Abstract
Key message
The efficiency of phenotype-based assessments of plant variety protection and registration could be improved by the integration of DNA-based testing. We review the current and proposed models in the era of next-generation breeding.
Abstract
The current plant variety protection system relies on morphological description of plant varieties. Distinctness, uniformity, and stability (DUS) assessments determine whether a new variety is distinguishable from common knowledge varieties and exhibits sufficient phenotypic uniformity and stability during two independent growing cycles. However, DUS assessment can be costly, time-consuming and often restricted to a relatively small number of traits that can be influenced by environmental conditions. This calls for the adoption of a DNA-based system which is endorsed by the International Union for the Protection of New Varieties of Plants (UPOV). This could enable examiners to deploy trait-specific DNA markers in DUS testing as well as using such genetic markers to manage reference collections. Within UPOV’s system, breeders can freely use protected varieties in breeding programs. However, breeders of protected varieties may seek sharing in ownership of essentially derived varieties once it is proven that they, with the exception of a few distinctive DUS trait(s), conform to parental varieties in essential characteristics. As well as their complementary role in DUS testing, DNA markers have been known as a good replacement of morphological traits in defining boundaries between independently and essentially derived varieties. With the advent of new breeding technologies that allow minor modification in varieties with outcomes of specific merit or utility, detecting distinctness between varieties may become increasingly challenging. This, together with the ever-increasing number of varieties with which to compare new candidate varieties, supports the potential utility of using DNA-based approaches in variety description.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
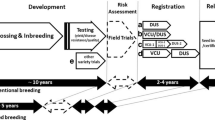
Breeding new plant varieties is a costly task in terms of skill, infrastructure and genetic resources. It can also be time-consuming, taking for example around 10 years to release a barley variety via pure line selection. Plant breeders have various options to protect the ownership of their new varieties in order to prevent unauthorized utilization. Such protection affords exclusive rights and is a means for recouping research and development costs via the collection of royalties, so supporting further breeding activities.
Among different mechanisms, plant variety protection (PVP) via the International Union for the Protection of New Varieties of Plants (UPOV) represents a harmonized system for the awarding of intellectual property rights (IPR) among its 75 member states and organizations (as of February 2019) for recognizing plant breeder’s rights/plant variety rights (PBR/PVR). Under the 1991 Act (latest revision) of the UPOV convention, PBR are granted for a period of not less than 20 years to protected varieties (UPOV 1991). In some countries, such as the USA, Japan, and Australia, breeders can also apply for a grant of plant or utility patent rights, if new varieties satisfy the required criteria. Alternative forms of protection for plant varieties and attaining IPR exist, such as contract law, registered trademarks, and trade secrets. The latter is particularly suitable for protection of the parental lines of hybrid varieties (Curtis and Nilsson 2012). In addition, breeders can benefit from self-regulatory protection systems, such as hybrid seed and variety-specific genetic use restriction technologies (reviewed by Lombardo 2014), which require growers to buy seed for each growing season.
Within the framework of UPOV’s PVP system, new varieties must satisfy three technical criteria to be eligible for protection. They should be distinct (D) from any other common knowledge varieties (CKVs) in at least one morphological trait; sufficiently uniform (U) in relevant characteristics depending on reproduction system of species; and stable (S) in expression of traits after two independent growing cycles. A total of 325 test guidelines are currently available in different plant species (as of February, 2019) for conducting such “DUS” tests. In some species, electrophoresis of isozymes (in sunflower, soybean, and maize) or storage proteins (glutenins in wheat and hordeins in barley) have been incorporated as additional characteristics in test guidelines. Although DUS testing is the pivotal component of UPOV’s PVP system, it is also a compulsory requisite for registering arable crops, vegetables, and fruits in the European Union’s Common Catalogue, prior to marketing. Moreover, agricultural crop varieties should exhibit “Value for Cultivation and Use” (VCU) in comparison with existing check cultivars across multi-environmental trials (Cooke and Reeves 2003). Thus, while the “utility” of new crop varieties is evaluated by the VCU process, variety descriptions, as the outcome of DUS tests, describe their morphological “identity”. In 2008–2009, the European Commission opened legal registration of agricultural and vegetable “conservation varieties” in the Common Catalogue, helping to address the gradual genetic erosion that has historically taken place as landraces were in more recent decades supplanted by more genetically uniform varieties. For example, “list C” of the French Official Catalogue currently maintains registration records of nine potato landraces and one maize open-pollinated population. However, the inherent heterogeneity of landraces demands applying a looser uniformity (i.e. U within DUS tests) standard for registration. For instance, up to 10% aberrant plant frequency may be tolerated for a landrace variety. Moreover, they are exempted from VCU assessment unless an exceptional merit or utility is claimed. Despite registration of landraces that authorize limited marketing of seeds, they are excluded from scope of protection under PVP system of UPOV convention as they cannot fulfil novelty criteria (not released more than 1 year before in the country). In contrast, extant varieties (including landraces and farmer’s varieties) are subject of protection in India (not a UPOV member) under the Protection of Plant Varieties & Farmer’s Rights (PPV&FR) Act of 2001.
The increased availability and cost-efficiency of DNA-based markers in recent years makes them attractive options to explore their use to supplement, or even ultimately replace, existing morphological- and protein-based approaches. Indeed, exploring such options is the focus of UPOV’s working group on “Biochemical and Molecular Techniques and DNA Profiling in Particular” (BMT). So far, two models, i.e. “characteristic-specific molecular markers” and “combining phenotypic and molecular distances in the management of variety collections” have been favoured (UPOV 2013a). Given the importance of this issue, the Community Plant Variety Office (CPVO), which administers the system of IPR within the European Union and is itself a member of the UPOV Convention, has also established a think-tank of experts for the “Integration of Molecular Data into DUS Testing” (IMODDUS) and acts to define relevant funding topics under the pan-European “Horizon 2020” research programme.
In this review, we summarise the literature over the last 2 decades regarding the application of molecular markers in DUS testing and the identification of EDVs, highlight two new models favoured by UPOV, and discuss future prospects and challenges associated with new breeding technologies for both DUS testing and the consideration of DNA-based variety description.
Why are DNA marker approaches being evaluated for possible future use in DUS testing?
DUS examiners are always looking for ways of improving the current system of variety protection and registration, which currently relies predominantly on morphological traits. In the current system, morphological traits may be influenced by environmental factors, including between site and between year factors, which complicate evaluation (Nuel et al. 2001; Cooke and Reeves 2003; Singh et al. 2004). This is especially true for the evaluation of disease resistance traits (which are obligatory in some vegetable varieties, e.g. tomato. Arens et al. 2010), or traits of a quantitative nature (e.g. the time of flowering) which comprise the majority of descriptors in many outbreeding species, such as ryegrass (Wang et al. 2016). The measurement of traits is also time-consuming and requires skill and adequate experience (Reid and Kerr 2007) and in some cases, special conditions or trial designs are required to conduct the tests (e.g. for assessment of the requirement for vernalization in barley and wheat). Moreover, establishing DUS criteria in some crops, such as ryegrass (Wang et al. 2016) and alfalfa (Annicchiarico et al. 2016), is arduous due to the high level of intra-population variation within cultivars.
In contrast, DNA markers are free from environmental effects (Noli et al. 2008) and could be applied to seed or early stages of plant growth. Therefore, they have the potential to greatly reduce the duration and cost of DUS tests, given the development of an appropriate regulatory system with which to govern their use. With ever-increasing numbers of plant varieties, conducting DUS trials is getting increasingly expensive (Lombard et al. 2000; Reid and Kerr 2007; Gunjaca et al. 2008; Wang et al. 2016). For instance, in the case of maize in China, there were a total of 6291 registered varieties as of 2013, and these all need to be grown out in DUS tests alongside the new candidate varieties for every DUS assessment, highlighting the laborious and costly process of field assessment (Tian et al. 2015). Thus, DNA markers could be an efficient approach for pre-screening or grouping reference varieties before sowing in the field. Thus, only those reference varieties sufficiently similar to the candidates under test would be utilised, so reducing overall costs (Lombard et al. 2000; Tommasini et al. 2003; Noli et al. 2008; Smỳkal et al. 2008). Moreover, they may be helpful where current DUS traits fail to differentiate varieties (Bernet et al. 2003; Gunjaca et al. 2008; Bonow et al. 2009).
However, the potential use of DNA-based approaches should be balanced against the proven efficacy of current DUS systems, and the associated cost and disruption that would be associated with the validation and implementation of a significantly altered system. Indeed, the use of DNA markers for DUS testing has been faced with cautious consideration by the International Seed Federation (ISF), a non-governmental organization that represents the interests of the seed industry at a global level. In their view, the sole use of anonymous DNA-based markers (with no relevance to phenotypic DUS characteristics) would jeopardize the essence of PBR, by theoretically reducing the minimum distance between varieties to a difference of only one DNA base pair. However, utilization of markers under currently recommended models is supported in such way that they be included in DUS test guidelines after reaching a harmonized method of publicly available markers (ISF 2012).
