Abstract
In recent decades, electrochemical DNA sensor has been demonstrated as up-and-coming alternative clinical diagnostic equipment through studying biological recognition matters and transforming them into a sensitive electrochemical output signals. This is because the conventional diagnostic strategies, for example, optical methods, have certain limitations including expensive, laborious, time-consuming, low sensitivity, and selectivity. Nucleic acid amplification strategies and principle of electrochemical signaling are two the most important aspects that should be considered before constructing highly sensitive and selective electrochemical sensor. In this chapter, we show how we are able to construct and investigate biosensing systems that utilize electrochemical signaling. Representative assays demonstrating the performance and merits of each nucleic acid amplification strategies are showed along with discussions on the limits of each strategy. Starting off from their sensing principles and significant characters, the adoption of nucleic acid amplification strategies in biosensing and their future outlooks are also provided.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
For sensitive biosensing, optical signals, such as fluorescence and colorimetry, are often used to quantitative detection of the target concentration. Conventional optical-based nucleic acid amplification methods not only demand accurate and cost equipment but also require sophisticated numerical algorithms to interpret the experimental data. In recent years, massive creative designs of electrochemical DNA sensors based on nucleic acid amplification techniques have been reported for trace target detection. These types of devices combine nucleic acid amplification methods with electrochemical transducers to produce an amplified signal, and thus offer a precise, inexpensive, and user-friendly platform for various practical applications, including environmental monitoring and clinical diagnosis.
A classic electrochemical DNA sensor studies the variation of electrical current that generated by redox reactions on the sensor surface. This method possesses several merits, including rapidity, simplicity, high sensitivity, and inexpensive. In particular, an electronic signal is directly yielded from electrochemical reactions; thus, costly equipment for signal transduction can be avoided. On account of the immobilization of functional DNA probes on electrode surfaces, miniaturized, portable, and sensitive biosensing systems can be created accordingly.
2 Preparation of Electrochemical DNA Sensor and Its Redox Labeling Methods
2.1 Preparation of Electrochemical DNA Sensor
For a classical electrochemical DNA sensor, gold electrode is one of the most frequently used working electrodes, and the thiolated DNA probes can be readily attached on the surface by stable gold–sulfur bonds (Gorodetsky et al. 2008). By phosphoramidite chemistry, the modification of terminal thiolated DNA can be simply realized when chemical synthesizes the probe. When the clean gold surface is immersed in a thiolated DNA solution, the immobilization of DNA can be realized readily on gold electrode surface, producing a self-assembled monolayer (SAM). The gold surface that immobilized with mixed SAM from alkanethiol alcohols and thiolated DNA generate a significantly better quality (less nonspecific adsorption and uniform packing) than that immobilized purely of thiolated DNA. The alkanethiol (e.g., 6-Mercaptohexanol)-DNA mixed SAM is traditionally used (White et al. 2008). Long alkanethiols, such as 11-mercaptoundecanol, are also found to passivate gold electrode surfaces. And the stability has been demonstrated after long-term dry storage (Lai et al. 2006).
In recent years, researchers devoted many efforts to explore new immobilization methods and supporting materials for the fabrication of DNA biosensors. In 2008, a DNA electrochemical biosensor based on covalent immobilization of ssDNA by coupling of sol-gel and self-assembled methods was fabricated (Fig. 7.1a). 3-Glycidoxypropyltrimethoxysiloxane (GPTMS) and 3-mercaptopropyltrimethoxysiloxane (MPTMS) were employed to prepare functional sol-gel film, which could assemble on the gold electrode surface by the sulfur-Au chemistry (Li et al. 2008b). Through co-condensation between silanols, GPTMS sol-gel interconnected into MPTMS sol-gel and offered epoxide groups for covalent immobilization of aminated ssDNA by epoxide/amine coupling reaction. The as-prepared biosensor showed promising performance, including good selectivity, high sensitivity, and regeneration ability. Furthermore, covalent immobilization of aminated ssDNA by diazotization coupling on self-assembled 4-ATP monolayer was developed (Fig. 7.1b) (Li et al. 2009a). The 4-ATP could stably assemble on the gold surface by sulfur–gold interaction. Through diazotization interaction, diazo-ATP for covalent immobilization of ssDNA was achieved, providing remarkable conductibility for electron transfer. Other materials, such as conducting polymers (Li et al. 2008c; Nie et al. 2009), also have been demonstrated to prepare functional DNA sensor with significant sensitivity and selectivity, which could be used for the detection of DNA hybridization events.
a Schematic representation for the construction of electrochemical DNA biosensor based on covalent attachment of DNA on self-assembled MPTMS-GPTMS sol-gel-modified gold electrode. Reprinted from Li et al. (2008b). Copyright 2008, with permission from Elsevier. b Fabrication of electrochemical DNA biosensor based on covalent immobilization of DNA on self-assembled diazo-ATP-modified gold electrode using diazotization coupling. Reprinted from Li et al. (2009a). Copyright 2009, with permission from Elsevier
2.2 Redox Labeling Methods for Electrochemical DNA Sensor
On the basis of the redox reactions of electroactive probes that confined to the gold electrode surface, sensitive electrochemical signaling is readily accomplished (Wang et al. 2008). In the past two decades, various redox labeling strategies have been reported for electrochemical DNA sensors. Redox reporter can be used to label DNA probe by an intercalation or covalent linker. To indirectly investigate the electrochemical performance of DNA SAM, the solution-diffused redox reporters are frequently employed. Therefore, the variations in the electrochemical response reflect the target concentration in the system.
2.2.1 Intercalation-Based Methods
Intercalation is the insertion of a reporter between the adjacent base pairs of a DNA duplex through a noncovalent interaction. The reporters, named intercalators, are mostly aromatic, planar, and polycyclic. Intensively investigated DNA intercalators include methylene blue (MB), Meldola’s blue (MDB), and some transition metal complexes. Typically, thiolated dsDNA with mismatched or matched sequences are first attached on gold electrode and then incubated with redox-active intercalator at micromolar concentration. In this case, the electrochemical reporter can be easily addressable, producing electrochemical signals.
