Abstract
Drass is the coldest inhabited place in India and the second coldest, inhabited place in the world, after Siberia. Using the 16SrDNA amplicon pyrosequencing, bacterial diversity patterns were cataloged across the Drass cold desert. In order to identify the ecotype abundance across cold desert environment, bacterial diversity patterns of Drass were further compared with the bacterial diversity of two other cold deserts, i.e., Antarctic and Arctic. Acidobacteria, Proteobacteria, Actinobacteria, Bacteroidetes, Cyanobacteria and Gemmatimonadetes were among the highly abundant taxonomic groups present across all the three cold deserts and were designated as the core phyla. However, Firmicutes, Nitrospirae, Armatimonadetes (former candidate division OP10), Planctomycetes, TM7, Chloroflexi, Deinococcus-Thermus, Tenericutes and candidate phyla WS3 were identified as rare phyla in Drass, Antarctic and Arctic samples. Differential abundance patterns were also computed across all the three samples, i.e., Acidobacteria (32.1 %) were dominant in Drass and Firmicutes (52.9 ± 17.6 %) and Proteobacteria (42 ± 1.3 %) were dominant in Antarctic and Arctic reference samples, respectively. Alpha diversity values Shannon’s (H) and Simpson’s (1-D) diversity indices were highest for Antarctic samples, whereas richness estimators (ACE and Chao1) were maximum for Drass soil suggesting greater species richness in bacterial communities in Drass than the Antarctic and Arctic samples.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Himalayan cold deserts are characterized by a fragile ecosystem and a complex climate due to dramatic seasonal shifts in physical and biochemical properties. Bordered by the Karakoram chain of mountains in the north and the Himalayas in the south (Rawat and Adhikari 2005), Ladakh represents a high-altitude cold desert. The Karakoram range blocks most of the monsoon bearing clouds, making the region arid (Namgail 2009). Based on the cloning-based cultivation-independent methods (Shivaji et al. 2011; Srinivas et al. 2011; Kistler et al. 2013), recent studies have reported the bacterial diversity of Himalayan regions. Though cloning-based cultivation-independent approaches overcome the limitations of cultivation-based methods, owing to the methodology-based bias, e.g., PCR and sequencing depth (Vaz-Moreira et al. 2011), these techniques are not efficient enough to catalog the rare microbial taxons. High-throughput sequencing of total environmental DNA or marker genes, e.g., 16S rRNA, circumvents cloning by taking advantage of a highly efficient in vitro DNA amplification method. This approach provides deeper insights into the taxonomical and functional dynamics of the in situ microbial diversity (Dowd et al. 2008; Rhoads et al. 2012; Kaur et al. 2015).
The primary goal of the present study was to explore the bacterial community dynamics of cold desert of Drass and its further comparison with other cold deserts, e.g., Antarctic and Arctic. Comparative analysis has been done to identify the differentially abundant bacterial taxa in cold desserts with different geographical conditions, i.e., ecotypes. Here, we provide a basic framework to analyze the bacterial diversity patterns enriched across cold desert environment. However, this framework would be benefitted via deeply sequenced longitudinal time series datasets.
Materials and methods
Sampling site
Drass is a small town in the Kargil district of Ladakh region (J&K, India) at 34.45°N, 75.77°E with an average elevation of 3280 meters (10,764 feet). Soil samples (n = 10) were collected from different locations in the Drass region. One centimeter of the surface soil was removed with a sterile spatula before collection of soil samples. Soil was collected with sterile spatula and transferred into sterile polythene bags. The temperature of the soil ranged from 10 to 15 °C at the time of sampling. The soil samples were transported to the laboratory under ice and then stored at −20 °C (Shivaji et al. 2011).
Physiochemical soil analysis
Soil samples were thawed, air-dried and passed through a 2-mm sieve and pooled as one composite sample and sent for analysis to QC research lab IIIM Jammu, India, for analysis of pH, water holding capacity and various metal ions concentration in the soil. All determinations were conducted using triplicate samples.
Metagenomic DNA extraction and amplification
Manual metagenomic DNA extraction protocols developed by Zhou et al. 1996; Wechter et al. 2003; Brady 2007; Amorim et al. 2008; Pang et al. 2008; Liles et al. 2009; Inceoglu et al. 2010 were applied for environmental DNA extraction. DNA extracted from multiple methods (n = 3) was pooled and diluted (1/100 dilutions) with final concentration of 20 ng/µl and used as template for PCR amplification. Small region (V1–V3) of the 16S rRNA gene was amplified from the total soil DNA by PCR using universal primers set 28F and 519 R (Nossa et al. 2010; Fan et al. 2012).
Pyrosequencing
Bacterial tag-encoded FLX amplicon pyrosequencing (bTEFAP) (Dowd et al. 2008) was performed at the Research and Testing Laboratory, Lubbock, TX, USA (www.researchandtesting.com).
