Abstract
A halophilic archaeal strain, YJ-18-S1T, was isolated from Yangjiang marine solar saltern, Guangxi Province, China. Cells were pleomorphic, stained Gram-negative and formed red-pigmented colonies on agar plates. Strain YJ-18-S1T was able to grow at 20–55 °C (optimum 37 °C), at 0.9–4.8 M NaCl (optimum 2.6 M NaCl), at 0.005–1.0 M MgCl2 (optimum 0.3 MgCl2) and at pH 5.5–8.5 (optimum pH 7.0). The cells were lysed in distilled water, and the minimal NaCl concentration to prevent cell lysis was found to be 5 % (w/v). The major polar lipids of the strain were phosphatidic acid, phosphatidylglycerol, phosphatidylglycerol phosphate methyl ester, phosphatidylglycerol sulfate and sulfated mannosyl glucosyl diether. The 16S rRNA gene and rpoB′ gene of strain YJ-18-S1T were phylogenetically related to the corresponding genes of Halorubrum members (94.3–98.0 and 86.7–96.1 % similarities, respectively). The DNA G+C content of strain YJ-18-S1T was 66.2 mol%. The phenotypic, chemotaxonomic and phylogenetic properties suggested that strain YJ-18-S1T (=CGMCC 1.12554T = JCM 30030T) represents a new species of Halorubrum, for which the name Halorubrum rutilum sp. nov. is proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diverse halophilic archaea, members of the class Halobacteria, are always found to be present in marine solar salterns (Ghai et al. 2011; Narasingarao et al. 2012; Oren 2014). The past several years have witnessed the description of novel species and genera based on the haloarchaeal strains isolated from these kinds of hypersaline environments, such as Halobellus inordinatus (Qiu et al. 2013a), Natronomonas gomsonensis (Kim et al. 2013), Halobacterium rubrum (Han and Cui 2014), Haloarchaeobius litoreus (Zhang and Cui 2014b), Salinigranum rubrum (Cui and Zhang 2014) and Halovenus salina (Infante-Domínguez et al. 2015). During our survey on halophilic archaeal diversity of the Yangjiang marine solar saltern (Guangxi Province, China), we obtained a halophilic archaeal isolate, YJ-18-S1T, that was most closely related to the members of Halorubrum, as judged from 16S rRNA gene sequences.
The genus Halorubrum, belonging to the family Haloferacaceae (Gupta et al. 2015), was first established by McGenity and Grant (1995) and currently contains 30 named species (Fig. 1) (Yim et al. 2014; Zhang and Cui 2014a; Han and Cui 2015; Kondo et al. 2015). Among these Halorubrum species, five of them (H. alkaliphilum, H. gandharaense, H. luteum, H. tibetense and H. vacuolatum) are haloalkaliphilic species, while the remaining 25 members are neutrophilic. The polar lipids of the neutrophilic members of Halorubrum are phosphatidylglycerol, phosphatidylglycerol phosphate methyl ester, phosphatidylglycerol sulfate and sulfated mannosyl glucosyl diether. The alkaliphilic species lack phosphatidylglycerol sulfate and glycolipids (Oren et al. 2009). In this study, we characterized strain YJ-18-S1T as a new species of the genus Halorubrum.
