Abstract
Lactic acid bacteria (LAB) can take advantage of fermentable carbohydrates to produce lactic acid. They are proverbially applied in industry, agricultural production, animal husbandry, food enterprise, pharmaceutical engineering and some other important fields, which are closely related to human life. For performing the probiotic functions, LAB have to face the low pH environment of the gastrointestinal tract. Therefore, acid resistance of LAB is of great importance not only for their own growth, but also for fermentation and preparation of probiotic products. Recent research studies on acid resistance mechanisms of LAB are mainly focused on neutralization process, biofilm and cell density, proton pump, protection of macromolecules, pre-adaptation and cross-protection, and effect of solutes. In this context, biotechnological strategies such as synthetic biology, genome shuffling, high pressure homogenization and adaptive laboratory evolution were also used to improve the acid resistance of LAB to respond to constantly changing low pH environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactic acid bacteria (LAB) belong to a large category of Gram-positive bacteria that can be found in daily environments as well as in the human gastrointestinal tract (GIT) (Gaspar et al. 2013). Besides, some LAB are economic fermentative bacteria with probiotic properties in GIT (Wu et al. 2009; Koponen et al. 2012; Zhai et al. 2014).
LAB are usually faced with stable low pH or sudden and transient acid stress (Koponen et al. 2012; Wu et al. 2009; Broadbent et al. 2010). During their growth, cellular machinery excretes lactic acid that can be imported back. It lowers the cytoplasmic pH, which can inhibit the growth of cells and may even lead to death (Koponen et al. 2012; Wu et al. 2012a). Several mechanisms are involved in the acid resistance regulation of LAB, including central metabolic pathways, proton pump, changes of cell membrane composition and cell density, DNA and protein damage repair, as well as neutralization processes (Cotter and Hill 2003; Koponen et al. 2012; Wu et al. 2014; Liu et al. 2015).
This article discusses the mechanisms used by LAB for adaptation in low pH environment, and techniques of biotechnology, which have been utilized to enhance the acid resistance of LAB.
Mechanisms of acid resistance
Neutralization processes
Arginine dihydrolase system (ADS)
Microorganisms produce alkaline substances such as urea, arginine and ammonia, which neutralize the acids. Urease hydrolyzes urea into carbon dioxide and ammonia. Arginine dihydrolase also called arginine deiminase is the part of the pathway, named after them as arginine dihydrolase system (ADS) which catalyzes the conversion of arginine into ornithine, ammonia and carbon dioxide.
Generally, ADS contains three enzymes in LAB, including arginine dihydrolase (EC 3.5.3.6), ornithine transcarbamylase (EC 2.1.3.3), and carbamate kinase (EC 2.7.2.2). They are encoded by an operon, arcABC (Maghnouj et al. 1998; Arena et al. 1999, 2002; Champomier et al. 1999). Compared with the wild-type strain Lactobacillus casei Zhang, its acid-resistant mutant displayed higher intracellular aspartate and arginine levels in the acidic environment (Wu et al. 2013a). During acid stress, aspartate enhances the flux of metabolites towards arginine biosynthesis. The acid resistance ability of L. casei was enhanced by the addition of 50 mM arginine or aspartate (Wu et al. 2012a). It indicated that arginine and aspartate are related to acid resistance regulation of strains (Wu et al. 2012a, 2013a; Zhang et al. 2012).
Furthermore, agmatine deiminase system and tyrosine catabolic pathway, which are encoded by the same locus in the chromosome, are beneficial to acid resistance of Lactobacillus brevis (Lucas et al. 2007). It was speculated that ammonia was formed from agmatine through the agmatine deiminase system to neutralize protons in cells to maintain intracellular pH homoeostasis in L. brevis, when encountered with low pH environment (Lucas et al. 2007).
