Abstract
Alzheimer’s disease is a complex and heterogeneous, severe neurodegenerative disorder and the predominant form of dementia, characterized by cognitive disturbances, behavioral and psychotic symptoms, progressive cognitive decline, disorientation, behavioral changes, and death. Genetic background of Alzheimer’s disease differs between early-onset familial Alzheimer’s disease, other cases of early-onset Alzheimer’s disease, and late-onset Alzheimer’s disease. Rare cases of early-onset familial Alzheimer’s diseases are caused by high-penetrant mutations in genes coding for amyloid precursor protein, presenilin 1, and presenilin 2. Late-onset Alzheimer’s disease is multifactorial and associated with many different genetic risk loci (>20), with the apolipoprotein E ε4 allele being a major genetic risk factor for late-onset Alzheimer’s disease. Genetic and genomic studies offer insight into many additional genetic risk loci involved in the genetically complex nature of late-onset Alzheimer’s disease. This review highlights the contributions of individual loci to the pathogenesis of Alzheimer’s disease and suggests that their exact contribution is still not clear. Therefore, the use of genetic markers of Alzheimer’s disease, for monitoring development, time course, treatment response, and prognosis of Alzheimer’s disease, is still far away from the clinical application, because the contribution of genetic variations to the relative risk of developing Alzheimer’s disease is limited. In the light of prediction and prevention of Alzheimer’s disease, a novel approach could be found in the form of additive genetic risk scores, which combine additive effects of numerous susceptibility loci.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Alzheimer’s Disease
Alzheimer’s disease is a complex and heterogeneous brain disorder that can be classified, according to its stages, into dementia in Alzheimer’s disease (F.00) and Alzheimer’s disease (G.30), according to ICD-10. Namely, it is a severe neurodegenerative disease and the predominant form of dementia (50–75%), but when behavioral and psychotic symptoms of dementia (BPSD) develop during the course of Alzheimer’s disease, it has to be treated as a severe mental, i.e., psychiatric disorder. These neuropsychiatric symptoms include depression, apathy, anxiety, irritability, agitation, euphoria, hallucinations, disinhibition, aberrant motor behavior, elation, delusions, and sleep or appetite changes; and they can occur in the early as well as in the middle and late stages of Alzheimer’s disease [1].
The first sign of dementia in Alzheimer’s disease is the gradual worsening of the ability to remember new information. However, during the course of Alzheimer’s disease, multiple cognitive domains are disrupted [2, 3]. The cognitive disturbances affect universal domains such as attention, working memory, executive function, procedural learning and memory, speed of processing, fear-extinction learning and semantic memory, and some higher domains that include episodic memory, social cognition, theory of mind, verbal learning, memory, and language (i.e., use and understanding) [3, 4]. Alzheimer’s disease is a slow, irreversible, progressive, complex, and lethal disorder, which represents a major health problem and fatal global epidemic worldwide [3]. It is characterized by progressive cognitive decline, disorientation, behavioral changes, and death. A latency phase of the Alzheimer’s disease is without clinical symptoms although the pathophysiological processes are active [2]. The clear etiology of Alzheimer’s disease is still unknown. However, the main risk factors are older age, genetic predisposition (especially the apolipoprotein E (ApoE) ε4 genotype), gender (female predominance), and presence of the mild cognitive impairment, but there are also modifiable factors such as cardiovascular risk factors, hypertension, diabetes, obesity, smoking, and high cholesterol levels [3]. Insulin signaling dysfunction and brain glucose metabolism disturbances are hallmarks of Alzheimer’s disease, and therefore recently Alzheimer’s disease was suggested to be considered as type 3 diabetes [5].
Genetic Background of Alzheimer’s Disease
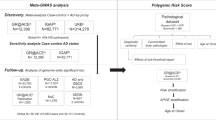
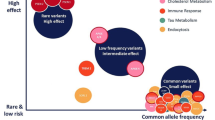
Alzheimer’s disease can be divided into autosomal dominant Alzheimer’s disease (or early-onset familial Alzheimer’s disease), other cases of early-onset Alzheimer’s disease, and late-onset Alzheimer’s disease [6]. Genetic background of Alzheimer’s disease differs between early-onset familial AD, other cases of early-onset Alzheimer’s disease, and late-onset Alzheimer’s disease. Early-onset familial Alzheimer’s disease, with a prevalence less than 1%, is caused by high-penetrant mutations in genes coding for amyloid precursor protein (APP), presenilin 1 (PSEN1), and presenilin 2 (PSEN2). Late-onset Alzheimer’s disease is multifactorial and associated with many different genetic risk loci (> 20), with the ApoE ε4 allele being a major genetic risk factor for late-onset Alzheimer’s disease. Genome-wide association studies (GWAS) offered insight into many additional genetic risk loci involved in the genetically complex nature of late-onset Alzheimer’s disease. This review focuses on the recent data from comprehensive meta-analysis and GWAS. However, it should be highlighted that the exact contributions of individual loci to the pathogenesis of Alzheimer’s disease still remain unclear to date.
Early-Onset Familial Alzheimer’s Disease
The discovery of the association between mutations in APP, presenilin PSEN1 and PSEN2 genes and the development of early-onset familial Alzheimer’s disease provided knowledge about the molecular mechanisms underlying the Alzheimer’s disease pathogenesis.
Amyloid Precursor Protein
The enzymatic cleavage of APP can lead to the formation of amyloid β-peptide (Aβ), which can be 38 to 43 amino acids long. Cleavage of APP by α- and γ-secretases results in the generation of nonpathogenic peptides, secreted form of APP (sAPPα) and C-terminal fragments. This pathway is known as nonamyloidogenic or constitutive pathway. Amyloidogenic pathway involves the proteolysis of APP by β- and γ-secretase, resulting in the formation of sAPPα, C-terminal fragments, and Aβ. We differentiate two main forms of Aβ, Aβ1–40, and Aβ1–42. Amyloid plaques are most commonly formed from more amyloidogenic Aβ1–42 form.
According to Alzheimer Disease and Frontotemporal Dementia Mutation Database (http://www.molgen.ua.ac.be/ADmutations/) and Alzforum (https://www.alzforum.org/mutations/app), there are around 35 different APP mutations that have been associated with Alzheimer’s disease pathogenesis. These mutations include APP gene locus duplications and different point mutations in coding region of APP gene, resulting in an amino acid substitution. Duplication of the whole gene/locus lead to elevated levels of APP and Aβ, and increase the ratio of Aβ1–42 to Aβ1–40. Missense mutations can have different effects, depending on their position. If these mutations cause amino acid substitution near the β-proteolytic cleavage site (N-terminal of Aβ), they usually lead to increased β-secretase cleavage, increased total Aβ production, and increased aggregation and fibril formation (Table 1). Missense mutations in the Aβ sequence in general increase Aβ aggregation and fibril formation (Table 1). If the missense mutation is near the C-terminal of Aβ, then it will increase the relative production of Aβ1–42, compared to Aβ1–40 (Table 1).
Presenilin 1 and Presenilin 2
PSEN1 and PSEN2 are two homologous multi-transmembrane proteins that share around 67% of the sequence [7], and they represent the catalytic core of γ-secretase complex. These proteins are also involved in the cleavage of some other proteins, like cadherins, low-density lipoprotein receptor (LDLR)-related proteins, Notch-1 and ErbB4 [8,9,10,11]. At the cell level, presenilins can be found in the nuclear membrane, endoplasmic reticulum, the trans-Golgi network and at the plasma membrane. They are also widely expressed throughout the organism.
Mutations in PSEN1 and PSEN2 genes are the most frequent known cause of early-onset familial Alzheimer’s disease, with emphasis on PSEN1 gene. Mutations in these two genes usually cause an impairment in γ-secretase activity and lead to an increase in the ratio between Aβ1–42 and Aβ1–40, as a consequence of Aβ1–42 overproduction or Aβ1–40 underproduction, or as a combination of both (Table 1). PSEN1 mutations were associated with the earliest disease onset ages, with an average age of onset around 43 years, from 25 until 65 years of age [12]. In APP mutation carriers the disease starts on average 8.4 years earlier (age of onset between 35 and 65 years of age), and in PSEN2 mutation carriers on average 14.2 years earlier, with a much older age of onset (between 45 and 70 years of age) [12].
Late-Onset Alzheimer’s Disease
Most of the genes that have been associated with late-onset Alzheimer’s disease, detected through different candidate genes studies and GWAS, are involved in cholesterol and lipid metabolism (genes coding for ApoE (APOE), sortilin-related receptor-1 (SORL1), ATP-binding cassette subfamily A member 7 (ABCA7), and clusterin (CLU)), immune system and inflammation (genes coding for complement C3b/C4b receptor 1 (CR1), CD33 antigen, membrane-spanning 4-domains, subfamily A member (MS4A), triggering receptor expressed on myeloid cells 2 (TREM2), member of the major histocompatibility complex class II HLA-DRB5/HLA-DRB1, and a SH2-containing inositol 5-phosphatase 1 (INPP5D)), and/or endosome cycling (genes coding for bridging integrator protein-1 (BIN1), CD2-associated protein (CD2AP), phosphatidylinositol binding clathrin assembly protein (PICALM), ephrin type-A receptor 1 (EPHA1)). However, there are also studies that implicate some other genes, whose function is not so well known and described, with Alzheimer’s disease pathology, like genes coding for thioredoxin domain-containing protein 3 (NME8), CUGBP Elav-like family member 1 (CELF1), cas scaffolding protein family member 4 (CASS4), protein-tyrosine kinase 2-beta (PTK2B), zinc finger CW-type PWWP domain protein 1 (ZCWPW1), fermitin family homolog 2 (FERMT2), sodium/potassium/calcium exchanger 4 (SLC24A4), and Ras and Rab interactor 3 (RIN3) [13,14,15,16,17,18,19]. In this chapter, some of the most interesting genes found to be associated with late-onset Alzheimer’s disease will be described, with their possible involvement in certain biological pathways and mechanisms that might be relevant for Alzheimer’s disease pathology.
Apolipoprotein E
One of the major risk loci for late-onset Alzheimer’s disease is the ε4 allele of APOE gene, gene coding for the main apolipoprotein in the central nervous system. This glycoprotein plays an important role in lipid transport, and it has an undeniable role in growth, repair, reorganization, and maintenance of neurons. ApoE facilitates the cellular uptake of lipoproteins by binding to the members of LDLR family, or it takes part in the activation of signaling pathways involved in modulating lipid homeostasis [20]. Two amino acid substitutions at the positions 112 and 158 lead to three possible ApoE isoforms, ApoE2, ApoE3, ApoE4, which are encoded by three common alleles (ε2, ε3, ε4). The ε4 allele has been associated with Alzheimer’s disease, and it is considered as a most important risk factor in the case of late-onset Alzheimer’s disease (Table 1). The carriers of APOE ε4 allele have an earlier age of onset of Alzheimer’s disease, and they also tend to have more pronounced accumulation of neurofibrillary tangles and amyloid plaques [21]. However, APOE ε2 allele was associated with reduced risk of developing Alzheimer’s disease, with reduced accumulation of neurofibrillary tangles and amyloid plaques [22, 23], but also with significantly larger regional cortical thicknesses and volumes in subjects with cognitive impairment or Alzheimer’s disease [24]. The amino acid substitution at the position 158 (arginine to cysteine) impairs the binding of ApoE2 to LDLR and its ability to promote clearance of TG‐rich lipoprotein remnant particles. ApoE4 is characterized by an amino acid substitution at the position 112 (cysteine to arginine) that affects the stability of the N‐terminal domain helix bundle and C-terminal domain, resulting in enhanced lipid-binding ability of ApoE4 [20] and less efficient clearance of soluble Aβ, amyloid plaques and/or neurofibrillary tangles [25].