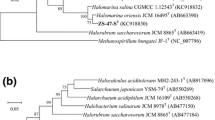
Maximum-likelihood phylogenetic tree reconstructions based on 16S rRNA gene (a) and rpoB′ gene (b) sequences, showing the relationships between strains YJ-18-S1T and related members within the family Halobacteriaceae. Bootstrap values (%) are based on 1000 replicates and are shown for branches with more 70 % bootstrap support. Bar represents expected substitutions per nucleotide position
Materials and methods
Isolation and cultivation of halophilic archaeal strain
Strain YJ-18-S1T was isolated from a sediment sample from Yangjiang marine solar saltern, at Yangjiang, Guangxi Province, China (21°31′48′N, 111°28′5′E; elevation, sea level), in 2012. The sediment sample was picked up from the surface of the bed of a crystallizer pond in which the brine had a temperature of 28 °C at the time of sampling, a pH of 7.2 and salinity of 28 °Bé. The neutral haloarchaeal medium (NHM) was used for the isolation procedure and contained the following ingredients (g/L): yeast extract (Oxoid) 0.05, fish peptone (Sinopharm Chemical Reagent Co., Ltd.) 0.25, sodium pyruvate 1.0, KCl 5.4, K2HPO4 0.3, CaCl2 0.29, NH4Cl 0.27, MgSO4·7H2O 26.8, MgCl2·6H2O 23.0, NaCl 184.0 (pH adjusted to 7.0–7.2 with 1 M NaOH solution). One gram of the sediment sample was suspended in 9 ml of NHM, and the turbid liquid was serially diluted in liquid NHM and spread onto NHM agar plates. The inoculated plates were incubated for 3 months at 37 °C. After this initial cultivation, colonies were successively re-streaked on NHM agar plates at least three times to obtain pure colonies.
Phenotypic determination
The type strains H. aidingense CGMCC 1.2670T, H. halophilum JCM 18963T, H. kocurii CGMCC 1.7018T, H. lacusprofundi CGMCC 1.3490T, H. lipolyticum CGMCC 1.5332T, H. rubrum CGMCC 1.12124T, H. saccharovorum CGMCC 1.2147T were selected as reference strains in phenotypic tests. These reference strains were routinely grown aerobically at 37 °C in NHM.
Determination of morphology and growth characteristics, nutrition, miscellaneous biochemical tests and sensitivity to antimicrobial agents were performed for all species in NHM, according to the proposed minimal standards for description of new taxa in the order Halobacteriales (Oren et al. 1997). The Gram staining was performed as described previously (Dussault 1955). Cell morphology and motility in exponentially growing liquid cultures were examined using a light microscope equipped with phase-contrast optics. The minimum salt concentration preventing cell lysis was determined by suspending cells in serial dilutions of sterile saline with NaCl concentrations ranging from 0 to 150 g/L, and the stability of the cells was detected by light microscopic examination. Growth and gas formation with nitrate as an electron acceptor were tested in 9-mL stoppered tubes (with Durham tubes) completely filled with liquid NHM and to which NaNO3 (5 g/L) had been added. The formation of gas from nitrate was detected by the presence of gas bubbles in the Durham tubes, and the formation of nitrite was monitored colorimetrically. Anaerobic growth in the presence of l-arginine or DMSO (5 g/L) was tested in completely filled 9-mL stoppered tubes. Starch hydrolysis was determined on NHM agar plates supplemented with 2 g/L soluble starch and detected by flooding the plates with Lugol’s iodine solution. Gelatin hydrolysis was performed by growing colonies on NHM agar plates amended with 5 g/L gelatin and detected by flooding the plates with Frazier’s reagent (McDade and Weaver 1959). Tests for catalase and oxidase activities were performed as described by Gonzalez et al. (1978). Esterase activity was detected as described by Gutiérrez and González (1972). Production of H2S was tested by growing the isolates and reference strains in a tube containing NHM liquid medium supplemented with 5 g/L sodium thiosulfate and detected using a filter-paper strip impregnated with lead acetate (Cui et al. 2007). To test for growth on single carbon sources, fish peptone and sodium pyruvate were omitted from the NHM and the compound to be tested was added at a concentration of 5 g/L. Antimicrobial susceptibilities were determined on NHM agar plates with antimicrobial compound disks.
Chemotaxonomic characterization
Strain YJ-18-S1T and H. saccharovorum CGMCC 1.2147T were routinely grown aerobically at 37 °C in NHM. Halophilic archaeal polar lipids were extracted using a chloroform–methanol system and analyzed using one- and two-dimensional TLC, as described previously (Cui et al. 2010). Two specific detection spray reagents, phosphate stain reagent for phospholipids and α-naphthol stain for glycolipids, were used. The general detection reagent, sulfuric acid–ethanol (1:2, by vol.), was also used to detect total polar lipids. The presence of phospholipids and glycolipids on the two-dimensional TLC was confirmed by comparison with one-dimensional TLC on which the polar lipid profile of reference strains was developed.
