Abstract
Single-cell sequencing is widely used in biological and medical studies. However, its application with multiple samples is hindered by inefficient sample processing, high experimental costs, ambiguous identification of true single cells, and technical batch effects. Here, we introduce sample-multiplexing approaches for single-cell sequencing in transcriptomics, epigenomics, genomics, and multiomics. In single-cell transcriptomics, sample multiplexing uses variants of native or artificial features as sample markers, enabling sample pooling and decoding. Such features include: (1) natural genetic variation, (2) nucleotide-barcode anchoring on cellular or nuclear membranes, (3) nucleotide-barcode internalization to the cytoplasm or nucleus, (4) vector-based barcode expression in cells, and (5) nucleotide-barcode incorporation during library construction. Other single-cell omics methods are based on similar concepts, particularly single-cell combinatorial indexing. These methods overcome current challenges, while enabling super-loading of single cells. Finally, selection guidelines are presented that can accelerate technological application.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cells are the basic structural and functional units of life, and multicellular organisms are composed of individual cells with varied gene expression profiles and functions. Single-cell RNA sequencing (scRNA-seq) has been developed rapidly since its inception [1]. In contrast to traditional bulk transcriptional profiling, scRNA-seq enables transcriptomic analysis of numerous individual cells in parallel [2,3,4], thereby helping to elucidate cellular heterogeneity and characterize rare cells in research related to tumors [5, 6], stem cells [7], the immune microenvironment [8], neurobiology [9,10,11], reproduction [12], and embryonic development [13,14,15,16,17]. Commercial scRNA-seq platforms such as 10 × Genomics Chromium [18] and BD Rhapsody [19, 20] routinely provide high throughput (103–105 cells) [21], which has catalyzed the development of numerous cell atlas projects, such as The Human Cell Atlas [22], The Fly Cell Atlas [23], and Plant Cell Atlas [24].
While the throughput has increased and the per-cell cost has fallen with rapid advancements in scRNA-seq, several major challenges still remain when processing multiple samples, which impede further adoption and evolution. In general, performing scRNA-seq for multiple individual samples is labor-intensive and requires expensive reagents and consumables. In the early stage, scientists eliminated false singlets (negatives and multiplets) subjectively using the unique molecular identifier (UMI) “cut-off,” resulting in the retention of false single cells and the loss of true cells with RNA contents that are too high or low. Afterward, more and more R packages have been developed to remove negatives or multiplets, such as scrublet [25], doubletFinder [26], and scuttle. However, these statistical methods can only partially solve the problem. Additionally, technical batch effects can mask biological signals when scRNA-seq profiles are integrated [27]. Fortunately, effective solutions to mitigate these challenges have been successfully applied.
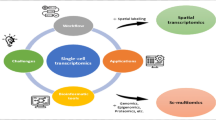
Here, we principally discuss sample-multiplexing approaches for scRNA-seq benefiting from bioinformatic and biochemical techniques. These methods overcome common challenges facing scRNA-seq and enable the super-loading of single cells in a single run. More single cells can be obtained by super-loading, which is especially important for the detection and characterization of some rare cell subsets. However, uncontrolled overloading when too many samples are multiplexed will lead to an ultra-high rate of multiplets, which although can be removed when the data is obtained but will consume sequencing power. Although scRNA-seq is the most sophisticated solution, other methods of single-cell omics are gradually being developed with comparable strengths in terms of their applications. Therefore, we also discuss other sample-multiplexing methods, such as single-cell assay for transposase-accessible chromatin with high-throughput sequencing (scATAC-seq), single-cell whole-genome sequencing (scWGS), single-cell DNA-methylation analysis, single-cell high-throughput chromosome conformation capture (scHi-C), and single-cell multi-omics (scMulti-omics) (Fig. 1, Table 1). Finally, we provide selection guidelines and application models for these multiplexing methods.
A Hierarchy of sample-multiplexing approaches used for single-cell sequencing. The different shapes represent different strategies, which are also mentioned in (B). B Timeline of sample-multiplexing approaches for single-cell sequencing. Omics targeted are distinguished by various colors. The strategies are represented by different shapes
Sample multiplexing for single-cell transcriptome sequencing
According to the central dogma of molecular biology, RNA bridges the gap between DNA and proteins, thereby reflecting the nature and genetic profiles of cells. Proteins are difficult to amplify and sequence, while high-throughput scWGS and scWES, requiring a large amount of sequencing, are too expensive to apply. Therefore, scRNA-seq is currently the most popular single-cell omics tool. Here, we present five categories of scRNA-seq sample-multiplexing approaches (Fig. 2).
Schematic overview of five sample-multiplexing strategies used for scRNA-seq. A Natural genetic variation. Without additional labeling, computational demultiplexing is conducted based on SNPs. B Nucleotide-barcode anchoring on cellular or nuclear membranes. The example shown here is Cell Hashing, where oligo-tagged antibodies (hashtags) bind to ubiquitously expressed cell-surface proteins. Oligos with a poly (A) tail are captured along with mRNA. Cells can be assigned to their sample of origin based on different barcodes in the hashtags. C Nucleotide-barcode internalization into the cytoplasm or nucleus. Barcoded DNA traverses the cellular or nuclear membrane by liposomal transfection or directly diffuses into the nuclei. SBO: short barcode oligonucleotide. D Vector-based barcode expression in cells. E Nucleotide-barcode incorporation during library construction
Natural genetic variation
Demultiplexing scRNA-seq data by genetic variations with reference genotypes
Genetic variation, particularly single-nucleotide polymorphisms (SNPs), distinguishes different individuals, qualifying SNPs as natural barcodes for assigning cells to donors and identifying doublets from different donors (Fig. 2A). Multiple methods for demultiplexing cells from different individuals based on SNPs have been developed in recent years. Kang et al. [28] introduced demuxlet, the first method for harnessing genetic barcodes to identify cells. The demuxlet method assigns each cell to its individual source by calculating the maximum likelihood of obtaining RNA-seq reads with overlapping SNP sets, the genotypes of which are given in parallel as a reference for each individual. Using simulated data, demuxlet showed that 50 genetic variations are sufficient to assign cells for up to 64 individuals. In the same study, the performance of demuxlet was evaluated by analyzing pooled peripheral blood mononuclear cells (PBMCs) from eight patients with lupus.
