Abstract
Nowadays, the bacteriocin industries have seen significant growth, supplanting chemical preservatives in its ability to improve the shelf-life and safety of food. The increasing customer desire to use natural preservatives has fueled advancing bacteriocin research. The objective of this study was to identify lactic acid bacteria (LAB) that produce bacteriocin-like inhibitory substance (BLIS) and have strong anti-listerial activity. We have identified and analyzed a LAB obtained from chhurpi samples, a popular milk-derived product in the Himalayan regions of India and Nepal. The strain was studied and identified based on its morphological, biochemical, and physiological characteristics. Furthermore, the molecular 16s-rDNA analysis suggests that the strain was Lactococcus sp. RGUAM1 (98.2% similar to Lactococcus lactis subsp. hordniae NBRC 100931T). The isolated strain can produce a potent BLIS, which has shown efficacy against three gram-positive bacteria responsible for food spoilage, such as Listeria monocytogenes (MTCC 657), Staphylococcus aureus subsp. aureus (MTCC 87), Lactobacillus plantarum (MTCC 1407), Lactobacillus paraplantarum (MTCC 12904). The scanning electron microscope (SEM) image illustrates that the crude cell-free supernatant (CFS) disrupts the cell envelope, leading to the release of cellular contents and the clustering of cells. In addition, this BLIS can easily withstand a wide range of pH (2–12), temperature (up to 100 °C for 15 min), bile salt (0.3% W/V), salinity (4% W/V), and enzyme activity of 1600 AU/ml against Listeria monocytogenes. Our research offers a robust framework and valuable insights into bio-preservation and its potential applications in diverse food products.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Food is an organic item that can spoil due to physical, chemical or microbial factors, leading to severe health hazards. According to the World Health Organization (WHO), contaminated food causes the deaths of 420,000 people annually (Mohammad et al. 2018). Therefore, food safety and stability are of utmost importance as a global concern. It is essential to use preservation techniques to maintain the quality and extend the shelf life of food products. The use of bio-preservatives in extending the shelf-life of food is a new and emerging area of research. For this purpose, fermented food products, beneficial bacteria such as lactic acid (LAB) and their metabolites are generally selected to control spoilage and render pathogens inactive (Colombo et al. 2018; Phillips 2022; Sharma et al. 2020). LAB are capable of producing various volatile or non-volatile extrolites such as secondary metabolites, organic acids, vitamins, etc. LAB-originated bioactive compounds like bacteriocin have broad bactericidal activity and considered as good bio-preservative agents due to their non-toxic, thermostable, non-immunogenic nature (Manna and Mondal 2023; Shafique et al. 2022). Most of the LAB bacteriocins act on pathogen cells by interrupting peptidoglycan synthesis or by binds with the Manose phosphotransferase system and forms pores in the bacterial cell membrane through barrel-stave-like pores or carpet mechanism (Scheme 1). Additionally, bacteriocin can disrupt DNA, RNA, and protein biosynthesis (Huang et al. 2021; Pérez-Ramos et al. 2021). Several reports have showed that antimicrobial metabolites produced by Lactococcus lactis exhibit broad inhibitory property towards pathogenic bacteria (Navale et al. 2023). Nisin, a bacteriocin used as a food bio-preservative from long time, was discovered in 1928 and approved by the WHO in 1969. Initially, it was the only bacteriocin used in food bio-preservation. Nisin inhibits growth of pathogenic bacterial cells through wedge-like pattern, which causes leakage of cellular components (Bartenslager 2020; Sharma et al. 2021). Furthermore, other bacteriocins, such as pediocin and micocin, are also used along with nisin (Aljohani et al. 2023). Therefore, our study was focused on the isolation and identification of a BLIS producing Lactococcus sp. RGUAM1 from the traditional cottage cheese “soft chhurpi” (a fermented milk product). In addition, we are trying to explore the partial characterization of BLIS and its impact on various potent pathogens.
Materials and methods
Bacterial strain and culture conditions
The following indicator bacterial strains were considered to identify potent BLIS-producing LAB: Escherichia coli (MTCC 77), Bacillus cereus (MTCC 6728), Klebsiella pneumoniae subsp. pneumoniae (MTCC 39), Staphylococcus aureus subsp. aureus (MTCC 87), Salmonella enterica ser. typhi (MTCC 3216), Enterococcus faecalis (MTCC 3159), Streptococcus mutans (MTCC 497), Yersinia enterocolitica (MTCC 859), and Shigella flexneri (MTCC 1457) were grown in Nutrient media (HiMedia, India); Listeria monocytogenes (MTCC 657) were grown in BHI (Brain Heart Infusion) media; Lactobacillus plantarum (MTCC 1407), Lactobacillus plantarum, Lactobacillus paraplantarum (MTCC 12904) were grown in MRS (de-Man–Rogosa–Sharpe) media (HiMedia, India); Vibrio cholerae (MTCC 3904) were grown in LB (Luria Broth) media (HiMedia, India) as instructed by the source of purchase. The strains were maintained in lyophilized frozen stocks at − 20 °C when not used regularly.