Phylogenetic and genotypic analysis
Halophilic archaeal genomic DNA was prepared as described previously (Cui et al. 2011), and the 16S rRNA genes were amplified with the forward primer 0018F (5′-ATTCCGGTTGATCCTGCC-3′) and reverse primer 1518R (5′-AGGAGGTGATCCAGCCGC-3′), then cloned and sequenced according to a previous protocol (Cui et al. 2009). The rpoB′ gene was amplified using the primer pair HrpoB2 1420F (5′-TGTGGGCTNGTGAAGAACTT-3′) and HrpoA 153R (5′-GGG TCCATCAGCCCCATGTC-3′) (Minegishi et al. 2010), and the PCR product was sequenced using the following primers: HrpoB2 1420F, HrpoA 153R and B1-628F (5′-CCNGCNGSVCAGAACTTC-3′). These sequences were aligned using the ClustalW program integrated in the MEGA 5 software (Tamura et al. 2011), and the phylogenetic trees were reconstructed using maximum-likelihood, maximum-parsimony and neighbor-joining algorithms in the MEGA 5 software. The DNA G+C content was determined from the midpoint value (T m) of the thermal denaturation method (Marmur and Doty 1962) at 260 nm with a Beckman Coulter DU800™ spectrophotometer equipped with a high-performance temperature controller.
Results and discussion
Cells of strain YJ-18-S1T were observed to be motile and pleomorphic rods when grown in NHM liquid medium (Supplementary Fig. S1). They stained Gram-negative, and the colonies were observed to be red-pigmented. Strain YJ-18-S1T was found to be able to grow at 20–55 °C (optimum 37 °C), at 0.9–4.8 M NaCl (optimum 2.6 M NaCl), at 0.005–1.0 M MgCl2 (optimum 0.3 MgCl2) and at pH 5.5–8.5 (optimum pH 7.0). The cells were lysed in distilled water, and the minimal NaCl concentration to prevent cell lysis was found to be 5 % (w/v). The strain was able to grow under anaerobic conditions using nitrate or l-arginine, but not with DMSO. It was found to be positive for H2S formation and indole formation. Strain YJ-18-S1T did not hydrolyze starch, gelatin, Tween 80 or casein. Strain YJ-18-S1T was sensitive to the following antimicrobial compounds (µg per disk, unless otherwise indicated): novobiocin (30), bacitracin (0.04 IU per disk), rifampin (5), nitrofurantoin (300), mycostatin (100) and trimethoprim (5). It was resistant to the following antimicrobial compounds: erythromycin (15), penicillin G (10 IU per disk), ampicillin (10), chloramphenicol (30), neomycin (30), norfloxacin (10), ciprofloxacin (5), streptomycin (10), kanamycin (30), tetracycline (30), vancomycin (30), gentamicin (10) and nalidixic acid (30). The main phenotypic characteristics differentiating strain YJ-18-S1T from the related members of the genus Halorubrum are anaerobic growth with nitrate, reduction of nitrate to nitrite, utilization of specific carbon sources, indole formation, hydrolysis of casein, gelatin, starch and Tween 80 (Table 1). More detailed results of phenotypic features of strain YJ-18-S1T are given in the species description.
The major polar lipids of strain YJ-18-S1T were identified as phosphatidylglycerol (PG), phosphatidylglycerol phosphate methyl ester (PGP-Me), phosphatidylglycerol sulfate (PGS) and a major glycolipid chromatographically identical to sulfated mannosyl glucosyl diether (S-DGD-3) (Supplementary Fig. S2). Since the polar lipid profile of strain YJ-18-S1T was identical to those of the neutrophilic members of the genus Halorubrum (Oren et al. 2009), the major polar lipid composition supports the classification of strain YJ-18-S1T in the genus Halorubrum.
