Abstract
Spermatogenesis is a complex process through which spermatogonial stem cells undergo mitosis, meiosis, and cell differentiation to generate mature spermatozoa. During this process, male germ cells experience several translational modifications. One of the major post-translational modifications in eukaryotes is the ubiquitination of proteins, which targets proteins for degradation; this enables control of the expression of enzymes and structural proteins during spermatogenesis. It has become apparent that ubiquitination plays a key role in regulating every stage of spermatogenesis starting from gonocytes to differentiated spermatids. It is understood that, where there is ubiquitination, deubiquitination by deubiquitinating enzymes (DUBs) also exists to counterbalance the ubiquitination process in a reversible manner. Normal spermatogenesis is dependent on the balanced actions of ubiquitination and deubiquitination. This review highlights the current knowledge of the role of DUBs and their essential regulatory contribution to spermatogenesis, especially during progression into meiotic phase, acrosome biogenesis, quality sperm production, and apoptosis of germ cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intracellular proteins are targeted for degradation by covalent attachment of ubiquitin via a non-lysosomal proteolytic pathway in the ubiquitin proteasome system (UPS). Ubiquitination is a post-translational modification process that regulates diverse arrays of cellular and biological processes such as transcriptional regulation, immune response, embryonic development, cell cycle control, apoptosis, oncogenesis, preimplantation, and intracellular signaling pathways [1]. Ubiquitin is a small molecule that is highly conserved from yeast to humans. It is attached to its substrates as a monomer or polymer. Attachment of a single ubiquitin molecule to one lysine residue within a substrate protein is termed monoubiquitination; this contributes to DNA repair, vesicle sorting, signal transduction, and receptor endocytosis [2–5]. If a chain of ubiquitin molecules is attached to a specific lysine residue within the substrate protein, this is referred to as polyubiquitination, which is important in protein degradation and signal transduction [6].
Ubiquitin contains seven lysine residues, namely lysine-6, -11, -27, -29, -33, -48, and -63, each of which can be used to create ubiquitin–ubiquitin linkages called polyubiquitin chains [7]. Ubiquitin chains are arranged in several different ways that modify target substrates in specific ways. For example, the lysine 63-linked polyubiquitin chain serves primarily as a signal for protein activation or signal transduction. In contrast, the lysine 48-linked polyubiquitin chain is known to signal proteasomal degradation. Ubiquitination is orchestrated by a cascade of enzymes, namely ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3), which transfer ubiquitin onto protein substrates. The ubiquitin conjugation process starts with the activation of the ubiquitin molecule by an E1 enzyme through the formation of an ATP-dependent thiol ester bond between the C-terminus of ubiquitin and the active cysteine site of the E1 enzyme. Next, ubiquitin is transferred to the E2 enzyme through a thioester-linked E2-ubiquitin intermediate. E3 enzyme recruits substrate protein and then interacts with the E2-ubiquitin intermediate and catalyzes the transfer of ubiquitin to a lysine residue on the targeted protein. Finally, lysine-48-linked polyubiquitinated proteins are channeled for ATP-dependent hydrolysis by the 26S proteasome [1]. The complete ubiquitination process is illustrated in Fig. 1. There are two E1 enzymes in mammalian cells (Uba1 and Uba6) and approximately 30 genes encode E2 enzymes, each of which can interact with approximately 800 E3 enzymes [1].
The ubiquitin proteasome system. The process of ubiquitination is regulated by an organized milieu of E1, E2, and E3 enzymes that mediate ligation of ubiquitin to lysine residues in target proteins. Proteins can be either monoubiquitinated or polyubiquitinated. Lysine 48-linked polyubiquitination chain-attached proteins are targeted to the 26S proteasome for degradation. DUB enzymes are involved in recycling of ubiquitin molecules
Deubiquitinating enzymes and their classification
Deubiquitinating enzymes (DUBs) belong to a large family of proteases that reverse the ubiquitination process. DUBs bind to the ubiquitin-based isopeptide bond, thus counteracting ubiquitin–protein ligase activity. DUBs are mainly involved in disassembling ubiquitin–substrate and ubiquitin–ubiquitin covalent links, thus they play a critical role in regulation of the proteasomal pathway. DUBs are involved in four main cellular activities: (1) processing of ubiquitin precursors, (2) recycling of ubiquitin molecules during ubiquitination, (3) editing of ubiquitin chains, and (4) reversal of ubiquitin conjugation [1, 8] (Fig. 2). About 100 DUB genes have been identified from the human genome. These DUBs are cysteine proteases, which are classified into six families: (1) ubiquitin C-terminal hydrolases (UCH), (2) ubiquitin-specific processing proteases (USP), (3) Jab1/Pab1/MPN domain-containing metalloenzymes (JAMM), (4) Otu-domain ubiquitin aldehyde-binding proteins (OTU), (5) Ataxin-3/Josephin, and (6) monocyte chemotactic protein-induced proteases (MCPIPs). Among these, the largest family is the USPs, comprising more than 50 members, each characterized by conserved domains and catalytic sites [1, 9–14].
Although the physiological role of deubiquitination is not well understood, DUB activity and specificity are ensured by protein–protein interactions via protein complexes that interact with DUBs, subcellular localization, variation in expression levels, and differential activity during the various cell cycle phases [1, 8]. Thus, DUBs are involved in the regulation of a variety of cellular functions such as apoptosis, cell cycle progression, protection of proteins from degradation, proteasome- or lysosome-dependent protein degradation, gene expression, chromosome segregation, DNA repair, kinase activation, and localization and degradation of signaling intermediates [1, 8, 12–14].
