Abstract
Key message
The resistance genes Rph22 and Rym16 Hb transferred into barley from Hordeum bulbosum have been separated from a large yield penalty locus that was present in the original introgression line ‘182Q20’.
Abstract
The Hordeum bulbosum introgression line ‘182Q20’ possesses resistance to barley leaf rust (Rph22) and Barley mild mosaic virus (Rym16 Hb) located on chromosome 2HL. Unfortunately, this line also carries a considerable yield penalty compared with its barley genetic background ‘Golden Promise’. Quantitative trait locus (QTL) mapping of the components of yield (total yield, thousand grain weight, hectolitre weight, percentage screenings and screened yield) was performed using 75 recombinant lines derived from the original ‘182Q20’ introgression line. A QTL for the yield penalty was located in the proximal region of the introgressed segment. Marker assisted selection targeting intraspecific recombination events between overlapping H. bulbosum introgression segments was used to develop the lines ‘372E’ and ‘372H’ which feature genetically small introgressions around Rph22. Further yield trials validated the separation of both Rph22 and Rym16 Hb from the proximal yield penalty. These results, combined with molecular markers closely linked to Rph22 and Rym16 Hb, make these resistance genes more attractive for barley breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The use of wild relatives for the genetic improvement of cultivated species is an important source of novel alleles and traits. However, the introgression of these target traits into commercial germplasm can often take many years to accomplish. Hordeum bulbosum L. is a member of the secondary gene pool of cultivated barley (Hordeum vulgare L.) and has been used mostly in barley improvement as a means to produce doubled haploids through chromosome elimination (Kasha and Kao 1970). In addition, H. bulbosum is considered a non-host to many pathogens which are virulent upon barley, and several resistance genes have been successfully introgressed from H. bulbosum into cultivated barley (Fetch et al. 2009; Johnston et al. 2013; Pickering et al. 1995, 1998, 2000, 2006; Ruge-Wehling et al. 2006; Ruge et al. 2003; Scholz et al. 2009; Shtaya et al. 2007; Toubia-Rahme et al. 2003; Walther et al. 2000; Xu and Kasha 1992). The development of crops which are resistant to diseases is a major goal of plant breeding in almost all commercial crop species. To provide a tangible benefit, crop resistance not only needs to provide yield stability under biotic stress, but also be durable. The test for durability of disease resistance requires that a given resistance has remained effective despite being challenged by the disease over a long period of time and over a large geographical area (Johnson 1984). Major resistance (R) genes have been used for many decades by plant breeders to protect cultivars from particular disease pathotypes in a gene-for-gene manner (Flor 1956). Unfortunately, pathogens often overcome newly deployed R genes within a few years of widespread cultivation in regions conducive to disease (Clifford 1985). Once a resistance gene has been overcome, virulent pathotypes can spread rapidly across or between continents via long distance air dispersal of fungal spores, thus compromising previously resistant crops in distant areas (Brown and Hovmøller 2002). With limited sources of new disease resistance genes, there is a need to develop more durable solutions. Partial, quantitative or adult plant resistance (APR) genes fall into a second category of plant disease resistance that is currently receiving a resurgence of interest especially against the cereal rusts (Case et al. 2014; Derevnina et al. 2013; Herrera-Foessel et al. 2014; Hulbert and Pumphrey 2014; Singh et al. 2013a; Ziems et al. 2014). For the purposes of this paper, we will use the term partial resistance as inclusive of APR. This type of resistance is often conferred by many genes of small effect, which together act to reduce disease severity despite a susceptible infection type (Parlevliet 1975, 1976, 1978). For barley leaf rust (Puccinia hordei Otth.), there have been twenty quantitative trait loci (QTL) uncovered which contribute to this ‘slow rusting’ type resistance (Marcel et al. 2007, 2008; Qi et al. 1998, 2000). The best understood ‘slow rusting’ system is from the cultivar ‘Vada’, which when crossed with the susceptible line ‘L94’ resulted in the identification and genetic mapping of six QTL conditioning this response (Qi et al. 1998). In contrast to R genes against barley leaf rust, which result in a hypersensitive response, the ‘slow rusting’ resistance reduces disease severity by limiting the pre-haustorial establishment of some fungal infection units (reduced infection frequency) and by delaying the development of sporification in those infection units which are able to establish (increased latency period) (Niks 1986). The ‘slow rusting’ resistance found in ‘Vada’ has proven durable despite widespread cultivation for several decades (Parlevliet 2002). However, partial resistance can be very difficult to manipulate in a breeding program, as many small effect genes need to be maintained for the resistance to be effective. The use of partial resistance in breeding has been made easier by the cloning of genes such as Lr34 (Krattinger et al. 2009) and Yr36 (Fu et al. 2009) in wheat, thus providing perfect markers for marker assisted selection (MAS). These efforts have also revealed that the genes underlying partial resistance are likely to be diverse in type and function (Fu et al. 2009). However, most of the genes/QTL for partial resistance have not been closely linked to genetic markers to aid their incorporation into breeding lines. Even if markers were available, the time and expense involved is likely to be prohibitive for the degree of resistance gain from each of these small effect genes.
Rph22 is a large effect partial resistance gene that was introgressed into cultivated barley from the wild species H. bulbosum. Rph22 is likely to be conferred by a single dominant gene located on the distal end of chromosome 2HL (Johnston et al. 2013). The presence of this single locus results in an increased latency period and reduced infection frequency that is superior to the degree of resistance found in the cultivar ‘Vada’, which is conditioned by at least six separate QTL (Qi et al. 1998). Partial resistance genes are also known as APR genes as their effect on extending fungal latency period increases over the course of plant development (Parlevliet 1975). The introgression line ‘182Q20’, derived from the backcross of diploid barley cultivar ‘Golden Promise’, to a partially fertile triploid interspecific hybrid between ‘Golden Promise’ and the tetraploid H. bulbosum genotype A17-1 (Johnston et al. 2013) was first identified in the field because of its ‘slow rusting’ response to natural infections of barley leaf rust. This line was subsequently shown using genomic in situ hybridization (GISH) and molecular markers to possess a single introgression from H. bulbosum which covers approximately 6 % of the physical length (IBSC physical map, 2012) and 24 % of the genetic length (POPSEQ map, Mascher et al. 2013) of chromosome 2H (N. Wendler, pers. comm.). Because of the large effect on latency period and infection frequency of Rph22, the presence of this gene can also be readily detected at the seedling stage (Pickering et al. 2004a). Molecular mapping of Rph22 has revealed a small overlapping genetic interval and the same phenotypic mechanism as Rphq2 (Johnston et al. 2013), the largest effect and most stable QTL from ‘Vada’ (Marcel et al. 2007). It seems likely that Rph22 and Rphq2 are paralogs of the same ancestral gene or members of a gene cluster. Rph22 is an attractive gene for barley breeding as it confers a large effect ‘slow rusting’ resistance which, because of its mechanism, may prove to be more durable than hypersensitive R genes currently available for the control of leaf rust. In addition, the incorporation of one large effect partial resistance gene is technically much simpler than tracking multiple genes of lesser effect.