MB, an organic molecule that belongs to the phenothiazine family, is a widely used redox indicator in electrochemical DNA sensors (Liu et al. 2012). It generates different voltammetric signals in the presence of ssDNA or dsDNA. As an indicator of the extent of DNA sensor, the widespread use of MB is owing to the relatively low potential value of its electrochemical reaction. For example, the oxidation potential is around −0.20 V versus Ag/AgCl depending on the experimental conditions used (Loaiza et al. 2007). Thus, it allows minimization of potential electroactive interferences. In comparison with the electrochemical signals obtained in the presence of ssDNA probes, MB exhibits higher electrochemical signals after the hybridization process. These enhanced signals might be attributed to the interaction of MB with G bases, as well as to its intercalation in the dsDNA, if there are at least two G-C base pairs (Bang et al. 2005). To investigate DNA-mediated charge transport, intercalated redox reporters are extensively employed. By using the intercalator, the density or amount of self-assembled DNA on the electrode surface can be accurately examined and ssDNA and dsDNA can be accurately differentiated. In addition, the DNA-mediated electron transfer at long range can be greatly influenced by coupling of intercalators into the p-stack of DNA. The electrochemical signals generated from redox reporters have been proved sensitive to subtle perturbations in the base stacking and DNA sequences. Meanwhile, the mismatches, protein binding, and structural damages all lead to less efficient charge flow within DNA.
Another classic electrochemical indicator, MDB, can intercalate into the dsDNA by the planar phenoxazine ring in its structure (Reid et al. 2002). The electrochemical active center of MDB is not fully enveloped by the bulky DNA when the aromatic ring of MDB intercalates into dsDNA. Thus, it is still available for redox activity, and it can be employed as a hybridization indicator for electrochemical biosensing assays (Pedrero et al. 2011; Radoi and Compagnone 2009).
Transition metal complexes exhibit significant potential and have been demonstrated as the mostly used electroactive indicators for bimolecular analysis. We developed a complex diaquabis [N-(2-pyridinylmethyl) benzamide-2-N, O]-cadmium(II) dinitrate {[CdL2(H2O)2](NO3)2, where L = N-(2-pyridinylmethyl) benzamide}(Zhang et al 2007). The proposed [CdL2]2+ had excellent electrochemical activity and could intercalate into the double helix of dsDNA. Using [CdL2]2+ as an electroactive indicator, the detection of human hepatitis B virus DNA was investigated by differential pulse voltammetry (DPV). The copper (II) complex of Luteolin C30H18CuO12 (CuL2) was demonstrated to intercalate into double helices of the dsDNA (Niu et al. 2009). Using CuL2 as an electroactive indicator, ssDNA fragment of human hepatitis B virus could be selectively detected. A complex of rutin(R) C54H58MnO32 (abbreviated by MnR2) was reported and also used as the electrochemical indicator for the detection of DNA hybridization event (Niu et al. 2008). We summarized the classic transition metal complexes and their interaction to DNA in a review, which is helpful for the reader to understand (Li et al. 2008a).
2.2.2 Covalently Tethered Redox-Active Methods
Using flexible alkyl linkages, various reporters, including MB and ferrocene (Fc), have been tethered covalently to the distal terminus of DNA. By varying the surface density or orientation of DNA SAM, the position of redox reporters can be easily regulated due to the highly restricted mobility of the redox reporters. When the length of alkyl linkages is limited, the only surface-controlled process is the redox reaction of reporters without the consideration of diffusive processes. In this case, the interpretation of obtained results is simplified. It should be noted that only one redox reporter is present on each DNA construct. Thus, it is a powerful tool to examine DNA density on the electrode surface in a 1:1 ratio with the reporter.
The reaction protocol has been successfully developed for covalent tethering of MB and Fc (Willner and Zayats 2007; Gao et al. 2014). The preparation of amino-labeled DNA probes and subsequent immobilization of the redox reporter by the amino functionality to the terminal phosphodiester are often involved. For MB, the NHS-ester of N-(carboxypropyl)methylene blue has been the gold-standard route. For Fc, N-hydroxysuccinimide (NHS) ester of ferrocenecarboxylic acid is typically used. It should be prepared freshly right before tethering to the amino-labeled DNA probe because the activated ester is unstable.
2.2.3 Solution-Diffused Methods
Owing to the strong electrostatic interactions, the common negatively charged redox reporters, which are freely diffusing in aqueous solution, cannot reach the DNA-immobilized electrode surface. For instance, the electronegative DNA phosphate backbone can electrostatically repel the negatively charged [Fe(CN)6]3−/4− ions. Thus, the redox reporters cannot get into close proximity with the electrode surface (Gao et al. 2015). This results in a high electron transfer resistance, which can be monitored through recording electrochemical impedance spectroscopy (EIS). However, the diffusion of [Fe(CN)6]3−/4− ions toward the sensor surface can be improved upon treatment or binding to positively charged proteins, which leads to a decrease in the electron transfer resistance (Zhou et al. 2012).
[Ru(NH3)6]3+/2+ is another solution-based redox system and often employed as a stable and accurate reporter to determine the DNA density on the electrode surface (Gao et al. 2013). The [Ru(NH3)6]3+ ions diffuse to the sensor surface and electrostatically bind to the electronegative DNA backbone. Their characteristic surface redox reaction can be adapted to discriminate the contribution of solution-diffused redox centers: DNA dehybridization is detected by a reduced signal from the electrostatically bound [Ru(NH3)6]3+. Besides, the surface densities of both double- and single-stranded DNA can be detected by integration of the reduction peak of [Ru(NH3)6]3+ to [Ru(NH3)6]2+. Following the treatment of [Ru(NH3)6]3+ with a concentration of 5.0 mM or lower ionic strength, the surface concentration of [Ru(NH3)6]3+, ΓRu, can be calculated by the following equation (Steel et al. 1998):
where Q represents the charge obtained by integrating the reduction peak area of surface-bound [Ru(NH3)6]3+, n is the number of electrons involved in the redox reaction (which is 1), F represents Faraday’s constant, and A is the electrode area. This equation assumes as follows: (1) The interaction between [Ru(NH3)6]3+ is purely electrostatic; (2) full saturation of the DNA-modified surface with [Ru(NH3)6]3+ occurs; (3) there is complete exchange of [Ru(NH3)6]3+ with compensation by Na+ ions; (4) every phosphate molecule is accessible for electrostatic binding to the redox cations.
Under saturation conditions (i.e., where the highest possible concentration of surface-bound [Ru(NH3)6]3+, ΓRu, was achieved), the measured ΓRu value can be converted to the surface density of DNA (Cheng et al. 2007), ΓDNA, using the following equation:
where z represents the valence of the redox cation and m is the number of nucleotides in the DNA.
