Abstract
So far, the majority of in vitro toxicological experiments are conducted after an acute 24 h treatment that does not represent a realistic human chemical exposure. Recently, new in vitro approaches have been proposed to study the chemical toxicological effect over several days in order to be more predictive of a representative exposure scenario. In this study, we investigated the genotoxic potential of chemicals (direct or bioactived clastogen, aneugen and apoptotic inducer) with the γH2AX and pH3 biomarkers, in the human liver-derived HepaRP cell line. We used different treatment durations, with or without a three-day recovery stage (release period), before genotoxicity measurement. Data were analysed with the Benchmark Dose approach. We observed that the detection of clastogenic compounds (notably for DNA damaging agents) was more sensitive after three days of repeated treatment compared to one or three treatments over 24 h. In contrast, aneugenic chemicals were detected as genotoxic in a similar manner whether after a 24 h exposure or a three-day repeated treatment. Globally, the release period decreases the genotoxicity measurement substantially. For DNA damaging agents, after high concentration treatments, γH2AX induction was always observed after a three-day release period. In contrast, for DNA topoisomerase inhibitors, no effect could be observed after the release period. In conclusion, in the HepaRP cell line, there are some important differences between a one-day acute and a three-day repeated treatment protocol, indicating that different cell treatment procedures may differentiate chemical genotoxic mechanisms of action more efficiently.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In vitro chemical toxicity measurement is usually performed after one day of acute exposure. However, the duration of exposure during the experiment may affect a compound’s potency to be detected as a toxic chemical. So far, only a limited number of studies have been conducted to investigate the impact of exposure duration on the toxic chemical potency with in vitro tests due to cell death at confluence for many cell lines used (Jennen et al. 2010; Bell et al. 2017; Gupta et al. 2021; Duivenvoorde et al. 2021).
As an alternative, the HepaRG human cell line (isolated from a patient with hepatocarcinoma) is widely used in toxicological experiments because of its strong similarity to human hepatocytes with metabolic capacities and the ability to apply repeated treatment (Aninat et al. 2006; Cerec et al. 2007; Guillouzo et al. 2007; Quesnot et al. 2016). However, the HepaRG cell bank is limited. Consequently, epigenetic reprogramming of HepaRG cells was used to generate the HepaRP cell line, with P450 enzyme levels comparable to those expressed in primary human hepatocyte cultures and the ability to remain differentiated for several days at confluence (Brun et al. 2023).
Several compounds classified as genotoxic can lead to DNA damage, increase mutation load, and consequently induce a carcinogenesis process as well as other health outcomes such as premature aging or infertility (Srinivasan et al. 1997; Friedberg 2003; Wheeldon et al. 2020). The aim of this study was to compare the toxic potential (cytotoxicity and genotoxicity) induced by eight compounds with different modes of action (MoA) after a single or repeated exposure in human HepaRP cells. We also chose as toxic control tunicamycin, a cytotoxic compound that does not induce genotoxic damage (Kim et al. 2018). We decided to test three groups of genotoxicants that induce DNA damage by different MoA: benzo[a]pyrene and aflatoxin B1, which lead to DNA adducts whereas mitomycin C leads to DNA cross link (Brüsehafer et al. 2014; Shah et al. 2016; Theumer et al. 2018). We also tested the genotoxic DNA topoisomerase inhibitors etoposide and camptothecin (Baldwin and Osheroff 2005; Pommier 2006). Finally, we chose two aneugen agents taxol and vinblastine (Vogel and Nivard 1993; Jordan and Wilson 1998).
In this study, HepaRP cells were exposed to each of the eight different compounds for one acute period or for repeated treatments. We also evaluated the effect of a three-day recovery period after the last chemical treatment. Genotoxicity was evaluated using the In-Cell Western (ICW) technique with the γH2AX and pH3 genotoxicity biomarkers. γH2AX is known to be induced after various types of DNA damage and is now used as a classical biomarker for clastogenic chemical detection (Burma et al. 2001; Bonner et al. 2008; Kopp et al. 2019). Aneugenic chemical treatment blocks the cells in mitosis inducing histone H3 phosphorylation (Banerjee and Chakravarti 2011). Using a combination of the γH2AX and pH3 biomarkers, clastogenic and aneugenic chemicals can be efficiently distinguished from cytotoxic compounds (Khoury et al. 2016a, b). The benchmark dose (BMD) modelling approach can be used as an alternative to the lowest observed genotoxic effect level (LOGEL). BMD modelling permits to fit mathematical functions to all of the concentration–response data in order to derive the BMD with an interval of confidence (Wills et al. 2016; Wheeldon et al. 2020; Sanders et al. 2022). This mathematical method can be applied to in vitro genotoxicity data for risk assessment and chemical prioritization (Beal et al. 2023). The BMD approach was used in this study to compare the effect of the different treatment protocols tested.