The complex of Barley mild mosaic virus (BaMMV) and Barley yellow mosaic virus (BaYMV) is transmitted by the soil-borne plasmodiophorid Polymyxa graminis and has been a problem for winter grown barley crops in Asia and Europe since the late 1970s and 1980s (Huth and Lesemann 1978; Hill and Evans 1980; Lapierre 1980; Kobayashi et al. 1987; Ruan and Jin 1983; Rubies-Autonell et al. 1995; Katis et al. 1997). To date, 18 resistance loci have been identified against these viruses, with 15 recessive genes and one dominant gene located in H. vulgare (Ordon et al. 2005; Kai et al. 2012), and two dominant genes (Rym14 Hb and Rym16 Hb) from H. bulbosum (Ruge et al. 2003; Ruge-Wehling et al. 2006). Development of resistant cultivars is the only effective tool for controlling the effect of these viruses. In addition to Rph22, the H. bulbosum introgression line ‘182Q20’ also carries the resistance gene Rym16 Hb which confers resistance to all known isolates of this virus complex in Germany (Habekuß et al. 2008). The resistance gene Rym16 Hb has been previously mapped to chromosome 2HL (Ruge-Wehling et al. 2006) in different H. bulbosum introgression mapping populations. Mapping of Rym16 Hb using the “182Q20_F4_Popn” would provide a greater mapping resolution and thus is seen as a useful population for examining both these resistance genes.
Some sources of durable disease resistance are known to be pleiotropic with traits that have a negative effect on yield, such as the leaf tip necrosis (Ltn) with Lr34. Indeed, the incorporation of Lr34 (and Ltn) was shown to give a 5 % reduction in yield under disease-controlled conditions (Singh and Huerta-Espino 1997). Sourcing novel resistance genes from wild relatives can also be associated with problems of linkage drag, resulting in the transfer of additional deleterious (or undomesticated) alleles/genes along with the target trait. Rph22 and Rym16 Hb are no exception, as the original H. bulbosum introgression line ‘182Q20’ has an approximately 25 % lower yield than its genetic background ‘Golden Promise’ under fungicide control (Pickering et al. 2004b). Interestingly, there was no appreciable difference in yield between ‘182Q20’ and ‘Golden Promise’ in the presence of natural leaf rust infection (Pickering et al. 2004b). This paper describes the experiments to locate Rym16 Hb genetically in the mapping population established for Rph22 (Johnston et al. 2013) and to determine whether the yield penalty is a consequence of (or pleiotropic to) these resistance genes or whether it is the result of linkage drag of other alleles/genes from the wild species H. bulbosum.
Materials and methods
Mapping of Barley mild mosaic virus resistance (BaMMV)
The “182Q20_F4_Popn”, previously developed for mapping Rph22 (Johnston et al. 2013) was also used to genetically map Rym16 Hb. This population consists of 176 lines, with between one and five independent F 4 homozygous sister lines derived from each of the original 76 F 2 recombinant lines as described in Johnston et al. (2013).
Six seeds of each of 155 lines from the “182Q20_F4_Popn” (21 lines were not included because of low amounts of seed from genotyped seed lots) plus cv. ‘Golden Promise’, introgression lines ‘182Q20’ and ‘372E’ were sown in the greenhouse. In addition, 17 seeds from the susceptible standard cv. ‘Maris Otter’ were also included in the experiment. At the 2–3 leaf stage, the plants were transferred to a climatic chamber with 12 °C and 16 h photoperiod (16 kLx). Between four and six plants (depending on germination), from each line were screened for their response to mechanical inoculation with the isolate BaMMV-ASL1 as described by Habekuß et al. (2008). Five weeks after inoculation, the number of plants with mosaic symptoms was scored and DAS-ELISA using polyclonal antibodies prepared by Frank Rabenstein (JKI, Institute for Epidemiology and Pathogen Diagnostics, Quedlinburg) was carried out. The infection rate (%) was calculated as the number of infected plants/number of inoculated plants.
Plant materials and field trial designs
A total of four field trials were carried out, trial 1 (2010–11) and trial 2 (2011–12) were designed to map the QTL responsible for the yield penalty whilst trials 3 (2012–13) and 4 (2013–14) were used to examine the potential to separate Rph22 and Rym16 Hb from the yield penalty QTL. All field trials were conducted near Lincoln, Canterbury, New Zealand. Field trials 1 and 2 featured 13 plots of each of two parent lines (high yielding H. vulgare cv. ‘Golden Promise’, low yielding H. bulbosum introgression line ‘182Q20’) along with two plots each of 75 recombinant lines (one homozygous F 4 representative derived from each of the original 76 F 2 recombinants as previously described in Johnston et al. (2013), with one left out because of insufficient seed). The layout of trial 1 was derived from four 11 × 11 Latin squares. The same trial design was employed for field trial 2, but with a new randomization.
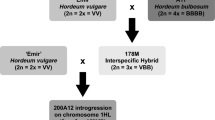
A crossing strategy was developed to reduce the size of the H. bulbosum introgression around Rph22 and the yield penalty locus. Crosses were made between a line which had the proximal region of the original introgression including the locus of interest and a line which had the distal region of the original introgression including the locus of interest. These combinations are shown in Fig. 1 with grouped triplets featuring the two parental lines and the resulting line selected from the progeny of that cross. For example, the line ‘372H’ (targeting Rph22) was derived from a cross between ‘IL_161’ and ‘IL_069’. By selecting lines with recombination events close to the target locus, it was possible to minimize extraneous regions of the introgression. Previously developed PCR markers (Johnston et al. 2009, 2013) were used to identify intraspecific recombination events (within the overlapping introgressions) in the F 2 lines. These F 2 recombinants of interest would be homozygous for the H. bulbosum genotype at the target region, heterozygous for one end of the introgression and homozygous for the barley genotype at the opposite end of the introgression. Individual F 3 plants which possessed homozygous introgression genotypes of reduced size were then selected from the selfed progeny and coded ‘372E’, ‘372H’, ‘372Q’ and ‘372W’. All lines discussed in this paper are available for distribution under MTA agreement.
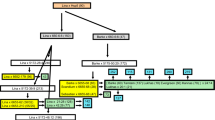
Genotypes of key introgression lines (ILs) and parents (from Johnston et al. 2013), that were used in field trials and for crossing to develop barley lines ‘372E’, ‘372H’, ‘372Q’ and ‘372W’. Lines are shown in crossing groups, i.e., ‘372E’ was developed from a cross between ‘IL_069’ with ‘IL_055’. Only a single marker from each unique genetic locus is shown in the order of genetic linkage from the distal end of chromosome 2HL (left) to the proximal end of the original introgression from ‘182Q20’ (right). Genotypes are displayed as V for the homozygous Hordeum vulgare allele and B for the homozygous Hordeum bulbosum allele. For Rph22 and Rym16 Hb, phenotypes are shown as R for resistant and S for susceptible
Trial 3 included the newly developed ‘372E’, five key introgression lines (‘IL_016’, ‘IL_041’, ‘IL_069’, ‘IL_101’, and ‘IL_161’) selected based on their genotype for containing different regions of the original introgression (Fig. 1) and the two parental lines ‘182Q20’ and ‘Golden Promise’. This trial was sown using a design modified from an 8 × 8 Latin square, with four of the eight plots of ‘372E’ replaced by two additional plots of each of the two parents (because of insufficient seed from the glasshouse increase of ‘372E’).