Isolation and purification of bacteria from soft chhurpi
We followed the standard protocol of Chatterjee et al. (2021) and Vanniyasingam et al. (2019) with a few modifications to isolate and purify potent BLIS producing LAB from soft chhurpi procured from local market of Sikkim, India (27° 19′ 42.1′′ N 88° 37′ 04.4′′ E). We inoculated the set amount of chhurpi samples into 5 ml MRS broth (pH 6.5) and incubated the solution anaerobically at 37 °C for overnight. After incubation, the grown sample was centrifuged at 6000g for 10 min to collect the CFS (cell-free supernatant). The CFS was tested as a spot (5 µl) on a BHI agar plate containing a lawn of 0.8% agar of BHI media and seeded with overnight grown culture of L. monocytogenes (MTCC 657). After that, the plates were incubated at 37 °C overnight. The potent BLIS-producing cell appeared as a clear hollow zone. We reassessed the samples to purify the producer strain if a clear zone appeared. We poured the sample onto an MRS plate, and after incubation, each colony that appeared was picked up, mixed with sterilized MRS broth, and incubated again overnight. The colony-containing broths were centrifuged at 6000g for 10 min and spotted on the lawn plate of indicator bacteria. Subsequently, the plates were subjected to overnight incubation in anaerobic condition (CO2 gas jar) at 37 °C, and if the CFS produced by the producer colony resulted in a clear hollow zone, the potent culture was stored in 25% glycerol stock at − 20 °C for further characterization of bacteriocin.
Inoculum preparation
To prepare the inoculum, begin by transferring a single colony into a 50 ml tube containing 10 ml of MRS broth with a pH of 6.5. Following that, it was incubated at 37 °C for 24 h without agitation. To prepare the inoculum, 1% (v/v) of the primary culture was added to a 50 mL tube with 10 mL MRS media. Then, the tube was incubated anaerobically at 37 °C for 24 h. The inoculum utilized in all tests had an optical density of 0.03, measured at 600 nm.
Identification and biochemical characterization of isolated LAB
Vitek-2 compact automated system-based bacterial identification
The bacterial culture was suspended in a polystyrene tube containing 3 ml of 0.9% saline solution. The solution was vortex thoroughly to homogenize it uniformly, and the turbidity of the solution was adjusted with VITEK Densichek (BioMérieux) to match the McFarland 0.5–0.63 standard. Afterwards, the VITEK® 2 GP identification card was placed in the polystyrene tube, and the VITEK 2 system reported the results automatically.
16s rDNA sequence analysis
The isolated strains were cultured in 50 ml of MRS broth at 37 °C overnight. The genomic DNA of the isolate was isolated using the phenol: chloroform extraction technique, followed by Park (2007). The amplification of the 16S ribosomal DNA (rDNA) by Thermal Cycler (BIO-RAD, Hercules, CA, USA) was conducted using the following Universal primers: 27F (5′-AGA GTT TGA TCA TGG CTC AG-3′) and 1492R (5′-AGA GTT TGA TCA TGG CTC AG-3′) (Cho et al. 2023). It was first denatured at 94 °C for 2 min. After that, it was heated to 50 °C for 1 min, annealed at 94 °C for 30 s, and stretched at 72 °C for 1 min. For 7 min, the final expansion was done at 72 °C. The determination of the 16 s rDNA was conducted by Barcode Bioscience Co., located in Bangalore, India. The isolate’s 16S rDNA gene was submitted to the EZBioCloud server (https://www.ezbiocloud.net/) to find the most similar type strains. The ClustalW program in MEGAX software was used to obtain multiple sequence alignments for the closest-type strains of isolated strains downloaded from the EZBioCloud server. MEGA X was used to make a phylogenetic tree using the neighbor-joining method for the alignment file obtained from ClustalW. All of the programs were run with the default set programs. The number of replications for the bootstrap test was modified to 1000, and the Kimura 2-parameter model was chosen as the replacement model (Some et al. 2020).
Determination of growth kinetics and bacteriocin production
The bacterial growth kinetics were examined by introducing the inoculum (1% v/v) of overnight-grown producer culture in a 250-ml flask containing 100 ml of MRS broth. Then, the OD will be measured at 600 nm in a colorimeter for bacterial growth arriving at the stationary phase, estimated at 2-h intervals after the incubation and corresponding to the medium pH, and the antimicrobial activity of crude CFS was tested.
Carbohydrate utilization test
The test for carbohydrate utilization was conducted through the VITEK-2 automated machine.
Lactic acid production test
The isolated strain was streaked on 0.5% calcium carbonate (CaCO3) containing MRS agar (1.5%) plate for the production of hollow zone due to the formation of calcium lactate after incubation for 24 h at 37 °C.
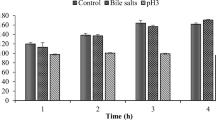
Effect of bile salt
The test was performed with the method followed by Song et al. (2015). The overnight grown culture of isolate was inoculated (1%) in 10 ml fresh MRS broth containing 0.3%, 0.5%, and 1% (w/v) of Ox Gall bile (SRL) at 37 °C. Then the bacterial count was performed for bile tolerance by comparing the initial plate count at 0 h and the final plate count after 24 h.