Demultiplexing scRNA-seq data by genetic variations without reference genotypes
While the above-mentioned methods enable demultiplexing of mixed samples, they require additional genotyping information to assign individual cells back to the donors. These factors limit the utility of genotype-specific methods, as genotype data might be unavailable, or insufficient tissue may be available for DNA extraction. Recently, approaches were developed that do not require a genotype reference. Xu et al. [29] introduced scSplit, a hidden-state model approach for demultiplexing individual samples from mixed scRNA-seq data without extra genotype information. ScSplit identifies putative variant sites from scRNA-seq data and models the allelic counts into clusters using an expectation–maximization framework. This tool requires only FASTQ files and a cell barcode list as inputs. Huang et al. [30] presented Variational Inference for Reconstructing Ensemble Origins (Vireo), a principled Bayesian method for demultiplexing pooled samples independently of the genomic information. Vireo can also leverage genotype information when available (termed Vireo-GT). They evaluated the performance of Vireo-GT for singlet assignment. As expected, Vireo-GT performed slightly better than Vireo when using 16 genetically distinct scRNA-seq samples (AUCVireo = 0.999, AUCVireo-GT = 1.000). More recently, the computational tool, Souporcell, was designed for clustering scRNA-seq data by genotype without reference genotypes [31]. Souporcell fits a mixture model using a deterministic annealing variant of the expectation–maximization algorithm to cluster cells. The advantage of mixture model clustering over hard clustering is that cells can be assigned to different clusters, benefiting both doublet calling and ambient RNA estimations. Souporcell achieved high accuracy in terms of genotype clustering, doublet detection, and ambient RNA estimation. This method even surpassed demuxlet (the gold-standard method requiring genotype information a priori) in cell assignments and doublet accuracies. When more challenging scenarios involving multiple cell types are involved, 21 cells that demuxlet had labeled as maternal or fetal appeared in the cell clusters of the counterpart individual by Souporcell. Based on the reference genotypes, they are of an error by demuxlet assignments rather than that by Souporcell. Notably, Souporcell includes a low signal-to-noise ratio due to decreased numbers of UMIs per cell and high numbers of donors, resulting in increased local maxima.
Nucleotide-barcode anchoring on cellular or nuclear membranes
Oligo-tagged antibodies
Cells with higher expression levels of specific surface proteins can be labeled with more corresponding antibodies. Therefore, the cellular indexing of transcriptomes and epitopes by sequencing (CITE-seq) [32] and RNA-expression and protein-sequencing assay (REAP-seq) [33] approaches can simultaneously measure gene- and protein-expression levels in single cells with oligo-tagged antibodies. These antibodies link ssDNA instead of the fluorophores applied to Fluorescence Activating Cell Sorter (FACS), and barcoded ssDNA integrated with antibodies will be captured along with mRNAs. Stoeckius et al. used oligo-tagged antibodies (hashtags) that bound to ubiquitous cell-surface membrane proteins to mark and pool experimental samples, demultiplexed the data according to the barcoded antibody signals, and assigned cells to the original samples (Fig. 2B). This method was named “Cell Hashing,” as it is based on the concept of hash functions in computer science to index datasets with specific features. Eight human PBMC samples were labeled, pooled, and analyzed [34]. Class-I major histocompatibility complex, β-2-microglobulin, and the sodium–potassium ATPase-subunit CD298 are broadly expressed on the surface of human cells [34], and combining these two corresponding antibodies fulfills the purpose of double assurance for labeling all cells successfully.
The performance of single-nucleus RNA sequencing (snRNA-seq) has been compared with that of scRNA-seq [35,36,37]. Although snRNA-seq usually captures fewer genes, it has significant advantages in addressing complex tissues that are challenging to dissociate or are frozen, or comprise large or irregularly shaped cells, while avoiding bias caused by the loss of certain cells during single-cell suspension preparation [38,39,40]. Following Cell Hashing, Nucleus Hashing was developed for multiplex snRNA-seq and to profile human brain cortex samples [41]. The Nucleus Hashing approach utilizes DNA-barcoded antibodies to target the nuclear pore complex, which has a relatively conserved sequence between species and expands the application range.
Commercial oligo-tagged antibodies produced by BioLegend and BD are currently available and should streamline such research. However, some cells may not express the nearly ubiquitous targeted surface proteins, resulting in labeling and decoding failures.
Lipid-tagged indices
Lipid and cholesterol are basic cell-membrane components that are required for all life. Lipid- and cholesterol-modified oligonucleotide (LMOs and CMOs, respectively) scaffolds, used as “anchors,” can be rapidly and stably integrated into live cell membranes [42]. McGinnis et al. adapted LMOs and CMOs into MULTI-seq, a multiplexing method for scRNA-seq and snRNA-seq using lipid-tagged indices. Briefly, conjugation with a 5′ lignoceric acid amide and a 3′ palmitic acid amide increased the hydrophobicity of LMOs and enabled them to stably associate with membranes. MULTI-seq sample barcodes are constituted of a 3′ poly-A capture sequence, an 8-bp sample barcode, and a 5′ polymerase chain reaction (PCR) handle. In their study, McGinnis et al. used MULTI-seq to reveal the dynamics of T-cell activation with Jurkat cells, to perform a 96-plex perturbation assay with primary human mammary epithelial cells, and to perform multiplex analysis of cryopreserved primary tumors and metastatic lungs dissected from the patient-derived xenograft (PDX) mice of triple-negative breast cancer [43].
For practical applications, one type of LMO scaffold is sufficient to assemble with different DNA barcodes, increasing the method’s capability for multiplexing numerous samples in parallel. Furthermore, this method is not limited by species or genetic background differences and has little impact on cell viability and endogenous gene expression patterns because the labeling is extremely mild and requires only 5 min.
Concanavalin A (ConA)-based sample barcoding
Glycoproteins are ubiquitous on the plasma membrane. Based on the glycoprotein-binding ability of Concanavalin A, a ConA-based sample-barcoding strategy (CASB) was developed by Fang et al. Three components, i.e., biotinylated single-stranded DNA (ssDNA; as barcodes), streptavidin, and biotinylated Concanavalin A, self-assemble into the CASB complex. CASB has been leveraged to dissect the transcriptomic dynamics of MDA-MB-231 cells perturbed with five compounds and to demonstrate the IFN-γ-mediated epigenomic and transcriptomic changes in HAP1 cells by combining scRNA-seq and scATAC-seq [44].
CASB enables cell and nucleus labeling independently of the genetic background. In addition, the CASB complex binds to the target quickly and stably, even at low temperatures, without degrading the sample integrity. ssDNA barcode synthesis and complex assembly are extraordinarily convenient, flexible, and easy to be adapted to different single-cell sequencing workflows. Furthermore, CASB combines simultaneously combinatorial barcoding and sequential split-pool barcoding, making it capable of multiplexing more samples and further eliminating multiplets than previous methods.
ClickTags
Methyltetrazine-modified oligonucleotides can be coupled with surface proteins on methanol-fixed cells via a two-step chemical reaction using inverse electron-demand Diels–Alder chemistry and the heterobifunctional amine-reactive cross-linker NHS-trans-cyclooctene. Using these modified oligonucleotides as “ClickTags,” Gehring et al. achieved highly multiplexed scRNA-seq with fixed cells. They applied this strategy in a 96-plex perturbation experiment using neural stem cells treated with various concentrations of decitabine and Scriptaid, epidermal growth factor and basic fibroblast growth factor, retinoic acid, and bone morphogenic protein 4 [45].
However, the method is currently only suitable for methanol-fixed cells (rather than live cells) because competitive NHS-ester hydrolysis in aqueous buffer tends to result in a poor signal-to-noise ratio with scRNA-seq libraries.