Complete 16S rRNA gene sequence comparisons indicated that strain YJ-18-S1T has one kind of 16S rRNA gene sequence (1472 bp in length). The 16S rRNA gene of the strain was phylogenetically related to H. lipolyticum CGMCC 1.5332T (98.0 % similarity), H. saccharovorum CGMCC 1.2147T (97.8 % similarity), H. kocurii CGMCC 1.7018T (97.8 % similarity) and other members of genus Halorubrum, showing 94.3–97.7 % similarity. These 16S rRNA gene similarities are well lower than the recently recommended thresholds (98.65 %) to separate two prokaryotic species (Kim et al. 2014). Phylogenetic tree reconstructions using the maximum-likelihood (ML) algorithm revealed that strain YJ-18-S1T tightly clustered with the current 30 members of Halorubrum (Fig. 1a). This phylogenetic position was also confirmed in other trees generated using the maximum-parsimony (MP) and neighbor-joining (NJ) algorithms (Supplementary Fig. S3a and Fig. S4a).
The rpoB′ gene of strain YJ-18-S1T was closely similar to the corresponding gene of H. halophilum JCM 18963T (96.1 % similarity), H. saccharovorum CGMCC 1.2147T (96.0 % similarity) and other members of genus Halorubrum, showing 86.7–95.8 % similarity. In phylogenetic tree reconstructions using the rpoB′ (Fig. 1b), strain YJ-18-S1T tightly clustered with the members of Halorubrum. The phylogenetic position was also confirmed in trees generated using the maximum-parsimony (MP) and neighbor-joining (NJ) algorithms (Supplementary Fig. S3b and Fig. S4b).
The 16S rRNA gene and rpoB′ gene-based phylogenetic analysis results supported the placement of strain YJ-18-S1T in the genus Halorubrum.
The DNA G+C content of strain YJ-18-S1T was determined to be 66.2 mol%, within the range of the DNA G+C content of the members of the genus Halorubrum (60.2–71.2 mol%; Oren et al. 2009). The DNA G+C content of strain YJ-18-S1T was higher than those of H. lipolyticum CGMCC 1.5332T (65.9 mol%) (Cui et al. 2006), H. halophilum JCM 18963T (64.6 mol%) (Yim et al. 2014), H. rubrum CGMCC 1.12124T (64.9 mol%) (Qiu et al. 2013b), H. lacusprofundi CGMCC 1.3490T (65.3 mol%) (McGenity and Grant 1995), H. aidingense CGMCC 1.2670T (64.2 mol%) (Cui et al. 2006) but lower than those of H. saccharovorum CGMCC 1.2147T (71.2 mol%) (Tomlinson and Hochstein 1976; McGenity and Grant 1995) and H. kocurii CGMCC 1.7018T (69.4 mol%) (Gutiérrez et al. 2008).
Based on these phenotypic, chemotaxonomic and phylogenetic properties, a novel species of the genus Halorubrum is proposed to accommodate the strain, Halorubrum rutilum sp. nov. Characteristics that distinguish strain YJ-18-S1T from the related members of the genus Halorubrum are shown in Table 1.
Description of Halorubrum rutilum sp. nov.
Halorubrum rutilum (ru.ti’lum. L. neut. adj. rutilum red-colored, pertaining to the colony pigmentation of the type strain).
Cells are motile, pleomorphic rods under optimal growth conditions and Gram-negative. Colonies on agar plates containing 2.6 M NaCl are red, elevated and round. The type strain is chemoorganotrophic and aerobic. Growth occurs at 20–55 °C (optimum 37 °C), at 0.9–4.8 M NaCl (optimum 2.6 M NaCl), at 0.005–1.0 M MgCl2 (optimum 0.3 MgCl2) and at pH 5.5–8.5 (optimum pH 7.0). Cells were lysed in distilled water, and the minimal NaCl concentration to prevent cell lysis is 5 % w/v. Catalase and oxidase positive. Anaerobic growth occurs in the presence of nitrate and arginine but not with DMSO. Nitrate reduction to nitrite and gas formation from nitrate were observed. H2S formation and indole formation are positive. The type strain does not hydrolyze casein, starch, gelatin or Tween 80. The following substrates are utilized as single carbon and energy sources for growth: d-glucose, d-mannose, l-sorbose, sucrose, glycerol, d-sorbitol, acetate, pyruvate, dl-lactate, succinate, l-malate, fumarate and citrate. The following substrates are utilized as single carbon, nitrogen or energy sources for growth: l-arginine, l-aspartate and l-glutamate. No growth occurs on d-galactose, d-fructose, d-ribose, d-xylose, maltose, lactose, d-mannitol, glycine, l-alanine, l-lysine or l-ornithine. The major polar lipids are phosphatidic acid (PA), phosphatidylglycerol (PG), phosphatidylglycerol phosphate methyl ester (PGP-Me), phosphatidylglycerol sulfate (PGS) and sulfated mannosyl glucosyl diether (S-DGD-3). The DNA G+C content of the type strain was 66.2 mol% (T m).