Microbial adhesion to hydrocarbon assay
Determining microbial adhesion to hydrocarbons (MATH) was conducted using the methodology outlined in Krausova et al. (2019). This approach measures the suspension’s affinity for xylene, a hydrocarbon solvent. Lactococcus sp. RGUAM1 was grown in MRS broth (pH 6.5) overnight at 37 °C and then centrifuged. The cells underwent two rounds of washing with phosphate-buffered saline (pH 7). Following the formation of the bacterial suspension, the absorption was quantified utilizing a spectrophotometer (Double Beam LI-2700, Lasany UV–Vis Spectrophotometer) with 600 nm wavelength. 5 mL of bacterial solution was blended with 1 mL of xylene and then incubated at 37 °C for 1 h. The ultimate determination of absorbance was conducted at a wavelength of 600 nm. The affinity of the suspension for xylene was determined by employing the subsequent formula:
A0 is the OD value of 600 nm initial suspension; A is the OD value of 600 nm suspension after mixed hydrocarbons.
Hemolytic activity assay
To assess the pathogenicity of the isolated Lactococcus sp. RGUAM1, we conducted a hemolytic activity assay. We streaked the test strain and Staphylococcus aureus subsp. aureus MTCC 87 (positive control) on nutrient agar plates supplemented with 5% (w/v) sheep blood and then incubated overnight at 37 °C. We will observe three possible hemolytic activity types: α hemolysis (partial hydrolysis), β hemolysis (clear zone of hydrolysis), and γ hemolysis (no zone around the colonies) (Parveen Rani et al. 2016).
EPS production test
To assess the bacterial exopolysaccharide production, we measured the amount of exopolysaccharides produced by bacteria in a 50 ml liquid medium of MRS (pH 6.5) supplemented with 2% sucrose as a carbon source. The bacteria were cultured anaerobically at 37 °C for three days. After incubation, cells were collected using 1 mm EDTA by adding 500 μl. The mixture was then vigorously mixed until it became uniform and then centrifuged at 9000g for 10 min. The supernatant was isolated from the bacterial cell sediment and mixed with a cold acetone solution at a 1:3 ratio. Subsequently, the combination underwent centrifugation at 15,000g for 5 min. The biomass is deposited as exopolysaccharide, washed with distilled water, and dried at 60 °C for 24 h or until a constant dry weight is achieved (Subair 2015).
NaCl tolerance test
We tested the ability of isolates to grow under NaCl concentrations by adding 1% (v/v) of isolates to 5 mL of MRS broth with varying NaCl concentrations (2%, 4%, 6.5%, 8%, and 10% w/w). These samples were then incubated at 37 °C for 24 h. After incubation the visible growth were observed.
Antimicrobial assessment of BLIS
The antimicrobial properties of the isolates were determined by spot-on lawn assay using different indicator strains:
-
The CFS was neutralized using 1N NaOH.
-
The lawn plate of different indicators (L. monocytogenes, L. planterum, L. paraplantarum, V. cholerae, S. typhy, E. faecalis, Y. enterocolitica, S. flexneri, S. mutans, B. cereus, K. pneumoniae,) with 0.8% agar containing specified media (mentioned by the supplier) was prepared.
-
The CFS was spotted on a lawn plate and incubated at 37 °C for 24 h.
After confirming the spot-on lawn assay result, sensitive indicator strains were analyzed under SEM. In this study, L. monocytogenes (MTCC 657) and S. aureus (MTCC 87) were incubated with CFS for 2 and 5 h. Similarly, L. plantarum (MTCC 1407) also underwent incubation with crude CFS for 1 h and 3 h. The cells were collected after centrifuged at 6000g for 10 min using a cold centrifuge. Subsequently, the cells were subjected to three rounds of washing with a phosphate-buffered saline (0.1 M at pH 7). The cells were then treated with a 2.5% v/v glutaraldehyde (HiMedia, India) and incubated overnight at 4 ºC. Subsequently, the dehydration process was carried out using ethanol solutions of varying concentrations (40%, 50%, 60%, 70%, 80%, 90%, 95%, and 100% v/v). Subsequently, gold coating was applied over the cells, which were examined using a scanning electron microscope (SEM).
Antibiotic sensitivity of isolate
The susceptibility of isolated strain to six distinct classes of antibiotics was assessed using the disc diffusion test followed Jawan et al. (2021). To perform antibiotic susceptibility test, firstly prepare lawn plate of isolated strain using MRS soft agar. After that, antibiotic discs were placed on a plate and incubated at 37 °C for 24 h. The impact of commercially available discs containing levofloxacin (5 mcg), erythromycin (15 mcg), ampicillin (10 mcg), vancomycin (30 mcg), imipenem (10 mcg), and amikacin (30 mcg) on the isolate was investigated by testing. Next, the measurement of inhibitory zones was conducted and, after that, classified into two categories: sensitive (≥ 21 nm) and resistant (≥ 15 nm). The zone diameter for each medication was interpreted according to the guidelines provided by the Clinical and Laboratory Standards Institute. (CLSI 2014; Ripamonti et al. 2011).