Spermatogenesis
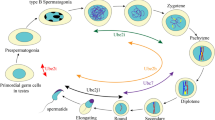
Spermatogenesis is a complex and well-organized process in which a diploid spermatogonium develops into a specialized haploid male gamete, a spermatozoon, through successive rounds of cell division and differentiation. The spermatogenic process involves three phases of development: (1) mitosis: spermatogonial stem cells undergo self-renewal and differentiation, (2) meiosis: primary spermatocytes undergo two meiotic cycles in which a new genome is generated through spermatocyte DNA recombination, resulting in haploid spermatids and (3) spermiogenesis: extensive cell morphological alterations and differentiation of the spermatid into highly specialized spermatozoa, also known as sperm cells, before they are released into the lumen [15]. In humans, the entire process of spermatogenesis takes approximately 74 days [16]. However, spermatogenesis varies widely among individuals; the whole process can take much less time than 74 days [17]. In humans, four types of spermatogonia have been reported: A long, A dark, A pale, and B [15, 18]. Type B spermatogonia undergo mitosis to give rise to primary spermatocytes [19]. In mice, spermatogenesis occurs in the seminiferous tubules, where differentiation of germ cells occurs at different stages in multiple layers. During differentiation, spermatocytes relocate towards the lumen of seminiferous tubules. The basal layers of seminiferous tubules hold all stages of spermatogonia: As (Asingle; singly isolated cells), Apr (Apaired; cysts of interconnected cell pairs), and Aal (Aaligned; interconnected cells in syncytial cysts of 4, 8, 16, and occasionally 32 cells). In addition, differentiation progeny A1, A2, A3, A4, In, and B spermatogonia are also located in the basal layer. At stage VII of the seminiferous epithelial cycle of 8.6 days, the germ cells translocate to the adluminal compartment when entering meiosis, becoming spermatocytes, after which they translocate to the second layer. Post-meiotic round and elongating spermatids migrate through the third and fourth stratified cell layers and finally, after 35 days, the mature sperms are released into the lumen [20–22].
DUBs in spermatogenesis
A growing body of evidences indicates that the UPS plays an important role in cell remodeling and protein turnover in gonocytes [23]. Various UPS enzymes have been reported to be functional at different stages of spermatogenesis, from gonocytes to differentiated spermatids. The expression level of UPS enzymes and their subcellular localization are critical for normal spermatogenesis, while inactivation of these genes results in abnormal progression or the developmental arrest of germ cells, leading to infertility [23]. UPS has been reported to have a number of physiological functions at every stage of spermatogenesis such as gonocyte and spermatogonial development, regulating meiosis (meiotic recombination, meiotic sex chromosome inactivation, meiotic progression), and spermiogenesis (nucleosome remodeling, acrosome biogenesis, sperm tail biogenesis, spermatid maturation) [23].
Although knowledge of the physiological functions of the UPS in spermatogenesis has increased, there is still limited information about the functions of DUBs in spermatogenesis. Recent studies on DUBs have convincingly demonstrated that these enzymes play an important role in the process of spermatogenesis. There are two classes of DUBs that have activity at the testicular level: USP and UCH members.
The DUB candidates USP2, USP8, USP9Y, USP14, USP26, USP42, UCH-L1, UCH-L3, UCH-L4, UCH-L5, and CYLD have been identified to date; these show varied expression levels during each stage of spermatogenesis. These DUBs are involved in gonocyte recruitment, cell cycle progression, regulation of the meiotic phase, spermiogenesis, acrosome biogenesis, and germ cell apoptosis [23]. In this review, we compile reported evidence and current knowledge about the functions of DUBs and their importance during spermatogenesis.
USP2/UBP-testis
USP2 has two isoforms, USP2a and USP2b, which were first observed in the rat testis [24, 25]. Both isoforms, which have an identical sequence but a distinct N-terminus region, are germ cell specific and developmentally regulated. These isoforms are induced in late elongating spermatids and show differential localization. USP2a is expressed in step 16–19 spermatids and is found in the nucleus, while USP2b is expressed in step 18–19 spermatids, is confined to the extranuclear space, and is found in residual bodies [24]. In a later study, the same group generated USP2 knockout mice to determine the function of USP2 during spermatogenesis. USP2 knockout mice had a normal appearance, weight, number of testicular spermatids, and epididymal spermatozoa, but showed some abnormal aggregations of elongating spermatids and formation of multinucleated cells in some tubules. USP2 knockout mice had a severe fertility defect due to failure of the spermatozoa to fertilize the egg when compared with normal mice [26]. The expression pattern of USP2 in late elongating spermatids and the severe infertility of USP2 knockout mice indicate that regulation of ubiquitination of proteins by USP2 is an important regulatory mechanism and is essential for fertilization.
USP8/mUBPy
USP8 (previously named UBPy, ubiquitin-specific processing protease-y) was originally identified as a putative protein encoded by a cDNA expressed in human myeloblasts (hUBPy) [27]. Mouse Ubpy is highly expressed in brain and testis [28]. Ubpy is considered to be a marker of acrosome biogenesis from the endocytic pathway because of its main functions of endosomal sorting, vesicle trafficking, and maintenance of endocytic vesicles [29, 30]. Inactivation of Ubpy results in embryonic lethality [30]. Ubpy shows a strong interaction with Msj-1, a mouse testis-specific DnaJ protein that is highly expressed in haploid male germ cells and has a significant role in acrosome formation [31–33]. Transcript and protein levels of Msj-1 were significantly lower in wobbler mice (natural mutant mouse with motoneuron degeneration and defective spermiogenesis) testes than wild-type mice testes, indicating that it functions in testicular metabolism [34]. However, the mRNA level of Ubpy, which is a deubiquitinating enzyme for Msj-1, was higher in wobbler mice than in normal mice [35].
