Abstract
A novel iron-oxidizing chemolithoautotrophic bacterium, strain ET2T, was isolated from a deep-sea sediment in a hydrothermal field of the Bayonnaise knoll of the Izu-Ogasawara arc. Cells were bean-shaped, curved short rods. Growth was observed at a temperature range of 15–30 °C (optimum 25 °C, doubling time 24 h) and a pH range of 5.8–7.0 (optimum pH 6.4) in the presence of NaCl at a range of 1.0–4.0 % (optimum 2.75 %). The isolate was a microaerophilic, strict chemolithoautotroph capable of growing using ferrous iron and molecular oxygen (O2) as the sole electron donor and acceptor, respectively; carbon dioxide as the sole carbon source; and either ammonium or nitrate as the sole nitrogen source. Phylogenetic analysis based on the 16S rRNA gene sequence indicated that the new isolate was related to the only previously isolated Mariprofundus species, M. ferrooxydans. Although relatively high 16S rRNA gene similarity (95 %) was found between the new isolate and M. ferrooxydans, the isolate was distinct in terms of cellular fatty acid composition, genomic DNA G+C content and cell morphology. Furthermore, genomic comparison between ET2T and M. ferrooxydans PV-1 indicated that the genomic dissimilarity of these strains met the standard for species-level differentiation. On the basis of its physiological and molecular characteristics, strain ET2T (= KCTC 15556T = JCM 30585 T) represents a novel species of Mariprofundus, for which the name Mariprofundus micogutta is proposed. We also propose the subordinate taxa Mariprofundales ord. nov. and Zetaproteobacteria classis nov. in the phylum Proteobacteria.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
“Candidatus (Ca.) Zetaproteobacteria” have recently been recognized as a cosmopolitan group in global marine microbial ecosystems and likely play key roles in biogeochemical processes in many iron-rich redox-cline environments such as sediments and rocks in deep-sea hydrothermal fields and in shallow waters (Emerson et al. 2010). In addition, some “Ca. Zetaproteobacteria” members have been found in association with metallic corrosion moieties (McBeth et al. 2011). However, knowledge of the physiological and molecular characteristics of “Ca. Zetaproteobacteria” is limited because such information has been obtained from only a few isolates from the Loihi Seamount hydrothermal field (Emerson et al. 2007).
The marine iron-oxidizing chemolithoautotrophic bacterial strains PV-1T and JV-1 were isolated by Emerson and Moyer (2002), Emerson et al. (2007) at the Loihi Seamount. Originally, strains PV-1T and JV-1 were classified as Gammaproteobacteria. Later, detailed phylogenetic analysis showed that strains PV-1T and JV-1 possibly represent a novel class of Proteobacteria, “Ca. Zetaproteobacteria”, and belonged to a new species of the new genus Mariprofundus, named Mariprofundus ferrooxydans (Emerson et al. 2007; Moreira et al. 2014). Notably, strain PV-1T forms ribbon-like iron oxyhydroxide stalks, an extracellular material secreted from the cell surface in a helical shape, whereas strain JV-1 does not form stalks. Stalks were first found in Gallionella sp. (Ehrenberg 1836) and became a defining feature. The genus Gallionella belongs to Betaproteobacteria. The genus Leptothrix also excretes extracellular material and is well known as a betaproteobacterial iron-oxidizing bacteria. While Gallionella forms helical stalks, Leptothrix forms tubular sheaths (van Veen et al. 1978; Emerson and Ghiorse 1992, 1993). Although these genera are found in the same environment, they are easily discerned based on their morphological differences under a microscope. Mariprofundus ferrooxydans PV-1 produces helical “stalks” (Emerson and Moyer 2002; Chan et al. 2011; Emerson et al. 2007) that morphologically resemble the stalks of Gallionella species (Ehrenberg 1836; Ghiorse 1984; Hallbeck and Pedersen 1990, 1991). Indeed, similar ribbon-like iron oxyhydroxide structures have been found globally in various iron-dominated deposits in deep-sea hydrothermal fields (Kennedy et al. 2003). Helical stalks in deep-sea hydrothermal environments were long regarded as potential products of deep-sea Gallionella populations (Halbach et al. 2001) until the isolation of M. ferrooxydans (Emerson and Moyer 2002; Emerson et al. 2007).
Recent culture-dependent and culture-independent microbiological characterizations have revealed the ecological concurrence between the distribution of “Ca. Zetaproteobacteria” members and multi-stalked iron-dominating deposits in low-temperature hydrothermal fields such as the Loihi Seamount (Emerson et al. 2007), the Northern Mariana arc (Davis and Moyer 2008), the Southern Mariana trough (Kato et al. 2009), the Vailulu’u seamount (Staudigel et al. 2006), the Kermadec arc (Hodges and Olson 2009) and the South Tonga arc submarine volcanoes (Forget et al. 2010). Mineralogical characterization has suggested that such natural biogenic iron oxyhydroxides, including the stalk structures, could be produced by in situ “Ca. Zetaproteobacteria” populations represented by M. ferrooxydans (Makita et al. 2016). Furthermore, using spatial autocorrelation analysis and molecular variance (AMOVA) analysis, McAllister et al. (2011) revealed the significant biogeographic diversity of “Ca. Zetaproteobacteria”. Although the genome sequence analysis of M. ferrooxidans PV-1T has been completed (Singer et al. 2011), the diversity, physiology and ecology of “Ca. Zetaproteobacteria” are still uncertain because very few strains are available for studies.