Nucleotide-barcode internalization to the cytoplasm or nucleus
Liposomal transfection with short barcoding oligos
Similar to bio-membranes, liposomes are widely used as vectors to transport drugs and exogenous nucleic acids into cells in research and clinical applications. Shin et al. established a universal sample-barcoding method using liposomal transfection with short barcode oligonucleotides (SBOs; Fig. 2C). SBOs consisting of a barcode and a poly-A sequence were implemented to label various samples. They applied this method in a 5-plex time-course experiment using K562 cells treated with a BCR-ABL-targeting drug (imatinib) and in a 48-plex assay performed to screen 45 selected drugs [46].
This method is characterized by a simple experimental process and low-cost sample labeling. SBO synthesis is flexible and convenient, and the number of labeled samples can be largely expanded; therefore, this strategy is very suitable for conducting numerous parallel experiments, such as broad-spectrum drug-screening assays. As this method is based on liposomal transfection, it can be universally adapted to various species and potentially applied to the nucleus. However, additional perturbations may occur when SBOs are transfected using Lipofectamine 3000 at 4 h before library construction.
Staining nuclei with polyadenylated ssDNA oligonucleotides
Evidence suggests that ssDNA selectively diffuses into the nuclei of permeabilized cells instead of intact cells (Fig. 2C). Therefore, Srivatsan et al. used a straightforward approach with unmodified polyadenylated ssDNA oligonucleotides (hash oligos) to mark nuclei followed by single-cell combinatorial indexing RNA-seq (sci-RNA-seq). With this nuclear hashing strategy (known as the “sci-Plex” strategy), the combination of two oligos (one for plates and another for wells) contributed to the exponential scalability of sample indexing. Using sci-Plex, they acquired data from approximately 650,000 single cells across nearly 5000 independent samples in one experiment by screening three cancer cell lines exposed to 188 compounds at different doses [47].
The labeling capacity of sci-Plex is extremely high. Nonetheless, it functions only in the nuclei and remains limited by significantly low mRNA-capture efficiency in sci-RNA-seq.
Vector-based barcode expression in cells
Lentiviral barcodes facilitate cell tagging, pooling, and tracking
In previous studies, virus transduction along with high-throughput scRNA-seq was successfully used for lineage tracing. Subsequently, Guo et al. adapted this method to develop a novel lentiviral barcode-based multiplexing approach, called CellTag Indexing (Fig. 2D). With the design, 8-nt CellTags representing distinct sample indices are located in the 3′ untranslated region of the green fluorescent protein gene (which enables measurement of the transduction efficiency), followed by an SV40 polyadenylation signal sequence. Barcoded viruses are used to transduce the cells to be tagged. Guo et al. used this method to profile cultured cells in vitro and track cell engraftment and differentiation in vivo [48].
CellTag Indexing has been used to successfully achieve long-term stable tracing and multiplexing of live cells. However, it is incompatible with clinical samples and frozen tissues because CellTag barcodes need to be expressed in immortalized cells. Additionally, compared to other labeling technologies, the tagging method is time-consuming due to the need for transducing cells with viruses and expressing viral RNA.
Clustered regularly interspaced short palindromic repeats (CRISPR) and short-hairpin RNA (shRNA) enable multiplexed single-cell screening
Genetic screens enable gene-function analysis and accelerate biological discoveries, while confronting trade-offs between the number of perturbations probed and the complexity of phenotypes evaluated. To bridge this gap, Dixit et al. developed Perturb-seq by combining scRNA-seq and CRISPR-based perturbations. In their experiments, cells were infected with a pool of lentiviruses that expressed single-guide RNAs and guide barcodes to be captured together with mRNA during reverse transcription. They applied Perturb-seq to profile 200,000 cells, focusing on transcription factors that regulate the responses of dendritic cells to lipopolysaccharide [49]. In a companion study, Perturb-seq was used with CRISPR interference to investigate the mammalian unfolded-protein response [50]. Similarly, Jaitin et al. developed CRISP-seq to dissect immune circuits [51], and Datlinger et al. developed CROP-seq (also known as CRISPR droplet sequencing) [52]. Consequently, Daniel and Allon co-authored a paper entitled, “Genetic screening enters the single-cell era” [53].
shRNA is more stable and lasts longer in cells than small interfering RNA and can be expressed from lentiviruses. Thus, Aarts et al. integrated scRNA-seq with a shRNA screen to investigate senescence-based restraint of cell-fate conversion when expression of the transcription factors OCT4, SOX2, KLF4, and c-MYC reprograms somatic cells into induced pluripotent stem cells. Their findings explained why mTOR plays a dual role and how it imposes opposing effects on reprogramming by regulating senescence [54].
A large number of exploratory conditions are analyzed in parallel with CRISPR or shRNA screens and high-throughput scRNA-seq. Therefore, CRISPR and shRNA screens are essentially a replacement of, or kind of alternative of sample-multiplexing strategies. In addition, different combinations of gene knockout can be used as different sample barcodes thus adapted to conventional sample multiplexing for scRNA-seq.
Incorporating nucleotide barcodes during library construction
Barcode assembly for targeted sequencing (BART-Seq)
Uzbas et al. developed BART-Seq as a sample-multiplexing technique for targeted transcriptome or genome sequencing at single-cell level or in bulk cells. Based on their design, in 96-well plates, forward and reverse primers for targeted genes were hybridized with partially complementary ssDNA barcodes, followed by fill-in reactions and strand removal, resulting in the formation of dual barcoded primers. The target regions of genes captured by these barcoded primers were subjected to PCR and next-generation sequencing. They used BART-Seq to profile thousands of single human pluripotent stem cells exposed to different maintenance media and activation of the Wnt/β-catenin pathway [55].
In terms of targeted sequencing, BART-Seq is cost-effective, highly suitable for transcriptome and genome analysis of genes of interest, and can be generalized for other omics methods in the future. Moreover, BART-seq can be used to analyze a broader range of RNA species (more than just mRNAs), including non-polyadenylated long non-coding RNAs. However, when detecting limited genes, BART-Seq fails to cover the whole transcriptome or genome. Furthermore, UMIs have not been adapted to BART-Seq, which hinders counting of the absolute number of transcripts.
Single-cell combinatorial fluidic indexing (scifi)-RNA-seq
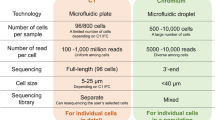
Loading a single-cell suspension into a microfluidic device at very low concentrations reduces the probability that two cells enter the same droplet (i.e., a doublet) as much as possible, resulting in waste as most “empty emulsion droplets” enclose fully functional barcoded microbeads and reverse transcription reagents, but no cells, thereby failing to capture any valid single-cell transcriptomes. For example, ~ 90–99% of gel bead-in-emulsions contain no cells when generated using the 10 × Genomics Chromium Single Cell 3' Reagent Kit v2, with a pool of ~ 750,000 barcodes to separately index each cell’s transcriptome, and possibly more exist with the v3 and v3.1 kits, with ~ 3,500,000 10 × barcodes. Inspired by combined indexing protocols [10,11,12], Paul et al. established the ultra-high-throughput scifi-RNA-seq system. Whole-transcriptome preindexing (round 1) was conducted by capturing barcoded primers and conducting reverse transcription inside permeabilized cells or nuclei, in “split pools,” within 96-well or 384-well plates (Fig. 2E). Following preindexing, cells or nuclei were pooled, and standard droplet-based scRNA-seq was performed with overloading and another round of indexing (round 2); thus, most droplets encapsulated several cells or nuclei (Fig. 3). Subsequently, many effective droplets, including those containing multiplets or singlets, were obtained because the combination of the two barcodes (round 1 and round 2) reached a huge quantity that almost all single cells received a unique identity tag. Paul et al. applied scifi-RNA-seq in various human and mouse cell lines, primary human T cells, and a highly multiplexed CRISPR screen for T-cell receptor activation [56].