Malolactic fermentation
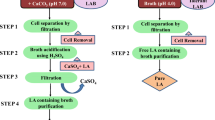
Another promising acid resistance mechanism is malolactic fermentation (Solieri et al. 2010; Broadbent et al. 2010). Malolactic fermentation (MLF) carried out by a variety of LAB, including Oenococcus oeni, and the members of the genera of Lactobacillus, and Leuconostoc etc (Solieri et al. 2010; Broadbent et al. 2010; Bravo-Ferrada et al. 2013). MLF is a decarboxylation of l-malate to yield l-lactic acid. Carbon dioxide is liberated in the process which neutralizes the protons and decreases their concentration (Sumby et al. 2014). MLF improved the survival ability of L. casei ATCC 334 in low pH environment. Reportedly, addition of 30 mM malate can enhance the acid resistance of the strain at pH 2.5 (Broadbent et al. 2010).
Biofilm and cell density
Generally, a group of microorganisms with a protective slimy sheath which is made up of DNA, proteins, and polysaccharides, constitute biofilm. It is the first barrier of the cell and has the ability to withstand environmental disruption. Modifying physicochemical properties of biofilm has proved to be an important survival strategy for many microorganisms (Hall-Stoodley and Stoodley 2009). When faced with low pH environment, there is a rise of membrane mobility and ratio of unsaturated fatty acids, and their mean chain length indicating that changing fluidity of biofilm, distribution of fatty acids, and integrity of cells may be potential methods for LAB to enhance acid resistance (Wu et al. 2012b). Studies have shown that ratio of cyclopropane fatty acids (CFA) remarkably changed in strains under low pH environment, indicating that CFA has a potential role in strains to cope with acidic environment (Broadbent et al. 2010; Wu et al. 2012b).
On the contrary, cyclopropanation of unsaturated fatty acid is found non-essential for acid resistance of Lactobacillus lactis subsp. cremoris and solely CFA could not preserve mobility level of biofilm (To et al. 2011). Hence, it is necessary to further investigate the specific protective effects of CFA under acid stress.
Biofilm formation is not only affected by changing pH, osmotic pressure, carbohydrate concentration in the environment, but also regulated by signaling molecules (Costerton et al. 1995). In Lactobacillus bulgaricus, central metabolic network genes bring about rerouting of pyruvate metabolism to induce modifications of fatty acid composition, and then influence mobility of biofilm, which will help them to overcome various low pH conditions (Fernandez et al. 2008).
Proton pump
F1-F0-ATPase
F1-F0-ATPase can hydrolyze or synthesize intracellular ATP through F1 protein, and transport proton through F0 complex. It is a substantial component of acid tolerance, and its activity is positively related with the more acid resistance in LAB (Kajfasz and Jr 2011). The F1-F0-ATPases of some LAB have been well identified, including Lactococcus lactis, Lactobacillus helveticus, Lactobacillus acidophilus, Lactobacillus rhamnosus, and O. oeni etc (Yokota et al. 1995; Yamamoto et al. 1996; Tourdot-Marechal et al. 1999; Kullen and Klaenhammer 1999; Koponen et al. 2012). Transcriptional levels of atp which encodes F1-F0-ATPase in L. acidophilus were found high when encountered with low pH environment (Kullen and Klaenhammer 1999). Similarly, low pH induced the expression of F1-F0-ATPase genes in L. rhamnosus GG (Koponen et al. 2012). The mutations of F1-F0-ATPases lead to lower survival of LAB at low pH (Yokota et al. 1995; Yamamoto et al. 1996; Tourdot-Marechal et al. 1999). The acid-resistant derivative strain L. casei Lbz-2 displayed greater H+-ATPase activity than its wild-type L. casei Zhang (Wu et al. 2012a). At the same time, it showed a higher intracellular pH than L. casei Zhang at low pH (Wu et al. 2012a).
Amino acid decarboxylation
Another mechanism associated with proton depletion is the amino acid decarboxylation-antiporter reaction (Azcarate-Peril et al. 2004). It can maintain intracellular pH homoeostasis in a decarboxylation reaction by consuming protons. For example, glutamate decarboxylase (GAD) can catalyze the conversion of glutamate to γ-aminobutyrate (GABA), and results in the raising of the intracellular pH (Feehily and Karatzas 2013). The expression of genes encoding GAD in L. acidophilus is strongly increased in gastric juice (Wilson et al. 2014). Four putative genes involved with decarboxylation reactions from L. acidophilus NCFM were disrupted by means of insertional inactivation, including an ornithine decarboxylase, an amino acid permease, a glutamate-aminobutyrate antiporter, and a transcriptional regulator (Azcarate-Peril et al. 2004). All mutants were more sensitive to low pH than the parental strain. The results indicated that the decarboxylation reaction played an important role in the improvement of acid resistance of strains. On the same note acid resistance of L. lactis was enhanced by heterologous expression of histidine decarboxylation pathway with the aid of histidine in acidic environment (Trip et al. 2012).
