Abstract
An OsWRKY11 gene, which encodes a transcription factor with the WRKY domain, was identified as one of the genes that was induced by both heat shock and drought stresses in seedlings of rice (Oryza sativa L.). To determine if overexpression of OsWRKY11 confers heat and drought tolerance, OsWRKY11 cDNA was fused to the promoter of HSP101 of rice and introduced into a rice cultivar Sasanishiki. Overexpression of OsWRKY11 was induced by heat treatment. After heat pretreatment, the transgenic lines showed significant heat and drought tolerance, as indicated by the slower leaf-wilting and less-impaired survival rate of green parts of plants. They also showed significant desiccation tolerance, as indicated by the slower water loss in detached leaves. Our results indicate that the OsWRKY11 gene plays a role in heat and drought stress response and tolerance, and might be useful for improvement of stress tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Drought stress has been found to be one of the major causes of reduced crop yield (Ozturk et al. 2002), and great efforts have been made to breed drought-tolerant crop varieties. As the most important world food crop, cultivated rice (Oryza sativa L.) demands tremendous amounts of water during growth, which results in a number of production challenges. Rice is also a popular model plant for studies of monocots. Improvements in the tolerance of cereal plants to abiotic stress are important when the efficiency of food production is to be increased. Breeding of transgenic rice cultivar with drought tolerance can help increase and stabilize crop yield under stress environments.
Many transgenic approaches have been carried out to increase biotic and abiotic stress tolerance (see Bajaj and Mohanty 2005 for a review). One successful approach to increase abiotic stress tolerance is overexpression of certain stress-inducible transcription factors, such as the following: CBF1/DREB1B, CBF2/DREB1C, CBF3/DREB1A (Thomashow 1999); DREB1A (Kasuga et al. 1999); DREB2A (Sakuma et al. 2006); EmBP1, OSBZ8, TAF1 (Zhu 2002); SNAC1 (Hu et al. 2006). However, employment of a constitutive promoter such as CaMV35S to drive genes for transcription factors may present some risk of deleterious effect on plant growth (Romero et al. 1997; Kasuga et al. 1999). Overexpression of the OsDREB1A proteins has been reported to cause growth retardation under unstressed control conditions in transgenic rice plants (Ito et al. 2006). Similar phenomena concerning improvement of stress tolerance and growth retardation have been observed in transgenic Arabidopsis overexpressing CBF1/DREB1B, CBF2/DREB1C, or CBF3/DREB1A (Thomashow 1999; Kasuga et al. 1999) and in tobacco (Kasuga et al. 2004). It has been reported that the stress-inducible rd29A promoter instead of the constitutive promoter minimizes this negative effect on plant growth (Kasuga et al. 1999, 2004). It is, therefore, desirable to use stress-inducible promoters to produce stress tolerant plants.
To search for drought and heat-inducible transcription factors in rice, we previously focused on rice orthologs of Arabidopsis genes for transcription factors known to be induced by drought or heat (Seki et al. 2002; Oono et al. 2003; Rizhsky et al. 2004). Using hydroponically grown seedlings, OsWRKY11 (DDBJ accession no. AK108745; previous name was WRKY16 in accession no. AY341856) was identified to show enhanced expression by placing the plants at 38°C for 1 h (heat treatment), by leaving the plants out of water for 10 h (drought treatment) and by combined heat/drought treatment for 0.5 h (Shiroto et al. 2004).
The WRKY genes encode a large group of transcription factors. There are over 70 WRKY genes in Arabidopsis and over 100 members in rice (Wu et al. 2005; Zhang and Wang 2005; Liu et al. 2007; Ramamoorthy et al 2008). This family is defined by a domain of 60 amino acids containing the amino acid sequence WRKY at its amino-terminal end and a putative zinc finger motif at its carboxy-terminal end. It binds specifically to the DNA sequence motif (T)(T)TGAC(C/T), which is known as the W box. WRKY genes are known to participate in various developmental and physiological programs, including disease resistance (Chen and Chen 2000; Dellagi et al. 2000; Du and Chen 2000; Eulgem et al. 2000; Kim et al. 2000; Asai et al. 2002), senescence (Hinderhofer and Zentgraf 2001; Robatzek and Somssich 2001), biotic and abiotic stress responses (Yoda et al. 2002; Dong et al. 2003; Huang and Duman 2002; Hara et al. 2000; Rizhsky et al. 2002, 2004; Pnueli et al. 2002; Seki et al. 2002), and growth and developmental processes (Lagacé and Matton 2004; Johnson et al. 2002; Sun et al. 2003; Xu et al. 2004). To the best of our knowledge, however, no information is available regarding the relationship between overexpression of WRKY genes and heat and drought tolerance.
To determine if overexpression of OsWRKY11 confers heat and drought tolerance, OsWRKY11 cDNA was fused to the promoter of HSP101 of rice and introduced into rice. The promoter of HSP101 was employed in this study, because the HSP101 gene has been reported to be induced by heat shock (Agarwal et al. 2003). Although the promoter activity of HSP101 has not been reported, it was expected that the promoter was stress-inducible and could be used to minimize possible deleterious effects of OsWRKY11 expression under unstressed conditions. In this study, we first examined activation of HSP101 promoter by heat and drought stress, and the phenotype of transgenic plants overexpressing OsWRKY11. Then, we evaluated survival of green parts of plants after heat and drought treatment, and water loss of detached aerial parts during desiccation. Enhanced heat and drought tolerance was observed.
