Abstract
Key message
A major QTL (QLr.cau-2BL) for APR to leaf rust was detected on 2BL; an SSR marker was developed to closely link with QLr.cau-2BL and validated for effectiveness of MAS.
Abstract
The wheat landrace Hongmazha (HMZ) possesses adult plant resistance (APR) to leaf rust. To detect and validate quantitative trait locus (QTL) for the APR, four wheat populations were assessed for leaf rust severity in a total of eight field and greenhouse experiments. The mapping population Aquileja × HMZ (120 recombinant inbred lines, RILs) was genotyped using 90 K SNP markers. A major QTL (QLr.cau-2BL) was detected between the markers IWB3854 and IWB21922 on chromosome 2BL. IWB3854 and IWB21922 were positioned at approximately 531.14 Mb and 616.48 Mb, respectively, on 2BL of IWGSC RefSeq v1.0 physical map. Based on the sequences between 531.14 and 616.48 Mb on 2BL of IWGSC RefSeq v1.0, 415 simple sequence repeat (SSR) markers were developed. These markers and 28 previously published SSR makers were screened; the resulted polymorphic markers were used to genotype the relatively larger population RL6058 × HMZ (371 RILs). QLr.cau-2BL was mapped within a 1.5 cM interval on 2BL map of RL6058 × HMZ, and a marker (Ta2BL_ssr7) was identified to closely link with QLr.cau-2BL. Effectiveness of selection for QLr.cau-2BL based on Ta2BL_ssr7 was validated using two populations (RL6058 × HMZ F2:3 and Jimai22 × HMZ BC4F2:3). In addition, polymorphism at Ta2BL_ssr7 was detected among a panel of 282 commercial wheat cultivars. We believe, therefore, that Ta2BL_ssr7 should be useful for introducing QLr.cau-2BL into commercial wheat cultivars and for accumulating QLr.cau-2BL with other APR QTL.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaf rust (caused by Puccinia triticina Erikss.) is an important disease of wheat (Triticum aestivum L) globally; yield losses due to leaf rust can exceed 50% under favorable conditions if fungicides are not applied (Huerta-Espino et al. 2011). In China, the world’s largest wheat producer, leaf rust is widespread, occurring on around 15 million ha annually (Liu and Chen 2012). Cultivation of wheats with resistance to leaf rust can reduce the need for fungicides and thus is an economical and environmentally friendly strategy to control the disease.
Currently, 79 designated leaf rust resistance (Lr) genes have been described (McIntosh et al. 2017; Qureshi et al. 2018). Most of these genes confer all-stage resistance that is effective throughout the life of the plant. All-stage resistance is often race-specific and vulnerable to adaptation by virulent variants in P. triticina population, becoming ineffective within limited years in commercial wheat fields (McDonald and Linde 2002). Compared to commercial cultivars, some landraces and wild relatives are richer sources of all-stage resistance (Qureshi et al. 2018). Another category of resistance, adult plant resistance (APR), is insignificant at the seedling growth stage, but is expressed at the post-seedling stage (Caldwell et al. 1957). Certain Lr genes for APR (e.g., Lr12 and Lr22b) are race-specific, showing contrasting resistant/susceptible infection types (ITs) to different P. triticina races at the adult plant stage (Huerta-Espino et al. 2011; Park and McIntosh 1994). Quantitative trait loci (QTL) associated with APR confer primarily low disease characterized by one or more of the resistance components such as longer latent period, fewer uredinia, and smaller uredinium size (Lagudah 2011; Huerta-Espino et al. 2011). Some APR including Lr34 and Lr46 (Krattinger et al. 2009; Singh et al. 1998) have proven to condition durable resistance (Johnson 1981). The cloned Lr34 and Lr67 illustrate novel resistance mechanisms (Krattinger et al. 2009; Moore et al. 2015), which are distinct from the mechanism underlying race-specific resistance genes (Dangl et al. 2013). It is generally considered that durable resistance can be achieved by accumulating APR QTL, and such an accumulation can be facilitated by marker-assisted selection (MAS) (Lagudah 2011; Singh et al. 2000).
According to a recent review by Silva et al. (2018), 249 QTL for APR to leaf rust were previously reported. Such numerous QTL should provide a convenience for a breeding program to choose appropriate APR resources, although some of them may be redundant. These QTL differ in effect size; percentage of phenotypic variance explained (PVE) by a single QTL ranges from 1 to 73%. Some QTL could be significantly effective in only one of the test environments; some, such as Lr34, are quite stable with consistent effectiveness across multiple environments and across different genetic backgrounds. From the viewpoint of practical breeding, it is easier to use major QTL (e.g., PVE > 15%) with high stability than minor QTL. It is essential to find new APR QTL for enhancing the genetic diversity of leaf rust resistance deployed over the vast and agro-ecologically diverse wheat-planting regions in the world. The recently published IWGSC RefSeq v1.0 sequences (IWGSC 2018) combined with the developed high-throughput genotyping platforms such as 90 K SNP array (Wang et al. 2014) constitute a powerful tool for detecting new QTL and for developing markers closely linked with QTL.
The wheat landrace HMZ was ever commercially cultivated on substantial scale in Northwest China before 1965. Although showing susceptible IT at both the seedling and adult plant growth stages, HMZ has consistently displayed lower leaf rust severities (< 35%) at the adult plant stage since 2002 in our field screening nurseries located in different provinces. The objectives of this study were to detect QTL associated with the APR in HMZ, to develop SSR markers closely linked with the detected major QTL, and to validate the developed marker.
Materials and methods
Wheat materials
Four wheat populations were used to detect and/or validate QTL, namely, Aquileja × Hongmazha (HMZ) with 120 RILs, RL6058 × HMZ with 371 RILs, 40 F2:3 families from Aquileja × HMZ, and 40 BC4F2:3 families from Jimai22 × HMZ with Jimai22 as recurrent parent. HMZ is a Chinese winter wheat landrace with the accession number K-61293 at http://wheatpedigree.net/. The winter wheat cultivar Aquileja (PI 393993; Tevere/Giuliari//Gallini) originates from Italy. The spring wheats Thatcher (CItr 10003; Marquis/Iumillo//Marquis/Kanred; Clark 1935) and RL6058 (Thatcher*6/PI 58548) came from North America. The Chinese winter wheat Jimai22 (935024 /935106) is a commercial cultivar. Aquileja × HMZ and RL6058 × HMZ were developed from F2 to F6 by single seed descent and the subsequent generations (F8, F9 and F10) were used in this study. HMZ possesses APR to leaf rust, RL6058 carries Lr34, and the other three wheats (Aquileja, Jimai22 and Thatcher) are susceptible to the P. triticina race THTT used in this study. Thatcher was used as a susceptible control throughout the experiments. A set of 282 commercial wheat cultivars (Online Resource 1 Table S1), collected from some important wheat production provinces of China, were used to test polymorphism of an SSR marker developed in this study as detailed below.