Protection and repair of cellular macromolecules
Low pH environment brings a selection pressure on LAB for survival fitness. As cytoplasmic pH decreases, the mechanisms to protect the major biological molecules such as DNA and proteins kick in Wu et al. (2012a).
The uvrA gene codes for subunit A of the ultraviolet excinuclease ABC complex, which involves in the nucleotide excision repair mechanism. Its transcriptional activity was activated by exposure to ultraviolet radiation. The expression of uvrA was significantly induced during acid-adaptation in L. helveticus (Cappa et al. 2005). Similarly, a moderate ultraviolet irradiation improved an acid tolerance in Lactococcus lactis subsp. lactis (Hartke et al. 1995). One can conclude that UvrA and nucleotide excision repair pathways have vital functions in the repair process of DNA damage caused by acid and are guarantees for the strains to favorably adapt to the acidic environment (Hartke et al. 1995; Cappa et al. 2005).
Although their precise role in acid adaptation of LAB is not fully understood, some general stress proteins were detected as more abundant in acid-stressed LAB, such as chaperones (DnaK, GrpE, GroEL, and GroES), and small heat shock proteins (Hsp1, Hsp3, and Shsp) etc (De Angelis and Gobbetti 2004; Fernandez et al. 2008; Lee et al. 2008; Wu et al. 2009, 2011, 2012a; Heunis et al. 2014). It is suggested that the molecular chaperones like DnaK can enhance biosynthesis of F1-F0-ATPase, which help the cell to remove protons to maintain intracellular pH homoeostasis (Kim and Batt 1993; Walker et al. 1999). For example, dnaK from E. coli was introduced into L. lactis NZ9000, and it enhanced acid resistance of strain in acidic environment (Abdullah et al. 2010). Heterogeneous expression of a small heat shock protein, Shsp, encoded by shsp from Streptococcus thermophilus in L. lactis ML23 had resulted in higher survival under acid stress (Tian et al. 2012). DNA repair protein RecO, encoded by recO in L. casei, once expressed in L. lactis NZ9000 increased acid resistance of strain under acidic conditions (Wu et al. 2012a, 2013b).
Likewise, accumulation of trehalose and glutathione protects the cellular proteins from acid stress. For example, the trehalose accumulation is response to acid stress in Propionibacterium freudenreichii (Cardoso et al. 2004). In order to study the potential effects of trehalose, its de novo biosynthetic pathway of P. freudenreichii was introduced into L. lactis, as expected, the recombinant strain exhibited higher acid resistance than the control strain (Carvalho et al. 2011). gshA and gshB are related to glutathione biosynthesis in E. coli. Their heterologous expression in L. lactis NZ9000, enhanced the acid resistance of the strain indicating the relationship between glutathione and acid resistance of strains (Zhang et al. 2007). Reportedly, addition of 3–6 mM glutathione can protect LAB from low pH (Kim et al. 2012).
Pre-adaptation and cross-protection
Pre-adaptation is the process of treating a strain to lethal or sub-lethal doses of the stress for a limited time, which accentuates the recovery of the strain when exposed to the natural stress. The mechanism behind pre-adaptation is not well understood. L. casei ATCC 334 was pre-treated for 20 min at a pH value of 4.5, which enhanced the resistance of the strain for low pH (Broadbent et al. 2010). Cross-protection works on the principle that interrelated responses are generated by different stress conditions. In other words, different stimuli like heat, oxygen, cold and low pH may generate similar responses. For example, heat pre-treatment induced an acid resistance response (ATR) in Lactobacillus plantarum which promoted its growth under low pH (De Angelis et al. 2004). Wang et al. obtained high acid tolerance strains by ultraviolet irradiation and heat pre-treatments (Wang et al. 2007). In summary, pre-adaptation and cross-protection are effective methods to strengthen the resistance of LAB against acidic environments. However, exact molecular mechanisms need further research.
