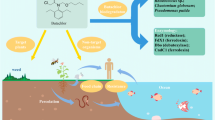
Abstract
Basically, nitro functional group-containing chemicals have been used to synthesize various useful products like dyes, pesticides and solvents, and also military products and so on. Hence, many nitroaromatics (including nitrophenols) have been continuously released into the environment and appear in the soil and water. Some are known to be toxic due to their great impact on living systems (especially on health). Most such chemicals (nitroaromatic compounds) are listed as priority chemicals by the Environmental Protection Agency (EPA). The vast use of such chemicals and their toxic effects had led to the study of the degradation of nitro group-containing chemicals by microbes (an easily available and cost-effective treatment). In view of this, we discuss the degradation of a few nitro group-containing compounds and herbicide(s) by microorganisms from published literature, and we consider the future perspective.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
16.1 Introduction
Over the last several decades, natural and manmade chemicals, including antimicrobial agents, antibiotics, personal care products, pesticides, herbicides, chlorinated aromatics and nitroaromatics, among others, have been used for various purposes in day-to-day life (Hoskeri et al. 2011, 2014; Megadi et al. 2010; Mulla et al. 2011a, b, 2016a, b; Tallur et al. 2015; Talwar et al. 2014). The long-term use of consumer products has released these chemicals to the environment, where they continually show up at levels from less than 1 nanogram to more than 1 microgram per litre (Mulla et al. 2012, 2014, 2016c, d, 2018). In recent years, several (including chloro and nitro group-containing aromatics) were found to be toxic to living beings—from humans to aquaculture organisms (Kovacic and Somanathan 2014; Edalli et al. 2018; Mulla et al. 2014, 2016c, d, 2017; Tallur et al. 2015). Various researchers from many countries have thus been prompted to investigate decontamination methods that would be effective on such chemicals (Arora et al. 2014, 2017; Burkul et al. 2015; Li and Yang 2018; Osin et al. 2018). Among the avenues considered, of the use of non-harmful biological mediators to detoxify toxic chemicals in wastewater treatment plants gained significantly more importance. In this chapter, we discuss some nitro group-containing compounds as used for degradation of toxic substances in microbes.
16.2 Uses of and Environmental Pollution by Nitro Group-Containing Compounds
In general, nitro group-containing chemicals go into the synthesis of certain useful products, for example, explosives, dyestuffs, insecticides, herbicides and rubbers (Arora et al. 2014; Douglas et al. 2011; Haizhen et al. 2009; Khalid et al. 2009; Spain 1995; Wan et al. 2007; Ye et al. 2004). In addition, they are also found in the preparation of products like medicines, fuels for vehicles, wall paints, solvents for polish, electronic batteries, coloured glass, etc. (Beard and Noe 1981; Dunlap 1982; Hirai 1999; Plunkett 1966; Windholz et al. 1976; Ware 1994).
On the other hand, most of nitro group-containing chemicals (including nitrophenols), are stable, but, some possess toxic properties (especially carcinogenicity) (Ju and Parales 2010; Kovacic and Somanathan 2014; Nishino et al. 2000; Padda et al. 2003; Purohit and Basu 2000). Hence, several (including nitrophenols) are included in the U.S. EPA list (Ju and Parales 2010). Some of these nitro group-containing chemicals have also been detected in food products. Likewise, mutagenic property-containing nitroaromatic compounds have been found in the atmosphere, especially by way of cigarette smoke and vehicle fuels (Kinouchi and Ohnishi 1983). Most of these have similar functional properties that make them beneficial for industrial use, on the one hand, while causing them to be harmful to the health of living systems. In general, the widespread use of nitroaromatics impacts the environment by contaminating soil and water (including groundwater) (Rieger and Knackmuss 1995; Ju and Parales 2010). Likewise, nitro functional group-containing insecticides are purposely applied to crops, leaving surrounding fields open to possible contamination. Additionally, inadequate management and/or loading practices by both manufacturers and customers results in their unintended injection into the environment (Mulla et al. 2014). For example, a Toxics Release Inventory report 2002 reported that nearly 11500 kg of 2,4-Dichloro-1-(4-nitrophenoxy)-benzene, 5100 kg of nitrobenzene and 1100 kg of 2,4-dinitrotoluene have been discharged into territory of United States (White and Claxton 2004). Ecological contamination from explosives manufacturers also occurs in Germany (Spain et al. 2000). The toxicological effects of nitro group-containing chemicals are well discussed in the literature (Kovacic and Somanathan 2014).
16.3 Biodegradation of Nitro Group-Containing Chemicals
Among multipurpose industrial organic compounds, nitro group-containing chemicals are among those most used in pesticides, pharmaceuticals, pigments, dyes, etc. (Haghighi-Podeh and Bhattacharya 1996). These chemicals may stay in the soil as the by-products of insecticides like parathion, methyl-parathion and other more complex nitroaromatics (including herbicides) through hydrolysis. Hence, many researchers with the aim of detoxification of such chemicals have demonstrated that various organisms are able to utilize different types of nitrophenols as a growth substrate. Generally, the nitro group-containing chemical degradation process involved oxidation (monooxygenase and/or dioxygenase) and/or reduction (reductase) or Meisenheimer complex generation.
