Abstract
Environmental stresses such as waterlogging, drought, salinity, high and low temperature, and elevated CO2 levels affect plant development and pose an increasing warning to sustainable agriculture. Environmental stresses have become a matter of contention due to concerns about the outcomes of climate change on plant resources, genetic diversity, and world food safety. Plant responses to these stress factors are highly intricate and include modifications at the cellular, molecular, and genetic levels. Now, it has been scientifically proven that plant responds differently to multiple stresses as compared to individual stresses. As the changing climate will expose the plants to interactive effects simultaneously, therefore there is a dire need for further research in plant developmental responses to these stress factors; otherwise, this will have a negative effect on sustainable agriculture. Therefore, in this chapter, recent advancement in mechanisms and perspectives of plant adaptation and tolerance to several environmental stresses have been discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Bioinformatics
- Climate change
- Environmental stresses
- Plant physiology
- Phytohormones
- Stress tolerance
- Transcription factors
5.1 Plant Responses to Environmental Stress Factors: An Overview
Plant responses to various environmental stresses are extremely variable and highly complicated, which may include changes at the physiological, cellular, and transcriptional level. Recent studies indicate that plant responds differently to multiple stresses as they respond to individual stress, by the activation of specific gene expressions, which are needed in particular environmental stress. Plants have developed to grow in their environment where plants are frequently subjected to various environmental conditions, as they are sessile and have well-emerged mechanisms that allowed them to recognize minute environmental changes to avoid damage and to conserve energy for growth and development. Plants can activate specific responses to confer stress, so it became difficult to study multiple responses at a time by available techniques. This phenomenon is quite accurate for both biotic and abiotic stresses. There is a dire need for vast research in plant studies in order to evaluate the multiple stresses and how plants can tolerate these stresses (Atkinson and Urwin 2012; Zhu 2016). In this regard, conventional and modern breeding platforms can help us to select plants for the improvement of dsirable traits; such as 1) development of environmental stress resistance plants, 2) to increase sustainable yield production, and 3) to enhance the content of micronutrients to develop nutritious crops.
5.1.1 Multiple Environmental Stresses: A Challenge in Agriculture
Essentially, plants require water, carbon, and vital mineral nutrients for growth and normal development (Cramer et al. 2011). Most of the time, plants do not grow in standard conditions, which limit their genetic potential to achieve maximum height and reproductive stage (Bray et al. 2000). This fact is observed by comparing the maximum crop yield and the average crop yield. This difference in yield is due to suboptimal environmental conditions, which induce damaging the physiological changes which occur within the plants, called stresses (Simontacchi et al. 2015; Shao et al. 2008; Zhu 2016).
Abiotic stresses such as cold, high temperature, drought, salinity, and nutrient imbalance are those environmental stresses which limit the plant growth and production below the threshold level. In addition to this, plants must have to defend themselves against various pathogens and pests, which may include fungi, bacteria, nematodes, and herbivore pests. Each plant activates a specific response mechanism to a particular stress to avoid damage (Hammond-Kosack 2000). Plant responses can be elastic (reversible) and plastic (irreversible). The duration of stress can also vary which directly affects the plant response. The plant reaction toward stress factors depends on the specific tissue or organ, which is affected by the corresponding stress factor. For instance, the transcriptional responses are different and depend on stress, which is either tissue- or cell-specific in roots. Water stress, for example, affects the plant cell wall enzymatically and nonenzymatically which may inhibit plant growth (Cramer et al. 2011). Under field conditions, drought and temperature are the major principal stresses and have a remarkable effect on different crops (Raza et al. 2019a).
5.2 How Can Climate Change Affect Plant Developmental Responses?
Climate change is one of the main reasons which widely affects plant growth and reproduction. This changing climatic conditions and increasing population led to the demand of stress-tolerant plant species to meet the environmental conditions and energy demand (Takeda and Matsuoka 2008; Raza et al. 2019a; Newton et al. 2011).
As plants are sessile, they have to tolerate fluctuating environmental conditions in order to survive. The climatic catastrophe, which may be in the form of drought, salinity, elevated temperature, and elevated CO2 , represents a concerned issue toward the sustainable development of agriculture. Naturally, plants are blessed with the ability to sense climate change and adapt accordingly. With changing the environment, plants evolved and have developed precise molecular and cellular mechanisms that enable them to survive in harsh conditions. However, unfortunately, there is not enough research on how plants evolved currently at present, and this knowledge gap should be minimized to develop the plant species, which can tolerate not only individual stresses but also multiple stresses (Ahuja et al. 2010). Recent researches in “omics” approach led to better understanding of transcriptome, proteome, and metabolomics of plants by linking with stress perceptions and responses. These approaches are also studied in other crop and woody plant species besides a model plant Arabidopsis (Coolen et al. 2016; Varoquaux et al. 2019; Razzaq et al. 2019; Zhang et al. 2019; You et al. 2019).
5.2.1 Elevated CO2 Levels Affect Plant Development and Morphology
The CO2 levels in the atmosphere have elevated to 440 ppm from 280 ppm due to increased industrial activity (Meehl et al. 2007). Increased levels of CO2 directly affect plant developmental processes and photosynthetic exchange of gas. Plants are also affected indirectly by CO2 as it is the cause of the greenhouse effect that causes global warming and other drastic changes in climate. Studies in C3 plants have found that the elevation in CO2 level contributes to increased aboveground biomass. In grassland species, the aboveground biomass has been found to increase by 33%, but it also depends on nutrient and water availability in soil. In dry and hot areas, there is low aboveground biomass due to low water and nitrogen availability (Reich et al. 2014). Studies also indicated that root biomass also increased due to increased CO2 levels (Madhu and Hatfield 2013). The increased production of shoot biomass led to an increase in seed yield, as is observed in most plant species such as soybean, peanut, wheat, bean, and rice (Hatfield et al. 2011). Soybean, the model plant, has shown increased leaf length and the number of leaf nodes, increased seed yield and pod number, and elongated root (Bishop et al. 2015; Gray et al. 2016).
5.2.2 Effects of Elevated Temperature
The global average temperature has been increased to 0.85 from 1880 to 2012, due to increasing concentrations of CO2 and other greenhouse gases (Hartmann et al. 2013). By the end of the century, there is a prediction of increasing global mean temperature from 1.0 to 3.7 °C. This increasing temperature makes the plants undergo heat stress due to changing frequency, intensity, and duration of heat waves. The effects of elevation in CO2 are quite consistent across many plant species, while the elevation in temperature causes the plants to behave differently in different regions across the globe. In the Arctic, for instance, the surface temperature increases much faster as compared to other regions (Gray et al. 2016). The elevation in temperature affects plant development based on their growing regions. The yield of some plants increased, while the yield of the same plants decreased when grown in different regions. For example, soybean had a maximum yield in the Midwestern USA as compared to low yield in the Southern USA (Hatfield et al. 2011). The plant physiology and development are also affected by temperature, which in turn affects the yield of the plant. This reduction in plant yield is also due to decreasing the activity of carbon assimilation enzyme Rubisco either by a reduction in activity of Rubisco actives or by reduction in the generation of ribulose-1,5 biphosphate (RuBP). The C3 and C4 plants have different photosynthetic functions and behave accordingly in their optimal range of temperatures (Sage et al. 2008). This mechanism is quite clear that plant development depends on species-specific temperature optimum.
5.2.3 Effects of Drought Stress on Plant Physiology
Climate change is one of the major reasons for water stress. This increasing water stress is affecting plant growth in different regions of the world. It is predicted that the drought will increase in those areas which are already under water stress. Drought can cause damaging hydrologic imbalances due to prolonged dry weather affecting plant development and morphology (Gray et al. 2016). Drought stress can be observed in different plant organs and tissues. This stress often leads to elongation of root and reduction in shoot growth. This means that root elongates at the expense of shoot. For instance, the two grass species (H. lanatus and A. pratensis) showed increased nutrient and metabolite content in roots as compared to shoots (Gargallo-Garriga et al. 2014). This property proves the functional equilibrium theory in which plants will try to optimize the limiting resource by shifting allocation among tissues. This adaptive behavior will help the plants grow in water-deficit areas by the reduction in transpiration rates as plants invest more in roots than in shoots.