Characterization of antimicrobial compound (CFS)
Determination of BLIS concentration
A serial twofold dilution was made with Whatman®-filter (0.22 µm) sterilized bacteriocin solution and sterile water. From each dilution, 5 µl of diluted samples were spotted on a BHI lawn plate (0.8% agar) seeded with L. monocytogenes (MTCC 657), followed by Nielsen et al. (1990). Plates were incubated overnight at 37 °C. In the current study, one arbitrary unit (AU) is quantified using a volume of 5 µl from the most diluted bacteriocin solution that elicits a discernible zone of inhibition on the surface of the indicator organism. The resulting measurement is expressed as AU per milliliter (AU/ml).
Sensitivity towards protein-digesting enzyme
The nature of crude CFS was identified for proteinaceous in nature by using proteolytic enzymes: Trypsin (SRL, India), Chymotrypsin, proteinase K, Protease, Pepsin (HiMedia, India). The stock solution was produced at a concentration of 2 mg/ml. The final concentration of the enzyme treatment after preparation was 1 mg/ml. Next, the enzymatically treated antimicrobial activity was evaluated by spot-on-lawn plate using L. monocytogenes (MTCC 657) as the indicator organism.
Effect of temperature
The effect of temperature on CFS stability was determined by exposing it to different temperatures and time durations. The temperatures used were 100 °C for 10 min and 15 min, 60 °C for 10 min and 15 min, and 37 °C for 10 min and 30 min. The bacteriocin activity was then tested using a spot-on lawn assay against L. monocytogenes (MTCC 657).
Effect of surfactant
The crude CFS was incubated with different surfactants at a final volume of 1% (v/v), including EDTA, SDS, Tween 80, and Tween 20 at 37 °C for 2 h. The antimicrobial activity of untreated CFS was measured and used as a positive control. Then, the activity was tested on spot-on lawn assay against L. monocytogenes (MTCC 657).
Effect of organic solvent
The effect of different organic solvents (Ethanol, Isopropanol, Acetone, methanol) on neutralized CFS was determined at a ratio of 1:1 (v/v). Then, the sample was kept at a temperature of 37 °C for 30 min. After incubation, the mixture was tested on spot-on lawn assay against specific indicator strain. The activity of untreated crude CFS was considered as a positive control. Then, the activity was tested on spot-on lawn assay against L. monocytogenes (MTCC 657).
Effect of different pH
To study the effect of different pH (2–12) on CFS were tested using 1 M HCl and 1 M NaOH and incubated for 2 h at 37 °C (Powell et al. 2007). After incubation, the CFS was adjusted to pH 7 using NaOH and HCl solution. Then, the activity was tested as a zone of inhibition against specific indicator strain on the spot of lawn assay. Then, the activity was tested on spot-on lawn assay against L. monocytogenes (MTCC 657).
Statistical analysis
The studies were conducted in triplicate, and the mean ± standard deviation (SD) data are reported in tables and figures. Microsoft Excel 2021 was used for data analysis and graphical representation.
Results and discussion
Identification of strain
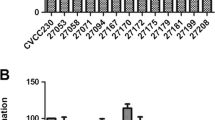
Vitek-2 automated system was identified the strain as Lactococcus sp. (94% probability) based on its biochemical characteristics. Moreover, 16s rDNA sequence revealed that the isolated strain was 98.2% similar (NCBI accession no. OQ947165) to its closely related species Lactococcus lactis subsp. hordniae NBRC 100931T (Fig. 1). Furthermore, the SEM pictures revealed that the isolated strain was coccobacillus in shape and its average diameter was 988 nm (Fig. 2). This aligns with the size range of Lactococcus lactis strains, which generally vary from 0.5 to 1.5 µm (Khelissa et al. 2021).
Growth kinetics and BLIS production
The process of BLIS synthesis commenced promptly after the incubation period, resulting in a steady and optimal production level after 8 h of cultivation in MRS broth (Fig. 3). Initially, the pH level of the media was 6.5. However, during the stationary growth phase, the pH level drops to 4.5 due to the production of lactic acid in the medium. Other LAB bacteriocin-producing strains have also reported similar results, i.e., Lactobacillus plantarum ST16Pa (Du et al. 2018), Lactococcus lactis F01 (Fotso Techeu et al. 2022), and Lactobacillus paraplantarum BT‐11 isolated from raw buffalo milk (Kalhoro et al. 2019). The isolated strain produces bacteriocin in the early log phase as a primary metabolite.