Quality filtering and phylogenetic analysis
Raw sequence data were processed and analyzed using the Quantitative Insights into Microbial Ecology (QIIME version 1.5.0) pipeline, with default settings (Caporaso et al. 2010). Reads were removed after further analysis if at least one of the following criteria was met: (1) the presence of homopolymers with more than 8 bp and (2) length shorter than 200 bp. Quality-filtered reads were denoised to remove sequencing errors by flowgram clustering. The resulting (trimmed and clean) sequences were then filtered of any non-bacterial ribosome sequences set at default parameters (Gontcharova et al. 2010). The 454 sequence data comprise both identical reads and reads that start at the same position in the genome but have different lengths or vary slightly due to pyrosequencing errors. These erroneous reads can lead to an overestimation of several operational taxonomic units in a sample. Uchime tool was used to remove the chimeric sequences (Edgar et al. 2011). Quality trimmed sequences were clustered into operational taxonomic units (OTUs) using UClust (Edgar 2010) with a cutoff value of 99 % sequence identity. Candidate OTUs were assigned to phylogeny using RDP (Cole et al. 2009; Krober et al. 2009) scheme set at 80 % confidence value. Rarefaction curves were calculated by using RDP pyrosequencing pipeline (Cole et al. 2009).
Reference datasets
A total of four reference datasets were obtained from NCBI. Two reference datasets were selected randomly from the Antarctic study with accession nos. SRX206452 and SRX206985, and remaining two reference datasets were selected randomly from the Arctic study with accession no SRX017110. The source of Antarctic and Arctic samples was from Grove Mountains, East Antarctic and Foreland of MidreLoven glacier.
Statistical analysis
For each metagenome, a subset of 1000 candidate OTUs were randomly selected to avoid the biases caused by differential sampling size (Kirchman et al. 2010). Inter-sample alpha diversity comparisons (Kirchman et al. 2010) were performed using a Vegan package (Oksanen et al. 2013). Sequences were aligned for phylogenetic reconstruction using MAFFT algorithm that appeared in the version 1.7.2 of UGENE as the External Tool. Phylogenetic trees were built from the aligned sequence profiles using FASTTREE 2.1 (Price et al. 2010). Distance matrices were constructed from each phylogeny, and pairwise Mantel test (1000 permutations, two tailed: p value) was performed using PASSAGE-2 (Rosenberg and Anderson 2011). To assess the uncertainty in hierarchical cluster analysis over samples, bootstrap resampling (10,000 iterations) analysis implemented in Pvclust (Felsenstein 1985) was applied. Pvclust provides the bootstrap probability (BP) value from the ordinary bootstrap resampling (Felsenstein 1985) and the approximately unbiased (AU) probability value from multi-scale bootstrap resampling (Shimodaira and Hasegawa 2001; Suzuki and Shimodaira 2006). Cluster dendrogram was constructed with AU/BP values (%). A heat map showing the pairwise Pearson coefficient correlations among the samples was plotted using heat map function implemented in R.
Nucleotide sequence accession number
The 16S rRNA gene sequences derived from pyrosequencing of Drass soil have been deposited in the European Nucleotide Archive under accession number PRJEB5191.
Results
Soil of Drass cold desert is sandy, slightly alkaline (pH 7.4), nutrient poor with 0.72 % organic content and 12 ds/m electric conductivity (EC). Physicochemical properties of composite soil sample from Drass are given in Table 1.
Only three DNA extraction protocols, developed by Zhou et al. (1996); Wechter et al. (2003) and Pang et al. (2008), worked efficiently on the soil of Drass. DNA extracted using these three protocols were pooled and used as a template for PCR amplification of V1–V3 region of 16S rDNA. The amplicons were pyrosequenced, and 3000 sequence reads were generated. Downstream quality filtering resulted in 2819 high-quality sequences with average read length of ≥200 bp. Phylogenetic analysis revealed Acidobacteria, Proteobacteria and Actinobacteria as highly abundant in the soil of Drass representing 39 % of the total bacterial sequences. In addition, Chloroflexi, Bacteroidetes, Verrucomicrobia, Planctomycetes, Firmicutes, Nitrospira, Armatimonadetes (former candidate division (OP10), Gemmatimonadetes and Cyanobacteria were also identified at the relatively low abundance (<10 %), along with candidate phyla WS3 and TM7 and several unclassified bacteria (Fig. 1).