A review of the literature finds that amplified fragment length polymorphisms (AFLPs) and simple sequence repeats (SSRs) represent the most-used marker technologies for evaluation of the potential of DNA-based approaches as complementary traits in DUS testing of agricultural crops, vegetables, fruit and forest trees (Table 1). Of these, SSRs have the advantage of being highly polymorphic and originate from defined chromosomal locations. Single nucleotide polymorphisms (SNPs) have more recently been used in distinctness testing of maize and alfalfa varieties. Compared to SSRs, bi-allelic SNPs are more abundant in plant genomes, making them the marker of choice for DNA fingerprinting in modern plant breeding and plant research programs.
Assessment of distinctness (D)
While the AFLP-based studies use markers that originate from genomic regions not a priori known to be linked to genetic loci controlling DUS characteristics, studies using other marker types have been more targeted. For example, expressed sequence tag (EST)-based SSRs derived from transcribed genomic regions have been used that represent markers within genes with speculated relevance to DUS traits. Bonow et al. (2009) used 13 SSRs from the region upstream of MADS-box genes and 37 EST-SSRs for the identification of 37 closely related rice varieties in Brazil. The resulting genotypic data allowed differentiation of japonica and indica rice subspecies. Moreover, as per application of multivariate analyses (e.g. cluster and principal component analysis, PCA), varieties were grouped according to some attributes. Namely, SSRs separated rice varieties into basmati and non-basmati groups as well as showing congruency with pedigree and breeding history of 14 rice varieties (Sarao et al. 2009). Similarly, SSR markers showed better performance than DUS traits in distinguishing and fitting with pedigree of 41 maize inbred lines (Gunjaca et al. 2008). In rapeseed, two AFLP primer combinations could distinguish and separate 83 cultivars based on seasonal growth habit (winter versus spring), breeding company, and country of origin (France or Germany) using PCA and analysis of molecular variance (AMOVA) (Lombard et al. 2000). Other studies in rapeseed also identified SSRs that delineated varieties according to their type (winter, spring, and forage). Moreover, genetic distance calculated on markers revealed better consistency correlated with pedigree of varieties (Tommasini et al. 2003). In sugar beet, cluster analysis based on Nei’s genetic distances based on AFLP data on 15 varieties grouped them according to their breeding programme origins (De Riek et al. 2001). Finally, AFLPs have been found to allow clustering of 66 poplar commercial clones, consisting of various species and hybrids, to their correct taxonomic classification (Fossati et al. 2005). These results suggest that selected markers can be effective in grouping varieties according to their population structure.
While these studies demonstrate the potential of DNA markers as complementary traits to classical morphological descriptors, other advantages, such as use in the management of reference collections for improving the quality of field comparisons, has also been highlighted (Lombard et al. 2000; Tommasini et al. 2003).
Assessment of uniformity (U) and stability (S)
Relatively few studies have investigated the utility of molecular markers for the assessment of uniformity and stability. Investigation using 55 SSRs on DNA extracted from 20 individuals of a single rice cultivar obtained from multiple sources (nucleus, breeder, foundation, certified, and farmer’s saved seed) found no variation between 100 plants (20 individuals × 5 seed classes), indicating SSRs potentially represent a useful tool for uniformity and stability tests (Singh et al. 2004). AFLPs have also been used for uniformity assessment in morphologically heterogeneous rapeseed inbred lines, where similar detection rates for off-types were obtained using PCA and Rogers’ distances, compared to those observed in the field (Lombard et al. 2002). Also, 15 SSRs have been used for the uniformity assessment of 10 rapeseed varieties, where different values of intra-varietal diversity were detected depending on variety tested and SSR used (Tommasini et al. 2003). Cooke et al. (2003) used SSRs for assessment of uniformity in wheat and tomato varieties—in spite of observing non-uniformity, 24 out of 45 wheat varieties met the uniformity standards by using seven to nine SSR loci. Also, nine of the ten tomato varieties were uniform using six SSR loci. Higher levels of uniformity were observed using SSRs than morphological traits in 3 (out of 4) hybrid varieties of Brassica chinensis (Yim et al. 2009). Furthermore, 9 polymorphic SSRs were used to assess the utility of genetic markers for the assessment of distinctness in grapevine and have also been assessed for application to uniformity and stability testing. These markers were successful in assessments in nearly all 171 variety-SSR combinations examined, except in one combination where the number of off-types did not meet the uniformity standard (Vélez and Ibánez 2012). In a study investigating stability testing in wheat, Wang et al. (2013) developed an SSR-based index as a replacement for field-based stability tests. In their method, varieties with a homozygous SSR loci ratio (SSR-HLR) more than 95% and less than 91% were deemed to be stable and unstable, respectively, while those between these two thresholds were entered into field tests for morphological assessment. Based on SSR-HLR, the stability test of 633 regional varieties was enabled by a set of 80 polymorphic markers. More recently, Cockram (unpublished) demonstrated that Kompetitive Allele-Specific PCR (KASP) SNP-based markers diagnostic for selected DUS traits in barley could be used to robustly detect off-types in DNA representing pools of ten individuals. This approach highlights how SNP-based markers could potentially be used as a like-for-like replacement for morphological traits—at least in those cases where genetics underlying the traits are well described. In the medium term, this could potentially be of relevance to morphological traits or grouping characteristics which are difficult, costly, or time-consuming to phenotype.
BMT models for application of DNA markers in DUS assessment
Characteristic-specific molecular markers
The use of gene- or allele-specific markers that accurately predict states of expression of phenotypic traits used in conventional DUS testing is known within UPOV’s BMT framework as the “Model 1” approach.
In the last 3 decades, evolution of DNA markers has had huge impact on molecular-assisted plant breeding. Appropriate genetic resources, such as classical bi-parental linkage mapping (Collard et al. 2005), association mapping (AM), multifounder advanced generation inter cross (MAGIC) and nested association mapping (NAM) populations (reviewed by Cockram and Mackay 2018), can be used to identify genetic markers that predict the phenotypic expression of traits controlled by a small number of genes of large effect, as well as by quantitative trait loci (QTL). These approaches, combined with advances in the availability of crop genome reference assemblies (even for large genome polyploid species such as wheat, IWGSC 2018), promote acceleration of the identification of linked or causative polymorphisms underlying phenotypes of interest, including DUS traits. Such markers represent valuable tools for indirect selection of phenotypes through marker-assisted selection (MAS) programmes and also pyramiding desirable genes in a single genotype.
Where available, functional markers (FMs) would be the best choice for MAS and DUS testing under Model 1, as they are derived from the underlying causative polymorphism(s) controlling the effect of a given genetic locus. Therefore, FMs could be highly predictive of trait expression and overcome the problem of using linked markers that are prone to recombination between marker and the target gene or causative polymorphism(s). In addition, linked markers may not always work in different genetic backgrounds (Collard et al. 2005), thus limiting their utility in MAS programmes and DUS testing.
For a review of FMs for agronomic and quality traits, disease resistance and abiotic stress tolerance in wheat and other major grain crops, see Liu et al. (2012) and Kage et al. (2015), respectively. Here, we highlight some breeder-friendly FMs in rice developed for grain length (short versus long), endosperm type (glutinous versus non-glutinous), amylose content of endosperm (different classes), gelatinization temperature (low versus high), and aroma in decorticated grain (fragrant versus non-fragrant), which could be practically utilized for the measurement of DUS characteristic numbers 58 and 62–65 (UPOV 2004) whose measurements are examiner dependent (fragrance) or expensive (amylase content). In addition, FMs aimed at DUS testing for disease resistance to pathogens in tomato (UPOV 2013b) and 13 morphological traits in barley (UPOV 2018) are also presented below.
In rice, Fan et al. (2009) validated a FM for grain length based on a C/A SNP in second exon of the gene, GRAIN SIZE 3 (GS3). The derived cleaved amplified polymorphic sequence (CAPS) marker could predict with 100% accuracy the long (> 9.0 mm) and short grain (< 8.5 mm) genotypes. This marker (which requires digesting the amplicon with the enzyme PstI) was later converted to an allele-specific primer (ASP) assay (Ramkumar et al. 2010). A tightly linked microsatellite to the Waxy gene and a G/T SNP at the putative leader intron 5′ splice site have been used to differentiate amylose classes of rice varieties (Ayres et al. 1997). Moreover, a dCAPS marker was developed for differentiating non-waxy (glutinous) indica and japonica varieties (Yamanaka et al. 2004). CAPS and dCAPS markers were developed in rice for the Starch Synthase IIa gene conferring gelatinization temperature of starch (Lu et al. 2010). Bradbury et al. (2005) developed an allele-specific amplification (ASA) assay for discriminating fragrant and non-fragrant rice varieties across a wide range of samples based on an 8-bp deletion and three SNPs in exon 7 of the Betaine Aldehyde Dehydrogenase 2 (BAD2) gene. This marker was later improved to a single co-dominant FM (Sakthivel et al. 2009). In another complementary study, FMs were developed based on a 7-bp deletion in exon 2 of the gene, as well as previously discovered 8-bp deletion (Shi et al. 2008). New mutations were found based on deletion of 803 bp between exons 4 and 5 of the gene, resulting in development of new FM for detecting fragrant varieties (Shao et al. 2011). Moreover, under a pseudonym (Os2AP) for the same gene, a 3-bp insertion allele in exon 13 was converted to a new FM for the detection of aromatic rice varieties from Myanmar (Myint et al. 2012).