Spectrum of antimicrobial activity
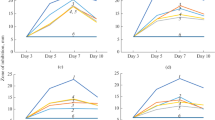
Application of crude CFS of overnight grown culture on different pathogenic strains has inhibited the growth of L. planterum (MTCC 1407), L. paraplantarum (MTCC 12904), L. monocytogenes (MTCC 657), and S. aureus (MTCC 87) (Table 1 and supplementary table T1). The SEM image analysis shows that the cell membrane integrity of pathogenic bacteria (treated cell) significantly changes in the presence of crude CFS in comparison to control (indicator bacterial cells without crude CFS) (Fig. 4). Here, L. plantarum MTCC 1407 membrane has partially disrupted after 12 h of incubation (Fig. 4A1) and complete disruption takes place after 24 h (Fig. 4A2). Similar results were also found in case of L. monocytogens MTCC 657 (Fig. 4B1, B2) and S. aureus MTCC 87 (Fig. 4C1, C2). Previous work has shown comparable investigations demonstrating the inhibitory effects of L. paraplantarum BT‐11 on the development of L. monocytogenes, S. aureus, and Salmonella typhi (Kalhoro et al. 2019). Kalhoro et al. (2023) showed that antimicrobial compounds from buffalo milk effectively suppress foodborne pathogens, including L. monocytogenes, S. typhimurium, S. aureus, and Escherichia coli, due to their antibacterial properties. Pathogenic bacteria are responsible for many foodborne diseases from consuming contaminated food (Chen et al. 2022). Similar studies by Cossettini et al. (2022) showed that E. coli, Salmonella typhi, and Listeria monocytogenes have caused numerous outbreaks and fatalities. The isolate has been found to effectively inhibit the growth of L. monocytogenes and S. typhi by disrupting the cell envelope, known as potent foodborne pathogens.
The scanning electron microscope image of Lactococcus sp. RGUAM1-BLIS mediated time-dependent killing of potent pathogens through cell membrane disruption. [A0—Lactobacillus plantarum MTCC 1407 (Control), A1—Lactobacillus plantarum MTCC 1407 after 12 h of treatment, A2—Lactobacillus plantarum MTCC 1407 after 24 h of treatment; B0—Listeria monocytogenes MTCC 657 (Control), B1—Listeria monocytogenes MTCC 657 after 12 h of treatment, B2—Listeria monocytogenes MTCC 657 after 24 h of treatment; C0—Staphylococcus aureus MTCC 87 (Control), C1—Staphylococcus aureus MTCC 87 after 12 h of treatment, C2—Staphylococcus aureus MTCC 87 after 24 h of treatment]
Biochemical characterization of isolate
The biochemical characterization of the strain is summarized in Table 2. The isolated organism was a gram-positive, catalase-negative lactic acid producing bacteria. The isolated bacteria have been found to exhibit γ hemolytic activity, in comparison to the positive control strain that shows β hemolysis (Supplementary Fig. S1). In vitro assessment of hemolytic activity for probiotics is one of the safety necessities used to assess potential probiotic strains (FAO 2002). Our results are supported by Kim et al. (2022) in which Lactococcus lactis IDCC 2301 exhibited γ hemolytic activity and thus confirmed that, this isolate does not cause any health hazard. Performing antibiotic susceptibility testing is paramount to safely utilizing isolated strains within the food sector. The isolated strain exhibits sensitivity to levofloxacin (DNA replication inhibitor), erythromycin and amikacin (protein synthesis inhibitor), and vancomycin (Cell wall synthesis inhibitor) antibiotics. But the isolated strain was resistant to ampicillin and imipenem (cell wall synthesis inhibitor). Similar studies by Morel et al. (2020) shown that the L. lactis Gh1’s resistance to various antimicrobial agents allows for evaluating its suitability for human and animal consumption in terms of safety. The isolated bacterial strain can utilize different carbon sources, such as lactose, xylose, maltose, D-glucose, raffinose, D-trehalose, melibiose, sucrose, and D-mannose, as energy sources. It can tolerate a growth temperature of 25–37 ºC. The mesophilic bacterium has been found to thrive particularly well at 30 ºC and 37 ºC (Chen et al. 2015). The study investigated the hydrophobicity of Lactococcus sp. RGUAM1’s cell surface was found to be 86% when exposed to xylene. The variability in hydrophobicity, as determined by the MATH technique, may be attributed to the impact of different strains, the period of culture, the specific cultural medium, the presence of acids, and the choice of solvent (Darmastuti et al. 2021). The EPS production capability by isolated bacteria is measured as the dry weight of the EPS. The EPS produced by isolate was 3.17 ± 0.02 mg/ml. The isolated strain grows well in MRS broth with 2% and 4% NaCl, which can resist high osmotic pressure in the gastrointestinal system and maintain a relative osmotic pressure equilibrium. The salt tolerance of bacteria will facilitate to counteract the negative effects of high osmotic pressure inside the gastrointestinal tract (Xu et al. 2019). The isolated strain can grow with a 0.3% bile salt concentration. However, it cannot survive in the presence of 0.5% and 1.0% bile salt concentrations. The ability to tolerate bile salt in the medium is a crucial characteristic of probiotic bacteria (Kumar & Kumar 2015). Similar studies by Ramalho et al. (2019) suggest that L. lactis can tolerate high levels of salt (6% NaCl) and bile salts (0.3%) in a medium, making it suitable for use in food preservation.
