Abstract
Key message
One major and three minor QTLs for resistance to pre-harvest sprouting (PHS) were identified from a white wheat variety “Danby.” The major QTL on chromosome 3A is TaPHS1, and the sequence variation in its promoter region was responsible for the PHS resistance. Additive × additive effects were detected between two minor QTLs on chromosomes 3B and 5A, which can greatly enhance the PHS resistance.
Abstract
Pre-harvest sprouting (PHS) causes significant losses in yield and quality in wheat. White wheat is usually more susceptible to PHS than red wheat. Therefore, the use of none grain color-related PHS resistance quantitative trait loci (QTLs) is essential for the improvement in PHS resistance in white wheat. To identify PHS resistance QTLs in the white wheat cultivar “Danby” and determine their effects, a doubled haploid population derived from a cross of Danby × “Tiger” was genotyped using genotyping-by-sequencing markers and phenotyped for PHS resistance in two greenhouse and one field experiments. One major QTL corresponding to a previously cloned gene, TaPHS1, was consistently detected on the chromosome arm 3AS in all three experiments and explained 21.6–41.0% of the phenotypic variations. A SNP (SNP−222) in the promoter of TaPHS1 co-segregated with PHS in this mapping population and was also significantly associated with PHS in an association panel. Gene sequence comparison and gene expression analysis further confirmed that SNP−222 is most likely the causal mutation in TaPHS1 for PHS resistance in Danby in this study. In addition, two stable minor QTLs on chromosome arms 3BS and 5AL were detected in two experiments with allele effects consistently contributed by Danby, while one minor QTL on 2AS was detected in two environments with contradicted allelic effects. The two stable minor QTLs showed significant additive × additive effects. The results demonstrated that pyramiding those three QTLs using breeder-friendly KASP markers developed in this study could greatly improve PHS resistance in white wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pre-harvest sprouting (PHS) in wheat (Triticum aestivum L.) occurs when physiologically matured spikes are exposed to a long wet period in the field before harvest (Cabral et al. 2014; Li et al. 2004; Mares and Mrva 2014). PHS may cause significant losses of grain yield and quality due to degraded starch and protein in germinated kernels (Flintham 2000; Shorinola et al. 2016). In the USA, PHS caused million dollars of losses to wheat growers in Washington State in 2013 alone (Steber et al. 2014). The average annual losses due to PHS are approximately $100 million in Canada and more than $1 billion worldwide (DePauw et al. 2012).
Compared with red wheat, white wheat usually has sweeter taste, higher flour yield and lighter color for its end-use products, making it more attractive for making Asian noodle, steamed bread and many whole wheat flour-based products (Fakthongphan et al. 2016). Although hard red winter wheat predominates wheat production in the US Great Plains, hard white winter wheat production acreage has been increasing since the late 1980s due to strong international market demands (Fakthongphan et al. 2016). However, PHS in white wheat has been a major barrier for the expansion of its production area. Breeding cultivars with resistance to PHS are an effective method to reduce the losses from sprouted grains in wheat production (Barrero et al. 2015; Gao and Ayele 2014). Therefore, improvement in PHS resistance in white wheat is critical for increasing its production in the US Great Plains to meet the growing international and domestic market demands.
Seed dormancy (SD) refers to the temporary resistance to germination for a viable seed under favorable environmental conditions. Lack of adequate SD is the major factor for PHS (Li et al. 2004). Besides SD, grain color and wheat spike structure, such as the presence of awns, ear nodding angle and glume tenacity, also affect PHS (King and Richards 1984; Mares and Mrva 2014). Red wheat cultivars are typically more resistant than white wheat cultivars (Groos et al. 2002). The association between PHS and grain color might be due to either tight genetic linkage between genes of PHS resistance and grain color or pleiotropic effects of the grain color genes (Flintham 2000; Lin et al. 2016). Although red wheat is more resistant to PHS than white wheat, breeding programs have made great improvements for PHS resistance in white wheat. Several white wheat cultivars released for the Great Plains, including “Rio Blanco,” “Trego” (Martin et al. 2001), “Danby”, “Snowmass” (Haley et al. 2011), “Clara CL” (Martin et al. 2014), “Antero” (Haley et al. 2014) and “Sunshine” (Haley et al. 2017), have at least moderate PHS resistance. These cultivars share a similar source of PHS resistance (http://wheatpedigree.net). Other genetic sources of PHS resistance have also been found in white wheat, including Aus1408, “Clark’s Cream,” NY6432-18, “Halberd,” “Konde,” “Kumpa” and “Swindy” (Jiménez et al. 2017; Mares and Mrva 2001; Mares et al. 2005; Paterson and Sorrells 1990). Combining these different resistant sources may greatly enhance the PHS resistance in new cultivars.
PHS can be evaluated in both field and controlled environments. Field evaluation of PHS resistance depends on the presence of weather conditions conducive for seed sprouting after the physiological maturity; thus, the PHS phenotypic data may not be repeatable in different environments (Graybosch et al. 2013; Kato et al. 2001). Therefore, PHS is usually evaluated under controlled environments. Evaluating sprouting rate of whole spikes in a misting chamber is one of the most common methods, but seed germination tests in petri dishes (Clarke et al. 2005) and indirect assays using flour falling number tests (Barnard and Bona 2004; Hareland 2003) are also common methods for PHS evaluation. However, all PHS phenotyping methods are time-consuming and labor intensive. Marker-assisted selection (MAS) is a desirable alternative that can reduce phenotyping cost, improve selection accuracy and shorten breeding cycles.
Markers tightly linked to PHS resistance QTLs are essential for MAS (Gao et al. 2013; Kulwal et al. 2012; Mares and Mrva 2014). Many PHS resistance QTLs have been reported, and they are located on almost all 21 wheat chromosomes (Anderson et al. 1993; Flintham et al. 2002; Groos et al. 2002; Kato et al. 2001; Li et al. 2004; Mares and Mrva 2001; Roy et al. 1999; Zanetti et al. 2000; Zhu et al. 2016). Among them, the QTLs on chromosome arms 3AS and 4AL showed major effects on PHS and have been investigated intensively (Albrecht et al. 2015; Bi et al. 2014; Cao et al. 2016; Fakthongphan et al. 2016; Lei et al. 2013; Lin et al. 2015; Liu et al. 2008, 2011; Lohwasser et al. 2013; Miao et al. 2013; Miura et al. 2002; Mori et al. 2005; Rasul et al. 2009; Xiao et al. 2012). Recently, the underlying genes for those two PHS resistance QTLs have been cloned and their causal mutations have been identified (Barrero et al. 2015; Liu et al. 2013; Nakamura et al. 2011; Torada et al. 2016). Two independent studies reported cloning of the QTL on chromosome arm 3AS that was designated as TaPHS1 by Liu et al. (2013) and as MFT by Nakamura et al. (2011). Among three different causal SNPs identified (Nakamura et al. 2011; Liu et al. 2013), a SNP in the promoter region was associated with the SD of a red wheat that was grown at a low temperature (13 °C) (Nakamura et al. 2011), whereas two SNPs in the gene-coding region were reported to be responsible for PHS resistance in a white wheat cultivar, Rio Blanco, grown under normal temperatures (Liu et al. 2013). Liu et al. (2015) suggested that TaPHS1 might have played a critical role during wheat domestication.