Main processes during library construction of the “sci family.” Each round of indexing provides an opportunity for sample multiplexing to occur, especially in round 1. With the inherent ability of sample multiplexing, combinatorial indexing occupies an important position in multiplexed single-cell sequencing
Using scifi-RNA-seq, 151,788 single cells were identified in four pooled human cell lines, a 15-fold increase over the recommended output on the 10 × Genomics Chromium system, further demonstrating that 1.53 million nuclei could be loaded into a single channel if necessary [54]. Undoubtedly, scifi-RNA-seq has built-in support for sample multiplexing because of the different reverse transcription primers used in the round-1 barcoding process. Additionally, scifi-RNA-seq technology can potentially function as an alternative system for scRNA-seq, including subnanoliter well plates, and also enhance the throughput of other single-cell omics sequencing approaches. Notably, permeabilized cells or nuclei (but not live cells) are required for scifi-RNA-seq.
Single-cell combinatorial indexing (sci)-RNA-seq and split-pool ligation-based transcriptome sequencing (SPLiT-seq)
In sci-RNA-seq, fixed and permeabilized cells (or nuclei) are implemented in two or three rounds of split-pool barcoding in 96- or 384-well plates (Fig. 3); therefore, large-scale combinatorial barcodes are sufficient to label a considerable number of single cells. Using this method, approximately 50,000 Caenorhabditis elegans cells (covering their somatic cell compositions) were investigated [57]. This technology has since been upgraded to sci-RNA-seq3, where a “mouse organogenesis cell atlas” was constructed within the critical window profiling of approximately 2 million cells [58]. Similar to sci-RNA-seq, SPLiT-seq involves four rounds of combinatorial barcoding (Fig. 3), and 156,049 single nuclei from mouse brains and spinal cords were analyzed [59].
When performing sci-RNA-seq or SPLiT-seq, each round of indexing gives sample multiplexing a chance, especially in round 1. In addition, neither method requires expensive platforms or regents. However, sci-RNA-seq and SPLiT-seq capture fewer genes and currently require more experimental time relative to popular commercial scRNA-seq technologies, which makes them less preferable options.
Sample multiplexing for single-cell epigenome and genome sequencing
Below, we present sample-multiplexing approaches for single-cell epigenome and genome sequencing together, which share considerable similarities.
Sample multiplexing for scATAC-seq
Chromatin accessibility is vital for regulating gene expression. ATAC-seq provides information regarding cis-acting DNA elements, including promoters, enhancers, and silencers, and the binding of trans-acting factors to DNA in open chromatin regions [60, 61]. Here, we introduce three strategies for sample multiplexing in scATAC-seq.
sci-ATAC-seq
Most single-cell technologies are based on the principle that single cells are individually compartmentalized, and the early library construction proceeds in independent biochemical reaction systems that each cell owns, making it difficult to simultaneously handle massive numbers of single cells per assay. Fortunately, single-cell combinatorial indexing as adapted to scATAC-seq (termed sci-ATAC-seq) overcomes this limitation. In this method, nuclei are molecularly tagged with barcoded Tn5 transposases in each of the wells, pooled, and then randomly redistributed into a second set of wells so that a second barcode is introduced during PCR (Fig. 3). Over 15,000 GM12878 and HL60 cells have been profiled using sci-ATAC-seq [62].
This method enables sample multiplexing but captures a relatively limited number of fragments from nucleosome-free regions per nucleus compared to common droplet-based scATAC-seq technology.
Droplet-based scATAC-seq (dscATAC-seq) combined with combinatorial indexing (dsciATAC-seq)
To boost the breadth and depth of chromatin accessibility profiling, Lareau et al. developed dscATAC-seq technology, where a microfluidic device was utilized to simultaneously encapsulate PCR reagents and barcoded beads into a single droplet with nuclei that were transposed using the Tn5 transposon to link sequencing adaptors into open chromatin regions. Based on this technology, they further designed dsciATAC-seq by combining dscATAC-seq with combinatorial indexing through barcoded transpositions, which is conceptually parallel to scifi-RNA-seq, thereby massively scaling up throughput (up to 105 single cells/experiment; Fig. 3). Subsequently, they applied dscATAC-seq or dsciATAC-seq to assay 46,653 cells from an adult mouse brain, or 136,463 resting and stimulated human bone marrow-derived cells, respectively [63].
dscATAC-seq produced high-quality data with 105 nuclear fragments/cell, while dsciATAC-seq increased the cell throughput tenfold using 24 transposon barcodes, with accessibility to larger output with 48 or 96 barcodes. Additionally, barcoded Tn5 transposon provides an opportunity to multiplex samples for scATAC-seq [63].
CASB for scATAC-seq
CASB has also been designed to suit sample multiplexing for both scATAC-seq and scRNA-seq. Indeed, the CASB barcodes remained abundant and showed minimal cross-contamination after a reaction of 1 h transposition at 37 °C, without influencing the epigenomic profiles. In the workflow, a 222-nt ssDNA barcode, with S5-ME and S7-ME adapter sequences attached by primers during scATAC-seq library amplification, flanked a sequence containing sample barcodes [44].
Wang et al. developed SNuBar to multiplex scATAC-seq and scATAC and RNA co-assay with unmodified oligonucleotides [64]. Furthermore, in Wang’s study, SNPs-based multiplexing was successfully used to verify the reliability of SNuBar, and a high correlation was acquired between two methods, which inspires us that sample multiplexing depending on natural genetic variation is also suitable for single-cell epigenome and genome sequencing.
Sample multiplexing for scWGS
Whole-genome sequencing facilitates genome assembly from various species and provides information pertaining to genetic variations, including single-nucleotide variants (SNVs), small indels, and copy-number variants (CNVs) [65]. scWGS is advancing in two directions: deep sequencing for detecting SNVs and shallow sequencing to identify CNVs and aneuploidy [66].
sci-ATAC-seq only targets open chromatin regions (1–4% of the genome), which is desirable for epigenetic profiling. However, this property or restriction is confusing when detecting single-cell CNVs because of biological bias and severely limits read counts (~ 3000/per cell). Therefore, Vitak et al. designed two strategies to unbind nucleosomes from genomic DNA without disturbing the nuclear integrity. One is lithium-assisted nucleosome depletion (LAND), which employs the chaotropic agent lithium diiodosalicylate to disrupt DNA–protein interactions, thereby releasing DNA from histones. The other is cross-linking with sodium dodecyl sulfate (xSDS), utilizing the detergent to denature histones and dissociate them from DNA (cross-linking is necessary before denaturation because SDS disrupts the nuclear integrity). They further developed a scWGS method for CNV detection incorporating combinatorial indexed sequencing, termed SCI-seq, which is highly consistent with sci-ATAC-seq during all experimental steps, except for the nucleosome removal step (Fig. 3). Subsequent assessment revealed substantially better coverage uniformity using xSDS than LAND. SCI-seq was used to analyze 16,698 single cells from cultured cell lines, primate frontal cortex tissue, and two human adenocarcinomas [66].
