Abstract
Key message
Analysis of 387 sugarcane clones using Bru 1 diagnostic markers revealed two possible sources of Bru 1 in Chinese cultivars: one from Saccharum spontaneum and another from Saccharum robustum of New Guinea.
Abstract
Sugarcane brown rust (SBR) is an important fungal disease in many sugarcane production areas around the world, and can cause considerable yield losses in susceptible sugarcane cultivars. One major SBR resistance gene, named Bru1, initially identified from cultivar R570, was shown to be a major SBR resistance source in most of the sugarcane producing areas of the world. In this study, by using the two Bru1-associated markers, R12H16 and 9O20-F4, we surveyed the presence of Bru1 in a Chinese sugarcane germplasm collection of 387 clones, consisting of 228 hybrid cultivars bred by different Chinese sugarcane breeding establishments, 54 exotic hybrid cultivars introduced from other countries and 105 clones of sugarcane ancestral species. The Bru1-bearing haplotype was detected in 43.4% of Chinese sugarcane cultivars, 20.4% of exotic hybrid cultivars, and only 3.8% of ancestral species. Among the 33 Chinese cultivars for which phenotypes of resistance to SBR were available, Bru1 was present in 69.2% (18/26) of the resistant clones. Analyses of the allelic sequence variations of R12H16 and 9O20-F4 suggested two possible sources of Bru1 in Chinese cultivars: one from S. spontaneum and another from S. robustum of New Guinea. In addition, we developed an improved Bru1 diagnostic marker, 9O20-F4-HaeIII, which can eliminate all the false results of 9O20-F4-RsaI observed among S. spontaneum, as well as a new dominant Bru1 diagnostic marker, R12E03-2, from the BAC ShCIR12E03. Our results provide valuable information for further efforts of breeding SBR-resistant varieties, searching new SBR resistance sources and cloning of Bru1 in sugarcane.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane is an economically important crop which accounts for as high as 79% of the world’s sugar production (Liu et al. 2015) and up to 90% of that in China (Que et al. 2014; Wei et al. 2015). It is a high-biomass perennial C4 grass and can also be used as renewable sources of raw materials for ethanol biofuel production and other industrial utilizations (bagasse, pulp, molasses, rum, etc.) (Canilha et al. 2012; Chandel et al. 2012). Sugarcane belongs to the genus Saccharum of the family Poaceae (Gramineae) within the tribe Andropogoneae and subtribe Saccharinae. The genus is widespread across tropical, subtropical and warm temperate regions, and comprises six commonly recognized species: Saccharum spontaneum L., Saccharum robustum Brandes and Jeswiet ex Grassl, Saccharum officinarum L., Saccharum barberi Jesw., Saccharum sinense Roxb. and Saccharum edule Hassk. (Daniels and Roach 1987). Saccharum spontaneum is a wild species with the largest geographic distribution, from Japan and New Guinea to the Mediterranean and Africa, with India as its probable center of origin (Panje and Babu 1960; Daniels et al. 1975). It encompasses the most diverse euploid and aneuploid forms of the genus with 2n = 40–128 (Panje and Babu 1960; Lu et al. 1994a). Saccharum robustum is another wild species indigenous to New Guinea and the adjacent islands of Melanesia. It encompasses both the euploid (2n = 60, 80) and aneuploid (2n = 63–200) forms and is thought to be evolved from introgression of S. spontaneum with other wild relatives (Daniels and Roach 1987). Saccharum officinarum is a cultivated species (the archetypal sweet chewing ‘noble’ canes) indigenous to New Guinea and thought to be derived from S. robustum 2n = 80 euploid form. Saccharum barberi is a cultivated ‘species’ indigenous to India with 2n = 81–124 aneuploids, while S. sinensis is a cultivated ‘species’ indigenous to India with 2n = 111–120 aneuploids. Both are thought to be derived from natural hybridization between S. officinarum (introgressed from New Guinea) and S. spontaneum (local forms) in China and India, respectively (Price 1968; Daniels and Daniels 1975; D’Hont et al. 2002). Saccharum edule is a marginal ‘species’ indigenous to Melanesia and Polynesia where it was cultivated as a traditional vegetable (edible aborted flower). It is sterile with 2n = 60, 70 or 80 and was suggested to be derived from S. robustum by mutations (Grivet et al. 2005). The general relationships between the different Saccharum species in sugarcane have been confirmed by the polymorphisms of nuclear (Glaszmann et al. 1990; Lu et al. 1994a) and cytoplasmic DNA (D’Hont et al. 1993). Despite of their complex genomic background, the Saccharum species (except S. edule) are general male and female fertile. Saccharum and its allied genera Erianthus sect. Ripidium, Miscanthus sect. Diandra, Narenga Bor and Sclerostachya (Hack.) A. Camus together constitute an inter-crossable interbreeding group which is termed ‘Saccharum complex’, of which the members are thought to have evolved through natural hybridization and polyploidization events (Mukherjee 1957; Daniels and Roach 1987; Amalraj and Balasundaram 2006) although this has been discussed by Grivet et al. (2005).
Sugarcane is a very successful example in the use of alien genetic resources. The discovery of sexual fertility in sugarcane in 1888 in Java, and the ravages of the ‘sereh’ disease at the same period, had stimulated the artificial hybridizations between S. officinarum and S. spontaneum clones by the breeders in India and Indonesia. The hybrid progenies had been repeatedly backcrossed with S. officinarum to minimize the negative effect of the wild parent, a procedure known as ‘nobilization’ in sugarcane (Bremer 1961a; Roach 1972). For the first and second nobilization generations, the female parent (S. officinarum) and the male parent (S. spontaneum or F1 hybrid) transmit 2n and n chromosomes, respectively, to their progenies. It is from the third nobilization generation (BC2) that the chromosomal transmission becomes normal n + n (Bremer 1961b; Bhatt and Gill 1985). So, modern sugarcane cultivars are often aneuploid and highly polyploid with 2n = 100–130, of which 80–90% came from S. officinarum and 10–20% from S. spontaneum (Lu et al. 1994b; D’Hont et al. 1996; Piperidis et al. 2010). One nobilized cultivar, POJ2878, showed such a success that in 1929 it occupied 90% of the sugarcane cultivated area in Java and was introduced to most sugarcane research stations of the world. Until recent years, sugarcane breeding had been essentially limited to intercross certain nobilized hybrids derived from a very small number of S. spontaneum parents and select the elite clones among the produced progeny (Bremer 1961a; Zhang et al. 2009; Zhou 2013; Luo et al. 2014; Todd et al. 2015).