Materials and methods
Chemicals and reagents
All tested compounds [benzo[a]pyrene (BaP), aflatoxin B1 (AFB1), mitomycin C (MMC), etoposide (ETO), camptothecin (CPT), paclitaxel (Taxol), vinblastine (VIN), tunicamycin (TUNI)] were purchased from Sigma-Aldrich (France) and prepared in dimethyl sulfoxide (DMSO). Penicilin, trypsin, PBS, RNAse A and Triton X-100 were purchased from Sigma Aldrich. The blocking solution (Maxblock Blocking Medium) was purchased from Active Motif (Belgium).
Cell culture
The HepaRP cell line was provided by Biopredic International (St Grégoire, France). These cells are derived from a human hepatic cancer cell line (HepaRG) and express a large panel of P450 cytochromes and detoxification phase II enzymes (Brun et al. 2023). Briefly, undifferentiated HepaRP cells were cultured in growth medium (Wiliam E medium, 10% AD310 serum and 1% glutamax) for 14 days and then differentiated in differentiation medium (William E, 10% serum AD320, 1% glutamax) for another 14 days in 5% CO2 at 37 °C. For differentiation, an increasing percentage of DMSO was added every other day (0.2%, 0.4%, 0.8% and 1.4%). Once the differentiation was completed, the cells were trypsinated, counted and seeded to 40,000 cells/well in a 96 transparent black bottom plate that allows fluorescence analysis.
Cell treatments
Cells were treated with eight compounds at seven different concentrations separated by a one third dilution. In addition, different treatment times were studied (Fig. 1): one treatment over 24 h, three treatments over 24 h and three repeated treatments every 24 h over 72 h. To study the medium-term effect of these treatments, we also performed a period of cell release (addition of a proliferation medium without the compound) for three days. This release period may allow the cells to repair the cellular damage or die in case of over-damage, before the genotoxicity test is carried out.
Genotoxicity analysis with the γH2AX/pH3 assay
The γH2AX/pH3 assay was mostly performed as described before (Khoury et al. 2016a, b; Kopp et al. 2020). At the end of the treatment or after the release period, the cells were washed in phosphate buffered saline (PBS) and fixed with 4% paraformaldehyde for 20 min at room temperature and washed with PBS. The cells were permeabilized with 0.2% Triton X-100 in the PBS for 5 min and washed with the PST wash solution (Triton 0.2%, SVF 0.2% in PBS). Cells were blocked with MAXblock Blocking medium supplemented with 0.1% RNase for 1 h at room temperature, and a washing step was done with the PST solution. Cells were incubated with the primary antibody solution (mouse monoclonal antibody anti-pH3 (Clone CM312, Sigma) and a rabbit polyclonal antibody anti-γH2AX (Clone 20E3, Cell Signaling)) in PST buffer for 120 min. The cells were washed by three successive 5 min washes with the PST solution. Then, secondary detection was carried out using an infrared fluorescent dye conjugated to goat anti-mouse antibody (CF770, Biotium) and a donkey anti-rabbit antibody (CF680, Biotium). For DNA labeling, Sybergold (Invitrogen) was used in conjugation with the secondary antibodies for 1 h. Finally, three successive final washes were performed. DNA, γH2AX and pH3 were visualized simultaneously using the Sapphire Biomolecular Imager (Azure Biosystem, United States).