3 Electrochemical Sensors Based on Nucleic Acid Amplification
The signal amplification methods can be divided into two groups based on whether the temperature changes during reactions (Patterson et al. 2013). One category is thermocycling amplification methods. For example, polymerase chain reaction (PCR), which is recognized as a traditional signal amplification strategy for sensitive and selective detection of nucleic acids at extremely low concentrations. The other group called isothermal amplification method that has been established since the early 1990s.
3.1 Electrochemical Sensors Based on Thermocycling PCR Amplification Systems
The electrochemical real-time method is first reported by Hsing and co-workers using solid-phase PCR with a simple experimental setup (Yeung et al. 2006) (Fig. 7.2a). To extend electrode-bound primers, a reaction mix containing ferrocene-modified dUTP is employed. Thus, any amplification is measurable as an increase in ferrocene redox current. They further implemented this strategy within an integrated PCR chip with microfabricated electrodes and realized a detection limit as low as 3 × 103 copies and a 1000-fold dynamic range (Yeung et al. 2008). Although pioneering, this method still has several disadvantages relative to standard optical approaches. For example, compared to standard optical approaches which can realize low detection limits of three DNA copies, its sensitivity and dynamic range are poorly presented (Bustin et al. 2009). This discrepancy presumably arises because of the inefficiency of solid-phase polymerization (Pemov et al. 2005). The degradation or loss of primers during electrochemical measurement indicates a second limitation. In this case, they were only able to measure at between four and six amplification cycles, thus reduces the precision of detection.
a The electrochemical real-time PCR system using surface-bound primers that are elongated by amplification. Reprinted with the permission from Yeung et al. (2006). Copyright 2006 American Chemical Society. b A direct method for the electrochemical detection of PCR products measures electroactive nucleotides using redox mediators. Reprinted with the permission from Defever et al. (2009). Copyright 2009 American Chemical Society. c The electrochemical eTaq PCR platform uses a redox label-modified probe. Reprinted from Luo et al. (2011). Copyright 2011, with permission from Elsevier. d The use of electroactive redox reporters that intercalate in dsDNA is a direct electrochemical method for monitoring PCR in real time. Reprinted from Patterson et al. (2013). Copyright 2013, with permission from Elsevier
The development of second-generation techniques which are more analogous to current optical methods is motivated by the limited sensitivity and precision of the above generation electrochemical PCR. The first-generation techniques are reported to monitor amplifications by testing either the loss of electroactive intercalating reporters or the consumption of electroactive nucleobases upon binding or incorporating into dsDNA.
Defever and co-workers, the first group to explore solution-phase electrochemical PCR, used electrochemical method to directly monitor the concentration of free nucleotides in solution as they are consumed during amplification (Defever et al. 2009) (Fig. 7.2b). They first developed the electrochemical approach capable of monitoring PCR at each cycle, significantly improving the system accuracy. The redox reactivity of 7-deaza-dGTP, the guanidine analogue, was also investigated. They indicated that when its base incorporates into a dsDNA, it can be effectively reduced. The detection limits as low as 283 bp sequence from human cytomegalovirus (HCMV) and 2300 bp sequence from the bacterial genome of Achromobacter xylosoxidans were obtained with a wide dynamic range over eight orders of magnitude.
Inspired by the TaqMan method, an electrochemical analog, which named ‘eTaq’ and supported sequence-specific, electrochemical qPCR, was reported by Luo and co-workers (Luo et al. 2011) (Fig. 7.2c). In this method, eTaq probe is modified with an electroactive reporter, such as MB or Fc. The diffusion of the reporter to a negatively charged indium tin oxide electrode was decreased due to the high molecular weight and negative charge. Upon hydrolysis into less highly charged, lower-molecular-weight mononucleotides diffusion of the reporter to the electrode surface is increased, improving the observed electrochemical signal. In consequence, they employed this method to detect a 137 bp segment of human genomic DNA with low background and excellent signal-to-noise ratio. However, the authors stated that the precision, dynamic range, and detection limit of the strategy have not been well investigated. For example, the detection of a single, relatively high target concentration about 1.6 × 106 copies is presented. And, this method has not yet been investigated in the context of a fully integrated truly continuous system, which is the second concern. Manual pipetting was used to transfer the amplicons from the solution onto the electrode chip at specific cycles, although their preliminary results showed feasibility.
To conduct optical qPCR, DNA-intercalating fluorescent dyes are usually employed. Thus, in electrochemical qPCR systems, the employment of electroactive intercalators has not surprisingly been discovered (Fig. 7.2d) (Patterson et al. 2013; Wilhelm and Pingoud 2003). When a low-molecular-weight redox reporter attaches to a high-molecular-weight amplicon, the diffusivity is reduced, which is commonly explored and demonstrated. The reduced diffusivity decreases the efficiency with which the reporter approaches the electrode surface and transfers an electron. Several intercalating reporters have been well exploited, to further improve this strategy, by Defever and co-workers. And, they showed that Os[(bpy)2DPPZ]2+ (DPPZ is dipyrido[3,2-a:2′,3′-c]phena-zine) displayed the critical attributes of good thermal stability, strong binding efficiency to double-stranded amplicons, low inhibition efficiency for PCR, and high sensitivity. To examine PCR at each cycle by using this reporter, quantitative detection of the same target from HCMV was processed. They finally achieved a low detection limit of 103 copies with a wide dynamic range over six orders of magnitude (Defever et al. 2011). An intercalation-based microfluidic qPCR system has been developed by Won and co-workers. They realized accurate detection of initial copy numbers of Chlamydia trachomatis DNA by a homemade microelectrode-patterned glass chip using MB as an intercalating reporter (Won et al. 2011). A low detection limit of 103 copies target DNA was obtained with a dynamic range spanning four orders of magnitude. In support of the proposed signaling principle, Won et al. have demonstrated that the transport of MB to the electrode decreases by more than an order of magnitude upon intercalation into a 612 bp dsDNA.
3.2 Electrochemical Sensor Based on Isothermal Amplification
Although PCR presents significant biosensing performance and is by far the most well-established approach to amplify nucleic acids, it still has many limitations, including high cost, easy contamination, and time-consuming (Pedrero et al. 2011). In addition, the requirement of a sophisticated thermal cycler greatly restricts the applications of PCR in the point-of-care measurements and resource-limited settings. Alternatively, isothermal amplification that combines with electrochemical methods indicates an extensive attraction and great opportunity for biosensing applications.