Here, we successfully isolated a novel iron-oxidizing strain belonging to “Ca. Zetaproteobacteria” from deep-sea sediment at a hydrothermal field in the Bayonnaise knoll at the Izu-Ogasawara arc. The new isolate was characterized as a mesophilic, neutrophilic, microaerophilic strict chemolithoautotroph. The phylogenetic analysis of its 16S rRNA gene sequence, molecular characteristics and morphological properties indicated that the isolate has features distinct from those of the previously described M. ferrooxidans. Through phenotypic and genomic comparison with related species, we propose a novel species of the genus Mariprofundus, named Mariprofundus micogutta sp. nov. Moreover, the taxonomic name Zetaproteobacteria had not been formally published to date. Therefore, taxonomic ranks between class and order are also proposed in this study.
Materials and methods
Sample collection, enrichment and purification
A deep-sea hydrothermal sediment sample was collected at the Bayonnaise knoll at the Izu-Ogasawara arc (31º57.432′N, 139º44.736′E; 772 m deep) using the JAMSTEC remotely operated vehicle (ROV) Hyper-Dolphin (dive #1649), in April 2014. Many dead chimney structures were present, but active chimney sites hosted diffusing hydrothermal fluids (>130 °C). The active chimneys were grey and orange and less than 20 cm tall. A sample of orange seafloor sediment was collected using an M-type sampler (Masuda et al. 2005). Immediately after recovery of the ROV, the subsample sediment was suspended in filter-sterilized (0.22 µm pore size) natural seawater containing 0.05 % (w/v) sodium sulphide in a 100-ml glass bottle and the bottle was tightly sealed with a butyl rubber cap under N2 atmosphere.
In the laboratory after the cruise, the suspended slurry was inoculated onto an O2 and Fe(II) gradient medium developed by Emerson and Moyer (1997, 2002) and modified in this study for use as an enrichment. In this method, opposite concentration gradients of molecular oxygen (O2) and Fe(II) were formed by two solid medium layers (top and bottom) in the test tube (Fig. S1). The basal medium, a modified medium for ASW (Emerson and Moyer 2002; Emerson and Floyd 2005), was composed of 27.5 g NaCl, 5.38 g MgCl2·6H2O, 6.78 g MgSO4·7H2O, 0.72 g KCl, 0.2 g NaHCO3, 1.4 g CaCl2·2H2O, 1 g NH4Cl and 10 ml trace element solution (MD-TMS; ATCC catalogue no. MD-TMS) per litre of distilled water. After autoclaving, 1 g NaHCO3 and 1 ml vitamin solution (MD-VS; ATCC catalogue no. MD-VS) were added aseptically. The top layer, consisting of agarose (0.15 % (v/v) Agarose S; NIPPON GENE, Tokyo, Japan)-stabilized medium, and the bottom layer, consisting of agarose (1.3 % v/v)-stabilized FeS, were prepared separately. The FeS was prepared according to Hanert (2006) and added as the Fe(II) source. The medium was prepared by purging the ASW medium containing NaHCO3 and vitamin solution with CO2 for an appropriate time to reduce the pH to 6.2–6.5, as determined with a pH meter. Microbial growth was visually confirmed by the formation of a brown colony band in the medium after culturing in the dark for approximately 10 days at 25 and 37 °C.
From a colony on solid gradient medium, which only grew at 25 °C, a pure culture was obtained with the dilution-to-extinction technique (Makita et al. 2012) using liquid medium at 25 °C. For the dilution series, 10 ml of liquid in 20 ml (16.5 φ × 16 cm) test tubes (Iwaki Glass, Tokyo, Japan) and screw caps with butyl rubber septa were used. Liquid medium was obtained by removing agarose from the top layer of the gradient medium described above. The medium was prepared under a gas phase of 80 % N2 and 20 % CO2. The final pH of medium was brought to approximately pH 6.4 by gas purging (measured with a pH meter). Immediately prior to inoculation, a sterile syringe with a 27-gauge needle was used to deliver approximately 200 µm FeCl2 from the stock solution and O2 (100 %, 100–300 µl, final O2 concentrations; 1–3 %) to the headspace. The FeCl2 stock solution was prepared according to Emerson and Moyer (2002). Unless otherwise indicated, growth tests were carried out with the above liquid medium at pH 6.4 and 25 °C. Non-motile bean shapes and curved rods dominated in the cultures. The purified strain was designated as strain ET2T. Purity was confirmed by microscopic observation, repeated partial sequencing of the 16S rRNA gene using several PCR primer sets (Lane 1991) and genome analysis.