In tomato, Arens et al. (2010) validated existing tightly linked sequence characterized amplified regions (SCAR) markers to a tomato mosaic virus (ToMV) resistance gene (Tm1) and a Fusarium resistance gene (I), in addition to developing robust tetra-primer amplification refractory mutation system (ARMS)-PCR and CAPS markers for identifying resistance to Verticillium (Ve1, Ve2), Fusarium (I2), tomato mosaic virus (Tm2, Tm22), and Meloidogyne incognita nematode (Mi1-2), all of which represent obligatory traits in DUS testing of tomato candidate varieties. High correlation was found between the results of these markers and biological assays, highlighting their potential use as a complement or alternative to bioassay tests.
In barley, PCR-based markers have been introduced for prediction of seasonal growth habit based on insertion/deletion (InDel) polymorphisms at the vernalization requirement locus, VRN-H1 (Cockram et al. 2009). This assay, together with primers targeting candidate ZCCT-H genes at the VRN-H2 vernalization locus (Karsai et al. 2005), can predict winter or spring phenotype. Recent molecular characterization of alternative (also termed “facultative”) seasonal growth habit and flowering time in barley now provided valuable tools for identification of all three growth habits in barley for variety registration purposes (Cockram et al. 2015). Furthermore, in a genome-wide association study (GWAS) of 32 DUS characteristics with 1536 EST-based SNPs in a panel of ~ 500 UK barley cultivars, significant associations were found for 15 traits (Cockram et al. 2010). These positive associations were then validated in 169 UK barley varieties using the KASP genotyping platform, providing a suite of 25 genetic markers for predicting phenotypic expression across 15 DUS traits (Cockram et al. 2012).
With the recent advancements in several genomics platforms, several genes of economic importance have been cloned which somehow needs to be converted into functional-based assays. Even the available functional markers may complement for number of traits in the current scenario with the morphological DUS characterization.
Also, the importance of specific markers could be recognized in green forensic investigations. Cristo-Araújo et al. (2017) showed that how estimation of Reynold’s coancestry coefficient between peach palm landrace accessions, as genotyped by 9 SSR markers, resulted in detection of origin of smuggled seeds of a popular landrace. Considering possible use of DNA markers in differentiating plant landraces dates back to the studies of Konieczny and Ausubel (1993), where ecotype-specific CAPS markers were introduced for distinguishing Arabidopsis thaliana Columbia and Landsberg erecta ecotypes. Likewise, ISSR markers were investigated for possible identification of 11 lentil landraces specific to different regions of Italy, of which a high VCU landrace “Castelluccio” is protected under EU’s “Geographical Indication” (GI) classification (Sonnante and Pignone 2007). In another study, unique SSR markers were identified for evaluation of 104 Perilla frutescens herbaceous landrace accessions collected from different geographical regions of Korea and Japan (Park et al. 2008). Also, whole-genome sequencing of African sorghum (Sorghum bicolor) revealed landrace-specific SNP alleles, suggesting their usefulness in differentiation of modern and landrace materials (Mace et al. 2013). Development of such specific markers suggests that they could be utilized in protection of farming community from biopiracy, smuggling, and any fraudulent marketing of recognized landraces.
Combining phenotypic and molecular distances in the management of variety collections
Developed by experts of the French Group for the Study and Control of Varieties and Seeds (GEVES), reduction in the number of candidate and reference collection varieties before conducting direct comparisons in the field is the key feature of BMT “Model 2” (UPOV 2013a). Its aim is to use only those reference varieties that are appropriately similar to candidate varieties for field evaluation, based on a combination of phenotypic (measured by GAIA software) and molecular (Roger’s) distances. The GAIA distance between varieties is calculated from a combination of phenotypic differences, in which characteristics are weighed according to their variability and susceptibility to the environment. Therefore, the determination of “distinct plus” varieties, which are deemed to be robustly eliminated from field evaluations, is done by plotting distances of molecular against morphological data. All pairs of varieties with a GAIA distance ≥ 6 are classified as “super distinct varieties” and excluded from DUS trial. Moreover, those pairs with a morphological distance > 2 plus a Roger’s distance ≥ 0.2 are also eliminated from field comparisons. In recent years, the use of SSRs has been replaced by a selected set of 312 SNPs for management of the maize reference collection comprising 4000 inbred lines and hybrids (UPOV 2014a). Similarly in potato, decisions on reducing the volume of field comparisons have been based on combination of morphological traits and 9 SSR markers. In that respect, a candidate/reference variety pair with a City block (Manhattan) distance > 0.1 plus a Jaccard distance > 0.2 are considered as “distinct plus” and exempted from second year of DUS testing (UPOV 2011).
In barley, the feasibility of this approach was assessed by using a large set of markers (3072 SNPs) in 431 winter and spring barley varieties phenotyped by 28 DUS traits (Jones et al. 2013). In spite of achieving high levels of correlations between molecular and morphological distances, it was not possible to demonstrate “distinct plus” varieties on a basis of one-to-one correspondence. In a subsequent study, correlation values were further improved by implementing ridge regression among molecular distances, and a reliable value (r > 0.60) was achieved once over 500 SNPs were used in the approach (Jones and Mackay 2015).
In earlier studies, despite detecting low to moderate levels of correlations between DUS traits and DNA markers, application of this approach was envisaged. For instance, seed storage proteins of wheat were used for grouping CKVs prior to field testing. Compared with morphological traits, they showed advantages in terms of both reducing costs and improving the quality of PVP (Jones et al. 2003). Nuel et al. (2001) proposed a statistical approach for predicting phenotypic distances based on molecular data (generated using restriction fragment length polymorphism, RFLP). They applied their model to a panel of 144 maize inbred lines characterized by RFLPs and morphological quantitative traits, resulting in a reduction of field comparisons to only those most similar cultivars. In durum wheat, moderate correlations were found between morphological and molecular marker (AFLPs, SSRs) similarity distances in a set of 69 advanced breeding lines, for which a set of 28 SSRs (one per chromosome arm) was introduced as a pre-screening tool. It was determined that field comparison could be avoided for pairs of varieties, provided that they were polymorphic at more than 13 loci (Noli et al. 2008). In soybean, in spite of detecting weak correlations (0.11–0.24) between similarity coefficient matrices of DUS traits and molecular markers (SSRs, AFLPs, RAPDs), establishing minimum genetic distances was favoured by a combination of morphological and SSR markers (Giancola et al. 2002). Correlation between distance matrices derived from 32 morphological and 28 SSR markers was low (0.2) between 41 maize inbred lines (Gunjaca et al. 2008). Similar results were found in rapeseed (Tommasini et al. 2003), in which no significant correlation was found between SSRs and phenotypes. In durum wheat, despite estimation of high correlation between genetic similarities of AFLPs and SSRs, lower values were obtained between morphological and either of markers (r = 0.46 and 0.56), respectively (Maccaferri et al. 2007a). In pepper, similar low correlations were found between SSRs and morphological traits (Kwon et al. 2005). Besides applying this model for distinctness testing, Wang et al. (2013) developed a procedure for reducing the number of comparator varieties for assessment of stability of wheat varieties before field comparisons. Based on the homozygous SSR loci ratio (SSR-HLR), only those varieties with a ratio between 91 and 95 are entered to the field, while pairs of varieties with a ratio > 95 and < 91 are considered stable and unstable, respectively.
“Essentially derived”: the other side of the PVP coin
Classical and creative breeding of new varieties, for which the UPOV convention was intended to support from the outset, relies on free access of researchers to protected germplasm as material for breeding new varieties. However, this principle, known as “breeder’s exemption” within the PVP system, may be faced with prohibitions in claims for utility patents (Lesser and Mutschler 2004).
In some instances, the developed varieties may conform to existing protected varieties from which they would be classified as “essentially derived”. The “essential derivation” concept was introduced as a process in which most of the essential characteristics of the initial (founder) variety remain intact in the newly bred variety. The so-called essentially derived varieties (EDVs) may be obtained using breeding techniques that result in only a minor or “cosmetic” modification, like mutation, backcrossing, and genetic engineering. The scope of PBR also extends to EDVs, as they are dependent on the founder variety and cannot be exploited without consent of the initial breeder, hence entitling some royalty sharing. Aimed at protecting the rights of the initial breeder, it is stipulated that “cascade of derivations” are possible, rather than the alternative option, “cascade of dependence”. Therefore, where EDV-2 is derived from EDV-1, PBR associated with the initial variety applies to both derived varieties.
By definition, a variety is deemed to be EDV if it is clearly distinct from an initial variety from which is predominantly derived, and conforms to it in the expression of essential characteristics (UPOV 1991). Therefore, distinctness testing and assessing conformity of varieties can be considered as “two sides of the same coin”. The technical concept of essential derivation is parallel to “substantial equivalence” as the backbone of risk assessment of genetically modified organisms (GMOs), which should be substantially equivalent to their non-GMO counterpart except the engineered transgene. Accordingly, EDVs share their essential characteristics with the initial parental variety except for the distinctive trait.
For practical purposes, a boundary of ownership (threshold) has to be defined around the initial variety, where a new variety falls within this boundary, it would be deemed to be essentially derived from it. On the other hand, if exceeded, it would be considered as an independently derived variety. This boundary has to be broad enough not to erode the rights of the initial breeder, but narrow enough to allow the use of protected varieties to perform innovative breeding work. While the UPOV convention has handed the essential derivation dispute over breeders, the Australian PBR Act defines a role for national authority as responsible for defining essential characteristics and administering EDVs (PBRA 1994).