During the first wave of spermatogenesis, Ubpy protein is present in the spermatids of wobbler mice in comparison to normal mice. Indeed, Ubpy is detected mainly in the soluble fraction in normal mice, while Ubpy is primarily present in the insoluble protein fraction in wobbler mouse [35]. In wild-type mice, Ubpy is expressed on the surface of acrosomic vesicles while in wobbler mice, the acrosomic vesicles do not fuse to form a functional acrosome, and there is diffuse expression signal of Ubpy in the cytoplasmic or perinuclear area of round spermatids [36]. In the wild-type testis, Vsp54 protein is expressed in several acrosomic vesicles that follow the route of Ubpy labeled vesicles [36]. However, wobbler mice have a mutated Vsp54 protein that is unable to coalesce into larger vesicles; this causes both Vsp54- and Ubpy-coated vesicles to diffuse and remain as scattered small vesicles in the cytoplasm [37]. Further, Ubpy has a microtubule-interacting and transport domain (MIT) that associates with both the spermatid endosomal sorting complex that is necessary for transport-0 (ESCRT-0), as well as microtubule structures [36]. Thus, Ubpy plays a key role in acrosome biogenesis by regulating both the endosomal pathway and the microtubule cytoskeleton.
USP9Y
The long arm of the human Y chromosome carries a number of genes that regulate spermatogenesis; several types of recurrent Yq deletions are associated with spermatogenic failure [38]. Deletion of azoospermia factor (AZF) regions (classically divided into three AZF regions, i.e., AZFa, AZFb, and AZFc) on the Y chromosome is associated with severe spermatogenetic failure. USP9Y spans 170 kb of DNA, comprises 46 exons, and lies within a small part of the AZFa interval. Deletions affecting USP9Y were originally reported to be strongly associated with mild-to-severe azoospermia or oligospermia [39, 40].
Later, it was reported that partial deletion of USP9Y in men is associated with the milder phenotype of infertility, suggesting a minor role for USP9Y in spermatogenesis [41, 42]. USP9Y may act as a fine tuner to improve the efficiency of human spermatogenesis rather than having an essential function such as acquisition of sperm-fertilizing ability or sperm maturation [42]. Recently, Luddi et al., reported no association between USP9Y and severe oligospermia or azoospermia in a clinical study [43], consistent with the previously reported marginal role of USP9Y in spermatogenesis [42]. Further, complete deletion of USP9Y does not cause spermatogenic defects, nor does it preclude the natural conception of children [43]. Thus, USP9Y is currently considered to have a nonessential role in normal sperm production and fertility in humans.
USP14
USP14 is one of three proteasome-associated DUBs that are mainly involved in removal of ubiquitin from proteasomal substrates prior to their degradation [44]. Homozygous ataxia (ax J) mice show decreased expression of Usp14 and have severe neuromuscular defects; these mice eventually die at the age of 2 months [45, 46]. Apart from its association with developmental abnormalities, downregulation of Usp14 expression has been found to affect reproductive capabilities in ax J mice. The weight of the ax J testis is half that of the wild-type testis, and the sperm number of ax J mice is 100-fold lower than that of wild-type mice. In addition, decapitated and two-tailed spermatozoa are common features of sperm from ax J mice. Histological examination of ax J testes showed that defects in the spermatid elongation process and the presence of degenerating germ cells resulted in minimal sperm release from the seminiferous tubule when compared to wild-type controls. Histological examination of Usp14-deficient testes revealed abnormal spermatogenesis and the presence of degenerating germ cells. Thus, Usp14 plays a key role in the male reproductive system, especially in spermatid differentiation during spermiogenesis [47].
USP26
USP26, which is an X-linked gene, was initially isolated from mouse spermatogonia, and its expression was reported to be testis specific in mice and humans [48–51]. The expression of USP26 is high in spermatogonia types A and B, pre-leptotene spermatocytes, round spermatids, and at the blood-testis barrier [51]. Initially, USP26 was reported to be a regulator of androgen receptor (AR) hormone-induced regulation of spermatogenesis and steroid production based on in vitro studies [49]. Nuclear USP26 was found to interact with AR through three nuclear receptor interaction motifs to regulate AR ubiquitination, transcriptional activity, and protein stability [49].
Several research groups have investigated the association between testis-specific expression of USP26 and male infertility. Approximately, 20 alterations in USP26 associated with male infertility have been reported [52]. Among these, three mutations that are frequently reported in both fertile and infertile groups are c.363_364insACA, c.494T>C, and c.1423C>T; these are associated with azoospermia and severe oligozoospermia. However, the results from different research groups are conflicting; some studies reported a significant association between these cluster of mutations and male fertility [53–55], while others did not find any association [56–58]. In a genome-wide study on single nucleotide polymorphism (SNP) associations with published male infertility genes, 14 of 147 SNPs showed a significant association with male fertility, including USP26 [59]. Recently, Zhang et al. conducted an enzymatic and meta-analyses of ten case–control studies and found no influence of the reported mutations on the enzymatic activity of USP26 and no significant association between USP26 variants and male infertility [60]. Thus, whether men carrying mutations in USP26 have an increased risk of infertility remains unknown. Further investigation of the biological function of USP26 is required to further elucidate the role of this protein in male infertility.
USP25, USP42, and USP44
There are several USPs that are expressed at high levels in the testis, but their physiological functions in spermatogenesis are not well understood. USP25 is expressed at a high level in adult mouse testis. USP25 positive cells were observed in spermatocytes I and II in the testis, which is where meiosis takes place and several cytoplasmic changes occur, suggesting that this protein may play a role in regulating protein turnover [61]. USP42 is expressed in mouse brain, lung thymus, and testis. The expression level of this protein increased from the second week after birth in the round-spermatid stage and gradually decreased during the condensing-spermatid stage during spermatogenesis [62]. Transcript levels of USP44 were also shown to be elevated in mouse testis [63]. However, the biological function of these DUBs during spermatogenesis has not yet been studied.
UCH
Ubiquitin C-terminal hydrolases (Uchs) are members of the DUB family. Among mammalian Uchs, Uchl-1 and Uchl-3 have been extensively studied and have high sequence similarity [64]. Uchl-1 and Uchl-3 have different expression patterns, indicating distinct functions during different stages of spermatogenesis [65]. Uchl-3 is highly expressed in meiotic pachytene spermatocytes and post-meiotic spermatids, indicating that it functions in the meiotic differentiation of spermatocytes into spermatids [66]. Uchl-1 is highly expressed in spermatogonia. The expression level of Uchl-1 can be up- or downregulated to determine spermatogonial self-renewal or differentiation. The undifferentiated spermatogonia marker Plzf is expressed in cells that express high levels of Uchl-1, while the differentiated spermatogonia marker c-Kit is expressed in cells with a low level of Uchl-1 [67].