3.2.1 Quantitative Electrochemical Methods Using Helicase-Dependent Amplification
The mechanism of helicase-dependent amplification (HDA) is similar to that of PCR, accelerating adaptation in current biosensing approaches. Generally, three steps are included in HAD reaction that are template separation, primer hybridization, and primer extension (Qi et al. 2018). The first step is template separation. DNA helicase is used to separate dsDNAs, and each of them is covered by ssDNA-binding proteins. For the second step, two specific primers attach to each ssDNA, respectively, and finally extend their sequence to form the corresponding dsDNAs in the presence of DNA polymerase. By taking the newly generated dsDNAs as the substrates, the next cycle of HDA reaction starts with the use of DNA helicase. As a result, the continuous cycle processes lead to an exponential amplification of the target. To dissociate the two DNA strands, instead of using high temperatures, a thermostable DNA helicase is employed in this method, which processes this step enzymatically (Deng and Gao 2015; Barreda-Garcia et al. 2015; Vincent et al. 2004).
Taking advantage of the similarities between HDA and PCR, Kivlehan et al. proposed Os[(bpy)2DPPZ]2+ as an intercalating redox reporter to establish the first isothermal real-time electrochemical system (Kivlehan et al. 2011). Using a disposable microplate with embedded electrodes, continuous measurements were conducted every 38 s throughout the reaction, achieving excellent temporal resolution. However, they mention that HAD is inhibited by the proposed electrochemical reporter, and thus almost twice the time is needed for the determination of the same initial targets number as their optical method (e.g., ~28 min for fluorescence versus ~45 min for electrochemistry at 105 copies).
In addition to homogeneous detection, Lobo-Castanon et al. proposed an HDA-based heterogenous-phase strategy by determining the unpurified HDA amplicons to electrochemical detect transgene extracted from the Cauliflower Mosaic Virus 35 S Promoter (CaMV35S) (Moura-Melo et al. 2015) (Fig. 7.3). The HDA products attached to the signaling probes that were modified with fluorescein isothiocyanate (FITC) and further bound to the thiolated P35S-capture probes. To combine with the FITC, the anti-FITC-peroxidase (POD) fragments were added. By catalyzing the oxidation of 3,3′,5,5′-tetramethylbenzidine (TMB) under the condition of H2O2, the electrochemical signal was detected, which performed highly sensitive and selective detection of target CaMV35S with a low detection limit down to 30 copies.
HDA-based heterogenous-phase electrochemical biosensor for transgene detection. Reprinted with the permission from Moura-Melo et al. (2015). Copyright 2015 American Chemical Society
Despite the merits of HDA, optimization of experimental conditions including primer sets and buffer composition, to ensure the HDA reaction can be conducted effectively, is often ineluctable. However, PCR experiments are often used for the optimization. As a result, more investigations should be carried out to achieve the whole potential of HDA. Moreover, the development of helicases and ssDNA-binding proteins should be focused in future researches. Finally, the improvement of the efficiency of the existing helicases and ssDNA-binding proteins should not be ignored, which are important to the identification of the rate-limiting step in HDA.
3.2.2 Quantitative Electrochemical Sensor Using Loop-Mediated Isothermal Amplification
Loop-mediated isothermal amplification (LAMP) is another widely used isothermal strategy to be integrated into an electrochemical platform and employs strand displacing polymerase and a loop-forming primers to produce large, concatemeric amplicons comprising multiple copies of the amplification analytes (Notomi et al. 2000). LAMP has demonstrated to be an effective and alternative protocol to PCR because of its significant amplification yields and specificity. For instance, a high amplicon yield can be obtained by LAMP method (>500 ng/ml, even ~100 times greater than PCR) (Nagamine et al. 2002b). Similarly, the specificity of LAMP is remarkable greater than that of PCR, because it needs that four to six distinct primer probes simultaneously recognize six to eight different regions within the target (Nagamine et al. 2002a). Despite the complexity, commercially available master mixes and readily accessible primer design software have greatly decreased the overheads and cost of LAMP, contributing to its increasing popularity (Mori and Notomi 2009). Given these strengths, it is reasonable that, in recent years, LAMP has been integrated with electrochemical method to develop a real-time biosensing platform combining the very real benefits of both.
LAMP has recently been used to combine with electrochemical method for the detection of various analytes. The electrochemical detection is conducted by detecting redox molecules in the absence or presence of amplicons or by immobilizing redox-labeled LAMP amplicons on the surface of electrode. The LAMP amplicons can be measured after the amplification reactions or be detected in real time. The expected electrochemical signals can be monitored by square wave voltammetry (SWV), linear sweep voltammetry (LSV), differential pulse voltammetry (DPV), and the change of conductivity.
Nagatani and co-workers first presented a semi-real-time electrochemical monitoring approach based on the reverse transcription LAMP using a screen-printed electrode for the detection of influenza virus RNA (Nagatani et al. 2011). They used MB as an electroactive DNA intercalator to combine with amplified DNA, and SWV was used as the electrochemical signal in the RT-LAMP biosensing system. Based on the RT-LAMP, amplification process of DNA was successfully monitored by recording and analyzing the MB electrochemical signal with only one screen-printed working electrode. The obtained electrochemical current was greatly related to the concentration of target RNA and the extent of DNA amplification. This approach avoids potential cross-contamination because both the amplification and the detection processes are possible in a single tube, and laborious probe immobilization is not needed. Moreover, this approach offers a novel aspect to fabricate sensitive electrochemical handheld biosensors.
Recently, Marchal and co-workers reported a sensitive and selective electrochemical LAMP approach to determinate single DNA copies by using one nonintercalating reporter (Ru(NH3) 3+6 ) and three intercalating redox reporters (MB, [Os(bpy)2dppz]2+, and PhP), respectively. Compared with the traditional PCR and fluorescence LAMP assays, they carefully studied the sensing performance that monitored by each of the four electrochemical reporters (Martin et al. 2016). More intercalating redox reporters were used, with the progress of the reaction, to combine with the LAMP amplicons. Besides, in the presence of Ru(NH3) 3+6 , it prefers to bind with the by-product pyrophosphate anion, which can be released significantly in the reaction. The two distinct combination assays between nonintercalating and intercalating probes with LAMP products lead to a reduced current along with the amplification. Thus, the proposed approach provided an accurate and reliable merit for real-time electrochemical LAMP.