Mission Bio Tapestri platform enables simultaneous single-cell DNA and protein analysis using oligo-tagged antibodies provided by BioLegend. Thus, we suggest that this strategy may be applicable to multiplexed scDNA-seq according to the concept of Cell Hashing.
Sample multiplexing for single-cell DNA-methylation analysis
DNA methylation regulates gene expression by recruiting relevant proteins or by suppressing transcription factor binding to DNA, and 5-methylcytosine is the most common type of DNA methylation in plants and animals [67, 68].
Based on the principle that sodium bisulfite (BS) converts cytosine (but not methylcytosine) to uracil in genomic DNA [69], DNA methylation can be detected at single-base resolution from a limited fraction of the genome to the whole-genome scale using bisulfite sequencing (WGBS) [70], in bulk or at single-cell level (scWGBS) [71,72,73,74,75,76]. To increase the cell-count throughput of scWGBS, Mulqueen et al. established sci-MET, which is highly similar to SCI-seq, with the two largest differences occurring during library construction (i.e., transposition with adaptors depleted of cytosines and BS conversion; Fig. 3). They applied sci-MET to discriminate three human cell lines (after mixing them) and to analyze mouse cortical tissue, which mainly comprised excitatory and inhibitory neuronal cells [77].
Sample multiplexing for scHi-C
Hi-C helps shed light on three-dimensional genome architectures and DNA interactions in eukaryotes. For example, topologically associating domains are visible by Hi-C analysis and frequently involve dynamic promoter–enhancer interactions that influence gene expression [78,79,80,81].
As a type of single-cell combinatorial-indexing technology, sciHi-C has successfully bridged the gap between high-throughput and single-cell chromosome conformation analyses. These experimental procedures were derived mostly from traditional Hi-C assays (such as fixation, digestion, proximity ligation, affinity purification, and library amplification), while utilizing biotinylated bridge adaptors and custom-barcoded Illumina Y-adaptors to link the first and second round of barcodes in two 96-well plates, respectively, mirroring a combinatorial indexing design (Fig. 3). In the proof of concept, sciHi-C was leveraged to generate six libraries including 10,696 single cells from mixed mouse cells (primary mouse embryonic fibroblasts and the “Patski” embryonic fibroblast line) and human cells (HeLa S3, HAP1, K562, and GM12878) [82].
Sample multiplexing for scMulti-omics
Multiomics considers multiple types of molecules simultaneously and generates a comprehensive understanding of biological processes [83, 84]. Here, we introduce some sample-multiplexing ideas for scMulti-omics.
First, Cell Hashing plus CITE-seq (or REAP-seq) enables researchers to pool samples and to simultaneously quantitate mRNA transcripts and proteins in the same cells [32,33,34]. Furthermore, REAP-seq can be merged with expanded CRISPR-compatible cellular indexing of transcriptomes and epitopes by sequencing (ECCITE-seq), which simultaneously profiles proteins, mRNAs, T cell receptors, and CRISPR perturbations [85]. Secondly, sci-CAR jointly profiles single-cell chromatin accessibility and mRNA (CAR) with an inherent ability of combinatorial indexing for sample multiplexing [86] (Fig. 3). Additionally, using the 10 × Genomics Chromium microfluidics platform, Mimitou et al. developed ATAC with select antigen profiling by sequencing (ASAP-seq) to detect proteins, chromatin accessibility, and mitochondrial DNA mutations in thousands of single cells, and gene-expression levels (beyond those three levels) can be measured via DOGMA-seq (named for DNA, RNA, and protein spanning the central dogma of gene regulation) [87]. Similarly, Swanson et al. developed integrated cellular indexing of chromatin landscape and epitopes (ICICLE-seq) and TEA-seq (named for transcriptomics, epitopes, and chromatin accessibility) [88]. Both teams’ methods are amenable to sample multiplexing with barcoded antibodies. Finally, theoretically natural genetic variations can be used to demultiplex scMulti-omics profiles derived from mixed non-isogenic samples.
Most of the multiplexing strategies reviewed above can potentially be integrated with various scMulti-omics research techniques to meet experimental needs.
Selection guidelines and applications of sample-multiplexing approaches for single-cell sequencing
The suitability of a given sample-multiplexing method depends on several factors, namely:
-
(1)
whether the focus of the study is on transcriptomics, epigenomics, genomics, or multiomics;
-
(2)
experimental materials, e.g., Cell Hashing is designed for live cells, ClickTags for fixed cells, Nucleus Hashing and sci-Plex for nuclei, and MULTI-seq and CASB for both cells and nuclei;
-
(3)
the number of samples pooled together, e.g., oligo-tagged antibodies is sufficient for labeling a low number of samples, whereas some sci technologies can label thousands of samples;
-
(4)
the cell count per sample and the median number of genes captured per cell. A limited number of cells per sample with a low median number of captured genes/cell may be acquired using sci-Plex, sci-RNA-seq, or SPLiT-seq, despite their capacity for analyzing a massive number of samples;
-
(5)
research needs. The CRISPR system and shRNAs are appropriate tools for screening gene functions broadly, whereas CellTag is a favorable choice for studying stem cell differentiation; and
-
(6)
extra disturbances. Various labeling strategies and experimental times result in different levels of extra disturbances. For instance, it takes 4 h to conduct liposomal transfection with short barcoding oligos (probably followed by severe impacts), whereas using natural genetic variations minimizes extra disturbances to almost zero without additional labeling procedures. In addition, the factors such as the cost, experimental platform, processing time, and labor are also well worth considering.
Some sample-multiplexing technologies among the mentioned have greatly facilitated single-cell studies. For the purpose of appropriate use, we still need to be aware of their respective technical highlights and potential caveats (Table 2). For example, demuxlet has the prominent advantage that avoids additional process of labeling, and additional damage of the cells, but is inapplicable when samples share the same genetic background or reference genotypes cannot be obtained. ScSplit, Vireo, and Souporcell have further reduced sequencing costs depending on machine learning rather than reference genotypes, yet researchers still need additional information when they assign cell clusters to the specific samples.
In general, Cell Hashing can label almost all cell types [89]. However, it has been found that combining the two corresponding antibodies (against CD298 and 2M2) is ineffective for labeling a few human tumor cell lines that have been cultured for multiple generations in vitro. It should be noted that the expression of membrane proteins used for sample barcoding in some stem cells is unclear. BioLegend’s antibody combination for mouse Cell Hashing performs poor at marking egg cells, some stem cells, C3 and B16-BL6 melanoma cells. In addition, attention should also be paid to the presence of membrane protein damage in samples that are sensitive to digestion with trypsin. Therefore, it may be a good choice to use FACS to verify whether cells can be labeled before Cell Hashing is applied, and different antibodies may be chosen to label a minority of special cell types.
