NCTC 12201 to Staphylococcus aureus. FEMS Microbiol Lett 72(2):195–198
Hegstad K, Mikalsen T, Coque TM, Werner G, Sundsfjord A (2010) Mobile genetic elements and their contribution to the emergence of antimicrobial resistant Enterococcus faecalis and Enterococcus faecium. Clin Microbiol Infect 16(6):541–554. doi:10.1111/j.1469-0691.2010.03226.x
de Niederhäusern S, Bondi M, Messi P, Iseppi R, Sabia C, Manicardi G, Anacarso I (2011) Vancomycin-resistance transferability from VanA enterococci to Staphylococcus aureus. Curr Microbiol 62(5):1363–1367. doi:10.1007/s00284-011-9868-6
Brehm-Stecher BF, Johnson EA (2004) Single-cell microbiology: tools, technologies, and applications. Microbiol Mol Biol Rev 68(3):538–559. doi:10.1128/mmbr.68.3.538-559.2004
Jorgensen JH, Ferraro MJ (2009) Antimicrobial susceptibility testing: a review of general principles and contemporary practices. Clin Infect Dis 49(11):1749–1755. doi:10.1086/647952
Palladino S, Kay ID, Flexman JP, Boehm I, Costa AM, Lambert EJ, Christiansen KJ (2003) Rapid detection of vanA and vanB genes directly from clinical specimens and enrichment broths by real-time multiplex PCR assay. J Clin Microbiol 41(6):2483–2486
Waldeisen JR, Wang T, Mitra D, Lee LP (2011) A real-time PCR antibiogram for drug-resistant sepsis. PLoS One 6(12), e28528. doi:10.1371/journal.pone.0028528
Gousia P, Economou V, Bozidis P, Papadopoulou C (2015) Vancomycin-resistance phenotypes, vancomycin-resistance genes, and resistance to antibiotics of enterococci isolated from food of animal origin. Foodborne Pathog Dis. doi:10.1089/fpd.2014.1832
Machen A, Drake T, Wang YF (2014) Same day identification and full panel antimicrobial susceptibility testing of bacteria from positive blood culture bottles made possible by a combined lysis-filtration method with MALDI-TOF VITEK mass spectrometry and the VITEK2 system. PLoS One 9(2), e87870. doi:10.1371/journal.pone.0087870
Jung JS, Popp C, Sparbier K, Lange C, Kostrzewa M, Schubert S (2014) Evaluation of matrix-assisted laser desorption ionization-time of flight mass spectrometry for rapid detection of beta-lactam resistance in Enterobacteriaceae derived from blood cultures. J Clin Microbiol 52(3):924–930. doi:10.1128/jcm.02691-13
Lange C, Schubert S, Jung J, Kostrzewa M, Sparbier K (2014) Quantitative matrix-assisted laser desorption ionization-time of flight mass spectrometry for rapid resistance detection. J Clin Microbiol 52(12):4155–4162. doi:10.1128/jcm.01872-14
Gauthier C, St-Pierre Y, Villemur R (2002) Rapid antimicrobial susceptibility testing of urinary tract isolates and samples by flow cytometry. J Med Microbiol 51(3):192–200
Jarzembowski T, Jozwik A, Wisniewska K, Witkowski J (2010) Flow cytometry approach study of Enterococcus faecalis vancomycin resistance by detection of Vancomycin@FL binding to the bacterial cells. Curr Microbiol 61(5):407–410. doi:10.1007/s00284-010-9628-z
Seo JY, Kim PW, Lee JH, Song JH, Peck KR, Chung DR, Kang CI, Ki CS, Lee NY (2011) Evaluation of PCR-based screening for vancomycin-resistant enterococci compared with a chromogenic agar-based culture method. J Med Microbiol 60(Pt 7):945–949. doi:10.1099/jmm.0.029777-0
Chan WS, Chan TM, Lai TW, Chan JF, Lai RW, Lai CK, Tang BS (2015) Complementary use of MALDI-TOF MS and real-time PCR-melt curve analysis for rapid identification of methicillin-resistant staphylococci and VRE. J Antimicrob Chemother 70(2):441–447. doi:10.1093/jac/dku411