Trial 4 included the smallest introgression line around Rph22 coded ‘372H’ (which had superseded the previous line ‘372E’) and additional lines ‘372Q’ and ‘372W’ which were also developed using the same system of overlapping crosses in an attempt to better resolve the genetic location of the yield penalty QTL. This trial featured two blocks, each containing three replicates of eight lines (six introgression lines ‘372H’, ‘372Q’, ‘372W’, ‘IL_016’, ‘IL_041’, ‘IL_069’ and the two parents). The two blocks featured different fungicide regimes (treated and untreated), each laid out using a Latinized resolvable block design generated with CycDesigN (CycSoftware 2009). As there was insufficient seed of the line ‘372Q’ for six plots, this line was replaced by ‘Golden Promise’ in the untreated block.
All trials were managed to maximize yield potential. Each trial received two applications of fungicides, Proline® (prothioconazole, Bayer CropScience) and Proline® + Amistar® (Azoxystrobin, Syngenta) at the recommended rates and two applications of urea at the rate of 150 kg per hectare (a total of 138 units nitrogen per hectare). However, trial 4 featured a block which was not treated with fungicide, to examine the effect of Rph22 in the presence of natural leaf rust infection. Plot sizes for all trials were 5.2 × 1.3 m (6.75 m2) and agronomic data were collected from all field trials including total plot yield, thousand grain weight (TGW), hectolitre weight, percentage screenings (percentage of a grain sub-sample which fell through a 2.4-mm slotted screen) and screened yield (total plot yield less the weight due to screenings). Lodging was observed in all four trials but was only formally recorded in trial 2 (2011–12). Lodging score (proportion of each plot that was still standing, i.e., 10 = all standing, 0 = completely lodged) was recorded for the 2011–12 field trial only on two separate dates.
Statistical analysis
Spatial patterns in each trial were explored using the methods described by Verbyla et al. (1999). These analyses showed that there were trends across each trial. The trends were accounted for within a mixed model analysis fitted with restricted maximum likelihood (REML) (Payne et al. 2012). Lines were included in these analyses as fixed effects, and factors to adjust for spatial patterning as random effects. Adjusted means for each line and associated standard errors were obtained from the results of these analyses.
Genotypic data from 21 marker loci (a single marker from each unique recombination interval) were combined with the agronomic data from trials 1 and 2. Separate analyses were carried out for each marker locus. The REML analysis was extended by partitioning the line effects into B (allele derived from H. bulbosum introgression line parent ‘182Q20’) versus V (allele derived from H. vulgare parent ‘Golden Promise’) for the marker, and lines within each of these groups. These analyses gave exactly the same adjusted line means, but allowed estimation of the mean for lines with B or V and a comparison of the B mean with the V mean. The approximate F-probabilities for these comparisons were obtained with denominator degrees of freedom for the F-statistic calculated using the method of Kenward and Roger (1997). The −log10 of the F probability is comparable with the logarithm of odds (LOD) score commonly used in QTL analyses.
Results
The resistance reactions of 155 lines from the “182Q20_F4_Popn” were evaluated in comparison to susceptible and resistant control lines after mechanical BaMMV inoculation. All tested plants of the susceptible control lines were infected; cv. ‘Golden Promise’ (5 infected plants/5 inoculated plants), ‘372E’ (5/5) and cv. ‘Maris Otter’ (17/17). The infection rate of the resistant control ‘182Q20′ was 0 % (0/6). Of the 155 tested lines from the “182Q20_F4_Popn”, 72 lines displayed a resistant reaction type with no visible leaf symptoms and no serological detection of virus. The remaining 83 lines showed a susceptible reaction type (infection rates of 33–100 %). The 155 lines tested here were homozygous lines derived from 74 unique F 2 recombinants (Johnston et al. 2013). This resulted in a total of 36 resistant families and 38 susceptible families and consistent with Rym16 Hb being conditioned by a single locus. Only two lines (‘IL_021’ and ‘IL_141’) gave inconsistent results within their families. Both these lines were initially classified as resistant whilst sister lines (‘ILs_019’, ‘020’, ‘022’ and ‘IL_142’, respectively) were all susceptible. Assuming that ‘IL_021’ and ‘IL_141’ were escapes (actually susceptible), the best fit of these data was for the resistance gene Rym16 Hb to co-segregate with the marker H35_17700 (k03475) near the distal end of the introgression (Figs. 1, 2).
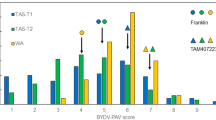
Marker/trait associations from the 2010–11 (trial 1) and 2011–12 (trial 2) barley field trials (75 introgression lines). The blue and green bars on the linkage map indicate the position of the resistance loci Rph22 and Rym16 Hb, respectively. The thick bars displaying the location of quantitative trait loci (QTL) on the linkage map show the area covered by a drop of one −log10(P) (equivalent to LOD) from the peak and the thin lines show an additional drop of one −log10(P). Similar colours have been used to group each component of yield trait for each field trial. Lodging was recorded in 2011–12 (trial 2) only. Numbers to the left of the linkage map indicate intervals between markers in cM (colour figure online)
For the agronomic yield traits measured, the introduction of the H. bulbosum introgression in ‘182Q20’ led to a less desirable mean when compared to the genetic background, cv. ‘Golden Promise’ (Table 1), i.e., lower total yield, TGW, screened yield, lodging score and higher percentage screenings. However, hectolitre weight was variable between the parental lines in trials 1 and 2. Spatial adjustments were applied to the means of the agronomic data to account for trends across the field trials, however, in most cases these adjustments made only minor changes to those means. Some lines had yield components superior to those of cv. ‘Golden Promise’ and poorer than those of ‘182Q20’, although none was significantly different (p > 0.05) (data not shown). As would be expected, total (raw) yield was correlated with screened yield for field trials 1 and 2 (Table 2 and 3), but it was not well correlated with the other variables. Screened yields were well correlated with TGW, and with percentage screenings, but not with hectolitre weights. In all field trials, there was a considerable degree of post-anthesis lodging observed because of relatively high yields combined with strong winds and the poor lodging resistance of the parental cultivars ‘Golden Promise’ and ‘182Q20’. In each trial, all plots were harvested using a small plot combine and manual lifting of stems to ensure that all the seed was harvested despite the crop lodging.