Essential derivation can be assessed through variety origin, breeding methods, heterosis, and phenotypic and/or genotypic characteristics. In spite of technical discrepancies regarding assessment of conformity by phenotypic characteristics or molecular data, genetic distances based on DNA markers have been commonly studied and are strongly supported by ISF (ISF 2012). Heckenberger (2004) suggested a redefinition of the term essential characteristics used in the definition of essential derivation in the UPOV convention, by which marker information can also play the same role as phenotypic traits for essential characteristics. It is now widely accepted that DNA markers can, when used appropriately, provide a largely unbiased estimate of relatedness and similarity between two varieties due to their lack of sensitivity to environmental factors—in contrast with even highly heritable phenotypic traits that often offer only an estimate of the true relatedness. Also, in spite of conventional traits controlled only by a low number of genetic loci, DNA-based approaches can be selected to ensure equal and dense genome coverage. Currently, AFLPs and SSRs represent the most widely used markers in published datasets for defining EDV thresholds on the basis of calculating genetic similarity between varieties.
Thresholds for identifying EDVs
A number of criteria for choosing markers in developing an EDV protocol were discussed by Van Eeuwijk and Law (2004), including marker unbiasedness, precision, and genome coverage. For the latter case, to determine whether markers should be distributed equidistantly across genome or be confined to specific regions controlling differences in characteristics, computer simulations of the distribution of genetic similarities between maize parental inbred lines and their progenies derived from F2 and different backcross populations were undertaken. The results showed that with increasing number and length of chromosome, when using dense and uniform marker distributions, there was a decrease in standard deviations and overlaps of distributions in the studied materials. In that respect, Leigh et al. (2003) suggested that marker system and statistical method should be specified on a crop-by-crop basis for the establishment of EDV thresholds. This recommendation was based on comparing AFLP, SSR, inter-retrotransposon amplified polymorphism (IRAP), and sequence-specific amplified polymorphism (S-SAP) markers in a set of 40 registered barley varieties. Despite discrepancies in distances with each marker between variety groups (UK versus Finnish), similar patterns of diversity (based on Jaccard similarity measures) were observed through combined principal co-ordinate analysis (PCoA) plots.
Van Eeuwijk and Law (2004) defined three approaches for defining thresholds, namely “tail”, “calibration”, and “pedigree” principles, recently reviewed and elaborated on by Noli et al. (2013). So far, the threshold for a number of crops (perennial ryegrass, maize, oilseed rape, cotton, and lettuce) has been adopted by ISF according to these approaches (https://www.worldseed.org/our-work/trade-rules/#essential-derivation, accessed 26 April 2019). Based on tail principle, a percentile point as threshold is chosen in the upper tail end of the distribution of similarities within the reference collection. The composition of the collection is required to be known, as selection of threshold in a collection of no ED pairs is statistically less complicated than calculation on a mixture of non-ED and ED similarity distribution. Application of this approach in lettuce led to the introduction of a threshold of 0.96, based on Jaccard similarity coefficient for three known groups of varieties (ISF 2004). Maccaferri et al. (2007b) compared molecular markers (68 SSRs, 217 AFLPs) with pedigree information for establishing ED in a panel of 134 durum wheat varieties. Although pedigree-based coefficients did not match with the registered parentages of 58 of the accessions, SSRs could identify “breeding lineages” in the collection. In another study, a set of 60 durum wheat genotypes with different levels of relatedness were genotyped by 13 AFLP primer combinations and 109 SSR loci for EDV identification. A threshold of 0.96 Jaccard similarity calculated from AFLPs was proposed based on tail principle (Noli et al. 2012). This principle was also applied for defining EDV thresholds based on Dice similarity in Calluna vulgaris, a popular pot plant in Europe (Borchert et al. 2008).
During implementation of the calibration principle, the threshold is defined on pedigree information or known relationships of closely related genotypes. Van Eeuwijk and Law (2004) adopted this approach for barley and decided to place the threshold at the level of second backcross. Recently, a high correlation between coefficient of relatedness and genetic (3072 SNPs) or morphological (28 DUS traits) distances were observed, using 431 barley varieties grouped according to their pedigree relationships, suggesting both or either type of data could be suitable for essential derivation issues (Jones et al. 2013). Using this approach, genetic similarities among 83 rose varieties were studied using AFLP markers for establishing essential derivation, where mutants (also known as “sport”) as putative EDVs were distinguished at a threshold of 0.95 from original varieties (Vosman et al. 2004). Earlier studies on the ability of morphological traits and seed storage proteins (Gilliland et al. 2000) and AFLP markers (Roldán-Ruiz et al. 2000) for estimating the genetic conformity between related ryegrass (Lolium perenne L.) varieties revealed congruent results via multivariate clustering methods. A set of 107 SSR and 48 SCAR markers were used in cucumber, representing a species with narrow genetic base, to define threshold values for identification of EDVs. Results based on genetic distances showed some levels of polymorphisms in three sets of backcross families, restraining the establishment of thresholds, while identifying SNPs as better discriminators (Staub et al. 2005).
In the pedigree principle, a threshold is defined in such a way that observed genetic similarities are related to identity by descent (IBD) probabilities. Therefore, utilization of genome-wide markers with known map position would help implement this approach. Using this method, morphological traits, heterosis, and molecular markers (AFLPs, SSRs) have been used for identification of EDVs in maize inbred lines (Heckenberger et al. 2002, 2003, 2005a, b). Based on comparison of these various data for estimating genetic distances, a complementary application of AFLPs and SSRs was recommended for uncovering close pedigree relationships and distinguishing EDVs from IDVs (Heckenberger et al. 2006). These studies depicted three zones in maize based on already defined boundaries or thresholds of genetic conformity between varieties. Similar to traffic lights, the zones of “non-distinctness or indisputable essential derivation” (red), “uncertainty” (orange), and “non-derivation or independence” (green), define boundaries between varieties. It is assumed that once putative EDVs fall into the orange zone, a reversal of burden of proof occurs and their breeders should prove the independency of them for the initial variety. In recent years, ISF updated SSR-based zones (namely < 82% similarity (green), 82–90% (orange), and > 90 (red)) with SNP-based EDV thresholds. Therefore, genetic similarity < 91% and ≥ 95% between the initial variety and EDV is considered as non-derivation (green) and definitely derivation (red), respectively. Accordingly, putative EDVs with genetic similarity ≥ 91 to < 95% reside in orange zone (ISF 2014; UPOV 2014b). The list of 3072 SNP markers used for the recent guideline in maize is available on the ISF website (https://www.worldseed.org/wp-content/uploads/2015/10/Annex_2_SNP_Markers_EDV_Maize_2014.pdf, accessed 26 April 2019).
In another study, Kahler et al. (2010) introduced a set of 285 SSRs for utilization in variety (including EDV) identification in US maize. The markers constitute 150 SSRs already validated by the American Seed Trade Association (ASTA) and 135 loci, selected totally according to their informativeness, allele size, fragment expression, and ease of scoring on gel-based systems. Comparing the discriminatory power of these markers among 30 US inbred lines with other results of loci recommended by the French Association of Maize Breeders (SEPROMA) showed high congruence, suggesting the suitability of both ASTA and SEPROMA protocols for determining essential derivation in maize.
In cases where molecular thresholds are not defined, morphological differences may be used for resolving varietal disputes. This was illustrated by litigation of Danziger, the initial breeder, against the gypsophila (commonly known as Baby’s Breath) variety “Blancanieves”, bred by Astee Flowers. The two breeders independently conducted genetic similarity assessments, with Danziger finding the founder variety “Million Stars” conforms 97.7% (similarity for 209 out of 214 markers tested) to the putative EDV “Blancanieves”. However, the second breeder identified lower relatedness (91%) via analysis of 133 markers. The court declined to use the genotypic data and referred to the phenotypic differences (evidenced by second breeder) between two disputed varieties to reject Danziger claim of EDV ownership. In that case, distinguishing morphological traits such as plant height, branching, length of the flower stem, and flower diameter were sufficient to prove that “Blancanieves” was not essentially derived from “Million Stars”. Thus, the court dismissed genotypic tests owing to complexities in comparing quantitative values and even differences in number of markers tested by breeders (Janis and Smith 2007).
Future prospects
Challenges of new breeding technologies for DUS testing
DUS testing is prerequisite for authenticating a unique “identity” before protection and registration of new varieties of plants. Moreover, agricultural crops are required to have a specific “utility” or VCU adequacy. A recent study on the socio-economic impact of plant breeding (period 2001–2015) shows that a good part (75%) of overall productivity growth in the European Union is due to improved cultivars for nine major crops, with an annual 1.24% yield increase (Noleppa 2016). This significant contribution could be higher if cultivars with sufficient VCU were not eliminated by DUS testing. In France, 26% of alfalfa (Medicago sativa L. ssp. sativa) varieties were rejected during 2001–2010 due to lack of distinctness from CKVs. Moreover, the elimination rate was much higher (21%) compared to those (3%) with insufficient VCU during a 2-year period (Annicchiarico et al. 2016). Similar figures were recorded for Lolium spp. and white clover (Trifolium repens L.), where just over one-tenth of cultivars did not satisfy DUS criteria during 2000–2008, peaking at 19% (Gilliland and Gensollen 2010).