16.3.1 Microbial Degradation of Mononitrophenols
The bacterial culture Pseudomonas putida B2 utilizes both ortho- and meta-nitrophenol as a growth substrate and degrades both substrates by producing different pathways (Zeyer and Kearney 1984). In Alcaligenes sp. NyZ215 (Xiao et al. 2007) and Pseudomonas putida B2, enzyme monooxygenase initiates oxidative mechanism of 2-nitrophenol and is transformed to catechol with the elimination of nitrite ions. Catechol is further degraded by catechol 1,2-dioxygenase and finally enters into TCA cycle (Zeyer and Kearney 1984; Zeyer et al. 1986) (Fig. 16.1).
On the other hand, there are two different degradative pathways for 3-nitrophenol. However, both initiate with reduction of 3-nitrophenol to form 3-hydroxylaminophenol (Fig. 16.2).
During the degradation of 3-nitrophenol in Pseudomonas putida B2, the compound is initially transformed to 3-hydroxylaminophenol by reductase enzyme and subsequently transformed to 1,2,4-trihydroxybenzene with the release of ammonia (Meulenberg et al. 1996) (Fig. 16.2). In contrast, in Ralstonia eutropha JMP134, 3-nitrophenol is transformed to 3-hydroxylaminophenol through aminohydroquinone molecule (Fig. 16.2), and ammonia is released in the ring-cleavage pathway (Schenzle et al. 1997). Moreover, the enzymes involved during the degradation of 3-nitrophenol in strain JMP134 also help the bacterium to utilize 2-chloro-5-nitrophenol as the growth substrate (Schenzle et al. 1999).
On the other hand, various microbial strains are isolated and identified on the basis of their capacity to utilize 4-nitrophenol as a growth substrate. In most of cases it has been observed that 4-nitrophenol is degraded into either only 4-nitrocatechol by oxidative mechanism and/or further altered to 1,2,4-trihydroxybenzne with the release of nitrite (Fig. 16.3) in different genus bacteria like Achromobacter xylosoxidans Ns, Arthrobacter chlorophenolicus A6, Arthrobacter sp. CN2, Arthrobacter sp. strain JS443, Arthrobacter sp. Y1, Arthrobacter protophormiae RKJ100, Bacillus sphaericus JS905, Burkholderia cepacia RKJ200, Ralstonia sp. SJ98, Rhodococcus opacus AS2, Rhodococcus erythropolis AS3, Rhodococcus imtechensis strain RKJ300, Rhodococcus opacus SAO101 and Serratia sp. strain DS001 (Arora et al. 2014; Gosh et al. 2010; Jain et al. 1994; Ju and Parales 2010; Kadiyala and Spain 1998; Kitagawa et al. 2004; Li et al. 2008; Pakala et al. 2007; Unell et al. 2008; Wang et al. 2016).
Kadiyala and Spain (1998) demonstrated that enzymes like oxygenase and flavoprotein reductase are involved in the first two oxidation steps of 4-nitrophenol to 2-hydroxyl-1,4-quinone by the release of nitrite in Bacillus sphericus JS905 (Fig. 16.3). However, bacterial strains like Arthrobacter aurescens TW17, Arthrobacter sp. YUP-3, Pseudomonas putida JS444, Pseudomonas sp. strain WBC-3, Methylobacterium sp. C1, Moraxella sp., Rodococcus opacus SAO101 and Rodococcus sp. PN1 (Hanne et al. 1993; Kitagawa et al. 2004; Nishino and Spain 1993; Spain and Gibson 1991; Takeo et al. 2008; Tian et al. 2018; Yue et al. 2018; Zhang et al. 2009) transformed 4-nitrophenol to benzoquinone with the help of enzyme monooxygenase. The by-product further converted to hydroquinone molecule (Spain and Gibson 1991) (Fig. 16.3). Furthermore, the pathways are similar at the ring cleavage, where hydroquinone and 1,2,4-trihydroxybenzene molecules individually transform into maleylacetate and finally enter into TCA cycle (Fig. 16.3). Hanne et al. (1993) studied the degradation of 4-nitrophenol by Nocardia sp. strain TW2 in the presence of different chemical inducers, and their results suggest that 1,2,4-trihydroxybenzene and hydroquinone pathways are expressed differently. Likewise, a similar process was also observed in both Rhodococcus strain PN1 and QAO101. Hence, an in-depth study of these bacterial strains is essential in order to know the exact mechanism of regulation of 4-nitrophenol catabolism. Yue et al. (2018) studied the bioaugmentation of Methylobacterium sp. C1 towards 4-nitrophenol removal, and their results suggest that Methylobacterium sp. C1 with other two bacterial cultures (Bacillus megaterium T1 and Bacillus cereus G5) biofilm were shown to be highly resistant as well as more efficient for the removal of 4-nitrophenol (up to 0.6 g/L). On the other hand, Min et al. (2017a) demonstrated by applying Burkholderia sp. strain SJ98 to artificially contaminated soil (4-nitrophenol, 3-methyl-4-nitrophenol and 2-chloro-4-nitrophenol), and their results suggest that the bacterial culture efficiently degrades all three compounds, thereby decreasing the concetration of nitrophenol, leading significantly to enhancement of several genera of rich microorganisms like Nonomuraea, Kribbella and Saccharopolyspora. Recently, Subashchandrabose et al. (2018) demonstrated 4-nitrophenol degradation by Rhodococcus wratislaviensis strain 9, and their results showed the organism is able to degrade 900 μM of 4-nitrophenol within 14 h. Likewise, 4-nitroanisole degradation in two individual bacterial strains (AS2 and AS3) was studied, showing that 4-nitrophenol as a key intermediate occurs during degradation (Schäfer et al. 1996) (Fig. 16.3).