Assessment on disease severities of RIL populations
Leaf rust severities of wheat RIL populations were evaluated in seven field experiments involving three locations (Gansu province, Wushan county, 34°42′15′′N, 104°40′08″E, elevation 1650 m; Henan province, Suiping county, 33°6′30′′N, 113°54′48″E, 150 m; and Shandong province, Tai’an district, 36°18′09′′N, 117°13′05″E, 90 m) and three autumn-sown wheat crop seasons (2016–2017, 2017–2018 and 2018–2019; abbreviated as 2017, 2018 and 2019, respectively). The experiments were conducted for Aquileja × HMZ in Gansu and Shandong in 2017 and 2018, and for RL6058 × HMZ in Gansu, Henan and Shandong in 2019. Each of the seven location × year combinations was considered as an individual “environment.” Sowing was done in mid-September in Gansu and in early October in Henan and Shandong. In each environment, experiment was in a randomized complete block design with three replicates. An individual plot consisted of a single 1-m-long row with 25 cm between adjacent rows. Each plot was sown with approximately 40 seeds of a RIL. A set of parents and susceptible control wheat Thatcher were included after every 60 RIL rows. Two rows of Thatcher perpendicular and adjacent to the plot rows were sown along each field block to serve as spreader plants of inoculums. The test plants were covered with plastic films during winter months for facilitating the plants to overwinter.
Although leaf rust epidemic can be induced by naturally occurring P. triticina urediniospores in all of the seven test environments, artificial inoculation was applied to create higher and more homogeneous disease pressure. The P. triticina race THTT was used as inoculum, which has been prevalent in the P. triticina population in China (Liu and Chen 2012). THTT has the avirulence/virulence spectrum Lr9, Lr19, Lr24, Lr28, Lr38, Lr39, Lr42, Lr45/Lr1, Lr2a, Lr2b, Lr2c, Lr3, Lr3bg, Lr3ka, Lr10, Lr11, Lr14a, Lr14b, Lr15, Lr16, Lr17a, Lr18, Lr20, Lr21, Lr23, Lr25, Lr26, Lr29, Lr30, Lr32, Lr33, Lr36, Lr50, LrB (Li et al. 2010). At the stem elongation stage of the test plants, inoculation was done on clear afternoons when there was a high likelihood of overnight dew. The spreader plants were sprayed with fresh THTT urediniospores suspended in water containing 0.04% Tween 20. The inoculated plants were then covered with plastic films to ensure long-lasting dew, and 18 h later the films were removed. About 2 weeks after inoculation when some urediniospores were obviously released from the spreader plants, the test plants were frequently misted after sunset to facilitate re-infection before disease recording was completed.
Leaf rust severity in the form of percentage of infected leaf area was scored using the modified Cobb scale (Peterson et al. 1948) on 5 to 12 flag leaves each plot entry. Recording was done three times when the leaf rust severities on Thatcher flag leaves reached 10 − 30%, 50 − 60%, and 80 − 90%, respectively. The area under the disease progress curve (AUDPC) was calculated based the three time recordings for each plot entry with the formula AUDPC = Σ[(xi + x(i+1))/2] × (t(i+1) − ti), where xi and x(i+1) are scores for a plot entry on date ti and date t(i+1), respectively, and t(i+1) − ti = the number of days between date t(i+1) and date ti. QTL mapping was done on AUDPC means calculated for each RIL by averaging over the three replicates within each experiment.
Identification of P. triticina races
To obtain some information on number and virulence of P. triticina races in the experimental plots, race identification was done on samples from the experiment in Shandong in 2018 following the procedure in Long and Kolmer (1989). Briefly, Thatcher leaves bearing uredinia were collected from the experimental plots. Single uredinia were isolated, and then the spores were increased. The resulted urediniospores were used to inoculate the wheat host differentials that consisted of 16 near-isogenic lines of Thatcher with respective single resistance genes Lr1, Lr2a, Lr2c, Lr3, Lr9, Lr16, Lr24, Lr26, Lr3ka, Lr11, Lr17, Lr30, LrB, Lr10, Lr14a, and Lr18. Ten to 12 days after incubation, IT was recorded as either high (IT 3–4) or low (IT 0 to 2 +). Races were designated following the three letter code system of Long and Kolmer (1989), with an additional fourth letter that described the high or low IT to Thatcher wheat lines with genes LrB, Lr10, Lr14a, and Lr18 (Long et al. 2002).
Assessment on disease severities of F2:3 populations
To examine effectiveness of selection based on an SSR marker developed in this study, the selected 40 Aquileja × HMZ F2:3 families (12 plants each family; 480 plants in total) and 40 Jimai22 × HMZ BC4F2:3 families (18 plants each family; 720 plants in total) were assessed for leaf rust severity in a greenhouse. Sowing was done in early October 2019 in 1.2-m-long rows with 25 cm spacing between adjacent rows. Each row had six seeds 20 cm apart. The families within each cross were arranged in a randomized complete block design with two (for Aquileja × HMZ) or three (for Jimai22 × HMZ) blocks. Two sets of parents and Thatcher were included in every block. From late November 2019 to mid-February 2020, heaters of the greenhouse were turned off. This resulted in the low temperatures ranging from − 2 °C (the lowest, night) to 9 °C (the highest, daytime) for vernalizing the seedlings. Afterwards, the greenhouse was running at 18–25 °C daily under natural daylight supplemented with high-pressure sodium lamps (8000–10,000 lx at top leaves) in a 16 h photoperiod. In early March, the plants were inoculated with THTT as mentioned above. The plants were frequently misted after sunset to facilitate re-infection. Leaf rust severity was scored on flag leaves when Thatcher flag leaves showed a severity around 90%. For each F2:3 or BC4F2:3 family, mean disease severity was calculated by averaging over the 12 or 18 plants within the family.