Recent demands for sustainability in agriculture has also incentivised breeders to develop vegetable varieties that uptake and use nutrients more efficiently while producing high biomass (Ferrante et al. 2017). While disease resistance has been included in vegetable DUS test guidelines, inclusion of agronomic traits is not recommended due to significant interaction with environment, and hence complicating measures of distinctness. Therefore, a spinach variety that utilizes nitrogen more efficiently than check varieties, while indistinguishable for other characters, would be a challenge for variety protection and registration.
Agronomic traits such as number of seeds per spikelet and thousand-seed weight in ryegrass (Wang et al. 2016), and sugar content, yield, Na, and K content in sugar beet roots (https://www.naktuinbouw.com/agriculture/variety-description/sugar-beet-beta-vulgaris-l, accessed 26 April 2019) have been included in variety descriptions. These are examples of compensating for the low potential of morphological traits in distinguishing cultivars with sufficient VCU. As a solution, a marker-based registration has been suggested as a way to provide a connection between VCU and DUS traits (Cooke and Reeves 2003).
Recently, plant breeders have been enabled to precisely manipulate gene sequences in order to create new alleles, or remove undesirable ones, using programmable nucleases. This technology, known as gene/genome-editing, may revolutionize plant breeding by allowing development of plants with enhanced or novel traits, e.g. for combating present biotic and abiotic stresses, as well as making herbicide-resistant crops (Nogué et al. 2016). Among the various tools available, clustered regularly interspaced short palindromic repeats (CRISPR) is currently particularly favoured, owing to its high efficiency and simplicity. Despite the feasibility of engineering specific DUS phenotype combinations to satisfy DUS testing purposes (Liu and Moschou 2018), and so complicating current EDV regulations, agronomically important traits will be the predominant target of CRISPR-assisted plant breeding.
Conventional breeding that is implicitly considered “creative” by the UPOV convention generally relies on genetic recombination. This process allows for selection of promising varieties via creation of new allelic and phenotypic combinations. The generation of such novel combinations is also enhanced in multi-parent advanced generation recurrent selection (MAGReS), where the genetic backgrounds of multiple parents are thoroughly shuffled via intercrossing (Huang et al. 2015). On the other hand, genotypic distances between varieties are impaired by genome-editing, in which modification can occur at the level of a few nucleotides. For instance, an elite maize cultivar could be modified to be tolerant to glyphosate by editing only two nucleotides of the EPSPS gene using CRISPR technology. If the founder variety would be considered as the line protected by PBR, the new CRISPR-plant would be subject to an EDV claim as it wholly conforms and retains the essential characteristics of it, with the exception of a 2-bp difference. However, this minor modification that results in adding value for cultivation (herbicide tolerance) to the initial variety could also be considered as a distinctive trait in DUS testing. In that respect, satisfying distinctness with specific markers targeting SNPs would result in an unequivocal decision of herbicide tolerance phenotype, analogous to phenotype derived by spraying in the field (UPOV 2013a).
A redefinition of essential derivation concept is inevitable to fortify the “breeder’s exemption” principle within the UPOV convention. In this regard, the Australian PVP Act, in which a newly bred variety from the initial variety can be declared EDV unless it adds a VCU trait as an essential characteristic, could be paradigmatic—since, it is the responsibility of the putative EDV breeder to explain the VCU of the distinctive trait (e.g. a shorter internode length in turf grass could be defined as increasing thatch and hence resistance to wear).
Shifting from phenotype- to DNA-based variety description: advantages and drawbacks
Variety descriptions as the outcome of DUS testing have multiple functions. They provide a unique “identity” to new varieties as an essential requirement of PVP, as well as variety registration. In addition, they can be utilized by variety maintainers to ensure variety purity and by field inspectors during the seed certification process (van Wijk and Louwaars 2014). It is now more than half a century since the PVP system was introduced via the UPOV convention. So far, a huge number of varieties have been developed and released for many uses, including food, feed, and fibre. Within this timeframe, distinguishing candidate varieties from large number of CKVs with narrow genetic base has become increasingly challenging for DUS testing. Towards reducing DUS testing costs, limiting the number of varieties required for side-by-side comparison in the field is highly favoured by the BMT (Model 2) and IMODDUS working groups.
With the rapidly reducing costs of next-generation sequencing (NGS) and genotyping-by-sequencing (GBS) platforms, plant breeders are shifting from conventional to next-generation breeding approaches (Barabaschi et al. 2016). The increasing availability and cost-efficient access to SNP markers and sequencing data will facilitate dissection of marker-trait associations in many crops (Liu et al. 2012). Such tools would also be used in genomic selection, in which breeding values of individuals are accurately predicted by their genotypes over multiple generations in the absence of phenotyping (Lin et al. 2014). Indeed, the timeframe between sequential crossing cycles remains an inherent major constraint for increasing the rate of genetic improvement. Therefore, combining genomic selection with approaches to speed up breeding cycles, such as “speed breeding” (Watson et al. 2018; Voss-Fels et al. 2019) or the potential of “in vitro cycling”, where generations are advanced via marker-assisted selection on gametes without progressing individuals to adult plants, (De la Fuente et al. 2013), would be highly favourable. Such advances mean it could be possible that varieties will be released at a faster rate than before (Li et al. 2018). This supports the case for a considerable reduction in duration of DUS testing, as it is conceivable that releasing a crop cultivar would take much less than the current 10–15 years. Against a background of increased use of emerging technologies such as GBS, GS, accelerated generation cycling approaches and phenomics as the integrated hallmarks of “next-generation breeding”, renewed focus on detailed investigation of molecular-based or molecular-assisted DUS testing processes could be timely.
Centrally maintained databases of DNA sequences of CKVs have been constructed for the determination of distinctness in certain species like potato, tomato, rose, Phalaenopsis, and Gypsophila (Kees van Ettekoven, personal communication). It is conceivable that SSR databases for soybean (Oliveira et al. 2010), wheat (Röder et al. 2002; Jaiswal et al. 2017), tomato (Bredemeijer et al. 2002; Iquebal et al. 2013), rose (Smulders et al. 2009), potato (Reid et al. 2011), and lettuce (Hong et al. 2015) will be switched to SNPs due to their high abundance and genome coverage. While these databases could be utilized during infringement of PBRs and EDV disputes, some challenges remain in addressing uniformity and stability of plant varieties. Increasing numbers of aberrant individuals in a submitted sample for uniformity testing is of utmost concern. In that case, considering a new definition of “plant variety” with reference to characteristics of genomic tools in “next-generation breeding” is a timely topic for all those involved with the development, evaluation, and release of new crop varieties.
Author contribution statement
SHJ designed the outline and wrote the manuscript. JC and LTH revised the manuscript. All authors approved the final version.