Comparative account of bacterial diversity across Drass, Antarctic and Arctic soil samples
The 16S rRNA gene sequences from the present study (Drass soil) and those retrieved from NCBI (Arctic and Antarctic soil) were clustered at 99 % sequence identity (Egge et al. 2013) (Table 2). A significant proportion (2.2, 2.8 and 1.9 %) of the total sequences from Drass and Antarctic (ANT1 and ANT2) samples was observed as chimeric (Uchime), and samples were filtered before clustering. No chimeric sequences were detected in Arctic samples. The rarefaction curves constructed at the 1 % genetic distance exhibited a steeper slope for the Drass soil than Antarctic and Arctic soil samples, hence demonstrating a greater genetic richness in the Drass soil (Fig. 2a). Drass sample, however, needs extensive sampling to represent the taxonomic diversity. Furthermore, Shannon’s (H) and Simpson’s diversity indices were highest for Antarctic samples, whereas ACE and Chao1 were maximum for Drass soil suggesting higher species richness across Drass soil sample than the Antarctic and Arctic soil samples (Table 3). Beta-diversity and hierarchical clustering analysis revealed that the total bacterial community structure in the Drass is more similar to Arctic than Antarctic soil samples (Fig. 2b–d).
a Rarefaction analysis performed on samples Drass (DR), Antarctic 1 (ANT1), Antarctic 1 (ANT2), Arctic 1 (ARC1) and Arctic 2 (ARC2). b Phylogenetic correlation of microbial communities of Drass, Antarctic and Arctic. A subset of 1000 randomly selected OTUs from each metagenome was used to construct an Euclidean distance matrix. Matrices were pairwise compared using Mantel test (1000 permutation, 0.05 as standard P value), and Pearson correlation values were calculated. Symbols used: Drass (DR), Antarctic 1 (ANT1), Antarctic 1 (ANT2), Arctic 1 (ARC1) and Arctic 2 (ARC2). c Heat map: Pearson correlation distances, ranging from 0 to 2, were normalized to a scale of 0–1, where 1 represents the furthest distance, or the least similar samples. Symbols used: Drass (DR), Antarctic 1 (ANT1), Antarctic 1 (ANT2), Arctic 1 (ARC1) and Arctic 2 (ARC2). d To assess the uncertainty in hierarchical cluster analysis over samples Drass (DR), Antarctic 1 (ANT1), Antarctic 1 (ANT2), Arctic 1 (ARC1) and Arctic 2 (ARC2). Bootstrap resampling (10,000 iterations) was applied via the R package Pvclust. The uncentered Pearson correlation is used as the distance metric with average linkage. The numbers above each edge show the probability of nodes below that edge occurring as a cluster in resampled trees, via ordinary bootstrap resampling (BP, green) or multi-scale bootstrap resampling (AU, red) (color figure online)
Abundant versus rare OTU analysis
Core/abundant phyla in Drass soil, Antarctic samples and Arctic samples
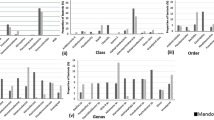
OTUs represented in all the five samples, i.e., Drass (DR), Antarctic (ANT1 and ANT2) and Arctic (ARC1 and ARC2), were collectively termed as core phyla (Serkebaeva et al. 2013) affiliated with the Acidobacteria, Proteobacteria, Actinobacteria, Verrucomicrobia, Bacteroidetes, Cyanobacteria and Gemmatimonadetes (Fig. 3a). The relative percentage abundance of phyla, class and orders mentioned in the below paragraph is given in Table 2.
Interestingly, Acidobacteria dominated the bacterial diversity of Drass sample with Gp6, Gp4 and GP7 being the abundant classes. Relative abundance patterns of Class Gp6 and Gp4 was lower in the Antarctic and Arctic samples with GP7 as entirely absent. Proteobacteria were abundant across all the three sites; however, they were comparatively more predominant in Arctic soil samples than Drass and Antarctic samples. Order Rhizobiales, Sphingomonadales, Caulobacterales of the Alphaproteobacteria, Burkholderiales of the Betaproteobacteria and Pseudomonadales and Xanthomonadales of the Gammaproteobacteria were represented among all the three sites. Enterobacteriales (Gammaproteobacteria) were represented among Antarctic samples only. Order Myxococcales (Deltaproteobacteria) were not represented in Antarctic and Arctic subsamples.
There were no significant differences in the abundance of Actinobacteria across Drass, Antarctic and Arctic soil samples. Unclassified order Actinomycetales was present across all the soil samples. However, order Solirubrobacterales and Acidimicrobiales were observed in Drass only and Arctic soil samples. Interestingly, Gaiellales were detected in Arctic samples only. At genus level, Aciditerrimonas, Conexibacter, Patulibacter, Rubrobacter, Propionibacterium, Marmoricola, Nocardioides, Aeromicrobium, Arthrobacter, Leifsonia, Amycolatopsis, Corynebacterium and Solirubrobacter were uniformly distributed in the Drass soil sample. Bacteroidetes were less abundant in Drass than Antarctic and Arctic samples. Order Sphingobacteriales was commonly represented across Drass, Antarctic and Arctic samples. However, Cytophagales were detected across Arctic samples only. Members of the Verrucomicrobia were more abundant in Drass than the Antarctic soil and Arctic soil samples. Order Opitutales was present in higher abundance across Drass sample, but not detected in the Antarctic and Arctic samples. Similarly, Order Verrucomicrobiales not detected in Drass was relatively abundant in Antarctic samples and detected in Arctic samples. Phylum Cyanobacteria were significantly less abundant in Drass than in Antarctic and Arctic samples. Class Chloroplast was abundant across Antarctic samples, and class Cyanobacteria was abundant across Arctic samples. Phylum Gemmatimonadetes represented by single genus, i.e., Gemmatimonas, was predominant across Drass sample and relatively very less in Antarctic and Arctic samples.