Danby has been the most popular white wheat cultivar in the Great Plains for over 10 years, and it has a high level of PHS resistance. However, the PHS resistance in Danby is not yet fully understood. The objectives of this study were to identify PHS resistance QTLs in Danby, determine their effects and develop user-friendly DNA markers for MAS.
Materials and methods
Plant materials
A population of 211 doubled haploid (DH) lines was developed by Heartland Plant Innovations, Manhattan, KS, from a cross between Danby and “Tiger” (Martin et al. 2013), a PHS-susceptible white wheat cultivar, using the wheat–maize wide hybridization method. This population and its parents were evaluated for PHS in three environments, two in a greenhouse and one in a field. In addition, an association-mapping population of 167 US winter wheat cultivars and elite breeding lines evaluated for PHS in multiple environments (Lin et al. 2016, see panel list and PHS data in supplementary table) was used for marker validation.
PHS phenotyping in greenhouse and field experiments
To evaluate PHS resistance, the DH population and its parents were grown in a greenhouse at Kansas State University, Manhattan, KS, in 2014 and 2015, and in a field at Hays, KS, in 2015. The association-mapping population was evaluated for PHS in both greenhouse (2012 and 2013) and field (2013 and 2014) experiments at Manhattan, KS, and in the field (2013 and 2014) experiments at Hays, KS (Lin et al. 2016). In the greenhouse experiments, five plants per line were grown in a 13×13 cm plastic pot, and pots were arranged in a randomized complete block design with two replications. The greenhouse was set at 22 °C day/17 °C night with 12-h supplemental light. In the field experiments, all the lines were planted in 1.2-m-long single-row plots with two replications using a randomized complete block design. Around 35 seeds were sowed in each plot.
Five spikes were harvested from different plants in each line at their physiological maturity when both the peduncle and spike turned yellow. Harvested spikes were air-dried for 5 days at room temperature and then stored in a freezer at − 20 °C to maintain their dormancy. After all were harvested, they were air-dried again for 6 days at room temperature and then incubated in a misting chamber for 7 days (Liu et al. 2008). Germinated and non-germinated kernels in each tested spike were counted, and the mean percentage of geminated kernels was calculated for each line for subsequent analysis.
Analysis of variance and heritability
Analysis of variance was conducted by SAS program (SAS 9.4, SAS Institute, 2012) using a general linear model (GLM) yijkl = µ + Gi + Ej + Bk(j) + GEij + eijkl, where Gi is the effect of the ith genotype, Ej is the effect of the jth environment, Bk(j) is the blocking effect, GEij is the interaction effect between genotype and environment, and eijkl is the random error in individual plots (Imtiaz et al. 2008). To account for the missing data, type III sum of squares were used. The broad sense heritability across three experiments was estimated using the following equation (Toojinda et al. 1998): H2 = Vg/(Vg + Vge/e + Ve/re), where Vg is the genotypic variance, Vge is the variance of genotype by environment, Ve is the error variance, r is the number of replications, and e is the number of environments.
Assays of genotyping-by-sequencing and simple sequence repeat markers
Genomic DNA was isolated from leaf tissues collected at three-leaf stage using a BioSprint 96 DNA Plant Kit (Qiagen, Hilden, Germany) for genotyping-by-sequencing (GBS) and simple sequence repeats (SSR) markers. For GBS, each DNA sample was normalized to 20 ng/µl for library construction following the protocol described by Poland et al. (2012). Briefly, DNA samples from both DH lines and parents were digested using PstI and MspI restriction enzymes and ligated to forward and reverse adapters. The ligated DNA samples from the parents and all DH lines were pooled into a single tube for PCR amplification. The PCR products were cleaned up and sequenced using Illumina HiSeq 2000 (Illumina, Inc., CA, USA). SNPs were called using a population-based custom Java script and TASSEL (Bradbury et al. 2007). Raw sequence reads were parsed and assigned to samples using barcodes and trimmed to 64 bps in length. To identify SNPs in the DH population, all pairs of tags were evaluated first for one or two base-pair differences. Bi-allelic SNPs were determined by querying the filtered tags for pairs of sequences (Poland et al. 2012) if they differed in only one or two SNPs, were detected in at least 20% genotypes of the population and could pass a Fisher exact independence test. SNPs were discarded if 10% or more DH lines were heterozygotes in the population. Only SNPs with less than 20% missing data were used for map construction.
Thirteen polymorphic SSR markers were randomly selected from a wheat consensus map (Somers et al. 2004) for validating chromosome locations that were assigned based on GBS-SNPs. A 10-µl PCR mix for a SSR marker contained 20–40 ng DNA, 0.4 mM each of reverse and M13-tailed forward primers, 0.4 mM fluorescence-labeled M13 primer, 0.08 mM of each dNTP, 1.2 µl 10X PCR buffer, 1 mM MgCl2 and 0.6 units of Taq polymerase. PCR was performed using a touch-down program (Liu et al. 2008) in a DNA Engine® Peltier Thermal Cycler (Bio-Rad Lab, Hercules, CA, USA). Four different plates of PCR products labeled with one of the four dyes (FAM, VIC, NED and PET) were pooled into one plate using a Biomek NXP liquid handling system (Beckman Coulter Inc., CA, USA). The pooled PCR products were analyzed in an ABI Prism 3730 DNA Sequencer (Applied Biosystems, Foster City, CA, USA), and their fragment lengths and polymorphisms of amplicons were scored using GeneMarker (SoftGenetics LLC, State College, PA, USA).
Linkage map construction and QTL analysis
A linkage map was constructed with GBS-SNP and SSR markers using JoinMap 4.1 (Van Ooijen 2006) and the Kosambi mapping function (Kosambi 1943). A minimum logarithm of odds (LOD) score of 5 and a maximum recombination frequency of 0.35 were set to identify linkage groups. Chromosome names and genetic locations of QTLs on the wheat reference genome were assigned by blasting the GBS tags of mapped SNPs to the flow-sorted Chinese Spring survey sequences (Mayer et al. 2014) and the PopSeq sequence data (Chapman et al. 2015) using a web-based blasting tool (http://129.130.90.211/wpdb/gbsloc). WinQTLCart 2.5 (Wang et al. 2007) was used for QTL analysis using composite interval mapping function (Silva et al. 2012). Significant QTLs were claimed if the LOD scores were above the threshold that was derived from 1000 permutations (Doerge and Churchill 1996). To investigate the combined effects of the identified QTLs, all DH lines were grouped based on different allele combinations of the QTLs. The closest marker to each QTL was selected to represent that QTL. The mean sprouting rates of each allelic group were compared using Tukey’s multiple comparison (Altman 1991).