Abbreviations
- CKV:
-
Common knowledge varieties
- DUS:
-
Distinctness, uniformity, and stability
- EDV:
-
Essentially derived variety
- IPR:
-
Intellectual property rights
- PBR:
-
Plant breeder’s rights
- PVP:
-
Plant variety protection
- SNP:
-
Single nucleotide polymorphism
- SSR:
-
Simple sequence repeat
- UPOV:
-
International Union for the Protection of New Varieties of Plants
References
Annicchiarico P, Nazzicari N, Ananta A, Carelli M, Wei Y, Brummer EC (2016) Assessment of cultivar distinctness in alfalfa: a comparison of genotyping-by-sequencing, simple-sequence repeat marker, and morphophysiological observations. Plant Genome 9(2):1–12
Arens P, Mansilla C, Deinum D, Cavellini L, Moretti A, Rolland S, van der Schoot H, Calvache D, Ponz F, Collonnier C, Mathis R, Smilde D, Caranta C, Vosman B (2010) Development and evaluation of robust molecular markers linked to disease resistance in tomato for distinctness, uniformity and stability testing. Theor Appl Genet 120:655–664
Ayres NM, McClung AM, Larkin PD, Bligh HFJ, Jones CA, Park WD (1997) Microsatellites and a single-nucleotide polymorphism differentiate apparent amylose classes in an extended pedigree of US rice germ plasm. Theor Appl Genet 94:773–781
Barabaschi D, Tondelli A, Desiderio F, Volante A, Vaccino P, Valè G, Cattivelli L (2016) Next generation breeding. Plant Sci 242:3–13
Bernet GP, Bramardi S, Calvache D, Carbonell EA, Asins MJ (2003) Applicability of molecular markers in the context of protection of new varieties of cucumber. Plant Breed 122:146–152
Bonow S, Von Pinho EVR, Vieira MGC, Vosman B (2009) Microsatellite markers in and around rice genes: applications in variety identification and DUS testing. Crop Sci 49:880–886
Borchert T, Krueger J, Hohe A (2008) Implementation of a model for identifying essentially derived varieties in vegetatively propagated Calluna vulgaris varieties. BMC Genet 9:1
Bradbury LMT, Henry RJ, Jin Q, Reinke RF, Waters DLE (2005) A perfect marker for fragrance genotyping in rice. Mol Breed 16:279–283
Bredemeijer G, Cooke R, Ganal M, Peeters R, Isaac P, Noordijk Y, Rendell S, Jackson J, Röder M, Wendehake K (2002) Construction and testing of a microsatellite database containing more than 500 tomato varieties. Theor Appl Genet 105:1019–1026
Chen Y, Dai X, Hou J, Guan H, Wang Y, Li Y, Yin T (2016) DNA fingerprinting of oil camellia cultivars with SSR markers. Tree Genet Genomes 12:7
Choi S, Sim S, Hong J, Choi K, Jin M, Park B, Kim D, Kwon Y (2016) Genetic characterisation of commercial Chinese cabbage varieties using SSR markers. Seed Sci Technol 44:595–608
Cockram J, Mackay I (2018) Genetic mapping populations for conducting high-resolution trait mapping in plants. In: Blakebrough N, Ghose TK, Fiechter A (eds) Advances in biochemical engineering/biotechnology. Springer, Berlin
Cockram J, Norris C, O’Sullivan DM (2009) PCR-based markers diagnostic for spring and winter seasonal growth habit in barley. Crop Sci 49:403–410
Cockram J, White J, Zuluaga DL, Smith D, Comadran J, Macaulay M, Luo Z, Kearsey MJ, Werner P, Harrap D et al (2010) Genome-wide association mapping to candidate polymorphism resolution in the unsequenced barley genome. Proc Nat Acad Sci USA 107:21611–21616
Cockram J, Jones H, Norris C, O’Sullivan DM (2012) Evaluation of diagnostic molecular markers for DUS phenotypic assessment in the cereal crop, barley (Hordeum vulgare ssp. vulgare L.). Theor Appl Genet 125:1735–1749
Cockram J, Horsnell R, E-h Soh, Norris C, O’Sullivan DM (2015) Molecular and phenotypic characterization of the alternative seasonal growth habit and flowering time in barley (Hordeum vulgare ssp. vulgare L.). Mol Breed 35:1–11
Collard B, Jahufer M, Brouwer J, Pang E (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196
Cooke RJ, Reeves JC (2003) Plant genetic resources and molecular markers: variety registration in a new era. Plant Genet Resour 1:81–87
Cooke RJ, Bredemeijer GMM, Ganal MW, Peeters R, Isaac P, Rendell S, Jackson J, Roder MS, Korzun V, Wendehake K, Areshchenkova T, Dijcks M, Laborie D, Bertrand L, Vosman B (2003) Assessment of the uniformity of wheat and tomato varieties at DNA microsatellite loci. Euphytica 132:331–341
Cristo-Araújo M, Molles DB, Rodrigues DP, Clement CR (2017) Genetic analysis identifies the region of origin of smuggled peach palm seeds. Forensic Sci Int 273:e15–e17
Curtis F, Nilsson M (2012) Collection systems for royalties in wheat: an international study. Bio-Sci Law Rev 12:215
De la Fuente G, Frei UK, Lübberstedt T (2013) Accelerating plant breeding. Trends Plant Sci 18:667–672
De Riek J, Calsyn E, Everaert I, Van Bockstaele E, De Loose M (2001) AFLP based alternatives for the assessment of distinctness, uniformity and stability of sugar beet varieties. Theor Appl Genet 103:1254–1265
Delfini J, Moda-Cirino V, de Fátima Ruas C, dos Santos Neto J, Ruas PM, Buratto JS, Ruas EA, Gonçalves LSA (2017) Distinctness of Brazilian common bean cultivars with carioca and black grain by means of morphoagronomic and molecular descriptors. PLoS ONE 12:e0188798
Fan C, Yu S, Wang C, Xing Y (2009) A causal C-A mutation in the second exon of GS3 highly associated with rice grain length and validated as a functional marker. Theor Appl Genet 118:465–472
Ferrante A, Nocito FF, Morgutti S, Sacchi GA (2017) Plant breeding for improving nutrient uptake and utilization efficiency. In: Tei F, Nicola S, Benincasa P (eds) Advances in research on fertilization management of vegetable crops. Springer, Berlin, pp 221–246
Fossati T, Zapelli I, Bisoffi S, Micheletti A, Vietto L, Sala F, Castiglione S (2005) Genetic relationships and clonal identity in a collection of commercially relevant poplar cultivars assessed by AFLP and SSR. Tree Genet Genomes 1:11–20
Gao L, Jia J, Kong X (2016) A SNP-based molecular barcode for characterization of common wheat. PLoS ONE 11(3):e0150947
Giancola S, Poltri SM, Lacaze P, Hopp HE (2002) Feasibility of integration of molecular markers and morphological descriptors in a real case study of a plant variety protection system for soybean. Euphytica 127:95–113
Gilliland TJ, Gensollen V (2010) Review of the protocols used for assessment of DUS and VCU in Europe—perspectives. In: Huyghe C (ed) Sustainable use of genetic diversity in forage and turf breeding. Springer, Berlin, pp 261–275
Gilliland T, Coll R, Calsyn E, De Loose M, Van Eijk M, Roldán-Ruiz I (2000) Estimating genetic conformity between related ryegrass (Lolium) varieties. 1. Morphology and biochemical characterisation. Mol Breed 6:569–580
Gunjaca J, Buhinicek I, Jukic M, Sarcevic H, Vragolovic A, Kozic Z, Jambrovic A, Pejic I (2008) Discriminating maize inbred lines using molecular and DUS data. Euphytica 161:165–172
Hall BD, Fox R, Zhang Q, Baumgarten A, Nelson B, Cummings J, Drake B, Phillips D, Hayes K, Beatty M (2016) Comparison of genotypic and expression data to determine distinctness among inbred lines of maize for granting of plant variety protection. Crop Sci 56:1443–1459
Heckenberger M (2004) Identification of essentially derived varieties in maize (Zea mays L.) using molecular markers, morphological traits, and heterosis. Ph.D. thesis, University of Hohenheim
Heckenberger M, Bohn M, Ziegle JS, Joe LK, Hauser JD, Hutton M, Melchinger AE (2002) Variation of DNA fingerprints among accessions within maize inbred lines and implications for identification of essentially derived varieties. Mol Breed 10:181–191
Heckenberger M, van der Voort JR, Peleman J, Bohn M (2003) Variation of DNA fingerprints among accessions within maize inbred lines and implications for identification of essentially derived varieties: II. Genetic and technical sources of variation in AFLP data and comparison with SSR data. Mol Breed 12:97–106
Heckenberger M, Bohn M, Melchinger A (2005a) Identification of essentially derived varieties obtained from biparental crosses of homozygous lines. Crop Sci 45:1120–1131
Heckenberger M, Bohn M, Frisch M, Maurer H, Melchinger A (2005b) Identification of essentially derived varieties with molecular markers: an approach based on statistical test theory and computer simulations. Theor Appl Genet 111:598–608
Heckenberger M, Muminović J, van der Voort JR, Peleman J, Bohn M, Melchinger A (2006) Identification of essentially derived varieties obtained from biparental crosses of homozygous lines. III. AFLP data from maize inbreds and comparison with SSR data. Mol Breed 17:111–125
Hong J-H, Kwon Y-S, Mishra RK, Kim DH (2015) Construction of EST-SSR databases for effective cultivar identification and their applicability to complement for lettuce (Lactuca sativa L.) distinctness test. Am J Plant Sci 6:113
Huang BE, Verbyla KL, Verbyla AP, Raghavan C, Singh VK, Gaur P, Leung H, Varshney RK, Cavanagh CR (2015) MAGIC populations in crops: current status and future prospects. Theor Appl Genet 128:999–1017
Ibàñez J, Vélez MD, de Andrés MT, Borrego J (2009) Molecular markers for establishing distinctness in vegetatively propagated crops: a case study in grapevine. Theor Appl Genet 119:1213–1222
Iquebal MA, Arora V, Verma N, Rai A, Kumar D (2013) First whole genome based microsatellite DNA marker database of tomato for mapping and variety identification. BMC Plant Biol 13:197