The γH2AX/pH3 assay can determine cytotoxicity and genotoxicity in a single experiment. To determine cytotoxicity, the DNA content (linked to the number of cells) recorded in the treated cells was compared to the DNA content of the DMSO (control) treated cells and expressed as a percentage and was expressed as relative cell count (% RCC). To determine genotoxicity, γH2AX and pH3 fluorescence in treated wells was divided by untreated well fluorescence to determine the change in phosphorylation level of the selected histone. The treated conditions were compared with the DMSO (control).
BMD analysis
The data obtained in the different experiments were analyzed with the BMD covariate method using the PROAST webtool (version 70.1- https://proastweb.rivm.nl/) and following the manual provided by the European Food Safety Authority (Committee et al. 2017). In this study, we choose a BMR value of 0.5, which corresponds to a 50% increase over the background signal as recommended for in vitro genotoxicity data analysis (Wills et al. 2016; Wheeldon et al. 2020; Sanders et al. 2022).
Statistical analysis
The results are presented as a mean ± standard deviation from the mean (SEM) of at least three separate experiments. Statistically significant increases in biomarkers, following treatment, were compared with the vehicle (DMSO) control using two-sided Student's t-test (*P ≤ 0.05; **P ≤ 0.01). A result was considered positive if there was a 1.5-fold increase of the considered biomarker and a level of cytotoxicity above 50% compared to the DMSO control. The result after 72 h treatment was also compared with the γH2AX and pH3 induction after the 24 h treatment (a and b respectively).
Results
DNA damage after one or three treatments over 24h
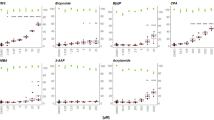
In order to study the genotoxicity of the eight selected compounds, we evaluated the induction of the γH2AX and pH3 biomarkers. We first tested the standard one treatment over 24 h protocol (Fig. 1A). The TUNI compound (a non-genotoxic cytotoxic compound) did not induce any change of the studied biomarkers (supplementary data, Fig. S1A) but a decrease in the viability at the highest concentration was observed. BaP treatment at 1, 3 and 10 µM (Fig. 2A) gave a significant increase of γH2AX compared to DMSO. In a similar way, AFB1 treatment increased γH2AX significantly at concentrations of 0.1, 0.3 and 1 µM. In addition, we observed a decrease of cell viability from 3 µM AFB1 (Fig. 2B). For MMC (Fig. 2C), γH2AX was significantly increased from 1 to 10 µM. For ETO (Fig. 3A), we observed a significant increase of the γH2AX biomarker at 0.1 µM whereas for the CPT (Fig. 3B), we observed a significantly increase of γH2AX at 0.01 µM. At the same time, we used the pH3 marker to study mitotic cell arrest. We observed a significant increase of pH3 from 0.003 µM for taxol (Fig. 4A), compared to DMSO, as well as for the VIN treatment (Fig. 4B). As anticipated for an aneugenic chemical, we did not observe a significant difference of γH2AX compared to the control.
In vitro cytotoxicity and genotoxicity of three genotoxic compounds (benzo[a]pyrene, aflatoxin B1 and mitomycin C) tested for γH2AX and pH3 after one or three repeated treatments. Panel A: Benzo[a]pyrene; Panel B: Aflatoxin B1; Panel C: Mitomycin C. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
In vitro cytotoxicity and genotoxicity of two genotoxic compounds (etoposide and camptothecin) tested for γH2AX and pH3 after one or three repeated treatments. Panel A: Etoposide; Panel B: Camptothecin. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
In vitro cytotoxicity and genotoxicity of two aneugenic compounds (taxol and vinblastin) tested for γH2AX and pH3 after one or three repeated treatments. Panel A: Taxol; Panel B: Vinblastin. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
We then compared the single 24 h acute treatment with a protocol including three treatments over a day (Fig. 1B), with the concentrations previously used for the single dose treatment. After three treatments over 24 h, we did not observe a significant difference for the cytotoxicity and of γH2AX and pH3 biomarkers compared to a single treatment over 24 h (supplementary data Fig S2).