Morphology
Cells were routinely examined under a phase-contrast Olympus BX53 microscope with an Olympus DP72 colour CCD camera (Olympus Co. Ltd., Tokyo, Japan). Transmission electron micrographs (TEMs) of negatively stained cells were obtained as previously described (Zillig et al. 1990). The cells grown in liquid medium were harvested by centrifugation at the mid-exponential growth phase and stained with 1 % (w/v) neutral phosphotungstic acid for observation with a Tecnai-20 (FEI Company, Japan) electron microscope at an acceleration voltage of 120 kV. For scanning electron microscopic (SEM) observation, cells were prefixed 2 h in 2.5 % (w/v) glutaraldehyde in liquid medium at 25 °C. After being washed in fresh liquid media, cell suspensions were adhered to the poly-l-lysine (Sigma)-coated tip for 30 min at room temperature. Following this treatment, the cells were fixed in 2 % (w/v) osmium tetroxide dissolved in phosphate-buffered saline (PBS, pH 7.2; Nakarai Tesque, Japan). After rinsing with distilled water, conductive staining was performed by incubation with 0.2 % aqueous tannic acid (pH 6.8) for 30 min. The samples were washed with distilled water, treated with 1 % aqueous osmium tetroxide for 1 h, then dehydrated in a graded ethanol series and dried in a desiccator. The samples were finally coated with osmium using an osmium plasma coater (POC-3; Meiwafosis) and observed with a JSM-6700F field emission scanning electron microscope (JEOL, Japan) operated at 5 kV. Acid treatment (0.1 M HCl, 10 min) was performed to observe the association of cells with iron oxyhydroxides. Energy-dispersive spectrometry (EDS) was used to estimate the elemental composition of extracellular materials. The SEM–EDS and TEM–EDS analyses were conducted at an acceleration voltage of 15 keV for elemental analysis. EDS analyses were conducted using a JSM-6700F field emission SEM (JEOL, Japan) equipped with energy-dispersive X-ray spectroscopy (EDS, JED-2300; JEOL, Japan) and a Tecnai-20 (FEI Company, Japan) electron microscope equipped with EDAX EDS (EDAX Inc., USA).
Growth characteristics
The growth of strain ET2T was examined by direct cell counting after staining with 4,6-diamidino-2-phenylindole (DAPI) using a phase-contrast Olympus BX53 microscope equipped for epifluorescence (Porter and Feig 1980). Before observation, acid treatment (1 M HCl of final concentration, 5 min) was performed to remove iron oxyhydroxides from cells in culture. To determine the temperature, pH and NaCl ranges for growth, strain ET2T was grown in temperature-controlled dry ovens using 20-ml (16.5 φ × 16 cm) test tubes (Iwaki Glass, Tokyo, Japan) containing 10 ml of liquid medium. When the pH range for growth was determined, the pH of the liquid medium was adjusted to various values with 10 mM acetate/acetic acid buffer (pH 4–5), MES (pH 5–6), PIPES (pH 6–7), HEPES (pH 7–7.5), Tris (pH 7.5–8), TAPS (pH 8–8.5), CHES (pH 8.5–9.7) and CAPS (pH 9.7–10) at 25 °C. The pH was readjusted with HCl or NaOH immediately before inoculation. The NaCl requirement for growth was determined using various concentrations of NaCl (0–7.0 %, w/v) in the medium.
The new isolate was tested for its ability to grow with various combinations of electron donors and acceptors. ASW supplemented with 0.1 % (w/v) NaHCO3 and vitamin solution (MD-VS) was used as the basal medium. To examine growth with electron acceptors other than O2, Fe2+ was used as an electron donor under N2/CO2 (80:20) gas phase (250 kPa). The potential electron acceptors S0 (3 %, w/v), Na2S2O3·5H2O (0.1 %, w/v), Na2SO3·5H2O (0.01–0.1 %, w/v), NaNO3 (0.1 %, w/v) and NaNO2 (0.1 %, w/v) were tested. For testing growth on S0 (3 %, w/v), Na2S2O3·5H2O (0.1 %, w/v) as an electron donor, N2/CO2/O2 (77:20:3) was used as the gas phase (250 kPa). We also tested whether strain ET2T could use zero-valent iron (Fe0) for growth. Autoclaved Fe0 particles were utilized as an iron source instead of the FeCl2 solution. Approximately 50 mg of Fe0 particles was added to 10 ml of liquid medium.
To examine heterotrophic growth, experiments were conducted using liquid medium without NaHCO3 under a N2/O2 (97:3) gas phase (250 kPa). Each of the following potential carbon sources was tested at concentrations of 0.01 and 0.1 % (w/v): casamino acids, D(+)-glucose, maltose, formate, acetate, fumarate, lactate, citrate, pyruvate, tryptone peptone (Difco) and yeast extract (Difco).
To determine potential nitrogen sources for growth, utilization of N2 as the nitrogen source was examined with liquid medium lacking all nitrogen compounds under a N2/CO2/O2 (77:20:3) gas phase (250 kPa). After ET2T was confirmed not to use N2 as a nitrogen source, 0.1 % (w/v) NH4Cl, NaNO2, NaNO3 or casamino acids were added to liquid media lacking all other nitrogen compounds under a N2/CO2/O2 (77:20:3) gas phase (250 kPa). The O2 preference for growth was determined by measuring the dissolved O2 concentration at the location of colony band formation in the solid gradient culture using an oxygen sensor (Oxy-N; Unisense, Aarhus, Denmark).
The sensitivity of growth of strain ET2T to antibiotics such as chloramphenicol (50 and 100 μg/ml), streptomycin (50 and 100 μg/ml), kanamycin (50 and 100 μg/ml), ampicillin (50 and 100 μg/ml) and rifampicin (50 and 100 μg/ml) was tested in liquid medium at 25 °C.
Fatty acid analysis
The cellular fatty acid composition was analysed via gas chromatography–mass spectrometer (GC/MS). Cells grown in liquid medium at 25 °C for 5 days (late exponential growth phase) were collected by centrifugation, and the iron hydroxide particles with cells were dissolved with a solution of dithionate (50 g/l) in 0.2 M citrate and 0.35 M acetic acid (Kostka and Luther 1994). The cells were collected by centrifugation and washed with a 2.75 % NaCl solution before lyophilization. The lyophilized cells (1.5 mg) were placed in test tubes containing 3 ml anhydrous methanolic HCl and heated at 100 °C for 3 h. Extraction and analysis of fatty acid methyl esters were as described elsewhere (Komagata and Suzuki 1987; Suzuki et al. 2005). The fatty acid methyl esters (FAMEs) were extracted thrice with n-hexane. Concentrated FAMEs were analysed using a GC/MS (JMS-Q1500GC; JEOL, Japan).