Sugarcane brown rust (SBR) is currently one of the most damaging fungal diseases in many sugarcane production areas around the world (Dean et al. 1979; Comstock et al. 2008; Hoy and Hollier 2009; Pocovi et al. 2010; Wang et al. 2013; Peixoto-Junior et al. 2014; Li et al. 2017). It is caused by the fungus Puccinia melanocephala H. & P. Sydow, of which the windblown spores can infect the plant leaf tissue and cause reddish-brown lesions resulting in the reduction of photosynthesis and plant growth rates. SBR can cause considerable yield losses of as high as 50% in susceptible cultivars (Purdy et al. 1983; Comstock et al. 1992; Hoy and Hollier 2009). In China, the occurrence of SBR was first reported in Yunnan province in 1982, and subsequently in other sugarcane growing provinces such as Fujian, Guangdong, Sichuan, Jiangxi, Hainan, etc., where the SBR caused severe losses in yield and sucrose by 10–40% (Ruan et al. 1983; Wang et al. 2013; Li et al. 2017). Several widely cultivated Chinese elite (high-yield and high-sucrose content) sugarcane cultivars risked to be eliminated by their susceptibility to SBR, threatening the stability and sustainability of the sugarcane industry in China (Li et al. 2017).
The best strategy to manage SBR is the development of resistant varieties. Daugrois et al. (1996) identified, for the first time, a major dominant SBR resistance gene, named Bru1, in analyzing the self-progeny of sugarcane cultivar ‘R570’. Further study showed that Bru1 conferred a wide and durable resistance to various SBR isolates collected from different geographic origins (Asnaghi et al. 2001). Molecular markers tightly linked to Bru1 have been developed by both genetic and physical mapping approaches (Asnaghi et al. 2000, 2004; Le Cunff et al. 2008). Haplotype-specific chromosome walking using a Bru1-enriched BAC library revealed the presence of an insertion of unknown size on the Bru1-bearing haplotype in comparison with other hom(e)ologous haplotypes at the Bru1 locus (Le Cunff et al. 2008). Comparative mapping of same set of markers surrounding the insertion on different hom(e)ologous haplotypes of Bru1 locus showed that the insertion induced a reduction of recombination on the Bru1-bearing haplotype, which is probably the cause of the strong linkage disequilibrium (LD) observed in the Bru1 region among modern sugarcane cultivars (Costet et al. 2012). Two PCR markers, one named R12H16 which is located in the insertion and another named 9O20-F4 which flanks the insertion, were found completely associated with each another and only in SBR-resistant self-progeny or sugarcane cultivars, constituting then efficient diagnostic markers for the presence of Bru1 (Le Cunff et al. 2008; Costet et al. 2012). These two PCR markers have been successfully used for the detection of Bru1 and prediction of SBR-resistant phenotype in sugarcane germplasm collected in different countries or regions in the world. By using these two markers, Costet et al. (2012) analyzed 380 modern sugarcane cultivars and breeding materials covering the worldwide diversity that were maintained by CIRAD in Reunion and Guadeloupe, and found that Bru1 was present in 47% of the total tested clones corresponding to 86% of the resistant clones; Glynn et al. (2013) analyzed 1072 parental clones and 1527 genotypes of a CP breeding program maintained at the USDA-ARS sugarcane breeding station in Florida, and found that Bru1 was present in 27% of parental clones and 44% of genotypes in CP program, corresponding to 59 and 83% of the resistant clones in the two cases, respectively; Racedo et al. (2013) analyzed 319 sugarcane clones maintained by EEAOC (Estación Experimental Agroindustrial Obispo Colombres) at Tucuman in Argentina, and found that Bru1 was present in only 6.6% of the tested clones corresponding to 16.3% of resistant clones; Parco et al. (2014) reported that Bru1 was present in only 4.3% of the sugarcane cultivars and elite breeding clones maintained at the USDA-ARS Sugarcane Research Unit in Louisiana, corresponding to 12.5% of the resistant clones; Li et al. (2017) analyzed 101 common sugarcane breeding parents maintained by CNNSGR (Chinese National Nursery of Sugarcane Germplasm Resources) and found that Bru1 was present in 47.5% of tested clones corresponding to 62.3% of resistant clones. The absence of Bru1 in some of the resistant sugarcane clones indicated the presence of other sources of resistance genes (e.g., Bru2, Raboin et al. 2006) which could be useful for broadening the SBR resistance basis in modern sugarcane breeding programs.
The objectives of this study were to (1) survey the presence of the Bru1-bearing haplotype (chromosome segment), using available R12H16 and 9O20-F4 markers and new or improved diagnostic PCR markers for Bru1, in a Chinese sugarcane germplasm collection of 387 clones, consisting of 228 hybrid cultivars bred by different Chinese sugarcane breeding establishments, 54 exotic hybrid cultivars introduced from other countries of the world and 105 clones of sugarcane ancestral species; (2) analyze the allelic sequence variations of R12H16 and 9O20-F4 markers to investigate the origin of Bru1.
Materials and methods
Plant materials
The plants used in this study were divided into three panels. The first panel consisted of 228 sugarcane hybrid cultivars bred by different Chinese sugarcane breeding establishments situated in different sugarcane production regions (e.g., Fujian, Guangdong, Guangxi, Hainan, Jiangxi, Sichuan, Yunnan and Taiwan) of China (Table S1). The second panel consisted of 54 exotic sugarcane hybrid cultivars, of which 34 from USA, 7 from India, 3 from Indonesia/New Guinea, 2 from Thailand, 2 from Australia, 2 from Brazil, 2 from South Africa, 1 from France and 1 from Mexico (Table S2). The third panel consisted of 105 clones of sugarcane ancestral species, including 74 clones of S. spontaneum, 7 clones of S. robustum, 3 clones of S. barberi, 3 clones of S. sinense, 7 clones of S. officinarum, 3 clones of Erianthus arundinaceus Retz., 1 clone of Erianthus fulvus Nees ex Hack., 3 clones of Erianthus rockii Keng, 1 clone of Miscanthus floridulus (Labill.) Warb. ex Schum. & Laut., 1 clone of Miscanthus sinensis Anderss., and 2 clones of Narenga porphyrocoma (Hance ex Trin.) Bor (Table S3). Of the 74 clones of S. spontaneum, 69 are from different regions of China and 5 from India. Of the 7 clones of S. robustum, 5 are from Indonesia/New Guinea and 2 from China. The three clones of S. barberi are from India, while those of S. sinense are from China. The seven clones of S. officinarum are all from Indonesia/New Guinea. The clones of Erianthus, Miscanthus and Narenga are all indigenous to China. These plants were grown either in the Nursery of Sugarcane Germplasm Resources of Sugarcane or Research Institute of Fujian Agriculture and Forestry University (Fuzhou, Fujian Province, China) or in the National Nursery of Sugarcane Germplasm Resources (NNSGR) of Sugarcane Research Institute of Yunnan Academy of Agricultural Sciences (Kaiyuan, Yunan Province, China).