Kloss S, Kampe B, Sachse S, Rösch P, Straube E, Pfister W, Kiehntopf M, Popp J (2013) Culture independent Raman spectroscopic identification of urinary tract infection pathogens: a proof of principle study. Anal Chem 85(20):9610–9616. doi:10.1021/ac401806f
Münchberg U, Rösch P, Bauer M, Popp J (2014) Raman spectroscopic identification of single bacterial cells under antibiotic influence. Anal Bioanal Chem 406(13):3041–3050. doi:10.1007/s00216-014-7747-2
Harz M, Kiehntopf M, Stöckel S, Rösch P, Straube E, Deufel T, Popp J (2009) Direct analysis of clinical relevant single bacterial cells from cerebrospinal fluid during bacterial meningitis by means of micro-Raman spectroscopy. J Biophotonics 2(1-2):70–80. doi:10.1002/jbio.200810068
Mathey R, Dupoy M, Espagnon I, Leroux D, Mallard F, Novelli-Rousseau A (2015) Viability of 3h grown bacterial micro-colonies after direct Raman identification. J Microbiol Methods 109:67–73. doi:10.1016/j.mimet.2014.12.002
Schröder U-C, Ramoji A, Glaser U, Sachse S, Leiterer C, Csaki A, Hübner U, Fritzsche W, Pfister W, Bauer M, Popp J, Neugebauer U (2013) Combined dielectrophoresis-Raman setup for the classification of pathogens recovered from the urinary tract. Anal Chem 85(22):10717–10724. doi:10.1021/ac4021616
Maquelin K, Kirschner C, Choo-Smith LP, van den Braak N, Endtz HP, Naumann D, Puppels GJ (2002) Identification of medically relevant microorganisms by vibrational spectroscopy. J Microbiol Methods 51(3):255–271. doi:10.1016/s0167-7012(02)00127-6
Kirschner C, Maquelin K, Pina P, Ngo Thi NA, Choo-Smith LP, Sockalingum GD, Sandt C, Ami D, Orsini F, Doglia SM, Allouch P, Mainfait M, Puppels GJ, Naumann D (2001) Classification and identification of enterococci: a comparative phenotypic, genotypic, and vibrational spectroscopic study. J Clin Microbiol 39(5):1763–1770. doi:10.1128/JCM.39.5.1763-1770.2001
Rösch P, Harz M, Schmitt M, Peschke KD, Ronneberger O, Burkhardt H, Motzkus HW, Lankers M, Hofer S, Thiele H, Popp J (2005) Chemotaxonomic identification of single bacteria by micro-Raman spectroscopy: application to clean-room-relevant biological contaminations. Appl Environ Microbiol 71(3):1626–1637. doi:10.1128/AEM.71.3.1626-1637.2005
Harz M, Rösch P, Popp J (2009) Vibrational spectroscopy--a powerful tool for the rapid identification of microbial cells at the single-cell level. Cytometry A 75(2):104–113. doi:10.1002/cyto.a.20682
Kastanos EK, Kyriakides A, Hadjigeorgiou K, Pitris C (2010) A novel method for urinary tract infection diagnosis and antibiogram using Raman spectroscopy. J Raman Spectrosc 41(9):958–963. doi:10.1002/jrs.2540
Lu XN, Al-Qadiri HM, Lin MS, Rasco BA (2011) Application of mid-infrared and Raman spectroscopy to the study of bacteria. Food Bioprocess Tech 4(6):919–935. doi:10.1007/s11947-011-0516-8
Kloss S, Rösch P, Pfister W, Kiehntopf M, Popp J (2015) Toward culture-free Raman spectroscopic identification of pathogens in ascitic fluid. Anal Chem 87(2):937–943. doi:10.1021/ac503373r
Schröder U-C, Beleites C, Assmann C, Glaser U, Hübner U, Pfister W, Fritzsche W, Popp J, Neugebauer U (2015) Detection of vancomycin resistances in enterococci within 3 (1/2) hours. Sci Rep 5:8217. doi:10.1038/srep08217
Madiyar FR, Bhana S, Swisher LZ, Culbertson CT, Huang X, Li J (2015) Integration of a nanostructured dielectrophoretic device and a surface-enhanced Raman probe for highly sensitive rapid bacteria detection. Nanoscale 7(8):3726–3736. doi:10.1039/c4nr07183b