5.2.4 Multiple Environmental Stress Effects on Plant Development
There is limited research on how plants behave when a number of stresses are inflicted upon them at a time. Researchers have studied each stress independently but still challenging to comprehend multiple stresses. Climate change is one of the main reasons for the increasing number of stresses, and thus plants have to behave accordingly. In soybean, for instance, seed yield is increased by the elevation of CO2 which diminishes due to drought stress (Ruiz-Vera et al. 2018; Gray et al. 2016). In maize, high CO2 and elevated temperature do not contribute to an increase in seed yield (Ruiz-Vera et al. 2015, 2018). These effects demonstrated that the influence of one climate change factor is affected by the presence of another climate change factor on plant development and physiology. In order to comprehend the interactive effects of changing climate, there is a need to study how plants reacted to climate change at the molecular and cellular level.
5.3 How Transgenic Plants Respond to Different Environmental Stresses?
The transgenic approach has helped us to understand the ways of stress tolerance. Transgenic plants have developed to study the abiotic stress factors in greenhouses, growth rooms, and under a controlled environment. These studies have helped to understand the behavior of the number of transgenic plants in stress conditions like drought, salinity, and low and high temperature (Ashraf and Foolad 2007; Wang et al. 2016a; Gilliham et al. 2017). Some examples of transgenic plants showing resistance to different environmental stresses are described in Table 5.1.
There is a comparable difference which exists between conventional and transgenic approaches to tolerate water stress. The genes of significant metabolic and defensive pathways can be engineered through transgenic approaches such as osmoprotectant producing pathways and antioxidant defense approaches (Singh et al. 2015; Wang et al. 2016a). Microarray techniques are proving helpful in several stress-inducible genes, but their function at the molecular level is still unknown. Recently, the study was carried out to enhance the photosynthetic properties of different C3 crops either by the introgression of C4 genes or by the overexpression of C3 genes (Ashraf and Harris 2013).
Among other abiotic stresses, high salinity is exceptionally damaging for plants. In transgenic plants, compartmentation and diversion of harmful ions such as Na+ and Cl− from delicate areas of plants like mesophyll to apoplast or vacuole took place to avoid the deleterious effects of high salinity (Sperling et al. 2014). Transgenic plants, mostly halophytes, have increased salinity tolerance due to overexpression of the ion transporter gene (Flowers and Colmer 2008). The physiological and molecular responses are determinant of plant survival under low temperatures (John et al. 2016; Sergeant et al. 2014). Transgenic plants at transcriptional as well as translational levels show cold tolerance mechanisms. Biotechnological and molecular approaches are used to alter gene expression in such a way that it increases the concentration of several metabolites to avoid cold stress (John et al. 2016). The expression of altered genes, which have been evaluated most recently, is said to be controlled by core binding factor (CBF) or dehydration responsive element binding (DREB) binding factors (Agarwal et al. 2017).
High temperature, another abiotic factor, has to be tolerated by plants in this fluctuating environmental temperature (Mittler et al. 2012). High temperature directly affects the proteome, transcriptome, metabolome, and liposome which makes it challenging for the plants to survive. Molecular chaperones such as heat shock protein (HSP) are highly valuable in avoiding the harmful effect of high temperature (Xu et al. 2013). Five major HSPs play a central role in avoiding stress. These HSPs may also combine with other co-chaperones to repair damaged proteins by refolding, thereby protecting plant cellular functions. Any reduction in the HSPs is responsible for the abnormal development of plants (Kotak et al. 2007).
Pest-resistant crops are highly useful in limiting the biotic stress caused by pests and fungi. The development of these bio-friendly plants helps to improve plant tolerance to a certain fungicide or a pesticide (Mahmood et al. 2014). In the literature, it is well reported that resistance to pests can be brought about by enhancing the complex multigene enzymes activities such as glutathione-S-transferase, esterases, and cytochrome P450s (Bass and Field 2011).
Another factor, nutrient use efficiency (NUE), becomes reduced due to leaching, surface runoff, volatilization, or microbial consumption. Efficient means are required to maximize the nutrient use efficiency to diminish mineral losses (Salim and Raza 2020; Baligar and Fageria 2015). There are not enough molecular approaches used to date to improve the NUE of plants. However, despite the limiting knowledge, it has been acknowledged that transcription factors and associated kinases are improving the NUE of plants (Salim and Raza 2020; Canales et al. 2014).
Heavy metal toxicity leads to necrosis and stunted growth of plants, and it is one of the major determinants to assess agricultural productivity (Liu et al. 2014b). It not only affects the plant at the cellular level but also at the molecular level by impairing transcription and replication mechanisms (User 2013). Phytoremediation is a highly useful technique in eliminating environmental contaminants (Glick 2003). Moreover, rhizoremediation, including plants and also their rhizospheric microbes which naturally either exist or are introduced in plants, can help to eliminate or reduce contamination levels and encourage normal plant development (Rainbird et al. 2018).
5.4 Role of Transcription Factors (TFs) in Stress Management
Plants have developed a number of defense processes to tolerate abiotic stresses. It is becoming essential to assess these mechanisms to produce new varieties of genetically stress-tolerant plants. Due to the progress in novel genetic fields such as genomics, transcriptomics, and proteomics, now it is not only easy to dissect the whole defense mechanism but also have the ability to enhance the resistance in plants (Liu et al. 2014a). Molecular markers contribute to the utilization of candidate genes in the genetic engineering of crops (Raza et al. 2019b). As it becomes easy to study the stress signaling pathway, therefore, the generic signaling pathway consists of key steps such as signal perception, signal transduction, stress-responsive gene expression, concerning physiological processes, and metabolic reactions (Zhu 2016; Pérez-Clemente et al. 2013). Firstly, plants via sensors/receptors present in the cell wall or membrane receive the extracellular stimuli. The second messenger, which includes inositol phosphate, sugar, reactive oxygen species (ROS), calcium ions (Ca2+), cyclic nucleotides (cAMP and cGMP), and nitric oxide (NO), converts the extracellular signals into intracellular signals. Thereupon, these secondary messengers activate the required signaling pathway to transduce the signal (Newton et al. 2016; Bhargava and Sawant 2013). The phosphorylation and dephosphorylation of proteins are carried out with the help of protein kinase and phosphatases, respectively, in most of the signaling pathways and quite effective in signal relay mechanism (Jaiwal and Singh 2003). For example, in plant abiotic stress, the mitogen-activated protein kinases (MAPKs ) pathways and calcium-dependent protein kinases (CDPKs) pathways are known to play an important role (Simeunovic et al. 2016; Huang et al. 2012; Rayapuram et al. 2018). The protein kinases or phosphatases activate transcription factors at the end of phosphorylation chain and further bind TFs precisely to cis-elements in the promoter region of stress-related genes and modulate their transcription process (Simeunovic et al. 2016). At the same time, further upstream components regulate TFs to their transcript level and undergo a number of alterations at the posttranscriptional level, like ubiquitination and simulation. After posttranscriptional modification, it forms a multiplex regulatory network to inflect the response of stress-related genes that regulate physiological activities and metabolic reactions (Simeunovic et al. 2016; Zhu 2016; Mizoi et al. 2013). A model for plant abiotic stress responses is described in Fig. 5.1.
All these steps contribute to form a genetic pathway for abiotic stress-related signal transduction in plants. Various TFs have been identified and characterized through a considerable research, which are elaborated in abiotic stress responses in plants either in ABA-dependent pathways or in ABA-independent pathways (Simeunovic et al. 2016; Umezawa et al. 2006; Zhu 2016; Golldack et al. 2011). Several studies were carried out to engineer these TFs to overcome plant stress tolerance, and some worth noting results have been reported (Table 5.2).
5.5 Regulation of Environmental Responses in Plants by Oxidative Stress
The growth of plants depends on the exposing environment. Due to the continuous fluctuating environment, these plants are open to a number of stress factors and other extreme meteorological conditions such as drought and floods, heat and frost waves, and others, which are mainly encouraged by climate change (Raza et al. 2019a; Walter et al. 2013). The dose-response connection is a property of the stress factor. These stress factors inflict the stress on plants from a specific intensity level of stress; according to classical stress theory, low doses of stress does not cause much damage to plants, but instead, they can stimulate metabolism and growth—a phenomenon called eustress (Poschenrieder et al. 2013).
Oxidative stress, which is mainly a disproportion between oxidizing (ROS) and reducing (antioxidants) factors, is a common property of all environmental stresses (Potters et al. 2010). ROS are produced by aerobic plant metabolism as a by-product. These ROS are either excited or reduced forms of oxygen. Various forms of ROS, for example, superoxide (O˙−2), hydrogen peroxide (H2O2), and hydroxyl radical (HO˙), are obtained when atmospheric oxygen undergoes univalent reduction (Sgherri et al. 2018; Gill and Tuteja 2010).