The marker analyses for the components of yield in field trials 1 and 2 were carried out using the adjusted means, as the differences between the H. bulbosum and barley marker alleles were enhanced in the adjusted analysis and thus the associated p values were smaller. In both trials 1 and 2, a QTL affecting several components of yield was detected in the proximal region of the introgression between markers H35_1860 and H35_18000 (Fig. 2). Lines with barley alleles in this region were strongly associated with an increase in total yield, TGW, hectolitre and screened yield, and a decrease in percentage screenings compared with lines having the H. bulbosum alleles. The most significant differences between the marker alleles were seen in the data for screened yield and percentage screenings (Fig. 2). The QTL peaks for lodging (which was only recorded in trial 2, 2011–12) and hectolitre weight (2011–12) were located more centrally within the introgressed segment, with a peak at marker k04109 (Fig. 2), compared with the other traits. Both, the Rph22 and Rym16 Hb resistance loci are located near the distal end of the introgression and hence were not closely associated with any of the proximal QTL regions affecting the yield parameters in either of the field trials (Fig. 2).
To validate the separation of Rph22 and Rym16 Hb from the yield penalty QTL near Cly1, five key introgression lines and the newly developed line ‘372E’ were included in trial 3 with a higher rate of plot replication. Analysis of data from this third field trial revealed a clear grouping of the lines with one or other of the two parents. Yield component data for lines ‘372E’ and ‘IL_069’ both clustered with the parent cv. ‘Golden Promise’, whilst the yield component data for the remaining lines (‘IL_016’, ‘IL_041’, ‘IL_101’, ‘IL_161’) clustered with the parent line ‘182Q20′ (Fig. 3). This confirmed that the smaller distal introgression of ‘IL_069’ (containing both Rph22 and Rym16 Hb) and the reduced introgression around Rph22 in line ‘372E’ resulted in the same yield characteristics as those of cv. ‘Golden promise’ (Fig. 3). There were significant differences between the lines for all five variables (p < 0.001 for an overall test for line differences for variables other than hectolitre weight, where p = 0.024). A glasshouse pathology screen confirmed that ‘372E’ possessed the same ‘slow rusting’ response to leaf rust as ‘182Q20’ (data not shown).
Adjusted means for eight barley lines across five yield components, Yield, % screenings, screened yield (kg per plot), thousand grain weight (TGW, g per 1000 seed) and hectolitre weight (g) from field trial 3 (2012–13). Closed symbols indicate lines share similar yield parameters to ‘Golden Promise’ and open symbols to ‘182Q20’. Error bars are LSD 5 % to compare a line with a parent, with the shorter (lower) bar for comparisons of most lines with the parents, and the longer (upper) bar to compare line ‘372E’ with the parent (because of a lower number of replications in the trial)
Trial 4 featured two blocks that differed in their fungicide treatment. In the fungicide treated block, there was a clear demarcation between lines with either ‘Golden Promise’ or ‘182Q20’ yield component data. The line ‘372H’ carrying the smallest introgression around Rph22 plus lines ‘372Q’ and ‘IL_069’ (containing both Rph22 and Rym16 Hb) possessed the same yield parameters as cv. ‘Golden Promise’, whereas lines ‘372W’ ‘IL_016’ and ‘IL_041’ possessed the yield parameters of ‘182Q20’ (Fig. 4). In the block without fungicide treatment, there was a small amount of late leaf rust infection, which allowed confirmation of the same ‘slow rusting’ resistance in ‘372H’ as in ‘182Q20’. Under this disease pressure from barley leaf rust, lines ‘IL_069’ (both Rph22 and Rym16 Hb) and ‘372H’ (Rph22 only), which feature small distal introgressions, both outperformed cv. ‘Golden Promise’ for total yield, percentage screenings and screened yield (Fig. 4). The original introgression parent ‘182Q20’ had very high percentage screenings in this untreated block and consequently the lowest screened yield.
Relative adjusted mean screened grain yield (cv. ‘Golden Promise’ 100) from trial 3 (2012–13) and trial 4 (2013–14) featuring fungicide-treated (left, trials 3 and 4) and untreated (right, trial 4 only) blocks. Parental lines cv. ‘Golden Promise’ (GP light grey) and ‘182Q20’ (Q darker grey) are highlighted for reference purposes. Bars indicate 95 % confidence intervals. The confidence intervals differ between the trials due to differences in underlying random variation and the number of replicates (8 plots per line in trial 3, except ‘372E’ which had 4 plots; but only 3 plots per line in trial 4)
The lines ‘372E’, ‘372H’, ‘372Q’ and ‘372W’, with reduced introgressions around loci of interest were successfully developed from crosses between ILs that possessed proximal and distal introgressions which overlapped around the locus of interest. The use of markers across the entire introgression segment allowed lines resulting from only intraspecific recombination events within this overlapping region to be identified in the F 2 progeny. The number of these recombinants detected was dependent on the genetic size of the overlap based on the interspecific genetic map (Fig. 2). For instance; three intraspecific recombinants were detected from 184 ‘372H’ F 2 seedlings (~0.5 cM overlap, Figs. 1, 2), whilst eight intraspecific recombinants were detected from only 92 ‘372Q’ F 2 seedlings (~2.4 cM overlap, Figs. 1, 2). Subsequent selection of F 3 selfed seed from these intraspecific recombinants resulted in the identification of lines that were homozygous for these introgressions of reduced size. These lines possessed homozygous introgressions which spanned the marker intervals H35_19216 to H35_13826 (‘372E’), Rph22 to H35_13826 (‘372H’), k06104 to k00917 (‘372Q’) and k00917 to H35_15016 (‘372W’) (Fig. 1). The cleistogamy locus was phenotyped as a morphological marker as described previously (Johnston et al. 2013).
Discussion
Disease resistance is a key target trait for plant breeding and a major contributing factor to yield stability. Conventional breeding has historically focused on the selection of breeding lines displaying immunity or hypersensitive response under disease pressure. This type of selection favours the incorporation of major R genes, which give a clean crop in the field. However, these single dominant, major resistance genes are often rapidly overcome upon widespread cultivation. For example, from 235 worldwide isolates of wheat stripe rust, virulence was detected for all but two R genes tested (Yr5 and Yr15) (Sharma-Poudyal et al. 2012). In contrast, partial resistance genes, as a group, are generally considered to be a more durable mechanism of disease resistance because of their polygenic nature and non-race specificity (Brown 2002). For instance, there are several examples of partial resistance genes which have remained effective over long time periods, such as Sr2 derived from Triticum turgidum in the wheat variety ‘Hope’ (McFadden 1930), Lr13 from the wheat variety ‘Frontana’ (Dyck et al. 1966), Lr34/Yr18/Pm38 from the wheat variety ‘Terenzio’ (Dyck 1987) and mlo from mutants and Ethiopian landraces of barley (Jørgensen 1992). The classic example of partial resistance, Lr34/Yr18/Pm38 in wheat, is effective against multiple pathogens such as leaf rust, stripe rust, powdery mildew and also stem rust in some genetic backgrounds (Dyck 1987). However, even amongst partial resistance genes there are examples which are race specific, for instance Lr12 against wheat leaf rust (Park and McIntosh 1994). The two partial resistance genes cloned to date from wheat (Yr36 and Lr34/Yr18/Pm38) have revealed different classes of genes with presumably different mechanisms of action (Fu et al. 2009; Krattinger et al. 2009). It also seems probable that if partial resistance genes are varied in form and function, their individual durability will be similarly heterogeneous.