Compared to Cell Hashing, Nucleus Hashing is more friendly to complex tissues that are challenging to dissociate or are frozen, and the labeling target, nuclear pore complex has a relatively conserved sequence between species thus expanding the application. Nevertheless, Nucleus Hashing generally shows a less effectiveness of nucleus labeling than lipid-tagged methods, due to the destruction of nuclear pore complex during cell lysis and nucleus isolation.
MULTI-seq and 10 × Genomics CellPlex break the limits of species and cell types, but requires high quality of single cell (or nucleus) suspension. 10 × Genomics has reminded on its website that FACS or Fluorescence Activating Nucleus Sorter (FANS) to purify substandard samples before or after labeling is needed to improve the ratio of signal to noise. Recently, Viacheslav et al. compared antibody- and lipid-based multiplexing methods for single-cell RNA-seq, and drew a conclusion that antibody-based hashing is a more efficient protocol on human cell lines and PBMCs, while lipid hashing delivers a better result on nuclei [90]. In addition, our laboratory finds that antibody-based hashing is more stable and effective than lipid hashing when cell viability is low or apparent debris exists in single cell suspension, while lipid hashing suffers from the tendency of cross-contamination which may be attributed to the fluidity of lipids.
Sample-multiplexing approaches for single-cell sequencing have numerous applications, including: (1) massive screening of drugs or genes, (2) studying stem cell differentiation and lineage tracing, (3) constructing spatiotemporal cell atlases for various species, (4) discovering and identifying rare cell subpopulations, and (5) exploring tumor heterogeneity and the immune microenvironment (Fig. 4). These applications are becoming flexible and convenient owing to the native characteristics of sample multiplexing, such as the ability to overload single cells and to reduce experimental costs.
Concluding remarks
Here, we describe sample-multiplexing approaches for single-cell sequencing in transcriptomics, epigenomics, genomics, and multiomics, illustrate their respective strengths and disadvantages, and briefly provide selection guidelines and applications. Researchers are paying increasing attention to these methods, and innovations such as single-nucleus barcoding (SNuBar) [64] are being developed continuously. Sample multiplexing makes library construction more economically efficient while the cost of sequencing is still decreasing, which is very significant when a large number of samples or/and cells are desired in analysis, thereby sample multiplexing increases its impact on the broad fields of life science research.
Obviously, single-cell transcriptome sequencing provides the maturest solution to profile cell states and molecular characterizations by measuring mRNA information, and consequently relevant studies have been blossoming. Recently, Cheng et al. summarized sample-multiplexing approaches for scRNA-seq [91], and witnessed the thriving development in this field. Compared to single-cell transcriptome sequencing, there is a significant lag in experimental techniques and analytical pipelines for single-cell epigenome and genome sequencing. And in the latter case, the research costs are generally more expensive; the technological obstacles are even harder, so sample multiplexing is more desired. The good news is that we can see these challenges being overcome, and multiplexed single-cell epigenome and genome sequencing are driving to maturity out of their infancy. Especially, scATAC-seq matching scRNA-seq well has been widely applied and sample-multiplexing methods for it have emerged increasingly. In addition, Mission Bio Tapestri platform provides single-cell DNA-seq and DNA and protein co-assay, probably accessible to sample multiplexing based on the concept of Cell Hashing. Great importance should be attached to multiplexed scMulti-omics, which provides multidimensional and more comprehensive data, deepening the understanding of the central dogma associating with different physiological and pathological status of cells.
Notably, a dozen of popular technologies are available to explore spatial information of tissues, such as Slide-Seq [92], HDST [93], DBiT-seq [94], 10 × Genomics VISIUM, and BGI Stereomics. At present, there is no relevant literature of technology about sample multiplexing for spatial transcriptomics, which is of the future direction. In our laboratory, we have successfully implemented sample multiplexing on 10 × Genomics VISIUM by embedding multiple tissue samples together in OCT, or placing several tissue sections in one capture area. Further progress may include the adaptation of sample multiplexing to many emerging sequencing technologies, such as Cleavage Under Targets and Tagmentation (CUT&Tag) [95]. These elegant and powerful methods will fuel insights into complex biological processes at single-cell level.
Availability of data and material
Not applicable.
References
Tang F et al (2009) mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods 6:377–382. https://doi.org/10.1038/nmeth.1315
Shalek AK et al (2013) Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498:236–240. https://doi.org/10.1038/nature12172
Tang F et al (2011) Development and applications of single-cell transcriptome analysis. Nat Methods 8:S6-11. https://doi.org/10.1038/nmeth.1557
Deng Q et al (2014) Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science (New York, N.Y.) 343:193–196. https://doi.org/10.1126/science.1245316
Zhang X et al (2016) Single-cell sequencing for precise cancer research: progress and prospects. Can Res 76:1305–1312. https://doi.org/10.1158/0008-5472.Can-15-1907
Suvà ML, Tirosh I (2019) Single-cell RNA sequencing in cancer: lessons learned and emerging challenges. Mol Cell 75:7–12. https://doi.org/10.1016/j.molcel.2019.05.003
Wen L, Tang F (2016) Single-cell sequencing in stem cell biology. Genome Biol 17:71. https://doi.org/10.1186/s13059-016-0941-0
Chen H et al (2019) Revolutionizing immunology with single-cell RNA sequencing. Cell Mol Immunol 16:242–249. https://doi.org/10.1038/s41423-019-0214-4
Ofengeim D et al (2017) Single-cell RNA sequencing: unraveling the brain one cell at a time. Trends Mol Med 23:563–576. https://doi.org/10.1016/j.molmed.2017.04.006
Cuevas-Diaz Duran R et al (2017) Single-cell RNA-sequencing of the brain. Clin Transl Med 6:20. https://doi.org/10.1186/s40169-017-0150-9
Mu Q et al (2019) Deciphering brain complexity using single-cell sequencing. Genomics Proteomics Bioinformatics 17:344–366. https://doi.org/10.1016/j.gpb.2018.07.007
Tšuiko O et al (2020) Preimplantation Genetic Testing: single-cell technologies at the forefront of PGT and embryo research. Reproduction (Cambridge, England) 160:A19-a31. https://doi.org/10.1530/rep-20-0102
Shangguan Y et al (2020) Application of single-cell RNA sequencing in embryonic development. Genomics 112:4547–4551. https://doi.org/10.1016/j.ygeno.2020.08.007