DNA damage after three repeated treatments of 24h
Following the 24 h treatment experiment protocol, we performed three separated treatments over three consecutive days (Fig. 1C). TUNI compound did not give any significant results on the γH2AX or pH3 genotoxicity markers (supplementary data, Fig. S1B). After three repeated treatments over 72 h, we observed a significant increase of γH2AX from 0.1 to 10 µM of BaP compared to the control (Fig. 2A). For AFB1, we observed a significant increase in γH2AX from 0.1 µM (Fig. 2B). For the MMC we observed a significant increase for the γH2AX marker at 0.3 µM (Fig. 2C). ETO gave a significant increase in γH2AX at 0.1 µM (Fig. 3A) whereas for CPT, γH2AX induction was significant at 0.03 µM (Fig. 3B). For topoisomerase inhibitors, we observed a strong significant decrease in cell viability after three days of repeated treatments compared to the 24 h protocol. For VIN and taxol, we observed a significant increase for the pH3 biomarker and this effect was more pronounced after an acute 24 h treatment compared to a three-day repeated protocol (Fig. 4A, B).
Effect of a three-day release period on genotoxicity assessment at the end of the treatment
As expected, for all of the compounds tested in this study (excepted for TUNI), we observed an increase of the γH2AX or pH3 biomarkers (depending on the MoA), indicating genotoxicity induction. Following these DNA insults, the cells may activate some DNA repair pathways. To verify this hypothesis, we tested the effect of a three-day release period. At the end of the cell treatment (after a single or three treatments over 24 h or three days of repeated treatments), cells were incubated in fresh media without any compounds for three days (Fig. 1). For the apoptosis inducer TUNI (supplementary data Fig. 1B), we did not observe any effect of the release period. We observed a significant decrease in the γH2AX markers for BaP (Fig. 5A), AFB1 (Fig. 5B) and MMC (Fig. 5C), but a significant γH2AX induction was always observed at the highest concentrations tested. For ETO and CPT, we observed a significant difference in the DNA damage with a release period compared to no release. After the three-day release period, subsequently to topoisomerase inhibitors treatment, we did not observe any γH2AX induction in contrast to the observed effect at the end of the treatment (Fig. 6). For the two tested aneugens, taxol and VIN, the release period significantly decreases the pH3 induction observed at the end of the 24 h treatment (Fig. 7). Similar effects of the three-day release period were observed after a single or three treatments over 24 h (supplementary data Fig. S3 and S4).
In vitro cytotoxicity and genotoxicity of three clastogenic compounds (benzo[a]pyrene, aflatoxin B1 and mitomycin C) tested for γH2AX and pH3 after three repeated treatments with or without a release period. Panel A: Benzo[a]pyrene; Panel B: Aflatoxin B1; Panel C: Mitomycin C. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
In vitro cytotoxicity and genotoxicity of two clastogenic compounds (etoposide and camptothecin) tested for γH2AX and pH3 after three repeated treatments with or without a release period. Panel A: Etoposide; Panel B: Camptothecin. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
In vitro cytotoxicity and genotoxicity of two aneugenic compounds (taxol and vinblastin) tested for γH2AX and pH3 after treatment with or without a release period. Panel A: Taxol; Panel B: Vinblastin. Cytotoxicity is represented by the % RCC. Each value represents the mean ± SEM (n ≥ 3). Significant differences are noted (*P ≤ 0.05; **P ≤ 0.01; a ≤ 0.05 compared to γH2AX; b ≤ 0.05 compared to pH3)
BMD-covariate analysis classifies genotoxic substances by their mechanism of action
We used the BMD-covariate approach to compare the different compound effects depending on the treatment protocols used (Fig. 8). There was some overlap between the different exposure treatments, indicating no statistical difference between the treatment protocol used. However, we observed a significant lower benchmark concentration (BMC) (between 5 and 10 times lower) for three consecutive repeated treatments compared to one acute treatment for BaP, AFB1 and MMC (Fig. 8A). In contrast, for ETO and CPT, the BMC did not change between one or three repeated treatments (Fig. 8A). For taxol and VIN, we observed a similar BMC for one acute 24 h and three repeated treatments (Fig. 8B). For all the tested chemicals, we observed a significant increase of the BMC after the release period for the same final concentrations tested.
Discussion
The aim of this study was to compare the classical 24 h acute treatment protocol to a repeated exposure over one or three days and the effect of a release period on genotoxicity measurement. We investigated the toxic effects of genotoxic compounds with different MoA: direct and bioactivated clastogenic chemicals, aneugenic compounds and an apoptotic inducer compound. For this purpose, we used the HepaRP cell line, because classical adherent cell models, like HepG2 cells, die after reaching confluence (Khoury et al. 2016a).