Nucleic acid analyses
Cells in the late exponential growth phase were harvested by centrifugation, and the cell and iron oxide pellet was washed three times with PBS. DNA was extracted using the MO BIO PowerMaxSoil DNA isolation kit (Carlsbad, CA, USA).
The 16S rRNA gene was amplified by PCR using primers of Bac 27F and 1492R (Lane 1991). Amplified 16S rRNA gene fragments directly served as a template for sequencing analysis. The sequence of the 16S rRNA gene was determined using the deoxynucleotide chain-termination method with a 3730XL DNA sequencer (Applied Biosystems, Carlsbad, CA, USA). Sequence similarity analysis of 16S rRNA genes was conducted using BLAST (Altschul et al. 1997; Benson et al. 1998). The phylogenetic trees were constructed by the maximum likelihood (ML) method in the MEGA 5.0 package (Tamura et al. 2011), using the parameters for Jukes–Cantor model distance. Bootstrap confidence of branching was calculated using 1000 replications for the ML tree.
The genome of ET2T was sequenced at FASMAC Co. Ltd., (Atsugi, Japan) using the Illumina MiSeq platform (San Diego, CA, USA). Genome assembly was performed using SPAdes version 3.5.0 (Bankevich et al. 2012), and the average coverage of pair-end reads was 100-fold. Gene annotations of the ET2T assemblies were performed using Prokka version 1.11 (Seemann 2014). Protein annotation was further supported by KASS (KEGG Automatic Annotation Server) (Moriya et al. 2007). Average nucleotide identity (ANI) values were calculated with the OrthoANI algorithm (Lee et al. 2015). Digital DNA–DNA hybridization (DDH) values were determined using the in silico genome-to-genome distance calculator (GGDC2.1; http://ggdc.dsmz.de/distcalc2.php) using the alignment method blast+ (Meier-Kolthoff et al. 2013). Orthologous genes between strain ET2T and M. ferrooxydans PV-1T were identified with a bidirectional BLASTP search, and average amino acid identity (AAI) as well as percentage of conserved proteins (POCP) (Qin et al. 2014) were estimated.
Nucleotide sequence accession number
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of M. micogutta strain ET2T is LC107871. The genome sequence data of this strain are publicly available under the accession numbers BDFD01000001-BDFD01000059.
Results and discussion
Enrichment, purification and morphology
Cell growth was observed in a solid gradient culture after 5–10-day incubation only at 25 °C. Rust-coloured bands, at an intermediate distance between the FeS plug in the bottom of the tube and the air interface at the surface of the gel (Fig. S1), appeared as a brown sparkling granule drops. Once a stable band formed, usually after 10 days, it was subsampled and diluted into a new gradient medium. After the culture was cultivated several times in gradient media, it was transferred to a liquid medium. The non-motile bean-shaped cells and curved rods were purified from the first enrichment using the dilution-to-extinction technique at 25 °C.
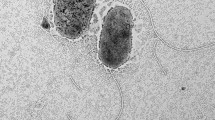
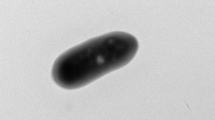
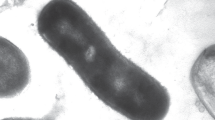
Cells of strain ET2T were gram-negative bean shapes or curved rods, approximately 0.5 μm wide and 1.0–1.6 μm long (Fig. 1a, b). Some cells had polar appendages, similar to prosthecate structures (Fig. 1b), that were 0.13 μm wide and 1.0–23 μm long. Cells occurred singly, and no sporulation was apparent under any culture condition tested.
Electron micrographs of cell strain ET2T. Scanning electron micrographs of a cell (a), cells producing polar prosthecates-like materials (b), cell with extracellular materials and iron oxide (c–f). Transmission electron micrographs of negatively stained single cell (g) and dividing cell (h) with extracellular iron oxyhydroxide filaments. Black arrows indicate extracellular materials. Scale bars indicate 0.1 μm (a), 1.0 μm (b, d–f), 10 μm (c), 0.2 μm (g) and 0.5 μm (h)
Strain ET2T produced many extracellular filamentous structures (Fig. 1c–f), which formed large structures by clumping together (Fig. 1f). These filamentous extracellular materials were morphologically different from the stalks formed by M. ferrooxydans (Emerson et al. 2007). SEM–EDS mapping analysis revealed major peaks for Fe and O and minor peaks for C in the extracellular materials (Fig. 2), suggesting that these materials mainly consist of iron oxides cemented together with trace carbon.