Peng G et al (2020) Using single-cell and spatial transcriptomes to understand stem cell lineage specification during early embryo development. Annu Rev Genomics Hum Genet 21:163–181. https://doi.org/10.1146/annurev-genom-120219-083220
Hedlund E, Deng Q (2018) Single-cell RNA sequencing: technical advancements and biological applications. Mol Aspects Med 59:36–46. https://doi.org/10.1016/j.mam.2017.07.003
Huang X et al (2018) High throughput single cell RNA sequencing, bioinformatics analysis and applications. Adv Exp Med Biol 1068:33–43. https://doi.org/10.1007/978-981-13-0502-3_4
Yasen A et al (2020) Progress and applications of single-cell sequencing techniques. Infect Genet Evol 80:104198. https://doi.org/10.1016/j.meegid.2020.104198
Zheng GX et al (2017) Massively parallel digital transcriptional profiling of single cells. Nat Commun 8:14049. https://doi.org/10.1038/ncomms14049
Shum EY et al (2019) Quantitation of mRNA transcripts and proteins using the BD Rhapsody™ single-cell analysis system. Adv Exp Med Biol 1129:63–79. https://doi.org/10.1007/978-981-13-6037-4_5
Svensson V et al (2018) Exponential scaling of single-cell RNA-seq in the past decade. Nat Protoc 13(4):599–604. https://doi.org/10.1038/nprot.2017.149
Valihrach L et al (2018) Platforms for single-cell collection and analysis. Int J Mol Sci. https://doi.org/10.3390/ijms19030807
Regev A et al (2017) The human cell Atlas. Elife. https://doi.org/10.7554/eLife.27041
Li H et al (2021) Fly Cell Atlas: a single-cell transcriptomic atlas of the adult fruit fly. J bioRxiv. https://doi.org/10.1101/2021.07.04.451050
Rhee SY et al (2019) Towards building a plant cell Atlas. Trends Plant Sci 24:303–310. https://doi.org/10.1016/j.tplants.2019.01.006
Wolock SL et al (2019) Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst 8(4):281–291. https://doi.org/10.1016/j.cels.2018.11.005 (e9)
McGinnis CS et al (2019) DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst 8(4):329-337.e4. https://doi.org/10.1016/j.cels.2019.03.003
Hicks SC et al (2018) Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics (Oxford, England) 19:562–578. https://doi.org/10.1093/biostatistics/kxx053
Kang HM et al (2018) Multiplexed droplet single-cell RNA-sequencing using natural genetic variation. Nat Biotechnol 36:89–94. https://doi.org/10.1038/nbt.4042
Xu J et al (2019) Genotype-free demultiplexing of pooled single-cell RNA-seq. Genome Biol 20:290. https://doi.org/10.1186/s13059-019-1852-7
Huang Y et al (2019) Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol 20:273. https://doi.org/10.1186/s13059-019-1865-2
Heaton H et al (2020) Souporcell: robust clustering of single-cell RNA-seq data by genotype without reference genotypes. Nat Methods 17:615–620. https://doi.org/10.1038/s41592-020-0820-1
Stoeckius M et al (2017) Simultaneous epitope and transcriptome measurement in single cells. Nat Methods 14:865–868. https://doi.org/10.1038/nmeth.4380
Peterson VM et al (2017) Multiplexed quantification of proteins and transcripts in single cells. Nat Biotechnol 35:936–939. https://doi.org/10.1038/nbt.3973
Stoeckius M et al (2018) Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol 19:224. https://doi.org/10.1186/s13059-018-1603-1
Wu H et al (2019) Advantages of single-nucleus over single-cell RNA sequencing of adult kidney: rare cell types and novel cell states revealed in fibrosis. J Am Soc Nephrol 30:23–32. https://doi.org/10.1681/asn.2018090912
Slyper M et al (2020) A single-cell and single-nucleus RNA-Seq toolbox for fresh and frozen human tumors. Nat Med 26:792–802. https://doi.org/10.1038/s41591-020-0844-1
Ding J et al (2020) Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol 38:737–746. https://doi.org/10.1038/s41587-020-0465-8
Habib N et al (2017) Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat Methods 14:955–958. https://doi.org/10.1038/nmeth.4407
Nagy C et al (2018) Single-nucleus RNA sequencing shows convergent evidence from different cell types for altered synaptic plasticity in major depressive disorder
Lake BB et al (2018) Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat Biotechnol 36:70–80. https://doi.org/10.1038/nbt.4038
Gaublomme JT et al (2019) Nuclei multiplexing with barcoded antibodies for single-nucleus genomics. Nat Commun 10:2907. https://doi.org/10.1038/s41467-019-10756-2
Weber RJ et al (2014) Efficient targeting of fatty-acid modified oligonucleotides to live cell membranes through stepwise assembly. Biomacromol 15:4621–4626. https://doi.org/10.1021/bm501467h
McGinnis CS et al (2019) MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat Methods 16:619–626. https://doi.org/10.1038/s41592-019-0433-8
Fang L et al (2021) CASB: a concanavalin A-based sample barcoding strategy for single-cell sequencing. Mol Syst Biol 17:e10060. https://doi.org/10.15252/msb.202010060
Gehring J et al (2020) Highly multiplexed single-cell RNA-seq by DNA oligonucleotide tagging of cellular proteins. Nat Biotechnol 38:35–38. https://doi.org/10.1038/s41587-019-0372-z
Shin D et al (2019) Multiplexed single-cell RNA-seq via transient barcoding for simultaneous expression profiling of various drug perturbations. Sci Adv 5:eaav2249. https://doi.org/10.1126/sciadv.aav2249
Srivatsan SR et al (2020) Massively multiplex chemical transcriptomics at single-cell resolution. Science (New York, N.Y.) 367:45–51. https://doi.org/10.1126/science.aax6234
Guo C et al (2019) Cell Tag Indexing: genetic barcode-based sample multiplexing for single-cell genomics. Genome Biol 20:90. https://doi.org/10.1186/s13059-019-1699-y
Dixit A et al (2016) Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167:1853-1866.e1817. https://doi.org/10.1016/j.cell.2016.11.038
Adamson B et al (2016) A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167:1867–1882. https://doi.org/10.1016/j.cell.2016.11.048 (e1821)
Jaitin DA et al (2016) Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-Seq. Cell 167:1883–1896. https://doi.org/10.1016/j.cell.2016.11.039 (e1815)
Datlinger P et al (2017) Pooled CRISPR screening with single-cell transcriptome readout. Nat Methods 14:297–301. https://doi.org/10.1038/nmeth.4177
Wagner DE, Klein AM (2017) Genetic screening enters the single-cell era. Nat Methods 14:237–238. https://doi.org/10.1038/nmeth.4196
Aarts M et al (2017) Coupling shRNA screens with single-cell RNA-seq identifies a dual role for mTOR in reprogramming-induced senescence. Genes Dev 31:2085–2098. https://doi.org/10.1101/gad.297796.117
Uzbas F et al (2019) BART-Seq: cost-effective massively parallelized targeted sequencing for genomics, transcriptomics, and single-cell analysis. Genome Biol 20:155. https://doi.org/10.1186/s13059-019-1748-6
Datlinger P et al (2021) Ultra-high-throughput single-cell RNA sequencing and perturbation screening with combinatorial fluidic indexing. Nat Methods 18:635–642. https://doi.org/10.1038/s41592-021-01153-z
Cao J et al (2017) Comprehensive single-cell transcriptional profiling of a multicellular organism. Science (New York, N.Y.) 357:661–667. https://doi.org/10.1126/science.aam8940
Cao J et al (2019) The single-cell transcriptional landscape of mammalian organogenesis. Nature 566:496–502. https://doi.org/10.1038/s41586-019-0969-x