Materials and methods
Construction of HSP101 promoter::GUS
Genomic DNA was extracted from leaves of O. sativa L. cv. Nipponbare using DNeasy (Qiagen, Tokyo, Japan). An approximately 1 kb-fragment containing the promoter region of HSP101 (DDBJ accession no. AJ316025) was amplified from the DNA using Ex Taq (TaKaRa-Bio, Ohtsu, Japan), the PCR enhancer system (Invitrogen, Carlsbud, USA), and the following primers: OsHSP101p-F2 primer (5′-GGCCAACATGATACAAGAAACCG-3′) and OsHSP101p-R1 primer (5′-CGTTGGTCTTGTGCGTGAAGTTG-3′). Cycling conditions were as follows: 35 cycles at 94°C for 30 s, at 58°C for 30 s, and at 68°C for 30 s. The amplified fragments were gel-purified using the MinElute Gel Extraction Kit (Qiagen, USA). A Hind III site and a Nhe I site (italicized) were added using Ex Taq (Takara, Kyoto, Japan), the PCR enhancer system (Invitrogen, Carlsbud, USA), and OsHSP101p-F primer (5′-AAGCTTCCTTCCGGCGATCTTGCAG-3′) and OsHSP101p-R primer (5′-GCTAGCTCCTCCTCTCCTCACACAATC-3′). The amplified fragment was ligated to pGEM-T Easy Vector Systems (Promega, Madison, USA).
The HSP101 promoter-GUS was taken as a Hind III-Sac I fragment and inserted in the same sites of pBI101-UbiHyg, which contains a maize ubiquitin promoter (Christensen et al. 1992) and a hygromycin phosphotransferase gene as a selection marker.
Construction of HSP101 promoter::OsWRKY11
A full length cDNA of OsWRKY11 in pCMVFL3 (DDBJ accession no. AK108745) was obtained from the Rice Genome Resource Center at the National Institute of Agrobiological Sciences, Japan (accession no. AK108745 in the rice full-length cDNA database, KOME, http://cdna01.dna.affrc.go.jp/cDNA/). The HSP101 promoter was taken as a Eco RI fragment in pGEM-T Easy Vector and inserted into the same site located in front of the OsWRKY11 cDNA. The HSP101 promoter-OsWRKY11 contains two Spe I sites, one derived from pGEM-T easy vector and the other from the 3′ end of the cDNA. The Spe I fragment was inserted into the Xba I site of pBI101H, which contained CaMV35S promoter–hygromycin phosphotransferase gene as a selection marker (Yokoi et al. 1998).
Production of transgenic plants
The final construct, HSP101 promoter::OsWRKY11 and HSP101 promoter::GUS, was transferred into Agrobacterium tumefaciens strain EHA105 (Hood et al. 1993) and used for Agrobacterium-mediated transformation (Yokoi et al. 1997). Primary tranformants, To plants, were grown in pots (4.5 cm in diameter and 5.7 cm in depth) in a greenhouse at 28/23°C (12/12 h) under natural day length conditions, as described by Shirasawa et al. (2006).
Southern blot analysis
Genomic DNA (2 μg) was extracted from leaves using DNeasy (QIAGEN K.K., Japan) and digested with HindIII and electrophoresed on a 0.8% agarose gel. DNA fragments were blotted onto nylon membrane (GE Healthcare, UK) and hybridized with a 708-bp HPT probe labeled with digoxigenin (Roche, Switzerland) using primers HPT-F (5′-AATGAGTTGGACCAGCAGAAG-3′) and HPT-R (5′-CATTCAGGTCAAACATAGGCC-3′), or with a GUS probe using primers G787-F (5′-TGTGAATTCGATATCTACCCGCTTCGCGTC-3′) and G1809-R (5′-GATGAATTCTCATTGTTTGCCTCCCTGCTG-3′). After hybridization, blots were washed at 65°C in 0.1× SSC and 0.1% SDS.
GUS assay
Eight independent transgenic plants with HSP101 promoter::GUS were grown in soil in pots at 22/18°C (12/12 h) for 2 weeks and were subjected to heat shock, cold, NaCl, or desiccation treatment. For heat shock treatment, the excised fully expanded leaf blades (0.5–1 cm in length), leaf sheaths, and roots were placed in water in microcentrifuge tubes and incubated in a water bath at 37, 42, or 45°C for 1 h. The control was simultaneously placed at room temperature for 1 h. For cold treatment, the leaves in the microcentrifuge tubes were placed in water at 4°C for 1 h. Desiccation treatment was carried out by leaving the leaf segments in the tubes with neither water nor a lid at 25°C for 1 h. For NaCl treatment, the excised leaves were placed in 250 mM NaCl solution in the microcentrifuge tubes and incubated at 25°C for 1 h. Histochemical staining for GUS activity was performed using 5-bromo-4-chloro-3-indolyl-β-d-glucuronide (X-gluc) as a chromogenic substrate (Jefferson et al. 1987). Tissues were vacuum infiltrated with X-Gluc reaction buffer (100 mM sodium phosphate buffer pH 7.2, 0.5 mM potassium ferrocyanide, 0.5 mM potassium ferricyanide, 0.1% Triton X-100 and 1 mg/ml X-gluc) and incubated at 37°C overnight. After incubation, chlorophyll was removed from green tissues by immersion in 70% ethanol, and tissue samples were observed on a Nikon stereomicroscope.