Clusterin
Clusterin is a highly glycosylated cell-aggregating factor that is involved in different processes, including complement inhibition, inflammation, apoptosis, and lipid transport [26]. As a chaperone, it could be involved in the amyloid aggregation and pathogenesis of Alzheimer’s disease [27]. Evidence suggests that clusterin forms complexes with Aβ in cerebrospinal fluid that are able to cross the brain–blood barrier [28]. Few GWAS studies suggested clusterin as a potential biomarker of Alzheimer’s disease [29, 30]. A single nucleotide polymorphism (SNP) in the CLU gene was suggested to be associated with Alzheimer’s disease pathology by affecting alternative splicing of CLU [31]. Other rare non-synonymous single nucleotide variations have also been identified, along with an in-frame 9-bp deletion, that could possibly/probably disturb clusterin structure and function (Table 1). Findings summarized in Table 1 include mutations that could affect the β-chain domain of clusterin or are positioned in the intron sequence with high regulatory potential [32].
Sortilin-Related Receptor-1
Sortilin-related receptor-1 (SORL1) is considered a member of low-density lipoprotein receptor family and a member of the vacuolar protein sorting 10 (Vps10) family of receptors. There are indications that SORL1 could be involved in APP processing and trafficking, and that it could be responsible for directing Aβ toward lysosomes [33]. However, as a member of LDLR family and an ApoE receptor, SORL1 also plays a role in lipid metabolism. SORL1 was first suggested as a potential risk factor for late-onset Alzheimer’s disease by Rogaeva and colleagues [34], and this was later confirmed by other more comprehensive studies [14, 35]. One of the possibilities is that the mutations in SORL1 gene (Table 1) affect SORL1 expression and BDNF-induced APP processing [36].
ATP-Binding Cassette Subfamily A Member 7
ATP-binding cassette subfamily A member 7 (ABCA7) belongs to a family of ABC transporters that are responsible for transporting various molecules across cellular membranes. The exact function of ABCA7 still unknown, but there are indications that this protein could play a role in lipid homeostasis and the immune system. Therefore, the mutations in ABCA7 gene could contribute to Alzheimer’s disease development by affecting its interaction with ApoE and lipid metabolism and/or by modulating the immune response and the clearance of amyloid plaques. ABCA7 was associated with Alzheimer’s disease in 2011 in a large-scale GWAS analysis [37]. The reported mutations in ABCA7 gene (Table 1) mostly lead to alterations in gene expression [38], but some rare loss-of-function mutations have also been reported [39].
Bridging Integrator Protein-1
Bridging integrator protein-1 (BIN1) is an amphiphysin involved in caspase-independent cell death pathways and clathrin-mediated endocytic pathway [29, 30]. Few GWAS have identified mutations in BIN1 gene (Table 1) associated with Alzheimer’s disease diagnosis [29, 30, 40]. The study by Chapuis and colleagues [41] suggested that increased BIN1 gene expression in dementia patients mediates Alzheimer’s disease risk by modulating tau pathology.
Phosphatidylinositol Binding Clathrin Assembly Protein
Phosphatidylinositol binding clathrin assembly protein (PICALM) is, similarly to BIN1, involved in clathrin-mediated endocytic pathway. Certain SNPs in PICALM gene (Table 1) were found to be associated with the risk of developing Alzheimer’s disease. There are even indications that this protein is involved in the internalization of APP and Aβ production [42], Aβ and tau clearance [43, 44].
Complement C3b/C4b Receptor 1
Complement C3b/C4b receptor 1 (CR1) is a glycoprotein belonging to the receptors of complement activation (RCA) family. CR1 regulates complement activation, but it is also participating in innate immune responses. It is expressed by many cell types, including erythrocytes, leukocytes, and dendritic cells. Different SNPs and an intragenic functional copy-number variation [45, 46] in CR1 gene (Table 1) have been associated with increased risk of developing Alzheimer’s disease. Mentioned copy-number variation in CR1 gene results in two different CR1 protein isoforms (CR1-F and CR1-S), differentiating in the number of C3b/C4b, and cofactor activity binding sites [45, 46], but the exact mechanism of the association with Alzheimer’s disease pathology is not known.
CD33
CD33 is a cell surface receptor that mediates cell–cell interaction. This member of the sialic acid-binding receptor family transmembrane proteins is an important mediator of cell growth and survival, and one of key players in clathrin-independent endocytic pathway and innate and adaptive immune system functions [37, 47]. There is evidence of a positive correlation between the expression of CD33 in microglial cells, amyloid plaque burden and decline in cognitive functions [48, 49]. Different GWAS found a significant association between certain SNPs (Table 1) and late-onset Alzheimer’s disease. One of these SNPs, rs3865444, was associated with the modifications in CD33 level and amyloid pathology [49, 50].
Membrane-Spanning 4-Domains Subfamily A Gene Cluster
Membrane-spanning 4-domains subfamily A gene cluster (MS4A) gene products are transmembrane proteins with at least four transmembrane domains. The genes belonging to this cluster family are not very well characterized, but they might play a role in immunity and intracellular protein trafficking in microglia [51, 52]. There are indications that genes within the MS4A gene cluster regulate soluble triggering receptor expressed on myeloid cells 2 (sTREM2) levels, linking this gene cluster family with Alzheimer’s disease pathogenesis [52]. Three members of MS4A family (MS4A4A, MS4A4E, MS4A6E) have been linked to Alzheimer’s disease by GWAS (Table 1), more precisely, SNPs rs670139 (MS4A4E), rs4938933 and rs1562990 (region between MS4A4E and MS4A4A), and rs610932 and rs983392 (MS4A6A) [53].
Triggering Receptor Expressed on Myeloid Cells 2
Triggering receptor expressed on myeloid cells 2 (TREM2) gene product is an important part of transmembrane receptor-signaling complex that is very abundant on the cell surface of microglia where it plays an important role in downregulation of inflammation, microglial survival and activation, and phagocytosis [54]. There is evidence of high involvement of TREM2 in Alzheimer’s disease pathology. Different TREM2 variants (Table 1) have been associated with Alzheimer’s disease by few meta-analysis and GWAS. These variants include missense mutation rs75932628, rs6916710, rs6922617 (Table 1). TREM2 mutations have been linked to extensive brain atrophy in Alzheimer’s disease patients and with other neuropathological phenotypes characteristic for Alzheimer’s disease [52].
CD2-Associated Protein
CD2-associated protein (CD2AP) is a cytoplasmic protein involved in cytoskeletal structure regulation, receptor-mediated endocytosis, intracellular trafficking, cytokinesis, cell adhesion, and apoptosis. Few GWAS pointed to CD2AP SNPs (rs9296559, rs9349407) as loci possibly associated with late-onset Alzheimer’s disease (Table 1). SNP rs9349407 could be associated with increased neuritic plaque burden in patients with diagnosed Alzheimer’s disease [55].
Ephrin Type-A Receptor 1
Ephrin type-A receptor 1 (EPHA1) is a tyrosine kinase family member important during the nervous system development and during the formation of synapse. It binds to membrane-bound ephrin-A family ligands leading to bidirectional signaling between two adjacent cells, directing cell adhesion and migration. EPHA1 could potentially have a role in the microglial immune response in Alzheimer’s disease. Two EPHA1 gene variants (rs11771145 and rs11767557) were associated with Alzheimer’s disease risk (Table 1).
Conclusion
The knowledge about the genetic background of early-onset familial Alzheimer’s disease allowed detection of mutations in APP, PSEN1, and PSEN2 as a predictive/diagnostic screening, but only for these rare cases of autosomal dominant Alzheimer’s disease. In the case of late-onset Alzheimer’s disease, there is still much of the heritability that remains unexplained, even though, with the help from GWAS, there are now more than 20 different identified loci that have been associated with late-onset complex Alzheimer’s disease. In the case of APOE, the evidence from an extensive meta-analysis shows that around 75% of individuals that carry one APOE ε4 allele never develop Alzheimer’s disease, and around 50% of individuals diagnosed with Alzheimer’s disease are not the carriers of this risk allele [56]. Therefore, the use of genetic variations identified by GWAS, for more effective diagnosis of Alzheimer’s disease, is still far away from the clinical application, because the contribution of these variations to the relative risk of developing Alzheimer’s disease is limited. In the light of prediction and prevention of Alzheimer’s disease, there is much more we can expect from the additive genetic risk scores, which combine additive effects of numerous susceptibility loci.
References
Cerejeira J, Lagarto L, Mukaetova-Ladinska EB. Behavioral and psychological symptoms of dementia. Front Neurol. 2012;3:73.
Reitz C (2016) Toward precision medicine in Alzheimer’s disease. Ann Transl Med. 2016;4(6):107.
Hampel H, O’Bryant S, Durrleman S, Younesi E, Rojkova K, Escott-Price V, et al. A precision medicine Initiative for Alzheimer’s disease: the road ahead to biomarker-guided integrative disease modeling. Climacteric. 2017;20(2):107–18.
Nikolac Perkovic M, Svob Strac D, Tudor L, Konjevod M, Nedic Erjavec G, Pivac N. Catechol-O-methyltransferase, cognition and Alzheimer’s disease. Curr Alzheimer Res. 2018;15(5):408–19.
Duarte A, Santos MS, Oliveira CR. Moreira P. Brain insulin signalling, glucose metabolism and females’ reproductive aging: a dangerous triad in Alzheimer’s disease. Neuropharmacology. 2018;136(Pt B):223–42.
Van Cauwenberghe C, Van Broeckhoven C, Sleegers K. The genetic landscape of Alzheimer disease: clinical implications and perspectives. Genet Med. 2015;18(5):421–30.
Levy-Lahad E, Wasco W, Poorkaj P, Romano DM, Oshima J, Pettingell WH, et al. Candidate gene for the chromosome 1 familial Alzheimer’s disease locus. Science. 1995;269(5226):973–7.
Kopan R, Goate A. A common enzyme connects Notch signaling and Alzheimer’s disease. Genes Dev. 2000;14(22):2799–806.
Marambaud P, Shioi J, Serban G, Georgakopoulos A, Sarner S, Nagy V, et al. A presenilin-1/γ-secretase cleavage releases the E-cadherin intracellular domain and regulates disassembly of adherens junctions. EMBO J. 2002;21(8):1948–56.