Rosenberg AB et al (2018) Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science (New York, N.Y.) 360:176–182. https://doi.org/10.1126/science.aam8999
Shashikant T, Ettensohn CA (2019) Genome-wide analysis of chromatin accessibility using ATAC-seq. Methods Cell Biol 151:219–235. https://doi.org/10.1016/bs.mcb.2018.11.002
Yan F et al (2020) From reads to insight: a hitchhiker’s guide to ATAC-seq data analysis. Genome Biol 21:22. https://doi.org/10.1186/s13059-020-1929-3
Cusanovich DA et al (2015) Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science (New York, N.Y.) 348:910–914. https://doi.org/10.1126/science.aab1601
Lareau CA et al (2019) Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat Biotechnol 37:916–924. https://doi.org/10.1038/s41587-019-0147-6
Wang K et al (2021) Simple oligonucleotide-based multiplexing of single-cell chromatin accessibility. Mol Cell 81:4319-4332.e4310. https://doi.org/10.1016/j.molcel.2021.09.026
Ng PC, Kirkness EF (2010) Whole genome sequencing. Methods Mol Biol (Clifton, N.J.) 628:215–226. https://doi.org/10.1007/978-1-60327-367-1_12
Vitak SA et al (2017) Sequencing thousands of single-cell genomes with combinatorial indexing. Nat Methods 14:302–308. https://doi.org/10.1038/nmeth.4154
Moore LD et al (2013) DNA methylation and its basic function. Neuropsychopharmacology 38:23–38. https://doi.org/10.1038/npp.2012.112
Gouil Q, Keniry A (2019) Latest techniques to study DNA methylation. Essays Biochem 63:639–648. https://doi.org/10.1042/ebc20190027
Frommer M et al (1992) A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc Natl Acad Sci USA 89:1827–1831. https://doi.org/10.1073/pnas.89.5.1827
Lister R et al (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462:315–322. https://doi.org/10.1038/nature08514
Smallwood SA et al (2014) Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat Methods 11:817–820. https://doi.org/10.1038/nmeth.3035
Farlik M et al (2015) Single-cell DNA methylome sequencing and bioinformatic inference of epigenomic cell-state dynamics. Cell Rep 10:1386–1397. https://doi.org/10.1016/j.celrep.2015.02.001
Farlik M et al (2016) DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell 19:808–822. https://doi.org/10.1016/j.stem.2016.10.019
Angermueller C et al (2016) Parallel single-cell sequencing links transcriptional and epigenetic heterogeneity. Nat Methods 13:229–232. https://doi.org/10.1038/nmeth.3728
Clark SJ et al (2017) Genome-wide base-resolution mapping of DNA methylation in single cells using single-cell bisulfite sequencing (scBS-seq). Nat Protoc 12:534–547. https://doi.org/10.1038/nprot.2016.187
Luo C et al (2017) Single-cell methylomes identify neuronal subtypes and regulatory elements in mammalian cortex. Science (New York, N.Y.) 357:600–604. https://doi.org/10.1126/science.aan3351
Mulqueen RM et al (2018) Highly scalable generation of DNA methylation profiles in single cells. Nat Biotechnol 36:428–431. https://doi.org/10.1038/nbt.4112
Lieberman-Aiden E et al (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science (New York, N.Y.) 326:289–293. https://doi.org/10.1126/science.1181369
Ramani V et al (2016) Understanding spatial genome organization: methods and insights. Genomics Proteomics Bioinformatics 14:7–20. https://doi.org/10.1016/j.gpb.2016.01.002
Eagen KP (2018) Principles of chromosome architecture revealed by Hi-C. Trends Biochem Sci 43:469–478. https://doi.org/10.1016/j.tibs.2018.03.006
Kong S, Zhang Y (2019) Deciphering Hi-C: from 3D genome to function. Cell Biol Toxicol 35:15–32. https://doi.org/10.1007/s10565-018-09456-2
Ramani V et al (2017) Massively multiplex single-cell Hi-C. Nat Methods 14:263–266. https://doi.org/10.1038/nmeth.4155
Hasin Y et al (2017) Multi-omics approaches to disease. Genome Biol 18:83. https://doi.org/10.1186/s13059-017-1215-1
Chappell L et al (2018) Single-sell (Multi)omics technologies. Annu Rev Genomics Hum Genet 19:15–41. https://doi.org/10.1146/annurev-genom-091416-035324
Mimitou EP et al (2019) Multiplexed detection of proteins, transcriptomes, clonotypes and CRISPR perturbations in single cells. Nat Methods 16:409–412. https://doi.org/10.1038/s41592-019-0392-0
Cao J et al (2018) Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science (New York, N.Y.) 361:1380–1385. https://doi.org/10.1126/science.aau0730
Mimitou EP et al (2021) Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat Biotechnol 39:1246–1258. https://doi.org/10.1038/s41587-021-00927-2
Swanson E et al (2021) Simultaneous trimodal single-cell measurement of transcripts, epitopes, and chromatin accessibility using TEA-seq. Elife. https://doi.org/10.7554/eLife.63632
Hartmann FJ et al (2018) A universal live cell barcoding-platform for multiplexed human single cell analysis. Sci Rep 8(1):10770. https://doi.org/10.1038/s41598-018-28791-2
Mylka V et al (2022) Comparative analysis of antibody- and lipid-based multiplexing methods for single-cell RNA-seq. Genome Biol 23(1):55. https://doi.org/10.1186/s13059-022-02628-8
Cheng J et al (2021) Multiplexing methods for simultaneous large-scale transcriptomic profiling of samples at single-cell resolution. Adv Sci (Weinheim, Baden-Wurttemberg, Germany) 8:e2101229. https://doi.org/10.1002/advs.202101229
Rodriques SG et al (2019) Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 363(6434):1463–1467. https://doi.org/10.1126/science.aaw1219
Vickovic S et al (2019) High-definition spatial transcriptomics for in situ tissue profiling. Nat Methods 16(10):987–990. https://doi.org/10.1038/s41592-019-0548-y
Liu Y et al (2020) High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183(6):1665-1681.e18. https://doi.org/10.1016/j.cell.2020.10.026
Kaya-Okur HS et al (2019) CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun 10:1930. https://doi.org/10.1038/s41467-019-09982-5
Acknowledgements
We thank Kai He from Southern Medical University, and Yanfang Lu from Henan Provincial People’s Hospital for suggestion in the preparation of this manuscript.
Funding
This work was supported by the grants from the Natural Science Foundation of Guangdong Province (Major Basic Cultivation Project 2018B030308004, and Major Projects of Basic and Applied Basic Research 2019B1515120033), the National Nature Science Foundation of China (32071452), the Open Fund Programs of Shenzhen Bay Laboratory (SZBL2020090501003), and the Pearl River Talents Program Local Innovative and Research Teams (2017BT01S131).
Author information
Authors and Affiliations
Contributions
All authors contributed to the review conception and design. Material preparation, data collection, and analysis were performed by YZ, SX, ZW, and JG. The first draft of the manuscript was written by YZ and SX, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts of interest to declare.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Xu, S., Wen, Z. et al. Sample-multiplexing approaches for single-cell sequencing. Cell. Mol. Life Sci. 79, 466 (2022). https://doi.org/10.1007/s00018-022-04482-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04482-0