Teramoto et al. (2004) studied how the fungal culture Phanerochaete chrysosporium can utilize 4-nitrophenol as a sole source of carbon and energy. However, basically under ligninolytic conditions, 4-nitrophenol transforms to 1,2-dimethoxy-4-nitrobenzene (DMNB) through by-product formation of 4-nitroanisole by the fungus (Fig. 16.4).
16.3.2 Microbial Degradation of Di- and Trinitrophenol
Very few known microbes are able to utilize both dinitrophenols such as 2,4-dinitrophenol and 2,6-dinitrophenol and trinitrophenols like picric acid (2,4,6-trinitrophenol) as growth substrate. Ecker et al. (1992) demonstrated a bacterium is able to utilize 2,6-dinitrophenol as a growth substrate. Interestingly, in Cupriavidus necator JMP134, the degradative pathway of 2,6-dinitrophenol is different than the 3-nitrophenol degradation pathway.
In bacterial strain JMP134, degradation initiates with the oxidation step to transform 2,6-dinitrophenol to 4-nitro pyrogallol with the help of a dioxygenase enzyme that releases the first nitrite ion from the benzene ring. In the next step, the by-product is further converted to 2-hydroxy-5-nitro muconate in a ring-cleavage and subsequently transforms to 2-hydroxy-5-nitropenta-2,4-dienoic acid by decarboxylation (Fig. 16.5). However, it was predicted that the other nitro radical present on the broken benzene ring might be released during subsequent steps. On the other hand, various genera of bacterial cultures like Anabaena variabilis, Anabaena cylindrica, Burkhorderia KU-46, Haloanaerobiumpraevalens DSM 2228, Janthinobacterium sp., Nocardioides sp. strain CB22–2, Methanococcus sp. B, Nocardioides simplex FJ2-1A, Rhodococcus erythropolis strain HL24–1, Rhodococcus erythropolis strain Hl24–2, Rhodococcus imtechensis strain RKJ300, Rodococcus sp. strain RB1, Rhodococcus sp. strain NJUST16 and Sporohalobacter marismortui ATCC 35420 (Behrend and Heesche-Wagner 1999; Blasco et al. 1999; Boopathy 1994; Gosh et al. 2010; Hess et al. 1993; Hirooka et al. 2006; Iwaki et al. 2007; Lenke and Knackmuss 1992; Oren et al. 1991; Rajan et al. 1996; Shen et al. 2009; Zin et al. 2018) utilize either 2,4-dinitrophenol and/or picric acid as a growth substrate. Initially, Knackmuss and research group studied the catabolic pathway of 2,4-dinitrophenol and picric acid in microorganism(s) (Ju and Parales 2010) (Fig. 16.6).
During the degradation process, a hydride-Meisenheimer complex is formed by the reduction of picric acid and produces 2,4-dinitrophenol with the release of a nitrite radical. Again, by hydride-Meisenheimer complex and reduction, hydrolytic cleavage of 2,4-dinitrophenol occurs, producing 4,6-dinitrohexanoate and finally entering into TCA cycle through various by-product formations. Yet, few of the responsible genes and enzymes of the degradative pathway have been determined with respect to their functional characterization and regulation mechanism (Arora et al. 2014; Ju and Parales 2010). Still, it is necessary to know the exact mechanism involved in catabolic regulation of 2,6-dinitrophenol, 2,4-dintrophenol and picric acid in different bacterial strains.
16.3.3 Microbial Degradation of Chloro and Nitro Group-Containing Chemicals
Very few microbes (Arthrobacter nitrophenolicus SJCon, Burkholderia sp. RKJ800, Burkholderia sp. strain SJ98, Cupriavidus sp. strain CNP-8 and Rhodococcus imtechensis RKJ300, etc.) are known to utilize 2-chloro-4-nitrophenol as a growth substrate (Arora et al. 2014, 2017; Min et al. 2018). A Rhodococcus imtechensis strain RKJ300 was isolated from pesticide-contaminated soil by enrichment with 4-nitrophenol as a growth substrate (Gosh et al. 2010). The organism also utilizes 2-chloro-4-nitrophenol as a growth substrate. In Cupriavidus sp. strain CNP-8 and Rhodococcus imtechensis strain RKJ300, the degradation of 2-chloro-4-nitrophenol initiates with oxidation to form chlorohydroquinone with the release of nitrite. In the next step, the metabolic by-product is dechlorinated and forms hydroquinone and subsequently converts to gamma-hydroxymuconic semialdyhyde and finally enters into TCA cycle (Fig. 16.7).
In Arthrobacter nitrophenolicus SJCon, 2-chloro-4-nitrophenol degraded to chlorohydroquinone and was further converted to maleylacetate by oxidation, which finally entered into TCA cycle. On the other hand, in Burkholderia sp. SJ98, 2-chloro-4-nitrophenol degraded into 4-nitrophenol with the release of chloride radicals, which was further sequentially converted into maleylacetate via 4-nitrocatechol and 1, 2, 4-benzenetriol and finally entered the TCA pathway (Fig. 16.7).