Taking a different approach, a microfluidic multiplex electrochemical LAMP (named μME-LAMP) system was developed by Kong and colleagues for the sensitive and real-time detection of multiple bacteria (Luo et al. 2014) (Fig. 7.4a). Multiplex assay was achieved with the addition of different samples into the isolated electrochemical microchambers. A lower electrochemical current was generated after performing the LAMP reaction than that generated before LAMP, owing to the intercalation of MB into the DNA duplex and the slow diffusion of MB to the electrode surface. The proposed μME-LAMP chip realized the detection limits as low as 17, 28, and 16 copies/µL for Haemophilus influenza, Mycobacterium tuberculosis, and Klebsiella pneumonia, respectively.
a Schematic of the principle of the microfluidic multiplex electrochemical LAMP (named μME-LAMP) system for real-time quantitative multiple bacteria. Reprinted from Luo et al. (2014). Copyright 2014, with permission from Elsevier. b MEQ-LAMP system for pathogenic DNA detection. Reproduced from Hsieh et al. (2012) by permission of John Wiley & Sons Ltd.
Taken advantages of LAMP, a microfluidic electrochemical quantitative (MEQ)-LAMP platform is developed for simple, sensitive, rapid, and quantitative determination of Salmonella genomic DNA at the point of care by Hsieh and co-workers. DNA amplification was electrochemically processed in real time utilizing MB to record measurements at 1-min intervals within a monolithic microfluidic device. By a single-step process, they realized a low detection limit as few as 16 copies of target sequence in less than an hour (Hsieh et al. 2012) (Fig. 7.4b). Compared with most other electrochemical detection systems, this method exhibits better sensitivity and faster response speed. As the sensor surface can be simply modified to contain multiple chambers for parallel determination of multiple targets, the proposed MEQ-LAMP strategy is readily compatible with multiplexing. The significant potential and the performance for expanded capability might inspire researchers to envision the MEQ-LAMP platform as a viable protocol for biosensing at the point of care.
Accurate calibration and control of temperature are demanded for quantitative detection because the activity of polymerase in LAMP is greatly influent by temperature. Therefore, create stable LAMP enzymes and other LAMP biochemical components is in great demand. In addition, nonenzymatic LAMP biosensing methods might also be achieved that could be practical and potential for remote settings. LAMP, as a powerful tool, has significant promise in the establishment of practical approaches in future.
3.2.3 Quantitative Electrochemical Sensor Using Rolling Circle Amplification
Rolling circle amplification (RCA), a sensitive and selective isothermal amplification technique, has been broadly employed in various domains owing to its high efficiency and simplicity (Gao et al. 2018). Various electrochemical RCA (ERCA) strategies have been reported to achieve ultrahigh sensitivity, such as hyperbranched rolling circle amplification (HRCA) and padlock exponential rolling circle amplification (P-ERCA). Lizardi and co-workers first proposed HRCA for the determination of single base mismatch in a small quantity of human genomic DNA (Lizardi 1998). In addition, HRCA is simplicity, less time-consuming, ultrahigh sensitivity, and cost-effective performance. Compared with conventional PCR assay, HRCA has the advantage that it can be carried out at constant temperature without the need of complex thermal cyclers. During the past decades, HRCA has been extensively used for the detection of viral RNA, DNA, metal ions, small molecules, and proteins.
In 2013, Lin et al. developed a label-free electrochemical aptasensor for the determination of platelet-derived growth factor B-chain (PDGF-BB) based on HRCA with high specificity and sensitivity (Wang et al. 2013) (Fig. 7.5a). First, cDNA was attached on the gold electrode surface by Au–S bond and it could partially hybridize with aptamer probe to form the DNA duplex. When PDGF-BB was added, it prefers to hybridize with the aptamer probe, removing the aptamer probe from the electrode surface. In this case, the cDNA presented free state and able to hybridize with a padlock probe. The padlock probe contains a small gap, and it could generate a circular padlock probe in the presence of E. coli DNA ligase. Primer 1 was hybridized with the circular padlock probe, leading to the RCA reaction by polymerase to generate long ssDNA with repeated region, providing as the templates to hybridize with primer 2. Therefore, primer 2 extended immediately forward with the addition of DNA polymerase. In this case, a number of dsDNA were produced on the electrode surface. Finally, the electrochemical indicator, MB, was used to insert into the dsDNA, generating significant electrochemical signals that were proportional to the amount of target. Thus, a label-free electrochemical aptasensor based on HRCA amplification was established, realizing a detection limit as low as 1.6 fM of PDGF-BB and high selectivity against other interferents, such as thrombin, lysozyme, and human serum albumin (HSA).
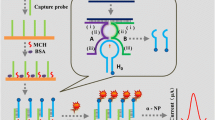
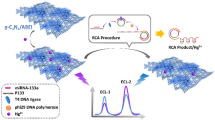
a HRCA-based electrochemical biosensor for PDGF-BB detection. Reproduced from Wang et al. (2013) by permission of The Royal Society of Chemistry. b Electrochemical DNA biosensor based on the cascade signal amplification of hairpin-mediated CSD and HRCA. Reprinted from Li et al. (2016). Copyright 2016, with permission from Elsevier. c Schematic illustration of P-ERCA assay and CoFe2O4 MNPs assisted no substrate nanoelectrocatalysis for miR-21 detection. Reprinted from Yu et al. (2017). Copyright 2017, with permission from Elsevier
Yi and colleagues created an electrochemical biosensor for sensitive determination of DNA fragments by integrating HRCA with hairpin-mediated circular strand displacement polymerization (CSD) (Li et al. 2016) (Fig. 7.5b). The hairpin probe that immobilized on the gold electrode surface can be opened to form the dsDNA, when target DNA was added. Then, the unfolded stem binds with biotin-labeled primer probe, resulting in the DNA strand polymerization with the aid of dNTPs and DNA polymerase. Therefore, the target DNA was released to trigger the next cycle of hybridization process with hairpin probes, resulting in the accumulation of biotin-labeled DNAs. Given the specific and strong streptavidin–biotin interaction, massive streptavidin molecules were attached on gold surface. With the addition of HRCA primers and circular templates, the HRCA reaction was occurred. Consequently, massive dsDNA with various lengths were generated. Finally, the biotin-labeled probes are selectively attached to the dsDNA, which further linked to the streptavidin–alkaline phosphatase (ST-AP). The recorded electrochemical signal that catalyzed by alkaline phosphatase (AP) was produced from the irreversible conversion of α-naphthyl phosphate (α-NP) to electroactive products, which was measured by DPV. The proposed strategy realized an ultrahigh sensitivity for DNA detection with a low detection limit down to 8.9 aM.