We used TUNI (apoptosis inducer) as a negative genotoxic control. As expected, we did not observe a significant induction of γH2AX or pH3 biomarkers after TUNI treatment (Khoury et al. 2020). We classically detected an increase of the γH2AX biomarker for the clastogenic genotoxins BaP, MMC, AFB1, CPT and ETO (Kopp et al. 2019). In contrast, as expected, cells blocked in mitosis after aneugenic compound treatment such as taxol (inhibition of the depolymerization of microtubules) or VIN (inhibition of the polymerization of microtubules) demonstrated an induction of pH3 as observed previously and there was no impact on the γH2AX biomarker (Parry et al. 1996; Bernacki et al. 2019).
In this study, we also compared the cell toxicity after one 24-h treatment or 72-h repeated treatment. We observed differences depending on the MoA of the tested compound (inhibitor of topoisomerase or DNA damaging agent). Direct DNA damaging compounds (BaP, AFB1 and MMC), induced equivalent cytotoxicity after 24 h or 72-h treatment. In contrast, for topoisomerase inhibitors, we observed more pronounced cytotoxicity after a three-day repeated treatment protocol compared to one day acute treatment confirming the results of previous studies (Kopp et al. 2020; Dural et al. 2020).
We also compared a 3 × 8 h treatment and the 24 h treatment. We wanted to test the hypothesis that if we treated the cells three times instead of once in the same day, the toxicity would be higher. However, in our study, we found no difference between the two treatment procedures for cytotoxicity or genotoxicity. The DNA damage is already present after the first treatment, and it seems there was no cumulative effect. Therefore, for the tested chemicals, three treatments in one day correspond to one acute treatment of 24 h for the same final concentration. It may be possible that less stable compound as alkylating agent or oxidative stress inducers produce different results.
DNA damaging agents AFB1, BaP and MMC induced important DNA damage resulting in γH2AX induction as expected. After a three-day release period, the cells treated with these chemicals at a low concentration did not display anymore DNA damage, presumably due to an efficient DNA repair process. However, at the highest tested concentrations, cells always demonstrated some DNA damage as indicated by an elevated γH2AX status, indicating that high DNA damage may persist despite the three-day DNA repair recovery period. Also, this treatment protocol did not induce cytotoxicity at this late time point, indicating that the γH2AX induction was not related to apoptosis. These DNA damaging agents can activate the pathway of senescence linked to the DNA damage response pathway (DDR). It has been demonstrated that if the DNA damage persists, the DDR stays active and may lead to cellular senescence and stop proliferation (Pospelova et al. 2009; Di Micco et al. 2021). Some studies have shown similar results with the marker of senescence Sen-B-Gal and the γH2AX biomarker (Bernadotte et al. 2016; Zhang et al. 2021). In contrast, after the three-day release period after cell treatment with the two DNA topoisomerase inhibitors ETO or CPT, cells did not present anymore DNA damage. This indicated that for these non-direct-DNA damaging agents, cells can either repair their DNA damage properly during the release period, or die. Finally, for aneugenic chemicals, taxol and VIN, we observed the same effect as with the direct DNA damaging agents. At low concentration, cells were able to resume their cell cycle during the release period. However, at higher concentrations, aneugenic chemicals induced an important cell cycle block in mitosis and cells did not totally resume their cell cycle despite the three-day release period.
In this study, for some compounds, we can detect a genotoxic effect at lower concentrations with a repeated treatment compared to one unique regular acute treatment. We know that there are many compounds in food and the environment that can be hazardous (Barsouk et al. 2021), but they are generally tested after an acute 24 h treatment. However, if the population is exposed to these same compounds several times at low doses, there can be an accumulation of DNA damage and possibly mutagenesis induction (Rumgay et al. 2022). Here we show the benefit of using a three-day repeated treatment versus a single 24 h treatment. Using the BMD approach, it was demonstrated that in vitro genotoxicity data may be used for risk assessment and chemical prioritization (Beal et al. 2023). The lowest observed BMC, caused by the repeated treatment protocol, should be taken into account in future studies.