Lleó A, Waldron E, von Arnim CA, Herl L, Tangredi MM, Peltan ID, et al. Low density lipoprotein receptor-related protein (LRP) interacts with presenilin 1 and is a competitive substrate of the amyloid precursor protein (APP) for gamma-secretase. J Biol Chem. 2005;280:27303–9.
Vidal GA, Naresh A, Marrero L, Jones FE. Presenilin-dependent gamma-secretase processing regulates multiple ERBB4/HER4 activities. J Biol Chem. 2005;280:19777–83.
Dai MH, Zheng H, Zeng LD, Zhang Y. The genes associated with early-onset Alzheimer’s disease. Oncotarget. 2017;9(19):15132–43.
Larsson M, Duffy DL, Zhu G, Liu JZ, Macgregor S, McRae AF, et al. GWAS findings for human iris patterns: associations with variants in genes that influence normal neuronal pattern development. Am J Hum Genet. 2011;89(2):334–43.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45(12):1452–8.
Liu Y, Yu JT, Wang HF, Hao XK, Yang YF, Jiang T, et al. Association between NME8 locus polymorphism and cognitive decline, cerebrospinal fluid and neuroimaging biomarkers in Alzheimer’s disease. PLoS ONE. 2014;9:e114777.
Ruiz A, Heilmann S, Becker T, Hernández I, Wagner H, Thelen M, et al. Follow-up of loci from the International Genomics of Alzheimer’s Disease Project identifies TRIP4 as a novel susceptibility gene. Transl Psychiatry. 2014;4:e358.
Rosenthal SL, Barmada MM, Wang X, Demirci FY, Kamboh MI. Connecting the dots: potential of data integration to identify regulatory SNPs in late-onset Alzheimer’s disease GWAS findings. PLoS ONE. 2014;9(4):e95152.
Allen M, Kachadoorian M, Carrasquillo MM, Karhade A, Manly L, Burgess JD, et al. Late-onset Alzheimer disease risk variants mark brain regulatory loci. Neurol Genet. 2015;1(2):e15.
Jiao B, Liu X, Zhou L. Polygenic analysis of late-onset Alzheimer’s disease from mainland China. PLoS ONE. 2015;10:e0144898.
Phillips MC. Apolipoprotein E isoforms and lipoprotein metabolism. IUBMB Life. 2014;66(9):616–23.
Lai MK, Tsang SW, Garcia-Alloza M, Minger SL, Nicoll JA, Esiri MM, et al. Selective effects of the APOE ε4 allele on presynaptic cholinergic markers in the neocortex of Alzheimer’s disease. Neurobiol Dis. 2006;22(3):555–61.
Corder EH, Saunders AM, Risch NJ, Strittmatter WJ, Schmechel DE, Gaskell PC Jr, et al. Protective effect of apolipoprotein E type 2 allele for late onset Alzheimer disease. Nat Genet. 1994;7(2):180–4.
Tiraboschi P, Hansen LA, Masliah E, Alford M, Thal LJ, Corey-Bloom J. Impact of APOE genotype on neuropathologic and neurochemical markers of Alzheimer disease. Neurology. 2004;62(11):1977–83.
Liu Y, Paajanen T, Westman E, Zhang Y, Wahlund LO, Simmons A, et al. APOE ε2 allele is associated with larger regional cortical thicknesses and volumes. Dement Geriatr Cogn Disord. 2010;30(3):229–37.
Deane R, Sagare A, Hamm K, Parisi M, Lane S, Finn MB, et al. ApoE isoform-specific disruption of amyloid beta peptide clearance from mouse brain. J Clin Invest. 2008;118(12):4002–13.
Matukumalli SR, Tangirala R, Rao CM. Clusterin: full-length protein and one of its chains show opposing effects on cellular lipid accumulation. Sci Reports. 2017;7:41235.
Li X, Ma Y, Wei X, Li Y, Wu H, Zhuang J, Zhao Z. Clusterin in Alzheimer’s disease: a player in the biological behaviour of amyloid-beta. Neurosci Bull. 2014;30(1):162–8.
Zlokovic BV. Cerebrovascular transport of Alzheimer’s amyloid beta and apolipoproteins J and E: possible anti-amyloidogenic role of the blood-brain barrier. Life Sci. 1996;59:1483–97.
Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML, et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat Genet. 2009;41(10):1088–93.
Lambert JC, Heath S, Even G, Campion D, Sleegers K, Hiltunen M, et al. Genome-wide association study identifies variants at CLU and CR29 associated with Alzheimer’s disease. Nat Genet. 2009;41(10):1094–9.
Szymanski M, Wang R, Bassett SS, Avramopoulos D. Alzheimer’s risk variants in the clusterin gene are associated with alternative splicing. Transl Psychiatry. 2011;1(7):e18.
Bettens K, Brouwers N, Engelborghs S, Lambert JC, Rogaeva E, Vandenberghe R, et al. Both common variations and rare non-synonymous substitutions and small insertion/deletions in CLU are associated with increased Alzheimer risk. Mol Neurodegener. 2012;16(7):3.
Offe K, Dodson SE, Shoemaker JT, Fritz JJ, Gearing M, Levey AI, et al. The lipoprotein receptor LR11 regulates amyloid beta production and amyloid precursor protein traffic in endosomal compartments. J Neurosci. 2006;26(5):1596–603.
Rogaeva E, Meng Y, Lee JH, Gu Y, Kawarai T, Zou F, et al. The neuronal sortilin-related receptor SORL1 is genetically associated with Alzheimer disease. Nat Genet. 2007;39(2):168–77.
Vardarajan BN, Zhang Y, Lee JH, Cheng R, Bohm C, Ghani M, et al. Coding mutations in SORL1 and Alzheimer disease. Ann Neurol. 2015;77:215–27.
Young JE, Boulanger-Weill J, Williams DA, Woodruff G, Buen F, Revilla AC, et al. Elucidating molecular phenotypes caused by the SORL1 Alzheimer’s disease genetic risk factor using human induced pluripotent stem cells. Cell Stem Cell. 2015;16(4):373–85.
Hollingworth P, Harold D, Sims R, Gerrish A, Lambert JC, Carrasquillo MM, et al. Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat Genet. 2011;43(5):429–35.
Vasquez JB, Fardo DW, Estus S. ABCA7 expression is associated with Alzheimer’s disease polymorphism and disease status. Neurosci Lett. 2013;556:58–62.
Steinberg S, Stefansson H, Jonsson T, Johannsdottir H, Ingason A, Helgason H, et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat Genet. 2015;47(5):445–7.
Seshadri S, Fitzpatrick AL, Ikram MA, DeStefano AL, Gudnason V, Boada M, et al. Genome-wide analysis of genetic loci associated with Alzheimer disease. JAMA. 2010;303(18):1832–40.
Chapuis J, Hansmannel F, Gistelinck M, Mounier A, Van Cauwenberghe C, Kolen KV, et al. Increased expression of BIN1 mediates Alzheimer genetic risk by modulating tau pathology. Mol Psychiatry. 2013;18(11):1225–34.
Xiao Q, Gil SC, Yan P, Wang Y, Han S, Gonzales E, et al. Role of phosphatidylinositol clathrin assembly lymphoid-myeloid leukemia (PICALM) in intracellular amyloid precursor protein (APP) processing and amyloid plaque pathogenesis. J Biol Chem. 2012;287:21279–89.
Zhao Z, Sagare AP, Ma Q, Halliday MR, Kong P, Kisler K, et al. Central role for PICALM in amyloid-β blood-brain barrier transcytosis and clearance. Nat Neurosci. 2015;18:978–87.
Moreau K, Fleming A, Imarisio S, Lopez Ramirez A, Mercer JL, Jimenez-Sanchez M, et al. PICALM modulates autophagy activity and tau accumulation. Nat Commun. 2014;5:4998.
Brouwers N, Van Cauwenberghe C, Engelborghs S, Lambert JC, Bettens K, Le Bastard N, et al. Alzheimer risk associated with a copy number variation in the complement receptor 1 increasing C3b/C4b binding sites. Mol Psychiatry. 2012;17(2):223–33.
Hazrati LN, Van Cauwenberghe C, Brooks PL, Brouwers N, Ghani M, Sato C, et al. Genetic association of CR45 with Alzheimer’s disease: a tentative disease mechanism. Neurobiol Aging. 2012;33(12):2949.e5–12.
Naj AC, Jun G, Beecham GW, Wang LS, Vardarajan BN, Buros J, et al. Common variants at MS4A4/MS4A6E, CD2AP, CD33 and EPHA1 are associated with late-onset Alzheimer’s disease. Nat Genet. 2011;43(5):436–41.
Karch CM, Jeng AT, Nowotny P, Cady J, Cruchaga C, Goate AM. Expression of novel Alzheimer’s disease risk genes in control and Alzheimer’s disease brains. PLoS ONE. 2012;7:e50976.
Griciuc A, Serrano-Pozo A, Parrado AR, Lesinski AN, Asselin CN, Mullin K, et al. Alzheimer’s disease risk gene CD33 inhibits microglial uptake of amyloid beta. Neuron. 2013;78(4):631–43.
Bradshaw EM, Chibnik LB, Keenan BT, Ottoboni L, Raj T, Tang A, et al. CD33 Alzheimer’s disease locus: altered monocyte function and amyloid biology. Nat Neurosci. 2013;16(7):848–50.
Zuccolo J, Deng L, Unruh TL, Sanyal R, Bau JA, Storek J, et al. Expression of MS4A and TMEM176 genes in human B lymphocytes. Front Immunol. 2013;4:195.
Deming Y, Filipello F, Cignarella F, Cantoni C, Hsu S, Mikesell R, et al. The MS4A gene cluster is a key regulator of soluble TREM2 and Alzheimer disease risk. BioRxiv. 2018. https://doi.org/10.1101/352179.
Antunez C, Boada M, González-Pérez A, Gayán J, Ramírez-Lorca R, Marín J, et al. The membrane-spanning 4-domains, subfamily A (MS4A) gene cluster contains a common variant associated with Alzheimer’s disease. Genome Med. 2011;3(5):33.
Ulrich JD, Ulland TK, Colonna M, Holtzman DM. Elucidating the Role of TREM2 in Alzheimer’s disease. Neuron. 2017;94:237–48.
Shulman JM, Chen K, Keenan BT, Chibnik LB, Fleisher A, Thiyyagura P, et al. Genetic susceptibility for Alzheimer disease neuritic plaque pathology. JAMA Neurol. 2013;70(9):1150–7.
Farrer LA, Cupples LA, Haines JL, Hyman B, Kukull WA, Mayeux R, et al. Effects of age, sex, and ethnicity on the association between apolipoprotein E genotype and Alzheimer disease. A meta-analysis. APOE and Alzheimer disease meta analysis consortium. JAMA. 1997;278(16):1349–56.
Hooli BV, Mohapatra G, Mattheisen M, Parrado AR, Roehr JT, Shen Y, et al. Role of common and rare APP DNA sequence variants in Alzheimer disease. Neurology. 2012;78(16):1250–7.