Region-specific variations in Uchl-1 and Uchl-3 expression have also been found during epididymal passage. Uchl-1 expression level is high in the caput epididymidis, which is the main maturation organ, while Uchl-3 expression is high in the cauda epididymidis, the main storage organ [68]. Overexpressing Uchl-1 in the testis of male mice resulted in arrest of spermatogenesis at the pachytene stage of spermatocytes due to an increase in the number of apoptotic spermatocytes. In addition, an elevated level of activated caspase-3 was observed, indicating that Uchl-1 is an apoptotic factor related to ubiquitin function during spermatogenesis [69]. Knocking out Uchl-1 resulted in an increased number of premeiotic germ cells between post-natal days 7 and 14, along with increased levels of apoptotic proteins such as TRP53, Bax, and caspase-3 [70]. In addition, the UCH isozymes, Uchl-1 and Uchl-4, are expressed in spermatogonia, whereas Uchl-3 and Uchl-5 are primarily expressed in spermatocytes and spermatids, reflecting their distinct functions during spermatogenesis [66]. Uchl-1 and Uchl-3 play an important role in mediating apoptotic stress during spermatogenesis.
CYLD
Germline mutations in the cylindromatosis (CYLD) gene have been reported in families with an autosomal dominantly inherited predisposition towards cylindromas, trichoepitheliomas, and/or spiradenomas [71–74]. Familial cylindromatosis (MIM# 132700) and multiple familial trichoepithelioma (MIM# 601606) are allelic diseases at the two ends of the spectrum of Brooke-Spiegler syndrome (MIM# 605041) [74]. The human CYLD gene is localized on chromosome 16q12.1 and encodes the cylindromatosis protein. CYLD has a catalytic domain at its C-terminus region, shares sequence homology with ubiquitin carboxy-terminal hydrolases, and belongs to the peptidase C19 family of proteins [73, 74]. Knockout mouse models have shown that the physiological functions of CYLD are regulation of immune function and tumorigenesis [75–81]. Inactivation of Cyld in mice makes them more susceptible to chemical tumorigenesis [82].
Apart from tumorigenesis, CYLD plays a critical role in germ cell apoptosis, spermatogenesis, and male fertility [83]. CYLD knockout mice failed to produce offspring, and severe abnormalities were observed in the seminiferous tubules, including reduced lumen size and tubule cellularity. CYLD knockout male mice had disorganized seminiferous epithelium and failure of germ cell development with attenuation of the early wave of germ cell apoptosis [83]. Early wave germ cell apoptosis is an organized process during spermatogenesis required to remove excess germ cells and maintain the balance between germ cells and Sertoli cells [84]. Loss of CYLD is associated with constitutive activation of NF-κB in germ cells, leading to aberrant expression of anti-apoptotic Bcl-2 members. Moreover, CYLD interacts physically with receptor-interacting protein 1 (RIP1) and prevents its ubiquitination by primarily targeting an ubiquitin-dependent activator of IKK and NF-κB. Inactivation of CYLD leads to accumulation of ubiquitinated RIP1, which in turn activates IKK and NF-κB [83]. Thus, CYLD is considered to be a crucial DUB that regulates germ cell apoptosis and spermatogenesis, suggesting that it has a key role in regulating RIP1 ubiquitination and NF-κB activation in testicular cells.
Concluding remarks
A growing body of evidence indicates that the UPS plays an important role during each step of spermatogenesis from the gonocyte stage to differentiated spermatids ready to be released into the epididymis. A great amount of attention has been focused on the role of the ubiquitination system in regulating spermatogenesis. However, reversal of ubiquitination by DUBs plays an equally important role in regulation of the expression of enzymes and/or their subcellular localization by preventing ubiquitination during spermatogenesis. DUB enzymes might process the removal of histones and regulate protein turnover during meiosis, which is a critical stage in spermatogenesis. DUBs frequently appear as members of large multi-protein complexes that directly or indirectly regulate ubiquitination of proteins during spermatogenesis. Though less abundant, there are still some loss-of-function studies that have demonstrated the physiological functions of DUBs in spermatogenesis. In this review, we focused on DUBs that directly regulate the process of spermatogenesis and are involved in male fertility; these DUBs are summarized in Table 1.
Although it is clear that DUBs play a major role in the regulation of spermatogenesis, in most cases, there is still a lack of information about the molecular mechanisms by which these proteins exert their effects. In addition, there are inconsistencies among published findings for DUBs and their role in spermatogenesis. Thus, identification of specific substrates for DUBs is important to confirm that the proteins implicated in loss-of-function studies play a direct role in spermatogenesis. Moreover, DUBs are comparatively few in number and specific in function. It is now understood that while there are several thousand proteins that are ubiquitinated and degraded during spermatogenesis, there are only few DUBs that counteract this ubiquitination. Thus, more detailed studies of DUBs that regulate germ cell proliferation and differentiation during spermatogenesis would be useful for developing effective DUB-targeted treatment approaches for male infertility.