Schenzle et al. (1999) demonstrated that Ralstonia eutropha JMP134 (also known more recently as Cupriavidus necator) utilizes 2-chloro-5-nitrophenol as a growth substrate. Initially, 2-chloro-5-nitrophenol is reduced to 2-chloro-5-hydroxylaminophenol, then follows an enzymatic Bamberger rearrangement forming 2-amino-5-chlorohydroquinone, which is further converted to aminohydroquinone with the release of chlorine (Fig. 16.8). Similarly, in Cupriavidus sp. strain CNP-8, 2-chloro-5-nitrophenol degradation is initiated by reduction and finally enters into TCA cycle via sequential transformation to different by-products (Min et al. 2017b).
16.3.4 Microbial Degradation of Nitro Group-Containing Herbicides
Different genera of bacterial cultures like Arthrobacter-like organisms, Corynebacterium simplex, a Pseudomonas-like strain and two unidentified bacterial strains isolated from soil showed the ability to degrade a well-known herbicide 3,5-dinitro-ortho-cresol (Gundersen and Jensen 1956; Jensen and Gundesen 1955; Jesnen and Lautrup-Larsen 1967; Tabak et al. 1964). Interestingly, in all of these studies the only intermediate by-product detected was a nitrite radical. Tewfik and Evans (1966) reported concisely on a Pseudomonas sp. that metabolized 3,5-dinitro-ortho-cresol by a reduction process. During degradation, both nitro functional groups were reduced to amino groups, which were further oxidatively deaminated to form 2,3,5-trihydroxytoluene before entering into the ring-cleavage pathway (Fig. 16.9). On the other hand, in Arthrobacter simplex, 3,5-dinitro-ortho-cresol initially was converted to 3-methyl-5-nitrocatechol and subsequently transformed to 2,3,5- trihydroxytoluene and finally entered into TCA cycle via ring fission (Tewfik and Evans 1966) (Fig. 16.9).
Meanwhile, the same research group also found that in the Corynebacterium simplex strain (reported earlier by Gundersen and Jensen 1956), 3,5-dinitro-ortho-cresol is degraded by essentially the same metabolic route; however, apart from that, reduced by-products probably were not formed as no nitroreductase activity could be confirmed in this organism (Fig. 16.9). Likewise, degradation of the other herbicide, 2-sec-butyl-4,6-dinitrophenol (dinoseb) by bacterial isolates was studied and reported (Stevens et al. 1991). However, the isolates in this case were unable to degrade the herbicide without a co-substrate. But, when the medium contained glucose/ammonium chloride with dinoseb under unshaken conditions, the medium turned a bright red colour, indicating formation of by-product(s). Various transformed products were detected and identified by GC-MS (Stevens et al. 1991). Similarly, Kaake et al. (1995) demonstrated a dinoseb degradation mechanism by bacterial cultures under reducing conditions, and they proposed a degradative pathway for dinoseb on the basis of reduction of nitro groups (attached on the benzene ring of dinoseb) with the formation of amino groups, which subsequently were replaced with hydroxyl groups.
16.4 Conclusions and Perspectives
The information reviewed in the present chapter represents the immense effort undertaken in the investigation to carry out the detoxification of nitro group-containing chemicals and herbicides. In recent times, the topic of biodegradation of such chemicals in biological living systems has emerged in the scientific community. The applications of microbes (individual pure and mixed cultures) towards catabolism of nitro group-containing chemicals and herbicides are being used now, especially in wastewater treatment plants, because, the method is freely available and cost-effective. Degradation mechanisms (especially genomic profile and enzymatic characterization) of several nitro group-containing chemicals have been comprehensively investigated in microbes (especially in bacteria). However, a few key questions still need to be clarified, among others: What is the behaviour of microbes towards a shock load of such chemicals? Can these microbes remediate such chemicals under various environmental conditions? Can a pure culture co-operate with other organism(s) (especially in a bioaugmentation process) during the degradation of such chemicals? The present substantial information for understanding the mechanisms of microbial degradation of nitro group-containing chemicals will help future studies.
References
Arora, P. K., Srivastav, A., & Singh, V. P. (2014). Bacterial degradation of nitrophenols and their derivatives. Journal of Hazardous Materials, 266, 42–59.
Arora, P. K., Srivastava, A., Garg, S. K., & Singh, V. P. (2017). Recent advances in degradation of chloronitrophenols. Bioresource Technology, 250, 902–909.
Beard, R. R., & Noe, J. T. (1981). In G. D. Clayton & F. E. Clayton (Eds.), Patty’s handbook of industrial hygiene and toxicology (Vol. 2A, 3rd ed., pp. 2413–2489). New York: Wiley-Interscience.
Behrend, C., & Heesche-Wagner, K. (1999). Formation of hydride-Meisenheimer complexes of picric acid (2,4,6-trinitrophenol) and 2,4-dinitrophenol during mineralization of picric acid by Nocardioides sp. strain CB22-2. Applied and Environmental Microbiology, 65, 1372–1377.
Blasco, R., Moore, E., Wray, V., Pieper, D. H., Timmis, K., & Castillo, F. (1999). 3-Nitroadipate, a metabolic intermediate for the mineralization of 2,4-dinitrophenol by a new strain of a Rhodococcus species. Journal of Bacteriology, 181, 149–152.
Boopathy, R. (1994). Transformation of nitroaromatic compounds by a methanogenic bacterium, Methanococcus sp. (strain B). Archives of Microbiology, 62, 167–172.
Burkul, R. M., Ranade, S. V., & Pangarkar, B. L. (2015). Removal of pesticides by using various treatment method: Review. International Journal of Emerging Trends in Engineering and Basic Sciences, 2, 88–91.