ISF, International Seed Federation (2004) Guidelines for the handling of a dispute on essential derivation in lettuce. https://www.worldseed.org/wp-content/uploads/2015/10/Guidelines_EDV_Lettuce_2004.pdf. Accessed 26 Apr 2019
ISF, International Seed Federation (2012) ISF view on intellectual property. https://www.worldseed.org/wp-content/uploads/2015/10/View_on_Intellectual_Property_2012.pdf. Accessed 26 Apr 2019
ISF, International Seed Federation (2014) Guidelines for the handling of a dispute on essential derivation in maize lines. https://www.worldseed.org/wp-content/uploads/2015/10/ISF_Guidelines_Disputes_EDV_Maize_2014.pdf. Accessed 26 Apr 2019
IWGSC, International Wheat Genome Sequencing Consortium (2018) Shifting the limits of wheat research and breeding using a fully annotated reference genome. Science 361:eear7191
Jaiswal S, Sheoran S, Arora V, Angadi UB, Iquebal MA, Raghav N, Aneja B, Kumar D, Singh R, Sharma P (2017) Putative microsatellite DNA marker-based wheat genomic resource for varietal improvement and management. Front Plant Sci 8:2009
Jamali S, Sadeghi L, Sadeghin-Motahhar S (2011) Identification and distinction of soybean commercial cultivars using morphological and microsatellite markers. Iran J Crop Sci 13:131–145
Jamali SH, Mohammadi SA, Sadeghzadeh B (2017) Association mapping for morphological traits relevant to registration of barley varieties. Span J Agric Res 15:0704
Janis MD, Smith S (2007) Technological change and the design of plant variety protection regimes. Chicago-Kent Law Rev 82(3):1557–1615
Jones H, Mackay I (2015) Implications of using genomic prediction within a high-density SNP dataset to predict DUS traits in barley. Theor Appl Genet 128:2461–2470
Jones H, Jarman R, Austin L, White J, Cooke R (2003) The management of variety reference collections in distinctness, uniformity and stability testing of wheat. Euphytica 132:175–184
Jones H, Norris C, Smith D, Cockram J, Lee D, O’Sullivan DM, Mackay I (2013) Evaluation of the use of high-density SNP genotyping to implement UPOV Model 2 for DUS testing in barley. Theor Appl Genet 126:901–911
Kage U, Kumar A, Dhokane D, Karre S, Kushalappa AC (2015) Functional molecular markers for crop improvement. Crit Rev Biotechnol 16:1–14
Kahler AL, Kahler JL, Thompson SA, Ferriss RS, Jones ES, Nelson BK, Mikel MA, Smith S (2010) North American study on essential derivation in maize: II. Selection and evaluation of a panel of simple sequence repeat loci. Crop Sci 50:486–503
Karsai I, Szűcs P, Mészáros K, Filichkina T, Hayes P, Skinner J, Láng L, Bedő Z (2005) The Vrn-H2 locus is a major determinant of flowering time in a facultative × winter growth habit barley (Hordeum vulgare L.) mapping population. Theor Appl Genet 110:1458–1466
Konieczny A, Ausubel FM (1993) A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant J 4(2):403–410
Kumar V, Rani A, Jha P, Rawal R, Husain S (2017) Molecular identification of dominant cultivars of soybean using simple sequence repeat markers. Proc Natl Acad Sci India Sect B Biol Sci 87:647–653
Kwon YS (2016) DNA fingerprinting analysis for soybean (Glycine max) varieties in Korea using a core set of microsatellite marker. J Plant Biotechnol 43:457–465
Kwon YS, Lee JM, Yi GB, Yi SI, Kim KM, Soh EH, Bae KM, Park EK, Song IH, Kim BD (2005) Use of SSR markers to complement tests of distinctiveness, uniformity, and stability (DUS) of pepper (Capsicum annuum L.) varieties. Mol Cells 19:428–435
Law JR, Donini P, Koebner RMD, James CR, Cooke RJ (1998) DNA profiling and plant variety registration. III: the statistical assessment of distinctness in wheat using amplified fragment length polymorphisms. Euphytica 102:335–342
Leigh F, Law JR, Lea VJ, Donini P, Reeves JC (2003) A comparison of molecular markers and statistical tools for diversity and EDV assessments. In: The wake of the double helix: from the green revolution to the gene revolution: proceedings of an international congress held in Bologna, May 27–31, pp 349–363
Lesser W, Mutschler M (2004) Balancing investment incentives and social benefits when protecting plant varieties. Crop Sci 44:1113–1120
Li H, Rasheed A, Hickey LT, He Z (2018) Fast-forwarding genetic gain. Trends Plant Sci 23:184–186
Lin Z, Hayes B, Daetwyler H (2014) Genomic selection in crops, trees and forages: a review. Crop Pasture Sci 65:1177–1191
Liu C, Moschou PN (2018) Phenotypic novelty by CRISPR in plants. Dev Biol 425:170–175
Liu Y, He Z, Appels R, Xia X (2012) Functional markers in wheat: current status and future prospects. Theor Appl Genet 125:1–10
Lombard V, Baril CP, Dubreuil P, Blouet F, Zhang D (2000) Genetic relationships and fingerprinting of rapeseed cultivars by AFLP: consequences for varietal registration. Crop Sci 40:1417–1425
Lombard V, Tireau B, Blouet F, Zhang D, Baril CP (2002) Usefulness of AFLP markers to estimate varietal homogeneity of rapeseed inbred line varieties in the context of plant registration and protection. Euphytica 125:121–127
Lombardo L (2014) Genetic use restriction technologies: a review. Plant Biotechnol J 12:995–1005
Lu Y, Xiao P, Shao Y, Zhang G, Thanyasiriwat T, Bao J (2010) Development of new markers to genotype the functional SNPs of SSIIa, a gene responsible for gelatinization temperature of rice starch. J Cereal Sci 52:438–443
Maccaferri M, Stefanelli S, Rotondo F, Tuberosa R, Sanguineti MC (2007a) Relationships among durum wheat accessions. I. Comparative analysis of SSR, AFLP, and phenotypic data. Genome 50:373–384
Maccaferri M, Sanguineti MC, Xie C, Smith JSC, Tuberosa R (2007b) Relationships among durum wheat accessions. II. A comparison of molecular and pedigree information. Genome 50:385–399
Mace ES, Tai S, Gilding EK, Li Y, Prentis PJ, Bian L, Campbell BC, Hu W, Innes DJ, Han X, Cruickshank A (2013) Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat Commun 4:2320
Mall N, Chawla H (2014) Suitability of SSR marker for establishing distinctiveness in French bean varieties (Phaseolus vulgaris L.). Legume Res Int J 37:353–358
Myint KM, Arikit S, Wanchana S, Yoshihashi T, Choowongkomon K, Vanavichit A (2012) A PCR-based marker for a locus conferring the aroma in Myanmar rice (Oryza sativa L.). Theor Appl Genet 125(5):887–896
Nogué F, Mara K, Collonnier C, Casacuberta JM (2016) Genome engineering and plant breeding: impact on trait discovery and development. Plant Cell Rep 35(7):1475–1486
Noleppa S (2016) The economic, social and environmental value of plant breeding in the European Union, an ex post evaluation and ex ante assessment. HFFA Research paper 03/2016. https://www.agrarzeitung.de/news/media/3/EU-Studie_zum_Nutzen_der_Pflanzenzu__chtung.pdf-22944.pdf. Accessed 26 Apr 2019
Noli E, Teriaca MS, Sanguineti MC, Conti S (2008) Utilization of SSR and AFLP markers for the assessment of distinctness in durum wheat. Mol Breed 22:301–313
Noli E, Teriaca MS, Conti S (2012) Identification of a threshold level to assess essential derivation in durum wheat. Mol Breed 29:687–698
Noli E, Teriaca MS, Conti S (2013) Criteria for the definition of similarity thresholds for identifying essentially derived varieties. Plant Breed 132:525–531
Nováková A, Šimáčková K, Bárta J, Čurn V (2009) Potato variety identification by molecular markers based on retrotransposon analyses. Czech J Genet Plant Breed 45:1–10
Nuel G, Baril C, Robin S (2001) Varietal distinctness assisted by molecular markers: a methodological approach. In: International symposium on molecular markers for characterizing genotypes and identifying cultivars in horticulture, vol 546, pp 65–71
Oliveira MB, Vieira ESN, Schuster I (2010) Construction of a molecular database for soybean cultivar identification in Brazil. Genet Mol Res 9:705–720
Park YJ, Dixit A, Ma KH, Lee JK, Lee MH, Chung CS, Nitta M, Okuno K, Kim TS, Cho EG, Rao VR (2008) Evaluation of genetic diversity and relationships within an on-farm collection of Perilla frutescens (L.) Britt. using microsatellite markers. Genet Resour Crop Evol 55(4):523–535
Patra N, Chawla HS (2010) Biochemical and RAPD molecular markers for establishing distinctiveness of basmati rice (Oryza sativa L.) varieties as additional descriptors for plant variety protection. Indian J Biotechnol 9:371–377
Phan NT, Kim M-K, Sim S-C (2016) Genetic variations of F1 tomato cultivars revealed by a core set of SSR and InDel markers. Sci Hortic 212:155–161
PBRA, Plant Breeder’s Rights Act (1994) Australian Government, No. 110. https://www.legislation.gov.au/Details/C2018C00361. Accessed 26 Apr 2019
Pourabed E, Jazayeri Noushabadi MR, Jamali SH, Moheb Alipour N, Zareyan A, Sadeghi L (2015) Identification and DUS testing of rice varieties through microsatellite markers. Int J Plant Genom. Article ID 965073
Ramkumar G, Sivaranjani AKP, Pandey MK, Sakthivel K, Shobha Rani N, Sudarshan I, Prasad GSV, Neeraja CN, Sundaram RM, Viraktamath BC (2010) Development of a PCR-based SNP marker system for effective selection of kernel length and kernel elongation in rice. Mol Breed 26(4):735–740
Rani A, Kumar V, Shukla S, Jha P, Rawal R (2016) DNA barcoding of Indian soybean varieties as constructed through SSR markers. Seed Sci Technol 44:357–369