SEM images and elemental maps showing the ET2T cell and extracellular materials with iron oxide. a SEM image showing the cell and extracellular materials. b SEM–EDS image of the cell with extracellular material from a. Also shown are elemental maps: c Carbon shown in red, d Fe shown in green and e both elements overlaid. Scale bars, 0.1 µm (a) and 2 µm (b–e)
Acid treatment of the filamentous materials with 0.1 M HCl resulted in near-complete dissolution, suggesting that they are composed of the same poorly crystalline iron oxyhydroxides as M. ferrooxydans PV-1T (Emerson et al. 2007). After acid treatment, residual filamentous extracellular materials were observed at several locations in the cell (Fig. 1g, h; black arrows). It has been reported that the stalk of PV-1 develops from a specific location in the cell (Emerson et al. 2007; Chan et al. 2011). TEM-EDS analysis revealed that the extracellular materials consisted mainly of carbon (Fig. S4). Interestingly, a polar appendage was no longer observed (possibly degraded by the acid treatment). The filaments produced by strain ET2T may be an excretion mechanism of iron minerals, as in the stalks of M. ferrooxydans PV-1 and Gallionella sp.
Growth characteristics
In the liquid media, orange-brown deposits were produced in the bottom of test tubes during growth. The growth curves of strain ET2T, shown in Fig. S3, indicate growth from 15 to 30 °C, with optimal growth at 25 °C. No growth was observed below 10 °C or above 35 °C. Growth occurred between pH 5.8 and pH 7.0, with an optimum at approximately pH 6.4. No growth was detected below pH 5.0 or above pH 8.3. The isolate showed an absolute requirement for NaCl at concentrations from 1.5 to 4.0 % with optimal growth at 2.75 %. No growth was observed below 1.0 % NaCl or above 4.5 % NaCl. The generation time and maximum cell yield at 25 °C were approximately 24 h and 3.0 × 107 cells/ml, respectively.
No combination other than Fe2+ and O2 supported the growth of strain ET2T, but growth was observed on zero-valent iron (Fe0). The isolate was unable to use sulphur compounds of thiosulfate, bisulphite and elemental sulphur as either electron donors or acceptors and did not use nitrite or nitrate as electron acceptors, indicating that the new isolate grows via simple energy metabolism of iron-oxidizing O2 reduction.
Strain ET2T could not utilize any organic substrates as carbon sources using Fe2+ and O2 as the electron donor and acceptor. In addition, no organic compounds sustained the growth of strain ET2T as the electron donors instead of Fe2+. The isolate ET2T utilizes ammonium and nitrate as the sole nitrogen sources and could not utilize nitrite, N2 or casamino acid. The optimal O2 concentration for growth of ET2T in the solid gradient medium was 2.6–2.8 mg/L, as determined by the oxygen sensor Oxy-N. Thus, strain ET2T represents microaerophilic growth.
Antibiotics such as chloramphenicol (50 and 100 μg/ml), streptomycin (50 and 100 μg/ml), kanamycin (50 and 100 μg/ml), ampicillin (50 and 100 μg/ml) and rifampicin (50 and 100 μg/ml) completely inhibited the growth of strain ET2T.
Genome sequencing
A draft genome of strain ET2T was obtained via genome sequencing and assembly. The assemblies consisted of 59 contigs (total length of 2.497 Mbp), and the G+C content was 48.8 %. The ET2T genome coded 2417 protein-coding sequences (CDSs). Two copies of the rRNA operons are encoded in the genome of strain ET2T. A complete CDS set for carrying out carbon fixation using the Calvin–Benson–Bassham cycle was identified.
16S rRNA gene phylogenetic analysis
The 16S rRNA gene sequence of strain ET2T was similar to the sequences of M. ferrooxydans strain PV-1T (95 %), Mariprofundus sp. strain JV-1 (95 %) (Emerson et al. 2007) and Mariprofundus sp. strain GSB-2 (94 %) (McBeth et al. 2011). The most similar sequences were environmental clones obtained from a colonization system deployed in an observatory at the Juan De Fuca ridge flank (FLOCS_1301A-GWPyrrhotite_C01; 97 %) (Baquiran et al. 2016) and from an inactive hydrothermal sulphide chimney at the mid-ocean ridge (MOR) spreading centre (3M34_081; 97 %) (Sylvan et al. 2012). The phylogenetic tree was constructed with 1329 homologous sequence positions and indicated that strain ET2T is distinct from previously isolated strains of M. ferrooxydans within “Ca. Zetaproteobacteria” (Fig. 3).
Genomic comparison with other “Ca. Zetaproteobacteria”
Compared with M. ferrooxydans PV-1T, strain ET2T harbours a smaller genome with lower G+C content (2.99 Mb and 54 %, respectively) (Singer et al. 2011) (Table 2). The 16S rRNA gene similarity between strain ET2T and M. ferrooxydans PV-1T fell within the common index for genus-level differentiation (90–96 %) (Gillis et al. 2001). Genomic comparison with ANI and DDH calculations, however, indicated that genomic similarities between strain ET2T and M. ferrooxydans strain PV-1T were sufficient only for the threshold of species-level differentiation (95–96 % ANI, 70 % eDDH (Richter and Rosselló-Móra. 2009); 71.6 % of ANI and 13.4–19.8 % of DDH). The POCP value between ET2T and M. ferrooxydans PV-1T was 67.7 %, and the AAI of the shared CDSs was 68.5 %. These genomic similarity values are higher than the threshold of genus-level differentiation (50 %) (Qin et al. 2014). Combined, these results suggest that strain ET2T should be characterized as a new species in the genus Mariprofundus within “Ca. Zetaproteobacteria”.
Fatty acid composition
The major cellular fatty acid components of strain ET2T were C16:1ω7c (51.5 %), C16:0 (27.1 %), C18:0 (8.1 %), C12:0 (7.7 %) and C16:1ω7t (5.6 %). Mariprofundus ferrooxydans strain PV-1T contains iso-C11:0-3OH as the most abundant component of cellular fatty acids (Emerson et al. 2007). Thus, the two strains differ greatly in fatty acid content (Table 1).