RT-PCR
Two-week-old seedlings grown in soil at 22/18°C were exposed to heat pretreatment at 37/22°C (12/12 h) for 2 weeks with water everyday. For no-heat pretreatment, plants were grown at 22/18°C. Induction of OsWRKY11 expression was determined by RT-PCR. Total RNA was isolated from leaves using RNeasy (Qiagen, Tokyo, Japan) according to the manufacturer’s instructions. Poly A RNA was isolated using the Dynabeads mRNA Purification Kit (Invitrogen, Carlsbud, USA). cDNA synthesis was accomplished using SuperScript III Reverse Transcriptase (Invitrogen, Carlsbud, USA) and then used as a template for PCR amplification with the following primers: 5′-AAATCGTCACGCCGGTGCAAG-3′ and 5′-TCAACCTCGCTCTTGGTGAGGAA-3′ for OsWRKY11; 5′-TACAACGGTTGGCGTCGCAC-3′ and 5′-AACTTGCGCACACGGTCCAG-3′ for tubulin alpha-1 chain (accession no. AK069140 in DDBJ database). PCR was carried out with Ex Taq polymerase (TaKaRa-Bio, Ohtsu, Japan) for 30 cycles of denaturation for 0.5 min at 94°C, annealing for 0.5 min at 66°C, and extension for 0.5 min at 72°C, followed by final extension for 5 min at 72°C. For quantitative RT-PCR analysis, PCR was performed using SYBR Premix Ex Taq polymerase (TaKaRa-Bio) and Thermal Cycler Dice Real Time System TP800 (TaKaRa-Bio) for 55 cycles of denaturation for 5 s at 95°C, annealing for 10 s at 55°C, and extension for 30 s at 72°C.
Evaluation of heat and drought tolerance of seedlings
Evaluation of tolerance against heat and drought stress was performed by determining the extent of recovery of whole plants from exposure to 37/22°C (12/12 h) without a water supply for 2 days. Prior to combined heat/drought treatment, 2-week-old seedlings were grown for 2 weeks at 37/22°C with a water supply. Then the plants were exposed to 37/22°C for 2 days without a water supply. The plants were returned to 22/18°C and grown with a water supply for 2 weeks. Arial parts were separated into green living parts and nongreen dead parts. Each dry weight (DW) was determined after drying at 80°C for 24 h. The percentage of surviving green parts was calculated from the following formula: (DW of green parts/DW of green parts plus dead parts) × 100. Heat and drought treatment was also carried out at 37/22°C for 2.5 days without a water supply. Survival of plants was observed after 2 weeks of recovery.
Evaluation of water loss in detached aerial parts
For heat pretreatment, 2-week-old seedlings grown in soil at 22/18°C (12/12 h) were exposed to heat pretreatment at 37/22°C for 2 weeks with water everyday. For the control condition with no-heat pretreatment, plants were grown at 22/18°C. After the pretreatment for 2 weeks, the aerial parts of transgenic plants and wild type (WT) were detached and weighed immediately and then placed on a laboratory bench at 25°C and weighed at designated times. Detached aerial parts were then dried at 80°C for 24 h to determine final DW. Water content was standardized as a percentage relative to the initial water content of aerial parts of the plant; it was calculated as follows: [(FWi − DW)/(FW0 − DW)] × 100, where FWi and FW0 are fresh weight for any given interval and original fresh weight, respectively. Four T2 plants per line were used for each test. All tests were simultaneously repeated three times.
Results
Activation of HSP101 promoter in transgenic rice plants with HSP101 promoter::GUS
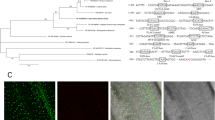
An approximately 1 kb-fragment containing the promoter region of HSP101 (DDBJ accession no. AJ316025) was amplified from rice genomic DNA and inserted in pBI101-UbiHyg, which contains a hygromycin phosphotransferase gene as a selection marker (Fig. 1). The GUS gene was fused to the promoter of HSP101 and introduced into an elite cultivar Sasanishiki. Excised leaf blades of the eight independent transgenic plants grown at 22/18°C (12/12 h) were subjected to histochemical GUS assay. Incubation of the leaf segments at 37, 42, or 45°C for 1 h induced a high level of GUS expression, while no GUS expression was detected at room temperature (Fig. 2a). Blue staining was stronger at 42 and 45°C. GUS expression was not observed in WT leaf blades. Strong GUS expression was also observed in leaf sheaths and roots, whereas weak blue staining was observed in the base of stems at room temperature. The GUS activity under other stresses, i.e., cold (4°C), 250 mM NaCl or drought for 1 h was not detected. Thus, the heat shock activation of the HSP101 promoter was confirmed.
The T-DNA region of HSP101 promoter::GUS and HSP101 promoter::OsWRKY11 used for transformation. HSP101 promoter was amplified from rice genome based on the sequence of DDBJ accession no. AJ316025. OsWRKY11 cDNA was a full length cDNA (accession no. AK108745). LB left border, NOS P nopaline synthase promoter, NPTII neomycin phosphotransferase, NOS T 3′ signal of nopaline synthase terminator, 35S PRO cauliflower mosaic virus 35S promoter, HPT hygromycin phosphotransferase
A transgenic line carrying a single copy of the GUS gene, which was revealed by Southern blot analysis (data not shown), was selected (line #g8). T2 plants homozygous for the introduced GUS gene were used for further analysis.
Production of transgenic rice plants carrying HSP101 promoter::OsWRKY11
OsWRKY11 cDNA (DDBJ accession no. AK108745) was fused to the promoter of HSP101 of rice (Fig. 1) and introduced into rice. Four independent transgenic lines (#ox2, #ox3, #ox4, and #ox5) were obtained. The number of copies of the T-DNA was determined by Southern blot analysis. We used Hind III, which cuts only once within the introduced T-DNA, and probed with the HPT gene (data not shown) so that the number of bands represented the copy number. The transgenic lines, #ox2 and #ox3, were identified to have a single copy of T-DNA, and #ox4 contained two copies and #ox5 carried four copies (Table 1).