Statistical analysis of disease data
All analyses were performed in SAS/STAT version 9.3 (SAS Institute Inc., Cary, NC, USA). Analysis of variance (ANOVA) was conducted based on AUDPC values across the four experiments for Aquileja × HMZ and across the three experiments for RL6058 × HMZ. A linear model was fitted using PROC GLM: Pijk = μ + Gi + Ej + Ej(Rk) + Gi × Ej + eijk, where Pijk is the phenotypic value, μ the population mean, Gi the effect of the ith genotype (RIL), Ej the effect of the jth environment, Ej(Rk) the effect of the kth replicate within the jth environment, Gi × Ej the ijth effect of the genotype-by-environment interaction, and eijk designated the residual. Broad-sense heritability (H2) on a RIL mean basis was estimated applying the variance component analysis of PROC VARCOMP with the restricted maximum likelihood method setting all effects as random; the calculation was based on the equation H2 = σ2G/[σ2G + σ2G×E/m + σ2error/(m × r)], where σ2G denotes the genotypic (i.e., RIL) variance, σ2G×E the genotype-by-environment interaction variance, σ2error the error variance, m the number of environments, and r the number of replicate within each environment (Holland et al. 2003). The correlation coefficients for AUDPC among different experiments were estimated applying PROC CORR (Pearson).
Genotyping, map construction and QTL analysis
The mapping population Aquileja × HMZ (120 RILs) was genotyped using the 90 K Illumina® iSelect wheat SNP array (Wang et al. 2014). One seed from F9 generation of each RIL was used to grow plant from which DNA was extracted using cetyl trimethyl ammonium bromide (CTAB) method (Saghai-Maroof et al. 1984). The 90 K SNP assays were performed by CapitalBio Technology (Beijing, China; http://www.capitalbiotech.com). SNP allele clustering and genotype calling were performed following the procedure of Wang et al. (2014). Data points of heterozygosity were treated as missing values. Genotypic data were filtered to retain only markers with missing values < 5% and allele frequency between 0.35 and 0.65. RILs with missing values > 5% were filtered out. After removing redundant markers identified applying the BIN function in QTL IciMapping V4.0 software (Li et al. 2007), the retained high-quality markers (each marker representing a unique locus) were grouped and ordered using JoinMap 4.0 (Stam 1993). Genetic distances (in cM) were estimated based on the Kosambi mapping function. A linkage group was split if there was a distance ≥ 20 cM between two adjacent markers. Linkage groups were then orientated and assigned to chromosomes through alignments of the sequences flanking SNP with chromosomal survey sequence map (IWGSC 2014).
The second mapping population RL6058 × HMZ (371 RILs) was genotyped using 12 SSR markers that were developed in the present study based on the 2BL sequences of IWGSC RefSeq v1.0 (IWGSC 2018) or selected from the previously documented resources (http://wheat.pw.usda.gov/GG2/index.shtml). PCR reaction system consisted of 1.0 μl 10 × PCR buffer (Mg2+plus), 0.1 μl 5 × Taq DNA polymerase, 1.0 μl forward primer (1 μM), 1.0 μl reverse primer (1 μM), 2.0 μl DNA template (20–60 ng/μl), 0.25 μl dNTPs (10 mM), and 4.65 μl ddH2O. While the normal PCR cycling (Röder et al. 1998) was performed for most of the markers, for certain markers, a touchdown program was used, which started with 94 °C for 5 min; then 10 cycles of 94 °C denaturing for 30 s, 60 °C annealing for 30 s (touchdown with − 0.5 °C each cycle), and 72 °C extending for 30 s; followed by 35 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s; and ended with 72 °C for 10 min. Amplicons were separated in 6% denaturing polyacrylamide gels and visualized with the silver staining method (Bassam et al. 1991). Linkage map was constructed using MAPMAKER/EXP 3.0 (Lander et al. 1987).
QTL detection was performed using the composite interval mapping (CIM) method in Windows QTL Cartographer 2.5 (Wang et al. 2010). The threshold LOD score was calculated by running the permutation program set with 3000 replicates at a type I error rate of α = 0.05. Permutation analysis was conducted for each of the seven experiments, resulting in threshold LOD values varying from 2.8 to 3.2; for simplicity, the highest threshold LOD value (3.2) was chosen as a uniform threshold for all experiments. Effect size of QTL was measured with determination coefficient R2 (i.e., PVE).
Results
Leaf rust severity of wheat RILs
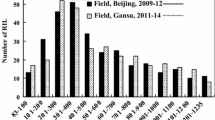
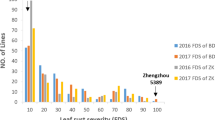
In each of the seven field experiments, leaf rust severities were approximately 90% on the susceptible control wheat Thatcher and less than 35% on the resistant landrace HMZ as exemplified by the typical symptoms shown in Fig. 1, indicating that disease pressure was high adequately for revealing resistance and susceptibility. To improve power of QTL detection, disease severities were measured in AUDPC. AUDPC values of RILs varied between 93.8 and 1133.8 in the population Aquileja × HMZ and from 23.3 to 1148.7 in RL6058 × HMZ. Mean AUDPC over RILs varied from 361 to 585 dependent on populations and environments (location × year combinations) (Table 1; Online Resource 1 Tables S2 and S3). Correlation coefficients for AUDPC among the experiments were all significant with P < 0.0001, ranging from 0.77 to 0.93 in Aquileja × HMZ and from 0.87 to 0.96 in RL6058 × HMZ. In both populations, variances due to RIL and environment were significant at α = 0.0001 and predominated over the variances due to RIL × environment, though the interaction variances were also significant (Table 2). Broad-sense heritability was high (0.92 in Aquileja × HMZ and 0.94 in RL6058 × HMZ). RILs continuously distributed over AUDPC scales (Fig. 2). These results indicate that the APR in HMZ was segregating in a quantitative nature in both RIL populations.