Phenotypic data of brown rust resistance
After carefully checking the conformance of nomenclature and Bru1 diagnostic marker response, we identified 21 exotic cultivars and 33 Chinese cultivars that have been previously phenotyped for resistance to brown rust in the studies of Costet et al. (2012), Racedo et al. (2013), Parco et al. (2014) and Li et al. (2017). These cultivars were phenotyped as HR (highly resistant), R (resistant), MR (moderately resistant), MS (moderately susceptible), S (susceptible) or HS (highly susceptible) in Tables S1–2 following the same criteria that were described in their previous studies.
Genomic DNA extraction
For each studied sugarcane clone, fully expanded young leaf tissues were collected directly from field-grown plants, numbered and stored temporarily at 4 °C just before DNA extraction. Total genomic DNA was isolated from about 100 mg of leaf tissue using a simplified CTAB method (Doyle and Doyle 1990). Quality and quantity of DNA was determined using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific Inc, Wilmington, DE, USA). According to the measured values, each extracted DNA sample was diluted to a concentration of 25 ng/µL in TE buffer and stored at −20 °C prior to PCR reaction use.
PCR detection of Bru1-bearing chromosomal segment
We followed the methods of Costet et al. (2012) for PCR detection of Bru1-bearing chromosomal segment. The primers for the two Bru1-associated markers, R12H16 (Fw: 5′-CTA CGA TGA AAC TAC ACC CTT GTC-3′, Rv: 5′-CTT ATG TTA GCG TGA CCT ATG GTC-3′) and 9O20-F4 (Fw: 5′-TAC ATA ATT TTA GTG GCA CTC AGC-3′, Rv: 5′-ACC ATA ATT CAA TTC TGC AGG TAC-3′), were synthesized by Sangon Biotech Co. Ltd. (Shanghai, China), and diluted to a concentration of 10 μM in TE buffer. PCR reactions were carried out on a Bio-Rad S1000 Thermal Cycler with a reaction volume of 20 μL including 2 μL of DNA template, 1 μL of forward primer (10 μM), 1 μL of forward primer (10 μM), 10 μL 2xTaq Plus PCR Master Mix (Tiangen Biotech Co. Ltd., Beijing, China) and 6 μL sterile distilled water. The PCR profile used was as following: one step of 94 °C for 4 min followed by 35 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s, and a final step at 72 °C for 5 min. Ten microlitres of PCR product of marker 9O20-F4 was digested with RsaI in a total volume of 20 μL. R12H16 amplicons and 9O20-F4/RsaI restriction fragments were resolved on 2% agarose gels in 1X TAE buffer and stained with 1X GoldView (Yeasen, Shanghai, China) for visualization and analysis in a Molecular Imager Gel Doc™ XR + System (Bio-Rad, USA). By analysis in silico of the amplicon sequences of 9O20-F4 from various clones (see below), we identified another restriction enzyme, named HaeIII, which can result in a new Bru1 diagnostic band of 389 bp by digesting the PCR products of 9O20-F4. The experimental procedure for 9O20-F4/HaeIII is the same as for 9O20-F4/RsaI.
Cloning and sequencing of PCR products of R12H16 and 9O20-F4
PCR amplification products of R12H16 and 9O20-F4 from 10 sugarcane hybrids, 2 S. sinensis, 1 S. barberi, 1 S. robustum, 9 S. spontaneum and 1 E. arundinaceus (selected according to their response to the two diagnostic markers see “Results”) were directly cloned into the pMD™18-T Vector using a TA Cloning Kit (Takara Biotech Co. Ltd., Dalian, China). After 24 h of incubation at 37 °C, individual bacterial colonies were picked into 1.5-mL Eppendorf tubes containing 500 µL of LB medium + 50 μg/mL of ampicillin, which were further incubated overnight at 37 °C and shaken at 220 rpm. Screening of positive bacterial clones with inserts was performed using M13-specific primers (Fw: 5′-CR12H16AC GAC GTT GTA AAA CGA C-3′, Rv: 5′-GGA TAA CAA TTT CAC ACA GG-3′) by PCR reaction in a 10-μL volume including 1 μL of bacterial culture, 0.25 μL of forward primer (10 μM), 0.25 μL of forward primer (10 μM), 5 μL 2xTaq Plus PCR Master Mix (Tiangen Biotech Co. Ltd., Beijing, China) and 3.5 μL sterile distilled water. For the cloning of 9O20-F4 PCR products of a few selected sugarcane genotypes, the positive PCR products of M13-specific primers from different bacterial clones were further submitted to RsaI digestion followed by electrophoresis in 2% agarose gel. The RsaI restriction patterns were used as references to help to select a maximum of haplotypes from the positive bacterial clones for sequencing. A sample of 50 μL of culture solution of each selected positive bacterial clone was sent to Sangon Biotech Co. Ltd. (Shanghai, China) for insert DNA sequencing by an ABI 3730XL DNA Analyzer. Three to twelve bacterial clones per each sugarcane clone were sequenced.
Multiple sequence alignment, phylogenetic tree construction and analysis of RFLP in silico
Multiple sequence alignment was performed using Clustal W program (Larkin et al. 2007) and edited with BioEdit software (Hall 1999) using default parameters in each case. Phylogenetic trees were generated with MEGA6.06 (Tamura et al. 2013) using the neighbor-joining (NJ) algorithm. Bootstrap analysis with 1000 replicates was used to evaluate the significance of the nodes. Pairwise gap deletion mode was used to ensure that the divergent domains could contribute to the topology of the NJ tree. Nucleotide homology searches were performed on NCBI using the BLASTn module. Analysis of restriction fragment length polymorphism (RFLP) in silico of 9O20-F4 amplicon sequences was performed using the Restriction Map function of BioEdit software.
Design of new gene-specific primers from BAC sequence of ShCIR12E03 and PCR analysis
BAC sequence of ShCIR12E03 was downloaded from the NCBI databases (FN431661.1). The two first genes on the BAC, one positioned from 1 to 2745 and predicted as a conserved hypothetical protein (named R12E03-1), another positioned from 12,457 to 14,685 (complement) and predicted as a zinc knuckle family protein (named R12E03-2), were both located in the insertion region (on the right extremity) of Bru1 locus (Le Cunff et al. 2008; Garsmeur et al. 2011). Gene-specific primers were designed using online Primer3Plus software (http://www.primer3plus.com) for gene R12E03-1 (Fw: 5′-TTG ATG AAC AAC AGT AAG TTC GTG-3′, Rv: 5′-ATA CAA ACA ACG GTT ACA GCA CAC-3′, product size = 600 bp) and R12E03-2 (Fw: 5′-GAT CAG ACA GCT CTT TCT CTT GAG-3′, Rv: 5′-CAA GTA TTA CAA TCG GTC TGG GTT-3′, product size = 589 bp), respectively. The PCR conditions and analysis methods used for these two pairs of primers were similar to those for R12H16 and 9O20-F4 except for the annealing temperature at 56 °C.