T1 plants segregated for the introduced T-DNA were grown to maturity under normal growth conditions. Three types of phenotypes were observed: dwarf with bent leaves, normal plant length with bent leaves and normal plant length with normal leaves. In case of the bent leaves, the edges of leaf blades were bent toward the abaxial side, i.e. toward the opposite direction observed in the typical wilted leaf-rolling response. The number of plants with each phenotype is shown in Table 1. Most of the OsWRKY11 transgenic plants show the phenotype of normal plant length with bent leaves; plant length was more than 94 cm. In contrast, #ox4 showed dwarf phenotype; the plant length was less than 85 cm, and the WT plant had a plant length of 110–128 cm. Nontransgenic plants segregating from each primary transformant showed a phenotype of normal plant length with normal leaves. These results suggested that introduction of HSP101 promoter::OsWRKY11 might have caused some deleterious effects on plant growth.
Transgenic lines with a single copy of the T-DNA (i.e., #ox2 and #ox3) were selected, and T2 plants homozygous for the T-DNA were used for further study. A transgenic line, #g8, which contained HSP101 promoter::GUS, was used as a negative control. In 2-week-old seedlings of T2 plants, plant length was identical among #ox2, #ox3, #g8, and WT. However, the plant length of #ox3 was slightly shorter when compared to that of #ox2, #g8, and WT in 4-week-old seedlings grown at 22/18°C (n > 26, P < 0.0001). The dwarf phenotype and bent leaves were manifested when the plants were subjected to heat treatment at 37/22°C (12/12 h) for 2 weeks (Fig. 3). Plant length of #ox3 was significantly shorter than that of #ox2, #g8, and WT (n > 28, P < 0.0001). Under normal growth conditions, the seed set percentage was 36.7 ± 5.8 for #ox2 and 98.0 ± 0.8 for #ox3.
Induction of OsWRKY expression by heat treatment
Two-week-old seedlings grown in soil at 22/18°C (12/12 h) were exposed to heat pretreatment at 37/22°C (12/12 h) for 2 weeks with water everyday. Plants were maintained at 37°C during the day and at 22°C at night. For the control condition of no-heat pretreatment, plants were grown at 22/18°C. Induction of OsWRKY11 expression was investigated in leaf blades using RT-PCR. A 164-bp band of OsWRKY11 was detected in #ox2 and #ox3 after heat treatment, but was not detected in #g8 or WT (data not shown). In the case of no-heat pretreatment, a band of OsWRKY11 was hardly detected in these PCR conditions.
The relative OsWRKY11 expression level was also investigated using real-time quantitative RT-PCR (Fig. 4). The value in Fig. 4 indicates the relative expression level compared to that of tubulin. The relative OsWRKY11 expression level of #ox2 and #ox3 with no-heat treatment (control condition) was the same as that of WT. In contrast, the values were 3.7 times higher in #ox2 and 3.0 times higher in #ox3 than in WT after the heat pretreatment. These results demonstrated that overexpression of WRKY11 was induced by heat treatment, but it was not detectable under unstressed normal conditions.
Relative OsWRKY11 expression level compared to that of tubulin alpha-1 chain detected by real-time quantitative RT-PCR in plants with no-heat treatment (C) at 22/18°C (12/12 h) or plants with heat treatment (H) at 37/22°C (12/12 h) for 2 weeks. Values indicate relative expression level against that of WT-C in three biological replicates from cDNA prepared from leaf blades. Overexpression of OsWRKY11 was induced by heat treatment. The expression level after no-heat treatment was the same as that in WT. #ox2 and #ox3, T2 plants with HSP101 promoter::OsWRKY11; #g8, HSP101 promoter::GUS
Heat and drought tolerance of seedlings
Evaluation of tolerance against combined heat/drought stress was performed by determining the extent of recovery of whole plants from exposure to 37/22°C (12/12 h) without a water supply for 2 days. Prior to the combined heat/drought treatment, 2-week-old seedlings were grown for 2 weeks at 37/22°C with a water supply for the induction of OsWRKY11 expression. Figure 5a shows the photographs of #ox2, #ox3, and WT after the combined heat/drought treatment for 2 days without a water supply. The OsWRKY11 transgenic lines #ox2 and #ox3 show a weaker leaf-wilting phenotype.
Photographs of Os WRKY11 transgenic lines #ox2 and #ox3, and WT after combined heat/drought treatment at 37/22°C (12/12 h) without water supply. Two-week-old seedlings were grown for 2 weeks at 37/22°C with water supply, and then the plants were exposed to combined heat and drought treatment. a The photographs were taken after combined heat/drought treatment for 2 days. The Os WRKY11 transgenic lines #ox2 and #ox3 show weaker leaf-wilting phenotype. b The photographs were taken after recovery at 22/18°C for 2 weeks following heat/drought treatment for 2.5 days. The Os WRKY11 transgenic lines #ox2 and #ox3 survived, whereas WT plants died after recovery at 22/18°C for 2 weeks. Bar 5 cm
After the plants were exposed to 37/22°C for 2 days without water supply, they were returned to 22/18°C and grown with water supply for 2 weeks. Then, the percentage of surviving green parts was investigated (Fig. 6). The value (%) was 81.7 ± 1.9 for #ox2, 94.1 ± 1.6 for #ox3, while it was 39.8 ± 11 for #g8 and 46.6 ± 7.8 for WT. The difference was significant based on a t test (P < 0.001).