Douglas, T. A., Walsh, M. E., McGrath, C. J., Weiss, C. A., Jaramillo, A. M., & Trainor, T. P. (2011). Desorption of nitramine and nitroaromatic explosive residues from soils detonated under controlled conditions. Environmental Toxicology and Chemistry, 30, 345–353.
Dunlap, K. L. (1982). In H. F. Mark, D. F. Othmer, C. G. Overberger, & G. T. Seaborg (Eds.), Kirk and Othmer’s encyclopaedia of chemical technology (Vol. 15, 3rd ed., pp. 916–932). New York: Wiley.
Ecker, S., Widmann, T., Lenke, H., Dickel, O., Fischer, P., Bruhn, C., & Knackmuss, H. -J. (1992). Catabolism of 2,6-dinitrophenol by Alcaligenes eutrophus JMP 134 and JMP 222. Archives of Microbiology, 158(2), 149–154.
Edalli, V. A., Patil, K. S., Le, V. V., & Mulla, S. I. (2018). An overview of aniline and chloroaniline compounds as environmental pollutants. Significances of Bioengineering & Biosciences, 1(4), 1–2. https://doi.org/10.31031/SBB.2018.01.000519.
Gosh, A., Khurana, M., Chauhan, A., Takeo, M., Chakraborti, A. K., & Jain, R. K. (2010). Degradation of 4-nitrophenol, 2-chloro-4-nitrophenol and 2,4-dinitrophenol by Rhodococcusimtechensis strain RKJ300. Environmental Science and Technology, 44, 1067–1077.
Gundersen, K., & Jensen, H. L. (1956). A soil bacterium decomposing organic nitro-compounds. Acta Agriculturae Scandinavica, 6, 110–114.
Haghighi-Podeh, M. R., & Bhattacharya, S. K. (1996). Fate and toxic effects of nitrophenols on anaerobic treatment systems. Water Science and Technology, 34, 345–350.
Haizhen, W., Chaohai, W., Yaqin, W., Qincong, H., & Shizhong, L. (2009). Degradation of o-chloronitrobenzene as the carbon & nitrogen sources by Pseudomonas putida OCNB-1. Journal of Environmental Sciences, 21, 89–95.
Hanne, L. F., Kirk, L. L., Appel, S. M., Narayan, A. D., & Bains, K. K. (1993). Degradation and induction specificity in actinomycetes that degrade p-nitrophenol. Applied and Environmental Microbiology, 59, 3505–3508.
Hess, T. F., Silverstein, J., & Schmidt, S. K. (1993). Effect of glucose on 2,4-dinitrophenol degradation kinetics in sequencing batch reactors. Water Environment Research, 65(1), 73–81.
Hirai, K. (1999). Structural evolution and synthesis of diphenyl ethers, cyclic imides, and related compounds. In P. Boger & K. Wakabayashi (Eds.), Peroxidizing herbicides (pp. 15–72). Berlin: Springer.
Hirooka, T., Nagase, H., Hirata, K., & Miyamoto, K. (2006). Degradation of 2,4-dinitrophenol by a mixed culture of photoautotrophic microorganisms. Biochemical Engineering Journal, 29(1), 157–162.
Hoskeri, R. S., Mulla, S. I., Shouche, Y. S., & Ninnekar, H. Z. (2011). Biodegradation of 4- chlorobenzoic acid by Pseudomonas aeruginosa PA01 NC. Biodegradation, 22, 509–516.
Hoskeri, R. S., Mulla, S. I., & Ninnekar, H. Z. (2014). Biodegradation of chloroaromatic pollutants by bacterial consortium immobilized in polyurethene foam and other matrices. Biocatalysis and Agricultural Biotechnology, 3, 390–396.
Iwaki, H., Abe, K., & Hasegawa, Y. (2007). Isolation and characterization of a new 2,4-dinitrophenol-degrading bacterium Burkholderia sp. strain KU-46 and its degradation pathway. FEMS Microbiology Letters, 274(1), 112–117.
Jain, R. K., Dreisbach, J. H., & Spain, J. C. (1994). Biodegradation of p-nitrophenol via 1,2,4-benzenetriol by an Arthrobacter sp. Applied and Environmental Microbiology, 60, 3030–3032.
Jensen, H. L., & Gundersen, K. (1955). Biological decomposition of aromatic nitro compounds. Nature, 175, 341.
Jensen, H. L., & Lautrup-Larsen, G. (1967). Microorganisms that decompose nitro-aromatic compounds, with special reference to dinitro-ortho-cresol. Acta Agriculturae Scandinavica, 17, 115–126.
Ju, K. S., & Parales, R. E. (2010). Nitroaromatic compounds, from synthesis to biodegradation. Microbiology and Molecular Biology Reviews, 74, 250–272.
Kaake, R. H., Crawford, D. L., & Crawford, R. L. (1995). Biodegradation of the nitroaromatic herbicide dinoseb (2-sec-butyl-4,6-dinitrophenol) under reducing conditions. Biodegradation, 6, 329–337.
Kadiyala, V., & Spain, J. C. (1998). A two-component monooxygenasecatalyzes both the hydroxylation of p-nitrophenol and the oxidative release of nitrite from 4-nitrocatechol in Bacillus sphaericus JS905. Applied and Environmental Microbiology, 64, 2479–2484.
Khalid, A., Arshad, M., & Crowley, D. E. (2009). Biodegradation potential of pure and mixed bacterial cultures for removal of 4-nitroaniline from textile dye wastewater. Water Research, 43, 1110–1116.