While most partial resistance genes characterized to date have only small effects on the phenotypic variance of disease resistance, Rph22 is a single locus which contributes a large effect partial resistance response. The genetic mapping of Rph22 and the identification of molecular markers closely linked to this resistance gene (Johnston et al. 2013) will allow it to be incorporated into modern barley varieties. However, because of a considerable yield penalty associated with the H. bulbosum introgression containing Rph22, this gene has not been an attractive target for barley breeders. In this paper, we have used the “182Q20_F4_Popn” mapping population (Johnston et al. 2013) to QTL map (single marker analysis), the individual components of the yield penalty (total yield, TGW, hectolitre weight, percentage screenings and screened yield) to the proximal end of the introgression and genetically distinct from both Rph22 and Rym16 Hb (Fig. 2). To validate the results from the mapping population, two introgression lines, ‘372E’ and ‘372H’, were developed using targeted intraspecific recombination within overlapping introgression segments. This strategy successfully avoided the problem of highly suppressed recombination that has been previously seen in a physically small introgression on chromosome 2HL (Johnston et al. 2013). The use of overlapping introgression segments exploits intraspecific recombination to obtain the desired recombination events in areas where interspecific recombination is suppressed. The resulting lines have introgression boundaries that are defined by the original lines used in the cross, thus removing the need to screen a large number of lines to locate rare recombination events which are suitably close to the target gene. Instead, the position of the introgression boundaries used in the cross will determine the extent of the introgression in the newly developed line. The two lines developed in this manner, ‘372E’ and ‘372H’, were subsequently shown to have yield parameters comparable to those of the barley parent cultivar ‘Golden Promise’, with the additional benefit of the large effect partial resistance response conditioned by Rph22. The line ‘372H’, retains only a very small introgression around Rph22 (Fig. 1) and would be a suitable donor line for the incorporation of Rph22 into advanced breeding lines using marker assisted selection. In a previous study (Pickering et al. 2004b), plots of the IL ‘182Q20’ gave a similar yield to ‘Golden Promise’ under natural leaf rust infection. In trial 4, without fungicide treatment, the three replicates of ‘182Q20’ gave unusually high percentage screenings (52–68 %) compared with means of 33 and 37 % in trials 1 and 2 (Table 1). As there was only a small amount of late leaf rust infection in this trial, it seems likely that some other unrecorded biotic stress may have resulted in the poor performance of ‘182Q20’. The newly developed ‘372H’ and ‘IL_069’ had mean screened yields slightly higher than ‘Golden Promise’ (Fig. 4), indicating that they did not suffer from the problems associated with the full introgression of ‘182Q20’ in trial 4. The yield comparisons performed in this study were against the parental genetic background ‘Golden Promise’, which is no longer a high-yielding cultivar by modern standards. Incorporation of Rph22 into elite barley germplasm will be required to determine if there is an identifiable cost to the inclusion of this partial resistance gene itself. However, by reducing the overall size of the introgression around the target trait, it is possible to minimize the likelihood that other alleles/genes derived from H. bulbosum may contribute additional yield and/or quality issues.
Another partial resistance gene against barley leaf rust, called Rph20, was genetically mapped to chromosome 5HS (Hickey et al. 2011) and has been linked via pedigree/molecular marker analysis as having the same common origin as the cultivar ‘Vada’ from the barley landrace H. laevigatum. The partial resistance in ‘Vada’, including Rphq2 (likely to be a paralog of Rph22) and Rphq4 (likely to be the same gene as Rph20), has been durable over a considerable period of time (Parlevliet 2002). By extrapolation, we postulate that Rph22 and Rph20 may represent durable components of the resistance in ‘Vada’ against barley leaf rust. Further characterization of these partial resistance genes across a range of plant developmental stages, genetic backgrounds and temperatures, will lead to a greater understanding of the mechanisms involved and how effective these genes will be under field conditions for controlling barley leaf rust (Singh et al. 2013b). Wang et al. (2010) were able to show the different effects that Rphq2, Rphq3 and Rphq4 (and their susceptible alleles) conferred at different growth stages, with Rphq2 more effective at seedling stages and Rphq4 more effective at adult stages. Combinations of QTL/genes for partial resistance to barley leaf rust are known to improve the overall effectiveness of the disease resistance (Parlevliet 1976). Rphq2 and Rphq4 can clearly have synergistic roles in plant defense at different plant development stages and perhaps using different pathways (Wang et al. 2010). With molecular markers linked to genes such as Rph22/Rphq2, Rph20/Rphq4, it is now possible to combine, track and confirm the effectiveness of these combinations. Additional partial resistance genes against barley leaf rust have also been identified from H. bulbosum introgressions on chromosomes 1HL and 5HL (Pickering et al. 2004b). Combinations of these resistance genes would also be beneficial to explore but will need considerable work in marker development and genetic mapping.
Resistance to BaMMV/BaYMV in most winter barley cultivars is conditioned by the recessive genes rym4 and rym5. However, pathotypes of the soil-borne barley mosaic virus complex have been identified which have overcome rym4 (BaYMV-2, Huth 1989) and rym5 (BaMMV-Sil, Hariri et al. 2003; Kanyuka et al. 2004 and BaMMV-Teik, Habekuß et al. 2008). Consequently, it is important to look for new, effective resistance genes from other sources including the secondary gene pool of barley. Two dominant resistant genes, namely Rym14 Hb located on chromosome 6HS and Rym16 Hb on chromosome 2HL, were found in H. bulbosum (Ruge et al. 2004; Ruge-Wehling et al. 2006). The “182Q20_F4_Popn” was used to map the resistance gene Rym16 Hb against Barley mild mosaic virus with a greater number of 2HL markers, and for its position relative to Rph22 to be determined. In this study, Rym16 Hb was shown to co-segregate with the molecular marker H35_17700 (k03475) near the distal end of the introgression (Fig. 2). Previously, Rym16 Hb was mapped as the most distal 2HL marker, 3.6 cM distal of the RFLP marker MWG949 and 5.5 cM distal of the STS marker MWG2076 (Ruge-Wehling et al. 2006). In this study, Rym16 Hb was located 0.5 cM distal of MWG2076 (which co-segregates with H35_15816, Fig. 2). The presence of two resistance genes (Rph22 and Rym16 Hb) against two different diseases located near the distal end of a single introgression shows the high value of H. bulbosum as a resource for barley improvement. As BaMMV is not a problem in New Zealand barley crops, neither ‘372H’ nor ‘372E’ were developed to carry the Rym16 Hb resistance gene. However, the line ‘IL_069’ carries both Rph22 and Rym16 Hb, possesses a minimal distal introgression, and was shown in trials 3 and 4 to have similar yield parameters to those of cv. ‘Golden Promise’. Another line of interest, ‘IL_094’, was shown to have the genetically smallest distal introgression containing Rym16 Hb, spanning the genetic interval between markers k08380 and H35_17700 and thus should also not carry the yield penalty.