Abbreviations
- DUBs:
-
Deubiquitinating enzymes
- UPS:
-
Ubiquitin proteasome system
- UCH:
-
Ubiquitin C-terminal hydrolases
- USP:
-
Ubiquitin-specific processing proteases
- JAMM:
-
Jab1/Pab1/MPN domain-containing metalloenzymes
- OTU:
-
Otu-domain ubiquitin aldehyde-binding proteins
- MCPIPs:
-
Monocyte chemotactic protein-induced proteases
- MIT:
-
Microtubule interacting and transport domain
- AZF:
-
Azoospermia factor
- AR:
-
Androgen receptor
- ax:
-
Ataxia
- SNPs:
-
Single nucleotide polymorphism
- CYLD:
-
Cylindromatosis
- RIP1:
-
Receptor-interacting protein 1
References
Amerik AY, Hochstrasser M (2004) Mechanism and function of deubiquitinating enzymes. Biochim Biophys Acta 1695:189–207
Sigismund S, Polo S, Di Fiore PP (2004) Signaling through monoubiquitination. Curr Top Microbiol Immunol 286:149–185
Sun L, Chen ZJ (2004) The novel functions of ubiquitination in signaling. Curr Opin Cell Biol 16(2):119–126. doi:10.1016/j.ceb.2004.02.005
Sadowski M, Suryadinata R, Tan AR, Roesley SN, Sarcevic B (2012) Protein monoubiquitination and polyubiquitination generate structural diversity to control distinct biological processes. IUBMB Life 64(2):136–142. doi:10.1002/iub.589
Ramanathan HN, Ye Y (2012) Cellular strategies for making monoubiquitin signals. Crit Rev Biochem Mol Biol 47(1):17–28. doi:10.3109/10409238.2011.620943
Komander D, Rape M (2012) The ubiquitin code. Annu Rev Biochem 81:203–229. doi:10.1146/annurev-biochem-060310-170328
Komander D (2009) The emerging complexity of protein ubiquitination. Biochem Soc Trans 37(Pt 5):937–953. doi:10.1042/BST0370937
Reyes-Turcu FE, Ventii KH, Wilkinson KD (2009) Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem 78:363–397. doi:10.1146/annurev.biochem.78.082307.091526
Baek KH (2006) Cytokine-regulated protein degradation by the ubiquitination system. Curr Protein Pept Sci 7:171–177
Baek KH, Kim MS, Kim YS, Shin JM, Choi KH (2004) DUB-1A, a novel subfamily member of deubiquitinating enzyme, is polyubiquitinated and cytokine inducible in B-lymphocytes. J Biol Chem 279:2368–2376
Lim KH, Ramakrishna S, Baek KH (2013) Molecular mechanisms and functions of cytokine-inducible deubiquitinating enzymes. Cytokine Growth Factor Rev 24(5):427–431. doi:10.1016/j.cytogfr.2013.05.007
Ramakrishna S, Kim KS, Baek KH (2014) Posttranslational modifications of defined embryonic reprogramming transcription factors. Cell Reprogramm 16(2):108–120. doi:10.1089/cell.2013.0077
Ramakrishna S, Suresh B, Baek KH (2011) The role of deubiquitinating enzymes in apoptosis. Cell Mol Life Sci 68(1):15–26. doi:10.1007/s00018-010-0504-6
Ramakrishna S, Suresh B, Baek KH (2015) Biological functions of hyaluronan and cytokine-inducible deubiquitinating enzymes. Biochim Biophys Acta 1855(1):83–91. doi:10.1016/j.bbcan.2014.11.006
Hess RA, Renato de Franca L (2008) Spermatogenesis and cycle of the seminiferous epithelium. Adv Exp Med Biol 636:1–15. doi:10.1007/978-0-387-09597-4_1
Heller CH, Clermont Y (1964) Kinetics of the germinal epithelium in man. Recent Prog Horm Res 20:545–575
Misell LM, Holochwost D, Boban D, Santi N, Shefi S, Hellerstein MK, Turek PJ (2006) A stable isotope-mass spectrometric method for measuring human spermatogenesis kinetics in vivo. J Urol 175(1):242–246. doi:10.1016/S0022-5347(05)00053-4 (discussion 246)
Schulze C (1979) Morphological characteristics of the spermatogonial stem cells in man. Cell Tissue Res 198(2):191–199
Clermont Y (1972) Kinetics of spermatogenesis in mammals: seminiferous epithelium cycle and spermatogonial renewal. Physiol Rev 52(1):198–236
de Rooij DG, Russell LD (2000) All you wanted to know about spermatogonia but were afraid to ask. J Androl 21(6):776–798
de Rooij DG (2001) Proliferation and differentiation of spermatogonial stem cells. Reproduction 121(3):347–354
Tegelenbosch RA, de Rooij DG (1993) A quantitative study of spermatogonial multiplication and stem cell renewal in the C3H/101 F1 hybrid mouse. Mutat Res 290(2):193–200
Bose R, Manku G, Culty M, Wing SS (2014) Ubiquitin-proteasome system in spermatogenesis. Adv Exp Med Biol 759:181–213. doi:10.1007/978-1-4939-0817-2_9
Lin H, Keriel A, Morales CR, Bedard N, Zhao Q, Hingamp P, Lefrancois S, Combaret L, Wing SS (2000) Divergent N-terminal sequences target an inducible testis deubiquitinating enzyme to distinct subcellular structures. Mol Cell Biol 20(17):6568–6578
Manku G, Wing SS, Culty M (2012) Expression of the ubiquitin proteasome system in neonatal rat gonocytes and spermatogonia: role in gonocyte differentiation. Biol Reprod 87(2):44. doi:10.1095/biolreprod.112.099143
Bedard N, Yang Y, Gregory M, Cyr DG, Suzuki J, Yu X, Chian RC, Hermo L, O’Flaherty C, Smith CE, Clarke HJ, Wing SS (2011) Mice lacking the USP2 deubiquitinating enzyme have severe male subfertility associated with defects in fertilization and sperm motility. Biol Reprod 85(3):594–604. doi:10.1095/biolreprod.110.088542