Kinouchi, T., & Ohnishi, Y. (1983). Purification and characterization of 1-nitropyrene nitroreductases from Bacteroidesfragilis. Applied and Environmental Microbiology, 46, 596–604.
Kitagawa, W., Kimura, N., & Kamagata, Y. (2004). A novel p-nitrophenol degradation gene cluster from a gram-positive bacterium, Rhodococcusopacus SAO101. Journal of Bacteriology, 186, 4894–4902.
Kovacic, P., & Somanathan, R. (2014). Nitroaromatic compounds: Environmental toxicity, carcinogenicity, mutagenicity, therapy and mechanism. Journal of Applied Toxicology, 34, 810–824.
Lenke, H., & Knackmuss, H. J. (1992). Initial hydrogenation during catabolism of picric acid by Rhodococcuserythropolis HL 24-2. Applied and Environmental Microbiology, 58, 2933–2937.
Li, Z., & Yang, P. (2018). Review on physicochemical, chemical, and biological processes for pharmaceutical wastewater. IOP Conference Series: Earth and Environmental Science, 113, 012185.
Li, Y. Y., Zhou, B., Li, W., Peng, X., Zhang, J. S., & Yan, Y. C. (2008). Mineralization of p-nitrophenol by a new isolate Arthrobacter sp. Y1. Journal of Environmental Science and Health. Part. B, 43, 692–697.
Megadi, V. B., Tallur, P. N., Mulla, S. I., & Ninnekar, H. Z. (2010). Bacterial degradation of Fungicide captan. Journal of Agricultural and Food Chemistry, 58, 12863–12868.
Meulenberg, R., Pepi, M., & de Bont, J. A. M. (1996). Degradation of 3-nitrophenol by Pseudomonas putida B2 occurs via 1,2,4-benzenetriol. Biodegradation, 7, 303–311.
Min, J., Wang, B., & Hu, X. (2017a). Effect of inoculation of Burkholderia sp. strain SJ98 on bacterial community dynamics and para-nitrophenol, 3-methyl-4-nitrophenol, and 2-chloro-4-nitrophenol degradation in soil. Scientific Reports, 7, 5983.
Min, J., Chen, W., Wang, J., & Hu, X. (2017b). Genetic and biochemical characterization of 2-chloro-5-nitrophenol degradation in a newly isolated bacterium, Cupriavidus sp. Strain CNP-8. Frontiers in Microbiology, 8, 1778.
Min, J., Wang, J., Chen, W., & Hu, X. (2018). Biodegradation of 2-chloro-4-nitrophenol via a hydroxyquinol pathway by a Gram-negative bacterium, Cupriavidus sp. strain CNP-8. AMB Express, 8, 43.
Mulla, S. I., Hoskeri, R. S., Shouche, Y. S., & Ninnekar, H. Z. (2011a). Biodegradation of 2-nitrotoluene by Micrococcus sp. strain SMN-1. Biodegradation, 22, 95–102.
Mulla, S. I., Manjunatha, T. P., Hoskeri, R. S., Tallur, P. N., & Ninnekar, H. Z. (2011b). Biodegradation of 3-nitrobenzoate by Bacillus flexus strain XJU-4. World Journal of Microbiology and Biotechnology, 27, 1587–1592.
Mulla, S. I., Talwar, M. P., Hoskeri, R. S., & Ninnekar, H. Z. (2012). Enhanced degradation of 3-nitrobenzoate by immobilized cells of Bacillus flexus strain XJU-4. Biotechnology and Bioprocess Engineering, 17, 1294–1299.
Mulla, S. I., Talwar, M. P., & Ninnekar, H. Z. (2014). Bioremediation of 2,4,6-Trinitrotoluene explosive residues. In S. N. Singh (Ed.), Biological remediation of explosive residues (Environmental science and engineering) (pp. 201–233). Cham: Springer.
Mulla, S. I., Bangeppagari, M. D., Mahadevan, G. D., Eqani, S. A. M. A. S., Sajjan, D. B., Tallur, P. N., Megadi, V. B., Harichandra, Z., & Ninnekar, H. Z. (2016a). Biodegradation of 3-chlorobenzoate and 3-hydroxybenzoate by polyurethane foam immobilized cells of Bacillus sp. OS13. Journal of Environmental Chemical Engineering, 4(2), 1423–1431.
Mulla, S. I., Sun, Q., Hu, A., Wang, Y., Ashfaq, M., Eqani, S. A. M. A. S., & Yu, C. P. (2016b). Evaluation of sulfadiazine degradation in three newly isolated pure bacterial cultures. PLoS One, 11, e0165013.
Mulla, S. I., Wang, H., Sun, Q., Hu, A., & Yu, C. P. (2016c). Characterization of triclosan metabolism in Sphingomonassp. strain YL-JM2C. Scientific Reports, 6, 21965.
Mulla, S. I., Hu, A., Wang, Y., Sun, Q., Huang, S. L., Wang, H., & Yu, C. P. (2016d). Degradation of triclocarban by a triclosan-degrading Sphingomonassp. strain YL-JM2C. Chemosphere, 144, 292–296.
Mulla, S. I., Ameen, F., Tallur, P. N., Bharagava, R. N., Bangeppagari, M., SAMAS, E., Bagewadi, Z. K., Mahadevan, G. D., Yu, C. P., & Ninnekar, H. Z. (2017). Aerobic degradation of fenvalerate by a gram-positive bacterium, Bacillus flexus strain XJU-4. 3 Biotech, 7, 320.