Sensitivity to enzymes, pH, temperature, surfactant, and organic solvents
The biochemical characterization of crude CFS is summarized in Table 3. The study included conducting experiments to investigate the impact of trypsin, chymotrypsin, proteinase K, protease, and pepsin on the antibacterial characteristics of crude bacteriocin. The study revealed that all tested enzymes eliminated the bacteriocin’s antimicrobial properties, leading to a 100% reduction in activity. Yildirim and Johnson (1998) reported a similar result, indicating that the bacteriocin produced by Bifidobacterium bifidum NCFB 1454 is susceptible to proteolytic degradation. So, the susceptibility of protein-degrading enzymes to the activity of crude CFS may be attributed to the proteinaceous character of the BLIS.
Moreover, the bacteriocin can withstand pH variations from 2 to 12 and temperatures up to 100 ºC for 15 min. Similar studies by Azhar et al. (2017) shows that, the antimicrobial compound produced by L. lactis A5 can withstand the high temperature (100 °C). De Vuyst and Vandamme (2012) suggest that thermal stability at high temperatures is attributed to several factors, including forming small spherical structures, hydrophobic regions, strong links between molecules, and a high glycine concentration. Generally, bacteriocin was an effective food preservative if it could withstand heat, as numerous food-processing steps entail heating (Ghrairi et al. 2008). Furthermore, the crude bacteriocin activity remains unaltered when exposed to surfactants like Tween 80, Tween 20, EDTA, and SDS and organic solvents such as Acetone, Ethanol, Methanol, and Butanol.
Conclusion
Based on this research, it has been determined that Lactococcus sp. RGUAM1 and its bacteriocin-like inhibitory substances (BLIS) are deemed secure and suitable for utilization in the food sector. This strain exhibits probiotic characteristics, including resistance to bile salts, adaptability to varying salt concentrations in the growing medium, utilization of diverse carbohydrates, and susceptibility to various antibiotics. The BLIS exhibit exceptional stability throughout various pH levels and temperatures, making them valuable preservatives. Using Lactococcus lactis strains with BLIS production capabilities might augment scientific comprehension and provide concrete benefits in several sectors, including food manufacturing, medicinal research, and poultry feed.
References
Aljohani AB, Al-Hejin AM, Shori AB (2023) Bacteriocins as promising antimicrobial peptides, definition, classification, and their potential applications in cheeses. Food Sci Technol. https://doi.org/10.1590/fst.118021
Azhar NS, Zin NHM, Hamid THTA (2017) Lactococcus lactis strain a5 producing nisin-like bacteriocin active against gram positive and negative bacteria. Trop Life Sci Res 28(2):107
Bartenslager A (2020) Investigating microbiomes and developing direct-fed microbials to improve cattle health
Chatterjee M, Jana SC, Raychaudhuri U (2021) Isolation, purification and characterization of a bacteriocin with broad spectrum activity from Lactococcus lactis JC10 from perishable papaya fruit. J Microbiol Biotechnol Food Sci 2021:655–660
Chen J, Shen J, Ingvar Hellgren L, Ruhdal Jensen P, Solem C (2015) Adaptation of Lactococcus lactis to high growth temperature leads to a dramatic increase in acidification rate. Sci Rep 5(1):14199. https://doi.org/10.1038/srep14199
Chen L, Sun L, Zhang R, Liao N, Qi X, Chen J (2022) Surveillance for foodborne disease outbreaks in Zhejiang Province, China, 2015–2020. BMC Public Health 22(1):1–9
Cho Y, Han HT, Kim T-R, Sohn M, Park Y-S (2023) Immunostimulatory activity of Lactococcus lactis LM1185 isolated from Hydrangea macrophylla. Food Sci Biotechnol 32(4):497–506. https://doi.org/10.1007/s10068-022-01199-5
CLSI (2014) Performance standards for antimicrobial susceptibility testing 24th informational supplement. Clinical and Laboratory Standards Institute, Wayne
Colombo M, Castilho NP, Todorov SD, Nero LA (2018) Beneficial properties of lactic acid bacteria naturally present in dairy production. BMC Microbiol 18:1–12
Cossettini A, Vidic J, Maifreni M, Marino M, Pinamonti D, Manzano M (2022) Rapid detection of Listeria monocytogenes, Salmonella, Campylobacter spp., and Escherichia coli in food using biosensors. Food Control 137:108962
Darmastuti A, Hasan PN, Wikandari R, Utami T, Rahayu ES, Suroto DA (2021) Adhesion properties of Lactobacillus plantarum Dad-13 and Lactobacillus plantarum Mut-7 on Sprague Dawley rat intestine. Microorganisms 9(11):2336