Naviglio S, Mattecucci C, Matoskova B, Nagase T, Nomura N, Di Fiore PP, Draetta GF (1998) UBPY: a growth-regulated human ubiquitin isopeptidase. EMBO J 17(12):3241–3250. doi:10.1093/emboj/17.12.3241
Gnesutta N, Ceriani M, Innocenti M, Mauri I, Zippel R, Sturani E, Borgonovo B, Berruti G, Martegani E (2001) Cloning and characterization of mouse UBPy, a deubiquitinating enzyme that interacts with the ras guanine nucleotide exchange factor CDC25(Mm)/Ras-GRF1. J Biol Chem 276(42):39448–39454. doi:10.1074/jbc.M103454200
Mizuno E, Kobayashi K, Yamamoto A, Kitamura N, Komada M (2006) A deubiquitinating enzyme UBPY regulates the level of protein ubiquitination on endosomes. Traffic 7(8):1017–1031. doi:10.1111/j.1600-0854.2006.00452.x
Niendorf S, Oksche A, Kisser A, Lohler J, Prinz M, Schorle H, Feller S, Lewitzky M, Horak I, Knobeloch KP (2007) Essential role of ubiquitin-specific protease 8 for receptor tyrosine kinase stability and endocytic trafficking in vivo. Mol Cell Biol 27(13):5029–5039. doi:10.1128/MCB.01566-06
Berruti G, Martegani E (2001) MSJ-1, a mouse testis-specific DnaJ protein, is highly expressed in haploid male germ cells and interacts with the testis-specific heat shock protein Hsp70-2. Biol Reprod 65(2):488–495
Berruti G, Martegani E (2002) mUBPy and MSJ-1, a deubiquitinating enzyme and a molecular chaperone specifically expressed in testis, associate with the acrosome and centrosome in mouse germ cells. Ann N Y Acad Sci 973:5–7
Berruti G, Martegani E (2005) The deubiquitinating enzyme mUBPy interacts with the sperm-specific molecular chaperone MSJ-1: the relation with the proteasome, acrosome, and centrosome in mouse male germ cells. Biol Reprod 72(1):14–21. doi:10.1095/biolreprod.104.030866
Meccariello R, Cobellis G, Berruti G, Junier MP, Ceriani M, Boilee S, Pierantoni R, Fasano S (2002) Mouse sperm cell-specific DnaJ first homologue: an evolutionarily conserved protein for spermiogenesis. Biol Reprod 66(5):1328–1335
Chianese R, Scarpa D, Berruti G, Cobellis G, Pierantoni R, Fasano S, Meccariello R (2010) Expression and localization of the deubiquitinating enzyme mUBPy in wobbler mouse testis during spermiogenesis. Gen Comp Endocrinol 166(2):289–295. doi:10.1016/j.ygcen.2009.09.014
Berruti G, Ripolone M, Ceriani M (2010) USP8, a regulator of endosomal sorting, is involved in mouse acrosome biogenesis through interaction with the spermatid ESCRT-0 complex and microtubules. Biol Reprod 82(5):930–939. doi:10.1095/biolreprod.109.081679
Paiardi C, Pasini ME, Gioria M, Berruti G (2011) Failure of acrosome formation and globozoospermia in the wobbler mouse, a Vps54 spontaneous recessive mutant. Spermatogenesis 1(1):52–62. doi:10.4161/spmg.1.1.14698
Krausz C, Forti G, McElreavey K (2003) The Y chromosome and male fertility and infertility. Int J Androl 26(2):70–75
Sun C, Skaletsky H, Birren B, Devon K, Tang Z, Silber S, Oates R, Page DC (1999) An azoospermic man with a de novo point mutation in the Y-chromosomal gene USP9Y. Nat Genet 23(4):429–432. doi:10.1038/70539
Brown GM, Furlong RA, Sargent CA, Erickson RP, Longepied G, Mitchell M, Jones MH, Hargreave TB, Cooke HJ, Affara NA (1998) Characterisation of the coding sequence and fine mapping of the human DFFRY gene and comparative expression analysis and mapping to the Sxrb interval of the mouse Y chromosome of the Dffry gene. Hum Mol Genet 7(1):97–107
Ferlin A, Arredi B, Speltra E, Cazzadore C, Selice R, Garolla A, Lenzi A, Foresta C (2007) Molecular and clinical characterization of Y chromosome microdeletions in infertile men: a 10-year experience in Italy. J Clin Endocrinol Metab 92(3):762–770. doi:10.1210/jc.2006-1981
Krausz C, Degl’Innocenti S, Nuti F, Morelli A, Felici F, Sansone M, Varriale G, Forti G (2006) Natural transmission of USP9Y gene mutations: a new perspective on the role of AZFa genes in male fertility. Hum Mol Genet 15(18):2673–2681. doi:10.1093/hmg/ddl198
Luddi A, Margollicci M, Gambera L, Serafini F, Cioni M, De Leo V, Balestri P, Piomboni P (2009) Spermatogenesis in a man with complete deletion of USP9Y. N Engl J Med 360(9):881–885. doi:10.1056/NEJMoa0806218
Hu M, Li P, Song L, Jeffrey PD, Chenova TA, Wilkinson KD, Cohen RE, Shi Y (2005) Structure and mechanisms of the proteasome-associated deubiquitinating enzyme USP14. EMBO J 24(21):3747–3756. doi:10.1038/sj.emboj.7600832
D’Amato CJ, Hicks SP (1965) Neuropathologic alterations in the ataxia (paralytic) mouse. Arch Pathol 80(6):604–612
Vaden JH, Bhattacharyya BJ, Chen PC, Watson JA, Marshall AG, Phillips SE, Wilson JA, King GD, Miller RJ, Wilson SM (2015) Ubiquitin-specific protease 14 regulates c-Jun N-terminal kinase signaling at the neuromuscular junction. Mol Neurodegener 10(1):3. doi:10.1186/1750-1326-10-3