Survival of green parts in seedlings after recovery for 2 weeks from exposure to combined heat and drought treatment. Two-week-old seedlings were grown for 2 weeks at 37/22°C with water supply, and then the plants were exposed to combined heat and drought treatment at 37/22°C (12/12 h) for 2 days without water supply. After recovery for 2 weeks, areal parts were separated to green living parts and nongreen dead parts, and each DW was determined. Values (mean ± SD) indicate the percentage of surviving green parts calculated using the following formula: (DW of green parts/DW of green parts plus dead parts) × 100 (n = 4). The Os WRKY11 transgenic lines #ox2 and #ox3 conferred significant heat and drought tolerance compared to WT and #g8, plants with HSP101 promoter::GUS
Figure 5b shows the photographs of #ox2, #ox3, and WT at 2 weeks of recovery after combined heat/drought treatment at 37/22°C for 2.5 days without a water supply. Prolonged heat and drought treatment caused all the WT plants to die, while the OsWRKY11 transgenic lines #ox2 and #ox3 survived. These results demonstrated that the OsWRKY11 transgenic lines #ox2 and #ox3 gained significant heat and drought tolerance.
Evaluation of water loss in detached aerial parts
We determined whether water loss was affected in OsWRKY11 transgenic plants #ox2 and #ox3 by comparing the rates of change of the fresh weight of the detached aerial parts of plants during dehydration. For heat pretreatment, 2-week-old seedlings grown in soil at 22/18°C were exposed to heat pretreatment at 37/22°C for 2 weeks with water everyday. For the control condition with no-heat pretreatment, plants were grown at 22/18°C. After pretreatment for 2 weeks, the aerial parts of the transgenic plants and WT were detached and placed on a laboratory bench at 25°C and weighed at designated times.
The time course of water loss of plants with the heat pretreatment is shown in Fig. 7a and that of plants with the no-heat pretreatment is shown in Fig. 7b. In plants of both cases, water loss in #ox2 and #ox3 was slower than that in WT or #g8. In the case of plants with the heat pretreatment, the time (min) at which 20% of water loss was achieved was 141 ± 13 (mean ± SD) for #ox2 and 121 ± 31 for #ox3, whereas it was 51 ± 13 for WT and 56 ± 32 for #g8. The difference was significant based on a t test (P < 0.01). In the case of plants with the no-heat pretreatment, the time (min) at which 20% of water loss was achieved was 69 ± 8 for #ox2 and 68 ± 13 for #ox3, whereas it was 34 ± 12 for WT and 35 ± 4 for #g8. The difference was significant based on a t test (P < 0.01). When we compared the ratio between the time of 20% water loss of plants with the heat treatment and those with the no-heat pretreatment, the ratio was 1.5 for WT, 1.6 for #g8, 2.1 for #ox2, and 1.8 for #ox3, indicating that heat pretreatment enhanced drought tolerance especially in the #ox2 and #ox3 plants. It is considered that the heat pretreatment induced the expression of the introduced OsWRKY11 in addition to induction of endogenous OsWRKY11. Crosstolerance of heat and drought stresses was implicated even in WT plants. This experiment was repeated two more times and reproducibility was confirmed (data not shown).
The time courses of water loss in detached aerial parts. The water loss in #ox2 and #ox3 was slower than in WT and #g8. The water loss of plants with heat pretreatment (H) was slower than that with no-heat pretreatment (C). #ox2 and #ox3, T2 plants with HSP101 promoter::OsWRKY11; #g8, HSP101 promoter::GUS. Values (mean ± SD) indicate relative water content calculated using the following formula: [(FWi − DW)/(FW0 − DW)] × 100, where FWi and FW0 are fresh weight for any given interval and original fresh weight, respectively, and DW is the dry weight (n = 15 for #ox2, #ox3, and WT; n = 14 for #g8)
Discussion
Advantage of using of HSP101 promoter
Overexpression of transcription factor genes has sometimes been reported to accompany growth defects resulting in reduced productivity. This is presumably due to constitutive activation of defense-response genes under the regulation of introduced transcription factors. Such growth defects have been reported in transgenic rice plants overexpressing OsDREB1A (Ito et al. 2006) and WRKY45 (Shimono et al. 2007). The WRKY45 gene is one of the benzothiadiazole (BTH)- and salicylic acid (SA)-inducible WRKY genes. Growth retardation has been observed in transgenic rice plants with WRKY45 driven by maize ubiquitin promoter (Shimono et al. 2007). It has been reported that growth conditions have a profound influence on the expression of pathogen-related genes during concurrent WRKY45 expression, which may in turn affect the growth of WRKY45-overexpressing plants. This is quite similar to our results of the appearance of dwarf phenotype in the transgenic OsWRKY11 lines. Dehydration-responsive element-binding protein 1 (DREB1)/C-repeat-binding factors (CBFs) are known to specifically interact with the DRE/CRT cis-acting element and to control the expression of many stress-inducible genes in Arabidopsis. Overexpression of the OsDREB1A under the control of CaMV35S promoter or maize ubiquitin promoter has also been reported to cause growth retardation under unstressed control conditions in transgenic rice (Ito et al. 2006). Some transgenic lines showed a dwarf phenotype even at the reproductive stage. Kasuga et al. (1999, 2004) previously showed that the stress-inducible rd29A promoter instead of the constitutive promoter minimizes this negative effect on plant growth. Ito et al. (2006) demonstrated that the Arabidopsis rd29A gene promoter did not function in rice leaves; therefore, strong stress-inducible promoters of rice were necessary.