Rare OTUs across Drass, Arctic and Antarctic samples
The OTUs detected in two samples only were referred to as rare OTUs (Serkebaeva et al. 2013). Firmicutes, Nitrospirae, Armatimonadetes (Former candidate division OP10), Planctomycetes, TM7, Chloroflexi, Deinococcus-Thermus, Tenericutes and Candidate WS3 were distinguished as rare genera (Fig. 3b). Brevibacillus was the most abundant genera of Firmicutes present in Drass sample. Similarly, Leuconostoc, Tumebacillus and Staphylococcus were the most abundant genera (rare) present across Antarctic samples. The overall proportion of phyla Nitrospira, Planctomycetes, Chloroflexi and Armatimonadetes was significantly higher in Drass than in Antarctic and Arctic samples. Tenericutes were detected in Antarctic samples only and Candidate phylum WS3 detected in Drass only in the present study.
Discussion
Extreme conditions are the characteristics of cold desert alpine soils due to temperature fluctuations, and this holds true for Drass as well. To our knowledge, this is the first report on the bacterial diversity (richness and distribution) of Drass by high-throughput tag-encoded FLX amplicon pyrosequencing and its comparison with the bacterial diversity reported from other two cold deserts, Antarctic and Arctic based on retrieved sequence sets from the NCBI.
Low organic content in Drass is due to lack of vegetation, poor microbial activities, coarse sediments and low humus content (Kastovska et al. 2005; Rawat and Adhikari 2005; Sagwal 1997; Charan et al. 2013). Organic carbon content, soil texture and salinity have a strong influence on the microbial community diversity (Fierer et al. 2003, 2007). Multiple DNA extraction protocols were employed since no single method of metagenomic DNA isolation is efficient enough to represent all the bacterial diversity (Delmont et al. 2011). Sequence analysis revealed the predominance of Acidobacteria in Drass soil despite slightly alkaline pH although expect to be more in Antarctic and Arctic soil with acidic pH (6–6.7) (Li et al. 2003; Kastovska et al. 2005). This could be due to low carbon availability, soil organic carbon (OC), soil C/N ratio, as the abundance of Acidobacteria and the organic carbon content of the soil have been negatively co-related (Fierer et al. 2007; Pointing et al. 2009; Zeglin et al. 2011; Lee et al. 2012; Naether et al. 2012; Bottos et al. 2014; Niederberger et al. 2015). In addition, the Class Gp6 and Gp4 of Acidobacteria are reported to be present at high pH (Jones et al. 2009) and that could be the reason for their abundance in Drass in comparison with the Antarctic and Arctic samples.
Actinobacteria form the core phyla in cold deserts under investigation, and being mycelia formers, they are adapted to oligotrophic environments where the hyphae allow the bacteria to retrieve water and nutrients via pores in soil (Keki et al. 2013). Lesser representation of Bacteroidetes and Cyanobacteria in the soil of Drass supports the fact that Drass soil is relatively more oligotropic than the Antarctic and Arctic soils. Bacteroidetes are positively correlated with the organic carbon content of the soil (Fierer et al. 2007). These are typically decomposers and commonly associated with substrates rich in organic carbon contributing to the turnover of polymeric carbon (Fazi et al. 2005; Aislabie et al. 2009; Zakhia et al. 2008) in the soil. Cyanobacteria are the predominant biota in cold polar environments and play an important role in the carbon and nitrogen economy of tundra and polar desert soils (Bonilla et al. 2005; Adams et al. 2006; Vincent 2007; Varin et al. 2012). This photosynthetic phylum is either reported to be present in low frequency or absent in the Himalayan regions (Pradhan et al. 2010; Shivaji et al. 2011; Stres et al. 2014). Gemmatimonadetes and Verrucomicrobia found in the present study were reported from other cold habitats, though at a low frequency (Steven et al. 2007; Wagner et al. 2009; Pradhan et al. 2010; Wu et al. 2012). The core phyla obtained in the present study have also been reported from other cold habitats. Acidobacteria have been reported from the Tibetan plateau glacier, China, Mount Everest, Nepal, John Evans glacier, Canada Bench Glacier, Alaska, Schirmacher Oasis soil and Antarctic Siberian tundra soil samples (Zhou et al. 1997, Shivaji et al. 2004; Skidmore et al.2005; Cheng and Foght 2007; Liu et al. 2009; Campbell et al. 2010; Wu et al. 2012; Kim et al. 2014). Proteobacteria have been reported to be the dominant in the cold soils of the Himalayan mountains (Pradhan et al. 2010; Shivaji et al. 2011, Srinivas et al. 2011); Malan ice core drilled from the Tibetan Plateau (Xiang et al. 2004); surface sediment from the Arctic Ocean (Xuezheng et al. 2014); High-Arctic snow over sea ice (Moller et al. 2013) and Antarctic samples (Tytgat et al. 2014). The dominant Order Myxococcales of Deltaproteobacteria detected in Drass have been reported in Siberia, Arctic. Bacteroidetes has been reported as one of the abundant phyla from cold habitats (Wu et al. 2012; Moller et al. 2013). Actinobacteria reported to be the abundant phyla in cold habitats (Wu et al. 2012; Tytgat et al. 2014) and predominant phyla in Roopkund glacier, Himalayas (Pradhan et al. 2010).