Neugebauer U, Schmid U, Baumann K, Holzgrabe U, Ziebuhr W, Kozitskaya S, Kiefer W, Schmitt M, Popp J (2006) Characterization of bacterial growth and the influence of antibiotics by means of UV resonance Raman spectroscopy. Biopolymers 82(4):306–311. doi:10.1002/bip.20447
Neugebauer U, Schmid U, Baumann K, Ziebuhr W, Kozitskaya S, Holzgrabe U, Schmitt M, Popp J (2007) The influence of fluoroquinolone drugs on the bacterial growth of S. epidermidis utilizing the unique potential of vibrational spectroscopy. J Phys Chem A 111(15):2898–2906. doi:10.1021/jp0678397
Jung GB, Nam SW, Choi S, Lee GJ, Park HK (2014) Evaluation of antibiotic effects on Pseudomonas aeruginosa biofilm using Raman spectroscopy and multivariate analysis. Biomed Opt Express 5(9):3238–3251. doi:10.1364/boe.5.003238
Moritz TJ, Polage CR, Taylor DS, Krol DM, Lane SM, Chan JW (2010) Evaluation of Escherichia coli cell response to antibiotic treatment by use of Raman spectroscopy with laser tweezers. J Clin Microbiol 48(11):4287–4290. doi:10.1128/JCM.01565-10
Heidari Torkabadi H, Bethel CR, Papp-Wallace KM, de Boer PA, Bonomo RA, Carey PR (2014) Following drug uptake and reactions inside Escherichia coli cells by Raman microspectroscopy. Biochemistry 53(25):4113–4121. doi:10.1021/bi500529c
Athamneh AI, Alajlouni RA, Wallace RS, Seleem MN, Senger RS (2014) Phenotypic profiling of antibiotic response signatures in Escherichia coli using Raman spectroscopy. Antimicrob Agents Chemother 58(3):1302–1314. doi:10.1128/aac.02098-13
Liu TY, Tsai KT, Wang HH, Chen Y, Chen YH, Chao YC, Chang HH, Lin CH, Wang JK, Wang YL (2011) Functionalized arrays of Raman-enhancing nanoparticles for capture and culture-free analysis of bacteria in human blood. Nat Commun 2:538. doi:10.1038/ncomms1546
Stöckel S, Walter A, Boßecker A, Meisel S, Ciobotă V, Schumacher W, Rösch P, Popp J (2011) Identification and characterization of microorganisms by vibrational spectroscopy. In: Popp J, Tuchin VV, Chiou A, Heinemann SH (eds) Handbook of biophotonics, vol 2, Photonics for Health Care. John Wiley & Sons, Weinheim, pp 105–142
McCreery RL (2000) Raman spectroscopy for chemical analysis, vol 157, Chemical analysis. John Wiley & Sons, New York
R Core Team (2014) R: a language and environment for statistical computing. R version 3.0.3 (2014-03-06). R Foundation for Statistical Computing, Vienna
Beleites C, Sergo V (2014) hyperSpec: a package to handle hyperspectral data sets in R. R package version 0.98-20140220
Wickham H (2009) ggplot2: elegant graphics for data analysis. Springer, New York. doi:10.1007/978-0-387-98141-3
Venables WN, Ripley BD (2002) Modern applied statistics with S. Statistics and computing, 4th edn. Springer, New York
Lasch P, Hermelink A, Naumann D (2009) Correction of axial chromatic aberrations in confocal Raman microspectroscopic measurements of a single microbial spore. Analyst 134(6):1162–1170. doi:10.1039/b822553b
Barker M, Rayens W (2003) Partial least squares for discrimination. J Chemometrics 17(3):166–173
Næs T, Mevik B-H (2001) Understanding the collinearity problem in regression and discriminant analysis. J Chemometrics 15(4):413–426
Beleites C (2014) cbmodels: Collection of "combined" models: PCA-LDA, PLS-LDA, etc. R package version 0.5-20140225
Mevik B-H, Wehrens R, Liland KH (2013) pls: Partial Least Squares and Principal Component regression. R package version 2.4-3
Beleites C, Neugebauer U, Bocklitz T, Krafft C, Popp J (2013) Sample size planning for classification models. Anal Chim Acta 760 (0):25-33. doi:http://dx.doi.org/10.1016/j.aca.2012.11.007