Our results indicated that the HSP101 promoter of rice is heat-inducible (Figs. 2, 4) and can be successfully used for the induction of OsWRKY11 expression to enhance heat and drought tolerance. Some leaky expression of OsWRKY11, although it was undetectable in leaf blades by RT-PCR, was likely to have occurred, because some plants, such as #ox3 and #ox4, showed a dwarf phenotype under unstressed normal conditions (Table 1). The HSP101 promoter might drive OsWRKY11 expression in the base of shoots, because weak GUS expression was observed in the base of stems without heat treatment (Fig. 2b). It is also possible that the CaMV 35S promoter in front of hygromycin phosphotransferase gene affected the HSP101 promoter activity. Leaky expressions of OsWRKY11 could also explain the fact that less impaired water loss of detached leaves was observed in plants without heat pretreatment (Fig. 7).
Involvement of OsWRKY11 in stress tolerance
Successful enhancement of drought tolerance by overexpression of transcription factors has been reported in rice. Ito et al. (2006) reported that overexpression of Arabidopsis DREB1 and rice ortholog OsDREB1 showed improved tolerance to drought, high-salt, and low-temperature stresses in transgenic rice. Hu et al. (2006) reported that overexpression of stress-responsive gene SNAC (STRESS-RESPONSIVE NAC 1) significantly enhances drought resistance in transgenic rice. Improvement of water use efficiency and drought tolerance in rice has been reported by expression of HARDY, an Arabidopsis drought and salt tolerance gene encoding an AP2/ERF-like transcription factor (Karaba et al. 2007). Xu et al. (2008) reported that overexpression of ZFP252, which encodes a TFIIIA-type zinc finger protein, in rice increased the amount of free proline and soluble sugar, elevated the expression of stress defense genes, and enhanced tolerance to salt and drought stresses.
WRKY proteins constitute a large family of transcription factors implicated in many different processes. WRKY genes play a variety of developmental and physiological roles in plants. The most reported studies for this superfamily of genes address their involvement in disease responses. Overexpression of AtWRKY18 in transgenic Arabidopsis plants results in enhanced expression of pathogenesis-related genes (Chen and Chen 2002). Enhancement of biotic stress by overexpressing WRKY genes has been reported. Shimono et al. (2007) reported that overexpression of BTH- and SA-inducible WRKY45 gene in rice markedly enhanced resistance to rice blast fungus. Overexpression of OsWRKY71 gene in rice resulted in enhanced resistance to virulent bacterial pathogens Xanthomonas oryzae pv. oryzae (Xoo) 13751 (Liu et al. 2007). However, employment of WRKY genes to enhance abiotic stresses has not yet been reported.
The relationship between WRKY and abiotic stress responses has been reported for some plant species. A barley gene coding for WRKY protein, Hv-WRKY38, has been reported to be involved in cold- and drought-response in barley (Mare et al. 2004). Involvement of an ABA-inducible WRKY gene in abiotic stresses has recently been reported in creosote bush (Larrea tridentate) (Zou et al. 2007). Qiu et al. (2004) reported that 10 of 13 OsWRKY genes (OsW8-9, 12–14, 16, 17, 21–24, 26, 30, and 45) were differentially regulated in plants treated by the four following abiotic stress factors: NaCl, PEG, cold (4°C), and heat (42°C). The resulting expression profiles exhibited significant differences in both the manner and timing of their response to the four different abiotic treatments. The difference of gene expression profiles suggested the different physiological functions among the WRKY genes.
Our results in the current study have demonstrated that the transgenic lines #ox2 and #ox3 showed significant desiccation tolerance, as indicated by the slower water loss in detached leaves (Fig. 7). After heat pretreatment, the transgenic lines #ox2 and #ox3 showed significant heat and drought tolerance, as indicated by the slower leaf-wilting and less-impaired survival rate of green parts (Figs. 5, 6). OsWRKY11 gene products may function in heat and drought stress response and tolerance. This is the first report of enhancement of abiotic stress by overexpression of WRKY genes.
Xie et al. (2005) undertook a comprehensive computational analysis of rice genomic sequences and predicted the structures of 81 OsWRKY genes, 48 of which are supported by full-length cDNA sequences. Recently, Ramamoorthy et al. (2008) have predicted 103 genes encoding WRKY transcription factors and reported comprehensive transcriptional profiles of the WRKY gene family in rice under various abiotic and phytohormone treatments. OsWRKY11 (LOC_Os01g43650) was shown to be relatively highly expressed in young and mature panicles and weakly expressed in young and mature leaves and roots under normal growth conditions. They reported that OsWRKY11 was not induced under abiotic (cold, drought, and salinity) or various hormone (ABA, IAA, gibberellic acid, methyl jasmonate, and salicylic acid) treatments. Ryu et al. (2006) carried out a systematic expression analysis of OsWRKY genes and identified that, among 45 tested genes, the expression of 15 genes, including OsWRKY11, was increased remarkably in an incompatible interaction between rice and rice blast fungus (Magnaporthe grisea). They also suggested that these genes are also involved in defense response to abiotic stress, such as humidity, touch stress induced by spraying, and altered diurnal rhythms due to prolonged darkness during the pathogen infection procedure. The induction of some OsWRKY expressions, including OsWRKY11, has been shown to be induced by wounding in clipped leaves. It is considered that OsWRKY11 is involved in biotic stress response to disease, as well in as heat and drought response. The transgenic plants #ox2 and #ox3 are expected to gain tolerance to such abiotic stresses.