The presence of Firmicutes in other cold habitats (Liu et al. 2009; Wagner et al. 2009) is well documented in the literature. Earlier studies on Himalayas (Pradhan et al. 2010; Shivaji et al. 2011) reported Firmicutes as the abundant phylum contrary to the present study. Negligible reads detected in Arctic subsamples is in accordance with Chu et al. (2010) who classified Firmicutes as the rarer phyla in Arctic soil. Armatimonadetes, Chloroflexi, Nitrospirae, candidate phylum TM7 and Planctomycetes are present in Drass soil at a low frequency as in other cold habitats (Shivaji et al. 2004; Cheng and Foght 2007; Wu et al. 2007; Li et al. 2008; Liu et al. 2009; Wagner et al. 2009; Pradhan et al. 2010; Shivaji et al. 2011; Stres et al. 2014). WS3 was detected in Drass soil only and completely absent in Antarctic and Arctic samples although few reports that justify the presence of Phylum WS3 in Antarctic and Arctic (Schutte et al. 2010; Tytgat et al. 2014) also. Tenericutes detected in Antarctic samples only in the present study have also been reported from few proglacial soils within the Himalayan regions (Srinivas et al. 2011; Stres et al. 2014).
Conclusion
On analyzing, and comparing the bacterial diversity of soil metagenome of Drass with Antarctic and Arctic metagenome, it was found that though most of the bacterial phyla present was common, but only the pattern of their dominance differed. Phylum Acidobacteria were dominant in the Drass soil, phylum Firmicutes were dominant in the Antarctic (not detected in Arctic soil), and Proteobacteria were dominant in the Arctic soil. Comparative sequence analysis also suggests that Drass soil is more arid than the Arctic and Antarctic soils and further resembles more to the Arctic than Antarctic soil or other Himalayan regions. Present study compliments the scientific reports that suggest that the bacterial community of any niche is specific, qualitatively and quantitatively designed by the combination of factors such as temperature, pH, salinity, water content, nutrients and carbon content.
References
Adams BJ, Bardgett RD, Ayres E, Wall DH, Aislabie J, Bamforth S, Bargagli R, Cary C, Cavacini P, Connell L, Convey P, Fell JW, Frati F, Hogg ID, Newsham KK (2006) Diversity and distribution Victoria Land biota. Soil Biol Biochem 38:3003–3018
Aislabie J, Jordan S, Ayton J, Klassen JL, Barker GM, Turner S (2009) Bacterial diversity associated with ornithogenic soil of the Ross Sea region, Antarctica. Can J Microbiol 55:21–36
Amorim JH, Macena TNS, Lacerda GV (2008) An improved extraction protocol for metagenomic DNA from a soil of the Brazilian Atlantic Rainforest. Genet Mol Res 7(4):1226–1232
Bonilla S, Villeneuve V, Vincent WF (2005) Benthic and planktonic algal communities in a high arctic lake: pigment structure and contrasting responses to nutrient enrichment. J Phycol 41:1120–1130
Bottos EM, Scarrow JW, Archer SD, McDonald IR, Cary SC (2014) Bacterial community structures of Antarctic soils. In: Cowan DA (ed) Antarctic terrestrial microbiology, vol 1. Springer, Berlin, pp 9–33
Brady SF (2007) Construction of soil environmental DNA cosmid libraries and screening for clones that produce biologically active small molecules. Nat Protoc 2(5):1297–1305
Campbell BJ, Polson SW, Hanson TE, Mack MC, Schuur EAG (2010) The effect of nutrient deposition on bacterial communities in Arctic tundra soil. Environ Microbiol 12(7):1842–1854
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Charan G, Bharti VK, Jadhav SE, Kumar S, Acharya S, Kumar P, Gogoi D, Srivastava RB (2013) Altitudinal variations in soil physico-chemical properties at cold desert high altitude. J Soil Sci Plant Nutr 13(2):267–277
Cheng SM, Foght JM (2007) Cultivation-independent and –dependent characterization of bacteria resident beneath John Evans Glacier. FEMS Microbiol Ecol 59:318–330
Chu H, Fierer N, Lauber CL, Caporaso JG, Knight R, Grogan P (2010) Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environ Microbiol 12:2998–3006