Kasuga K, Shimohata T, Nishimura A, Shiga A, Mizuguchi T, Tokunaga J, et al. Identification of independent APP locus duplication in Japanese patients with early-onset Alzheimer disease. J Neurol Neurosurg Psychiatry. 2009;80(9):1050–2.
McNaughton D, Knight W, Guerreiro R, Ryan N, Lowe J, Poulter M, et al. Duplication of amyloid precursor protein (APP), but not prion protein (PRNP) gene is a significant cause of early onset dementia in a large UK series. Neurobiol Aging. 2012;33(2):426.e13–21.
Thonberg H, Fallström M, Björkström J, Schoumans J, Nennesmo I, Graff C. Mutation screening of patients with Alzheimer disease identifies APP locus duplication in a Swedish patient. BMC Res Notes. 2011;4:476.
Wallon D, Rousseau S, Rovelet-Lecrux A, Quillard-Muraine M, Guyant-Maréchal L, Martinaud O, et al. The French series of autosomal dominant early onset Alzheimer’s disease cases: mutation spectrum and cerebrospinal fluid biomarkers. J Alzheimers Dis. 2012;30(4):847–56.
Brouwers N, Sleegers K, Van Broeckhoven C. Molecular genetics of Alzheimer’s disease: an update. Ann Med. 2008;40(8):562–83.
Kamino K, Orr HT, Payami H, Wijsman EM, Alonso ME, Pulst SM, et al. Linkage and mutational analysis of familial Alzheimer disease kindreds for the APP gene region. Am J Hum Genet. 1992;51(5):998–1014.
Mullan M, Crawford F, Axelman K, Houlden H, Lilius L, Winblad B, et al. A pathogenic mutation for probable Alzheimer’s disease in the APP gene at the N-terminus of beta-amyloid. Nat Genet. 1992;1(5):345–7.
Nilsberth C, Westlind-Danielsson A, Eckman CB, Forsell C, Axelman K, Luthman J, et al. The Arctic APP mutation (E693G) causes Alzheimer’s disease through a novel mechanism: increased amyloid β protofibril formation and decreased amyloid β levels in plasma and conditioned media. Neurobiol Aging. 2000;21 Suppl 1:S58.
Nilsberth C, Westlind-Danielsson A, Eckman CB, Condron MM, Axelman K, Forsell C, et al. The ‘Arctic’ APP mutation (E693G) causes Alzheimer’s disease by enhanced Abeta protofibril formation. Nat Neurosci. 2001;4(9):887–93.
Tomiyama T, Nagata T, Shimada H, Teraoka R, Fukushima A, Kanemitsu H, et al. A new amyloid beta variant favoring oligomerization in Alzheimer’s-type dementia. Ann Neurol. 2008;63:377–87.
Wakutani Y, Watanabe K, Adachi Y, Wada-Isoe K, Urakami K, Ninomiya H, et al. Novel amyloid precursor protein gene missense mutation (D678N) in probable familial Alzheimer’s disease. J Neurol Neurosurg Psychiatry. 2004;75(7):1039–42.
Wakutani Y. Gene symbol: APP. Disease: Familial Alzheimer’s disease. Hum Genet. 2005;117(2–3):299.
Ancolio K, Dumanchin C, Barelli H, Warter JM, Brice A, Campion D, et al. Unusual phenotypic alteration of beta amyloid precursor protein (betaAPP) maturation by a new Val-715 → Met betaAPP-770 mutation responsible for probable early-onset Alzheimer’s disease. Proc Natl Acad Sci USA. 1999;96(7):4119–24.
Blauwendraat C, Wilke C, Jansen IE, Schulte C, Simón-Sánchez J, Metzger FG, et al. Pilot whole-exome sequencing of a German early-onset Alzheimer’s disease cohort reveals a substantial frequency of PSEN2 variants. Neurobiol Aging. 2016;37(208):e11–7.
Brooks WS, Martins RN, De Voecht J, Nicholson GA, Schofield PR, Kwok JB, et al. A mutation in codon 717 of the amyloid precursor protein gene in an Australian family with Alzheimer’s disease. Neurosc Lett. 1995;199(3):183–6.
Brouwers N, Sleegers K, Engelborghs S, Bogaerts V, Serneels S, Kamali K, et al. Genetic risk and transcriptional variability of amyloid precursor protein in Alzheimer’s disease. Brain. 2006;129(Pt 11):2984–91.
Campion D, Brice A, Hannequin D, Charbonnier F, Dubois B, Martin C, et al. No founder effect in three novel Alzheimer’s disease families with APP 717 Val → Ile mutation. Clerget-darpoux. French Alzheimer’s Disease Study Group. J Med Genet. 1996;33(8):661–4.
Campion D, Dumanchin C, Hannequin D, Dubois B, Belliard S, Puel M, et al. Early-onset autosomal dominant Alzheimer disease: prevalence, genetic heterogeneity, and mutation spectrum. Am J Hum Genet. 1999;65(3):664–70.
Campion D, Flaman JM, Brice A, Hannequin D, Dubois B, Martin C, et al. Mutations of the presenilin I gene in families with early-onset Alzheimer’s disease. Hum Mol Genet. 1995;4(12):2373–7.
Clarimón J, Guerreiro R, Lleó A, Guardia C, Blesa F, Gómez-Isla T, et al. Genetic screening in a large cohort of early-onset Alzheimer’s disease patients from Spain: novel mutations in the amyloid precursor protein and presenilines. Alzheimer Dement. 2008;4 Supp 2:T583.
Cruts M, Dermaut B, Kumar-Singh S, Rademakers R, Van den Broeck M, Stögbauer F, et al. Novel German APP V715A mutation associated with presenile Alzheimer’s disease. Neurobiol Aging. 2002;23(1S):S327.
Cruts M, Dermaut B, Rademakers R, Van den Broeck M, Stogbauer F, Van Broeckhoven C. Novel APP mutation V715A associated with presenile Alzheimer’s disease in a German family. J Neurol. 2003;250(11):1374–5.
De Jonghe C, Kumar-Singh S, Cruts M, Kleinert R, Vanderstichele H, Vanmechelen EJM, et al. Unusual Aβ amyloid deposition in Alzheimer’s disease due tu an APP T714I mutation at the γ42-secretase site. Neurobiol Aging. 2000;21(1):S200.
De Jonghe C, Esselens C, Kumar-Singh S, Craessaerts K, Serneels S, et al. Pathogenic APP mutations near the gamma-secretase cleavage site differentially affect Abeta secretion and APP C-terminal fragment stability. Hum Mol Genet. 2001;10(16):1665–71.
Dobricic V, Stefanova E, Jankovic M, Gurunlian N, Novakovic I, Hardy J, et al. Genetic testing in familial and young-onset Alzheimer’s disease: mutation spectrum in a Serbian cohort. Neurobiol Aging. 2012;3(1481):e7–12.
Eckman CB, Mehta ND, Crook R, Perez-tur J, Prihar G, Pfeiffer E, et al. A new pathogenic mutation in the APP gene (I716V) increases the relative proportion of A beta 42(43). Hum Mol Genet. 1997;6(12):2087–9.
Edwards-Lee T, Ringman JM, Chung J, Werner J, Morgan A, St George Hyslop P, et al. An African American family with early-onset Alzheimer disease and an APP (T714I) mutation. Neurology. 2005;64(2):377–9.
Fidani L, Rooke K, Chartier-Harlin MC, Hughes D, Tanzi R, Mullan M, et al. Screening for mutations in the open reading frame and promoter of the beta-amyloid precursor protein gene in familial Alzheimer’s disease: identification of a further family with APP717 Val → Ile. Hum Mol Genet. 1992;1(3):165–8.
Finckh U, Muller-Thomsen T, Mann U, Eggers C, Marksteiner J, Meins W, et al. High prevalence of pathogenic mutations in patients with early-onset dementia detected by sequence analyses of four different genes. Am J Hum Genet. 2000;66(1):110–7.
Finckh U, Kuschel C, Anagnostouli M, Patsouris E, Pantes GV, Gatzonis S, et al. Novel mutations and repeated findings of mutations in familial Alzheimer disease. Neurogenetics. 2005;6(2):85–9.
Ghetti B, Hake AM, Murrell JR, Epperson F, Farlow MR, Vidal R, Spina S. Familial Alzheimer disease associated with the V717L amyloid precursor protein gene mutation: Neuropathological characterization. Alzheimers Dement. 2008;4 Suppl 4:T585.
Goate A, Chartier-Harlin MC, Mullan M, Brown J, Crawford F, Fidani L, et al. Segregation of a missense mutation in the amyloid precursor protein gene with familial Alzheimer’s disease. Nature. 1991;349(6311):704–6.
Godbolt AK, Beck JA, Collinge JC, Cipolotti L, Fox NC, Rossor MN. A second family with familial AD and the V717L APP mutation has a later age at onset. Neurology. 2006;66(4):611–2.
Guardia-Laguarta C, Pera M, Clarimón J, Molinuevo JL, Sánchez-Valle R, Lladó A, et al. Clinical, neuropathologic, and biochemical profile of the amyloid precursor protein I716F mutation. J Neuropathol Exp Neurol. 2010;69(1):53–9.
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E, et al. TREM2 variants in Alzheimer’s disease. N Engl J Med. 2013;368(2):117–27.
Hardy J, Mullan M, Chartier-Harlin M-C, Brown J, Goate A, Rossor M, Collinge J, et al. Molecular classification of Alzheimer’s disease. Lancet. 1991;337(8753):1342–3.
Janssen JC, Beck JA, Campbell TA, Dickinson A, Fox NC, Harvey RJ, et al. Early onset familial Alzheimer’s disease: mutation frequency in 31 families. Neurobiol Aging. 2002;23(1):S311.
Janssen JC, Beck JA, Campbell TA, Dickinson A, Fox NC, Harvey RJ, et al. Early onset familial Alzheimer’s disease: mutation frequency in 31 families. Neurology. 2003;60(2):235–9.
Jiao B, Tang B, Liu X, Xu J, Wang Y, Zhou L, et al. Mutational analysis in early-onset familial Alzheimer’s disease in Mainland China. Neurobiol Aging. 2014;35(8):1957.e1–6.
Kumar-Singh S, De Jonghe C, Cruts M, Kleinert R, Wang R, Mercken M, et al. Nonfibrillar diffuse amyloid deposition due to a gamma(42)-secretase site mutation points to an essential role for N-truncated abeta(42) in Alzheimer’s disease. Hum Mol Genet. 2000;9(18):2589–98.
Kwok JBJ, Li Q-X, Hallupp M, Milward L, Whyte S, Schofield PR. Novel familial early-onset Alzheimer’s disease mutation (Leu723Pro) in amyloid precursor protein (APP) gene increases production of 42(43) amino-acid isoform of amyloid beta peptide. Neurobiol Aging. 1998;19(4):S91.