Almost all environmental stresses are responsible for enhancing the rate of ROS. The most common processes observed are disruption of photosynthesis and reduction in stomatal conductance results in the making of ROS. Increase in photorespiration is due to the reduction in stomatal conductance which causes increases in 70% of H2O2. During a water shortage, excessive photorespiration causes stomata to close to reduce transpiration results in CO2 deficiency which leads to limited photosynthesis (Sgherri et al. 2018; Dat et al. 2000; Noctor et al. 2002; Nayyar and Gupta 2006).
Oxidative stress can be induced by some stress factors directly, such as transition metals, e.g., iron and copper, that activate the Haber–Weiss pathway and elevate the production of HO˙ (Bhattacharjee 2019; Ravet and Pilon 2013). In addition to metals, ozone can also induce ROS production directly which results in apoplast breakdown and formation of H2O2 and HO˙. Further, metabolic changes cause a secondary oxidative burst, which further increases the synthesis of ROS (Fiscus et al. 2005). In addition, these metabolic abnormalities induce oxidative stress indirectly mainly by UV-B radiation (Hideg et al. 2013). Nevertheless, many researchers believe that UV-B has the ability to convert H2O2 to HO˙, which directly leads to the damage of oxidative tissues (Czégény et al. 2014). Plants to avoid oxidative stress have evolved many approaches. Both enzymatic and non-enzymatic antioxidants are helpful to control ROS concentration, of which the two most important are superoxide dismutase (SOD) which catalyzes the production of H2O2 and catalase (CAT) which eliminates H2O2 in peroxisomes. The vital non-enzymatic antioxidants are tocopherols, ascorbate, glutathione (GSH), carotenoids, and flavonoids which have a significant part in ROS elimination and in repairing cell damage (Kacienė et al. 2015).
5.6 Phytohormones: Role in Plant Responses to Environmental Stresses
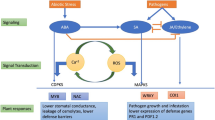
As it is evident, Ca2+- and ROS-mediated plant responses to environmental stresses do not contribute much as the stress response mechanism is highly complex. Phytohormones have the ability to mitigate stress response because of the highly intricate link among plant hormones and their capability to regulate physiological functions (Verma et al. 2016). A group of nine different plant hormones mediates the production of highly efficient response, and their signaling pathways are interconnected to help in the plant’s defense mechanism. The major phytohormones are auxins, gibberellin (GA), cytokinin (CK), abscisic acid (ABA), ethylene (ET) , salicylic acid (SA), jasmonate (JA), brassinosteroid (BRs), and strigolactones (STs) . For abiotic stresses and pathogens, ABA, SA, JA, and ethylene are recognized to have a significant role (Raza et al. 2019c; Nakashima and Yamaguchi-Shinozaki 2013). The levels of ABA are increased in abiotic stresses like salinity, drought, and low and high temperature (Raza et al. 2019c; Lata and Prasad 2011). Comparatively, other hormones such as SA, JA, and ethylene have a key role in biotic stresses as well, and their levels increase with the pathogen infection (Bari and Jones 2009). The possible role of hormones related to different stresses in plants is described in Fig. 5.2. However, the stress response process is not only limited to these hormones. Recent research in plant hormones has provided considerable evidence for the cross talk of ABA, SA, JA, and ET with auxins, GA, and CK in regulating plant defense responses (Bari and Jones 2009; Raza et al. 2019c; Nishiyama et al. 2013).
Hormonal cross-talk related to different environmental stresses in the plant with a relation to ABA. Abbreviations are explained in the text. Modified from Raza et al. (2019a)
5.7 Interaction of Plants with Other Organisms and Viruses in Stress Management
Microbes or microbial community plays a vital role to mitigate environmental constraints. The soil contains a pool of microbial population that comprises of bacteria, fungi, actinomycetes, protozoa, and algae. The maximum number of bacteria is found around the area of plant roots commonly called the rhizosphere, as compared to bulk soil. Bacteria can infect plants in three ways. Firstly, the bacteria may be harmful, beneficial, or neutral to plants. Secondly, plant growth-promoting bacteria (PGPB) includes both free-living and those that form symbiotic relationships with plants. Lastly, bacterial endophytes that colonize in plants interior and cyanobacteria make symbiotic relationship with fungi. The symbiotic relationship of PGPB with plants helps them to avoid environmental stresses and protects plants from dying. Over the past decade, bacteria belonging to the diverse genera have been stated to bestow host plants under different abiotic stress environments (Ho et al. 2017). Figure 5.3 shows the factors that affect the plant–microbe interaction during both abiotic and biotic stresses.
PGPB also help plants to combat biotic stresses in two different ways: directly by phytohormone production or by assisting the uptake of certain nutrients that promote plant growth or indirectly when PGPB minimizes or eliminates the harmful effects of pathogens. For example, the growth of phytopathogenic fungi is inhabited by 2,4-diacetyl phloroglucinol (DAPG) produced by P. Fluorescens (Voisard et al. 1994). The new approach to developing entomopathogenic bacteria has been in use to handle resistant pests. Several species are used as biological control of pests includes Aschersonia, Agerata, Verticillium, Sphaerostilbe, Podonectria, Myriangium, Hirsutella, Metarhizium, and Bacillus thuringiensis. Brevibacillus laterosporus is considered to be productive against insects like Coleoptera, Lepidoptera, nematodes, and phytopathogenic fungi (Boets et al. 2004; Saikia et al. 2011).
5.8 Usage of Bioinformatics Tools for Expression Analysis of Plant Transcriptome
Bioinformatics has been incorporated in different subjects of sciences, including plant breeding (Barh et al. 2013). Table 5.3 shows a list of bioinformatics tools for gene expression analysis based on statistical data analysis in plant breeding programs. The consistency and predictability of plant breeding programs, reduction in time, and the cost of stress-tolerant varieties are now possible with the help of OMICs tools (Van Emon 2015). These tools can enhance the nutritional content of food crops and maximize the agricultural production for food, feed, and energy (Davies 2010; Van Emon 2015). Transcriptomics, a subcategory of OMICs, appeals several scientists especially in plant breeding subjects (Shariatipour and Heidari 2017). It allows us to classify genes differentially present in diverse cell populations or in reaction to altered treatments. Microarray, a high-throughput technique, has the ability to measure gene expression and thereby produce functional data for numerous genes at once (Schulze and Downward 2001; Oktem et al. 2008). However, to overcome environmental distress and to improve plant varieties, it is crucial to investigate the role of bioinformatics.
5.9 Plant Evolution-Adaptive and Neutral Processes
The K-Pg dividing line is commonly linked with its related extinction events, which comprise the last five important mass extinction events in the Phanerozoic Eon (Rohde and Muller 2005). Several environmental issues, such as the rise in volcanism and global warming, were responsible for this mass extinction which leads to unfavorable conditions for the existence of the number of living organisms (Robertson et al. 2013).
In order to study the present-day challenges, the production of polyploids is being observed, which are often exposed to changing and unstable environment. In the Arctic, for example, there is an excess of latterly establishing polyploids. The adaptive and neutral processes have the ability to describe these phenomena and the consequences thereof for plant evolution. Both these processes help in the enhancement of polyploidy establishment under changing environment and distress conditions. An unreduced gamete formation might be increased by the adaptive processes in addition to other processes such as hybridization and extinction of the background diploid population. This is likely to lead to other polyploids being established even in the unavailability of any active adaptive benefit. On the other hand, transgressive segregation and genomic vulnerability of polyploids may lead toward the heterotic phenotypes, enhanced phenotypic irregularity, and plasticity. If it is considered useful, then it might be quickly selected under the changing environmental conditions, which is likely to lead to more polyploids being established even in the lake of enhanced polyploid formation (Vanneste et al. 2014). It is imperative that the environment plays a crucial role in polyploid formation regardless of which process contributes more as described in Fig. 5.4.