Sequence analysis and KASP marker development
A new gene-specific Kompetitive allele-specific (KASP) marker was designed for the possible causal SNP at position −222 (SNP−222) in the promoter of TaPHS1 based on published promoter sequence (Nakamura et al. 2011; Liu et al. 2013) to improve the application efficiency of the original marker developed by Liu et al. (2015). A new KASP marker was also developed for SNP−314; another SNP identified in the promoter region in this study. Forward and reverse primers were designed using Primer3 web version 4.0.0 (http://primer3plus.com/primer3web/primer3web_input.htm). KASP markers were assayed following the manufacturer’s instruction (http://www.lgcgroup.com/LGCGroup/media/PDFs/Products/Genotyping/KASP-genotyping-chemistry-User-guide.pdf). For those GBS-SNP markers tightly linked to the QTLs, primers were designed using the web-based primer design pipeline (http://polymarker.tgac.ac.uk/) that was developed specifically to design homoeolog-specific KASP assays for the polyploidy wheat genome (Ramirez-Gonzalez et al. 2015). The newly designed KASP primers were then tested for parental polymorphisms, and the polymorphic SNPs were genotyped in the mapping population. The KASP-SNP data were used to replace their corresponding GBS-SNPs, and the map was re-constructed for QTL validation. The KASP assay was performed in a 6-µl PCR mix that consisted of 2.9 µl of reaction mix (LGC Genomics, Beverly, MA, USA), 0.1 µl of primer assay mix and 3 µl of DNA at a concentration of 15 ng/µl. PCR was assayed following the manufacturer’s instruction (LGC Genomics) using an ABI 7900HT Real-Time PCR System (Life Technology, Grand Island, NY, USA).
Gene expression analysis
Seeds of Danby, Trego, Tiger and “Lakin” were germinated in separate petri dishes in three replications under room temperature for 5 days, and embryos were excised from the imbibed seeds for RNA isolation. RNA from the embryos was isolated and purified using the RNeasy plant kit with on-column DNase I treatment (Qiagen, Valencia, CA). Quantitative real-time PCR was conducted using TaPHS1-specific primers as described in Liu et al. (2013). Transcription levels were normalized using the 18S rRNA as an internal control to make comparisons among different genotypes.
Results
Phenotypic analysis
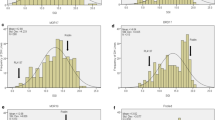
The DH population and its parents were evaluated for PHS resistance in three experiments. Danby displayed a significantly (P < 0.01) lower PHS rate (14.3%) than Tiger (64.9%) on an average across three experiments (Fig. 1). The population had the highest PHS rate in 2014 greenhouse experiment with a mean of 49.1% and a range from 0 to 98.6%, and the lowest PHS rate in 2015 field experiment with a mean of 26.9% and a range from 0 to 87.8%. The analysis of variance for PHS rates across the three experiments showed highly significant (P < 0.0001) variations not only for genotypes but also for environments (Table 1). Significant genotype × environment interactions were also detected. The broad sense heritability for PHS resistance was high (0.72) across three experiments.
Frequency distributions of pre-harvest sprouting (PHS) rates in the DH population. The PHS rates are the mean values of the two replications in each experiment. Black arrows point to the PHS rates of the parental lines. a PHS rate from 2014 greenhouse experiment. b PHS rate from 2015 greenhouse experiment. c PHS rate from 2015 field experiment
Genetic linkage map
A total of 5578 and 2580 GBS-SNPs were identified with less than 50 and 20% missing data, respectively. Those GBS-SNPs with less than 20% missing data were combined with 13 SSR markers to construct a linkage map. A total of 1811 GBS-SNPs and all 13 SSR markers were mapped on 31 linkage groups of 1476 cM covering all the 21 chromosomes of common wheat with some of them composed of more than one linkage group. The average length of linkage groups was 47.6 cM with an average marker interval of 1.2 cM. The marker distribution was uneven across three genomes, with 721 markers (39.5%) on A genome, 648 markers (35.5%) on B genome and 455 markers (24.9%) on D genome (Fig. 2). The A genome had the highest marker density (1.5 markers/cM), followed by the B genome (1.2 markers/cM) and D genome (0.8 markers/cM). The numbers of markers per chromosome varied greatly within a genome. In the A genome, chromosomes 2A and 5A had the most markers (165 markers each), while chromosome 1A had the fewest (16 markers); in the B genome, chromosome 3B had the most markers (163 markers) and 1B had the fewest (9 markers); in the D genome, chromosome 6D had 194 markers, while 4D had only two markers.
QTLs for PHS resistance
Composite interval mapping identified four QTLs for PHS resistance on chromosome arms 2AS (Qphs.hwwg-2A.1), 3AS (Qphs.hwwg-3A.1), 3BS (Qphs.hwwg-3B.1) and 5AL (Qphs.hwwg-5A.1) (Fig. 3, Table 2), and they were all consistently detected in at least two experiments. Three of them, Qphs.hwwg-3A.1, Qphs.hwwg-3B.1 and Qphs.hwwg-5A.1, were contributed by the resistant parent Danby, while Qphs.hwwg-2A.1 showed contradictory allelic effects between two experiments. Qphs.hwwg-3A.1 was identified in all the three experiments and explained 21.6 to 41.0% of the phenotypic variations (PVE). Qphs.hwwg-3B.1 had PVE of 4.7% in both 2014 greenhouse and 2015 field experiments. Qphs.hwwg-5A.1 explained 6.4 and 5.4% of PVE in the 2014 and 2015 greenhouse experiments, respectively. Qphs.hwwg-2A.1 was identified in the 2014 greenhouse and 2015 field experiments with PVE of 4.7 and 15.3%, respectively; however, Danby contributed the resistant allele in the 2014 greenhouse experiment and the susceptible allele in the 2015 field experiment.
Composite interval mapping of a major QTL on chromosome arm 3AS and two minor QTLs on chromosome arms 3BS and 5AL for pre-harvest sprouting resistance in 2014 greenhouse (2014GH), 2015 greenhouse (2015GH) and 2015 field (2015FLD) experiments. Line parallel to the X-axis is the threshold for claiming significant QTLs with a logarithm of odds at 2.5. Genetic distances of molecular markers are shown in centimorgans (cM) along the X-axis
Combination effects among the QTLs
Three resistance QTLs, Qphs.hwwg-3A.1 (A), Qphs.hwwg-3B.1 (B) and Qphs.hwwg-5A.1 (C) with the Danby alleles for resistance, were selected to investigate their combination effects. All the lines in the DH population were grouped into eight groups (abc, aBc, abC, aBC, Abc, AbC, ABc and ABC) based on allele combinations of those three QTLs, and the sprouting rates were compared among the groups (Fig. 4).
Comparisons of pre-harvest sprouting rates (%) among different QTL combinations. Letter combinations abc, abC, aBc, aBC, Abc, AbC, ABc, ABC were designated for eight possible allele combinations of the three consistent QTLs, Qphs.hwwg-3A.1 (a), Qphs.hwwg-3B.1 (b) and Qphs.hwwg-5A.1 (c). On the x-axis, capital letters, A, B and C, were designated as resistance alleles, and a, b and c were susceptibility alleles for the three QTLs, respectively. The number below the letters is the sample size for each genotype. Error bar is the standard error of the group mean
When Qphs.hwwg-3A.1 was absent, Qphs.hwwg-3B.1 (aBc) and Qphs.hwwg-5A.1 (abC) only significantly (P < 0.05) reduced the sprouting rate in 2015 field experiment and 2014 greenhouse experiment, respectively (Fig. 4). However, when those two minor QTLs were combined (aBC), the sprouting rates were significantly (P < 0.05) lower in all three experiments than those without QTLs (abc), demonstrating an additive × additive effect between the two minor QTLs. The two minor QTLs together reduced the sprouting rate from 78.8% (abc) to 51.8% (aBC) in the 2014 greenhouse experiment, from 53.0 to 34.4% in the 2015 greenhouse experiment and from 45.1 to 28.9% in the 2015 field experiment.