Kwok JB, Li QX, Hallupp M, Whyte S, Ames D, Beyreuther K, Masters CL, Schofield PR. Novel Leu723Pro amyloid precursor protein mutation increases amyloid beta42(43) peptide levels and induces apoptosis. Ann Neurol. 2000;47(2):249–53.
Matsumura Y, Kitamura E, Miyoshi K, Yamamoto Y, Furuyama J, Sugihara T. Japanese siblings with missense mutation (717Val → Ile) in amyloid precursor protein of early-onset Alzheimer’s disease. Neurology. 1996;46(6):1721–3.
Murrell J, Farlow M, Ghetti B, Benson MD. A mutation in the amyloid precursor protein associated with hereditary Alzheimer’s disease. Science. 1991;254(5028):97–9.
Murrell JR, Hake AM, Quaid KA, Farlow MR, Ghetti B. Early-onset Alzheimer disease caused by a new mutation (V717L) in the amyloid precursor protein gene. Arch Neurol. 2000;57:885–7.
Naruse S, Igarashi S, Kobayashi H, Aoki K, Inuzuka T, Kaneko K, et al. Mis-sense mutation Val → Ile in exon 17 of amyloid precursor protein gene in Japanese familial Alzheimer’s disease. Lancet. 1991;337(8747):978–9.
Park HK, Na DL, Lee JH, Kim JW, Ki CS. Identification of PSEN1 and APP gene mutations in Korean patients with early-onset Alzheimer’s disease. J Korean Med Sci. 2008;23(2):213–7.
Raux G, Guyant-Marechal L, Martin C, Bou J, Penet C, Brice A, et al. Molecular diagnosis of autosomal dominant early onset Alzheimer’s disease: an update. J Med Genet. 2005;42(10):793–5.
Sassi C, Guerreiro R, Gibbs R, Ding J, Lupton MK, Troakes C, et al. Exome sequencing identifies 2 novel presenilin 1 mutations (p.L166V and p.S230R) in British early-onset Alzheimer’s disease. Neurobiol Aging. 2014;35(10):2422.e13–6.
Sassi C, Guerreiro R, Gibbs R, Ding J, Lupton MK, Troakes C, et al. Investigating the role of rare coding variability in Mendelian dementia genes (APP, PSEN1, PSEN2, GRN, MAPT, and PRNP) in late-onset Alzheimer’s disease. Neurobiol Aging. 2014;35(12):2881.e1–6.
Sorbi S, Nacmias B, Forleo P, Piacentini S, Amaducci L, Provinciali L. APP717 and Alzheimer’s disease in Italy. Nat Genet. 1993;4(1):10.
Sorbi S, Nacmias B, Forleo P, Piacentini S, Latorraca S, Amaducci L. Epistatic effect of APP717 mutation and apolipoprotein E genotype in familial Alzheimer’s disease. Ann Neurol. 1995;38(1):124–7.
Tedde A, Nacmias B, Ciantelli M, Forleo P, Cellini E, Bagnoli S, et al. Identification of new presenilin gene mutations in early-onset familial Alzheimer disease. Arch Neurol. 2003;60(11):1541–4.
Terreni L, Fogliarino S, Franceschi, Forloni G. Novel pathogenic mutation in an Italian patient with familial Alzheimer’s disease detected in APP gene. Neurobiol Aging. 2002;23(1):S319.
Yoshioka K, Miki T, Katsuya T, Ogihara T, Sakaki Y. The 717Val → Ile substitution in amyloid precursor protein is associated with familial Alzheimer’s disease regardless of ethnic groups. Biochem Biophys Res Commun. 1991;178(3):1141–6.
Yoshizawa T, Komatsuzaki Y, Iwamoto H, Mizusawa H, Kanazawa I. Screening of the mis-sense mutation producing the 717Val → Ile substitution in the amyloid precursor protein in Japanese familial and sporadic Alzheimer’s disease. J Neurol Sci. 1993;117(1–2):12–5.
Zekanowski C, Styczynska M, Peplonska B, Gabryelewicz T, Religa D, Ilkowski J, et al. Mutations in presenilin 1, presenilin 2 and amyloid precursor protein genes in patients with early-onset Alzheimer’s disease in Poland. Exp Neurol. 2003;184(2):991–6.
Theuns J, Marjaux E, Vandenbulcke M, Van Laere K, Kumar-Singh S, Bormans G, et al. Alzheimer dementia caused by a novel mutation located in the APP C-terminal intracytosolic fragment. Hum Mutat. 2006;27(9):888–96.
Alberici A, Bonato C, Borroni B, Cotelli M, Mattioli F, Binetti G, et al. Dementia, delusions and seizures: storage disease or genetic AD. Eur J Neurol. 2007;14(9):1057–9.
Aldudo J, Bullido MJ, Arbizu T, Oliva R, Valdivieso F. Identification of a novel mutation (Leu282Arg) of the human presenilin 1 gene in Alzheimer’s disease. Neurosci Lett. 1998;240(3):174–6.
Aldudo J, Bullido MJ, Valdivieso F. DGGE method for the mutational analysis of the coding and proximal promoter regions of the Alzheimer’s disease presenilin-1 gene: two novel mutations. Hum Mutat. 1999;14(5):433–9.
Antonell A, Balasa M, Oliva R, Lladó A, Bosch B, Fabregat N, et al. A novel PSEN1 gene mutation (L235R) associated with familial early-onset Alzheimer’s disease. Neurosci Lett. 2011;496(1):40–2.
Aoki M, Abe K, Oda N, Ikeda M, Tsuda T, Kanai M, et al. A presenilin-1 mutation in a Japanese family with Alzheimer’s disease and distinctive abnormalities on cranial MRI. Neurology. 1997;48(4):1118–20.
Arai N, Kishino A, Takahashi Y, Morita D, Nakamura K, Yokoyama T, et al. Familial cases presenting very early onset autosomal dominant Alzheimer’s disease with I143T in presenilin-1 gene: implication for genotype-phenotype correlation. Neurogenetics. 2008;9(1):65–7.
Arango D, Cruts M, Torres O, Backhovens H, Serrano ML, Villareal E, et al. Systematic genetic study of Alzheimer disease in Latin America: mutation frequencies of the amyloid beta precursor protein and presenilin genes in Colombia. Am J Med Genet. 2001;103(2):138–43.
Athan ES, Williamson J, Ciappa A, Santana V, Romas SN, Lee JH, et al. A founder mutation in presenilin 1 causing early-onset Alzheimer disease in unrelated Caribbean Hispanic families. JAMA. 2001;286(18):2256–63.
Axelman K, Basun H, Lannfelt L. Wide range of disease onset in a family with Alzheimer disease and a His163Tyr mutation in the presenilin-1 gene. Arch Neurol. 1998;55(5):698–702.
Batelli S, Albani D, Prato F, Polito L, Franceschi M, Gavazzi A, Forloni G. Early-onset Alzheimer disease in an Italian family with presenilin-1 double mutation E318G and G394V. Alzheimer Dis Assoc Disord. 2008;22(2):184–7.
Boteva K, Vitek M, Mitsuda H, de Silva H, Xu PT, Small G, Gilbert JR. Mutation analysis of presenillin 1 gene in Alzheimer’s disease. Lancet. 1996;347(8994):130–1.
Brouwers N, Sleegers K, Theuns J, Engelborghs S, Bogaerts V, Serneels S, et al. Contribution of dementia genes to Alzheimer’s disease in Belgium. Alzheimer Dement. 2006;2(3):S191.
Cervenakova L, Brown P, Sandoval F, Foncin J-F, Garruto R, Polinsky RJ, et al. Identification of presenilin-1 gene point mutations in early-onset Alzheimer’s disease families. Am J Hum Genet. 1996;59(Suppl):A252.
Church A, Prescott J, Lillis S, Rees J, Chance P, Williamson K, Morris HR. A novel presenilin 1 mutation, I202F occurring at a previously predicted pathogenic site causing autosomal dominant Alzheimer’s disease. Neurobiol Aging. 2011;32(556):e1–2.
Clarimón J, Guerreiro R, Lleó A, Guardia C, Blesa F, Gómez-Isla T, et al. Genetic screening in a large cohort of early-onset Alzheimer’s disease patients from Spain: novel mutations in the amyloid precursor protein and presenilines. Alzheimers Dement. 2008;4(Supp 2):T583.
Clark RF, Hutton M, Fuldner RA, Froelich S, Karran E, Talbot C, et al. The structure of the presenilin 1 (S182) gene and identification of six novel mutations in early onset AD families. Nat Genet. 1995;11(2):219–22.
Coleman P, Kurlan R, Crook R, Werner J, Hardy J. A new presenilin Alzheimer’s disease case confirms the helical alignment of pathogenic mutations in transmembrane domain 5. Neurosci Lett. 2004;364(3):139–40.
Crook R, Ellis R, Shanks M, Thal LJ, Perez-Tur J, Baker M, et al. Early-onset Alzheimer’s disease with a presenilin-1 mutation at the site corresponding to the Volga German presenilin-2. Ann Neurol. 1997;42(1):124–8.
Cruts M, Backhovens H, Wang SY, Gassen GV, Theuns J, De Jonghe CD, Wehnert A, De Voecht J, De Winter G, Cras P, Bruyland M, Datson N, Weissenbach J, den Dunnen JT, Martin J-J, Hendriks L, Van Broeckhoven C. Molecular genetic analysis of familial early-onset Alzheimer’s disease linked to chromosome 14q24.3. Hum Mol Genet. 1995;4(12):2363–72.
Cruts M, van Duijn CM, Backhovens H, Van den Broeck M, Wehnert A, Serneels S, et al. Estimation of the genetic contribution of presenilin-1 and -2 mutations in a population-based study of presenile Alzheimer disease. Hum Mol Genet. 1998;7:43–51.
Doran M, Larner AJ. Familial Alzheimer’s disease due to presenilin-1 Y115C mutation. J Neurol. 2006;253 Suppl 2:ii91,2006.
Dowjat K, Kuchna I, Wisniewski K, Wisniewski T, Wegiel J. Another highly pathogenic Alzheimer presenilin-1 mutation in codon 117: genotype-phenotype comparison of P117S and P117L mutations. Neurobiol Aging. 2002;23(1S):S219.
Dowjat WK, Kuchna I, Wisniewski T, Wegiel J. A novel highly pathogenic Alzheimer presenilin-1 mutation in codon 117 (Pro117Ser): comparison of clinical, neuropathological and cell culture phenotypes of Pro117Leu and Pro117Ser mutations. J Alzheimers Dis. 2004;6(1):31–43.
Dumanchin C, Brice A, Campion D, Hannequin D, Martin C, Moreau V, et al. De novo presenilin 1 mutations are rare in clinically sporadic, early onset Alzheimer’s disease cases. French Alzheimer’s Disease Study Group. J Med Genet. 1998;35(8):672–3.
Edwards-Lee T, Wen J, Bell J, Hardy J, Chung J, Momeni P. A presenilin-1 mutation (T245P) in transmembrane domain 6 causes early onset Alzheimer’s disease. Neurosci Lett. 2006;398(3):251–2.