Phylogenetic tree based on the 16S rRNA gene sequences of representative strains and environmental clones within the “Ca. Zetaproteobacteria”, including strain ET2T. Aquifex pyrophilus strain Kol 5aT was used as an outgroup. Branch values indicate the bootstrap confidence of branching in per cent. Numbers in parentheses are GenBank/EMBL/DDBJ database accession numbers. Bar indicates 1 substitution per 100 nucleotide positions
Phenotypic comparison with Mariprofundus ferrooxydans
The 16S rRNA gene phylogeny and genomic comparison indicate that strain ET2T is a member of the genus Mariprofundus. Many physiological and metabolic properties are very similar between M. ferrooxydans and the new isolate. Mariprofundus ferrooxydans is a strict chemolithoautotroph, and it cannot grow on other inorganic energy sources, such as reduced sulphur compounds and H2, or on organoheterotrophic substrates. Strain ET2T is also a strict chemolithoautotroph that grows only with iron-oxidizing O2 reduction and inorganic carbon. However, strain ET2T shows different cellular fatty acid composition and genomic DNA G+C content from M. ferrooxydans strain PV-1T (Table 1). More importantly, strain ET2T has distinctive morphological features (Fig. 1; Table 1). Strain ET2T forms appendages similar to those of Caulobacter spp. (Abraham et al. 1999) and Hyphomicrobium spp. (Abraham and Rhode 2014) in the class Alphaproteobacteria and is the first example of a prosthecate bacterium in “Ca. Zetaproteobacteria”. The multiple iron oxyhydroxide filaments associated with ET2T are morphologically different from the stalks composed of organic compounds and iron oxyhydroxides in the M. ferrooxydans strain PV-1T. The cell division style is also quite different between strain ET2T and M. ferrooxydans strain PV-1T.
On the basis of these phylogenetic, genomic and phenotypic characteristics, we conclude that strain ET2T represents a novel species within the Mariprofundus and propose a novel species named Mariprofundus micogutta sp. nov. Furthermore, the class “Ca. Zetaproteobacteria” containing the family Mariprofundaceae (Moreira et al. 2014) is a novel and distinctive taxon in the phylum Proteobacteria. However, the taxonomic name Zetaproteobacteria had not been formally published to date. The presence of “Ca. Zetaproteobacteria” has been confirmed in recent years a various environments (Emerson et al. 2007; Davis and Moyer. 2008; Kato et al. 2009; Staudigel et al. 2006; Hodges and Olson 2009; Forget et al. 2010). Moreover, the analysis of their genome is also progressing (Emerson et al. 2010; Singer et al. 2011). For the development of further research for these iron utilization microorganisms, there is a need to formally define their class and order. Therefore, taxonomic ranks between class and order are also proposed in this study.
Description of Mariprofundus micogutta sp. nov
Mariprofundus micogutta (mic.o.gutta. L. intr. mico sparkle. twinkle; L. f. gutt.a drop, spots; the type strain produces sparkling spot-like colonies under culture).
Cells are bean-shaped or curved rods of approximately 1.0–2.0 μm in diameter. Cells produce polar prosthecates-like materials, and appendages are 0.13 μm wide and 1.0–23 μm long. The temperature range for growth is 15–30 °C (optimum 25 °C). The pH range for growth is pH 5.8–7.0 (optimum pH 6.4). The NaCl concentration range for growth is 1.0–4.0 % (optimum 2.75 %). Fe2+ and Fe0 are oxidized to Fe3+ during growth. Nitrate or ammonium is required as a nitrogen source. Organic compound is not available for their growth. The major fatty acids are C16:1ω7c (51.5 %), C16:0 (27.1 %), C18:0 (8.1 %), C12:0 (7.7 %) and C16:1ω7t (5.6 %). The G+C content of DNA is 48.8 mol %.
The type strain is ET2T (= KCTC 15556T = JCM 30585T), isolated from deep-sea sediment associated with hydrothermal activity at the Bayonnaise knoll of the Izu-Ogasawara Arc.
Description of Mariprofundales ord. nov
Mariprofundales (Mar.is.pro.fund’a.les. NL. masc. n. Mariprofundus type genus of the order; suff. -ales ending to denote an order; N. L. fem. pl. n. Mariprofundales the order of the genus Mariprofundus). The type genus is Mariprofundus (Emerson et al. 2007). The order is defined on the basis of a phylogenetic analysis of the 16S rRNA gene sequence analysis.
Description of Zetaproteobacteria classis nov
Zetaproteobacteria (Gr. n. zeta name of sixth letter of Greek alphabet; N.L. masc. n. Mariprofundus (Emerson et al. 2007) type genus of the family; N. L. fem. Pl. n. Mariprofundaceae (Moreira et al. 2014) the type family of the type order of the class; N. L. fem. pl. n. Mariprofundales the order of the class Zetaproteobacteria).
The class is defined on the basis of a phylogenetic analysis of 16S rRNA gene sequences of four isolated strains and uncultured representatives from various environments. The type genus: Mariprofundus. The type family: Mariprofundaceae. The type order: Mariprofundales.