Cole JR, Wang Q, Cardenas E, Fish J, Chai B (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:141–145
Delmont TO, Robe P, Cecillon S, Clark IM, Constancias F, Simonet P, Hirsch PR, Vogel TM (2011) Accessing the soil metagenome for studies of microbial diversity. Appl Environ Microbiol 77(4):1315–1324
Dowd SE, Callaway TR, Wolcott RD, Sun Y, McKeehan T (2008) Evaluation of the bacterial diversity in the feces of cattle using 16S rDNA bacterial tag-encoded FLX amplicon pyrosequencing (bTEFAP). BMC Microbiol 8:125
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16):2194–2200
Egge E, Bittner L, Andersen T, Audic S, de Vargas C, Edvardsen B (2013) 454 pyrosequencing to describe microbial eukaryotic community composition, diversity and relative abundance: a test for marine haptophytes. PLoS ONE 8(9):e74371
Fan L, McElroy K, Thomas T (2012) Reconstruction of ribosomal RNA genes from metagenomic data. PLoS ONE 7(6):e39948
Fazi S, Amalfitano S, Pernthaler J, Puddu A (2005) Bacterial communities associated with benthic organic matter in headwater stream microhabitats. Environ Microbiol 7:1633–1640
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fierer N, Schimel J, Holden P (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35:167–176
Fierer N, Bradford M, Jackson R (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Gontcharova V, Youn E, Wolcott RD, Hollister EB, Gentry TJ (2010) Black box chimera check (B2C2): a windows-based software for batch depletion of chimeras from bacterial 16S rRNA gene datasets. Open Microbiol J 4:47–52
Inceoglu O, Hoogwout EF, Hill P, Elsas JDV (2010) Effect of DNA extraction method on the apparent microbial diversity of soil. Appl Environ Microbiol 76(10):3378–3382
Jones RT, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453
Kastovska K, Elster J, Stibal M, Santruckova H (2005) Microbial assemblages in soil microbial succession after glacier retreat in Svalbard (high arctic). Microbiol Ecol 50:396–407
Kaur G, Sharma R, Singh K, Sharma P (2015) Delineating bacterial community structure of polluted soil samples collected from cancer prone belt of Punjab, India. 3 Biotech. doi:10.1007/s13205-014-0270-5
Keki Z, Grebner K, Bohus V, Marialigeti K, Toth EM (2013) Application of special oligotrophic media for cultivation of bacterial communities originated from ultrapure water. Acta Microbiol Immunol Hung 60(3):345–357
Kim HM, Jung JY, Yergeau E, Hwang CY, Hinzman L, Nam S, Hong SG, Kim OS, Chun J, Lee YK (2014) Bacterial community structure and soil properties of a subarctic tundra soil in Council, Alaska. FEMS Microbiol Ecol 89:465–475
Kirchman DL, Cottrell MT, Lovejoy C (2010) The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ Microbiol 12:1132–1143
Kistler JO, Booth V, Bradshaw DJ, Wade WG (2013) Bacterial community development in experimental gingivitis. PLoS ONE 8(8):e71227
Krober M, Bekel T, Diaz NN, Goesmann A, Jaenicke S (2009) Phylogenetic characterization of a biogas plant microbial community integrating clone library 16S-rDNA sequences and metagenome sequence data obtained by 454-pyrosequencing. J Biotechnol 142:38–49
Lee CK, Barbier BA, Bottos EM, McDonald IR, Cary SC (2012) The inter-valley soil comparative survey: the ecology of Dry Valley edaphic microbial communities. ISME J 6:1046–1057
Li XL, Liu XH, Ju YT, Huang FX (2003) Properties of soils in Grove Mountains, East Antarctica. Sci China (D) 46(7):683–693
Li Y, Li F, Zhang X, Qin S, Zeng Z, Dang H, Qin Y (2008) Vertical distribution of bacterial and archaeal communities along discrete layers of a deep-sea cold sediment sample at the East Pacific Rise (13°N). Extremophiles 12:573–585
Liles MR, Williamson LL, Rodbumrer J, Torsvik V, Parsley LC, Goodman RM, Handelsman J (2009) Isolation and cloning of high-molecular-weight metagenomic DNA from soil microorganisms. Cold Spring Harb Protoc 8:pdb.prot5271R
Liu Y, Yao T, Jiao N, Kang S, Xu B, Zeng Y, Huang S, Liu X (2009) Bacterial diversity in the snow over Tibetan Plateau Glaciers. Extremophiles 13:411–423