P. triticina races in the experimental plots
Eleven races were identified from 40 uredinial isolates collected from the field experimental plots of Shandong in 2018 (Table 3). Of the 40 samples, 18 were differentiated as the artificially inoculated race THTT, and 22 were grouped into 10 naturally occurring races. In light of the limited number of samples analyzed (22), the number of races (10) should not be understood as a comprehensive survey of all naturally occurring races in the plots; the actual race number might be more than 10. It can be inferred that multiple naturally occurring races might also exist in the other six experiments of this study.
QTL mapping using Aquileja × HMZ population
After filtering out the genotypic data that could not meet the quality control criteria as described above and removing redundant markers, 1517 high-quality unique marker loci were mapped on 27 linkage groups covering all 21 chromosomes. Six chromosomes (1D, 2D, 3A, 6B, 6D and 7D) were split into two groups. The total length of the genetic map was 3656.8 cM with an average distance of 2.4 cM (min = 0.1 and max = 19.8 cM) between two adjacent markers (Online Resource 1 Table S4).
The 120 Aquileja × HMZ RILs were scanned genome-wide with the 1517 markers to detect chromosome regions associated with the AUDPC values (Table 1; Online Resource 1 Table S2) for each of the four field experiments. A major QTL for APR to leaf rust was found on chromosome arm 2BL (designated as QLr.cau-2BL; Fig. 3a). For reading ease, only selected markers at about 10 cM distances are displayed in Fig. 3a, while more detailed information including all mapped unique markers with their positions can be found in Online Resource 1 Table S4. LOD peaks of QLr.cau-2BL were located between the markers IWB3854 and IWB21922. LOD values, ranging from 18.5 to 23.6 (Table 4), were significantly larger than the LOD threshold (3.2) in each of the four experiments (Fig. 3a). The parent HMZ contributed to the resistance (i.e., lower AUDPC) at QLr.cau-2BL, which explained from 47 to 57% of AUDPC variance.
Genetic maps of chromosome 2B (a) and 5DL (b) of Aquileja × Hongmazha, illustrating the logarithm of the odds (LOD) curves of QLr.cau-2BL (a) and QLr.cau-5DL (b). Marker names and intervals (in Kosambi cM) between adjacent markers are shown along the chromosome orientated with the telomere of 2BS and the centromere of 5DL to the left. Arrows suggest the positions of the centromeres inferred by aligning marker sequences to the chromosome survey sequence (IWGSC 2014). Horizontal lines indicate the threshold LOD of 3.2. Each of the small triangles along the x-axis represents a marker used for QTL mapping. For reading ease, only selected marker names at about 10 cM distances are displayed for 2B, while more detailed information including all mapped unique marker names with their positions can be found in Online Resource 1 Table S4
Two QTL (QLr.cau-5DL1 and QLr.cau-5DL2) were detected on 5DL (Fig. 3b). Resistance at both QTL was contributed by HMZ. Their effects on reducing AUDPC were significant at the LOD threshold (3.2) in only one or two of the four experiments (Fig. 3b) with small effect sizes (PVE = 2 to 7%; Table 4). These minor QTL need to be further tested. Only the major QTL QLr.cau-2BL is addressed in the remainder of the present paper.
Mapping of QLr.cau-2BL using RL6058 × HMZ population
The Aquileja × HMZ population was small (120 RILs). To improve mapping resolution and to examine QLr.cau-2BL in different genetic backgrounds, the relatively large population RL6058 × HMZ (371 RILs) was used for further mapping QLr.cau-2BL. Sequences of the flanking markers IWB3854 and IWB21922 on Aquileja × HMZ map (Fig. 3a) were aligned to the 2BL sequences of IWGSC RefSeq v1.0 (IWGSC 2018) at approximately 531.14 Mb and 616.48 Mb, respectively (Table 4). On the basis of the 2BL sequences in this interval in IWGSC RefSeq v1.0, 415 SSR markers were developed, which were named with the prefix “Ta2BL_ssr” (SSR marker of common wheat 2BL) followed by a consecutive number. Information about the 415 markers and 28 other SSR markers of 2BL selected from public domain (http://wheat.pw.usda.gov/GG2/index.shtml) was presented in Online Resource 1 Table S5. After screening, 12 polymorphic markers (Table 5) were used to genotype the 371 RL6058 × HMZ RILs and a map of QLr.cau-2BL region was constructed (Fig. 4). QTL analysis based on the AUDPC data (Table 1; Online Resource 1 Table S3) recorded in the three field experiments mapped QLr.cau-2BL within a 1.5 cM interval delimited by wmc499 and Ta2BL_ssr16 (Fig. 4). Ta2BL_ssr7 was 1.0 cM distal to wmc499 and 0.5 cM proximal to Ta2BL_ssr16. LOD values ranged from 31.6 to 35.6 (Table 4; Fig. 4) and PVE values ranged from 34 to 36%. As expected, APR was associated with the amplicon from HMZ (146 bp; so-called R-allele) at Ta2BL_ssr7 and RL6058 produced an alternative amplicon (131 bp; S-allele). The sequence of wmc499 was aligned to the 2BL sequences of IWGSC RefSeq v1.0 physical map at about 594.44 Mb, while Ta2BL_ssr7 and Ta2BL_ssr16 were located at approximately 597.88 Mb and 600.14 Mb, respectively (Tables 4 and 5).
Partial genetic map of 2BL of RL6058 × Hongmazha, illustrating the logarithm of the odds (LOD) value curves of QLr.cau-2BL. Marker names and intervals (in Kosambi cM) between adjacent markers are shown along the chromosome orientated with the centromere to the left. Horizontal line indicates the threshold LOD of 3.2
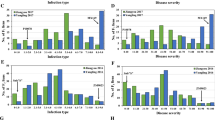
The 371 RL6058 × HMZ RILs, segregating for Lr34 as well as for QLr.cau-2BL, produced a 751 bp amplicon (R-allele) and an alternative 523 bp amplicon (S-allele) at the locus cssf5 that is a marker diagnostic for Lr34 (Lagudah et al. 2009). Selection based on Ta2BL_ssr7 and cssf5 for QLr.cau-2BL and Lr34, respectively, divided the 371 RILs into four groups, i.e., the RILs that carried resistant allele at both QLr.cau-2BL and Lr34, at QLr.cau-2BL alone, at Lr34 alone, and none at both, respectively. As shown in Fig. 5, AUDPC mean was significantly (P < 0.0001) higher in “None,” lower in “QLr.cau-2BL alone” and “Lr34 alone,” and further lower in “QLr.cau-2BL + Lr34” in each of the three experiments.