Use of protective substances
Addition of protective substances is a comparatively simple and direct method to reduce the damage caused by acidic environment. Many kinds of protective agents are used to resist damage caused by acidic environment in LAB, most of which include amino acids, fatty acids, and sugars. Addition of arginine and aspartate can improve acid resistance of L. casei Zhang under acidic environment (Zhang et al. 2012; Wu et al. 2013a). When Tween-80 was added to the culture of L. rhamnosus, the strain showed 1000-fold higher survival fitness than control (Corcoran et al. 2007). Addition of glutathione also protected LAB from low pH (Kim et al. 2012).
Use of high-throughput techniques for acid resistance in LAB
In recent years, a lot of genomics studies of LAB are completed and published. For example, largest and most diverse genus, Lactobacillus in LAB, contains about 214 genome sequencing projects in public databases (Stefanovic et al. 2017). The genomic studies of LAB have helped to understand their metabolic processes. The same information has been utilized for their application in industry (Zhu et al. 2009; Stefanovic et al. 2017). The advent of genomics provides a possibility to modify the strain for purposeful exploitation on the basis of a more knowledge-based approach (Stefanovic et al. 2017).
Synthetic biology is an interdisciplinary science, which combines many disciplines like genetic engineering, systems biology, biophysics, computer engineering and evolutionary biology (Zhu et al. 2012). Synthetic biology is widely applied in medical and food industries by means of building artificial biological systems. Concurrently, it accelerates our understanding of mechanisms of biology. It offers a new approach for improvement of the acid tolerance of LAB. For example, Bacillus coagulans SIM-7 DSM 14043 is a novel lactic acid producing strain with high acid tolerance (Michelson et al. 2006). Acid-resistant components of B. coagulans SIM-7 DSM 14043 can be sythesized through synthetic biology and transformed into other LAB. Synthetic biology has a great potential to enhance the acid resistance in LAB.
Genome shuffling is an efficient method for the rapid improvement of important microbial phenotypes. It consists of four steps, which are (i) construction of a mutant library by classical strain-improvement methods, (ii) screening of number of positive mutants, (iii) undertaking multiple rounds of protoplast fusion to generate many random mutants and (iv) screening of strains for the expected phenotypes (Stephanopoulos 2002). Obtaining multitrait phenotypes by means of traditional methods is difficult but genome shuffling can engineer such strains in less time (Stephanopoulos 2002). This approach has been used to improve acid tolerance in LAB (Patnaik et al. 2002; Wang et al. 2007; Triratna et al. 2011). New shuffled LAB can grow at substantially lower pH than does the wild-type strain (Patnaik et al. 2002; Wang et al. 2007; Triratna et al. 2011).
In food industry, some non-thermal pasteurization processes have been utilized. These methods like pulsed electric field (PEF) and high pressure homogenization (HPH) do not use heat; therefore, sensorial and nutritional properties of food products are not affected. Amazingly, these processes can improve functional properties of strains (Cueva 2009; Muramalla and Aryana 2011; Tabanelli et al. 2013).
PEF involves the application of pulses of high voltage (20–80 kV/cm) to fluid placed between two electrodes for less than one second (Cueva 2009). The effect of PEF treatment on acid tolerance of L. acidophilus LA-K was evaluated and the results indicated that specific PEF conditions can improve the acid tolerance of the strain (Cueva 2009). In HPH process, the samples in a liquid are homogenized in a range of different pressures. Generally, it is considered that HPH has a close relationship with improvement of sensorial or functional properties of fermented milks and cheeses (Patrignani et al. 2009). In recent years, HPH is used in modification of the functional properties of LAB like acid tolerance and bile tolerance (Muramalla and Aryana 2011). Acid tolerance of L. acidophilus LA-K had been enhanced using HPH at 13.8 MPa (Muramalla and Aryana 2011). The same effects were observed for L. delbrueckii ssp. bulgaricus LB-12 and S. salivarius ssp. thermophilus ST-M5 (Muramalla and Aryana 2011). Similarly, sub-lethal HPH-treated L. paracasei A13 exhibited higher acid resistance compared with controls (Tabanelli et al. 2013). These results indicate that PEF and HPH can be recommended for increasing probiotic characteristics of LAB, including acid tolerance.