This study was performed with a particular cell type (HepaRP) with specific genotoxic biomarkers (γH2AX and pH3). New experiments should be performed with other cell types, possibly in proliferation status, permitting repeated treatment protocol (like TK6 cells) and with other genotoxic endpoints (mutagenesis, micronucleus) to confirm the conclusion of your study.
The main strength of this study is that we presented an analysis of eight different chemicals with several genotoxic MoA. The new protocol used in this study with repeated treatment helps to differentiate and better understand some mechanisms of action in a more human realistic exposure scenario. Also, with repeated treatment, we gain a better evaluation of the minimal concentration inducing a genotoxic effect. The results of this study demonstrated the utility of performing a three-day repeated in vitro treatment for genotoxicity studies, as proposed in some previously published studies (Quesnot et al. 2016; Kopp et al. 2020; Dural et al. 2020; Sanders et al. 2022). Consequently, with HepaRP or HepaRG cell models, it should be pertinent to use, in parallel of the classical 24 h protocol, a repeated 72 h treatment period for genotoxicity assessment with the γH2AX/pH3 biomarkers, therefore generating more information about the genotoxic MoA. 3D cell culture is another cell model with easily repeatable cell treatment overtime without the confluence toxicity drawback (ter Braak et al. 2022). New experiments with 3D models should be performed to confirm the advantage to using repeated chemical treatment over an acute 24 h period.
Data availability
Not applicable.
References
Aninat C, Piton A, Glaise D et al (2006) Expression of cytochromes P450, conjugating enzymes and nuclear receptors in human hepatoma HepaRG cells. Drug Metab Dispos Biol Fate Chem 34:75–83. https://doi.org/10.1124/dmd.105.006759
Baldwin EL, Osheroff N (2005) Etoposide, topoisomerase II and cancer. Curr Med Chem Anti-Cancer Agents 5:363–372. https://doi.org/10.2174/1568011054222364
Banerjee T, Chakravarti D (2011) A peek into the complex realm of histone phosphorylation. Mol Cell Biol 31:4858–4873. https://doi.org/10.1128/MCB.05631-11
Barsouk A, Thandra KC, Saginala K et al (2021) Chemical risk factors of primary liver cancer: an update. Hepatic Med Evid Res 12:179–188. https://doi.org/10.2147/HMER.S278070
Beal MA, Audebert M, Barton-Maclaren T et al (2023) Quantitative in vitro to in vivo extrapolation of genotoxicity data provides protective estimates of in vivo dose. Environ Mol Mutagen 64:105–122. https://doi.org/10.1002/em.22521
Bell CC, Lauschke VM, Vorrink SU et al (2017) Transcriptional, functional, and mechanistic comparisons of stem cell-derived hepatocytes, HepaRG cells, and three-dimensional human hepatocyte spheroids as predictive in vitro systems for drug-induced liver injury. Drug Metab Dispos Biol Fate Chem 45:419–429. https://doi.org/10.1124/dmd.116.074369
Bernacki DT, Bryce SM, Bemis JC, Dertinger SD (2019) Aneugen molecular mechanism assay: proof-of-concept with 27 reference chemicals. Toxicol Sci 170:382–393. https://doi.org/10.1093/toxsci/kfz123
Bernadotte A, Mikhelson VM, Spivak IM (2016) Markers of cellular senescence: telomere shortening as a marker of cellular senescence. Aging 8:3–11. https://doi.org/10.18632/aging.100871
Bonner WM, Redon CE, Dickey JS et al (2008) GammaH2AX and cancer. Nat Rev Cancer 8:957–967. https://doi.org/10.1038/nrc2523
Brun C, Allain C, Ferron P-J et al (2023) Extended lifespan and improved genome stability in HepaRG-derived cell lines through reprogramming by high-density stress. Proc Natl Acad Sci 120:e2219298120. https://doi.org/10.1073/pnas.2219298120