Taking a different approach, P-ERCA has been recently established. Yu and colleagues developed a P-ERCA method for electrochemical determination of microRNA-21 (miR-21) using magnetic nanoparticles (MNPs) for nonsubstrate nanoelectrocatalysis (Yu et al. 2017) (Fig. 7.5c). The exponential growth of P-ERCA triggers was demonstrated during the continuous process of polymerization and cleavage. Meanwhile, the electrode surface was modified with capture probe I (CP I), which could hybridize with triggers and further produce the immobilization of CP II/Au@CoFe2O4/Tb-Gra composites on the surface. Finally, the expected signal was electrochemically measured from the MNPs-aided nonsubstrate nanoelectrocatalysis. The proposed method showed an excellent sensitivity, realizing a low detection limit of 0.3 fM for miR-21 detection. Taken together, the electrochemical P-ERCA approaches are universal and suitable for the detection of various targets, such as proteins and nucleic acids. The method has the advantages of low cost, rapid response, high sensitivity, simplicity, and specificity, which holds significant promise in the development of clinical diagnostics, environmental monitoring, and food safety.
Despite its significant performance, RCA-based electrochemical devices, or diagnostic protocols have yet to reach the commercial market. To date, the RCA amplification kit has been created only for research use, such as the Illustra TempliPhi DNA amplification kits from GE HealthCare (Buckinghamshire, UK). Developing RCA-based electrochemical sensors is of importance for point-of-care testing and holds great business opportunities in future.
3.2.4 Quantitative Electrochemical Sensor Using Nanomaterial-Assisted Amplification
To meet the growing requirements for ultrasensitive biosensing, scientists have established various strategies to improve the response of the electrochemical DNA sensor by modifying with functional nanomaterials. With the development of nanotechnology, significant attention has been paid to various nanomaterials, such as MNPs, quantum dots (QDs), metal nanoparticles, carbon-based nanomaterials, and polymeric NPs. Owing to the remarkable biological compatibility, chemical stability, nontoxicity, high surface area, remarkable conductivity, and catalytic activity, the introduction of nanomaterials has significantly enhanced the biosensing performance to obtain the amplified electrochemical signal and the stabilized recognition probes.
In recent years, many reviews on functional nanomaterials and their biosensing applications have been published. Herein, we simply concentrate on the recent development of nanomaterial-based signal amplification strategies, especially gold nanoparticle (AuNPs), in ultrasensitive DNA-based electrochemical biosensing. In 2008, Zhang and co-workers fabricated an electrochemical sensor based on a structure-switching aptamer and reporter DNA loaded on AuNPs for the detection of adenosine (Fig. 7.6a) (Zhang et al. 2008). The electroactive complex, [Ru(NH3)6]3+, acts as a signaling transducer. As a single AuNP can be loaded with hundreds of reporter DNA strands, this provides a great amplification of electrochemical signal for the detection of adenosine. Based on this method, the adenosine could be determined with a detection limit as low as 0.18 nM. Next, Hu et al. developed a sensitive electrochemical DNA sensor based on multifunctional encoded AuNPs and nanoporous gold (NPG) electrode (Fig. 7.6b) (Hu et al. 2008). Different from the above-mentioned AuNPs with one kind of DNA probe, the AuNPs we used here labeled with two kinds of DNA biobarcode. One is complementary to the target DNA, while the other is not, decreasing the cross-reaction of targets on the same AuNPs. Given the multifunctional encoded AuNPs and dual-amplification effects of the electrode, the proposed biosensor achieved a low detection limit of 28 aM DNA.
a Schematic representation of the adenosine detection based on AuNPs. Reprinted with the permission from Zhang et al. (2008). Copyright 2008 American Chemical Society. b Chronocoulometry detection of DNA hybridization by two steps of amplification. Reprinted with the permission from Hu et al. (2008). Copyright 2008 American Chemical Society
Taken the advantages of high loading capacity, programmable ability, and signal amplification ability, AuNPs were widely used for the fabrication of electrochemical sensors to detect various targets, such as DNA (Zhong et al. 2009; Hu et al. 2009; Li et al. 2009b), miRNA (Song et al. 2016), thrombin (Zhang et al. 2009a), human α-fetoprotein (Ding et al. 2009), and cancer cell (Ding et al. 2010). As multiple DNA can be attached to the gold surface to construct biobarcode AuNPs, amplified electrical biobarcode assay for one-pot detection of multiple targets was realized (Li et al. 2010a, b; Zhang et al. 2009b, 2011). In addition, other nanomaterials, such as functionalized metal–organic framework, also show significant sensing performance and promising for the detection of a wide range of targets (Shi et al. 2017).
3.2.5 Quantitative Electrochemical Sensor Using Enzyme-Free Nucleic Acid Amplification
Toehold-mediated strand displacement (TMSD) has proved to be a useful strategy for the design of enzyme-free isothermal amplification systems to selectively and sensitively detect a broad range of targets (Zhang and Seelig 2011). Firstly, a long ssDNA, in a typical TMSD, hybridizes to the toehold area of dsDNA, initiating the strand displacement rapidly and producing a new dsDNA with all complementary base pairs (Genot et al. 2011; Yurke et al. 2000). Hybridization chain reaction (HCR) is a prominent example of TMSD. A cascade of hybridization events, in a typical HCR, was occurred between alternating two hairpin probes induced by an initiator, producing a long-nicked dsDNA (Dirks and Pierce 2004). Not only linear HCR, branched HCR strategies for exponential amplification have also been constructed (Bi et al. 2017). For example, a TMSD-based nonlinear HCR designs that usually occur in an exponential manner are reported by Hsing and colleagues (Xuan and Hsing 2014) (Fig. 7.7a). Firstly, they added trigger into the system, which was used to hybridize with the toehold of Substrate-A and triggering the first TMSD reaction. In this case, the black probe could be displaced to open the first functional loop. A new toehold of the displaced strand was exposed, hybridizing with the Assistant-A and extending forward to open the second loop. Thus, By-product-A was released. Substrate-A has two exposed sequences, and it could further hybridize with the toehold of Substrate-B. Consequently, the dissociation of By-product-B and the hybridization of Assistant-B were proceeded. In this case, the product, consisting two ssDNA that are the same as the trigger DNA, is exposed and continually triggers TMSD reactions, which therefore leading to the exponentially growth of the branched DNA nanostructure.