Adaptive laboratory evolution (ALE) is a method that explores the natural adaptation of microorganisms over time to the artificial selection pressures given in the laboratory. The different techniques employed in ALE are DNA sequencing, high-throughput screening and gene manipulation (Portnoy et al. 2011). This approach is used in improvement of the acid tolerance of L. casei Zhang (Zhang et al. 2012). The evolved mutant lb-2 was obtained in 70 days with serially exposing exponentially growing strains to low pH conditions. The strain showed a 318-fold higher survival rate than the parental strain at pH 3.3 for 3 h (Zhang et al. 2012).
Conclusion
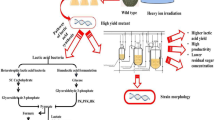
Lactic acid bacteria are always encountered with acidic environment and they have developed various mechanisms to improve their acid resistance (Fig. 1). Emergence of high-throughput techniques brings the improvement in acid resistance of LAB. LAB with high acid resistance had been generated by these approaches. It is important to fully understand the mechanisms of acid resistance in LAB as it will accentuate the benefits of probiotics for humankind.
Mechanisms of acid tolerance in LAB. ADP adenosine diphosphate, AI-2 auto-inducer 2, ALE adaptive laboratory evolution, ATP adenosine triphosphate, CFA cyclopropane fatty acids, Dnak molecular chaperone protein, GABA γ-aminobutyrate, GAD glutamate decarboxylase, HPH high pressure homogenization, LuxS S-ribosylhomocysteinelyase, Nth endonuclease, RecA DNA repair protein, RecO DNA repair protein, Shsp small heat shock protein, SmnA AP endonuclease, TCS two-component signal system, UvrA ultraviolet excinuclease
References
Abdullah AlM, Sugimoto S, Higashi C, Matsumoto S, Sonomoto K (2010) Improvement of multiple-stress tolerance and lactic acid production in Lactococcus lactis NZ9000 under conditions of thermal stress by heterologous expression of Escherichia coli dnaK. Appl Environ Microbiol 76(13):4277–4285
Arena ME, Saguir FM, de Nadra MC (1999) Arginine dihydrolase pathway in Lactobacillus plantarum from orange. Int J Food Microbiol 47:203–209
Arena ME, de Nadra MC, Muñoz R (2002) The arginine deiminase pathway in the wine lactic acid bacterium Lactobacillus hilgardii X 1 B: structural and functional study of the arcABC genes. Gene 301:61–66
Azcarate-Peril MA, Altermann E, Hoover-Fitzula RL, Cano RJ, Klaenhammer TR (2004) Identification and inactivation of genetic loci involved with Lactobacillus acidophilus acid tolerance. Appl Environ Microb 70(9):5315–5322
Bravo-Ferrada B, Hollmann A, Delfederico L, Valdés La Hens D, Caballero A, Semorile L (2013) Patagonian red wines: selection of Lactobacillus plantarum isolates as potential starter cultures for malolactic fermentation. World J Microbiol Biotechnol 29(9):1537–1549
Broadbent JR, Larsen RL, Deibel V, Steele JL (2010) Physiological and transcriptional response of Lactobacillus casei ATCC 334 to acid stress. J Bacteriol 192(9):2445–2458
Cappa F, Cattivelli D, Cocconcelli PS (2005) The uvrA gene is involved in oxidative and acid stress responses in Lactobacillus helveticus CNBL1156. Res Microbiol 156:1039–1047
Cardoso FS, Gaspar P, Hugenholtz J, Ramos A, Santos H (2004) Enhancement of trehalose production in dairy propionibacteria through manipulation of environmental conditions. Int J Food Microbiol 91(2):195–204
Carvalho AL, Cardoso FS, Bohn A, Neves AR, Santos H (2011) Engineering trehalose synthesis in Lactococcus lactis for improved stress tolerance. Appl Environ Microbiol 77(12):4189–4199