When the major resistance QTL, Qphs.hwwg-3A.1, was present, adding either Qphs.hwwg-3B.1 or Qphs.hwwg-5A.1 did not significantly (P > 0.05) reduce the PHS rates (Fig. 4). However, when both minor QTLs were present with Qphs.hwwg-3A.1 (ABC), PHS rates were significantly (P < 0.05) reduced compared to the group with only the Qphs.hwwg-3A.1 resistance allele (Abc) in all three experiments (Fig. 4). In the 2014 greenhouse experiment, the sprouting rate of the ABC group (20.3%) was half that for the Abc group (40.1%). Similar trends were observed in the 2015 greenhouse (12.1 vs 23.9%) and 2015 field experiment (5.4 vs 20.7%). Therefore, the addition of the two minor QTLs could greatly enhance the PHS resistance.
Sequence analysis and KASP marker development
Qphs.hwwg-3A.1 was mapped to the same position as TaPHS1 based on the common linked SSR marker Xbarc321 (Liu and Bai 2010; Liu et al. 2008; Mori et al. 2005; Nakamura et al. 2011). Two previously reported KASP markers, developed based on two causal SNP variations (SNP646 and SNP666) in TaPHS1 (Liu et al. 2013), were monomorphic between the two parents in this study. This result turned our attention to other reported SNP variations in TaPHS1. Comparison of the gene sequences of TaPHS1 between Danby and Tiger (Liu et al. 2013) identified seven SNPs at positions −823, −714, −712, −620, −573, −314, −222 in the promoter and one deletion (TGG) at +883 in the third intron of the coding region. Gene structure analysis and function prediction suggested that only SNP−222 and SNP−314 were functional variations. Two new KASP markers (KASP−222 and KASP−314) were developed based on their sequence information (Table 3). The newly designed KASP−222 marker is four base pairs shorter in forward primers than the previous one and has a completely different reverse primer. This newly designed KASP assay showed much better separation between the two alleles than previous reported marker (Liu et al. 2015). In the association analysis using the association panel, KASP−314 was not significantly (P = 0.17) associated with the PHS variations, while KASP−222 was significantly (P < 0.01) associated with the PHS variations (See SNP allele information in supplementary table). This KASP−222 marker was genotyped on the DH population (Fig. 5), mapped under the peak of Qphs.hwwg-3A.1 and explained the greatest phenotypic variation among all the markers mapped in the QTL region. These results indicate that SNP−222 in the promoter is most likely the causal variation of TaPHS1 in Danby.
KASP assays of the causal single nucleotide polymorphism (SNP) at the position of −222 bp in the promoter of TaPHS1 in a the double haploid mapping population and b an association panel of 167 wheat accessions. Green dots are the resistant genotypes with C nucleotide. Blue dots are the susceptible genotypes with T nucleotide. Red dots represent heterozygous genotypes. The black dots are blank controls, and cross symbols represent undetermined genotypes because of unsuccessful PCRs (color figure online)
To make another two stable QTLs, Qphs.hwwg-3B.1 and Qphs.hwwg-5A.1, available for marker-assisted selection, five user-friendly KASP markers were developed based on their flanking GBS-SNP sequences (Table 3). All the five KASP markers were genotyped and mapped to the corresponding QTL regions in the mapping population (Figs. 3 and 6).
KASP marker assays developed from the flanking GBS markers of Qphs.hwwg-3B.1 and Qphs.hwwg-5A.1 in the double haploid mapping population. KASP471 and KASP765 are flanking Qphs.hwwg-3B.1. KASP868, KASP907 and KASP8426 are linked to Qphs.hwwg-5A.1. Green dots show the resistant genotypes with T nucleotide. Blue dots show the susceptible genotypes with C nucleotide. Red dots represent heterozygous genotypes. The black dots are blank controls, and cross symbols represent undetermined genotypes because of unsuccessful PCRs (color figure online)
Differential gene expression of TaPHS1 between parental lines
To further confirm the gene function of TaPHS1, gene expression study was conducted for parental lines, Danby and Tiger, and other two control cultivars, Trego and Lakin. Same as Danby and Tiger, Trego and Lakin have the same resistance alleles at SNP+646 and SNP+666, but different alleles at SNP−222. Trego is a PHS-resistant cultivar and carries the resistance allele at SNP−222 as Danby, whereas Lakin is PHS-susceptible and has the susceptible allele at SNP−222 as Tiger. Quantitative RT-PCR results showed that the mRNA expression levels of TaPHS1 in the two resistant cultivars (Danby and Trego) were at least eight times higher than those in two susceptible cultivars Tiger and Lakin at 5 days after imbibing during seed germination (Fig. 7) (p < 0.01). The significant gene expression difference between the genotypes with the contrasting alleles at SNP−222 supported SNP−222 as the causal variation in TaPHS1 for PHS resistance in Danby.
Relative TaPHS1 expression levels in embryos of four wheat genotypes. Wheat cultivars Danby and Trego carry the resistant haplotype (RRR) in three SNPs, SNP−222, SNP+646 and SNP+666 of TaPHS1, respectively, whereas Tiger and Lakin carry the susceptible haplotype (SRR). The relative mRNA level in Lakin was set as a standard value. Error bars show standard deviation of three replications
Discussion
Using GBS-SNPs, we identified four QTLs for PHS resistance in this study. Qphs.hwwg-3A.1 showed a major effect on PHS resistance and was identified in all three experiments conducted. Kulwal et al. (2005) identified a major QTL for PHS on chromosome arm 3AL, which should be different from our Qphs.hwwg-3A.1 on 3AS. However, our QTL was co-localized with the previously cloned PHS resistance gene, TaPHS1 (Liu et al. 2013). This was expected since one of Danby’s parental lines (Trego) was derived from Rio Blanco that was used by Liu et al. (2013) for cloning TaPHS1. In the TaPSH1 cloning study, Liu et al. (2013) found three possible causal mutations at positions of −222, +646 and +666 in Rio Blanco, but claimed mutations at positions of +646 and +666 as main factors because of the insignificance of SNP−222 in an association panel. In this study, a Rio Blanco-derived cultivar Danby was used, and it has the same genotypes at all those three SNPs as Rio Blanco. However, Danby and PHS-susceptible parent Tiger in our study are monomorphic at both positions of +646 and +666. After a further analysis of the sequences between the two parental lines, SNP−222 was found polymorphic and considered the most likely causal variation in our study. This result was further confirmed in an association panel, which agrees with two previous reports (Nakamura et al. 2011; Chono et al. 2015), but disagrees with Liu et al. (2013). Liu et al. (2013) used a small subset of this association panel (83 accessions) and did not find a significant association between SNP−222 and PHS. This discrepancy among the studies might be due to the sampling bias. When the panel of genotypes investigated has allelic differences at SNP+646 and SNP+666, but not at SNP−222, the effect of SNP−222 may not be significant, whereas effect of SNP−222 can be significant when the genotypes studied carry contrasting alleles at SNP−222 but resistance alleles at both SNP+646 and SNP+666. Our gene expression study showed that SNP−222 increased TaPHS1 expression in Danby, but reduced the TaPHS1 expression in Tiger, which validated the effect of SNP−222 on regulating TaPHS1 expression (Nakamura et al. 2011). These results indicate that all these three mutations in TaPHS1 can change the expression levels of TaPHS1 (Liu et al. 2013; Nakamura et al. 2011), resulting in phenotypic variation in PHS. The causal SNPs may not be the same in different breeding populations depending on resistant sources and other parents used to develop the populations. Therefore, for marker-assisted breeding, all the three markers should be investigated to identify the key SNP that changes TaPHS1 expression as target marker for selection in a breeding population.