5.10 Conclusion and Future Perspectives
Environmental stresses are responsible for reversible and irreversible changes in the plants. These changes can be biochemical, physiological, and genomic types, including at transcriptional and posttranscriptional levels. Climate change is the major reason for the environmental distress and is responsible for increasing drought and salinity and low and high temperature. These changes minimize the plant plasticity for development and agricultural productivity. However, it has been reported that the world population projected to reach 9.8 billion in 2050, and 11.2 billion in 2100 (United Nation 2017). To overcome the food demands in coming years, it is very important to generate climate-resiliet plants. Plant biotechnology played a vital role in the establishment of stress-resistant plant varieties. However, care should be taken in creating transgenic varieties to introduce genes that have the ability to tolerate interactive stress factors, precisely at the entire plant level. Currently, there is less understanding of plant responses to harsh climate conditions in several experiments related to molecular changes. A more holistic approach is needed which requires data from several biological kinds of research to study individual and multiple stress conditions. Nevertheless, the integration of OMICS approaches could help us to identify stress related genes and regulators, which ultimately leads towards the manipulation of candidate genes for the development of climate-resilient plants. Additionally, a powerful genome editing tool such as CRISPR/Cas system together with conventional and modern breeding technologies should be utilize to cope with environmental stresses and to secure the world food security for sustainable agriculture.
References
Agarwal PK, Gupta K, Lopato S, Agarwal P (2017) Dehydration responsive element binding transcription factors and their applications for the engineering of stress tolerance. J Exp Bot 68:2135–2148
Ahuja I, de Vos RC, Bones AM, Hall RD (2010) Plant molecular stress responses face climate change. Trends Plant Sci 15:664–674
Alshareef NO, Wang JY, Ali S, Al-Babili S, Tester M, Schmöckel SM (2019) Overexpression of the NAC transcription factor JUNGBRUNNEN1 (JUB1) increases salinity tolerance in tomato. Plant Physiol Biochem 140:113–121
Amara I, Capellades M, Ludevid MD, Pagès M, Goday A (2013) Enhanced water stress tolerance of transgenic maize plants over-expressing LEA Rab28 gene. J Plant Physiol 170:864–873
Ashraf M, Foolad M (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216
Ashraf M, Harris P (2013) Photosynthesis under stressful environments: an overview. Photosynthetica 51:163–190
Atkinson NJ, Urwin PE (2012) The interaction of plant biotic and abiotic stresses: from genes to the field. J Exp Bot 63:3523–3543
Babitha K, Ramu S, Pruthvi V, Mahesh P, Nataraja KN, Udayakumar M (2013) Co-expression of AtbHLH17 and AtWRKY28 confers resistance to abiotic stress in Arabidopsis. Transgenic Res 22:327–341
Baligar VC, Fageria NK (2015) Nutrient use efficiency in plants: an overview. In: Nutrient use efficiency: from basics to advances. Springer, New Delhi, pp 1–14
Barh D, Zambare V, Azevedo V (2013) Omics: applications in biomedical, agricultural, and environmental sciences. CRC Press, Boca Raton. https://doi.org/10.1201/b14289
Bari R, Jones JD (2009) Role of plant hormones in plant defence responses. Plant Mol Biol 69:473–488
Bass C, Field LM (2011) Gene amplification and insecticide resistance. Pest Manag Sci 67:886–890
Bhargava S, Sawant K (2013) Drought stress adaptation: metabolic adjustment and regulation of gene expression. Plant Breed 132:21–32
Bhattacharjee S (2019) ROS and oxidative stress: origin and implication. In: Reactive oxygen species in plant biology. Springer, New Delhi, pp 1–31
Bishop KA, Betzelberger AM, Long SP, Ainsworth EA (2015) Is there potential to adapt soybean (Glycine max M err.) to future [CO2]? An analysis of the yield response of 18 genotypes in free-air CO2 enrichment. Plant Cell Environ 38:1765–1774
Boets A, Arnaut G, Van Rie J, Damme N (2004) Toxins. U.S. Patent No. 6,706,860, U.S. Patent and Trademark Office, Washington, DC
Borgohain P, Saha B, Agrahari R, Chowardhara B, Sahoo S, van der Vyver C, Panda SK (2019) SlNAC2 overexpression in Arabidopsis results in enhanced abiotic stress tolerance with alteration in glutathione metabolism. Protoplasma 256:1065–1077
Bray E, Bailey-Serres J, Weretilnyk E (2000) Biochemistry and molecular biology of plants. Am Soc Plant Physiol, Rockville, pp 1158–1203
Butt HI et al (2017) GaMYB85, an R2R3 MYB gene, in transgenic Arabidopsis plays an important role in drought tolerance. BMC Plant Biol 17:142
Byun MY et al (2015) Constitutive expression of DaCBF7, an Antarctic vascular plant Deschampsia antarctica CBF homolog, resulted in improved cold tolerance in transgenic rice plants. Plant Sci 236:61–74
Canales J, Moyano TC, Villarroel E, Gutiérrez RA (2014) Systems analysis of transcriptome data provides new hypotheses about Arabidopsis root response to nitrate treatments. Front Plant Sci 5:22
Cao Z-H, Zhang S-Z, Wang R-K, Zhang R-F, Hao Y-J (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdoMYB121 gene confering abiotic stress tolerance in plants. PLoS One 8:e69955
Chen Y, Huang L, Yan H, Zhang X, Xu B, Ma X (2016) Cloning and characterization of an ABA-independent DREB transcription factor gene, HcDREB2, in Hemarthria compressa. Hereditas 153:3
Cheng Z, Targolli J, Huang X, Wu R (2002) Wheat LEA genes, PMA80 and PMA1959, enhance dehydration tolerance of transgenic rice (Oryza sativa L.). Mol Breed 10:71–82
Cheng L, Yang J, Yin L, Hui L, Qian H, Li S-Y, Li L-J (2017) Transcription factor NnDREB1 from lotus improved drought tolerance in transgenic Arabidopsis thaliana. Biol Plant 61:651–658
Chu X et al (2015) The cotton WRKY gene GhWRKY41 positively regulates salt and drought stress tolerance in transgenic Nicotiana benthamiana. PLoS One 10:e0143022
Coolen S, Proietti S, Hickman R, Davila Olivas NH, Huang PP, Van Verk MC, Van Pelt JA, Wittenberg AH, De Vos M, Prins M, Van Loon JJ (2016) Transcriptome dynamics of Arabidopsis during sequential biotic and abiotic stresses. Plant J 86:249–267
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163. https://doi.org/10.1186/1471-2229-11-163
Czégény G, Wu M, Dér A, Eriksson LA, Strid Å, Hideg É (2014) Hydrogen peroxide contributes to the ultraviolet-B (280–315 nm) induced oxidative stress of plant leaves through multiple pathways. FEBS Lett 588:2255–2261
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143:1739–1751
Dat J, Vandenabeele S, Vranová E, Van Montagu M, Inzé D, Van Breusegem F (2000) Dual action of the active oxygen species during plant stress responses. Cell Mol Life Sci 57:779–795
Davies H (2010) A role for “omics” technologies in food safety assessment. Food Control 21:1601–1610
Downey T (2006) Analysis of a multifactor microarray study using Partek genomics solution. Methods Enzymol 411:256–270
Duan J, Cai W (2012) OsLEA3-2, an abiotic stress induced gene of rice plays a key role in salt and drought tolerance. PLoS One 7:e45117
Emig D, Salomonis N, Baumbach J, Lengauer T, Conklin BR, Albrecht M (2010) AltAnalyze and DomainGraph: analyzing and visualizing exon expression data. Nucleic Acids Res 38:W755–W762
Fiscus EL, Booker FL, Burkey KO (2005) Crop responses to ozone: uptake, modes of action, carbon assimilation and partitioning. Plant Cell Environ 28:997–1011
Floratos A, Smith K, Ji Z, Watkinson J, Califano A (2010) geWorkbench: an open source platform for integrative genomics. Bioinformatics 26:1779–1780