Reid A, Kerr EM (2007) A rapid simple sequence repeat (SSR)-based identification method for potato cultivars. Plant Genet Resour 5:7–13
Reid A, Hof L, Felix G, Rücker B, Tams S, Milczynska E, Esselink D, Uenk G, Vosman B, Weitz A (2011) Construction of an integrated microsatellite and key morphological characteristic database of potato varieties on the EU common catalogue. Euphytica 182:239–249
Röder M, Wendehake K, Korzun V, Bredemeijer G, Laborie D, Bertrand L, Isaac P, Rendell S, Jackson J, Cooke R (2002) Construction and analysis of a microsatellite-based database of European wheat varieties. Theor Appl Genet 106:67–73
Roldán-Ruiz I, Calsyn E, Gilliland T, Coll R, Van Eijk M, De Loose M (2000) Estimating genetic conformity between related ryegrass (Lolium) varieties. 2. AFLP characterization. Mol Breed 6:593–602
Rotondi A, Magli M, Ricciolini C, Baldoni L (2003) Morphological and molecular analyses for the characterization of a group of Italian olive cultivars. Euphytica 132:129–137
Sakthivel K, Rani NS, Pandey MK, Sivaranjani A, Neeraja C, Balachandran S, Madhav MS, Viraktamath B, Prasad G, Sundaram R (2009) Development of a simple functional marker for fragrance in rice and its validation in Indian Basmati and non-Basmati fragrant rice varieties. Mol Breed 24:185–190
Sarao NK, Vikal Y, Singh K, Joshi MA, Sharma RC (2009) SSR marker-based DNA fingerprinting and cultivar identification of rice (Oryza sativa L.) in Punjab state of India. Plant Genet Resour Char Util 1:3
Shao G, Tang A, Tang S, Luo J, Jiao G, Wu J, Hu P (2011) A new deletion mutation of fragrant gene and the development of three molecular markers for fragrance in rice. Plant Breed 130:172–176
Shi W, Yang Y, Chen S, Xu M (2008) Discovery of a new fragrance allele and the development of functional markers for the breeding of fragrant rice varieties. Mol Breed 22:185–192
Shukla S, Joshi D, Srivastava R, Qureshi M, Singh U (2011) Suitability of RAPD and ISSR to complement agro-morphological DUS descriptors for establishing distinctiveness in indigenous local strains of Kalanamak rice (Oryza sativa). Indian J Agric Sci 81:994
Singh RK, Sharma RK, Singh AK, Singh VP, Singh NK, Tiwari SP, Mohapatra T (2004) Suitability of mapped sequence tagged microsatellite site markers for establishing distinctness, uniformity and stability in aromatic rice. Euphytica 135:135–143
Smulders MJM, Esselink D, Voorrips RE, Vosman B (2009) Analysis of a database of DNA profiles of 734 Hybrid Tea Rose varieties. In: 23rd International Eucarpia symposium, section ornamentals: colourful breeding and genetics, vol 836, pp 169–175
Smỳkal P, Horacek J, Dostalova R, Hybl M (2008) Variety discrimination in pea (Pisum sativum L.) by molecular, biochemical and morphological markers. J Appl Genet 49:155–166
Sonnante G, Pignone D (2007) The major Italian landraces of lentil (Lens culinaris Medik.): their molecular diversity and possible origin. Genet Res Crop Evol 54(5):1023–1031
Sousa TV, Caixeta ET, Alkimim ER, de Oliveira ACB, Pereira AA, Zambolim L, Sakiyama NS (2017) Molecular markers useful to discriminate Coffea arabica cultivars with high genetic similarity. Euphytica 213:75
Staub J, Chung SM, Fazio G (2005) Conformity and genetic relatedness estimation in crop species having a narrow genetic base: the case of cucumber (Cucumis sativus L.). Plant Breed 124:44–53
Tian H-L, Wang F-G, Zhao J-R, Yi H-M, Wang L, Wang R, Yang Y, Song W (2015) Development of maizeSNP3072, a high-throughput compatible SNP array, for DNA fingerprinting identification of Chinese maize varieties. Mol Breeding 35:136
Tiwari JK, Ali N, Devi S, Kumar V, Zinta R, Chakrabarti SK (2018) Development of microsatellite markers set for identification of Indian potato varieties. Sci Hortic 231:22–30
Tommasini L, Batley J, Arnold GM, Cooke RJ, Donini P, Lee D, Law JR, Lowe C, Moule C, Trick M, Edwards KJ (2003) The development of multiplex simple sequence repeat (SSR) markers to complement distinctness, uniformity and stability testing of rape (Brassica napus L.) varieties. Theor Appl Genet 106:1091–1101
UPOV, International Union for the Protection of New Varieties of Plants (1991) International convention for the protection of new varieties of plants, Publication No. 221 (E), March 19, Geneva
UPOV, International Union for the Protection of New Varieties of Plants (2004) Guidelines for the conduct of tests for distinctness, uniformity and stability, rice (Oryza sativa L.). TG/16/8. https://www.upov.int/edocs/tgdocs/en/tg016.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2011) Combining morphological and molecular distance in the management of the reference collection of potato. BMT/13/10. https://www.upov.int/edocs/mdocs/upov/en/bmt_13/bmt_13_10.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2013a) Guidance on the use of biochemical and molecular markers in the examination of distinctness, uniformity and stability (DUS), TGP/15. https://www.upov.int/edocs/tgpdocs/en/tgp_15.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2013b) Guidelines for the conduct of tests for distinctness, uniformity and stability, tomato (Solanum lycopersicum L.). TG/44/11 Rev. https://www.upov.int/edocs/tgdocs/en/tg044.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2014a) The use of molecular markers (SNP) for maize DUS testing. BMT/14/10. https://www.upov.int/edocs/mdocs/upov/en/bmt_14/bmt_14_10.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2014b) Identification of SNP markers to aid assessment of essential derivation in maize. BMT/14/7 Rev. https://www.upov.int/edocs/mdocs/upov/en/bmt_14/bmt_14_7_rev.pdf. Accessed 26 Apr 2019
UPOV, International Union for the Protection of New Varieties of Plants (2018) Guidelines for the conduct of tests for distinctness, uniformity and stability, barley (Hordeum vulgare L. sensu lato). TG/19/10. https://www.upov.int/edocs/tgdocs/en/tg019.pdf. Accessed 26 Apr 2019
Van Eeuwijk F, Law J (2004) Statistical aspects of essential derivation, with illustrations based on lettuce and barley. Euphytica 137:129–137
Van Wijk A, Louwaars N (2014) Framework for the introduction of Plant Breeder’s Rights: guidance for practical implementation. Naktuinbouw, Roelofarendsveen, p 207
Vélez MD, Ibánez J (2012) Assessment of the uniformity and stability of grapevine cultivars using a set of microsatellite markers. Euphytica 186(2):419–432
Vosman B, Visser D, van der Voort JR, Smulders MJ, van Eeuwijk F (2004) The establishment of ‘essential derivation’ among rose varieties, using AFLP. Theor Appl Genet 109:1718–1725
Voss-Fels KP, Herzog E, Dreisigacker S, Sukumaran S, Watson A, Frisch M, Hayes B, Hickey LT (2019) “SpeedGS” to accelerate genetic gain in spring wheat. In: Miedaner T, Korzun V (eds) Application of genetic and genomic research in cereals. Woodhead Publishing, Elsevier, Sawston. https://doi.org/10.1016/B978-0-08-102163-7.00014-4
Wang LX, Li HB, Gu TC, Liu LH, Pang BS, Qiu J, Zhao CP (2013) Assessment of wheat variety stability using SSR markers. Euphytica 195:435–452
Wang L-X, Jun Q, Chang L-F, Liu L-H, Li H-B, Pang B-S, Zhao C-P (2015) Assessment of wheat variety distinctness using SSR markers. J Integr Agric 14:1923–1935
Wang J, Cogan NO, Forster JW (2016) Prospects for applications of genomic tools in registration testing and seed certification of ryegrass varieties. Plant Breed 135:405–412
Watson A, Ghosh S, Williams MJ, Cuddy WS, Simmonds J, Rey M-D, Asyraf Md Hatta M, Hinchliffe A, Steed A, Reynolds D et al (2018) Speed breeding is a powerful tool to accelerate crop research and breeding. Nat Plants 4:23–29
Yadav VK, Singh IS (2010) Comparative evaluation of maize inbred lines (Zea mays L.) according to DUS testing using morphological, physiological and molecular markers. Agric Sci 1:131–142
Yamanaka S, Nakamura I, Watanabe KN, Sato Y-I (2004) Identification of SNPs in the waxy gene among glutinous rice cultivars and their evolutionary significance during the domestication process of rice. Theor Appl Genet 108:1200–1204
Yim GR, Ding DR, Wang CP, Leong WH, Hong Y (2009) Microsatellite markers to complement distinctness, uniformity, stability testing of Brassica chinensis (Xiao baicai) varieties. Open Horticulture Journal 2:54–61
Acknowledgements
SHJ acknowledged a grant support No. 8229/253 from Seed and Plant Certification and Registration Institute. JC’s time was supported by Biotechnology and Biological Sciences Research Council (BBSRC) grant BB/L011700/1. LTH was supported by an Australian Research Council Early Career Discovery Research Award (Project Code DE170101296). The authors also thank Mr. Deon Goosen, Director of Commercial Engagement-Agricultural and Food Sciences, Uniquest, Australia, for his comments to improve this article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Rajeev K. Varshney.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jamali, S.H., Cockram, J. & Hickey, L.T. Insights into deployment of DNA markers in plant variety protection and registration. Theor Appl Genet 132, 1911–1929 (2019). https://doi.org/10.1007/s00122-019-03348-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03348-7