QTL for yield characteristics such as TGW have been previously mapped to chromosome 2HL for several barley mapping populations. Bezant et al. (1997) detected a QTL for TGW between markers bcd512b and bcd266 and three QTL for plant grain weight, ear grain weight and plot yield at the distal end of chromosome 2HL. Coventry et al. (2003) have reviewed a range of QTL contributing to grain yield, many of which are pleiotropic effects of plant development genes such as Ppd-H1, eps2, sgh1-3, sdw1/denso and Vrs1. In addition, there were several QTL mapped in three different mapping populations between bins 12 and 15 on chromosome 2HL ‘Blenheim’/’Kym’ (bins 12–14, thousand grain weight), ‘Igri’/’Danilo’ (bin 15, TGW) and ‘Sloop’/’Alexis’ (bin 13, screenings) by Coventry et al. (2003). The inheritance of genes/paralogs from H. bulbosum, within the ‘182Q20’ introgression on chromosome 2HL, resulted in a considerable deterioration in most of the agronomic traits measured (total yield, TGW, percentage screenings, screened yield and lodging). H. bulbosum is a wild species which possesses very slim seeds so these genes/paralogs on chromosome 2HL have obviously not been under the same degree of selective pressure as during the domestication and subsequent breeding of cultivated barley. The cleistogamy locus (Cly1 or HvAp2), which controls open/closed flowering (Nair et al. 2010), has been previously linked to QTL for TGW and lodging (Hori et al. 2005; Korff et al. 2006). The data from this study also support an association between the yield penalty inherited from H. bulbosum and the Cly1 locus. A broad QTL for reduction in yield attributes was initially mapped to the proximal end of the introgression using 75 lines from the “182Q20_F4_Popn” mapping population. This QTL location was then refined by the development of the additional introgression lines ‘372W’ and ‘372Q’ which possess introgressions that span different areas of this proximal region (Fig. 1). The line ‘372Q’ had yield characteristics similar to those of the H. vulgare parent cv. ‘Golden Promise’ (Fig. 4). However, the line ‘372W’, which carries an introgression spanning the marker interval k00917 to H35_15016 (including the Cly1 locus), had the same poor yield parameters as the original introgression line ‘182Q20’ (Fig. 4). This indicates that one locus or a cluster of tightly linked loci near Cly1 is responsible for the entire yield penalty. An additional locus involved in lodging resistance appears to be located more distally, but this is based on one year’s field trial only (2012–13). The initial concern was that lodging in the trials may contribute to measurable yield differences if there were appreciable differences in maturity between the lines. However, the mean lodging score was only correlated with hectolitre weight and less so with the other variables in 2011–12 (Table 3). Lodging can lead to ‘pinched’ grain resulting in smaller wrinkled seeds and thus have an effect on hectolitre weights. However, hectolitre weight was shown to be variable between seasons for the two parental lines and was not significantly different between ‘Golden Promise’ and ‘182Q20’ in trial 3 (2012–13, Fig. 3). Despite the lodging, we have managed to validate ‘Golden Promise’ yield characteristics in both ‘372E’ and ‘372H’ and thus the separation of Rph22 from the proximal yield penalty QTL. Even with only a small amount of late natural leaf rust infection, lines ‘372H’ and ‘IL_069’ both outperformed ‘Golden Promise’ due to the presence of the large-effect, partial leaf rust resistance of Rph22.
The distal end of barley chromosome 2HL is known to have high rates of recombination and to be gene rich (Chen et al. 2009; Künzel et al. 2000) and the corresponding region of wheat (2L1.0) covers only 5 % of the physical length but 68 % of the genetic length of that chromosome (Dilbirligi et al. 2005). Chromosome 2HL has also been the most abundant introgression location detected between H. vulgare and H. bulbosum (Johnston et al. 2009), which is most likely to be because of good rates of interspecific recombination, a high degree of co-linearity and the absence of critical barley genes which may not be compensated for by the introgressed segment. Additional traits or paralogs of interest may await discovery in these introgression lines which cover a gene-rich region of the barley genome.
Author contribution statement
Study was conceived by PAJ and RP, plants materials were initially developed by RP and MEF, marker genotyping was performed by VM and PAJ, linkage mapping by PAJ, plant crosses were performed by MEF, trial plans, statistical analysis and figures were done by RCB, and virus phenotyping was performed by AH. Manuscript was written by PAJ with contributions from AH and RCB. All authors contributed to editing.
References
Bezant J, Laurie D, Pratchett N, Chojecki J, Kearsey M (1997) Mapping QTL controlling yield and yield components in a spring barley (Hordeum vulgare L.) cross using marker regression. Mol Breed 3:29–38
Brown JKM (2002) Yield penalties of disease resistance in crops. Curr Opin Plant Biol 5:339–344
Brown JKM, Hovmøller MS (2002) Aerial dispersal of pathogens on the global and continental scales and its impact on plant disease. Science 297:537–541
Case AJ, Naruoka Y, Chen X, Garland-Campbell KA, Zemetra RS, Carter AH (2014) Mapping stripe rust resistance in a brundage × coda winter wheat recombinant inbred line population. PLoS One 9:e91758
Chen A, Brûlé-Babel A, Baumann U, Collins N (2009) Structure–function analysis of the barley genome: the gene-rich region of chromosome 2HL. Funct Integr Genomics 9:67–79
Clifford BC (1985) Barley leaf rust. In: Roelfs AP, Bushnell WR (eds) The cereal rusts volume II: diseases, distribution and control. Academic Press, New York
Coventry SJ, Barr AR, Eglinton JK, McDonald GK (2003) The determinants and genome locations influencing grain weight and size in barley (Hordeum vulgare L.). Aust J Agric Res 54:1103–1115
CycSoftware (2009) CycDesigN 4.0 A package for the computer generation of experimental designs, 4th edn. CycSoftware Ltd, Hamilton
Derevnina L, Singh D, Park RF (2013) Identification and characterization of seedling and adult plant resistance to Puccinia hordei in Chinese barley germplasm. Plant Breed 132:571–579
Dilbirligi M, Erayman M, Gill KS (2005) Analysis of recombination and gene distribution in the 2L1.0 region of wheat (Triticum aestivum L.) and barley (Hordeum vulgare L.). Genomics 86:47–54
Dyck PL (1987) The association of a gene for leaf rust resistance with the chromosome 7D suppressor of stem rust resistance in common wheat. Genome 29:467–469