Boxplots showing the effects of QTL combinations on disease severity in RL6058 × Hongmazha RIL population. The y-axis defines the score of disease severity (AUDPC) , and the x-axis defines four RIL groups with different QTL combinations, i.e., presence of resistance allele at both QLr.cau-2BL (represented by the 146 bp amplicon) plus Lr34 (751 bp), QLr.cau-2BL alone (146 bp), Lr34 alone (751 bp), and None (131 bp at QLr.cau-2BL and 523 bp at Lr34), respectively. Disease was assessed in the field of Gansu (a), Henan (b) and Shandong (c). *** Indicates significant difference at α = 0.0001 based on a Fisher’s least significant difference (LSD) test. NS means no significant difference. Within each box, the small diamond and the horizontal line indicate the mean and median AUDPC, respectively. The top and bottom edges of a box illustrate the 75th and 25th percentiles, respectively. Whiskers (vertical lines outside a box) extend to the extreme data points, and small circles denote outliers
MAS based on Ta2BL_ssr7
Effectiveness of selection for QLr.cau-2BL based on Ta2BL_ssr7 was examined with the tests on the plants of Aquileja × HMZ F2:3 and Jimai22 × HMZ BC4F2:3 in a greenhouse. DNA sample extracted from 1st-leaf stage F3 seedlings of each Aquileja × HMZ F2:3 family was used as template for PCR amplification with Ta2BL_ssr7 primers (Table 5). These families produced three genotypes in total, i.e., homozygous resistant (146 bp amplicon), homozygous susceptible (131 bp amplicon), and heterozygous (146 bp and 131 bp bands). Twenty of the F2:3 families were selected for homozygous R-allele and another 20 ones were selected for homozygous S-allele. Disease tests of the selected families indicate that leaf rust severity mean value was significantly (P < 0.0001) lower on R-group F2:3 plants than on S-group plants (Fig. 6a; Online Resource 1 Table S6).
Boxplots showing effects of the marker Ta2BL_ssr7 (representing QLr.cau-2BL) on disease severity in the F2:3 population of Aquileja × Hongmazha (a) and the BC4F2:3 population of Jimai22 × Hongmazha (b). The y-axis defines the score of disease severity, and the x-axis defines different groups of test plants. R-group carried the resistant allele in homozygous status at QLr.cau-2BL, and S-group carried the susceptible allele in homozygous status. *** Indicates a significant difference at α = 0.0001 based on a Fisher’s least significant difference (LSD) test. Refer to the caption of Fig. 5 for descriptions of box
Likewise, Jimai22 × HMZ BC4F2:3 families were segregating at Ta2BL_ssr7 for homozygous resistance (146 bp amplicon) and homozygous susceptibility (131 bp amplicon). Twenty BC4F2:3 families were selected for homozygous R-allele and another 20 ones were selected for homozygous S-allele. Leaf rust severity mean value was significantly (P < 0.0001) lower on R-group BC4F2:3 plants than on S-group plants (Fig. 6b; Online Resource 1 Table S7). These results show that selection for QLr.cau-2BL based on Ta2BL_ssr7 was well effective in both cases of F2:3 and BCF2:3.
A set of 282 commercial wheat cultivars was detected for polymorphism at Ta2BL_ssr7 (Online Resource 1 Table S1). Of these cultivars, Tao954-1and Tao7230 produced 146 bp amplicon similar to HMZ (R-allele), 220 showed a 131 bp amplicon (S-allele), and 60 displayed a 155 bp amplicon (another S-allele; disease data not shown).
Discussion
A QTL (QLr.cau-2BL) for APR to leaf rust was detected and an SSR marker (Ta2BL_ssr7) closely linked with QLr.cau-2BL was identified using four populations. We confirmed that Ta2BL_ssr7 is a robust marker for MAS. Our investigations during the period 2011–2017 indicated that 282 cultivars used in panel were commercially grown on large scales in China (with approximately 13.5 million ha in 2017; Online Resource 1 Table S1). Some of these cultivars were excellent in grain yield and adaptability, and these were widely grown; for instance, Jimai22 occupied more than a million ha in 2017. Of the 282 cultivars, some are still being commercially planted and the others are being used as parents for breeding new cultivars; however, 237 including Jimai22 were susceptible to leaf rust at both seedling and adult plant growth stages (Zhang’s Lab., unpublished data). Only two (Tao954-1and Tao7230) of the 282 cultivars produced the same 146 bp amplicon as HMZ (R-allele; Online Resource 1 Table S1) at Ta2BL_ssr7 and the others showed alternative amplicons (131 bp or 155 bp; S-allele). Tao954-1and Tao7230 were cultivated on limited scales (< 3000 ha annually). We therefore believe that QLr.cau-2BL has not yet been substantially utilized in wheat breeding in China. MAS based on Ta2BL_ssr7 should facilitate the use of QLr.cau-2BL in practical breeding.
RL6058 carries Lr34 and thus RL6058 × HMZ RILs were segregating for QLr.cau-2BL and Lr34. This provides us an opportunity for comparing the additive effect of these QTL. The selection based on Ta2BL_ssr7 and cssfr5 for QLr.cau-2BL and Lr34, respectively, divided the RL6058 × HMZ RILs into different groups (Fig. 5). Comparison in disease severity between groups indicates that QLr.cau-2BL could reduce leaf rust severity similar to Lr34 and acted in an additive manner (Fig. 5). Additive effect was also described for Lr34 with Lr68 (Herrera-Foessel et al. 2012). However, Lr34 has been reported to work with certain rust resistance QTL in non-additive manners. For instance, Kolmer et al. (2011) indicate that Lr34 can enhance the expression of an adult plant stem rust resistance QTL on 2BL in Thatcher, although Lr34 by itself does not condition stem rust resistance. Rosewarne et al. (2013) address the interaction of Yr18 with Yr29 and further indicate that in the presence of Lr34, Yr29 was less effective against stripe/yellow rust.