De Vuyst L, Vandamme EJ (2012) Bacteriocins of lactic acid bacteria: microbiology, genetics and applications. Springer
Du H, Yang J, Lu X, Lu Z, Bie X, Zhao H, Zhang C, Lu F (2018) Purification, characterization, and mode of action of plantaricin GZ1-27, a novel bacteriocin against Bacillus cereus. J Agric Food Chem 66(18):4716–4724
FAO (2002) WHO working group report on drafting guidelines for the evaluation of probiotics in food. London Ontario Canada 30(1):16–22
FotsoTecheu UD, Kaktcham PM, Momo HK, FokoKouam EM, TchamaniPiame L, Ngouenam RJ, ZambouNgoufack F (2022) Isolation, characterization, and effect on biofilm formation of bacteriocin produced by Lactococcus lactis F01 isolated from Cyprinus carpio and application for biopreservation of fish sausage. Biomed Res Int 2022:8437926
Ghrairi T, Frere J, Berjeaud J, Manai M (2008) Purification and characterisation of bacteriocins produced by Enterococcus faecium from Tunisian rigouta cheese. Food Control 19(2):162–169
Huang K, Zeng J, Liu X, Jiang T, Wang J (2021) Structure of the mannose phosphotransferase system (man-PTS) complexed with microcin E492, a pore-forming bacteriocin. Cell Discovery 7(1):20
Jawan R, Abbasiliasi S, Mustafa S, Kapri MR, Halim M, Ariff AB (2021) In vitro evaluation of potential probiotic strain Lactococcus lactis Gh1 and its bacteriocin-like inhibitory substances for potential use in the food industry. Probiotics Antimicrob Proteins 13:422–440. https://doi.org/10.1007/s12602-020-09690-3
Kalhoro MS, Visessanguan W, Nguyen LT, Anal AK (2019) Probiotic potential of Lactobacillus paraplantarum BT-11 isolated from raw buffalo (Bubalus bubalis) milk and characterization of bacteriocin-like inhibitory substance produced. J Food Process Preserv 43(8):e14015
Kalhoro MS, Anal AK, Kalhoro DH, Hussain T, Murtaza G, Mangi MH (2023) Antimicrobial activities and biopreservation potential of Lactic Acid Bacteria (LAB) from Raw Buffalo (Bubalus bubalis) Milk. Oxid Med Cell Longev. https://doi.org/10.1155/2023/8475995
Khelissa S, Chihib N-E, Gharsallaoui A (2021) Conditions of nisin production by Lactococcus lactis subsp. lactis and its main uses as a food preservative. Arch Microbiol 203:465–480. https://doi.org/10.1007/s00203-020-02054-z
Kim T, Mondal SC, Jeong CR, Kim SR, Ban OH, Jung YH, Yang J, Kim SJ (2022) Safety evaluation of Lactococcus lactis IDCC 2301 isolated from homemade cheese. Food Sci Nutr 10(1):67–74. https://doi.org/10.1002/fsn3.2648
Krausova G, Hyrslova I, Hynstova I (2019) In vitro evaluation of adhesion capacity, hydrophobicity, and auto-aggregation of newly isolated potential probiotic strains. Fermentation 5(4):100. https://doi.org/10.3390/fermentation5040100
Kumar A, Kumar D (2015) Characterization of Lactobacillus isolated from dairy samples for probiotic properties. Anaerobe 33:117–123. https://doi.org/10.1016/j.anaerobe.2015.03.004
Manna A, Mondal R (2023) Bacteriocin-mediated food preservation in conjugation with silver nanoparticles: a green approach. Food Chem Adv. https://doi.org/10.1016/j.focha.2023.100464
Mohammad A-M, Chowdhury T, Biswas B, Absar N (2018) Food poisoning and intoxication: a global leading concern for human health. Food safety and preservation. Elsevier, Amsterdam, pp 307–352
Morel CM, Lindahl O, Harbarth S, de Kraker ME, Edwards S, Hollis A (2020) Industry incentives and antibiotic resistance: an introduction to the antibiotic susceptibility bonus. J Antibiot 73(7):421–428. https://doi.org/10.1038/s41429-020-0300-y
Navale VD, Borade BR, Rama Krishna G, Vamkudoth KR, Kontham R (2023) Metabolites from Lactococcus lactis subsp. lactis: Isolation, structure elucidation, and antimicrobial activity. ACS Omega 8:36628–36635
Nielsen JW, Dickson JS, Crouse JD (1990) Use of a bacteriocin produced by Pediococcus acidilactici to inhibit Listeria monocytogenes associated with fresh meat. Appl Environ Microbiol 56(7):2142–2145. https://doi.org/10.1128/aem.56.7.2142-2145.1990
Park D (2007) Genomic DNA isolation from different biological materials. Protocol Nucleic Acid Anal Nonradioact Probes. https://doi.org/10.1385/1-59745-229-7:3
Parveen Rani R, Anandharaj M, Hema S, Deepika R, David Ravindran A (2016) Purification of antilisterial peptide (Subtilosin A) from novel bacillus tequilensis FR9 and demonstrate their pathogen invasion protection ability using human carcinoma cell line. Front Microbiol 7:1910