Crimmins S, Sutovsky M, Chen PC, Huffman A, Wheeler C, Swing DA, Roth K, Wilson J, Sutovsky P, Wilson S (2009) Transgenic rescue of ataxia mice reveals a male-specific sterility defect. Dev Biol 325(1):33–42. doi:10.1016/j.ydbio.2008.09.021
Zhang J, Tian H, Huo YW, Zhou DX, Wang HX, Wang LR, Zhang QY, Qiu SD (2009) The expression of Usp26 gene in mouse testis and brain. Asian J Androl 11(4):478–483. doi:10.1038/aja.2009.31
Dirac AM, Bernards R (2010) The deubiquitinating enzyme USP26 is a regulator of androgen receptor signaling. Mol Cancer Res 8(6):844–854. doi:10.1158/1541-7786.MCR-09-0424
Wang PJ, McCarrey JR, Yang F, Page DC (2001) An abundance of X-linked genes expressed in spermatogonia. Nat Genet 27(4):422–426. doi:10.1038/86927
Lin YW, Hsu TH, Yen PH (2011) Localization of ubiquitin specific protease 26 at blood-testis barrier and near Sertoli cell-germ cell interface in mouse testes. Int J Androl 34(5 Pt 2):e368–e377. doi:10.1111/j.1365-2605.2010.01130.x
Stouffs K, Tournaye H, Liebaers I, Lissens W (2009) Male infertility and the involvement of the X chromosome. Hum Reprod Update 15(6):623–637. doi:10.1093/humupd/dmp023
Lee IW, Kuan LC, Lin CH, Pan HA, Hsu CC, Tsai YC, Kuo PL, Teng YN (2008) Association of USP26 haplotypes in men in Taiwan, China with severe spermatogenic defect. Asian J Androl 10(6):896–904. doi:10.1111/j.1745-7262.2008.00439.x
Paduch DA, Mielnik A, Schlegel PN (2005) Novel mutations in testis-specific ubiquitin protease 26 gene may cause male infertility and hypogonadism. Reprod Biomed Online 10(6):747–754
Stouffs K, Lissens W, Tournaye H, Van Steirteghem A, Liebaers I (2005) Possible role of USP26 in patients with severely impaired spermatogenesis. Eur J Hum Genet 13(3):336–340. doi:10.1038/sj.ejhg.5201335
Ravel C, El Houate B, Chantot S, Lourenco D, Dumaine A, Rouba H, Bandyopadahyay A, Radhakrishna U, Das B, Sengupta S, Mandelbaum J, Siffroi JP, McElreavey K (2006) Haplotypes, mutations and male fertility: the story of the testis-specific ubiquitin protease USP26. Mol Hum Reprod 12(10):643–646. doi:10.1093/molehr/gal063
Stouffs K, Lissens W, Tournaye H, Van Steirteghem A, Liebaers I (2006) Alterations of the USP26 gene in Caucasian men. Int J Androl 29(6):614–617. doi:10.1111/j.1365-2605.2006.00708.x
Ribarski I, Lehavi O, Yogev L, Hauser R, Bar-Shira Maymon B, Botchan A, Paz G, Yavetz H, Kleiman SE (2009) USP26 gene variations in fertile and infertile men. Hum Reprod 24(2):477–484. doi:10.1093/humrep/den374
Aston KI, Krausz C, Laface I, Ruiz-Castane E, Carrell DT (2010) Evaluation of 172 candidate polymorphisms for association with oligozoospermia or azoospermia in a large cohort of men of European descent. Hum Reprod 25(6):1383–1397. doi:10.1093/humrep/deq081
Zhang W, Liu T, Mi YJ, Yue LD, Wang JM, Liu DW, Yan J, Tian QB (2015) Evidence from enzymatic and meta-analyses does not support a direct association between USP26 gene variants and male infertility. Andrology 3(2):271–279. doi:10.1111/andr.295
Valero R, Marfany G, Gonzalez-Angulo O, Gonzalez-Gonzalez G, Puelles L, Gonzalez-Duarte R (1999) USP25, a novel gene encoding a deubiquitinating enzyme, is located in the gene-poor region 21q11.2. Genomics 62(3):395–405. doi:10.1006/geno.1999.6025
Kim YK, Kim YS, Yoo KJ, Lee HJ, Lee DR, Yeo CY, Baek KH (2007) The expression of Usp42 during embryogenesis and spermatogenesis in mouse. Gene Expr Patterns 7(1–2):143–148. doi:10.1016/j.modgep.2006.06.006
Suresh B, Ramakrishna S, Lee HJ, Choi JH, Kim JY, Ahn WS, Baek KH (2010) K48- and K63-linked polyubiquitination of deubiquitinating enzyme USP44. Cell Biol Int 34(8):799–808. doi:10.1042/CBI20090144
Wilkinson KD, Lee KM, Deshpande S, Duerksen-Hughes P, Boss JM, Pohl J (1989) The neuron-specific protein PGP 9.5 is a ubiquitin carboxyl-terminal hydrolase. Science 246(4930):670–673
Kwon J (2007) The new function of two ubiquitin C-terminal hydrolase isozymes as reciprocal modulators of germ cell apoptosis. Exp Anim 56(2):71–77
Kwon J, Wang YL, Setsuie R, Sekiguchi S, Sakurai M, Sato Y, Lee WW, Ishii Y, Kyuwa S, Noda M, Wada K, Yoshikawa Y (2004) Developmental regulation of ubiquitin C-terminal hydrolase isozyme expression during spermatogenesis in mice. Biol Reprod 71(2):515–521. doi:10.1095/biolreprod.104.027565
Luo J, Megee S, Dobrinski I (2009) Asymmetric distribution of UCH-L1 in spermatogonia is associated with maintenance and differentiation of spermatogonial stem cells. J Cell Physiol 220(2):460–468. doi:10.1002/jcp.21789
Jervis KM, Robaire B (2001) Dynamic changes in gene expression along the rat epididymis. Biol Reprod 65(3):696–703
Wang YL, Liu W, Sun YJ, Kwon J, Setsuie R, Osaka H, Noda M, Aoki S, Yoshikawa Y, Wada K (2006) Overexpression of ubiquitin carboxyl-terminal hydrolase L1 arrests spermatogenesis in transgenic mice. Mol Reprod Dev 73(1):40–49. doi:10.1002/mrd.20364