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179:945–963
Gao F et al (2017) Overexpression of a tartary buckwheat R2R3-MYB transcription factor gene, FtMYB9, enhances tolerance to drought and salt stresses in transgenic Arabidopsis. J Plant Physiol 214:81–90
Gao H, Wang Y, Xu P, Zhang Z (2018) Overexpression of a WRKY transcription factor TaWRKY2 enhances drought stress tolerance in transgenic wheat. Front Plant Sci 9:997
Gargallo-Garriga A et al (2014) Opposite metabolic responses of shoots and roots to drought. Sci Rep 4:6829
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Gilliham M, Able JA, Roy SJ (2017) Translating knowledge about abiotic stress tolerance to breeding programmes. Plant J 90:898–917
Glick BR (2003) Phytoremediation: synergistic use of plants and bacteria to clean up the environment. Biotechnol Adv 21:383–393
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Gray SB et al (2016) Intensifying drought eliminates the expected benefits of elevated carbon dioxide for soybean. Nat Plants 2:16132
Gu L, Ma Q, Zhang C, Wang C, Wei H, Wang H, Yu S (2019) The cotton GhWRKY91 transcription factor mediates leaf senescence and responses to drought stress in transgenic Arabidopsis thaliana. Front Plant Sci 10:1352
Gunapati S et al (2016) Expression of GhNAC2 from G. herbaceum, improves root growth and imparts tolerance to drought in transgenic cotton and Arabidopsis. Sci Rep 6:24978
Guo H et al (2017a) Expression of the MYB transcription factor gene BplMYB46 affects abiotic stress tolerance and secondary cell wall deposition in Betula platyphylla. Plant Biotechnol J 15:107–121
Guo X, Zhang L, Zhu J, Wang A, Liu H (2017b) Christolea crassifolia HARDY gene enhances drought stress tolerance in transgenic tomato plants. Plant Cell Tissue Org Cult 129:469–481
Gupta SM, Pandey P, Negi P, Pande V, Grover A, Patade VY, Ahmed Z (2013) DRE-binding transcription factor gene (LlaDREB1b) is regulated by various abiotic stresses in Lepidium latifolium L. Mol Biol Rep 40:2573–2580
Habegger L, Sboner A, Gianoulis TA, Rozowsky J, Agarwal A, Snyder M, Gerstein M (2010) RSEQtools: a modular framework to analyze RNA-Seq data using compact, anonymized data summaries. Bioinformatics 27:281–283
Halbritter F, Vaidya HJ, Tomlinson SR (2012) GeneProf: analysis of high-throughput sequencing experiments. Nat Methods 9:7
Hammond-Kosack K (2000) Responses to plant pathogens. In: Biochemistry and molecular biology of plants. American Society of Plant Physiologists, Rockville, MD, pp 1102–1109
Hartmann DL, Klein Tank AMG, Rusticucci M, Alexander L, Brönnimann S, Charabi Y, Dentener F, Dlugokencky E, Easterling D, Kaplan A, Soden B, Thorne P, Wild M, Zhai PM (2013) Observations: atmosphere and surface supplementary material. In: Stocker TF, Qin D, Plattner G-K, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds) Climate change 2013: the physical science basis. Contribution of working group I to the fifth assessment report of the intergovernmental panel on climate change. www.climatechange2013.org; www.ipcc.ch
Hatfield JL, Boote KJ, Kimball BA, Ziska LH, Izaurralde RC, Ort D, Thomson AM, Wolfe D (2011) Climate impacts on agriculture: implications for crop production. Agron J 103:351–370
Hideg É, Jansen MA, Strid Å (2013) UV-B exposure, ROS, and stress: inseparable companions or loosely linked associates? Trends Plant Sci 18:107–115
Ho Y-N, Mathew DC, Huang C-C (2017) Plant-microbe ecology: interactions of plants and symbiotic microbial communities. In: Plant ecology-traditional approaches to recent trends. InTech, Rijeka, pp 93–119
Hossain MA, Cho J-I, Han M, Ahn C-H, Jeon J-S, An G, Park PB (2010) The ABRE-binding bZIP transcription factor OsABF2 is a positive regulator of abiotic stress and ABA signaling in rice. J Plant Physiol 167:1512–1520
Huang G-T et al (2012) Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep 39:969–987
Jaiwal PK, Singh RP (2003) Improvement strategies of leguminosae biotechnology. Springer Science & Business Media
Jiang M, Ye Z-H, Zhang H-J, Miao L-X (2019) Broccoli plants over-expressing an ERF transcription factor gene BoERF1 facilitates both salt stress and sclerotinia stem rot resistance. J Plant Growth Regul 38:1–13
Jisha V, Dampanaboina L, Vadassery J, Mithöfer A, Kappara S, Ramanan R (2015) Overexpression of an AP2/ERF type transcription factor OsEREBP1 confers biotic and abiotic stress tolerance in rice. PLoS One 10:e0127831
John R, Anjum N, Sopory S, Akram N, Ashraf M (2016) Some key physiological and molecular processes of cold acclimation. Biol Plant 60:603–618
Kacienė G, Žaltauskaitė J, Milčė E, Juknys R (2015) Role of oxidative stress on growth responses of spring barley exposed to different environmental stressors. J Plant Ecol 8:605–616
Kallio MA et al (2011) Chipster: user-friendly analysis software for microarray and other high-throughput data. BMC Genomics 12:507
Kang C, Zhai H, He S, Zhao N, Liu Q (2019) A novel sweetpotato bZIP transcription factor gene, IbbZIP1, is involved in salt and drought tolerance in transgenic Arabidopsis. Plant Cell Rep 38:1373–1382. https://doi.org/10.1007/s00299-019-02441-x
Khatri P, Bhavsar P, Bawa G, Draghici S (2004) Onto-Tools: an ensemble of web-accessible, ontology-based tools for the functional design and interpretation of high-throughput gene expression experiments. Nucleic Acids Res 32:W449–W456
Kim H-S, Lee JH, Kim JJ, Kim C-H, Jun S-S, Hong Y-N (2005) Molecular and functional characterization of CaLEA6, the gene for a hydrophobic LEA protein from Capsicum annuum. Gene 344:115–123
Kiranmai K et al (2018) A novel WRKY transcription factor, MuWRKY3 (Macrotyloma uniflorum lam. Verdc.) enhances drought stress tolerance in transgenic groundnut (Arachis hypogaea L.) plants. Front Plant Sci 9:346
Kolesnikov N et al (2014) ArrayExpress update—simplifying data submissions. Nucleic Acids Res 43:D1113–D1116
Kornobis E, Cabellos L, Aguilar F, Frías-López C, Rozas J, Marco J, Zardoya R (2015) TRUFA: a user-friendly web server for de novo RNA-seq analysis using cluster computing. Evol Bioinform Online 11:97–104. https://doi.org/10.4137/EBO.S23873
Kotak S, Larkindale J, Lee U, von Koskull-Döring P, Vierling E, Scharf K-D (2007) Complexity of the heat stress response in plants. Curr Opin Plant Biol 10:310–316
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748
Lemkin P, Thornwall G, Wagner U, Henninghausen L (1999) MAExplorer–microarray exploratory data analysis. Nat Genet 23:57–57
Li Y, Sun Y, Yang Q, Fang F, Kang J, Zhang T (2013) Isolation and characterization of a gene from Medicago sativa L., encoding a bZIP transcription factor. Mol Biol Rep 40:1227–1239
Li Y, Zhang H, Zhang Q, Liu Q, Zhai H, Zhao N, He S (2019) An AP2/ERF gene, IbRAP2-12, from sweetpotato is involved in salt and drought tolerance in transgenic Arabidopsis. Plant Sci 281:19–30
Liang Q-Y et al (2017) Chrysanthemum WRKY gene DgWRKY5 enhances tolerance to salt stress in transgenic chrysanthemum. Sci Rep 7:4799
Liao Y, Zhang JS, Chen SY, Zhang WK (2008) Role of soybean GmbZIP132 under abscisic acid and salt stresses. J Integr Plant Biol 50:221–230
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P (2015) The molecular signatures database hallmark gene set collection. Cell Syst 1:417–425
Liu C, Wu Y, Wang X (2012) bZIP transcription factor OsbZIP52/RISBZ5: a potential negative regulator of cold and drought stress response in rice. Planta 235:1157–1169
Liu C et al (2014a) OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol Biol 84:19–36