Results
Detection of Bru1-bearing haplotype among different panels of sugarcane germplasm
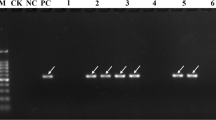
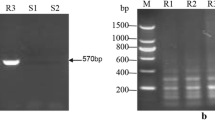
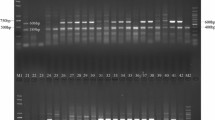
The presence of Bru1-bearing haplotype was diagnosticated by a 569-bp PCR product of marker R12H16 (Fig. 1a) or a 191-bp fragment produced after digestion of the PCR product of marker 9O24-F4 with RsaI (Fig. 1b, c). These two diagnostic bands were simultaneously detected in 99 of 228 (43.4%) Chinese sugarcane cultivars, 11 of 54 (20.4%) exotic sugarcane clones, and 4 of 105 (3.8%) ancestral species clones (Table S1–S3). Among the 228 Chinese sugarcane cultivars, Bru1 was steadily detected in those from Guangdong (18/54), Guangxi (24/42), Hainan (8/36), Fujian (17/32), Yunnan (15/29), Taiwan (11/17) and Jiangxi (6/8), excepting for Sichuan (0/10) (Table 1). For the 54 exotic sugarcane cultivars, Bru1 was detected in those from USA (3/34), India (1/7), Indonesia/New Guinea (3/3), Thailand (1/2), Australia (1/2), France (1/1) and Mexico (1/1), excepting for Brazil (0/2) and South Africa (0/2) (Table 2). The data of phenotypes of resistance to SBR were available for 33 Chinese sugarcane cultivars and 21 exotic cultivars (Table S1, 2), Bru1 was found to be inferred in 18/26 (69.2%) and 2/10 (20.0%) of the resistant clones in the two cases, respectively. For the panel of 105 ancestral species clones, 1 S. robustum (NG77-004), 1 S. barberi (Katha) and 2 S. sinensis (Songxi-zhuzhe and Youba) clones, were detected as positive simultaneously for R12H16 and 9O24-F4-RsaI (Table 3). All the seven S. officinarum clones were negative for the two diagnostic markers. However, in 11 of the 74 S. spontaneum clones, a fragment of size similar to the 191 bp diagnostic band of 9O24-F4-RsaI was detected, while the diagnostic 569 bp band of R12H16 was absent (Fig. 1c, Table S3). The three arundinaceus, one Erianthus fulvus, three Erianthus rockii, 1 Miscanthus floridulus, 1 Miscanthus sinensis, and two Narenga porphyrocoma clones were all negative for both the two diagnostic markers (Table S3).
Representative DNA profiles of PCR markers R12H16 (a), 9O20-F4 (b), 9O20-F4-RsaI (c), 9O20-F4-HaeIII (d), R12E03-1 (e) and R12E03-2 (f) in a subset of six sugarcane modern cultivars (lanes 1–6) and four S. spontaneum clones (lanes 7–10), separated by 2% agarose gel electrophoresis. M molecular weight marker, 1 FR93-435, 2 YZ91-510, 3 YZ03-194, 4 LC05-136, 5 YG35, 6 LC05-179, 7 FJ89-1-1, 8 YN-Heqing, 9 YN83-215, 10 GZ78-2-04. In each case, the Bru1-associated fragment is indicated by an arrow
Allelic variation of 9O24-F4
The marker 9O24-F4 amplified a mixture of DNA fragments which were more or less complex according to genotype (Fig. 1b). The PCR products of marker 9O24-F4 from 10 sugarcane hybrids (FR93-435, Kassoer, LC05-136, POJ2879, PT43-52, ROC22, YC05-179, YG35, YZ03-194 and YZ91-510), 2 S. sinensis (Songxi-zhuzhe and Youba), 1 S. barberi (Katha), 1 S. robustum (NG77-004), 9 S. spontaneum (FJ89-1-1, GD31, GD60, GZ78-1-31, GZ78-2-04, YN83-196, YN83-215, YN83-227 and YN-Heqing) and 1 E. arundinaceus (FJ-Banmao) were subjected to cloning and sequencing. A total of 59 amplicon sequences were obtained from the 24 genotypes above, with 1–7 amplicons per genotype (Fig. S1, Table S4). In addition, four amplicon sequences of the sugarcane cultivar R570 and 2 of the S. officinarum clone La Purple were obtained by in silico PCR from the NCBI databases. Figures S2 and S3 showed the alignments of seven amplicon sequences from POJ2878 and six from NG77-004, respectively, illustrating the complexity of the polyploid genome in sugarcane hybrids as well as in its wild relatives. The length of these amplicons varied from 214 to 947 bp. For each amplicon sequence, the most related BAC clone (with the highest sequence similarity) was identified by BLAST search against the NCBI data bases. The number of restriction sites and length of generated fragments by endonuclease RsaI and HaeIII were predicted by in silico analysis for each amplicon sequence and presented in Table S4. The Bru1-associated amplicons (389 bp in length) from different genotypes were identified by the presence of a 191-bp fragment after the digestion in silico by RsaI, as well as by their high sequence similarity (99–100%) to BAC clone ShCIR9O20. We noted that among the amplicons generated with S. spontaneum clone YN83-196, one had a size of 193 bp, and among the amplicons generated with S. spontaneum clones GZ78-2-04, YN83-215, YN83-227 and YN-Heqing, one had a size of 183-bp fragment upon RsaI digestion. Analysis of amplicon sequences showed that both 193- and 183-bp fragments were not associated with Bru1-haplotype, but these bands can be confused with the Bru1-associated 191-bp fragment in an agarose gel (Fig. 1c), and then give a false-positive result during the screening of Bru1 among S. spontaneum clones. This problem was resolved by application of restriction enzyme HaeIII instead of RsaI on the PCR products of 9O24-F4 which produced a new Bru1 diagnostic band of 389 bp (Fig. 1d), as the Bru1-associated amplicons do not possess a HaeIII cutting site while all other types of amplicons contain 1 or more restriction sites of HaeIII (Table S4).
Phylogenetic tree constructed on the basis of the alignment of 65 amplicon sequences of 9O24-F4 showed that the 14 191-bp Bru1-associated amplicons were clustered together and clearly separated from all other amplicon sequences (Fig. 2). The Bru1-associated amplicons identified from eight sugarcane hybrids (FR93-435, Kassoer, POJ2878, PT43-52, R570, ROC22, YZ03-194 and YZ91-510), 2 S. sinensis (Songxi-zhuzhe and Youba) and 1 S. barberi (Katha) have the same sequence with each other and differed from the two Bru1-associated amplicons of the S. robustum clone NG77-004 by a single SNP (Fig. 3a). These two NG77-004 Bru1-associated amplicons differed by two SNPs. Interestingly, the Bru1-associated amplicon from LC05-136 had a sequence identical to one of the two Bru1-associated amplicons identified from the S. robustum clone NG77-004.