The type strain is YJ-18-S1T (= CGMCC 1.12554T = JCM 30030T) and was isolated from Yangjiang marine solar saltern, Guangxi Province, China.
The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene and rpoB′ gene sequences of strain YJ-18-S1T are KC918819 and KT184687, respectively.
References
Cui H-L, Zhang W-J (2014) Salinigranum rubrum gen. nov., sp. nov., a new member of the family Halobacteriaceae isolated from a marine solar saltern. Int J Syst Evol Microbiol 64:1978–1983
Cui H-L, Tohty D, Zhou P-J, Liu S-J (2006) Halorubrum lipolyticum sp. nov. and Halorubrum aidingense sp. nov., isolated from two salt lakes in Xin-Jiang, China. Int J Syst Evol Microbiol 56:1631–1634
Cui H-L, Lin Z-Y, Dong Y, Zhou P-J, Liu S-J (2007) Halorubrum litoreum sp. nov., an extremely halophilic archaeon from a solar saltern. Int J Syst Evol Microbiol 57:2204–2206
Cui H-L, Zhou P-J, Oren A, Liu S-J (2009) Intraspecific polymorphism of 16S rRNA genes in two halophilic archaeal genera, Haloarcula and Halomicrobium. Extremophiles 13:31–37
Cui H-L, Gao X, Yang X, Xu X-W (2010) Halorussus rarus gen. nov., sp. nov., a new member of the family Halobacteriaceae isolated from a marine solar saltern. Extremophiles 14:493–499
Cui H-L, Yang X, Mou Y-Z (2011) Salinarchaeum laminariae gen. nov., sp. nov.: a new member of the family Halobacteriaceae isolated from salted brown alga Laminaria. Extremophiles 15:625–631
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70:484–485
Ghai R, Pašić L, Fernández AB, Martin-Cuadrado A-B, Megumi Mizuno C, McMahon KD, Papke RT, Stepanauskas R, Rodriguez-Brito B, Rohwer F, Sánchez-Porro C, Ventosa A, Rodríguez-Valera F (2011) New abundant microbial groups in aquatic hypersaline environments. Sci Rep 1:135
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov. an amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24:710–715
Gupta RS, Naushad S, Baker S (2015) Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbalesord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov. Int J Syst Evol Microbiol 65:1050–1069
Gutiérrez C, González C (1972) Method for simultaneous detection of proteinase and esterase activities in extremely halophilic bacteria. Appl Microbiol 24:516–517
Gutiérrez MC, Castillo AM, Pagaling E, Heaphy S, Kamekura M, Xue Y, Ma Y, Cowan DA, Jones BE, Grant WD, Ventosa A (2008) Halorubrum kocurii sp. nov., an archaeon isolated from a saline lake. Int J Syst Evol Microbiol 58:2031–2035
Han D, Cui H-L (2014) Halobacterium rubrum sp. nov., isolated from a marine solar saltern. Arch Microbiol 196:847–851
Han D, Cui H-L (2015) Halorubrum laminariae sp. nov., isolated from the brine of salted brown alga Laminaria. Antonie Van Leeuwenhoek 107:217–223
Infante-Domínguez C, Corral P, Sánchez-Porro C, Ventosa A (2015) Halovenus salina sp. nov., an extremely halophilic archaeon isolated from a saltern. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.000370
Kim T-Y, Kim S-J, Park S-J, Kim J-G, Cha I-T, Jung M-Y, Lee S-A, Roh SW, Yim KJ, Itoh T, Rhee S-K (2013) Natronomonas gomsonensis sp. nov., isolated from a solar saltern. Antonie Van Leeuwenhoek 104:627–635