Liu J-G, Qu P, Zhang W, Dong Y, Li L, Wang M-X (2014b) Variations among rice cultivars in subcellular distribution of Cd: the relationship between translocation and grain accumulation. Environ Exp Bot 107:25–31
Liu X, Song Y, Xing F, Wang N, Wen F, Zhu C (2016) GhWRKY25, a group I WRKY gene from cotton, confers differential tolerance to abiotic and biotic stresses in transgenic Nicotiana benthamiana. Protoplasma 253:1265–1281
Liu H, Zhou Y, Li H, Wang T, Zhang J, Ouyang B, Ye Z (2018) Molecular and functional characterization of ShNAC1, an NAC transcription factor from Solanum habrochaites. Plant Sci 271:9–19
Liu J, Chu J, Ma C, Jiang Y, Ma Y, Xiong J, Cheng Z-M (2019) Overexpression of an ABA-dependent grapevine bZIP transcription factor, VvABF2, enhances osmotic stress in Arabidopsis. Plant Cell Rep 38:587–596
Ma L-F, Li Y, Chen Y, Li X-B (2016) Improved drought and salt tolerance of Arabidopsis thaliana by ectopic expression of a cotton (Gossypium hirsutum) CBF gene. Plant Cell Tissue Org Cult 124:583–598
Madhu M, Hatfield J (2013) Dynamics of plant root growth under increased atmospheric carbon dioxide. Agron J 105:657–669
Mahmood Q, Bilal M, Jan S (2014) Herbicides, pesticides, and plant tolerance: an overview. In: Emerging technologies and management of crop stress tolerance, vol 1. Elsevier, pp 423–448
Manor O, Segal E (2015) GenoExp: a web tool for predicting gene expression levels from single nucleotide polymorphisms. Bioinformatics 31:1848–1850
Mao X, Chen S, Li A, Zhai C, Jing R (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PLoS One 9:e84359
Mao H, Yu L, Han R, Li Z, Liu H (2016) ZmNAC55, a maize stress-responsive NAC transcription factor, confers drought resistance in transgenic Arabidopsis. Plant Physiol Biochem 105:55–66
Mare C, Mazzucotelli E, Crosatti C, Francia E, Cattivelli L (2004) Hv-WRKY38: a new transcription factor involved in cold-and drought-response in barley. Plant Mol Biol 55:399–416
Meehl GA, Stocker TF, Collins WD, Friedlingstein P, Gaye AT, Gregory JM, Kitoh A, Knutti R, Murphy JM, Noda A, Raper SCB, Watterson IG, Weaver AJ, Zhao Z-C (2007) Global climate projections. In: Solomon S, Qin D, Manning M, Chen Z, Marquis M, Averyt KB, Tignor M, Miller HL (eds) Climate change 2007: the physical science basis. Contribution of working group I to the fourth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge, pp 747–845
Meng C, Cai C, Zhang T, Guo W (2009) Characterization of six novel NAC genes and their responses to abiotic stresses in Gossypium hirsutum L. Plant Sci 176:352–359
Mittler R, Finka A, Goloubinoff P (2012) How do plants feel the heat? Trends Biochem Sci 37:118–125
Mizoi J, Ohori T, Moriwaki T, Kidokoro S, Todaka D, Maruyama K, Kusakabe K, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2013) GmDREB2A; 2, a canonical DEHYDRATION-RESPONSIVE ELEMENT-BINDING PROTEIN2-type transcription factor in soybean, is posttranslationally regulated and mediates dehydration-responsive element-dependent gene expression. Plant Physiol 161:346–361
Mzid R et al (2018) The grapevine VvWRKY2 gene enhances salt and osmotic stress tolerance in transgenic Nicotiana tabacum. 3 Biotech 8:277
Nakashima K, Yamaguchi-Shinozaki K (2013) ABA signaling in stress-response and seed development. Plant Cell Rep 32:959–970
Nayyar H, Gupta D (2006) Differential sensitivity of C3 and C4 plants to water deficit stress: association with oxidative stress and antioxidants. Environ Exp Bot 58:106–113
Newton AC, Johnson SN, Gregory PJ (2011) Implications of climate change for diseases, crop yields and food security. Euphytica 179:3–18
Newton AC, Bootman MD, Scott JD (2016) Second messengers. CSH Perspect Biol 8:a005926
Nie J et al (2018) The AP2/ERF transcription factor CmERF053 of chrysanthemum positively regulates shoot branching, lateral root, and drought tolerance. Plant Cell Rep 37:1049–1060
Nishiyama R et al (2013) Arabidopsis AHP2, AHP3, and AHP5 histidine phosphotransfer proteins function as redundant negative regulators of drought stress response. Proc Natl Acad Sci U S A 110:4840–4845
Noctor G, Veljovic-Jovanovic S, Driscoll S, Novitskaya L, Foyer CH (2002) Drought and oxidative load in the leaves of C3 plants: a predominant role for photorespiration? Ann Bot 89:841–850
Oktem HA, Eyidogan F, Selçuk F, Oz MT, Teixeira da Silva JA, Yucel M (2008) Revealing response of plants to biotic and abiotic stresses with microarray technology. Genes Genomes Genomics 2:14–48
Pandey AS, Sharma E, Jain N, Singh B, Burman N, Khurana JP (2018) A rice bZIP transcription factor, OsbZIP16, regulates abiotic stress tolerance when over-expressed in Arabidopsis. J Plant Biochem Biotechnol 27:393–400
Papatheodorou I et al (2017) Expression Atlas: gene and protein expression across multiple studies and organisms. Nucleic Acids Res 46:D246–D251
Park B-J, Liu Z, Kanno A, Kameya T (2005) Genetic improvement of Chinese cabbage for salt and drought tolerance by constitutive expression of a B. napus LEA gene. Plant Sci 169:553–558
Park H-Y et al (2011a) AtERF71/HRE2 transcription factor mediates osmotic stress response as well as hypoxia response in Arabidopsis. Biochem Biophys Res Commun 414:135–141
Park S-C, Kim Y-H, Jeong JC, Kim CY, Lee H-S, Bang J-W, Kwak S-S (2011b) Sweetpotato late embryogenesis abundant 14 (IbLEA14) gene influences lignification and increases osmotic- and salt stress-tolerance of transgenic calli. Planta 233:621–634
Pérez-Clemente RM, Vives V, Zandalinas SI, López-Climent MF, Muñoz V, Gómez-Cadenas A (2013) Biotechnological approaches to study plant responses to stress. BioMed Res Int 2013:654120
Poschenrieder C, Cabot C, Martos S, Gallego B, Barceló J (2013) Do toxic ions induce hormesis in plants? Plant Sci 212:15–25
Potters G, Horemans N, Jansen MA (2010) The cellular redox state in plant stress biology—a charging concept. Plant Physiol Biochem 48:292–300
Qi X-N, Xiao Y-Y, Fan Z-Q, Chen J-Y, Lu W-J, Kuang J-F (2016) A banana fruit transcriptional repressor MaERF10 interacts with MaJAZ3 to strengthen the repression of JA biosynthetic genes involved in MeJA-mediated cold tolerance. Postharvest Biol Technol 120:222–231
Qin Y, Tian Y, Liu X (2015) A wheat salinity-induced WRKY transcription factor TaWRKY93 confers multiple abiotic stress tolerance in Arabidopsis thaliana. Biochem Biophys Res Commun 464:428–433
Rainbird B, Bentham RH, Soole KL (2018) Rhizoremediation of residual sulfonylurea herbicides in agricultural soils using Lens culinaris and a commercial supplement. Int J Phytoremediation 20:104–113
Ramegowda V, Senthil-Kumar M, Nataraja KN, Reddy MK, Mysore KS, Udayakumar M (2012) Expression of a finger millet transcription factor, EcNAC1, in tobacco confers abiotic stress-tolerance. PLoS One 7:e40397
Ravet K, Pilon M (2013) Copper and iron homeostasis in plants: the challenges of oxidative stress. Antioxid Redox Signal 19:919–932
Rayapuram N, Bigeard J, Alhoraibi H, Bonhomme L, Hesse AM, Vinh J, Hirt H, Pflieger D (2018) Quantitative phosphoproteomic analysis reveals shared and specific targets of Arabidopsis mitogen-activated protein kinases (MAPKs) MPK3, MPK4, and MPK6. Mol Cell Proteomics 17:61–80
Raza A, Razzaq A, Mehmood SS, Zou X, Zhang X, Lv Y, Xu J (2019a) Impact of climate change on crops adaptation and strategies to tackle its outcome: a review. Plants 8:34
Raza A, Mehmood SS, Shah T, Zou X, Yan L, Zhang X, Khan RS (2019b) Applications of molecular markers to develop resistance against abiotic stresses in wheat. In: Wheat production in changing environments. Springer, Singapore, pp 393–420