Our study also found two minor QTLs, Qphs.hwwg-3B.1 and Qphs.hwwg-5A.1, which were not identified in Rio Blanco (Liu et al. 2008). Those two minor QTLs might be inherited from other parents of Danby. Qphs.hwwg-3B.1 showed a minor effect but was repeatedly detected in both greenhouse and field experiments. Several previous studies have reported QTLs associated with PHS or SD on chromosome 3B, but they are all located on the long arm near the red grain color gene (R-B1) or Viviparous 1 (Cabral et al. 2014; Chang et al. 2010; Fofana et al. 2009; Groos et al. 2002; Lin et al. 2016; Mares et al. 2009; Somers et al. 2004; Yang et al. 2007a, b). In the current study, Qphs.hwwg-3B.1 was located in a region between 55.8 and 61.3 cM on 3BS of the Chinese Spring reference map after blasting the GBS tags of flanking SNPs against the flow-sorted Chinese Spring survey sequences (Mayer et al. 2014). This QTL region on 3BS is different from all previously reported QTLs. Therefore, Qphs.hwwg-3B.1 is more likely a novel PHS resistance QTL that does not relate to seed color, and it could be valuable for pyramiding with other QTLs to improve PHS resistance in white wheat.
Qphs.hwwg-5A.1 was another minor QTL that was consistently detected in the two greenhouse experiments. The peak of this QTL was 8.8 cM away from the distal end of the long arm of chromosome 5A on the Chinese Spring reference map based on the blasting result against the flow-sorted Chinese Spring survey sequences (Mayer et al. 2014). Several PHS-related QTLs were reported on chromosome 5A, but none of them were near the distal end of 5AL as they were either near the centromere (Iehisa et al. 2014; Nakamura et al. 2010) or on the short arm (Groos et al. 2002). In a cross between common wheat and spelt wheat (Tritcum Spelta), Zanetti et al. (2000) found a QTL for alpha-amylase activity on chromosome arm 5AL at the q locus, which is also far from Qphs.hwwg-5A.1. Genome-wide association studies identified two significant PHS-related QTLs on 5AL, but one was near the centromere (Zhu et al. 2016) and the other was more than 20 cM away from Qphs.hwwg-5A.1 (Lin et al. 2016). Therefore, Qphs.hwwg-5A.1 is also likely a novel QTL for PHS resistance.
Although, individually, these two QTLs showed a minor effect on PHS resistance, a combination of Qphs.hwwg-5A.1 and Qphs.hwwg-3B.1 greatly reduced the PHS with or without Qphs.hwwg-3A.1. These two minor QTLs, together with Qphs.hwwg-3A.1, reduced PHS sprouting by 50% or more in comparison with the genotype with Qphs.hwwg-3A.1 alone in all the three experiments. This significantly enhanced PHS resistance suggests a valuable additive × additive effect between these two minor QTLs. Gene interactions are critical in advanced quantitative genetic models, and assembly of favorable QTL or gene combinations is very important not only for crop breeding but also for understanding the genetic basis underlying crop adaptation and evolution (Allard 1996). QTL interactions have been reported for PHS resistance or SD in many crops including rice (Gu et al. 2004; Guo et al. 2004; Wang et al. 2014), barley (Hickey et al. 2012) and wheat (Imtiaz et al. 2008; Kumar et al. 2009; Liu et al. 2011; Mohan et al. 2009). However, QTL interactions between chromosomes 3B and 5A have not been previously documented for PHS resistance in wheat. Our study is the first to reveal the significant interactions between these two QTLs. The additive × additive effect detected in this study strongly suggests the genetically complex networks in wheat PHS regulation and the importance in pyramiding a specific combination of QTLs or genes in breeding.
Qphs.hwwg-2A.1 was detected in two environments but with contradicted allelic effects, in which Danby contributed the resistant allele in the greenhouse experiment but susceptible allele in the field experiment. Severe stripe rust infection in the 2015 field experiment was most likely responsible for the shift of allelic effects. Using the stripe rust data from the same population, a major resistance QTL contributed by Danby was mapped at the same location as Qphs.hwwg-2A.1 in the distal end of chromosome 2AS (unpublished data). We noticed that plants infected by stripe rust produced shriveled seeds, which might affect germination during the PHS test. Therefore, the PHS resistance allele on 2A contributed by Tiger in 2015 field experiment was most likely due to the rust susceptible allele in Tiger. It is also possible that the correlation between stripe rust susceptibility and PHS resistance is due to plant hormones triggered by rust infection that could suppress seed germination. Plant hormones such as abscisic acid (ABA) and gibberellic acid (GA), the most important regulator of seed dormancy, might be involved in the responses to many biotic stresses including disease and insect (de Zelicourt et al. 2016; Lee and Luan 2012; Pieterse et al. 2012; Skubacz et al. 2016; Verslues and Zhu 2005). Therefore, the effect of Qphs.hwwg-2A.1 on PHS resistance is unclear and needs to be further explored in experiments without confounding effects from stripe rust infection.
Conclusions
Our study identified one major PHS resistance QTL on chromosome arm 3AS and three minor ones on chromosome arms 2AS, 3BS and 5AL in a white wheat DH population by evaluating the population in both greenhouse and field experiments. We discovered that SNP−222 in the promoter region of TaPHS1 is most likely responsible for the major QTL in this mapping population. We developed a new KASP marker for this SNP for MAS. This KASP marker was validated in an association panel. Our gene expression result further validated the effect of SNP−222 in Danby. Two minor QTLs on 3BS and 5AL appear to be novel QTLs for PHS resistance and showed a valuable additive × additive effect when they were present together with or without Qphs.hwwg-3A.1. The results indicate that pyramiding these three QTLs can greatly reduce PHS. Five breeder-friendly KASP markers were developed for the two minor QTLs and validated in this study. These KASP markers will be valuable tools to assist breeders for genetic improvement in PHS resistance in white wheat.