QLr.cau-2BL showed consistence effect on reducing disease severity across all of the three wheat populations (Aquileja × HMZ, RL6058 × HMZ and Jimai22 × HMZ) and across all of the eight experiments (Table 3; Fig. 6). The wheat lines used in these crosses represent considerably diverse genetic backgrounds. Aquileja is an Italian winter wheat, and RL6058, a spring wheat, originates from North America. The Chinese winter wheat Jimai22 is morphologically and phenologically different from Aquileja, for instance, in plant height and maturity time. The eight experiments represent significantly different environments. For example, the test site in Gansu is geographically separated by more than 1100 km from the site in Shandong. In Gansu, the experimental plots were located at an elevation of 1650 m and wheat plants mature in the late July. The plots in Shandong were located at an elevation of 90 m and wheat matures in mid-June.
The APR in HMZ, primarily conferred by QLr.cau-2BL, might be effective against many P. triticina races than the artificially inoculated race THTT. We have observed HMZ for its APR in field nurseries in Gansu and Shandong since 2002. Leaf rust epidemic in the nurseries was induced by both artificially inoculated race (THTT) and naturally occurring urediniospores. We analyzed some uredinial samples once every three years or so and always found multiple races. For example, 11 races were identified from the experimental plots of Shandong in 2018 (Table 3). These races were collectively virulent to 27 Lr genes including 14 ones in the wheat differentials (Lr1, Lr2a, Lr2c, Lr3, Lr16, Lr26, Lr3ka, Lr11, Lr17, Lr30, LrB, Lr10, Lr14a and Lr18) and 13 present in other lines (Lr2b, Lr3bg, Lr14b, Lr15, Lr17a, Lr20, Lr21, Lr23, Lr25, Lr32, Lr33, Lr36 and Lr50) as tested by Li et al. (2010). In addition, THTT was virulent to Lr12, Lr13, Lr22b and Lr37 at the adult plant growth stage (Zhang’s Lab, unpublished data). Similarly, nine races were found from the plots of Gansu and Shandong in 2014 (Du et al. 2015). More races were comprehensively surveyed in China by other researchers. Liu and Chen (2012) reported that 79 P. triticina races were detected from 613 single-uredinial isolates collected between 2000 and 2006 from the provinces including Gansu, Henan, and Shandong. Kolmer (2015) and Ma (2020) also addressed the race diversity in these provinces. We believe that HMZ might have been subjected to inoculations by diverse races and the APR in HMZ was effective to these races. From 2002 to 2020, leaf rust severity was consistently less than 35% on HMZ, in contrast around 90% on the susceptible control wheat Thatcher in our field nurseries.
According to a recent review on QTL for APR to leaf rust (Silva et al. 2018), 14 QTL were previously reported to be located on 2BL. Ten of these QTL were mapped within intervals delimited by flanking markers and the others were tagged with single markers. These 14 QTL appear to be quite diverse with respect to effect magnitude, consistence of effectiveness, resistance component, and pleiotropy. Six QTL (QLr.osu-2B, QLrlp.osu-2B, QLr.inra-2B, QLr.cim-2B, QTL-2BL, and QLr.ifa-2BL) were mapped using biparental populations (Online Resource 1 Table S8), and the other eight were detected using wheat panels for genome-wide association study (GWAS). The resistance alleles at QLr.osu-2B and QLrlp.osu-2B originated from the wheat line CI 13227 and were located in the vicinity of SSR marker barc167 (Xu et al. 2005a, b). These two QTL had pleiotropic relationship for decreasing infection rate and prolonging latent period with PVE values up to 16.6% (Xu et al. 2005a, b). QLr.inra-2B from the French cultivar Apache could lengthen latent period (PVE > 20%) without significant effect on other resistance components such as infection efficiency and lesion size (Azzimonti et al. 2014). The minor QTL QLr.cim-2B from the cultivar Avocet-S was associated with low disease severity (Ren et al. 2017). QTL-2BL from the Mexican cultivar Opata 85 reduced disease severity with a PVE value of 12.0% (Nelson et al. 1997). QLr.ifa-2BL from the Austrian cultivar Capo was effective in multiple environments (overall PVE = 10.5%), and this QTL was coincided, in chromosome position, with a QTL for yellow rust resistance (Buerstmayr et al. 2014).
The eight GWAS QTL (QLr.IWA2025, QLr.IWA207, QLr.IWA5177, QLr.IWA2509, QLr.wpt-2BL, qNV.Lr-2B.2, qNV.Lr-2B.3, and Lr_Ho; Online Resource 1 Table S8) were mapped using diverse wheat panels involving landraces, core collections, registered cultivars, breeding lines, and synthetic hexaploid wheats (Gerard et al. 2018; Joukhadar et al. 2020; Riaz et al. 2018; Turner et al. 2017). All these QTL were associated with low disease severity. QLr.IWA2025, QLr.IWA207, QLr.IWA5177 and QLr.WA2509 showed PVE values up to 16.3% (Turner et al. 2017). QLr.wpt-2BL, identified based on analyses of 96 wheat accessions chosen from a larger panel (710 genotypes), expressed in all six experiments with PVE values from 7.8 to 11.7% (Gerard et al. 2018). qNV.Lr-2B.2 and qNV.Lr-2B.3, detected using 288 wheat lines originating from the “N. I. Vavilov Institute of Plant Genetic Resources,” were effective in multiple environments (Riaz et al. 2018). The minor QTL Lr_Ho was mapped with high precision applying a large wheat panel (2300 accessions), and this QTL was suggested to be a contribution to durable resistance (Joukhadar et al. 2020).
We inferred the physical positions of the 14 QTL by aligning their flanking/tagging marker sequences to the 2BL sequences of IWGSC RefSeq v1.0 (IWGSC 2018) (Fig. 7; Online Resource 1 Table S8). All of these QTL (except Qlr.inra-2B) are separated from QLr.cau-2BL by at least 70 Mb (Fig. 7). Qlr.inra-2B originates from the French wheat cultivar Apache (Azzimonti et al. 2014). HMZ produced a 146 bp amplicon (R-allele) at Ta2BL_ssr7, whereas Apache produced a 131 bp amplicon (S-allele) (Online Resource 1 Table S1). On the basis of these results, we believe that QLr.cau-2BL is highly likely to be a previously unreported QTL for APR to leaf rust, although further tests are required for a conclusive novelty.