The most similar protein to OsWRKY11 in Arabidopsis is AtWRKY28 encoded by AT4G18170 with a similarity of 88% in the WRKY domain and 48% in the whole proteins. OsWRKY11 contains a single WRKY domain with C2H2-type (C-X4–5-C-X22–24-H-X1–2-H) zinc finger motif and belongs to Group II of the WRKY family (Xie et al. 2005). OsWRKY11 contains the consensus coactivator motif, LXXLL, where L is Leu and X is any amino acid. OsWRKY11, therefore, is expected to bind specifically to the DNA sequence motif (T)(T)TGAC(C/T), which is known as the W box, in promoter regions and to activate certain gene expression. We are now investigating what kinds of genes are upregulated by OsWRKY11.
Conclusion
Drought and heat are major abiotic stresses on crop production associated with global warming. The HSP101 promoter was shown to be activated by heat but not by drought stress. Heat stress, however, often precedes drought stress. The HSP101 promoter, therefore, will be useful to confer heat and drought tolerance in field conditions, minimizing growth defects in unstressed conditions. Our data suggest that the OsWRKY11 gene, together with the HSP101 promoter, holds promising utility in improving drought and heat tolerance in rice.
References
Agarwal M, Sahi C, Katiyar-Agarwal S, Agarwal S, Young T, Gallie DR, Sharma VM, Ganesan K, Grover A (2003) Molecular characterization of rice hsp101: complementation of yeast hsp104 mutation by disaggregation of protein granules and differential expression in indica and japonica rice types. Plant Mol Biol 51:543–553
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL, Gomez-Gomez L, Boller T, Ausubel FM, Sheen J (2002) MAP kinase signaling cascade in Arabidopsis innate immunity. Nature 415:977–983
Bajaj S, Mohanty A (2005) Recent advances in rice biotechnology–towards genetically superior transgenic rice. Plant Biotech J 3:275–307
Chen C, Chen Z (2000) Isolation and characterization of two pathogen- and salicylic acid-induced genes encoding WRKY DNA-binding proteins from tobacco. Plant Mol Biol 42:387–396
Chen C, Chen Z (2002) Potentiation of developmentally regulated plant defense response by AtWRKY18, a pathogen-induced Arabidopsis transcription factor. Plant Physiol 129:706–716
Christensen AH, Sharrock RA, Quail PH (1992) Maize polyubiquitin genes: structure, thermal pertubation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol Biol 18:675–689
Dellagi A, Helibronn J, Avrova AO, Montesano M, Palva ET, Stewart HE, Toth IK, Cooke DE, Lyon GD, Birch PR (2000) A potato gene encoding a WRKY-like transcription factor is induced in interactions with Erwinia carotovora subsp. atroseptica and Phytophthora infestans and is coregulated with class I endochitinase expression. Mol Plant Microbe Interact 13:1092–1101
Dong J, Chen C, Chen Z (2003) Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol Biol 51:21–37
Du L, Chen Z (2000) Identification of genes encoding receptor-like protein kinases as possible targets of pathogen- and salicylic acid-induced WRKY DNA-binding proteins in Arabidopsis. Plant J 24:837–847
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206
Hara K, Yagi M, Kusano T, Sano H (2000) Rapid systemic accumulation of transcripts encoding tobacco WRKY transcription factor upon wounding. Mol Gen Genet 263:30–37
Hinderhofer K, Zentgraf U (2001) Identification of a transcription factor specifically expressed at the onset of leaf senescence. Planta 213:469–473
Hood E, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Nat Acad Sci USA 103:12987–12992
Huang T, Duman JG (2002) Cloning and characterization of a thermal hysteresis (antifreeze) protein with DNA-binding activity from winter bittersweet nightshade, Solanum dulcamara. Plant Mol Biol 48:339–350
Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, Seki M, Shinozaki K, Yaguchi-Shinozaki K (2006) Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol 47:141–153
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Johnson CS, Kolevski B, Smyth DS (2002) TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell 14:1359–1375
Karaba A, Dixit S, Greco R, Aharoni A, Trijatmiko KR, Marsch-Martinez N, Krishnan A, Nataraja KN, Udayakumar M, Pereira A (2007) Improvement of water use efficiency in rice by expression of HARDY, an Arabidopsis drought and salt tolerance gene. Proc Natl Acad Sci USA 104:15270–15275
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291
Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2004) A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low-temperature stress tolerance in tobacco by gene transfer. Plant Cell Physiol 45:346–350
Kim CY, Lee SH, Park HC, Bae CG, Cheong YH, Choi YJ, Han C, Lee SY, Lim CO, Cho MJ (2000) Identification of rice blast fungal elicitor-responsive genes by differential display analysis. Mol Plant Microbe Interact 13:470–474
Lagacé M, Matton DP (2004) Characterization of a WRKY transcription factor expressed in late torpedo-stage embryos of Solanum chacoense. Planta 219:185–189
Liu X, Bai X, Wang X, Chu C (2007) OsWRKY71, a rice transcription factor, is involved in rice defense response. J Plant Physiol 164:969–979
Mare C, Mazzucotelli E, Crosatti C, Francia E, Stanca AM, Cattivelli L (2004) Hv-WRKY38: a new transcription factor involved in cold- and drought-response in barley. Plant Mol Biol 55:399–416