Phylogenetic tree calculated by MEGA6.06 based on 65 amplicon sequences obtained by cloning and sequencing the PCR products of the diagnostic marker 9O20-F4 or by PCR in silico on 11 sugarcane cultivars (FR93-435, Kassoer, LC05-136, POJ2878, PT43-52, R570, ROC22, YC05-179, YG35, YZ03-194 and YZ91-510), 1 S. officinarum (La Purple), 1 S. barberi (Katha), 2 S. sinensis (Songxi-zhuzhe and Youba), 1 S. robustum (NG77-004), 8 S. spontaneum (GD31, GD60, GZ78-1-31, GZ78-2-04, YN83-196, YN83-215, YN83-227 and YN-Heqing), and 1 E. arundinaceus (FJ-Banmao)
Multiple sequence alignments of Bru1-associated amplicon sequences obtained by cloning and sequencing the PCR products of the diagnostic marker 9O20-F4 (a) and R12H16 (b) on nine sugarcane cultivars (FR93-435, Kassoer, LC05-136, POJ2878, PT43-52, ROC22, YC05-179, YZ03-194 and YZ91-510), 1 S. sinensis (Katha), 2 S. barberi (Songxi-zhuzhe and Youba) and 1 S. robustum clone (NG77-004). The amplicon sequences of R570 for the two markers were obtained by PCR in silico from the NCBI databases. The forward and reverse PCR primer sequences are framed, and the SNP positions are indicated by an arrow in the figure
Allelic variation of R12H16
The PCR products of marker R12H16 from eight sugarcane hybrids (FR93-435, Kassoer, LC05-136, POJ2878, PT43-52, ROC22, YZ03-194 and YZ91-510), 2 S. sinensis (Songxi-zhuzhe and Youba) and 1 S. barberi (Katha) and 1 S. robustum clone (NG77-004) were cloned and/or sequenced. Alignment of sequences showed that all the nine sugarcane hybrids (including R570), 2 S. sinensis and 1 S. barberi possessed an identical allele of 569 bp and differed from the S. robustum clone by a single nucleotide base substitution (Fig. 3b).
Development of new Bru1-associated markers from the insertion region of Bru1-bearing haplotype
According to the annotations and comparisons of hom(oe)ologous BAC sequence by Garsmeur et al. (2011), the two first annotated genes of the BAC clone ShCIR12E03, ShCIR12E03g_160 (conserved hypothetical protein) and ShCIR12E03g_180 (zinc knuckle family protein), were situated within the insertion region of the Bru1-bearing haplotype (CG VII-1) (Fig. S4). A new PCR-primer pair (marker R12E03-1), was developed from the sequence of ShCIR12E03g_160, and produced a monomorphic band of 600 bp among the sugarcane cultivars or wild relatives (Fig. 1e); while another new PCR-primer pair (marker R12E03-2), developed from the sequence of ShCIR12E03g_180, amplified a polymorphic band of 589 bp which was totally associated with the marker R12H16 (Fig. 1f). The result indicated that the gene ShCIR12E03g_160 is present in more than one copies in sugarcane hybrid cultivars while the gene ShCIR12E03g_180 is only present in one copy in the insertion region of Bru1-bearing haplotype chromosome. Blast search of these two genes against NCBI databases showed that both genes have their orthologs in multi-copies in the sorghum and rice genomes (Fig. S5).
Discussion
Modern sugarcane cultivars are highly polyploid and aneuploid interspecific hybrids, which are vegetatively propagated as clones by stem cuttings. The main components (80–90%) of the sugarcane cultivars genome (100–130 chromosomes) are derived from the domesticated sugar-producing species S. officinarum (2n = 8x = 80), while the a small portion (10–20%) is derived from the wild S. spontaneum species (2n = 5x ~ 16x = 40 ~ 128) by introgressions or ‘nobilizations’ (Sreenivasan et al. 1987; Lu et al. 1994b; D’Hont et al. 1996, 1998; Piperidis et al. 2010). But, it is this small portion of S. spontaneum chromosomes that has contributed many important agronomical traits to modern sugarcane cultivars, such as adaptation to environmental stresses, resistance to diseases and pests, growth vigor, ratooning performance, etc. (Panje 1972; Roach 1977). About 10% of the chromosomes of cultivar R570 (2n = 107–115) were identified as originating from S. spontaneum and about 10% were identified as recombinant chromosomes between the two species S. officinarum and S. spontaneum (D’Hont et al. 1996). Bru1 was mapped on one of the cosegregation groups (CGs) of the R570 homology group VII, named CG VII-1 (Daugrois et al. 1996; Asnaghi et al. 2000, 2004; Le Cunff et al. 2008), which was initially characterized as originated from S. officinarum by AFLP markers (Hoarau et al. 2001). Further studies revealed that this Bru1-bearing haplotype contained an insertion of unknown size in comparison with other hom(e)ologous haplotypes and probably originated by interspecific recombination between S. officinarum and S. spontaneum chromosomes (Le Cunff et al. 2008; Garsmeur et al. 2011; Zhang et al. 2016).
In this study, we firstly used the two Bru1-associated markers, R12H16 and 9O20-F4, to evaluate a Chinese sugarcane germplasm collection of 387 clones, consisting of 228 Chinese hybrid cultivars, 54 exotic cultivars and 105 clones of sugarcane ancestral species (Table S1–3). According to the double presence or not of the two markers, Bru1 was unambiguously detected in 43.4% of Chinese sugarcane cultivars bred by different Chinese sugarcane breeding stations (Tables 1, S1), indicating that Bru1 is widely distributed among the Chinese sugarcane cultivars. Bru1 was present in 69.2% of the resistant clone (18/26) among the 33 Chinese cultivars for which the phenotypes of resistance to SBR were available suggesting that it constitutes the main source of resistance to SBR in China. Our results are consistent with those of Li et al. (2017), who analyzed 101 common sugarcane breeding parents in China and found that Bru1 was detected in 47.5% of the tested clones and inferred in 62.3% of the brown rust-resistant clones. However, contradictory genotyping results were obtained for 7 of the 42 lines that were commonly analyzed in the two studies, indicating that possible errors occurred in the multiplication and maintenance of these sugarcane clones.