Ezquerra M, Carnero C, Blesa R, Oliva R. A novel presenilin 1 mutation (Leu166Arg) associated with early-onset Alzheimer disease. Arch Neurol. 2000;57(4):485–8.
Fang B, Jia L, Jia J. Chinese Presenilin-1 V97L mutation enhanced Abeta42 levels in SH-SY5Y neuroblastoma cells. Neurosci Lett. 2006;406(1–2):33–7.
Fang BY, Jia JP. The effect of two newly Chinese presenilin-1 mutations on the sensitivity to trophic factor withdrawal in human neuroblastoma cells. Zhonghua Yi Xue Za Zhi. 2007;87(5):336–40.
Finckh U, Alberici A, Antoniazzi M, Benussi L, Fedi V, Giannini C, Gal A, Nitsch RM, Binetti G. Variable expression of familial Alzheimer disease associated with presenilin 2 mutation M239I. Neurology. 2000;54(10):2006–8.
Forsell C, Froelich S, Axelman K, Vestling M, Cowburn RF, Lilius L, et al. A novel pathogenic mutation (Leu262Phe) found in the presenilin 1 gene in early-onset Alzheimer’s disease. Neurosci Lett. 1997;234(1):3–6.
Gallo M, Marcello N, Curcio SA, Colao R, Geracitano S, Bernardi L, et al. A novel pathogenic PSEN1 mutation in a family with Alzheimer’s disease: phenotypical and neuropathological features. J Alzheimers Dis. 2011;25(3):425–31.
Gliebus G, Rosso A, Lippa CF. Progranulin and beta-amyloid distribution: a case report of the brain from preclinical PS-1 mutation carrier. Am J Alzheimers Dis Other Demen. 2009;24(6):456–60.
Golan M, Lipczynska-Lojkowska W, Krzysko KA, Styczynska M, Luczywek E, Filipek S, et al. Two novel mutations in presenilin 1 (PSEN1) gene connected with atypical familial early-onset Alzheimer’s disease (EOAD). Alzheimer’s and Parkinson’s diseases: insights, progress and perspectives. In: 7th international conference AD/PD 2005 book of abstracts: 24, 2005.
Goldman JS, Reed B, Gearhart R, Kramer JH, Miller BL. Very early-onset familial Alzheimer’s disease: a novel presenilin 1 mutation. Int J Geriatr Psychiatry. 2002;17(7):649–51.
Gómez-Tortosa E, Barquero S, Barón M, Gil-Neciga E, Castellanos F, Zurdo M, et al. Clinical-genetic correlations in familial Alzheimer’s disease caused by presenilin 1 mutations. J Alzheimers Dis. 2010;19(3):873–84.
Guerreiro RJ, Baquero M, Blesa R, Boada M, Brás JM, Bullido MJ, et al. Genetic screening of Alzheimer’s disease genes in Iberian and African samples yields novel mutations in presenilins and APP. Neurobiol Aging. 2010;31(5):725–31.
Gustafson L, Brun A, Englund E, Hagnell O, Nilsson K, Stensmyr M, et al. A 50-year perspective of a family with chromosome-14-linked Alzheimer’s disease. Hum Genet. 1998;102(3):253–7.
Hamaguchi T, Morinaga A, Tsukie T, Kuwano R, Yamada M. A novel presenilin 1 mutation (L282F) in familial Alzheimer’s disease. J Neurol. 2009;256(9):1575–7.
Harvey RJ, Ellison D, Hardy J, Hutton M, Roques PK, Collinge J, Fox NC, Rossor MN. Chromosome 14 familial Alzheimer’s disease: the clinical and neuropathological characteristics of a family with a leucine → serine (L250S) substitution at codon 250 of the presenilin 1 gene. J Neurol Neurosurg Psychiatry. 1998;64(1):44–9.
Heckmann J, de Viliers C, Rutherfoord S, Ramesar R, Morris C, Low R, Kalaria R. Novel presenilin 1 mutation with profound neurofibrillary pathology in an indigenous South African family with early-onset Alzheimer’s disease. Brain. 2004;127(Pt 1):133–42.
Higuchi S, Yoshino A, Matsui T, Matsushita S, Satoh A, Iimura T, et al. A novel PS1 Mutation (W165G) in a Japanese family with early-onset Alzheimer’s disease. Alzheimers Rep. 2000;3(4):227–31.
Houlden H, Crook R, Dolan RJ, McLaughlin J, Revesz T, Hardy J. A novel presenilin mutation (M233V) causing very early onset Alzheimer’s disease with Lewy bodies. Neurosci Lett. 2001;313(1–2):93–5.
Hutton M, Busfield F, Wragg M, Crook R, Perez-Tur J, Clark RF, et al. Complete analysis of the presenilin 1 gene in early onset Alzheimer’s disease. NeuroReport. 1996;7(3):801–5.
Ikeda M, Sharma V, Sumi SM, Rogaeva EA, Poorkaj P, Sherrington R, et al. The clinical phenotype of two missense mutations in the presenilin I gene in Japanese patients. Ann Neurol. 1996;40(6):912–7.
Ikeda M, Yonemura K, Kakuda S, Tashiro Y, Fujita Y, Takai E, et al. Cerebrospinal fluid levels of phosphorylated tau and Aß1-38/Aß1-40/Aß1-42 in Alzheimer’s disease with PS1 mutations. Amyloid. 2013;20(2):107–12.
Ikeuchi T, Kaneko H, Miyashita A, Nozaki H, Kasuga K, Tsukie T, et al. Mutational analysis in early-onset familial dementia in the Japanese population. The role of PSEN1 and MAPT R406 W mutations. Dement Geriatr Cogn Disord. 2008;26(1):43–9.
Jacquier M, Arango D, Torres O, Cruts M, Serrano M, Matallana M, et al. Presenilin mutations in a Colombian familial and sporadic AD sample. Neurobiol Aging. 2000;21(1):S176.
Janssen JC, Lantos PL, Fox NC, Harvey RJ, Beck J, Dickinson A, et al. Autopsy-confirmed familial early-onset Alzheimer disease caused by the l153V presenilin 1 mutation. Arch Neurol. 2001;58(6):953–8.
Jia J, Xu E, Shao Y, Jia J, Sun Y, Li D. One novel presenilin-1 gene mutation in a Chinese pedigree of familial Alzheimer’s disease. J Alzheimers Dis. 2005;7(2):119–24.
Jiang HY, Li GD, Dai SX, Bi R, Zhang DF, Li ZF, et al. Identification of PSEN1 mutations p.M233L and p.R352C in Han Chinese families with early-onset familial Alzheimer’s disease. Neurobiol Aging. 2015;36(3):1602.e3–6.
Jin SC, Pastor P, Cooper B, Cervantes S, Benitez BA, Razquin C, Goate A, et al. Pooled-DNA sequencing identifies novel causative variants in PSEN1, GRN and MAPT in a clinical early-onset and familial Alzheimer’s disease Ibero-American cohort. Alzheimers Res Ther. 2012;4(4):34.
Jorgensen P, Bus C, Pallisgaard N, Bryder M, Jorgensen AL. Familial Alzheimer’s disease co-segregates with a Met146I1e substitution in presenilin-1. Clin Genet. 1996;50(5):281–6.
Kamimura K, Tanahashi H, Yamanaka H, Takahashi K, Asada T, Tabira T. Familial Alzheimer’s disease genes in Japanese. J Neurol Sci. 1998;160(1):76–81.
Kamino K, Sato S, Sakaki Y, Yoshiiwa A, Nishiwaki Y, Takeda M, et al. Three different mutations of presenilin 1 gene in early-onset Alzheimer’s disease families. Neurosci Lett. 1996;208(3):195–8.
Kasuga K, Ohno T, Ishihara T, Miyashita A, Kuwano R, Onodera O, et al. Depression and psychiatric symptoms preceding onset of dementia in a family with early-onset Alzheimer disease with a novel PSEN1 mutation. J Neurol. 2009;256(8):1351–3.
Kauwe JS, Jacquart S, Chakraverty S, Wang J, Mayo K, Fagan AM, et al. Extreme cerebrospinal fluid amyloid beta levels identify family with late-onset Alzheimer’s disease presenilin 1 mutation. Ann Neurol. 2007;61(5):446–53.
Kerchner GA, Holbrook K. Novel presenilin-1 Y159F sequence variant associated with early-onset Alzheimer’s disease. Neurosci Lett. 2012;531(2):142–4.
Kim HJ, Kim HY, Ki CS, Kim SH. Presenilin 1 gene mutation (M139I) in a patient with an early-onset Alzheimer’s disease: clinical characteristics and genetic identification. Neurol Sci. 2010;31(6):781–3.
Kim J, Bagyinszky E, Chang YH, Choe G, Choi BO, An SS, Kim S. A novel PSEN1 H163P mutation in a patient with early-onset Alzheimer’s disease: clinical, neuroimaging, and neuropathological findings. Neurosci Lett. 2012;530(2):109–14.
Klunemann HH, Rogaeva E, Neumann M, Kretzschmar HA, Kandel M, Toulina A, et al. Novel PS1 mutation in a Bavarian kindred with familial Alzheimer disease. Alzheimer Dis Assoc Disord. 2004;18(4):256–8.
Knight WD, Kennedy J, Mead S, Rossor MN, Beck J, Collinge J, Mummery C. A novel presenilin 1 deletion (p.L166del) associated with early onset familial Alzheimer’s disease. Eur J Neurol. 2007;14(7):829–31.
Kowalska A, Wender M, Florczak J, Pruchnik-Wolinska D, Modestowicz R, Szczech J, et al. Molecular genetics of Alzheimer’s disease: presenilin 1 gene analysis in a cohort of patients from the Poznan region. J Appl Genet. 2003;44(2):231–4.
Kowalska A, Pruchnik-Wolinska D, Florczak J, Modestowicz R, Szczech J, Kozubski W, et al. Genetic study of familial cases of Alzheimer’s disease. Acta Biochim Pol. 2004;51(1):245–52.
Kowalska A, Pruchnik-Wolinska D, Florczak J, Szczech J, Kozubski W, Rossa G, Wender M. Presenilin 1 mutations in Polish families with early-onset Alzheimer’s disease. Folia Neuropathol. 2004;42(1):9–14.
Kwok JB, Taddei K, Hallupp M, Fisher C, Brooks WS, Broe GA, et al. Two novel (M233T and R278T) presenilin-1 mutations in early-onset Alzheimer’s disease pedigrees and preliminary evidence for association of presenilin-1 mutations with a novel phenotype. NeuroReport. 1997;8(6):1537–42.
Kwok JB, Halliday GM, Brooks WS, Dolios G, Laudon H, Murayama O, et al. Presenilin-1 mutation L271V results in altered exon 8 splicing and Alzheimer’s disease with non-cored plaques and no neuritic dystrophy. J Biol Chem. 2003;278(9):6748–54.