a Enzyme-free exponential amplification method on the basis of TMSD-mediated nonlinear HCR. Reprinted with the permission from Xuan and Hsing (2014). Copyright 2014 American Chemical Society. b Principle of enzyme-free and immobilization-free exponential amplification assay based on nonlinear HCR for electrochemical detection of nucleic acids. Reprinted with the permission from Xuan et al. (2015). Copyright 2015 American Chemical Society. c A enzyme-free and solid-phase exponential amplification system based on nonlinear HCR for the detection of ATP. Reprinted from Ding et al. (2017), with kind permission from Springer Science + Business Media
An enzyme-free and immobilization-free electrochemical biosensing method based on nonlinear HCR system has been established for nucleic acid detection (Xuan et al. 2015) (Fig. 7.7b). The PNA probes labeled with ferrocene (Fc-PNAs) freely dispersed in solution in the absence of target, which can attach to the surface of negatively charged indium tin oxide (ITO) electrode and generate a significant electrochemical current. In the presence of the target, the nonlinear HCR was triggered and the branched DNA/PNA nanostructures were exponentially generated, and it hardly in close to the ITO electrode surface. Therefore, the reduction of the electrochemical signal was significantly amplified. It is worth noting that Fc-PNA was used to hybridize onto the dendritic DNA nanostructures with strong binding strength and high hybridization kinetics efficiency between DNA and PNA. The proposed method demonstrated a high sensitivity with a low detection limit of 100 fM.
Ding et al. designed an electrochemical biosensor, unlike the immobilization-free strategy that stated above, by attaching the capture probe onto the surface of gold electrode to detect ATP (Ding et al. 2017) (Fig. 7.7c). With the addition of ATP, it preferable to attach with its aptamer, resulting in the release of trigger DNA which could hybridize with the toehold of Substrate 1. Thus, the first TMSD reaction was triggered. Helper 1 was designed to hybridize with the region iii of Substrate 1-B and extended forward to generate the By-product-1. Substrate 1-A strand was designed with two identical sequences and combined with the two toeholds of Substrates 2, respectively. Then, By-product 2 was dissociated with the assistant of Helper 2. Finally, the product, consisting two target regions, was produced. Another cycle of reaction could be triggered, leading to the exponentially growth of DNA nanostructures. The resultant electrode surface was immobilized with DNA dendrimers that modified with terminal biotin by capture probe, which linked to ST-AP to produce significant electrochemical signal by the irreversible conversion of α-NP to electroactive products catalyzed by AP. This assay realized a detection limit as low as 5.8 nM of ATP.
Enzyme-free methods, compared with enzyme-assisted isothermal amplification methods, have the merit of simple and rapid response because of the inherent nonenzyme property (Gao et al. 2016). Moreover, enzyme-free strategies have excellent structural versatility, amplification efficiency, and controllable kinetics, which thus have significant promise in the application areas of biosensing, bioimaging, and biomedicine.
4 Conclusion
In this chapter, a series of nucleic acid amplification methods have been summarized and their electrochemical applications in fabrication of sensitive and selective biosensors during the recent years have been introduced. Given the significant merits of high amplification efficiency and biosensing sensitivity, nucleic acid amplification has been proved as a powerful protocol for electrochemical bioassays. With the development of bioanalytical chemistry, including DNAzymes and aptamers, electrochemical biosensors based on nucleic acid amplification techniques have been expanded to detect a wide range of targets, such as proteins, nucleic acids, ions, and small biomolecules. Furthermore, the combination of nucleic acid amplification with portable devices or microsystems has facilitated the development of clinical diagnosis with ultrahigh selectivity and sensitivity. In this research field, initial research was concentrated on the preparation of biosensing platforms that can perform concurrent and real-time electrochemical detection during the amplification process. In recent years, scientists have already developed promising electrochemical systems that deliver quantification capabilities and sensitivity that potentially rival bench-top optical systems. To date, the approach of integrating nucleic acid amplification with intercalating electroactive molecules within microfluidic electrochemical devices holds great potential and promising for practical applications. With the development of detection mechanisms and amplification techniques, coupled with straight-forward sample preparation and multiplexed assays, electrochemical strategies might bring new power for real-time detection to the clinical diagnostic at point-of-care in future (Patterson et al. 2013).
References
Bang GS, Cho S, Kim BG (2005) A novel electrochemical detection method for aptamer biosensors. Biosens Bioelectron 21:863–870
Barreda-Garcia S, Gonzalez-Alvarez MJ, de-Los-Santos-Alvarez N et al (2015) Attomolar quantitation of Mycobacterium tuberculosis by asymmetric helicase-dependent isothermal DNA-amplification and electrochemical detection. Biosens Bioelectron 68:122–128
Bi S, Yue S, Zhang S (2017) Hybridization chain reaction: a versatile molecular tool for biosensing, bioimaging, and biomedicine. Chem Soc Rev 46:4281–4298
Bustin SA, Benes V, Garson JA et al (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622
Cheng AK, Ge B, Yu HZ (2007) Aptamer-based biosensors for label-free voltammetric detection of lysozyme. Anal Chem 79:5158–5164
Defever T, Druet M, Rochelet-Dequaire M et al (2009) Real-time electrochemical monitoring of the polymerase chain reaction by mediated redox catalysis. J Am Chem Soc 131:11433–11441
Defever T, Druet M, Evrard D et al (2011) Real-time electrochemical PCR with a DNA intercalating redox probe. Anal Chem 83:1815–1821
Deng H, Gao Z (2015) Bioanalytical applications of isothermal nucleic acid amplification techniques. Anal Chim Acta 853:30–45
Ding C, Zhang Q, Zhang S (2009) An electrochemical immunoassay for protein based on bio bar code method. Biosens Bioelectron 24:2434–2440