Bru1 diagnostic markers can be used to trace the origin of Bru1 among the wild sugarcane relatives. The identification of disjunction cases between the two diagnostic markers (that have never been reported among modern sugarcane cultivars) in wild relatives can help to understand the origin and evolution of Bru1 among Saccharum species. In this study, we analyzed a total of 74 S. spontaneum clones, of which 69 were from different provinces of China and 5 were from India (Table S3). All these 74 clones were negative for the dominant marker R12H16 (a band of 569 bp), while 11 Chinese S. spontaneum clones can each produce a band with a size similar to the diagnostic 191-bp band upon digestion of the 9O20-F4 PCR products by RsaI. A closer examination of the amplicon sequences showed that these bands (183 or 193 bp) were not associated with the Bru1 haplotype (Table S4), indicating that while 9O20-F4-RsaI was perfectly efficient for Bru1 detection among modern sugarcane cultivars, it can give false diagnostic results for Bru1 detection among S. spontaneum or other wild relatives. To solve this problem, we developed an improved Bru1 diagnostic marker, 9O20-F4-HaeIII, which can produce a band of 389 bp and eliminate all the false-positive records of 9O20-F4-RsaI among the wild S. spontaneum clones (Fig. 1d). This marker, together with R12H16 and R12E03-2, another newly developed dominant Bru1 diagnostic marker (Fig. 1f), will be very useful for tracing the Bru1-associated chromosomal segment among the ‘Saccharum complex’.
Bru1 was detected in 20.4% of the exotic hybrids introduced from various countries/regions of the world, such as USA, India, Indonesia/New, Thailand, Australia, France and Mexico (Tables 2, S2) and in 3.8 of the clones from ancestral species tested, one S. robustum, two S. sinense and one S. barberi. It worth noting that one of the oldest sugarcane cultivar, POJ2878, as well as its progenitor Kassoer, a supposed natural hybrid between the S. officinarum clone Black Cheribon and the S. spontaneum clone Glagah (Bremer 1961a, b), were both detected as Bru1-positive by the two diagnostic markers, while its other progenitors Bandjarmasin Hitam, Loethers, Black Cheribon, EK28 as well as other S. officinarum clones were all detected as Bru1-negative (Table S3). This suggests that Bru1 in POJ2878 could originate from the wild S. spontaneum Glagah from Java, which was unfortunately not available from NNSGR in China for testing in this study.
In China, POJ2878 was introduced from Philippine by Guangdong Provincial Bureau of Agriculture in 1933–1934 and then widely cultivated in south of China and Taiwan for many years (Peng 1980, 1996; Zhang 2003). As in other regions of the world, POJ2878 was also widely used as an elite genitor in various sugarcane breeding programs in China. In accordance with this reality, Zhang et al. (2009) reviewed the pedigree of 186 Chinese sugarcane cultivars and found that 184 of them could be traced back to POJ2878 while the two remaining cultivars were traced back to POJ2725, a full sib variety of POJ2878. Some of these 186 sugarcane cultivars were also included in the present study. From our analysis of amplicon sequences of Bru1 diagnostic markers, we believed that the Bru1 in Chinese sugarcane varieties was essentially introduced through a few hybrid clones like POJ2878.
Our analyses of allelic variation of 9O20-F4 and R12H16 markers showed that their Bru1-associated amplicons are highly conserved in sequence between the Bru1-positive Chinese cultivars, exotic cultivars, S. sinensis, S. barberi and S. robustum clones (Fig. 3), suggesting a common ancestor of Bru1 among Saccharum species. Interestingly, with 9O20-F4 marker, we identified a Bru1-associated amplicon in the Chinese cultivar LC05-136 that shares the same sequence with one of two alleles identified in the S. robustum clone NG77-004 (Fig. 3a). This result suggested that the Bru1 in Chinese sugarcane cultivars could also originate from S. robustum, a hypothesis supported by the fact that some S. robustum clones were used in the Chinese sugarcane breeding programs (Zhang 1996).
The fact that the two markers 9O20-F4 and R12H16 were simultaneously detected in a single wild relative such as the S. robustum clone NG77-004, the S. barberi clone Katha, or the S. sinensis clones Songxi-zhuzhe and Youba, indicates that the insertion event on Bru1-associated haplotype happened far before its introgression into the modern sugarcane cultivars, and was not simply a result of the ‘nobilizations’ of S. spontaneum or intercrossing between sugarcane hybrids. In this study, all the 74 S. spontaneum clones were characterized as Bru1 negative, but Parco et al. (2014) identified one (from 51) S. spontaneum clone MPTH97-204 (from Thailand) which showed to be positive for both R12H16 and 9O20-F4 markers. These results suggested that Bru1-associated haplotype is present at a very low frequency among the wild S. spontaneum clones.
Interestingly, one (of three) S. barberi and two (of three) S. sinense clones were shown to be positive for both R12H16 and 9O20-F4 markers (Table S3), and possessed identical amplicon sequences compared with those of POJ2878 (Fig. 3). Lu et al. (1994a) have reported that there existed certain RFLP markers displayed by S. barberi and S. sinense that were not found in either supposed parental species (S. officinarum and S. spontaneum), but were found in an E. arundinaceus clone (IK76-48). The restricted size of the sample representative of the ancestral species used and the possibility of homoplasy of markers restrict the possible inference from these observations. However, associated to the highly diverged alleles observed for genes 9 and 11a (Garsmeur et al. 2011; Zhang et al. 2016) as well as the 9O20-F4 marker (on gene 10) on Bru1-associated BACs (Figs. 2, S2–4), these results may suggest that the Bru1-bearing fragment (including the insertion + flanking regions) come from an uncharacterized part of the Saccharum genus or from other genera of ‘Saccharum complex’. In this study, we screened seven Erianthus, two Miscanthus and two Narenga clones that were all indigenous to the southern subtropical region of China (Table S3), but none of them showed to be positive for the two Bru1 diagnostic markers. Further studies on a larger and geographically more representative collection of Saccharum (especially S. spontaneum, including Glagah) and other genera of ‘Saccharum complex’ will allow to confirm this hypothesis.
In conclusion, we have surveyed the presence of Bru1 in a Chinese sugarcane germplasm collection of 387 clones by using the Bru1 diagnostic markers, R12H16 and 9O20-F4, and found that the Bru1-bearing haplotype was present in 43.4% of Chinese sugarcane cultivars, 20.4% of exotic hybrid cultivars, and only 3.8% of the clones of ancestral species. Analyses of the allelic sequence variations of R12H16 and 9O20-F4 suggested two possible sources of Bru1 in Chinese cultivars: one from S. spontaneum and another from S. robustum of New Guinea. We also found that 9O20-F4-RsaI could give false diagnostic results when it was applied in wild relatives such as S. spontaneum. Then we developed an improved Bru1 diagnostic marker, 9O20-F4-HaeIII, which can eliminate all the false results of 9O20-F4-RsaI observed among the clones of S. spontaneum. We also developed a new dominant Bru1 diagnostic marker, R12E03-2, from the Bru1-associated BAC ShCIR12E03. Our results provide valuable information for further efforts of elucidating the origin of Bru1 among the ‘Saccharum complex’, cloning of Bru1, searching new SBR resistance sources and breeding SBR-resistant varieties in sugarcane.