Kwon J, Mochida K, Wang YL, Sekiguchi S, Sankai T, Aoki S, Ogura A, Yoshikawa Y, Wada K (2005) Ubiquitin C-terminal hydrolase L-1 is essential for the early apoptotic wave of germinal cells and for sperm quality control during spermatogenesis. Biol Reprod 73(1):29–35. doi:10.1095/biolreprod.104.037077
Welch JP, Wells RS, Kerr CB (1968) Ancell-Spiegler cylindromas (turban tumours) and Brooke-Fordyce trichoepitheliomas: evidence for a single genetic entity. J Med Genet 5(1):29–35
Young AL, Kellermayer R, Szigeti R, Teszas A, Azmi S, Celebi JT (2006) CYLD mutations underlie Brooke-Spiegler, familial cylindromatosis, and multiple familial trichoepithelioma syndromes. Clin Genet 70(3):246–249. doi:10.1111/j.1399-0004.2006.00667.x
Bignell GR, Warren W, Seal S, Takahashi M, Rapley E, Barfoot R, Green H, Brown C, Biggs PJ, Lakhani SR, Jones C, Hansen J, Blair E, Hofmann B, Siebert R, Turner G, Evans DG, Schrander-Stumpel C, Beemer FA, van Den Ouweland A, Halley D, Delpech B, Cleveland MG, Leigh I, Leisti J, Rasmussen S (2000) Identification of the familial cylindromatosis tumour-suppressor gene. Nat Genet 25(2):160–165. doi:10.1038/76006
Blake PW, Toro JR (2009) Update of cylindromatosis gene (CYLD) mutations in Brooke-Spiegler syndrome: novel insights into the role of deubiquitination in cell signaling. Hum Mutat 30(7):1025–1036. doi:10.1002/humu.21024
Mathis BJ, Lai Y, Qu C, Janicki JS, Cui T (2015) CYLD-mediated signaling and diseases. Curr Drug Targets 16(4):284–294
Massoumi R (2011) CYLD: a deubiquitination enzyme with multiple roles in cancer. Future Oncol 7(2):285–297. doi:10.2217/fon.10.187
Hovelmeyer N, Wunderlich FT, Massoumi R, Jakobsen CG, Song J, Worns MA, Merkwirth C, Kovalenko A, Aumailley M, Strand D, Bruning JC, Galle PR, Wallach D, Fassler R, Waisman A (2007) Regulation of B cell homeostasis and activation by the tumor suppressor gene CYLD. J Exp Med 204(11):2615–2627. doi:10.1084/jem.20070318
Lim JH, Stirling B, Derry J, Koga T, Jono H, Woo CH, Xu H, Bourne P, Ha UH, Ishinaga H, Xu H, Andalibi A, Feng XH, Zhu H, Huang Y, Zhang W, Weng X, Yan C, Yin Z, Briles DE, Davis RJ, Flavell RA, Li JD (2007) Tumor suppressor CYLD regulates acute lung injury in lethal Streptococcus pneumoniae infections. Immunity 27(2):349–360. doi:10.1016/j.immuni.2007.07.011
Reiley WW, Zhang M, Jin W, Losiewicz M, Donohue KB, Norbury CC, Sun SC (2006) Regulation of T cell development by the deubiquitinating enzyme CYLD. Nat Immunol 7(4):411–417. doi:10.1038/ni1315
Trompouki E, Tsagaratou A, Kosmidis SK, Dolle P, Qian J, Kontoyiannis DL, Cardoso WV, Mosialos G (2009) Truncation of the catalytic domain of the cylindromatosis tumor suppressor impairs lung maturation. Neoplasia 11(5):469–476
Zhang J, Stirling B, Temmerman ST, Ma CA, Fuss IJ, Derry JM, Jain A (2006) Impaired regulation of NF-kappaB and increased susceptibility to colitis-associated tumorigenesis in CYLD-deficient mice. J Clin Investig 116(11):3042–3049. doi:10.1172/JCI28746
Massoumi R, Chmielarska K, Hennecke K, Pfeifer A, Fassler R (2006) Cyld inhibits tumor cell proliferation by blocking Bcl-3-dependent NF-kappaB signaling. Cell 125(4):665–677. doi:10.1016/j.cell.2006.03.041
Wright A, Reiley WW, Chang M, Jin W, Lee AJ, Zhang M, Sun SC (2007) Regulation of early wave of germ cell apoptosis and spermatogenesis by deubiquitinating enzyme CYLD. Dev Cell 13(5):705–716. doi:10.1016/j.devcel.2007.09.007
Print CG, Loveland KL (2000) Germ cell suicide: new insights into apoptosis during spermatogenesis. BioEssays : news and reviews in molecular, cellular and developmental biology 22(5):423–430. doi:10.1002/(SICI)1521-1878(200005)22:5<423:AID-BIES4>3.0.CO;2-0
Kwon J, Kikuchi T, Setsuie R, Ishii Y, Kyuwa S, Yoshikawa Y (2003) Characterization of the testis in congenitally ubiquitin carboxy-terminal hydrolase-1 (Uch-L1) defective (gad) mice. Exp Anim 52(1):1–9
Osawa Y, Wang YL, Osaka H, Aoki S, Wada K (2001) Cloning, expression, and mapping of a mouse gene, Uchl4, highly homologous to human and mouse Uchl3. Biochem Biophys Res Commun 283(3):627–633. doi:10.1006/bbrc.2001.4841
Acknowledgments
We would like to thank all of Suri’s laboratory members for their helpful discussions. This study was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Korean government (MSIP) (No. 2012M3A9B4028738).
Author information
Authors and Affiliations
Corresponding authors
Additional information
B. Suresh and J. Lee contributed equally to this work.
Rights and permissions
About this article
Cite this article
Suresh, B., Lee, J., Hong, SH. et al. The role of deubiquitinating enzymes in spermatogenesis. Cell. Mol. Life Sci. 72, 4711–4720 (2015). https://doi.org/10.1007/s00018-015-2030-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-015-2030-z