Dyck PL, Samborski DJ, Anderson RG (1966) Inheritance of adult-plant leaf rust resistance derived from the common wheat varieties exchange and frontana. Can J Genet Cytol 8:665–671
Fetch T Jr, Johnston P, Pickering R (2009) Chromosomal location and inheritance of stem rust resistance transferred from Hordeum bulbosum into cultivated barley (H. vulgare). Phytopathology 99:339–343
Flor H (1956) The complementary genic systems in flax and flax rust. Adv Genet 8:29–54
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A kinase-START gene confers temperature-dependent resistance to Wheat Stripe Rust. Science 323:1357–1360
Habekuß A, Kühne T, Huth W, Rabenstein F, Ehrig F, Krämer I, Ordon F (2008) Identification of Barley mild mosaic virus isolates in Germany breaking rym5 resistance. J Phytopathol 156:36–41
Hariri D, Meyer M, Prud’homme H (2003) Characterization of a new Barley mild mosaic virus pathotype in France. Eur J Plant Pathol 109:921–928
Herrera-Foessel S, Singh R, Lillemo M, Huerta-Espino J, Bhavani S, Singh S, Lan C, Calvo-Salazar V, Lagudah E (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 127:781–789
Hickey L, Lawson W, Platz G, Dieters M, Arief V, Germán S, Fletcher S, Park R, Singh D, Pereyra S, Franckowiak J (2011) Mapping Rph20: a gene conferring adult plant resistance to Puccinia hordei in barley. Theor Appl Genet 123:55–68
Hill SA, Evans EJ (1980) Barley yellow mosaic virus. Plant Pathol 29:197–199
Hori K, Sato K, Nankaku N, Takeda K (2005) QTL analysis in recombinant chromosome substitution lines and doubled haploid lines derived from a cross between Hordeum vulgare ssp. vulgare and Hordeum vulgare ssp. spontaneum. Mol Breed 16:295–311
Hulbert S, Pumphrey M (2014) A time for more booms and fewer busts? unraveling cereal–rust interactions. Mol Plant Microbe Interact 27:207–214
Huth W (1989) Ein weiterer Stamm des Barley yellow mosaic virus aufgefunden. Nachrichtenbl Deut flanzenschutzd 41:6–7
Huth W, Lesemann DE (1978) Eine für die Bundesrepublik Deutschland neue Virose an Wintergerste. Nachrichtenbl Dtsch Pflanzenschutzdienstes 30:184–185
Johnson R (1984) A critical analysis of durable resistance. Annu Rev Phytopathol 22:309–330
Johnston P, Timmerman-Vaughan G, Farnden K, Pickering R (2009) Marker development and characterisation of Hordeum bulbosum introgression lines: a resource for barley improvement. Theor Appl Genet 118:1429–1437
Johnston PA, Niks RE, Meiyalaghan V, Blanchet E, Pickering R (2013) Rph22: mapping of a novel leaf rust resistance gene introgressed from the non host Hordeum bulbosum L. into cultivated barley (Hordeum vulgare L.). Theor Appl Genet 126:1613–1625
Jørgensen IH (1992) Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63:141–152
Kai H, Takata K, Tsukazaki M, Furusho M, Baba T (2012) Molecular mapping of Rym17, a dominant and Rym18 a recessive Barley yellow mosaic virus (BaYMV) resistance genes derived from Hordeum vulgare L. Theor Appl Genet 124:577–583
Kanyuka K, McGrann G, Alhudaib K, Hariri D, Adams MJ (2004) Biological and sequence analysis of a novel European isolate of Barley mild mosaic virus that overcomes the barley Rym5 resistance gene. Arch Virol 149:1469–1480
Kasha KJ, Kao KN (1970) High frequency haploid production in barley (Hordeum vulgare L.). Nature 225:874–876
Katis N, Tzavella-Klonari K, Adams MJ (1997) Occurrence of barley yellow mosaic and barley mild mosaic bymo viruses in Greece. Eur J Plant Pathol 103:281–284
Kenward MG, Roger JH (1997) Small sample inference for fixed effects from restricted maximum likelihood. Biometrics 53:983–997
Kobayashi SH, Yoshida S, Soutome S (1987) Breeding for resistance to yellow mosaic disease in malting barley. Proc 5th Int Barley Genet Symp Okayama Jpn Barley Genet V:667–672
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Künzel G, Korzun L, Meister A (2000) Cytologically integrated physical restriction fragment length polymorphism maps for the barley genome based on translocation breakpoints. Genetics 154:397–412
Lapierre H (1980) Nouvelles maladies à virus sur céréales d’hiver. Le Producteur Agricola Francais 270:11
Marcel TC, Aghnoum R, Durand J, Varshney RK, Niks RE (2007) Dissection of the barley 2L1.0 region carrying the ‘Laevigatum’ quantitative resistance gene to leaf rust using near-isogenic lines (NIL) and sub NIL. Mol Plant Microbe Interact 20:1604–1615
Marcel TC, Gorguet B, Ta MT, Kohutova Z, Vels A, Niks RE (2008) Isolate specificity of quantitative trait loci for partial resistance of barley to Puccinia hordei confirmed in mapping populations and near-isogenic lines. New Phytol 177:743–755
Mascher M, Richmond TA, Gerhardt DJ, Himmelbach A, Clissold L, Sampath D, Ayling S, Steuernagel B, Pfeifer M, D’Ascenzo M, Akhunov ED, Hedley PE, Gonzales AM, Morrell PL, Kilian B, Blattner FR, Scholz U, Mayer KFX, Flavell AJ, Muehlbauer GJ, Waugh R, Jeddeloh JA, Stein N (2013) Barley whole exome capture: a tool for genomic research in the genus Hordeum and beyond. Plant J 76:494–505
McFadden ES (1930) A successful transfer of emmer characters to Vulgare wheat. Agron J 22:1020–1034
Nair SK, Wang N, Turuspekov Y, Pourkheirandish M, Sinsuwongwat S, Chen G, Sameri M, Tagiri A, Honda I, Watanabe Y, Kanamori H, Wicker T, Stein N, Nagamura Y, Matsumoto T, Komatsuda T (2010) Cleistogamous flowering in barley arises from the suppression of microRNA-guided HvAP2 mRNA cleavage. Proc Natl Acad Sci 107:490–495
Niks RE (1986) Failure of haustorial development as a factor in slow growth and development of Puccinia hordei in partially resistant barley seedlings. Physiol Mol Plant Pathol 28:309–322
Ordon F, Ahlemeyer J, Werner K, Köhler W, Friedt W (2005) Molecular assessment of genetic diversity in winter barley and its use in breeding. Euphytica 146:21–28
Park RF, McIntosh RA (1994) Adult plant resistances to Puccinia recondita f. sp. tritici in wheat. N Z J Crop Hortic Sci 22:151–158
Parlevliet J (1975) Partial resistance of barley to leaf rust, Puccinia hordei. I. Effect of cultivar and development stage on latent period. Euphytica 24:21–27
Parlevliet J (1976) Partial resistance of barley to leaf rust, Puccinia hordei. III. The inheritance of the host plant effect on latent period in four cultivars. Euphytica 25:241–248
Parlevliet J (1978) Further evidence of polygenic inheritance of partial resistance in barley to leaf rust, Puccinia hordei. Euphytica 27:369–379
Parlevliet J (2002) Durability of resistance against fungal, bacterial and viral pathogens; present situation. Euphytica 124:147–156
Payne R, Welham S, Harding S (2012) A guide to REML in GenStat, 15th edn. VSN International, Oxford, p 94