Physical position of QLr.cau-2BL compared with the previously reported QTL for adult plant resistance (ARP) to leaf rust on 2BL (10-Mb tick size map). Black bar indicates an interval harboring a QTL flanked by two markers, and dot indicates a single marker representing a QTL. QLr.cau-2BL was written in increased font. Refer to Online Resource 1 Table S8 for detailed information including full references
In addition to the 14 APR QTL, two designated Lr genes were previously mapped on 2BL, i.e., Lr50 linked (6.7 cM) with the SSR marker gwm382 near the telomere of 2BL (Brown-Guerdira et al. 2003) and Lr58 that co-segregated with the SSR marker cfd50 in the distal region of 2BL (Kuraparthy et al. 2007). Both genes confer all-stage race-specific resistance and were transferred to T. aestivum from T. timopheevii and Aegilops triuncialis, respectively; thus, both genes should be genetically different from QLr.cau-2BL that is harbored in a T. aestivum landrace (i.e., HMZ).
Realizing that QLr.cau-2BL alone may provide inadequate level of APR if disease pressures are high, we are further checking the combinability of QLr.cau-2BL with APR including Lr46 and Lr67. The present study indicates that QLr.cau-2BL acted synergistically with L34 and consequently reduced leaf rust to an adequate level (Fig. 5). Fine mapping of QLr.cau-2BL is also underway.
Abbreviations
- 2BL:
-
The long arm of chromosome 2B
- ANOVA:
-
Analysis of variance
- APR:
-
Adult plant resistance
- AUDPC:
-
The area under the disease progress curve
- H 2 :
-
Broad-sense heritability
- HMZ:
-
Hongmazha, a wheat landrace
- IT:
-
Infection type
- LOD:
-
Logarithm of odds
- Lr gene:
-
Leaf rust resistance gene
- MAS:
-
Marker-assisted selection
- PVE:
-
Phenotypic variation explained
- QTL:
-
Quantitative trait locus / loci
- RIL:
-
Recombinant inbred line
- SNP:
-
Single nucleotide polymorphism
References
Azzimonti G, Marcel TC, Robert O, Paillard C, Lannou C, Goyeau H (2014) Diversity, specificity and impacts on field epidemics of QTL involved in components of quantitative resistance in the wheat leaf rust pathosystem. Mol Breed 34:549–567
Bassam BJ, Caetano-Anolles G, Gresshoff PM (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196:80–83
Brown-Guerdira GL, Singh S, Fritz AK (2003) Performance and mapping of leaf rust resistance to wheat from Triticum timopheevii subsp. ameniacum. Phytopathology 93:784–789
Buerstmayr M, Matiasch L, Mascher F, Vida G, Ittu M, Robert O, Holdgate S, Flath K, Neumayer A, Buerstmayr H (2014) Mapping of quantitative adult plant field resistance to leaf rust and stripe rust in two European winter wheat populations reveals colocation of three QTL conferring resistance to both rust pathogen. Theor Appl Genet 127:2011–2028
Caldwell RM, Schafer JF, Compton LE, Patterson FL (1957) A mature plant type of wheat leaf-rust resistance of composite origin. Phytopathology 47:691–692
Clark JA (1935) Registration of improved wheat varieties VIII. J Am Soc Agron 27(1):71
Dangl JL, Horvath DM, Staskawicz BJ (2013) Pivoting the plant immune system from dissection to deployment. Science 341:746–751
Du ZY, Che MZ, Li GH, Chen J, Quan W, Guo Y, Wang Z, Ren JD, Zhang HY, Zhang ZJ (2015) A QTL with major effect on reducing leaf rust severity on the short arm of chromosome 1A of wheat detected across different genetic backgrounds and diverse environments. Theor Appl Genet 128:1579–1594
Gerard GS, Kobiljskic B, Lohwasserd U, Borner A, Simon MR (2018) Genetic architecture of adult plant resistance to leaf rust in a wheat association mapping panel. Plant Pathol 67:584–594
Herrera-Foessel SA, Singh RP, Huerta-Espino J, Rosewarne GM, Periyannan SK, Viccars J, Calvo-Salazar V, Lan C, Lagudah ES (2012) Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theor Appl Genet 124:1475–1486
Holland JB, Nyquist WE, Cervantes-Martinez C (2003) Estimating and interpreting heritability for plant breeding: an update. Plant Breeding Reviews 22:9–112
Huerta-Espino J, Singh RP, Germán S, McCallum BD, Park RF, Chen WQ, Bhardwaj SC, Goyeau H (2011) Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179:143–160
International Wheat Genome Sequencing Consortium (IWGSC) (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788
International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:7191–7203
Johnson R (1981) Durable resistance: definition of, genetic control, and attainment in plant breeding. Phytopathology 71:567–568
Joukhadar R, Hollaway G, Shi F, Kant S, Forrest K, Wong D, Petkowski J, Pasam R, Tibbits J, Bariana H, Bansal U, Spangenberg G, Daetwyler H, Gendall T, Hayden M (2020) Genome-wide association reveals a complex architecture for rust resistance in 2300 worldwide bread wheat accessions screened under various Australian conditions. Theor Appl Genet 133:2695–2712
Kolmer JA (2015) Collections of Puccinia triticina in different provinces of China are highly related for virulence and molecular genotype. Phytopathology 105:700–706
Kolmer JA, Garvin DF, Jin Y (2011) Expression of a Thatcher wheat adult plant stem rust resistance QTL on chromosome arm 2BL is enhanced by Lr34. Crop Sci 51:526–533
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Kuraparthy V, Sood S, Chhuneja P, Dhaliwal HS, Kaur S, Bowden RL, Gill BS (2007) A cryptic wheat-Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci 47:1995–2003
Lagudah ES (2011) Molecular genetics of race non-specific rust resistance in wheat. Euphytica 179:81–91
Lagudah ES, Krattinger SG, Herrera-Foessel S, Singh RP, Huerta-Espino J, Spielmeyer W, Brown-Guedira G, Selter LL, Keller B (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Li HH, Ye GY, Wang JK (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Li ZF, Xia XC, He ZH, Li X, Zhang LJ, Wang HY, Meng QF, Yang WX, Li GQ, Liu DQ (2010) Seedling and slow rusting resistance to leaf rust in Chinese wheat cultivars. Plant Dis 94:45–53