References
Abraham WR, Rhode M (2014) The family Hyphomonadaceae. In: Rosenberg E et al (eds) The Prokaryotes. Springer, New York, pp 283–299
Abraham WR, Strömpl C, Meyer H, Lindholst S, Moore ER, Christ R, Vancanneyt M, Tindall BJ, Bennasar A, Smit J, Tesar M (1999) Phylogeny and polyphasic taxonomy of Caulobacter species. Proposal of Maricaulis gen. nov. with Maricaulis maris (Poindexter) comb. nov. as the type species, and emended description of the genera Brevundirnonas and Caulobacter. Int J Syst Bacteriol 49(3):1053–1073
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its application to single-cell sequencing. J Comput Biol 19:455–477. doi:10.1089/cmb.2012.0021
Baquiran JP, Ramírez GA, Haddad AG, Toner BM, Hulme S, Wheat CG, Edwards KJ, Orcutt BN (2016) Temperature and redox effect on mineral colonization in Juan de Fuca Ridge flank subsurface crustal fluids. Front Microbiol 7:396. doi:10.3389/fmicb.2016.00396 eCollection 2016
Benson DA, Boguski MS, Lipman DJ, Ostell J, Ouellette BF (1998) Genbank. Nucleic Acids Res 26:1–7
Chan CS, Fakra SC, Emerson D, Fleming EJ, Edwards KJ (2011) Lithotrophic iron-oxidizing bacteria produce organic stalks to control mineral growth: implications for biosignature formation. ISME J 5:717–727
Davis RE, Moyer C (2008) Extreme spatial and temporal variability of hydrothermal microbial mat communities along the Mariana Island Arc and southern Mariana back-arc system. J Geophys Res 113:B08S15. doi:10.1029/2007JB005413
Emerson D, Floyd MM (2005) Enrichment and isolation of iron-oxidizing bacteria at neutral pH. Methods Enzymol 397:112–123
Emerson D, Moyer C (1997) Isolation and characterization of novel iron-oxidizing bacteria that grow at circumneutral pH. Appl Environ Microbiol 63:4784–4792
Emerson D, Moyer C (2002) Neutrophilic Fe-oxidizing bacteria are abundant at the Loihi Seamount hydrothermal vents and play a major role in Fe oxide deposition. Appl Environ Microbiol 68:3085–3093
Emerson D, Rentz JA, Lilburn TG, Davis RE, Aldrich H, Chan C, Moyer CL (2007) A novel lineage of proteobacteria involved in formation of marine Fe-oxidizing microbial mat communities. PLoS One 2:e667
Emerson D, Fleming EJ, McBeth JM (2010) Iron-oxidizing bacteria: an environmental and genomic perspective. Annu Rev Microbiol 64:561–583
Emerson D, Field EK, Chertkov O, Davenport KW, Goodwin L, Munk C, Nolan M, Woyke T (2013) Comparative genomics of freshwater Fe-oxidizing bacteria: implications for physiology, ecology, and systematics. Front Microbiol 4:254. doi:10.3389/fmicb.2013.00254 eCollection2013
Forget NL, Murdock SA, Juniper SK (2010) Bacterial diversity in Fe-rich hydrothermal sediments at two South Tonga Arc submarine volcanoes. Geobiology 8:417–432. doi:10.1111/j.1472-4669.2010.00247.x
Gillis M, Vandamme P, De Vos P, Swings J, Kersters K (2001) Polyphasic taxonomy. In: Boone DR, Castenholz RW, Garrity GM (eds) Bergey’s manual of systematic bacteriology (The archaea and the deeply branching and phototrophic bacteria), vol 1, 2nd edn. Springer, New York, pp 43–48
Hallbeck L, Pedersen K (1990) Culture parameters regulating stalk formation and growth rate of Gallionella ferruginea. J Gen Microbiol 136:1675–1680
Hallbeck L, Pedersen K (1991) Autotrophic and mixotrophic growth of Gallionella ferruginea. J Gen Microbiol 137:2657–2661
Hallbeck L, Ståhl F, Pedersen K (1993) Phylogeny and phenotypic characterization of the stalk-forming and iron-oxidizing bacterium Gallionella ferruginea. J Gen Microbiol 139:1531–1535
Hanert HH (2006) The genus Gallionella. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stacke-Brandt E (eds) The Prokaryotes, vol 7. Springer, New York, pp 990–995
Hodges TW, Olson JB (2009) Molecular comparison of bacterial communities within iron-containing flocculent mats associated with submarine volcanoes along the Kermadec Arc. Appl Environ Microbiol 75:1650–1657
Kato S, Kobayashi C, Kakegawa T, Yamagishi A (2009) Microbial communities in iron-silica-rich microbial mats at deepsea hydrothermal fields of the Southern Mariana Trough. Environ Microbiol 11:2094–2111
Kato S, Chan C, Itoh T, Ohkuma M (2013) Functional gene analysis of freshwater iron-rich flocs at circumneutral pH and isolation of a stalk-forming microaerophilic iron-oxidizing bacterium. Appl Environ Microbiol 79:5283–5290
Kennedy CB, Scott SD, Ferris FG (2003) Ultra structure and potential sub-seafloor evidence of bacteriogenic iron oxides from Axial Volcano, Juan de Fuca Ridge, north-east Pacific Ocean. FEMS Microbiol Ecol 43:247–254. doi:10.1111/j.1574-6941.2003.tb01064.x
Komagata K, Suzuki K (1987) Lipid and cell-wall analysis in bacterial systematics. Method Microbiol 19:161–207