Author contribution statement
GZ and GB conceived the study. MS, ML, SL and TK conducted experiments. TR and JP contributed GBS data. HC and YL provided gene expression data. MS analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
References
Albrecht T, Oberforster M, Kempf H, Ramgraber L, Schacht J, Kazman E, Zechner E, Neumayer A, Hartl L, Mohler V (2015) Genome-wide association mapping of preharvest sprouting resistance in a diversity panel of European winter wheats. J Appl Genet 56:277–285
Allard RW (1996) Genetic basis of the evolution of adaptedness in plants. Euphytica 92:1–11
Altman DG (1991) Practical statistics for medical research. Chapman and Hall, London, pp 210–211
Anderson JA, Sorrells ME, Tanksley SD (1993) Rflp analysis of genomic regions associated with resistance to preharvest sprouting in wheat. Crop Sci 33:453–459
Barnard A, Bona L (2004) Sprout damage and falling number in South African and Hungarian wheats. Cereal Res Commun 32:259–264
Barrero JM, Cavanagh C, Verbyla KL, Tibbits JFG, Verbyla AP, Huang BE, Rosewarne GM, Stephen S, Wang PH, Whan A, Rigault P, Hayden MJ, Gubler F (2015) Transcriptomic analysis of wheat near-isogenic lines identifies PM19-A1 and A2 as candidates for a major dormancy QTL. Genome Biol 16:93. https://doi.org/10.1186/s13059-015-0665-6
Bi HH, Sun YW, Xiao YG, Xia LQ (2014) Characterization of DFR allelic variations and their associations with pre-harvest sprouting resistance in a set of red-grained Chinese wheat germplasm. Euphytica 195:197–207
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Cabral AL, Jordan MC, McCartney CA, You FM, Humphreys DG, MacLachlan R, Pozniak CJ (2014) Identification of candidate genes, regions and markers for pre-harvest sprouting resistance in wheat (Triticum aestivum L.). BMC Plant Biol 14:340. https://doi.org/10.1186/s12870-014-0340-1
Cao LZ, Hayashi K, Tokui M, Mori M, Miura H, Onishi K (2016) Detection of QTLs for traits associated with pre-harvest sprouting resistance in bread wheat (Triticum aestivum L.). Breed Sci 662:260–270
Chang C, Zhang HP, Feng JM, Yin B, Si HQ, Ma CX (2010) Identifying alleles of Viviparous-1B associated with pre-harvest sprouting in micro-core collections of Chinese wheat germplasm. Mol Breed 25:481–490
Chapman JA, Mascher M, Buluc A, Barry K, Georganas E, Session A, Strnadova V, Jenkins J, Sehgal S, Oliker L, Schmutz J, Yelick KA, Scholz U, Waugh R, Poland JA, Muehlbauer GJ, Stein N, Rokhsar DS (2015) A whole-genome shotgun approach for assembling and anchoring the hexaploid bread wheat genome. Genome Biol 16:26. https://doi.org/10.1186/s13059-015-0582-8
Chono M, Matsunaka H, Seki M, Fujita M, Kiribuchi-Otobe C, Oda S, Kojima H, Nakamura S (2015) Molecular and genealogical analysis of grain dormancy in Japanese wheat varieties, with specific focus on MOTHER OF FT AND TFL1 on chromosome 3A. Breed Sci 65:103–109
Clarke FR, Knox RE, DePauw RM (2005) Expression of dormancy in a spring wheat cross grown in field and controlled environment conditions. Euphytica 143:297–300
de Zelicourt A, Colcombet J, Hirt H (2016) The role of MAPK modules and ABA during abiotic stress signaling. Trends Plant Sci 21:677–685
DePauw RM, Knox RE, Singh AK, Fox SL, Humphreys DG, Hucl P (2012) Developing standardized methods for breeding preharvest sprouting resistant wheat, challenges and successes in Canadian wheat. Euphytica 188:7–14
Doerge RW, Churchill GA (1996) Permutation tests for multiple loci affecting a quantitative character. Genetics 142:285–294
Fakthongphan J, Bai G, Amand PS, Graybosch RA, Baenziger PS (2016) Identification of markers linked to genes for sprouting tolerance (independent of grain color) in hard white winter wheat (HWWW). Theor Appl Genet 129:419–430
Flintham JE (2000) Different genetic components control coat-imposed and embryo-imposed dormancy in wheat. Seed Sci Res 10:43–50
Flintham J, Adlam R, Bassoi M, Holdsworth M, Gale M (2002) Mapping genes for resistance to sprouting damage in wheat. Euphytica 126:39–45
Fofana B, Humphreys DG, Rasul G, Cloutier S, Brule-Babel A, Woods S, Lukow OM, Somers DJ (2009) Mapping quantitative trait loci controlling pre-harvest sprouting resistance in a red × white seeded spring wheat cross. Euphytica 165:509–521
Gao F, Ayele BT (2014) Functional genomics of seed dormancy in wheat: advances and prospects. Front Plant Sci 5:458. https://doi.org/10.3389/fpls.2014.00458
Gao X, Hu CH, Li HZ, Yao YJ, Meng M, Dong J, Zhao WC, Chen QJ, Li XY (2013) Factors affecting pre-harvest sprouting resistance in wheat (Triticum aestivum L.): a review. J Anim Plant Sci 23:556–565
Graybosch RA, St Amand P, Bai GH (2013) Evaluation of genetic markers for prediction of preharvest sprouting tolerance in hard white winter wheats. Plant Breed 132:359–366
Groos C, Gay G, Perretant MR, Gervais L, Bernard M, Dedryver F, Charmet D (2002) Study of the relationship between pre-harvest sprouting and grain color by quantitative trait loci analysis in a whitexred grain bread-wheat cross. Theor Appl Genet 104:39–47
Gu XY, Kianian SF, Foley ME (2004) Multiple loci and epistases control genetic variation for seed dormancy in weedy rice (Oryza sativa). Genetics 166:1503–1516
Guo LB, Zhu LH, Xu YB, Zeng DL, Wu P, Qian Q (2004) QTL analysis of seed dormancy in rice (Oryza sativa L.). Euphytica 140:155–162
Haley SD, Johnson JJ, Peairs FB, Stromberger JA, Heaton EE, Seifert SA, Kottke RA, Rudolph JB, Martin TJ, Bai G, Chen X, Bowden RL, Jin Y, Kolmer JA, Seifers DL, Chen M, Seabourn BW (2011) Registration of ‘Snowmass’ wheat. J Plant Reg 5:87–90
Haley SD, Johnson JJ, Peairs FB, Stromberger JA, Hudson-Arns EE, Seifert SA, Valdez VA, Kottke RA, Rudolph JB, Bai G, Chen X, Bowden RL, Jin Y, Kolmer JA, Chen M, Seabourn BW, Dowell FE (2014) Registration of ‘Antero’ Wheat. J Plant Reg 8:165–168
Haley SD, Johnson JJ, Peairs FB, Stromberger JA, Hudson-Arns EE, Seifert SA, Anderson VA, Bai G, Chen X, Bowden RL, Jin Y, Kolmer JA, Chen M, Seabourn BW (2017) Registration of ‘Sunshine’ hard white winter wheat. J Plant Reg 11:289–294
Hareland GA (2003) Effects of pearling on falling number and alpha-amylase activity of preharvest sprouted spring wheat. Cereal Chem 80:232–237
Hickey LT, Lawson W, Arief VN, Fox G, Franckowiak J, Dieters MJ (2012) Grain dormancy QTL identified in a doubled haploid barley population derived from two non-dormant parents. Euphytica 188:113–122