Liu TG, Chen WQ (2012) Race and virulence dynamics of Puccinia triticina in China during 2000–2006. Plant Dis 96:1601–1607
Long DL, Kolmer JA (1989) A North American system of nomenclature for Puccinia recondita f. sp. tritici. Phytopathology 79:525–529
Long DL, Kolmer JA, Leonard KJ, Hughes ME (2002) Physiologic specialization of Puccinia triticina on wheat in the United States in 2000. Plant Dis 86:981–986
Ma Y, Liu T, Liu B, Gao L, Chen W (2020) Population genetic structures of Puccinia triticina in five provinces of China. Eur J Plant Pathol 156:1135–1145
McDonald RA, Linde C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol 40:349–379
McIntosh RA, Dubcovsky J, Rogers JW, Morris C, Xia CX (2017) Catalogue of gene symbols for wheat: 2017 supplement. https ://shige n.nig.ac.jp/wheat /komug i/genes /macge ne/suppl ement 2017.pdf. Accessed from 23 Aug 2020
Moore JW, Herrera-Foessel S, Lan CX, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong X, Spielmeyer W, Talbot M, Bariana H, Patrick JW, Dodds P, Singh R, Lagudah E (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Nelson JC, Singh RP, Autrique JE, Sorrells ME (1997) Mapping genes conferring and suppressing leaf rust resistance in wheat. Crop Sci 37:1928–1935
Park RF, McIntosh RA (1994) Adult plant resistances to Puccinia recondita f. sp. tritici in wheat. N Z J Crop Hortic Sci 22:151–158
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity of leaves and stems of cereals. Can J Res 26:496–500
Qureshi N, Bariana H, Kumran VV, Muruga S, Forrest KL, Hayden MJ, Bansal U (2018) A new leaf rust resistance gene Lr79 mapped in chromosome 3BL from the durum wheat landrace Aus26582. Theor Appl Genet 131:1091–1098
Ren Y, Singh RP, Basnet BR, Lan CX, Huerta-Espino J, Lagudah ES, Ponce-Molina LJ (2017) Identification and mapping of adult plant resistance loci to leaf rust and stripe rust in common wheat cultivar Kundan. Plant Dis 101:456–463
Riaz A, Athiyannan N, Periyannan SK, Afanasenko O, Mitrofanova OP, Platz GJ, Aitken EAB, Snowdon RJ, Lagudah ES, Hickey LT, Voss-Fels KP (2018) Unlocking new alleles for leaf rust resistance in the Vavilov wheat collection. Theor Appl Genet 131:127–144
Röder MS, Korzun V, Wandehake K, Planschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rosewarne GM, Herrera-Foessel SA, Singh RP, Huerta-Espino J, Lan CX, He ZH (2013) Quantitative trait loci of stripe rust resistance in wheat. Theor Appl Genet 126:2427–2449
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Silva GBP, Zanella CM, Martinelli JA, Chaves MS, Hiebert CW, McCallum BD, Boyd LA (2018) Quantitative trait loci conferring leaf rust resistance in hexaploid wheat. Phytopathology 108:1344–1354
Singh RP, Mujeeb-Kazi A, Huerta-Espino J (1998) Lr46: a gene conferring slow-rusting resistance to leaf rust in wheat. Phytopathology 88:890–894
Singh RP, Huerta-Espino J, Rajaram S (2000) Achieving near-immunity to leaf and stripe rusts in wheat by combining slow rusting resistance genes. Acta Phytopathol Entomol Hung 35:133–139
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: JOINMAP. Plant J 3:739–744
Turner MK, Kolmer JA, Pumphrey MO, Bulli P, Chao S, Anderson JA (2017) Association mapping of leaf rust resistance loci in a spring wheat core collection. Theor Appl Genet 130:345–361
Wang S, Basten JC, Zeng ZB (2010) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC. http://statgen.ncsu.edu/qtlcart/WQTLCart.htm. Accessed from 24 Jul 2020
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, International Wheat Genome Sequencing Consortium, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Xu X, Bai G, Carver BF, Shaner GE, Hunger RM (2005a) Molecular characterization of slow leaf-rusting resistance in wheat. Crop Sci 45:758–765
Xu XY, Bai GH, Carver BF, Shaner GE, Hunger RM (2005b) Mapping of QTL prolonging the latent period of Puccinia triticina infection in wheat. Theor Appl Genet 110:244–251
Acknowledgement
We thank Junzhi Wang, Xinfa Mou, Xinhu Wang, Zonghai Bai, and Yanrun Lin for providing excellent technical assistance during the field work. We thank Dr. Ziyi Du for providing the picture of Figure 1. We acknowledge the enormous help of numerous graduate, undergraduate, and high school students who were associated with the Program for Accumulating Quantitative Resistance to Diseases, Plant Pathology Department, China Agricultural University. This study was supported by the China Postdoctoral Science Foundation (2020M670538), National Key Research and Development Program of China (2016YFD02000 and 2018YFD0200500), and Beijing Academy of Agriculture and Forestry Sciences Sci-Tech Innovation Capacity Building Program (KJCX20200115).
Author information
Authors and Affiliations
Contributions
ZZ and WQ conceived the study. YZ and ZW designed and managed the experiments, developed the SSR markers of 2BL, and constructed the genetic maps. YZ, ZW, WQ, JF, JR, ZZ and XJ performed the disease experiments and/or SSR genotyping. XZ statistically analyzed the disease data. ZZ and XZ collected the disease data from greenhouse experiment. ZZ, XZ, QW and JF developed the RILs. YZ and ZW drafted the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The authors declare that the experiments comply with the current laws of China.
Additional information
Communicated by Urmil Bansal.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Zhang, Y., Wang, Z., Quan, W. et al. Mapping of a QTL with major effect on reducing leaf rust severity at the adult plant growth stage on chromosome 2BL in wheat landrace Hongmazha. Theor Appl Genet 134, 1363–1376 (2021). https://doi.org/10.1007/s00122-021-03776-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03776-4