Champomier MC, Zúñiga M, Morel-Deville F, Pérez-Martínez G, Zagorec M, Ehrlich SD (1999) Relationships between arginine degradation, pH and survival in Lactobacillus sakei. FEMS Microbiol Lett 180:297–304
Corcoran BM, Stanton C, Fitzgerald GF, Ross RP (2007) Growth of probiotic lactobacilli in the presence of oleic acid enhances subsequent survival in gastric juice. Microbiology 153(1):291–299
Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995) Microbial biofilms. Annu Rev Microbiol 49:711–745
Cotter PD, Hill C (2003) Surviving the acid test: responses of gram-positive bacteria to low pH. Microbiol Mol Biol R 67(3):429–453
Cueva OA (2009) Pulsed electric field influences on acid tolerance, bile tolerance, protease activity and growth characteristics of Lactobacillus acidophilus La-k. Dissertation, Louisiana State University
De Angelis M, Gobbetti M (2004) Environmental stress responses in Lactobacillus: a review. Proteomics 4(1):106–122
De Angelis M, Di Cagno R, Huet C, Crecchio C, Fox PF, Gobbetti M (2004) Heat shock response in Lactobacillus plantarum. Appl Environ Microbiol 70(3):1336–1346
Feehily C, Karatzas KAG (2013) Role of glutamate metabolism in bacterial responses towards acid and other stresses. J Appl Microbiol 114(1):11–24
Fernandez A, Ogawa J, Penaud S, Boudebbouze S, Ehrlich D, van de Guchte M, Maguin E (2008) Rerouting of pyruvate metabolism during acid adaptation in Lactobacillus bulgaricus. Proteomics 8(15):3154–3163
Gaspar P, Carvalho AL, Vinga S, Santos H, Neves AR (2013) From physiology to systems metabolic engineering for the production of biochemicals by lactic acid bacteria. Biotechnol Adv 31(6):764–788
Hall-Stoodley L, Stoodley P (2009) Evolving concepts in biofilm infections. Cell Microbiol 11(7):1034–1043
Hartke A, Bouche S, Laplace J-M, Benachour A, Boutibonnes P, Auffray Y (1995) UV-inducible proteins and UV-induced crossprotection against acid, ethanol, H2O2 or heat treatments in Lactococcus lactis subsp. lactis. Arch Microbiol 163:329–336
Heunis T, Deane S, Smit S, Dicks LM (2014) Proteomic profiling of the acid stress response in Lactobacillus plantarum 423. J Proteome Res 13(9):4028–4039
Kajfasz JK, Jr RGQ (2011) Responses of lactic acid bacteria to acid stress. In: Tsakalidou E, Papadimitrious K (eds) Stress responses of lactic acid bacteria. Springer, New York, pp 23–53
Kim SG, Batt CA (1993) Cloning and sequencing of the Lactococcus lactis subsp. lactis groESL operon. Gene 127:121–126
Kim JE, Eom HJ, Kim Y, Ahn JE, Kim JH, Han NS (2012) Enhancing acid tolerance of Leuconostoc mesenteroides with glutathione. Biotechnol Lett 34(4):683–687
Koponen J, Laakso K, Koskenniemi K, Kankainen M, Savijoki K, Nyman TA et al (2012) Effect of acid stress on protein expression and phosphorylation in Lactobacillus rhamnosus GG. J Proteomics 75(4):1357–1374
Kullen MJ, Klaenhammer TR (1999) Identification of the pH-inducible, proton-translocating F1-F0-ATPase (atpBEFHAGDC) operon of Lactobacillus acidophilus by differential display: gene structure, cloning and characterization. Mol Microbiol 33(6):1152–1161
Lee K, Lee H, Pi K, Choi Y (2008) The effect of low pH on protein expression by the probiotic bacterium Lactobacillus reuteri. Proteomics 8(8):1624–1630
Liu Y, Tang H, Lin Z, Xu P (2015) Mechanisms of acid tolerance in bacteria and prospects in biotechnology and bioremediation. Biotechnol Adv 33(7):1484–1492
Lucas PM, Blancato VS, Claisse O, Magni C, Lolkema JS, Lonvaud-Funel A (2007) Agmatine deiminase pathway genes in Lactobacillus brevis are linked to the tyrosine decarboxylation operon in a putative acid resistance locus. Microbiology 153(7):2221–2230