Pickering R, Hill A, Michel M, Timmerman-Vaughan G (1995) The transfer of a powdery mildew resistance gene from Hordeum bulbosum L. to barley (H. vulgare L.) chromosome 2 (2I). Theor Appl Genet 91:1288–1292
Pickering R, Steffenson B, Hill A, Borovkova I (1998) Association of leaf rust and powdery mildew resistance in a recombinant derived from a Hordeum vulgare × Hordeum bulbosum hybrid. Plant Breed 117:83–84
Pickering R, Johnston P, Timmerman-Vaughan G, Cromey M, Forbes E, Steffenson B, Fetch T Jr, Effertz R, Zhang L, Murray B, Proeseler G, Habekuß A, Kopahnke D, Schubert I (2000) Hordeum bulbosum—a new source of disease and pest resistance genes for use in barley breeding programmes. Barley Genet Newsl 30:6–9
Pickering R, Johnston P, Ruge B (2004a) Importance of the secondary genepool in barley genetics and breeding I. Cytogenetics and molecular analysis. Czech J Genet Plant Breed 40:73–78
Pickering R, Niks RE, Johnston PA, Butler RC (2004b) Importance of the secondary genepool in barley genetics and breeding. II. Disease resistance, agronomic performance and quality. Czech J Genet Plant Breed 40:79–85
Pickering R, Ruge-Wehling B, Johnston P, Schweizer G, Ackermann P, Wehling P (2006) The transfer of a gene conferring resistance to scald (Rhynchosporium secalis) from Hordeum bulbosum into H. vulgare chromosome 4HS. Plant Breed 125:576–579
Qi X, Niks R, Stam P, Lindhout P (1998) Identification of QTLs for partial resistance to leaf rust (Puccinia hordei) in barley. Theor Appl Genet 96:1205–1215
Qi X, Fufa F, Sijtsma D, Niks R, Lindhout P, Stam P (2000) The evidence for abundance of QTLs for partial resistance to Puccinia hordei on the barley genome. Mol Breed 6:1–9
Ruan Y, Jin D (1983) On Barley yellow mosaic virus (BaYMV). Acta Phytopathol Sin 13:49–55
Rubies-Autonell C, Toderi G, Marenghi A, Vallega V (1995) Effects of soil tillage and crop rotation on BaYMV and BaMMV mixed infection. Agronomie 15:511–512
Ruge B, Linz A, Pickering R, Proeseler G, Greif P, Wehling P (2003) Mapping of Rym14 Hb, a gene introgressed from Hordeum bulbosum and conferring resistance to BaMMV and BaYMV in barley. Theor Appl Genet 107:965–971
Ruge B, Linz A, Pickering R, Proeseler G, Greif P, Wehling P (2004) Mapping of Rym14 Hb, a gene introgressed from Hordeum bulbosum and conferring resistance to BaMMV and BaYMV. Theor Appl Genet 107:965–971
Ruge-Wehling B, Linz A, Habehuß A, Wehling P (2006) Mapping of Rym16 Hb, the second soil-borne virus-resistance gene introgressed from Hordeum bulbosum. Theor Appl Genet 113:867–873
Scholz M, Ruge-Wehling B, Habekuß A, Schrader O, Pendinen G, Fischer K, Wehling P (2009) Ryd4Hb: a novel resistance gene introgressed from Hordeum bulbosum into barley and conferring complete and dominant resistance to the barley yellow dwarf virus. Theor Appl Genet 119:837–849
Sharma-Poudyal D, Chen XM, Wan AM, Zhan GM, Kang ZS, Cao SQ, Jin SL, Morgounov A, Akin B, Mert Z, Shah SJA, Bux H, Ashraf M, Sharma RC, Madariaga R, Puri KD, Wellings C, Xi KQ, Wanyera R, Manninger K, Ganzález MI, Koyda M, Sanin S, Patzek LJ (2012) Virulence characterization of international collections of the wheat stripe rust pathogen, Puccinia striiformis f. sp. tritici. Plant Dis 97:379–386
Shtaya M, Sillero J, Flath K, Pickering R, Rubiales D (2007) The resistance to leaf rust and powdery mildew of recombinant lines of barley (Hordeum vulgare L.) derived from H. vulgare × H. bulbosum crosses. Plant Breed 126:259–267
Singh RP, Huerta-Espino J (1997) Effect of leaf rust resistance gene Lr34 on grain yield and agronomic traits of spring wheat. Crop Sci 37:390–395
Singh A, Knox RE, DePauw RM, Singh AK, Cuthbert RD, Campbell HL, Singh D, Bhavani S, Fetch T, Clarke F (2013a) Identification and mapping in spring wheat of genetic factors controlling stem rust resistance and the study of their epistatic interactions across multiple environments. Theor Appl Genet 126:1951–1964
Singh D, Macaigne N, Park RF (2013b) Rph20: adult plant resistance gene to barley leaf rust can be detected at early growth stages. Eur J Plant Pathol 137:719–725
Toubia-Rahme H, Johnston P, Pickering R, Steffenson B (2003) Inheritance and chromosomal location of Septoria passerinii resistance introgressed from Hordeum bulbosum into Hordeum vulgare. Plant Breed 122:405–409
Verbyla AP, Cullis BR, Kenward MG, Welham SJ (1999) The analysis of designed experiments and longitudinal data by using smoothing splines. J Roy Stat Soc Ser C Appl Stat 48:269–311
von Korff M, Wang H, Leon J, Pillen K (2006) AB-QTL analysis in spring barley: II. Detection of favourable exotic alleles for agronomic traits introgressed from wild barley (H. vulgare ssp. spontaneum). Theor Appl Genet 112:1221–1231
Walther U, Rapke H, Proeseler G, Szigat G (2000) Hordeum bulbosum—a new source of disease resistance—transfer of resistance to leaf rust and mosaic viruses from H. bulbosum into winter barley. Plant Breed 119:215–218
Wang L, Wang Y, Wang Z, Marcel T, Niks R, Qi X (2010) The phenotypic expression of QTLs for partial resistance to barley leaf rust during plant development. Theor Appl Genet 121:857–864
Xu J, Kasha KJ (1992) Transfer of a dominant gene for powdery mildew resistance and DNA from Hordeum bulbosum into cultivated barley (H. vulgare). Theor Appl Genet 84:771–777
Ziems LA, Hickey LT, Hunt CH, Mace ES, Platz GJ, Franckowiak JD, Jordan DR (2014) Association mapping of resistance to Puccinia hordei in Australian barley breeding germplasm. Theor Appl Genet 127:1199–1212
Acknowledgments
The authors wish to acknowledge the efforts of Andy Hay, Russell Harrison-Kirk and their team for sowing, managing and harvesting the field trials, Dr. Soonie Chng for glasshouse leaf rust screening of line ‘372E’ and Donna Gibson for formatting of Fig. 1. Also thanks go to Dr Samantha Baldwin, Andy Hay and Dr. Bill Griffin for critical reading of this manuscript. Funding for this research was initially provided by the New Zealand Foundation for Research, Science and Technology contract number C06X0810 and more recently supported by Plant and Food Research Core funding.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Komatsuda.
R. Pickering: retired, formerly of The New Zealand Institute for Plant and Food Research Limited.
Rights and permissions
About this article
Cite this article
Johnston, P.A., Meiyalaghan, V., Forbes, M.E. et al. Marker assisted separation of resistance genes Rph22 and Rym16 Hb from an associated yield penalty in a barley: Hordeum bulbosum introgression line. Theor Appl Genet 128, 1137–1149 (2015). https://doi.org/10.1007/s00122-015-2495-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2495-z