Iehisa JC, Matsuura T, Mori IC, Takumi S (2014) Identification of quantitative trait locus for abscisic acid responsiveness on chromosome 5A and association with dehydration tolerance in common wheat seedlings. J Plant Physiol 171:25–34
Imtiaz M, Ogbonnaya FC, Oman J, van Ginkel M (2008) Characterization of quantitative trait loci controlling genetic variation for preharvest sprouting in synthetic backcross-derived wheat lines. Genetics 178:1725–1736
Jiménez N, Mares D, Mrva K, Lizana C, Contreras S, Schwembe AR (2017) Susceptibility to preharvest sprouting of Chilean and Australian elite cultivars of common wheat. Crop Sci 57:462–474
Kato K, Nakamura W, Tabiki T, Miura H, Sawada S (2001) Detection of loci controlling seed dormancy on group 4 chromosomes of wheat and comparative mapping with rice and barley genomes. Theor Appl Genet 102:980–985
King RW, Richards RA (1984) Water-uptake in relation to pre-harvest sprouting damage in wheat: ear characteristics. Aust J Agric Res 35:327–336
Kosambi DD (1943) The estimation of map distances from recombination values. Ann of Eugen 12:172–175
Kulwal PL, Kumar N, Gaur A, Khurana P, Khurana JP, Tyagi AK, Balyan HS, Gupta PK (2005) Mapping of a major QTL for pre-harvest sprouting tolerance on chromosome 3A in bread wheat. Theor Appl Genet 111:1052–1059
Kulwal P, Ishikawa G, Benscher D, Feng ZY, Yu LX, Jadhav A, Mehetre S, Sorrells ME (2012) Association mapping for pre-harvest sprouting resistance in white winter wheat. Theor Appl Genet 125:793–805
Kumar A, Kumar J, Singh R, Garg T, Chhuneja P, Balyan HS, Gupta PK (2009) QTL analysis for grain colour and pre-harvest sprouting in bread wheat. Plant Sci 177:114–122
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60
Lei L, Zhu XK, Wang SW, Zhu MR, Carver BF, Yan LL (2013) TaMFT-A1 is associated with seed germination sensitive to temperature in winter wheat. PLoS ONE 8:e73330
Li C, Ni P, Francki M, Hunter A, Zhang Y, Schibeci D, Li H, Tarr A, Wang J, Cakir M, Yu J, Bellgard M, Lance R, Appels R (2004) Genes controlling seed dormancy and pre-harvest sprouting in a rice-wheat-barley comparison. Funct Integr Genom 4:84–93
Lin M, Cai SB, Wang S, Liu SB, Zhang GR, Bai GH (2015) Genotyping-by-sequencing (GBS) identified SNP tightly linked to QTL for pre-harvest sprouting resistance. Theor Appl Genet 128:1385–1395
Lin M, Zhang DD, Liu SB, Zhang GR, Yu JM, Fritz AK, Bai GH (2016) Genome-wide association analysis on pre-harvest sprouting resistance and grain color in US winter wheat. BMC Genom 17:794. https://doi.org/10.1186/s12864-016-3148-6
Liu SB, Bai GH (2010) Dissection and fine mapping of a major QTL for preharvest sprouting resistance in white wheat Rio Blanco. Theor Appl Genet 121:1395–1404
Liu SB, Cai SB, Graybosch R, Chen CX, Bai GH (2008) Quantitative trait loci for resistance to pre-harvest sprouting in US hard white winter wheat Rio Blanco. Theor Appl Genet 117:691–699
Liu SB, Bai GH, Cai SB, Chen CX (2011) Dissection of genetic components of preharvest sprouting resistance in white wheat. Mol Breed 27:511–523
Liu S, Sehgal SK, Li J, Lin M, Trick HN, Yu J, Gill BS, Bai G (2013) Cloning and characterization of a critical regulator for preharvest sprouting in wheat. Genetics 195:263–273
Liu SB, Sehgal SK, Lin M, Li JR, Trick HN, Gill BS, Bai GH (2015) Independent mis-splicing mutations in TaPHS1 causing loss of preharvest sprouting (PHS) resistance during wheat domestication. New Phytol 208:928–935
Lohwasser U, Arif MAR, Borner A (2013) Discovery of loci determining pre-harvest sprouting and dormancy in wheat and barley applying segregation and association mapping. Biol Plant 57:663–674
Mares DJ, Mrva K (2001) Mapping quantitative trait loci associated with variation in grain dormancy in Australian wheat. Aust J Agric Res 52:1257–1265
Mares DJ, Mrva K (2014) Wheat grain preharvest sprouting and late maturity alpha-amylase. Planta 240:1167–1178
Mares D, Mrva K, Cheong J, Williams K, Watson B, Storlie E, Sutherland M, Zou Y (2005) A QTL located on chromosome 4A associated with dormancy in white- and red-grained wheats of diverse origin. Theor Appl Genet 111:1357–1364
Mares D, Rathjen J, Mrva K, Cheong J (2009) Genetic and environmental control of dormancy in white-grained wheat (Triticum aestivum L.). Euphytica 168:311–318
Martin TJ, Sears RG, Seifers DL, Harvey TL, Witt MD, Schlegel AJ, McCluskey PJ, Hatchett JH (2001) Registration of ‘Trego’ wheat. Crop Sci 41:929–930
Martin TJ, Zhang G, Fritz AK, Miller R, Chen M (2013) Registration of ‘Tiger’ wheat. J Plant Reg 7:201–204
Martin TJ, Zhang G, Fritz AK, Miller R, Chen M (2014) Registration of ‘Clara CL’ wheat. J Plant Reg 8:38–42
Mayer KFX, Rogers J, Dolezel J, Pozniak C, Eversole K, Feuillet C, Gill B, Friebe B, Lukaszewski AJ, Sourdille P, Endo TR, Dolezel J, Kubalakova M, Cihalikova J, Dubska Z, Vrana J, Sperkova R, Simkova H, Rogers J, Febrer M, Clissold L, McLay K, Singh K, Chhuneja P, Singh NK, Khurana J, Akhunov E, Choulet F, Sourdille P, Feuillet C, Alberti A, Barbe V, Wincker P, Kanamori H, Kobayashi F, Itoh T, Matsumoto T, Sakai H, Tanaka T, Wu JZ, Ogihara Y, Handa H, Pozniak C, Maclachlan PR, Sharpe A, Klassen D, Edwards D, Batley J, Olsen OA, Sandve SR, Lien S, Steuernagel B, Wulff B, Caccamo M, Ayling S, Ramirez-Gonzalez RH, Clavijo BJ, Steuernagel B, Wright J, Pfeifer M, Spannagl M, Mayer KFX, Martis MM, Akhunov E, Choulet F, Mayer KFX, Mascher M, Chapman J, Poland JA, Scholz U, Barry K, Waugh R, Rokhsar DS, Muehlbauer GJ, Stein N, Gundlach H, Zytnicki M, Jamilloux V, Quesneville H, Wicker T, Mayer KFX, Faccioli P, Colaiacovo M, Pfeifer M, Stanca AM, Budak H, Cattivelli L, Glover N, Martis MM, Choulet F, Feuillet C, Mayer KFX, Pfeifer M, Pingault L, Mayer KFX, Paux E, Spannagl M, Sharma S, Mayer KFX, Pozniak C, Appels R, Bellgard M, Chapman B, Pfeifer M, Pfeifer M, Sandve SR, Nussbaumer T, Bader KC, Choulet F, Feuillet C, Mayer KFX, Akhunov E, Paux E, Rimbert H, Wang SC, Poland JA, Knox R, Kilian A, Pozniak C, Alaux M, Alfama F, Couderc L, Jamilloux V, Guilhot N, Viseux C, Loaec M, Quesneville H, Rogers J, Dolezel J, Eversole K, Feuillet C, Keller B, Mayer KFX, Olsen OA, Praud S, IWGSC (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345(6194):1251788