Raza A, Mehmood SS, Tabassum J, Batool R (2019c) Targeting plant hormones to develop abiotic stress resistance in wheat. In: Wheat production in changing environments. Springer, Singapore, pp 557–577
Razzaq A, Sadia B, Raza A, Khalid Hameed M, Saleem F (2019) Metabolomics: a way forward for crop improvement. Meta 9:303
Reich PB, Hobbie SE, Lee TD (2014) Plant growth enhancement by elevated CO2 eliminated by joint water and nitrogen limitation. Nat Geosci 7:920
Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP (2006) Gene pattern 2.0. Nat Genet 38:500–501
Robertson DS, Lewis WM, Sheehan PM, Toon OB (2013) K-Pg extinction: reevaluation of the heat-fire hypothesis. J Geophys Res Biogeosci 118:329–336
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26
Rohde RA, Muller RA (2005) Cycles in fossil diversity. Nature 434:208–210
Ruiz-Vera UM, Siebers MH, Drag DW, Ort DR, Bernacchi CJ (2015) Canopy warming caused photosynthetic acclimation and reduced seed yield in maize grown at ambient and elevated [CO2]. Glob Chang Biol 21:4237–4249
Ruiz-Vera UM, Siebers MH, Jaiswal D, Ort DR, Bernacchi CJ (2018) Canopy warming accelerates development in soybean and maize, offsetting the delay in soybean reproductive development by elevated CO2 concentrations. Plant Cell Environ 41:2806–2820
Sage RF, Way DA, Kubien DS (2008) Rubisco, Rubisco activase, and global climate change. J Exp Bot 59:1581–1595
Saikia R, Gogoi D, Mazumder S, Yadav A, Sarma R, Bora T, Gogoi B (2011) Brevibacillus laterosporus strain BPM3, a potential biocontrol agent isolated from a natural hot water spring of Assam, India. Microbiol Res 166:216–225
Salim N, Raza A (2020) Nutrient use efficiency (NUE) for sustainable wheat production: a review. J Plant Nutr 43:297–315
Schulze A, Downward J (2001) Navigating gene expression using microarrays—a technology review. Nat Cell Biol 3:E190
Segal E, Yelensky R, Kaushal A, Pham T, Regev A, Koller D, Friedman N (2004) GeneXPress: a visualization and statistical analysis tool for gene expression and sequence data. In: Proceedings of the 11th international conference on intelligent systems for molecular biology (ISMB)
Segall RS, Zhang Q, Pierce R (2010) Data mining supercomputing with SAS JMP® genomics. In: Proceedings of 14th world multi-conference on systemics, cybernetics and informatics: WMSCI
Seo JS et al (2012) Expression of the Arabidopsis AtMYB44 gene confers drought/salt-stress tolerance in transgenic soybean. Mol Breed 29:601–608
Sergeant K, Kieffer P, Dommes J, Hausman J-F, Renaut J (2014) Proteomic changes in leaves of poplar exposed to both cadmium and low-temperature. Environ Exp Bot 106:112–123
Sgherri C, Pinzino C, Quartacci MF (2018) Reactive oxygen species and photosynthetic functioning: past and present. In: Reactive oxygen species in plants: boon or bane—revisiting the role of ROS. Wiley, Chichester, pp 137–155
Shao H-B, Chu L-Y, Jaleel CA, Zhao C-X (2008) Water-deficit stress-induced anatomical changes in higher plants. C R Biol 331:215–225
Shariatipour N, Heidari B (2017) Application of bioinformatics in plant breeding programmes. BAOJ Bioinform 1:1–10
Sharov AA, Schlessinger D, Ko MS (2015) ExAtlas: an interactive online tool for meta-analysis of gene expression data. J Bioinform Comput Biol 13:1550019
Shen J, Lv B, Luo L, He J, Mao C, Xi D, Ming F (2017) The NAC-type transcription factor OsNAC2 regulates ABA-dependent genes and abiotic stress tolerance in rice. Sci Rep 7:40641
Simeunovic A, Mair A, Wurzinger B, Teige M (2016) Know where your clients are: subcellular localization and targets of calcium-dependent protein kinases. J Exp Bot 67:3855–3872
Simontacchi M, Galatro A, Ramos-Artuso F, Santa-María GE (2015) Plant survival in a changing environment: the role of nitric oxide in plant responses to abiotic stress. Front Plant Sci 6:977
Singh M, Kumar J, Singh S, Singh VP, Prasad SM (2015) Roles of osmoprotectants in improving salinity and drought tolerance in plants: a review. Rev Environ Sci Biotechnol 14:407–426
Sperling O, Lazarovitch N, Schwartz A, Shapira O (2014) Effects of high salinity irrigation on growth, gas-exchange, and photoprotection in date palms (Phoenix dactylifera L., cv. Medjool). Environ Exp Bot 99:100–109
Tak H, Mhatre M (2013) Cloning and molecular characterization of a putative bZIP transcription factor VvbZIP23 from Vitis vinifera. Protoplasma 250:333–345
Takasaki H et al (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Gen Genomics 284:173–183
Takeda S, Matsuoka M (2008) Genetic approaches to crop improvement: responding to environmental and population changes. Nat Rev Genet 9:444–457
Tang Y et al (2017) JcDREB2, a physic nut AP2/ERF gene, alters plant growth and salinity stress responses in transgenic rice. Front Plant Sci 8:306
Tang Y et al (2019) Overexpression of a MYB family gene, OsMYB6, increases drought and salinity stress tolerance in transgenic rice. Front Plant Sci 10:168
Tárraga J et al (2008) GEPAS, a web-based tool for microarray data analysis and interpretation. Nucleic Acids Res 36:W308–W314
Tu M, Wang X, Huang L, Guo R, Zhang H, Cai J, Wang X (2016) Expression of a grape bZIP transcription factor, VqbZIP39, in transgenic Arabidopsis thaliana confers tolerance of multiple abiotic stresses. Plant Cell Tissue Org Cult 125:537–551
Ullah A, Sun H, Yang X, Zhang X (2018) A novel cotton WRKY gene, GhWRKY6-like, improves salt tolerance by activating the ABA signaling pathway and scavenging of reactive oxygen species. Physiol Plant 162:439–454
Umezawa T, Fujita M, Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 17:113–122
United Nation (2017) World population projected to reach 9.8 billion in 2050 and 11.2 billion in 2100. 01, 2017, [online] Available https://www.un.org/development/desa/en/news/population/world-population-prospects-2017.html
User I (2013) Biochemical studies on the effects of zinc and lead on oxidative stress, antioxidant enzymes and lipid peroxidation in okra (Hibiscus esculentus cv. Hassawi). Sci Int 1:29–38
Van Emon JM (2015) The omics revolution in agricultural research. J Agric Food Chem 64:36–44
Vanneste K, Maere S, Van de Peer Y (2014) Tangled up in two: a burst of genome duplications at the end of the Cretaceous and the consequences for plant evolution. Philos Trans R Soc Lond B Biol Sci 369:20130353
Varoquaux N, Cole B, Gao C, Pierroz G, Baker CR, Patel D, Madera M, Jeffers T, Hollingsworth J, Sievert J, Yoshinaga Y (2019) Transcriptomic analysis of field-droughted sorghum from seedling to maturity reveals biotic and metabolic responses. Proc Natl Acad Sci 116:27124–27132
Verma V, Ravindran P, Kumar PP (2016) Plant hormone-mediated regulation of stress responses. BMC Plant Biol 16:86
Voisard C et al (1994) Biocontrol of root diseases by Pseudomonas fluorescens CHA0: current concepts and experimental approaches. In: Molecular ecology of rhizosphere microorganisms: biotechnology and the release of GMOs. Wiley, pp 67–89
Walter J, Jentsch A, Beierkuhnlein C, Kreyling J (2013) Ecological stress memory and cross stress tolerance in plants in the face of climate extremes. Environ Exp Bot 94:3–8
Wan L et al (2014) Identification of ERF genes in peanuts and functional analysis of AhERF008 and AhERF019 in abiotic stress response. Funct Integr Genomics 14:467–477
Wang Y, Jiang J, Zhao X, Liu G, Yang C, Zhan L (2006) A novel LEA gene from Tamarix androssowii confers drought tolerance in transgenic tobacco. Plant Sci 171:655–662
Wang Q, Guan Y, Wu Y, Chen H, Chen F, Chu C (2008) Overexpression of a rice OsDREB1F gene increases salt, drought, and low temperature tolerance in both Arabidopsis and rice. Plant Mol Biol 67:589–602