Maghnouj A, De Sousa TF, Stalon V, Vander Wauven C (1998) The arcABDC gene cluster, encoding the arginine deiminase pathway of Bacillus licheniformis, and its activation by the arginine repressor ArgR. J Bacteriol 180:6468–6475
Michelson T, Kask K, Jõgi E, Talpsep E, Suitso I, Nurk A (2006) L(+)-Lactic acid producer Bacillus coagulans SIM-7 DSM 14043 and its comparison with Lactobacillus delbrueckii ssp. lactis DSM 20073. Enzym Microb Technol 39(4):861–867
Muramalla T, Aryana KJ (2011) Some low homogenization pressures improve certain probiotic characteristics of yogurt culture bacteria and Lactobacillus acidophilus LA-K. J Dairy Sci 94(8):3725–3738
Patnaik R, Louie S, Gavrilovic V, Perry K, Stemmer WPC, Ryan CM, Cardayre S (2002) Genome shuffling of Lactobacillus for improved acid tolerance. Nat Biotechnol 20:707–712
Patrignani F, Burns P, Serrazanetti D, Vinderola G, Reinheimer JA, Lanciotti R et al (2009) Suitability of high pressure-homogenized milk for the productionof probiotic fermented milk containing Lactobacillus paracasei and Lactobacillus acidophilus. J Dairy Res 76:74–82
Portnoy VA, Bezdan D, Zengler K (2011) Adaptive laboratory evolution-harnessing the power of biology for metabolic engineering. Curr Opin Biotechnol 22(4):590–594
Solieri L, Genova F, De Paola M, Giudici P (2010) Characterization and technological properties of Oenococcus oeni strains from wine spontaneous malolactic fermentations: a framework for selection of new starter cultures. J Appl Microbiol 108(1):285–298
Stefanovic E, Fitzgerald G, McAuliffe O (2017) Advances in the genomics and metabolomics of dairy lactobacilli: a review. Food Microbiol 61:33–49
Stephanopoulos G (2002) Metabolic engineering by genome shuffling. Nat Biotechnol 20(7):666–668
Sumby KM, Grbin PR, Jiranek V (2014) Implications of new research and technologies for malolactic fermentation in wine. Appl Microbiol Biotechnol 98(19):8111–8132
Tabanelli G, Patrignani F, Vinderola G, Reinheimer JA, Gardini F, Lanciotti R (2013) Effect of sub-lethal high pressure homogenization treatments on the in vitro functional and biological properties of lactic acid bacteria. LWT Food Sci Technol 53(2):580–586
Tian H, Tan J, Zhang L, Gu X, Xu W, Guo X, Luo Y (2012) Increase of stress resistance in Lactococcus lactis via a novel food-grade vector expressing a shsp gene from Streptococcus thermophilus. Braz J Microbiol 43(3):1157–1164
To TMH, Grandvalet C, Tourdot-Maréchal R (2011) Cyclopropanation of membrane unsaturated fatty acids is not essential to the acid stress response of Lactococcuslactis subsp. cremoris. Appl Environ Microbiol 77(10):3327–3334
Tourdot-Marechal R, Fortier LC, Guzzo J, Lee B, Divies C (1999) Acid sensitivity of neomycin-resistant mutants of Oenococcus oeni: a relationship between reduction of ATPase activity and lack of malolactic activity. FEMS Microbiol Lett 178:319–326
Trip H, Mulder NL, Lolkema JS (2012) Improved acid stress survival of Lactococcus lactis expressing the histidine decarboxylation pathway of Streptococcus thermophilus CHCC1524. J Biol Chem 287(14):11195–11204
Triratna L, Saksono B, Sukmarini L, Suparman A (2011) Genome-shuffling-improved acid tolerance and lactic acid production in Lactobacillus plantarum for commercialization. Microbiol Indones 5(1):4
Walker DC, Girgis HS, Klaenhammer TR (1999) The groESL chaperone operon of Lactobacillus johnsonii. Appl Environ Microbiol 65:3033–3041