Edelstein EM, Rosenzweig MS, Daneo-Moore L, Higgins ML (1980) Unit cell hypothesis for Streptococcus faecalis. J Bacteriol 143(1):499–505
Koch AL, Higgins ML (1984) Control of wall band splitting in Streptococcus faecalis. J Gen Microbiol 130(4):735–745
Shlaes DM, Bouvet A, Devine C, Shlaes JH, al-Obeid S, Williamson R (1989) Inducible, transferable resistance to vancomycin in Enterococcus faecalis A256. Antimicrob Agents Chemother 33(2):198–203
Swenson JM, Clark NC, Sahm DF, Ferraro MJ, Doern G, Hindler J, Jorgensen JH, Pfaller MA, Reller LB, Weinstein MP et al (1995) Molecular characterization and multilaboratory evaluation of Enterococcus faecalis ATCC 51299 for quality control of screening tests for vancomycin and high-level aminoglycoside resistance in enterococci. J Clin Microbiol 33(11):3019–3021
The European Committee on Antimicrobial Susceptibility Testing (2015) Breakpoint tables for interpretation of MICs and zone diameters. http://www.eucast.org
Rybak MJ, Lomaestro BM, Rotschafer JC, Moellering RC, Craig WA, Billeter M, Dalovisio JR, Levine DP (2009) Vancomycin therapeutic guidelines: a summary of consensus recommendations from the infectious diseases society of america, the american society of health-system pharmacists, and the society of infectious diseases pharmacists. Clin Infect Dis 49(3):325–327. doi:10.1086/600877
Große C, Bergner N, Dellith J, Heller R, Bauer M, Mellmann A, Popp J, Neugebauer U (2015) Label-free imaging and spectroscopic analysis of intracellular bacterial infections. Anal Chem 87(4):2137–2142. doi:10.1021/ac503316s
Naumann D, Labischinski H, Rönspeck W, Barnickel G, Bradaczek H (1987) Vibrational spectroscopic analysis of LD-sequential, bacterial cell wall peptides: an IR and Raman study. Biopolymers 26(6):795–817. doi:10.1002/bip.360260603
Notingher I, Verrier S, Romanska H, Bishop AE, Polak JM, Hench LL (2002) In situ characterisation of living cells by Raman spectroscopy. Spectroscopy 16(2):43–51. doi:10.1155/2002/408381
Beleites C, Baumgartner R, Bowman C, Somorjai R, Steiner G, Salzer R, Sowa MG (2005) Variance reduction in estimating classification error using sparse datasets. Chemometrics Intellig Lab Syst 79(1–2):91–100. doi:10.1016/j.chemolab.2005.04.008
Kohavi R (1995) A study of cross-validation and bootstrap for accuracy estimation and model selection. In: Proceedings of the 14th international joint conference on Artificial intelligence, Montreal, Quebec, Canada. Morgan Kaufmann Publishers Inc., San Francisco, CA, USA, pp 1137-1143
Acknowledgments
Financial support of the BMBF via the Integrated Research and Treatment Center “Center for Sepsis Control and Care” (FKZ 01EO1002) and via the Carl Zeiss Stiftung is highly acknowledged. We thank A. Saupe for the VITEK® measurements as well as Martin Gnauck and Steffen Wolf for recording the scanning electron microscope (SEM) image (graphical abstract).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in the topical collection Raman4Clinics with guest editors Jürgen Popp and Christoph Krafft.
Cora Assmann and Johanna Kirchhoff contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 116 kb)
Rights and permissions
About this article
Cite this article
Assmann, C., Kirchhoff, J., Beleites, C. et al. Identification of vancomycin interaction with Enterococcus faecalis within 30 min of interaction time using Raman spectroscopy. Anal Bioanal Chem 407, 8343–8352 (2015). https://doi.org/10.1007/s00216-015-8912-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-8912-y