La Bella V, Liguori M, Cittadella R, Settipani N, Piccoli T, Manna I, Quattrone A, Piccoli F. A novel mutation (Thr116Ile) in the presenilin 1 gene in a patient with early-onset Alzheimer’s disease. Eur J Neurol. 2004;11(8):521–4.
Lee P, Medina L, Ringman JM. The Thr354Ile substitution in PSEN1: disease-causing mutation or polymorphism? Neurology. 2006;66(12):1955–6.
Lee JH, Kahn A, Cheng R, Reitz C, Vardarajan B, Lantigua R, Medrano M, et al. Disease-related mutations among Caribbean Hispanics with familial dementia. Mol Genet Genomic Med. 2014;2(5):430–7.
Lendon CL, Martinez A, Behrens IM, Kosik KS, Madrigal L, Norton J, et al. E280A PS-1 mutation causes Alzheimer’s disease but age of onset is not modified by ApoE alleles. Hum Mutat. 1997;10(3):186–95.
Lindquist S, Schwartz M, Batbayli M, Waldemar G, Nielsen J. Genetic testing in familial AD and FTD: Mutation and phenotype spectrum in a Danish cohort. Clin Genet. 2009;76(2):205–9.
Lladó A, Sánchez-Valle R, Rey MJ, Mercadal P, Almenar C, López-Villegas D, Fortea J, Molinuevo JL. New mutation in the PSEN1 (E120G) gene associated with early onset Alzheimer’s disease. Neurologia. 2010;25(1):13–6.
Lleo A, Blesa R, Queralt R, Ezquerra M, Molinuevo JL, Pena-Casanova J, Rojo A, Oliva R. Frequency of mutations in the presenilin and amyloid precursor protein genes in early-onset Alzheimer disease in Spain. Arch Neurol. 2002;59(11):1759–63.
Lohmann E, Guerreiro RJ, Erginel-Unaltuna N, Gurunlian N, Bilgic B, Gurvit H, et al. Identification of PSEN1 and PSEN2 gene mutations and variants in Turkish dementia patients. Neurobiol Aging. 2012;33(8):1850.e17–27.
Luedecke D, Becktepe JS, Lehmbeck JT, Finckh U, Yamamoto R, Jahn H, Boelmans K. A novel presenilin 1 mutation (Ala275Val) as cause of early-onset familial Alzheimer disease. Neurosci Lett. 2014;566:115–9.
Miklossy J, Taddei K, Suva D, Verdile G, Fonte J, Fisher C, et al. Two novel presenilin-1 mutations (Y256S and Q222H) are associated with early-onset Alzheimer’s disease. Neurobiol Aging. 2003;24(5):655–62.
Miravalle L, Murrell JR, Takao M, Glazier B, Piccardo P, Vidal R, Ghetti B. Genetic mutations associated with presenile dementia. Neurobiol Aging. 2002;23(1):S322.
Moehlmann T, Winkler E, Xia X, Edbauer D, Murrell J, Capell A, et al. Presenilin-1 mutations of leucine 166 equally affect the generation of the Notch and APP intracellular domains independent of their effect on Abeta 42 production. Proc Natl Acad Sci USA. 2002;99(12):8025–30.
Morelli L, Prat MI, Levy E, Mangone CA, Castano EM. Presenilin 1 Met146Leu variant due to an A → T transversion in an early-onset familial Alzheimer’s disease pedigree from Argentina. Clin Genet. 1998;53(6):469–73.
Norton JB, Cairns NJ, Chakraverty S, Wang J, Levitch D, Galvin JE, Goate A. Presenilin1 G217R mutation linked to Alzheimer disease with cotton wool plaques. Neurology. 2009;73(6):480–2.
Palmer MS, Beck JA, Campbell TA, Humphries CB, Roques PK, Fox NC, Harvey R, Rossor MN, Collinge J. Pathogenic presenilin 1 mutations (P436S & I143F) in early-onset Alzheimer’s disease in the UK. Hum Mutat. 1999;13(3):256.
Pantieri R, Pardini M, Cecconi M, Dagna-Bricarelli F, Vitali A, Piccini A, et al. A novel presenilin 1 L166H mutation in a pseudo-sporadic case of early-onset Alzheimer’s disease. Neurol Sci. 2005;26(5):349–50.
Poorkaj P, Sharma V, Anderson L, Nemens E, Alonso ME, Orr H, et al. Missense mutations in the chromosome 14 familial Alzheimer’s disease presenilin 1 gene. Hum Mutat. 1998;11(3):216–21.
Portet F, Dauvilliers Y, Campion D, Raux G, Hauw JJ, Lyon-Caen O, et al. Very early onset AD with a de novo mutation in the presenilin 1 gene (Met 233 Leu). Neurology. 2003;61(8):1136–7.
Queralt R, Ezquerra M, Castellvi M, Lleo A, Blesa R, Oliva R. Detection of the presenilin 1 gene mutation (M139T) in early-onset familial Alzheimer disease in Spain. Neurosci Lett. 2001;299(3):239–41.
Ramirez-Duenas MG, Rogaeva EA, Leal CA, Lin C, Ramirez-Casillas GA, Hernandez-Romo JA, et al. A novel Leu171Pro mutation in presenilin-1 gene in a Mexican family with early onset Alzheimer disease. Ann Genet. 1998;41(3):149–53.
Reznik-Wolf H, Treves TA, Davidson M, Aharon-Peretz J, St George Hyslop PH, Chapman J, et al. A novel mutation of presenilin 1 in familial Alzheimer’s disease in Israel detected by denaturing gradient gel electrophoresis. Hum Genet. 1996;98(6):700–2.
Ringman JM, Gylys KH, Medina LD, Fox M, Kepe V, Flores DL, et al. Biochemical, neuropathological, and neuroimaging characteristics of early-onset Alzheimer’s disease due to a novel PSEN1 mutation. Neurosci Lett. 2011;487(3):287–92.
Rogaev EI, Sherrington R, Rogaeva EA, Levesque G, Ikeda M, Liang Y, et al. Familial Alzheimer’s disease in kindreds with missense mutations in a gene on chromosome 1 related to the Alzheimer’s disease type 3 gene. Nature. 1995;376(6543):775–8.
Rogaeva EA, Fafel KC, Song YQ, Medeiros H, Sato C, Liang Y, et al. Screening for PS1 mutations in a referral-based series of AD cases: 21 novel mutations. Neurology. 2001;57(4):621–5.
Romero I, Jorgensen P, Bolwig G, Fraser PE, Rogaeva E, Mann D, et al. A presenilin-1 Thr116Asn substitution in a family with early-onset Alzheimer’s disease. NeuroReport. 1999;10(11):2255–60.
Rossor MN, Fox NC, Beck J, Campbell TC, Collinge J. Incomplete penetrance of familial Alzheimer’s disease in a pedigree with a novel presenilin-1 gene mutation. Lancet. 1996;347(9014):1560.
Sherrington R, Rogaev EI, Liang Y, Rogaeva EA, Levesque G, Ikeda M, et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer’s disease. Nature. 1995;375(6534):754–60.
Shrimpton AE, Schelper RL, Linke RP, Hardy J, Crook R, Dickson DW, et al. A presenilin 1 mutation (L420R) in a family with early onset Alzheimer disease, seizures and cotton wool plaques, but not spastic paraparesis. Neuropathology. 2007;27(3):228–32.
Smith MJ, Gardner RJ, Knight MA, Forrest SM, Beyreuther K, Storey E, et al. Early-onset Alzheimer’s disease caused by a novel mutation at codon 219 of the presenilin-1 gene. NeuroReport. 1999;10(3):503–7.
Sorbi S, Tedde A, Nacmias B, Ciantelli M, Caffarra P, Ghidoni E, et al. Novel presenilin 1 and presenilin 2 mutations in early-onset Alzheimer’s disease families. Neurobiol Aging. 2002;23(1):S312.
Sugiyama N, Suzuki K, Matsumura T, Kawanishi C, Onishi H, Yamada Y, et al. A novel missense mutation (G209R) in exon 8 of the presenilin 1 gene in a Japanese family with presenile familial Alzheimer’s disease. Hum Mutat. 1999;14(1):90.
Tanahashi H, Kawakatsu S, Kaneko M, Yamanaka H, Takahashi K, Tabira T. Sequence analysis of presenilin-1 gene mutation in Japanese Alzheimer’s disease patients. Neurosci Lett. 1996;218(2):139–41.
Tedde A, Bartoli A, Piaceri I, Ferrara S, Bagnoli S, Serio A, et al. Novel presenilin 1 mutation (Ile408Thr) in an Italian family with late-onset Alzheimer’s disease. Neurosci Lett. 2016;610:150–3.
Terreni L, Valeria C, Calella AM, Gavazzi A, Alberoni M, Grimaldi LM, et al. A novel missense mutation (L219F) in exon 8 of the presenilin 1 gene in an Italian family with presenile familial Alzheimer’s disease. Neurobiol Aging. 2000;21(1):176–7.
Tiedt HO, Lueschow A, Winter P, Müller U. Previously not recognized deletion in presenilin-1 (p.Leu174del.) in a patient with early-onset familial Alzheimer’s disease. Neurosci Lett. 2013;544:115–8.
Ting SK, Benzinger T, Kepe V, Fagan A, Coppola G, Porter V, et al. A novel PSEN1 mutation (I238M) associated with early-onset Alzheimer’s disease in an African-American woman. J Alzheimers Dis. 2014;40(2):271–5.
Wasco W, Pettingell WP, Jondro PD, Schmidt SD, Gurubhagavatula S, Rodes L, et al. Familial Alzheimer’s chromosome 14 mutations. Nat Med. 1995;1(9):848.
Wisniewski T, Dowjat WK, Buxbaum JD, Khorkova O, Efthimiopoulos S, Kulczycki J, et al. A novel Polish presenilin-1 mutation (P117L) is associated with familial Alzheimer’s disease and leads to death as early as the age of 28 years. NeuroReport. 1998;9(2):217–21.
Yagi R, Miyamoto R, Morino H, Izumi Y, Kuramochi M, Kurashige T, et al. Detecting gene mutations in Japanese Alzheimer’s patients by semiconductor sequencing. Neurobiol Aging. 2014;35(7):1780.e1–5.
Yasuda M, Maeda K, Ikejiri Y, Kawamata T, Kuroda S, Tanaka C. A novel missense mutation in the presenilin-1 gene in a familial Alzheimer’s disease pedigree with abundant amyloid angiopathy. Neurosci Lett. 1997;232(1):29–32.
Yasuda M, Maeda K, Hashimoto M, Yamashita H, Ikejiri Y, Bird TD, et al. A pedigree with a novel presenilin 1 mutation at a residue that is not conserved in presenilin 2. Arch Neurol. 1999;56(1):65–9.
Yasuda M, Maeda S, Kawamata T, Tamaoka A, Yamamoto Y, Kuroda S, et al. Novel presenilin-1 mutation with widespread cortical amyloid deposition but limited cerebral amyloid angiopathy. J Neurol Neurosurg Psychiatry. 2000;68(2):220–3.