Brüsehafer K, Rees BJ, Manshian BB et al (2014) Chromosome breakage induced by the genotoxic agents mitomycin C and cytosine arabinoside is concentration and p53 dependent. Toxicol Sci off J Soc Toxicol 140:94–102. https://doi.org/10.1093/toxsci/kfu058
Burma S, Chen BP, Murphy M et al (2001) ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J Biol Chem 276:42462–42467. https://doi.org/10.1074/jbc.C100466200
Cerec V, Glaise D, Garnier D et al (2007) Transdifferentiation of hepatocyte-like cells from the human hepatoma HepaRG cell line through bipotent progenitor. Hepatol Baltim Md 45:957–967. https://doi.org/10.1002/hep.21536
Committee ES, Hardy A, Benford D et al (2017) Update: use of the benchmark dose approach in risk assessment. EFSA J 15:e04658. https://doi.org/10.2903/j.efsa.2017.4658
Di Micco R, Krizhanovsky V, Baker D, d’Adda di Fagagna F (2021) Cellular senescence in ageing: from mechanisms to therapeutic opportunities. Nat Rev Mol Cell Biol 22:75–95. https://doi.org/10.1038/s41580-020-00314-w
Duivenvoorde LPM, Louisse J, Pinckaers NET et al (2021) Comparison of gene expression and biotransformation activity of HepaRG cells under static and dynamic culture conditions. Sci Rep 11:10327. https://doi.org/10.1038/s41598-021-89710-6
Dural E, Shah U-K, Pritchard D et al (2020) The effect of chronic dosing and p53 status on the genotoxicity of pro-oxidant chemicals in vitro. Mutagenesis 35:479–489. https://doi.org/10.1093/mutage/geaa024
Friedberg EC (2003) DNA damage and repair. Nature 421:436–440. https://doi.org/10.1038/nature01408
Guillouzo A, Corlu A, Aninat C et al (2007) The human hepatoma HepaRG cells: a highly differentiated model for studies of liver metabolism and toxicity of xenobiotics. Chem Biol Interact 168:66–73. https://doi.org/10.1016/j.cbi.2006.12.003
Gupta R, Schrooders Y, Hauser D et al (2021) Comparing in vitro human liver models to in vivo human liver using RNA-Seq. Arch Toxicol 95:573–589. https://doi.org/10.1007/s00204-020-02937-6
Jennen DGJ, Magkoufopoulou C, Ketelslegers HB et al (2010) Comparison of HepG2 and HepaRG by whole-genome gene expression analysis for the purpose of chemical hazard identification. Toxicol Sci off J Soc Toxicol 115:66–79. https://doi.org/10.1093/toxsci/kfq026
Jordan MA, Wilson L (1998) Microtubules and actin filaments: dynamic targets for cancer chemotherapy. Curr Opin Cell Biol 10:123–130. https://doi.org/10.1016/s0955-0674(98)80095-1
Khoury L, Zalko D, Audebert M (2016a) Evaluation of four human cell lines with distinct biotransformation properties for genotoxic screening. Mutagenesis 31:83–96. https://doi.org/10.1093/mutage/gev058
Khoury L, Zalko D, Audebert M (2020) Evaluation of the genotoxic potential of apoptosis inducers with the γH2AX assay in human cells. Mutat Res Genet Toxicol Environ Mutagen 852:503165. https://doi.org/10.1016/j.mrgentox.2020.503165
Khoury L, Zalko D, Audebert M (2016b) Complementarity of phosphorylated histones H2AX and H3 quantification in different cell lines for genotoxicity screening. Arch Toxicol. https://doi.org/10.1007/s00204-015-1599-1
Kim SH, Kwon D-Y, Kwak J-H et al (2018) Tunicamycin-induced ER stress is accompanied with oxidative stress via abrogation of sulfur amino acids metabolism in the liver. Int J Mol Sci 19:4114. https://doi.org/10.3390/ijms19124114
Kopp B, Khoury L, Audebert M (2019) Validation of the γH2AX biomarker for genotoxicity assessment: a review. Arch Toxicol 93:2103–2114. https://doi.org/10.1007/s00204-019-02511-9
Kopp B, Le Hégarat L, Audebert M (2020) Differential toxic effects of food contaminant mixtures in HepaRG cells after single or repeated treatments. Mutat Res Genet Toxicol Environ Mutagen 850–851:503161. https://doi.org/10.1016/j.mrgentox.2020.503161