Moller AK, Soborg DA, Abu Al-Soud W, Sorensen SJ, Kroer N (2013) Bacterial community structure in High-Arctic snow and freshwater as revealed by pyrosequencing of 16rRNA genes and cultivation. Polar Res 32:17390
Naether A, Foesel BU, Naegele V, Wust PK, Weinert J, Bonkowski M, Alt F, Oelmann Y, Polle A, Lohaus G, Gockel S, Hemp A, Kalko EKV, Linsenmair KE, Pfeiffer S, Renner S, Schoning I, Weisser WW, Wells K, Fischer M, Overmann J, Friedrich MW (2012) Environmental factors affect acidobacterial communities below the subgroup level in grassland and forest soils. Appl Environ Microbiol 78(20):7398–7406
Namgail T (2009) Mountain ungulates of the Trans-Himalayan region of Ladakh, India. Int J Wilder 15:35–40
Niederberger TD, Sohm JA, Gunderson TE, Parker AE, Tirindelli J, Capone DG, Carpenter EJ, Cary SC (2015) Microbial community composition of transiently wetted Antarctic Dry Valley soils. Front Microbiol 6(9). doi:10.3389/fmicb.2015.00009
Nossa CW, Oberdorf WE, Yang L, Aas JA, Paster BJ, Desantis TZ, Brodie EL, Malamud D, Poles MA, Pei Z (2010) Design of 16S rRNA gene primers for 454 pyrosequencing of the human foregut microbiome. World J Gastroenterol 16(33):4135–4144
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2013) Vegan: community ecology package. R package version 2.0-7
Pang M, Abdullah N, Lee C, Ng CC (2008) Isolation of high molecular weight DNA from forest topsoil for metagenomic analysis. Asia Pac J Mol Biol Biotechnol 16:35–41
Pointing SB, Chan Y, Lacap DC, Lau MCY, Jurgens JA, Farrell RL (2009) Highly specialized microbial diversity in hyper-arid polar desert. Proc Natl Acad Sci USA 106:19964–19969
Pradhan S, Srinivas TN, Pindi PK, Kishore KH, Begum Z, Singh PK, Singh AK, Pratibha MS, Yasala AK, Reddy GS, Shivaji S (2010) Bacterial biodiversity from Roopkund Glacier, Himalayan mountain ranges, India. Extremophiles 14:377–395
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5(3):e9490
Rawat GS, Adhikari BS (2005) Floristics and distribution of plant communities across moisture and topographic gradients in Tso Kar basin, Changthang plateau, eastern Ladakh. Arct Antarct Alp Res 37:539–544
Rhoads DD, Wolcott RD, Sun Y, Dowd SE (2012) Comparison of culture and molecular identification of bacteria in chronic wounds. Int J Mol Sci 13:2535–2550
Rosenberg MS, Anderson CD (2011) PASSaGE: pattern analysis, spatial statistics and geographic exegesis. version 2. Methods Ecol Evol 2:229–232
Sagwal SS (1997-12) Ladakh: ecology and environment. APH Publishing Corporation, India
Schutte UME, Abdo Z, Foster J, Ravel J, Bunge J, Solheim B, Forney LJ (2010) Bacterial diversity in a glacier foreland of the high Arctic. Mol Ecol 19:54–66
Serkebaeva YM, Kim Y, Liesack W, Dedysh SN (2013) Pyrosequencing-based assessment of the bacteria diversity in surface and subsurface peat layers of a Northern Wetland, with focus on poorly studied phyla and candidate divisions. PLoS ONE 8(5):e63994
Shimodaira H, Hasegawa M (2001) CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17:1246–1247
Shivaji S, Reddy GSN, Aduri RP, Kutty R, Ravenschlag K (2004) Bacterial diversity of a soil sample from Schirmacher Oasis, Antarctica. Cell Mol Biol 50:525–536
Shivaji S, Pratibha MS, Sailaja B, Hara Kishore K, Singh AK, Begum Z, Anarasi U, Prabagaran SR, Reddy GS, Srinivas TN (2011) Bacterial diversity of soil in the vicinity of Pindari glacier, Himalayan mountain ranges, India, using culturable bacteria and soil 16S rRNA gene clones. Extremophiles 15:1–22
Skidmore M, Anderson SP, Sharp M, Foght J, Lanoil BD (2005) Comparison of microbial community composition in two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl Environ Microbiol 71:6986–6997
Srinivas TNR, Singh SM, Pradhan S, Pratibha MS, Kishore KH, Singh AK, Begum Z, Prabagaran SR, Reddy GSN, Shivaji S (2011) Comparison of bacterial diversity in proglacial soil from Kafni Glacier, Himalayan Mountain ranges, India, with the bacterial diversity of other glaciers in the world. Extremophiles 15:673–690