Wang YH, Li Y, Pei XL, Yu L, Feng Y (2007) Genome-shuffling improved acid tolerance and l-lactic acidvolumetric productivity in Lactobacillus rhamnosus. J Biotechnol 129:510–515
Wilson CM, Loach D, Lawley B, Bell T, Sims LM, O Toole PW, Zomer A, Tannock GW (2014) Lactobacillus reuteri 100 – 23 modulates urea hydrolysis in the murine stomach. Appl Environ Microbiol 80(19):6104–6113
Wu R, Wang W, Yu D, Zhang W, Li Y, Sun Z et al (2009) Proteomics analysis of Lactobacillus casei Zhang, a new probiotic bacterium isolated from traditional home-made koumiss in Inner Mongolia of China. Mol Cell Proteomics 8(10):2321–2338
Wu R, Zhang W, Sun T, Wu J, Yue X, Meng H et al (2011) Proteomic analysis of responses of a new probiotic bacterium Lactobacillus casei Zhang to low acid stress. Int J Food Microbiol 147(3):181–187
Wu C, Zhang J, Chen W, Wang M, Du G, Chen J (2012a) A combined physiological and proteomic approach to reveal lactic-acid-induced alterations in Lactobacillus casei Zhang and its mutant with enhanced lactic acid tolerance. Appl Microbiol Biot 93(2):707–722
Wu C, Zhang J, Wang M, Du G, Chen J (2012b) Lactobacillus casei combats acid stress by maintaining cell membrane functionality. J Ind Microbiol Biotechnol 39(7):1031–1039
Wu C, Zhang J, Du G, Chen J (2013a) Aspartate protects Lactobacillus casei against acid stress. Appl Microbiol Biot 97(9):4083–4093
Wu C, Zhang J, Du G, Chen J (2013b) Heterologous expression of Lactobacillus casei RecO improved the multiple-stress tolerance and lactic acid production in Lactococcus lactis NZ9000 during salt stress. Bioresour Technol 143:238–241
Wu C, Huang J, Zhou R (2014) Progress in engineering acid stress resistance of lactic acid bacteria. Appl Microbiol Biot 98(3):1055–1063
Yamamoto N, Masujima Y, Takano T (1996) Reduction of membrane bound ATPase activity in a Lactobacillus helveticus strain with slower growth at low pH. FEMS Microbiol Rev 138:179–184
Yokota A, Amachi S, Ishii S, Tomita F (1995) Acid sensitivity of a mutant of Lactococcus lactis subsp. lactis C2 with reduced membrane bound ATPase activity. Biosci Biotechnol Biochem 59:2004–2007
Zhai Z, Douillard FP, An H, Wang G, Guo X, Luo Y, Hao Y (2014) Proteomic characterization of the acid tolerance response in Lactobacillus delbrueckii subsp. bulgaricus CAUH1 and functional identification of a novel acid stress-related transcriptional regulator Ldb0677. Environ Microbiol 16(6):1524–1537
Zhang J, Fu RY, Hugenholtz J, Li Y, Chen J (2007) Glutathione protects Lactococcus lactis against acid stress. Appl Environ Microbiol 73(16):5268–5275
Zhang J, Wu C, Du G, Chen J (2012) Enhanced acid tolerance in Lactobacillus casei by adaptive evolution and compared stress response during acid stress. Biotechnol Bioprocess Eng 17(2):283–289
Zhu Y, Zhang YP, Li Y (2009) Understanding the industrial application potential of lactic acid bacteria through genomics. Appl Microbiol Biotechnol 83:597–610
Zhu L, Zhu Y, Zhang Y, Li Y (2012) Engineering the robustness of industrial microbes through synthetic biology. Trends Microbiol 20(2):94–101
Acknowledgements
This work was supported by National Nature Science Foundation of China (Grant no. 31371827; 31471712).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Additional information
Communicated by Yusuf Akhter.
Rights and permissions
About this article
Cite this article
Wang, C., Cui, Y. & Qu, X. Mechanisms and improvement of acid resistance in lactic acid bacteria. Arch Microbiol 200, 195–201 (2018). https://doi.org/10.1007/s00203-017-1446-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-017-1446-2