Parry JM, Parry EM, Boumer R et al (1996) The detection and evaluation of aneugenic chemicals. Mutat Res Mol Mech Mutagen 353:11–46. https://doi.org/10.1016/0027-5107(95)00242-1
Pommier Y (2006) Topoisomerase I inhibitors: camptothecins and beyond. Nat Rev Cancer 6:789–802. https://doi.org/10.1038/nrc1977
Pospelova TV, Demidenko ZN, Bukreeva EI et al (2009) Pseudo-DNA damage response in senescent cells. Cell Cycle Georget Tex 8:4112–4118. https://doi.org/10.4161/cc.8.24.10215
Quesnot N, Rondel K, Audebert M et al (2016) Evaluation of genotoxicity using automated detection of γH2AX in metabolically competent HepaRG cells. Mutagenesis 31:43–50. https://doi.org/10.1093/mutage/gev059
Rumgay H, Arnold M, Ferlay J et al (2022) Global burden of primary liver cancer in 2020 and predictions to 2040. J Hepatol 77:1598–1606. https://doi.org/10.1016/j.jhep.2022.08.021
Sanders J, Thienpont A, Anthonissen R et al (2022) Impact of experimental design factors on the potency of genotoxicants in in vitro tests. Mutagenesis 37:248–258. https://doi.org/10.1093/mutage/geac025
Shah U-K, Seager AL, Fowler P et al (2016) A comparison of the genotoxicity of benzo[a]pyrene in four cell lines with differing metabolic capacity. Mutat Res Genet Toxicol Environ Mutagen 808:8–19. https://doi.org/10.1016/j.mrgentox.2016.06.009
Srinivasan A, King RD, Muggleton SH, Sternberg MJE (1997) The predictive toxicology evaluation challenge. In: Proceedings of the 15th international joint conference on Artifical intelligence - vol 1. Morgan Kaufmann Publishers Inc., San Francisco, pp 4–9
ter Braak B, Niemeijer M, Wolters L et al (2022) Towards an advanced testing strategy for genotoxicity using image-based 2D and 3D HepG2 DNA damage response fluorescent protein reporters. Mutagenesis 37:130–142. https://doi.org/10.1093/mutage/geab031
Theumer MG, Henneb Y, Khoury L et al (2018) Genotoxicity of aflatoxins and their precursors in human cells. Toxicol Lett 287:100–107. https://doi.org/10.1016/j.toxlet.2018.02.007
Vogel EW, Nivard MJ (1993) Performance of 181 chemicals in a Drosophila assay predominantly monitoring interchromosomal mitotic recombination. Mutagenesis 8:57–81. https://doi.org/10.1093/mutage/8.1.57
Wheeldon RP, Bernacki DT, Dertinger SD et al (2020) Benchmark dose analysis of DNA damage biomarker responses provides compound potency and adverse outcome pathway information for the topoisomerase II inhibitor class of compounds. Environ Mol Mutagen 61:396–407. https://doi.org/10.1002/em.22360
Wills JW, Johnson GE, Doak SH et al (2016) Empirical analysis of BMD metrics in genetic toxicology part I: in vitro analyses to provide robust potency rankings and support MOA determinations. Mutagenesis 31:255–263. https://doi.org/10.1093/mutage/gev085
Zhang M, Serna-Salas S, Damba T et al (2021) Hepatic stellate cell senescence in liver fibrosis: characteristics, mechanisms and perspectives. Mech Ageing Dev 199:111572. https://doi.org/10.1016/j.mad.2021.111572
Acknowledgements
We acknowledge Biopredic for the use of the HepaRP cell line and specific cell culture media. The authors acknowledge Juliette Cooke for English editing of the manuscript. This work was in part supported by the Genoshift project (ANR-20-CE34-0016).
Author information
Authors and Affiliations
Contributions
CR performed all of the experiments, data analysis and interpretation through discussions with GM and MA. The first draft of the manuscript was written by CR. All the authors discussed the results and contributed to manuscript edition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Recoules, C., Mirey, G. & Audebert, M. Effect of cell treatment procedures on in vitro genotoxicity assessment. Arch Toxicol 98, 1225–1236 (2024). https://doi.org/10.1007/s00204-024-03690-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-024-03690-w