Kim M, Oh H-S, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kondo Y, Minegishi H, Echigo A, Shimane Y, Kamekura M, Itoh T, Ohkuma M, Takahashi-Ando N, Fukushima Y, Yoshida Y, Usami R (2015) Halorubrum gandharaense sp. nov., a novel alkaliphilic haloarchaeon from commercial rock salt. Int J Syst Evol Microbiol 65:2345–2350
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J Mol Biol 5:109–118
McDade JJ, Weaver RH (1959) Rapid methods for the detection of gelatin hydrolysis. J Bacteriol 77:60–64
McGenity TJ, Grant WD (1995) Transfer of Halobacterium saccharovorum, Halobacterium sodomense, Halobacterium trapanicum NRC 34021 and Halobacterium lacusprofundi to the genus Halorubrum gen. nov., as Halorubrum saccharovorum comb. nov., Halorubrum sodomense comb. nov., Halorubrum trapanicum comb. nov., and Halorubrum lacusprofundi comb. nov. Syst Appl Microbiol 18:237–243
Minegishi H, Kamekura M, Itoh T, Echigo A, Usami R, Hashimoto T (2010) Further refinement of Halobacteriaceae phylogeny based on the full-length RNA polymerase subunit B’ (rpoB ’) gene. Int J Syst Evol Microbiol 60:2398–2408
Narasingarao P, Podell S, Ugalde JA, Brochier-Armanet C, Emerson JB, Brocks JJ, Heidelberg KB, Banfield JF, Allen EE (2012) De novo assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. ISME J 6:81–93
Oren A (2014) Taxonomy of halophilic Archaea: current status and future challenges. Extremophiles 18:825–834
Oren A, Ventosa A, Grant WD (1997) Proposed minimal standards for description of new taxa in the order Halobacteriales. Int J Syst Bacteriol 47:233–238
Oren A, Arahal DR, Ventosa A (2009) Emended descriptions of genera of the family Halobacteriaceae. Int J Syst Evol Microbiol 59:637–642
Qiu X-X, Mou Y-Z, Zhao M-L, Zhang W-J, Han D, Ren M, Cui H-L (2013a) Halobellus inordinatus sp. nov., from a marine solar saltern and an inland salt lake of China. Int J Syst Evol Microbiol 63:3975–3980
Qiu X-X, Zhao M-L, Han D, Zhang W-J, Cui H-L (2013b) Halorubrum rubrum sp. nov., an extremely halophilic archaeon from a Chinese salt lake. Antonie Van Leeuwenhoek 104:885–891
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tomlinson GA, Hochstein LI (1976) Halobacterium saccharovorum sp. nov., a carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 22:587–591
Yim KJ, Cha I-T, Lee H-W, Song HS, Kim K-N, Lee S-J, Nam Y-D, Hyun D-W, Bae J-W, Rhee S-K, Seo M-J, Choi J-S, Choi H-J, Roh SW, Kim D (2014) Halorubrum halophilum sp. nov., an extremely halophilic archaeon isolated from a salt-fermented seafood. Antonie Van Leeuwenhoek 105:603–612
Zhang W-J, Cui H-L (2014a) Halorubrum salinum sp. nov., isolated from a marine solar saltern. Arch Microbiol 196:395–400
Zhang W-J, Cui H-L (2014b) Haloarchaeobius litoreus sp. nov., isolated from a marine solar saltern. Antonie Van Leeuwenhoek 105:1085–1090
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 31370054), the 11th “Six Talents Peak” Project of Jiangsu Province (No. 2014-SWYY-021) and the Qinglan Project of Jiangsu Province and a project funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yin, S., Wang, Z., Xu, JQ. et al. Halorubrum rutilum sp. nov. isolated from a marine solar saltern. Arch Microbiol 197, 1159–1164 (2015). https://doi.org/10.1007/s00203-015-1159-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-015-1159-3