Kostka JE, Luther GW III (1994) Partitioning and speciation of solid phase iron in saltmarsh sediments. Geochim Cosmochim Acta 58:1701–1710
Lane DJ (1991) 16S/23S rRNA sequencing. In: Dworkin M, Falkow S, Stackebrandt E, Goodfellow M, Chichester (eds) Nucleic acid techniques in bacterial systematics. Wiley, Hoboken, pp 115–175
Lee I, Kim YO, Park SC, Chun J (2015) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol. doi:10.1099/ijsem.0.000760
Makita H, Nakagawa S, Miyazaki M, Nakamura K, Inagaki F, Takai K (2012) Thiofractor thiocaminus gen. nov., sp. nov., a novel hydrogen-oxidizing, sulfur-reducing epsilonproteobacterium isolated from a deep-sea hydrothermal vent chimney in the Nikko Seamount field of the northern Mariana Arc. Arch Microbiol 194(9):785–794. doi:10.1007/s00203-012-0814-1
Makita H, Kikuchi S, Mitsunobu S, Takaki Y, Yamanaka T, Toki T, Noguchi T, Nakamura K, Abe M, Hirai M, Yamamoto M, Uematsu K, Miyazaki J, Nunoura T, Takahashi Y, Takai K (2016) Comparative analysis of microbial communities in iron-dominated flocculent mats in Deep Sea hydrothermal environments. Appl Environ Microbiol. doi:10.1128/AEM.01151-16
Masuda H, Fryer P, Ishibashi J, Toki T, Shitashima K, Kimura H, Suzuki R, Sato S, Takemoto K, Kato S, Kobayashi C, Kuno M, Noguchi T, Aoki M (2005) Yokosuka YK05-09 Leg2 Cruise report. http://www.godac.jamstec.go.jp/catalog/data/doc_catalog/media/YK05-09_leg2_all.pdf
McAllister SM, Davis RE, McBeth JM, Tebo BM, Emerson D, Moyer CL (2011) Biodiversity and emerging biogeography of the neutrophilic iron-oxidizing Zetaproteobacteria. Appl Environ Microbiol 77(15):5445–5457. doi:10.1128/AEM.00533-11
McBeth JM, Little BJ, Ray RI, Farrar KM, Emerson D (2011) Neutrophilic iron-oxidizing “Zetaproteobacteria” and mild steel corrosion in nearshore marine environments. Appl Environ Microbiol 77:1405–1412
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. doi:10.1186/1471-2105-14-60
Moreira APB, Meirelles PM, Thompson F (2014) The family Mariprofundaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The Prokaryotes: Deltaproteobacteria and Epsilonproteobacteria, Springer, pp 403–413. doi:10.1007/978-3-642-39044-9_378
Moriya Y, Itoh M, Okuda S, Yoshizawa A, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:W182–W185
Porter KG, Feig YS (1980) The use of DAPI for identifying and counting microflora. Limnol Oceanogr 25:943–948
Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069
Singer E, Emerson D, Webb EA, Barco RA, Kuenen JG, Nelson WC, Chan CS, Comolli LR, Ferriera S, Johnson J, Heidelberg JF, Edwards KJ (2011) Mariprofundus ferrooxydans PV-1 the first genome of a marine Fe(II) oxidizing Zetaproteobacterium. PLoS One 6(9):e25386. doi:10.1371/journal.pone.0025386
Staudigel H, Hart SR, Pile A, Bailey BE, Baker ET, Brooke S, Connelly DP, Haucke L, German CR, Hudson I, Jones D, Koppers AA, Konter J, Lee R, Pietsch TW, Tebo BM, Templeton AS, Zierenberg R, Young CM (2006) Vailulu’u Seamount, Samoa: life and death on an active submarine volcano. Proc Natl Acad Sci USA 103(17):6448–6453. doi:10.1073/pnas.0600830103
Sylvan JB, Toner BM, Edwards KJ (2012) Life and death of deep-sea vents: bacterial diversity and ecosystem. Mbio 3(1):e00279. doi:10.1128/mBio.00279-11
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
van Veen WL, Mulder EG, Denema MH (1978) The Sphaerotilus-Leptothrix group of bacteria. Microbiol Rev 42:329–356
Zillig W, Holz I, Janekovic D, Janekovic D, Klenk HP, Imsel E, Trent J, Wunderl S, Forjaz VH, Coutinho R, Ferreira T (1990) Hyperthermus butylicus, a hyperthermophilic sulfur-reducing archaebacterium that ferments peptides. J Bacteriol 172:3959–3965
Acknowledgments
We thank the captain and crew of the R/V Natsushima, the operation team of the ROV Hyper-Dolphin and chief scientist Dr. Motohiro Shimanaga (Kumamoto University) for their valuable help in obtaining deep-sea hydrothermal vent samples during cruise NT14-06. We also thank Mr. Akihiro Tame for his excellent assistance in preparing samples for electron microscopy. We are grateful to Dr. Chong Chen for his help in improving this manuscript. This work was partially supported by JSPS KAKENHI Grant Number JP26820389.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Makita, H., Tanaka, E., Mitsunobu, S. et al. Mariprofundus micogutta sp. nov., a novel iron-oxidizing zetaproteobacterium isolated from a deep-sea hydrothermal field at the Bayonnaise knoll of the Izu-Ogasawara arc, and a description of Mariprofundales ord. nov. and Zetaproteobacteria classis nov.. Arch Microbiol 199, 335–346 (2017). https://doi.org/10.1007/s00203-016-1307-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-016-1307-4