Steven B, Briggs G, McKay CP, Pollard WH, Greer CW, Whyte LG (2007) Characterization of the microbial diversity in a permafrost sample from the Canadian High Arctic using culture-dependent and culture-independent methods. FEMS Microbiol Ecol 59(2):513–523
Stres B, Sul WJ, Murovec B, Tiedje JM (2014) Recently deglaciated high-altitude soils of the Himalaya: diverse environments, heterogenous bacterial communities and long-range dust inputs from the upper troposphere. PLoS ONE 8(9):e76440
Suzuki R, Shimodaira H (2006) Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22(12):1540–1542
Tytgat B, Verleyen E, Obbels D, Peeters K, De Wever A (2014) Bacterial diversity assessment in Antarctic terrestrial and aquatic microbial mats: a comparison between bidirectional pyrosequencing and cultivation. PLoS ONE 9(6):e97564
Varin T, Lovejoy C, Jungblut AD, Vincent WF, Corbeil J (2012) Metagenomic analysis of stress genes in microbial mat communities from Antarctica and the high Arctic. Appl Environ Microbiol 78(2):549–559
Vaz-Moreira I, Egas C, Nunes OC, Manaia CM (2011) Culture-dependent and culture-independent diversity surveys target different bacteria: a case study in a freshwater sample. Antonie Van Leeuwenhoek 100(2):245–257
Vincent WF (2007) Cold tolerance in cyanobacteria and life in the cryosphere. In: Seckbach J (ed) Algae and cyanobacteria in extreme environments. Springer, Heidelberg, pp 287–301
Wagner D, Kobabe S, Liebner S (2009) Bacterial community structure and carbon turnover in permafrost-affected soils of the Lena Delta, northeastern Siberia. Can J Microbiol 55:77–83
Wechter P, Williamson J, Robertson A, Kluepfel D (2003) A rapid, cost-effective procedure for the extraction of microbial DNA from soil. World J Microbiol Biotechnol 19:85–91
Wu X, Xi W, Ye W (2007) Bacterial community composition of a shallow hypertrophic freshwater lake in China, revealed by 16S rRNA gene sequences. FEMS Microbiol Ecol 61:85–96
Wu X, Zhang W, Liu G, Yang X, Hu P, Chen T, Zhang G, Li Z (2012) Bacterial diversity in the foreland of the Tianshan No. 1 glacier, China. Environ Res Lett 7:014038
Xiang SR, Yao TD, An LZ, Xu BQ, Li Z, Wu GJ, Wang YQ, Ma S, Chen XR (2004) Bacterial diversity in Malan ice core from the Tibetan Plateau. Folia Microbiol 49:269–275
Xuezheng L, Zhen W, Shuai C, Song W, Dan YI (2014) Bacterial diversity in Arctic marine sediment determined by culture-dependent and -independent approaches. Adv Polar Sci 25(1):46–53
Zakhia F, Jungblut AD, Taton A, Vincent WF, Wilmotte A (2008) Cyanobacteria in cold environments. In: Margesin R, Schinner F, Marx JC, Gerday C (eds) Psychrophiles: from biodiversity to biotechnology. Springer, Heidelberg, pp 121–135
Zeglin LH, Dahm CN, Barrett JE, Gooseff MN, Fitpatrick SK, Takacs-Vesbach CD (2011) Bacterial community structure along moisture gradients in the parafluvial sediments of two ephemeral desert streams. Microbiol Ecol 61:543–556
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Zhou J, Davey ME, Figueras JB, Rivkina E, Gilichinsky D, Tiedje JM (1997) Phylogenetic diversity of a bacterial community determined from Siberian tundra soil DNA. Microbiology 143:3913–3919
Acknowledgments
Authors are thankful to DBT Government of India (BT/PR11727/BCE/08/720/2008) for funding of this project. PG is thankful to CSIR (9/100/0177) 2K13-EMR-I, Government of India for CSIR-SRF. Ms. Sneha Ganjoo, project fellow in the school, is acknowledged for her suggestions in the preparation of Manuscript. Resources used in the Bioinformatic Centre funded by DBT Government of India are also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Rights and permissions
About this article
Cite this article
Gupta, P., Sangwan, N., Lal, R. et al. Bacterial diversity of Drass, cold desert in Western Himalaya, and its comparison with Antarctic and Arctic. Arch Microbiol 197, 851–860 (2015). https://doi.org/10.1007/s00203-015-1121-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-015-1121-4