Pérez-Ramos A, Madi-Moussa D, Coucheney F, Drider D (2021) Current knowledge of the mode of action and immunity mechanisms of LAB-bacteriocins. Microorganisms 9(10):2107. https://doi.org/10.3390/microorganisms9102107
Phillips MC (2022) Metabolic strategies in healthcare: a new era. Aging Dis 13(3):655. https://doi.org/10.14336/AD.2021.1018
Powell J, Witthuhn R, Todorov S, Dicks L (2007) Characterization of bacteriocin ST8KF produced by a kefir isolate Lactobacillus plantarum ST8KF. Int Dairy J 17(3):190–198. https://doi.org/10.1016/j.idairyj.2006.02.012
Ramalho JB, Soares MB, Spiazzi CC, Bicca DF, Soares VM, Pereira JG, Da Silva WP, Sehn CP, Cibin FW (2019) In vitro probiotic and antioxidant potential of Lactococcus lactis subsp. cremoris LL95 and its effect in mice behaviour. Nutrients 11(4):901
Ripamonti B, Agazzi A, Bersani C, De Dea P, Pecorini C, Pirani S, Rebucci R, Savoini G, Stella S, Stenico A (2011) Screening of species-specific lactic acid bacteria for veal calves multi-strain probiotic adjuncts. Anaerobe 17(3):97–105. https://doi.org/10.1016/j.anaerobe.2011.05.001
Shafique B, Ranjha MMAN, Murtaza MA, Walayat N, Nawaz A, Khalid W, Mahmood S, Nadeem M, Manzoor MF, Ameer K (2022) Recent trends and applications of nanoencapsulated bacteriocins against microbes in food quality and safety. Microorganisms 11(1):85. https://doi.org/10.3390/microorganisms11010085
Sharma R, Garg P, Kumar P, Bhatia SK, Kulshrestha S (2020) Microbial fermentation and its role in quality improvement of fermented foods. Fermentation 6(4):106. https://doi.org/10.3390/fermentation6040106
Sharma K, Kaur S, Singh R, Kumar N (2021) Classification and mechanism of bacteriocin induced cell death: a review: bacteriocin classification and their mode action. J Microbiol Biotechnol Food Sci 11(3):e3733–e3733. https://doi.org/10.15414/jmbfs.3733
Some S, Sarkar B, Biswas K, Jana TK, Bhattacharjya D, Dam P, Mondal R, Kumar A, Deb AK, Sadat A (2020) Bio-molecule functionalized rapid one-pot green synthesis of silver nanoparticles and their efficacy toward the multidrug resistant (MDR) gut bacteria of silkworms (Bombyx mori). RSC Adv 10(38):22742–22757. https://doi.org/10.1039/D0RA03451G
Song M, Yun B, Moon J-H, Park D-J, Lim K, Oh S (2015) Characterization of selected Lactobacillus strains for use as probiotics. Korean J Food Sci Anim Resour 35(4):551. https://doi.org/10.5851/kosfa.2015.35.4.551
Subair H (2015) Isolation and Screening Bacterial Exopolysaccharide (EPS) from potato rhizosphere in highland and the potential as a producer Indole Acetic Acid (IAA). Procedia Food Sci 3:74–81. https://doi.org/10.1016/j.profoo.2015.01.007
Vanniyasingam J, Kapilan R, Vasantharuba S (2019) Isolation and characterization of potential probiotic lactic acid bacteria isolated from cow milk and milk products. AGRIEAST. https://doi.org/10.4038/agrieast.v13i1.62
Xu Y, Tian Y, Cao Y, Li J, Guo H, Su Y, Tian Y, Wang C, Wang T, Zhang L (2019) Probiotic properties of Lactobacillus paracasei subsp. paracasei L1 and its growth performance-promotion in chicken by improving the intestinal microflora. Front Physiol 10:937
Yildirim Z, Johnson MG (1998) Characterization and antimicrobial spectrum of bifidocin B, a bacteriocin produced by Bifidobacterium bifidum NCFB 1454. J Food Prot 61(1):47–51. https://doi.org/10.4315/0362-028X-61.1.47
Acknowledgements
AM is thankful to S.V.M.C.M. scholarship (WBP211618382055), Govt. of West Bengal, India. The authors are thankful to the Mr. Sanjeet Manna and Central Instrumentation Facility (C.I.F.), Odisha University of Agriculture and Technology (O.U.A.T.), Odisha, India, for the scanning electron microscope (S.E.M.) analysis.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
A.M. and S.C.J. designed the initial study; A.M. was involved in writing, image presentation, and table preparation; S.C.J. was involved in manuscript refinement, important intellectual content discussion, and overall supervision; and A.M. and S.C.J. were engaged in reviewing and editing the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest regarding the publication of this article. The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Sunita Varjani.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Manna, A., Jana, S.C. Isolation and characterization of lactic acid bacteria producing a potent anti-listerial bacteriocin-like inhibitory substance (BLIS) from chhurpi, a fermented milk product. Arch Microbiol 206, 73 (2024). https://doi.org/10.1007/s00203-023-03797-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03797-1