Zekanowski C, Golan MP, Krzysko KA, Lipczynska-Lojkowska W, Filipek S, Kowalska A, et al. Two novel presenilin 1 gene mutations connected with frontotemporal dementia-like clinical phenotype: genetic and bioinformatic assessment. Exp Neurol. 2006;200(1):82–8.
De Jonghe C, Cruts M, Rogaeva EA, Tysoe C, Singleton A, Vanderstichele H, et al. Aberrant splicing in the presenilin-1 intron 4 mutation causes presenile Alzheimer’s disease by increased Abeta42 secretion. Hum Mol Genet. 1999;8(3):1539–40.
Guo J, Wei J, Liao S, Wang L, Jiang H, Tang B. A novel presenilin 1 mutation (Ser169del) in a Chinese family with early-onset Alzheimer’s disease. Neurosci Lett. 2010;468(1):34–7.
Tysoe C, Whittaker J, Xuereb J, Cairns NJ, Cruts M, Van Broeckhoven C, et al. A presenilin-1 truncating mutation is present in two cases with autopsy-confirmed early-onset Alzheimer disease. Am J Hum Genet. 1998;62(1):70–6.
Bernardi L, Tomaino C, Anfossi M, Gallo M, Geracitano S, Puccio G, et al. Late onset familial Alzheimer’s disease: novel presenilin 2 mutation and PS1 E318G polymorphism. J Neurol. 2008;255(4):604–6.
Beyer K, Lao JI, Fernandández-Novoa L, Cacabelos R. Identification of a novel mutation (V148I) in the TM2 domain of the presenilin 2 gene in a patient with late-onset Alzheimer disease. Neurob Aging. 1998;19(4):S87.
Ertekin-Taner N, Younkin LH, Yager DM, Parfitt F, Baker MC, Asthana S, et al. Plasma amyloid beta protein is elevated in late-onset Alzheimer disease families. Neurology. 2008;70(8):596–606.
Ezquerra M, Lleo A, Castellvi M, Queralt R, Santacruz P, Pastor P, et al. A novel mutation in the PSEN2 gene (T430 M) associated with variable expression in a family with early-onset Alzheimer disease. Arch Neurol. 2003;60(8):1149–51.
Gallo M, Tomaino C, Bernardi L, Maletta R, Anfossi M, Geracitano S, et al. PS1 polymporphism and a novel PS2 mutation in a patient with late-onset familial Alzheimer’s disease. Alzheimers Dement. 2008;4(4):Suppl:T585.
Guerreiro RJ, Beck J, Gibbs JR, Santana I, Rossor MN, Schott JM, et al. Genetic variability in CLU and its association with Alzheimer’s disease. PLoS ONE. 2010;5(3):e9510.
Lao JI, Beyer K, Fernandez-Novoa L, Cacabelos R. A novel mutation in the predicted TM2 domain of the presenilin 2 gene in a Spanish patient with late-onset Alzheimer’s disease. Neurogenetics. 1998;1(4):293–6.
Li D, Parks SB, Kushner JD, Nauman D, Burgess D, Ludwigsen S, et al. Mutations of presenilin genes in dilated cardiomyopathy and heart failure. Am J Hum Genet. 2006;79(6):1030–9.
Lindquist SG, Hasholt L, Bahl JM, Heegaard NH, Andersen BB, Nørremølle A, et al. A novel presenilin 2 mutation (V393 M) in early-onset dementia with profound language impairment. Eur J Neurol. 2008;15(10):1135–9.
Lleo A, Blesa R, Gendre J, Castellvi M, Pastor P, Queralt R, Oliva R. A novel presenilin 2 gene mutation (D439A) in a patient with early-onset Alzheimer’s disease. Neurology. 2001;57(10):1926–8.
Müller U, Winter P, Bolender C, Nolte D. Previously unrecognized missense mutation E126 K of PSEN2 segregates with early onset Alzheimer’s disease in a family. J Alzheimers Dis. 2014;42(1):109–13.
Niu F, Yu S, Zhang Z, Yi X, Ye L, Tang W, Qiu C, Wen H, Sun Y, Gao J, Guo Y. Novel mutation in the PSEN2 gene (N141Y) associated with early-onset autosomal dominant Alzheimer’s disease in a Chinese Han family. Neurobiol Aging. 2014;35(10):2420.e1–5.
Piscopo P, Talarico G, Crestini A, Gasparini M, Malvezzi-Campeggi L, Piacentini E, et al. A novel mutation in the predicted TMIII domain of the PSEN2 gene in an Italian pedigree with atypical Alzheimer’s disease. J Alzheimers Dis. 2010;20(1):43–7.
Piscopo P, Talarico G, Spadoni O, Malvezzi-Campeggi L, Crestini A, Gasparini M, et al. A novel Italian presenilin 2 mutation (S175Y). Alzheimers Dement. 2008;4(4):Suppl 2:T595.
Sleegers K, Roks G, Theuns J, Aulchenko YS, Rademakers R, Cruts M, et al. Familial clustering and genetic risk for dementia in a genetically isolated Dutch population. Brain. 2004;127(Pt7):1641–9.
Tomaino C, Bernardi L, Anfossi M, Costanzo A, Ferrise F, Gallo M, et al. Presenilin 2 Ser130Leu mutation in a case of late-onset “sporadic” Alzheimer’s disease. J Neurol. 2007;254(3):391–3.
Youn YC, Bagyinszky E, Kim H, Choi BO, An SS, Kim S. Probable novel PSEN2 Val214Leu mutation in Alzheimer’s disease supported by structural prediction. BMC Neurol. 2014;14:105.
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, Roses AD, Haines JL, Pericak-Vance MA. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science. 1993;13;261(5123):921–3.
Poirier J, Davignon J, Bouthillier D, Kogan S, Bertrand P, Gauthier S. Apolipoprotein E polymorphism and Alzheimer’s disease. Lancet. 1993;342(8873):697–9.
Rebeck GW, Reiter JS, Strickland DK, Hyman BT. Apolipoprotein E in sporadic Alzheimer’s disease: allelic variation and receptor interactions. Neuron. 1993;11(4):575–80.
Roses AD. Apolipoprotein E alleles as risk factors in Alzheimer’s disease. Annu Rev Med. 1996;47:387–400.
Saunders AM, Strittmatter WJ, Schmechel D, George-Hyslop PH, Pericak-Vance MA, Joo SH, et al. Association of apolipoprotein E allele epsilon 4 with late-onset familial and sporadic Alzheimer’s disease. Neurology. 1993;43(8):1467–72.
Saunders AM. Apolipoprotein E and Alzheimer disease: an update on genetic and functional analyses. J Neuropathol Exp Neurol. 2000;59(9):751–8.
Strittmatter WJ, Saunders AM, Schmechel D, Pericak-Vance M, Enghild J, Salvesen GS, Roses AD. Apolipoprotein E: high-avidity binding to β-amyloid and increased frequency of type 4 allele in late-onset familial Alzheimer disease. Proc Natl Acad Sci USA. 1993;90(5):1977–81.
Cuenco KT, Lunetta KL, Baldwin CT, McKee AC, Guo J, Cupples LA, et al. Association of distinct variants in SORL1 with cerebrovascular and neurodegenerative changes related to Alzheimer disease. Arch Neurol. 2008;65(12):1640–8.
Holton P, Ryten M, Nalls M, Trabzuni D, Weale ME, Hernandez D, et al. Initial assessment of the pathogenic mechanisms of the recently identified Alzheimer risk Loci. Ann Hum Genet. 2013;77(2):85–105.
Tycko B, Feng L, Nguyen L, Francis A, Hays A, Chung WY, et al. Polymorphisms in the human apolipoproteinJ/clusterin gene: ethnic variation and distribution in Alzheimer’s disease. Hum Genet. 1996;98(4):430–6.
Kok EH, Luoto T, Haikonen S, Goebeler S, Haapasalo H, Karhunen PJ. CLU, CR255 and PICALM genes associate with Alzheimer’s-related senile plaques. Alzheimers Res Ther. 2011;3(2):12.
Bertram L, Lange C, Mullin K, Parkinson M, Hsiao M, Hogan MF, et al. Genome-wide association analysis reveals putative Alzheimer’s disease susceptibility loci in addition to APOE. Am J Hum Genet. 2008;83(5):623–32.
Malik M, Simpson JF, Parikh I, Wilfred BR, Fardo DW, Nelson PT, Estus S. CD33 Alzheimer’s risk-altering polymorphism, CD33 expression, and exon 2 splicing. J Neurosci. 2013;33(33):13320–5.
Lacher SE, Alazizi A, Wang X, Bell DA, Pique-Regi R, Luca F, Slattery M. A hypermorphic antioxidant response element is associated with increased MS4A6A expression and Alzheimer’s disease. Redox Biol. 2018;14:686–93.
Cruchaga C, Kauwe JS, Harari O, Jin SC, Cai Y, Karch CM, et al. GWAS of cerebrospinal fluid tau levels identifies risk variants for Alzheimer’s disease. Neuron. 2013;78(2):256–68.
Jin SC, Benitez BA, Karch CM, Cooper B, Skorupa T, Carrell D, et al. Coding variants in TREM2 increase risk for Alzheimer’s disease. Hum Mol Genet. 2014;23(21):5838–46.
Jonsson T, Stefansson H, Steinberg S, Jonsdottir I, Jonsson PV, Snaedal J, et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N Engl J Med. 2013;368(2):107–16.
Schjeide BM, Schnack C, Lambert JC, Lill CM, Kirchheiner J, Tumani H, et al. The role of clusterin, complement receptor 1, and phosphatidylinositol binding clathrin assembly protein in Alzheimer disease risk and cerebrospinal fluid biomarker levels. Arch Gen Psychiatry. 2011;68(2):207–13.
Morgen K, Ramirez A, Frölich L, Tost H, Plichta MM, Kölsch H, et al. Genetic interaction of PICALM and APOE is associated with brain atrophy and cognitive impairment in Alzheimer’s disease. Alzheimers Dement. 2014;10(5 Suppl):S269–76.
Chen H, Wu G, Jiang Y, Feng R, Liao M, Zhang L, et al. Analyzing 54,936 samples supports the association between CD2AP rs9349407 polymorphism and Alzheimer’s disease susceptibility. Mol Neurobiol. 2015;52(1):1–7.
Wang HF, Tan L, Hao XK, Jiang T, Tan MS, Liu Y, et al. Effect of EPHA1 genetic variation on cerebrospinal fluid and neuroimaging biomarkers in healthy, mild cognitive impairment and Alzheimer’s disease cohorts. J Alzheimers Dis. 2015;44(1):115–23.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Nikolac Perkovic, M., Pivac, N. (2019). Genetic Markers of Alzheimer’s Disease. In: Kim, YK. (eds) Frontiers in Psychiatry. Advances in Experimental Medicine and Biology, vol 1192. Springer, Singapore. https://doi.org/10.1007/978-981-32-9721-0_3
Download citation
DOI: https://doi.org/10.1007/978-981-32-9721-0_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-32-9720-3
Online ISBN: 978-981-32-9721-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)