Mulla, S. I., Hu, A., Sun, Q., Li, J., Suanon, F., Ashfaq, M., & Yu, C. P. (2018). Biodegradation of sulfamethoxazole in bacteria from three different origins. Journal of Environmental Management, 206, 93–102.
Nishino, N., & Spain, J. C. (1993). Cell density-dependent adaptation of Pseudomonas putida to biodegradation of p-nitrophenol. Environmental Science & Technology, 27, 489–494.
Nishino, S. F., Spain, J. C., & He, Z. (2000). Strategies for aerobic degradation of nitroaromatic compounds by bacteria: Process discovery to field application. In J. C. Spain, J. B. Hugeghes, & H. J. Knackmuss (Eds.), Biodegradation of nitroaromatic compounds and explosives (pp. 7–61). New York: Lewis Publishing Co.
Oren, A., Gurevich, P., & Henis, Y. (1991). Reduction of nitrosubstituted aromatic compounds by the halophilic anaerobic eubacteria Haloanaerobiumpraevalens and Sporohalobactermarismortui. Applied and Environmental Microbiology, 57(11), 3367–3370.
Osin, O. A., Yu, T., Cai, X., Jiang, Y., Peng, G., Cheng, X., Li, R., Qin, Y., & Lin, S. (2018). Photocatalytic degradation of 4-nitrophenol by C, N-TiO2: degradation efficiency vs. embryonic toxicity of the resulting compounds. Frontiers in Chemistry, 6, 192.
Padda, R. S., Wang, C., Hughes, J. B., Kutty, R., & Bennett, G. N. (2003). Mutagenicity of nitroaromatic degradation compounds. Environmental Toxicology and Chemistry, 22, 2293–2297.
Pakala, S. B., Gorla, P., Pinjari, A. B., Krovidi, R. J., Baru, R., Yanamandra, M., Merrick, M., & Siddavattam, D. (2007). Biodegradation of methyl parathion and p-nitrophenol: evidence for the presence of p-nitrophenol 2-hydroxylase in a Gram-negative Serratiasp. strain DS001. Applied Microbiology and Biotechnology, 73, 1452–1462.
Plunkett, E. R. (1966). Handbook of industrial toxicology (pp. 152–153). New York: Chemical Publishing Co.
Purohit, V., & Basu, A. K. (2000). Mutagenicity of nitroaromatic compounds. Chemical Research in Toxicology, 13, 673–692.
Rajan, J., Valli, K., Perkins, R. E., Sariaslani, F. S., Barns, S. M., Reysenbach, A. L., Rehm, S., Ehringer, M., & Pace, N. R. (1996). Mineralization of 2,4,6-trinitrophenol (picric acid): characterization and phylogenetic identification of microbial strains. Journal of Industrial Microbiology & Biotechnology, 16, 319–324.
Rieger, P. G., & Knackmuss, H. J. (1995). Basic knowledge and perspectives on biodegradation of 2,4,6-trinitrotoluene and related nitroaromatic compounds in contaminated soil. In J. C. Spain (Ed.), Biodegradation of nitroaromatic compounds (Vol. 49, pp. 1–18). New York: Plenum Press.
Schafer, A., Harms, H., & Zehnder, A. J. (1996). Biodegradation of 4-nitroanisole by two Rhodococcusspp. Biodegradation, 7, 249–255.
Schenzle, A., Lenke, H., Fischer, P., Williams, P. A., & Knackmuss, H. J. (1997). Catabolism of 3-nitrophenol by Ralstoniaeutropha JMP134. Applied and Environmental Microbiology, 63, 1421–1427.
Schenzle, A., Lenke, H., Spain, J. C., & Knackmuss, H. J. (1999). Chemoselective nitro group reduction and reductive dechlorination initiate degradation of 2-chloro-5-nitrophenol by Ralstoniaeutropha JMP134. Applied and Environmental Microbiology, 65, 2317–2323.
Shen, J., Zhang, J., Zuo, Y., Wang, L., Sun, X., Li, J., Han, W., & He, R. (2009). Biodegradation of 2,4,6-trinitrophenol by Rhodococcus sp. isolated from a picric acid-contaminated soil. Journal of Hazardous Materials, 163, 1199–1206.
Spain, J. C. (1995). Biodegradation of nitroaromatic compounds. Annual Review of Microbiology, 49, 523–555.
Spain, J. C., & Gibson, D. T. (1991). Pathway for biodegradation of p-nitrophenol in a Moraxella sp. Applied and Environmental Microbiology, 57, 812–819.
Spain, J. C., Hughes, J. B., & Knackmuss, H. J. (Eds.). (2000). Biodegradation of nitroaromatic compounds and explosives. Boca Raton: CRC Press.
Stevens, T. O., Crawford, R. L., & Crawford, D. L. (1991). Selection and isolation of bacteria capable of degrading dinoseb (2-sec-butyl-4,6-dinitrophenol). Biodegradation, 2, 1–13.
Subashchandrabose, S. R., Venkateswarlu, K., Krishnan, K., Naidu, R., Lockington, R., & Megharaj, M. (2018). Rhodococcuswratislaviensis strain 9: An efficient p-nitrophenol degrader with a great potential for bioremediation. Journal of Hazardous Materials, 347, 176–183.