Oono Y, Seki M, Nanjo T, Narusaka M, Fujita M, Satoh R, Satou M, Sakurai T, Ishida J, Akiyama K, Iida K, Maruyama K, Satoh S, Yamaguchi-Shinozaki K, Shinozaki K (2003) Monitoring expression profiles of Arabidopsis gene expression during rehydration process after dehydration using ca 7000 full-length cDNA microarray. Plant J 34:868–887
Ozturk ZN, Talamé V, Deyholos M, Michalowski CB, Galbraith DW, Gozukirmizi N, Tuberosa R, Bohnert HJ (2002) Monitoring large-scale changes in transcript abundance in drought- and salt-stressed barley. Plant Mol Biol 48:551–573
Pnueli L, Hallak-Herr E, Rozenberg M, Cohen M, Goloubinoff P, Kaplan A, Mittler R (2002) Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J 31:319–330
Qiu Y, Jing S, Fu J, Li L, Yu D (2004) Cloning and analysis of expression profile of 13 WRKY genes in rice. Chin Sci Bull 49:2159–2168
Ramamoorthy R, Jiang SY, Kumar N, Venkatesh PN, Ramachandran S (2008) Comprehensive transcriptional profiles of the WRKY gene family in rice under various abiotic and phytohormone treatments. Plant Cell Physiol 49:865–879
Rizhsky L, Liang H, Mittler R (2002) The combined effect of drought stress and heat shock on gene expression in tobacco. Plant Physiol 130:1143–1151
Rizhsky L, Davletova S, Liang H, Mittler R (2004) The zinc finger protein Zat12 is required for cytosolic ascorbate peroxidase 1 expression during oxidative stress in Arabidopsis. J Biol Chem 279:11736–11743
Robatzek S, Somssich IE (2001) A new member of the Arabidopsis WRKY transcription factor family, AtWRKY6, is associated with both senescence- and defence-related processes. Plant J 28:123–133
Romero C, Bellés JM, Vayá JL, Serrano R, Culiáñez-Macià FA (1997) Expression of the yeast trehalose-6-phosphate synthase gene in transgenic tobacco plants: pleiotropic phenotypes include drought tolerance. Planta 201:293–297
Ryu HS, Han M, Lee SK, Cho JI, Ryoo N, Heu S, Lee YH, Bhoo SE, Wang GL, Hahn TR, Jeon JS (2006) A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defense response. Plant Cell Rep 25:836–847
Sakuma Y, Maruyama K, Qin F, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2006) Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc Natl Acad Sci USA 103:18822–18827
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Shimono M, Sugano S, Nakayama A, Jiang CJ, Ono K, Toki S, Takatsuji H (2007) Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19:2064–2076
Shirasawa K, Takabe T, Takabe T, Kishitani S (2006) Accumulation of glycinebetaine in rice plants that overexpress choline monooxygenase from spinach and evaluation of their tolerance to abiotic stress. Ann Bot (Lond) 98:565–571
Shiroto Y, Toriyama K, Kishitani S (2004) Gene expression analysis of transcription factors in rice plants subjected to drought, heat or combination of both stresses. Breed Res 6(Suppl 2):129
Sun C, Palmqvist S, Olsson H, Borén M, Ahlandsberg S, Jansson C (2003) A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell 15:2076–2092
Thomashow MF (1999) Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu Rev Plant Physiol Plant Mol Biol 50:571–599
Wu KL, Guo ZJ, Wang HH, Li J (2005) The WRKY family of transcription factors in rice and Arabidopsis and their origins. DNA Res 12:9–26
Xie Z, Zhang Z-L, Zou X, Huang J, Ruas P, Thompson D, Shen QJ (2005) Annotations and functional analyses of the rice WRKY Gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol 137:176–189
Xu YH, Wang JW, Wang S, Wang JY, Chen XY (2004) Characterization of GaWRKY1, a cotton transcription factor that regulates the sesquiterpene synthase gene (+)-δ-cadinene synthase-A. Plant Physiol 135:507–515
Xu DQ, Huang J, Guo SQ, Yang X, Bao YM, Tang HJ, Zhang HS (2008) Overexpression of a TFIIIA-type zinc finger protein gene ZFP252 enhances drought and salt tolerance in rice (Oryza sativa L.). FEBS Lett 582:1037–1043
Yoda H, Ogawa M, Yamaguchi Y, Koizumi N, Kusano T, Sano H (2002) Identification of early-responsive genes associated with the hypersensitive response to tobacco mosaic virus and characterization of a WRKY-type transcription factor in tobacco plants. Mol Gen Genet 267:154–161
Yokoi S, Tsuchiya T, Toriyama K, Hinata K (1997) Tapetum-specific expression of the Osg6B promoter-beta-glucuronidase gene in transgenic rice. Plant Cell Rep 16:363–367
Yokoi S, Higashi S, Kishitani S, Murata N, Toriyama K (1998) Introduction of the cDNA for Arabidopsis glycerol-3-phosphate acyltransferase (GPAT) confers unsaturation of fatty acids and chilling tolerance of photosynthesis on rice. Mol Breed 4:269–275
Zhang YJ, Wang LJ (2005) The WRKY transcription factor superfamily: its origin in eukaryotes and expansion in plants. BMC Evol Biol 5:1–12
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zou X, Shen QJ, Neuman D (2007) An ABA inducible WRKY gene integrates responses of creosote bush (Larrea tridentate) to elevated Co2 and abiotic stresses. Plant Sci 172:997–1004
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Ebinuma.
Rights and permissions
About this article
Cite this article
Wu, X., Shiroto, Y., Kishitani, S. et al. Enhanced heat and drought tolerance in transgenic rice seedlings overexpressing OsWRKY11 under the control of HSP101 promoter. Plant Cell Rep 28, 21–30 (2009). https://doi.org/10.1007/s00299-008-0614-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-008-0614-x