Miao XL, Zhang YJ, Xia XC, He ZH, Zhang Y, Yan J, Chen XM (2013) Mapping quantitative trait loci for pre-harvest sprouting resistance in white-grained winter wheat line CA 0431. Crop Pasture Sci 64:573–579
Miura H, Sato N, Kato K, Amano Y (2002) Detection of chromosomes carrying genes for seed dormancy of wheat using the backcross reciprocal monosomic method. Plant Breed 121:394–399
Mohan A, Kulwal P, Singh R, Kumar V, Mir RR, Kumar J, Prasad M, Balyan HS, Gupta PK (2009) Genome-wide QTL analysis for pre-harvest sprouting tolerance in bread wheat. Euphytica 168:319–329
Mori M, Uchino N, Chono M, Kato K, Miura H (2005) Mapping QTLs for grain dormancy on wheat chromosome 3A and the group 4 chromosomes, and their combined effect. Theor Appl Genet 110:1315–1323
Nakamura S, Chono M, Abe F, Miura H (2010) Mapping a diploid wheat abscisic acid 8′-hydroxylase homologue in the seed dormancy QTL region on chromosome 5Am. Euphytica 171:111–120
Nakamura S, Abe F, Kawahigashi H, Nakazono K, Tagiri A, Matsumoto T, Utsugi S, Ogawa T, Handa H, Ishida H, Mori M, Kawaura K, Ogihara Y, Miura H (2011) A wheat homolog of MOTHER OF FT AND TFL1 Acts in the regulation of germination. Plant Cell 23:3215–3229
Paterson AH, Sorrells ME (1990) Inheritance of grain dormancy in white-kernelled wheat. Crop Sci 30:25–30
Pieterse CMJ, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SCM (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 7:e32253
Ramirez-Gonzalez RH, Uauy C, Caccamo M (2015) PolyMarker: a fast polyploid primer design pipeline. Bioinformatics 31:2038–2039
Rasul G, Humphreys DG, Brule-Babel A, McCartney CA, Knox RE, DePauw RM, Somers DJ (2009) Mapping QTLs for pre-harvest sprouting traits in the spring wheat cross ‘RL4452/AC Domain’. Euphytica 168:363–378
Roy JK, Prasad M, Varshney RK, Balyan HS, Blake TK, Dhaliwal HS, Singh H, Edwards KJ, Gupta PK (1999) Identification of a microsatellite on chromosomes 6B and a STS on 7D of bread wheat showing an association with preharvest sprouting tolerance. Theor Appl Genet 99:336–340
Shorinola O, Bird N, Simmonds J, Berry S, Henriksson T, Jack P, Werner P, Gerjets T, Scholefield D, Balcarkova B, Valarik M, Holdsworth MJ, Flintham J, Uauy C (2016) The wheat Phs-A1 pre-harvest sprouting resistance locus delays the rate of seed dormancy loss and maps 0.3 cM distal to the PM19 genes in UK germplasm. J Exp Bot 67:4169–4178
Silva LC, Wang S, Zeng ZB (2012) Composite interval mapping and multiple interval mapping: procedures and guidelines for using Windows QTL Cartographer. Methods Mol Biol 871:75–119
Skubacz A, Daszkowska-Golec A, Szarejko L (2016) The role and regulation of ABI5 (ABA-Insensitive 5) in plant development, abiotic stress responses and phytohormone crosstalk. Front Plant Sci 7:1884. https://doi.org/10.3389/fpls.2016.01884
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Steber CM, Carter AH, and Pumphrey MO (2014) Reduce falling numbers risk. http://wagrains.org/on-going-falling-numbers-reseach. Accessed 28 June 2017
Toojinda T, Baird E, Booth A, Broers L, Hayes P, Powell W, Thomas W, Vivar H, Young G (1998) Introgression of quantitative trait loci (QTLs) determining stripe rust resistance in barley: an example of marker-assisted line development. Theor Appl Genet 96:123–131
Torada A, Koike M, Ogawa T, Takenouchi Y, Tadamura K, Wu JZ, Matsumoto T, Kawaura K, Ogihara Y (2016) A causal gene for seed dormancy on wheat chromosome 4A encodes a MAP kinase kinase. Curr Biol 26:782–787
Van Ooijen JW (2006) JoinMap® 4, software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen
Verslues PE, Zhu JK (2005) Before and beyond ABA: upstream sensing and internal signals that determine ABA accumulation and response under abiotic stress. Biochem Soc Trans 33:375–379
Wang S, Basten CJ, Zeng ZB (2007) Windows QTL Cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh, NC. http://statgen.ncsu.edu/qtlcart/WQTLCart.htm
Wang L, Cheng JP, Lai YY, Du WL, Huang X, Wang ZF, Zhang HS (2014) Identification of QTLs with additive, epistatic and QTL × development interaction effects for seed dormancy in rice. Planta 239:411–420
Xiao SH, Zhang HP, You GX, Zhang XY, Yan CS, Chen X (2012) Integration of marker-assisted selection for resistance to pre-harvest sprouting with selection for grain-filling rate in breeding of white-kernelled wheat for the Chinese environment. Euphytica 188:85–88
Yang Y, Ma YZ, Xu ZS, Chen XM, He ZH, Yu Z, Wilkinson M, Jones HD, Shewry PR, Xia LQ (2007a) Isolation and characterization of Viviparous-1 genes in wheat cultivars with distinct ABA sensitivity and pre-harvest sprouting tolerance. J Exp Bot 58:2863–2871
Yang Y, Zhao XL, Xia LQ, Chen XM, Xia XC, Yu Z, He ZH, Roeder M (2007b) Development and validation of a Viviparous-1 STS marker for pre-harvest sprouting tolerance in Chinese wheats. Theor Appl Genet 115:971–980
Zanetti S, Winzeler M, Keller M, Keller B, Messmer M (2000) Genetic analysis of pre-harvest sprouting resistance in a wheat × spelt cross. Crop Sci 40:1406–1417
Zhu YL, Wang SX, Zhang HP, Zhao LX, Wu ZY, Jiang H, Cao JJ, Liu K, Qin M, Lu J, Sun GL, Xia XC, Chang C, Ma CX (2016) Identification of major loci for seed dormancy at different post-ripening stages after harvest and validation of a novel locus on chromosome 2AL in common wheat. Mol Breed 36:174. https://doi.org/10.1007/s11032-016-0598-0
Acknowledgements
Contribution number from the Kansas Agricultural Experiment Station is 18-034-J. This project was financially supported by the Kansas Wheat Alliance and Kansas Wheat Commission. This work was also partly funded by the Hatch Grant 1001453 and National Research Initiative Competitive Grants 2011-68002-30029, 2017-67007-25939 and 2017-67007-25929 from the USDA National Institute of Food and Agriculture. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. USDA is an equal opportunity provider and employer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Communicated by Mark E. Sorrells.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shao, M., Bai, G., Rife, T.W. et al. QTL mapping of pre-harvest sprouting resistance in a white wheat cultivar Danby. Theor Appl Genet 131, 1683–1697 (2018). https://doi.org/10.1007/s00122-018-3107-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3107-5