Tabak, H. H., Chambers, C. W., & Kabler, P. W. (1964). Microbial metabolism of aromatic compounds I.: Decomposition of phenolic compounds and aromatic hydrocarbons by phenol-adapted bacteria. Journal of Bacteriology, 87, 910–919.
Takeo, M., Murakami, M., Niihara, S., Yamamoto, K., Nishimura, M., Kato, D., & Negoro, S. (2008). Mechanism of 4-nitrophenol oxidation in Rhodococcus sp. strain PN1: Characterization of the two-component 4-nitrophenol hydroxylase and regulation of its expression. Journal of Bacteriology, 190, 7367–7374.
Tallur, P. N., Mulla, S. I., Megadi, V. B., Talwar, M. P., & Ninnekar, H. Z. (2015). Biodegradation of cypermethrin by immobilized cells of Micrococcus sp. strain CPN 1. Brazilian Journal of Microbiology, 46, 667–672.
Talwar, M. P., Mulla, S. I., & Ninnekar, H. Z. (2014). Biodegradation of organophosphate pesticide quinalphos by Ochrobactrumsp. strain HZM. Journal of Applied Microbiology, 117, 1283–1292.
Teramoto, H., Tanaka, H., & Wariishi, H. (2004). Degradation of 4-nitrophenol by the lignin-degrading basidiomycete Phanerochaete chrysosporium. Applied Microbiology and Biotechnology, 66(3), 312–317.
Tewfik, M. S., & Evans, W. C. (1966). The metabolism of 3,5-dinitro-o-cresol (DNOC). The Biochemical Journal, 99, 31–3231.
Tian, J., An, X., Liu, J., Yu, C., Zhao, R., Wang, J., & Chen, L. (2018). Optimization of 4-nitrophenol degradation by an isolated bacterium Arthrobacter sp. and the novel biodegradation pathways under nutrition deficient conditions. Journal of Environmental Engineering, 144(4), 04018012.
Unell, M., Nordin, K., Jernberg, C., Stenström, J., & Jansson, J. K. (2008). Degradation of mixtures of phenolic compounds by Arthrobacter chlorophenolicus A6. Biodegradation, 19, 495–505.
Wan, N., Gu, J. D., & Yan, Y. (2007). Degradation of p-nitrophenol by Achromobacter xylosoxidans Ns isolated from wetland sediment. International Biodeterioration and Biodegradation, 59, 90–96.
Wang, J., Ren, L., Jia, Y., Ruth, N., Shi, Y., Qiao, C., & Yan, Y. (2016). Degradation characteristics and metabolic pathway of 4-nitrophenol by a halotolerant bacterium Arthrobacter sp. CN2. Toxicological and Environmental Chemistry, 98, 226–240.
Ware, G. W. (1994). The pesticide book (4th ed.). Fresno: Thompson Publications.
White, P. A., & Claxton, L. D. (2004). Mutagens in contaminated soil: A review. Mutation Research, 567, 227–345.
Windholz, M., Budavari, S., Stroumtsos, L. Y., & Fertig, M. (1976). Merck Index (9th ed., pp. 6408–6474). Whitehouse Station: Merck and Co., Inc.
Xiao, Y., Zhang, J. J., Liu, H., & Zhou, N. Y. (2007). Molecular characterization of a novel ortho-nitrophenol catabolic gene cluster in Alcaligenes sp. strain NyZ215. Journal of Bacteriology, 189, 6587–6593.
Ye, J., Singh, A., & Ward, O. P. (2004). Biodegradation of nitroaromatics and other nitrogen-containing xenobiotics. World Journal of Microbiology and Biotechnology, 20, 117–135.
Yue, W., Chen, M., Cheng, Z., Xie, L., & Li, M. (2018). Bioaugmentation of strain Methylobacterium sp. C1 towards p-nitrophenol removal with broad spectrum coaggregating bacteria in sequencing batch biofilm reactors. Journal of Hazardous Materials, 344, 431–440.
Zeyer, J., & Kearney, P. C. (1984). Degradation of o-nitrophenol and m-nitrophenol by a Pseudomonas putida. Journal of Agricultural and Food Chemistry, 32, 238–242.
Zeyer, J., Kocher, H. P., & Timmis, K. N. (1986). Influence of para-substituents on the oxidative metabolism of o-nitrophenols by Pseudomonas putida B2. Applied and Environmental Microbiology, 52(2), 334–339.
Zhang, J. J., Liu, H., Xiao, Y., Zhang, X. E., & Zhou, N. Y. (2009). Identification and characterization of catabolic para-nitrophenol 4-monooxygenase and para-benzoquinone reductase from Pseudomonas sp. strain WBC-3. Journal of Bacteriology, 191, 2703–2710.
Zin, S. M., Habib, S., Yasid, N. A., & Ahmad, S. A. (2018). A Review on Microbial Degradation of 2,4-Dinitrophenol. Journal of Environmental Microbiology and Toxicology, 6, 28–33.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mulla, S.I. et al. (2019). An Overview of Nitro Group-Containing Compounds and Herbicides Degradation in Microorganisms. In: Arora, P. (eds) Microbial Metabolism of Xenobiotic Compounds. Microorganisms for Sustainability, vol 10. Springer, Singapore. https://doi.org/10.1007/978-981-13-7462-3_16
Download citation
DOI: https://doi.org/10.1007/978-981-13-7462-3_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-7461-6
Online ISBN: 978-981-13-7462-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)