References
Amalraj VA, Balasundaram N (2006) On the taxonomy of the members of ‘Saccharum complex’. Genet Resour Crop Evol 53:35–41
Asnaghi C, Paulet F, Kaye C, Grivet L, Deu M, Glaszmann JC, D’Hont A (2000) Application of synteny across Poaceae to determine the map location of a sugarcane rust resistance gene. Theor Appl Genet 101:962–969
Asnaghi C, D’hont A, Glaszmann J, Rott P (2001) Resistance of sugarcane cultivar R 570 to Puccinia melanocephala isolates from different geographic locations. Plant Dis 85:282–286
Asnaghi C, Roques D, Ruffel S, Kaye C, Hoarau JY, Telismart H, Girard JC, Raboin LM, Risterucci AM, Grivet L, D’Hont A (2004) Targeted mapping of a sugarcane rust resistance gene (Bru1) using bulked segregant analysis and AFLP markers. Theor Appl Genet 108:759–764
Bhatt SR, Gill SS (1985) The implications of 2n egg gametes in nobilization and breeding of sugarcane. Euphytica 34:377–384
Bremer G (1961a) Problems in breeding and cytology of sugarcane. I. A short history of sugarcane breeding—the original forms of Saccharum. Euphytica 10(1):59–78
Bremer G (1961b) Problems in breeding and cytology of sugarcane. II. The sugar cane breeding from a cytological view-point. Euphytica 10(2):121–258
Canilha L, Chandel AK, Milessi TSS, Antunes FAF, Freitas WLC, Felipe MGA, Silva SS (2012) Bioconversion of sugarcane biomass into ethanol: an overview about composition, pretreatment methods, detoxification of hydrolysates, enzymatic saccharification and ethanol fermentation. J Biomed Biotechnol 8:989572
Chandel AK, Silva SS, Carvalho W, Singh OV (2012) Sugarcane bagasse and leaves: foreseeable biomass of biofuel and bio-products. J Chem Technol Biotechnol 87:1–20
Comstock JC, Shine J Jr, Raid RN (1992) Effect of rust on sugarcane growth and biomass. Plant Dis 76:175–177
Comstock JC, Sood SG, Glynn NC, McKemy Shine JJJ, Castlebury LA (2008) First report of Puccinia kuehnii, causal agent of orange rust of sugarcane, in the United States and Western Hemisphere. Plant Dis 92:175
Costet L, Le Cunff L, Royaert S, Raboin LM, Hervouet C, Toubi L, Telismart H, Garsmeur O, Rousselle Y, Pauquet J, Nibouche S, Glaszmann JC, Hoarau JY, D’Hont A (2012) Haplotype structure around Bru1 reveals a narrow genetic basis for brown rust resistance in modern sugarcane cultivars. Theor Appl Genet 125(5):825–836
D’Hont A, Grivet L, Feldmann P, Rao PS, Berding N (1996) Characterisation of the double genome structure of modern sugarcane cultivars (Saccharum spp) by molecular cytogenetics. Mol Gen Genet 250:405–413
D’Hont A, Ison D, Alix K, Roux C, Glaszmann JC (1998) Determination of basic chromosome numbers in the genus Saccharum by physical mapping of RNA genes. Genome 41:221–225
D’Hont A, Paulet F, Glaszmann JC (2002) Oligoclonal interspecific origin of ‘North Indian’ and ‘Chinese’ sugarcanes. Chromosome Res 10:253–262
Daniels J, Daniels CA (1975) Geographical, historical and cultural aspects of the origin of the Indian and Chinese sugarcanes S. barberi and S. sinense. Sugarcane Breed Newslett 36:4–23
Daniels J, Roach BT (1987) Taxonomy and evolution. In: Heinz DJ (ed) Sugarcane improvement through breeding. Elsevier Press, Amsterdam, pp 7–84
Daniels J, Smith P, Paton N, Williams CA (1975) The origin of the genus Saccharum. Sugarcane Breed Newsett 36:24–39
Daugrois JH, Grivet L, Roques D, Hoarau JY, Lombard H, Glaszmann JC, D’Hont A (1996) A putative major gene for rust resistance linked with a RFLP marker in sugarcane cultivar‘R570’. Theor Appl Genet 92:1059–1064
Dean JL, Tai PYP, Todd EH (1979) Sugarcane rust in Florida. Sugar J 42:10
D’Hont A, Lu YH, Feldmann P, Glaszmann JC (1993) Cytoplasmic diversity in sugarcane revealed by heterologous probes. Sugar Cane 1993(1):12–15
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Garsmeur O, Charron C, Bocs S, Jouffe V, Samain S, Couloux A, Droc G, Zini C, Glaszmann JC, Van Sluys MA, D’Hont A (2011) High homologous gene conservation despite extreme autopolyploid redundancy in sugarcane. New Phytol 189(2):629–642
Glaszmann JC, Lu YH, Lanaud C (1990) Variation of nuclear ribosomal DNA in sugarcane. J Genet Breed 44:191–198
Glynn NC, Laborde C, Davidson RW, Irey MS, Glaz B, D’Hont A, Comstock JC (2013) Utilization of a major brown rust resistance gene in sugarcane breeding. Mol Breed 31(2):323–331
Grivet L, Glaszmann JC, D’Hont A (2005) Molecular evidence of sugarcane evolution and domestication. In: Motley TJ, Zerega N, Cross H (eds) Darwin’s harvest: new approaches to the origins, evolution, and conservation of crops. Columbia University Press, New York, pp 49–66
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–99
Hoarau JY, Offmann B, D’Hont A, Risterucci AM, Roques D, Glaszmann JC, Grivet L (2001) Genetic dissection of a modern sugarcane cultivar (Saccharum spp.). I. Genome mapping with AFLP markers. Theor Appl Genet 103:84–97
Hoy JW, Hollier CA (2009) Effect of brown rust on yield of sugarcane in Louisiana. Plant Dis 93:1171–1174
Larkin MA, Blackshields G, Brown NP, Chenna R, Mc Gettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947
Le Cunff L, Garsmeur O, Raboin LM, Pauquet J, Telismart H, Selvi A, Grivet L, Philippe R, Begum D, Deu M, Costet L, Wing R, Glaszmann JC, D’Hont A (2008) Diploid/polyploid syntenic shuttle mapping and haplotype-specific chromosome walking toward a rust resistance gene (Bru1) in highly polyploid sugarcane (2n ~ 12x ~ 115). Genetics 180(1):649–660
Li WF, Wang XY, Huang YK, Zhang RY, Shan HL, Yin J, Luo ZM (2017) Molecular detection of Bru1 gene and identification of brown rust resistance in chinese sugarcane germplasm. Sugar Technol 19(2):183–190
Liu YQ, Li YP, Liang WH, Song QD, Qin XL, Ye L (2015) Current situation of sugar cane industry in the world. World Agriculture 2015(8):147–152