Wang L, Li X, Chen S, Liu G (2009) Enhanced drought tolerance in transgenic Leymus chinensis plants with constitutively expressed wheat TaLEA 3. Biotechnol Lett 31:313–319
Wang X, Chen X, Liu Y, Gao H, Wang Z, Sun G (2011) CkDREB gene in Caragana korshinskii is involved in the regulation of stress response to multiple abiotic stresses as an AP2/EREBP transcription factor. Mol Biol Rep 38:2801–2811
Wang M et al (2014a) SiLEA14, a novel atypical LEA protein, confers abiotic stress resistance in foxtail millet. BMC Plant Biol 14:290
Wang RK, Cao ZH, Hao YJ (2014b) Overexpression of a R2R3 MYB gene MdSIMYB1 increases tolerance to multiple stresses in transgenic tobacco and apples. Physiol Plant 150:76–87
Wang Z et al (2014c) Characterization of two highly similar CBF/DREB1-like genes, PhCBF4a and PhCBF4b, in Populus hopeiensis. Plant Physiol Biochem 83:107–116
Wang X et al (2015) Expression of TaWRKY44, a wheat WRKY gene, in transgenic tobacco confers multiple abiotic stress tolerances. Front Plant Sci 6:615
Wang H, Wang H, Shao H, Tang X (2016a) Recent advances in utilizing transcription factors to improve plant abiotic stress tolerance by transgenic technology. Front Plant Sci 7:67
Wang Y, Shu Z, Wang W, Jiang X, Li D, Pan J, Li X (2016b) CsWRKY2, a novel WRKY gene from Camellia sinensis, is involved in cold and drought stress responses. Biol Plant 60:443–451
Wang G-D, Liu Q, Shang X-T, Chen C, Xu N, Guan J, Meng Q-W (2018) Overexpression of transcription factor SlNAC35 enhances the chilling tolerance of transgenic tomato. Biol Plant 62:479–488
Wei Q et al (2017) A wheat R2R3-type MYB transcription factor TaODORANT1 positively regulates drought and salt stress responses in transgenic tobacco plants. Front Plant Sci 8:1374
Wolfien M et al (2016) TRAPLINE: a standardized and automated pipeline for RNA sequencing data analysis, evaluation and annotation. BMC Bioinform 17:21
Wu X, Shiroto Y, Kishitani S, Ito Y, Toriyama K (2009) Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep 28:21–30
Wu M, Liu H, Han G, Cai R, Pan F, Xiang Y (2017) A moso bamboo WRKY gene PeWRKY83 confers salinity tolerance in transgenic Arabidopsis plants. Sci Rep 7:11721
Wu J, Jiang Y, Liang Y, Chen L, Chen W, Cheng B (2019) Expression of the maize MYB transcription factor ZmMYB3R enhances drought and salt stress tolerance in transgenic plants. Plant Physiol Biochem 137:179–188
Xia J, Gill EE, Hancock RE (2015) NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc 10:823–844
Xiao B, Huang Y, Tang N, Xiong L (2007) Over-expression of a LEA gene in rice improves drought resistance under the field conditions. Theor Appl Genet 115:35–46
Xu J, Xue C, Xue D, Zhao J, Gai J, Guo N, Xing H (2013) Overexpression of GmHsp90s, a heat shock protein 90 (Hsp90) gene family cloning from soybean, decrease damage of abiotic stresses in Arabidopsis thaliana. PLoS One 8:e69810
Yang A, Dai X, Zhang W-H (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Yang G, Yu L, Zhang K, Zhao Y, Guo Y, Gao C (2017) A ThDREB gene from Tamarix hispida improved the salt and drought tolerance of transgenic tobacco and T. hispida. Plant Physiol Biochem 113:187–197
Yang S et al (2019a) A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice. BMC Plant Biol 19:260
Yang X, Kim MY, Ha J, Lee S-H (2019b) Overexpression of the soybean NAC gene GmNAC109 increases lateral root formation and abiotic stress tolerance in transgenic Arabidopsis plants. Front Plant Sci 10:1036
Yao W, Wang L, Zhou B, Wang S, Li R, Jiang T (2016) Over-expression of poplar transcription factor ERF76 gene confers salt tolerance in transgenic tobacco. J Plant Physiol 198:23–31
Ying S, Zhang D-F, Fu J, Shi Y-S, Song Y-C, Wang T-Y, Li Y (2012) Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic Arabidopsis. Planta 235:253–266
Ying L, Chen H, Cai W (2014) BnNAC485 is involved in abiotic stress responses and flowering time in Brassica napus. Plant Physiol Biochem 79:77–87
You J, Zhang Y, Liu A, Li D, Wang X, Dossa K, Zhou R, Yu J, Zhang Y, Wang L, Zhang X (2019) Transcriptomic and metabolomic profiling of drought-tolerant and susceptible sesame genotypes in response to drought stress. BMC Plant Biol 19:267
Zhang X, Ju H-W, Chung M-S, Huang P, Ahn S-J, Kim CS (2010) The RR-type MYB-like transcription factor, AtMYBL, is involved in promoting leaf senescence and modulates an abiotic stress response in Arabidopsis. Plant Cell Physiol 52:138–148
Zhang L, Zhao G, Xia C, Jia J, Liu X, Kong X (2012) Overexpression of a wheat MYB transcription factor gene, TaMYB56-B, enhances tolerances to freezing and salt stresses in transgenic Arabidopsis. Gene 505:100–107
Zhang L, Zhang L, Xia C, Zhao G, Liu J, Jia J, Kong X (2015a) A novel wheat bZIP transcription factor, TabZIP60, confers multiple abiotic stress tolerances in transgenic Arabidopsis. Physiol Plant 153:538–554
Zhang X, Liu X, Wu L, Yu G, Wang X, Ma H (2015b) The SsDREB transcription factor from the succulent halophyte Suaeda salsa enhances abiotic stress tolerance in transgenic tobacco. Int J Genomics 2015:875497
Zhang L, Zhang L, Xia C, Zhao G, Jia J, Kong X (2016) The novel wheat transcription factor TaNAC47 enhances multiple abiotic stress tolerances in transgenic plants. Front Plant Sci 6:1174
Zhang L et al (2017) A novel wheat C-bZIP gene, TabZIP14-B, participates in salt and freezing tolerance in transgenic plants. Front Plant Sci 8:710
Zhang J, Luo H, Guo Z (2018) Overexpression of Malus hupehensis MhSHN1 gene enhances salt and osmotic stress tolerance in transgenic tobacco plants. Russ J Plant Physiol 65:857–864
Zhang Y, Li D, Zhou R, Wang X, Dossa K, Wang L, Zhang Y, Yu J, Gong H, Zhang X, You J (2019) Transcriptome and metabolome analyses of two contrasting sesame genotypes reveal the crucial biological pathways involved in rapid adaptive response to salt stress. BMC Plant Biol 19:66
Zhao B-Y, Hu Y-F, Li J-j, Yao X (2016a) BnaABF2, a bZIP transcription factor from rapeseed (Brassica napus L.), enhances drought and salt tolerance in transgenic Arabidopsis. Bot Stud 57:12
Zhao X et al (2016b) The Miscanthus NAC transcription factor MlNAC9 enhances abiotic stress tolerance in transgenic Arabidopsis. Gene 586:158–169
Zhao H, Zhao X, Li M, Jiang Y, Xu J, Jin J, Li K (2018) Ectopic expression of Limonium bicolor (Bag.) Kuntze DREB (LbDREB) results in enhanced salt stress tolerance of transgenic Populus ussuriensis Kom. Plant Cell Tissue Org Cult 132:123–136
Zhao Y et al (2019) Over-expression of an R2R3 MYB Gene, GhMYB73, increases tolerance to salt stress in transgenic Arabidopsis. Plant Sci 286:28–36
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zhu N et al (2015) The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice. Plant Sci 236:146–156
Zhu M, Meng X, Cai J, Li G, Dong T, Li Z (2018) Basic leucine zipper transcription factor SlbZIP1 mediates salt and drought stress tolerance in tomato. BMC Plant Biol 18:83
Zytnicki M, Quesneville H (2011) S-MART, a software toolbox to aid RNA-Seq data analysis. PLoS One 6:e25988
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Raza, A., Ashraf, F., Zou, X., Zhang, X., Tosif, H. (2020). Plant Adaptation and Tolerance to Environmental Stresses: Mechanisms and Perspectives. In: Hasanuzzaman, M. (eds) Plant Ecophysiology and Adaptation under Climate Change: Mechanisms and Perspectives I. Springer, Singapore. https://doi.org/10.1007/978-981-15-2156-0_5
Download citation
DOI: https://doi.org/10.1007/978-981-15-2156-0_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-2155-3
Online ISBN: 978-981-15-2156-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)