Lu YH, D’Hont A, Walker DIT, Rao PS, Feldmann P, Glaszmann JC (1994a) Relationships among ancestral species of sugarcane revealed with RFLP using single copy maize nuclear probes. Euphytica 78:7–18
Lu YH, D’Hont A, Paulet F, Grivet L, Arnaud M, Glaszmann JC (1994b) Molecular diversity and genome structure in modern sugarcane varieties. Euphytica 78:216–226
Luo J, Pan YB, Xu L, Zhang H, Yuan Z, Deng Z, Chen R, Que Y (2014) Cultivar evaluation and essential test locations identification for sugarcane breeding in China. Sci World J 2014:302753
Mukherjee SK (1957) Origin and distribution of Saccharum. Bot Gaz 119:55–61
Panje RR (1972) The role of Saccharum spontaneum in sugarcane breeding. Proc Int Soc Sugar Cane Technol 14:217–223
Panje RR, Babu CN (1960) Studies in Saccharum spontaneum: distribution and geographic association of chromo-some numbers. Cytologia (Tokyo) 25:152–172
Parco AS, Avellaneda MC, Hale AH, Hoy JW, Kimbeng CA, Pontif MJ, Gravois KA, Baisakh N (2014) Frequency and distribution of the brown rust resistance gene Bru1 and implications for the Louisiana sugarcane breeding programme. Plant Breed 133(5):654–659
Peixoto-Junior RF, Creste S, Landell MGA, Nunes DS, Sanguino A, Campos MF, Vencovsky R, Tambarussi EV, Figueira A (2014) Genetic diversity among Puccinia melanocephala isolates from Brazil assessed using simple sequence repeat markers. Genet Mol Res 13(3):7852–7863
Peng SG (1980) The general situation of evolution of sugarcane varieties at home and abroad. J South Agric 10:12–15
Peng SG (1996) Varietal evolution of sugarcane in Taiwan. Southwest China J Agric Sci 9(1):117–124
Piperidis G, Piperidis N, D’Hont A (2010) Molecular cytogenetic investigation of chromosome composition and transmission in sugarcane. Mol Genet Genom 284(1):65–73
Pocovi MII, Rech GE, Collavino NG, Caruso GB, Rios R, Mariotti JA (2010) Molecular diversity of Puccinia melanocephala populations. J Phytopathol 158:769–775
Price S (1968) Cytology of Chinese and North Indian sugarcanes. Econ Bot 22:155–164
Purdy LH, Liu JL, Dean JL (1983) Sugarcane rust, a newly important disease. Plant Dis 67:1292–1295
Que Y, Xu L, Wu Q, Liu Y, Ling H, Liu Y, Zhang Y, Guo J, Su Y, Chen J, Wang S, Zhang C (2014) Genome sequencing of Sporisorium scitamineum provides insights into the pathogenic mechanisms of sugarcane smut. BMC Genom 15:996
Raboin LM, Oliveira KM, Lecunff L, Telismart H, Roques D, Butterfield M, Hoarau JY, D’Hont A (2006) Genetic mapping in sugarcane, a high polyploid, using bi-parental progeny: identification of a gene controlling stalk colour and a new rust resistance gene. Theor Appl Genet 112:1382–1391
Racedo J, Perera MF, Bertani R, Funes C, González V, Cuenya MI, D’Hont A, Welin B, Castagnaro AP (2013) Bru1 gene and potential alternative sources of resistance to sugarcane brown rust disease. Euphytica 191:429–436
Roach BT (1972) Nobilization of sugarcane. Proc Int Soc Sugar Cane Technol 14:206–216
Roach BT (1977) Utilization of Saccharum spontaneum in sugarcane breeding. Proc Int Soc Sugar Cane Technol 16:43–57
Ruan XY, Yan F, Sun CJ (1983) Occurrence of Puccinia erianthi on sugarcane in Yunnan province. Acta Mycologica Sinia 2:260–261
Sreenivasan TV, Ahloowalia BS, Heinz DJ (1987) Cytogenetics. In: Heinz DJ (ed) Sugarcane improvement through breeding. Elsevier, Amsterdam, pp 143–210
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Todd J, Glaz B, Burner D, Kimbeng C (2015) Historical use of cultivars as parents in Florida and Louisiana sugarcane breeding programs. Int Sch Res Not 2015:257417
Wang XY, Li WF, Huang YK, Lu X, Luo ZH, Yin J, Shan HL, Zhang RY (2013) Evaluation of sugarcane introgression lines for resistance to brown rust disease caused by Puccinia melanocephala. Trop Plant Pathol 38(2):097–101
Wei Q, Yang BL, Gao ZJ (2015) Analysis of current situation of sugar cane industry. J Agric Mech Res 4:247–254
Zhang XY (1996) Utilization of S. robustum descendants in sugarcane breeding. Sugarcane 3(1):14–18
Zhang YX (2003) Sugarcane breeding strategies in Taiwan. Sugarcane Canesugar 2003(1):22–25
Zhang Q, Qi YW, Zhang CM, Chen YS, Deng HH (2009) Pedigree analysis of genetic relationship among core parents of sugarcane in Mainland China. Guangdong Agric Sci 2009(10):44–48
Zhang J, Sharma A, Yu Q, Wang J, Li L, Zhu L, Zhang X, Chen Y, Ming R (2016) Comparative structural analysis of Bru1 region homeologs in Saccharum spontaneum and S. officinarum. BMC Genom 17:446
Zhou M (2013) Conventional sugarcane breeding in South Africa: progress and future prospects. Am J Plant Sci 4:189–196
Acknowledgements
This work was supported by a startup fund for distinguished scholars of Fujian Agriculture and Forestry University, and the Project of Education and Scientific Research of Young Teacher of Fujian (JA13102). The authors would like to thank Xin Lu for providing facilities and help in obtaining the leaf tissues from NNSGR.
Author information
Authors and Affiliations
Contributions
YL conceived and designed the experiments; HW, PC and YY conducted the experiments; HW and YL processed the data; YL wrote the manuscript; AD reviewed the manuscript; all authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The authors declare that the experiments presented in this publication comply with the current laws of China and France.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Communicated by Dr. Antonio Augusto Franco Garcia.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, HB., Chen, PH., Yang, YQ. et al. Molecular insights into the origin of the brown rust resistance gene Bru1 among Saccharum species. Theor Appl Genet 130, 2431–2443 (2017). https://doi.org/10.1007/s00122-017-2968-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2968-3