Ding C, Ge Y, Zhang S (2010) Electrochemical and electrochemiluminescence determination of cancer cells based on aptamers and magnetic beads. Chem Eur J 16:10707–10714
Ding X, Wang Y, Cheng W et al (2017) Aptamer based electrochemical adenosine triphosphate assay based on a target-induced dendritic DNA nanoassembly. Microchim Acta 184:431–438
Dirks RM, Pierce NA (2004) Triggered amplification by hybridization chain reaction. Proc Natl Acad Sci USA 101:15275–15278
Gao ZF, Gao JB, Zhou LY et al (2013) Rapid assembly of ssDNA on gold electrode surfaces at low pH and high salt concentration conditions. RSC Adv 3:12334–12340
Gao ZF, Ling Y, Lu L et al (2014) Detection of single-nucleotide polymorphisms using an ON-OFF switching of regenerated biosensor based on a locked nucleic acid-integrated and toehold-mediated strand displacement reaction. Anal Chem 86:2543–2548
Gao ZF, Chen DM, Lei JL et al (2015) A regenerated electrochemical biosensor for label-free detection of glucose and urea based on conformational switch of i-motif oligonucleotide probe. Anal Chim Acta 897:10–16
Gao ZF, Huang YL, Ren W et al (2016) Guanine nanowire based amplification strategy: enzyme-free biosensing of nucleic acids and proteins. Biosens Bioelectron 78:351–357
Gao ZF, Liu R, Wang J et al (2018) Controlling droplet motion on an organogel surface by tuning the chain length of DNA and its biosensing application. Chem 4(12):2929–2943
Genot AJ, Zhang DY, Bath J et al (2011) Remote toehold: a mechanism for flexible control of DNA hybridization kinetics. J Am Chem Soc 133:2177–2182
Gorodetsky AA, Buzzeo MC, Barton JK (2008) DNA-mediated electrochemistry. Bioconjug Chem 19:2285–2296
Hsieh K, Patterson AS, Ferguson BS et al (2012) Rapid, sensitive, and quantitative detection of pathogenic DNA at the point of care through microfluidic electrochemical quantitative loop-mediated isothermal amplification. Angew Chem Int Ed 51:4896–4900
Hu K, Lan D, Li X et al (2008) Electrochemical DNA biosensor based on nanoporous gold electrode and multifunctional encoded DNA-Au bio-bar codes. Anal Chem 80:9124–9130
Hu K, Liu P, Ye S et al (2009) Ultrasensitive electrochemical detection of DNA based on PbS nanoparticle tags and nanoporous gold electrode. Biosens Bioelectron 24:3113–3119
Kivlehan F, Mavre F, Talini L et al (2011) Real-time electrochemical monitoring of isothermal helicase-dependent amplification of nucleic acids. Analyst 136:3635–3642
Lai RY, Seferos DS, Heeger AJ et al (2006) Comparison of the signaling and stability of electrochemical DNA sensors fabricated from 6- or 11-carbon self-assembled monolayers. Langmuir 22:10796–10800
Li F, Chen W, Tang C et al (2008a) Recent development of interaction of transition metal complexes with DNA based on biosensor and its applications. Talanta 77:1–8
Li F, Chen W, Zhang S (2008b) Development of DNA electrochemical biosensor based on covalent immobilization of probe DNA by direct coupling of sol-gel and self-assembly technologies. Biosens Bioelectron 24:787–792
Li X, Xia J, Zhang S (2008c) Label-free detection of DNA hybridization based on poly(indole-5-carboxylic acid) conducting polymer. Anal Chim Acta 622:104–110
Li F, Chen W, Dong P et al (2009a) A simple strategy of probe DNA immobilization by diazotization-coupling on self-assembled 4-aminothiophenol for DNA electrochemical biosensor. Biosens Bioelectron 24:2160–2164
Li G, Li X, Wan J et al (2009b) Dendrimers-based DNA biosensors for highly sensitive electrochemical detection of DNA hybridization using reporter probe DNA modified with Au nanoparticles. Biosens Bioelectron 24:3281–3287
Li X, Liu J, Zhang S (2010a) Electrochemical analysis of two analytes based on a dual-functional aptamer DNA sequence. Chem Commun 46:595–597
Li X, Xia J, Li W et al (2010b) Multianalyte electrochemical biosensor based on aptamer- and nanoparticle-integrated bio-barcode amplification. Chem Asian J 5:294–300
Li X, Guo J, Zhai Q et al (2016) Ultrasensitive electrochemical biosensor for specific detection of DNA based on molecular beacon mediated circular strand displacement polymerization and hyperbranched rolling circle amplification. Anal Chim Acta 934:52–58
Liu A, Wang K, Weng S et al (2012) Development of electrochemical DNA biosensors. Trends Anal Chem 37:101–111
Liu J, Cui M, Niu L et al (2016) Enhanced peroxidase-like properties of graphene-hemin-composite decorated with Au nanoflowers as electrochemical aptamer biosensor for the detection of K562 leukemia cancer cells. Chem Eur J 22:18001–18008
Lizardi PM, Huang X, Zhu Z et al (1998) Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat Genet 19:225–232
Loaiza OA, Campuzano S, Pedrero M et al (2007) DNA sensor based on an Escherichia coli lac Z gene probe immobilization at self-assembled monolayers-modified gold electrodes. Talanta 73:838–844
Luo X, Xuan F, Hsing IM (2011) Real time electrochemical monitoring of PCR amplicons using electroactive hydrolysis probe. Electrochem Commun 13:742–745
Luo J, Fang X, Ye D et al (2014) A real-time microfluidic multiplex electrochemical loop-mediated isothermal amplification chip for differentiating bacteria. Biosens Bioelectron 60:84–91
Martin A, Grant KB, Stressmann F et al (2016) Ultimate single-copy DNA detection using real-time electrochemical LAMP. ACS Sens 1:904–912
Mori Y, Notomi T (2009) Loop-mediated isothermal amplification (LAMP): a rapid, accurate, and cost-effective diagnostic method for infectious diseases. J Infect Chemother 15:62–69
Moura-Melo S, Miranda-Castro R, De-Los-Santos-Alvarez N et al (2015) Targeting helicase-dependent amplification products with an electrochemical genosensor for reliable and sensitive screening of genetically modified organisms. Anal Chem 87:8547–8554
Nagamine K, Hase T, Notomi T (2002a) Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol Cell Probes 16:223–229
Nagamine K, Kuzuhara Y, Notomi T (2002b) Isolation of single-stranded DNA from loop-mediated isothermal amplification products. Biochem Biophys Res Commun 290:1195–1198
Nagatani N, Yamanaka K, Saito M et al (2011) Semi-real time electrochemical monitoring for influenza virus RNA by reverse transcription loop-mediated isothermal amplification using a USB powered portable potentiostat. Analyst 136:5143–5150
Nie G, Zhang Y, Guo Q et al (2009) Label-free DNA detection based on a novel nanostructured conducting poly(indole-6-carboxylic acid) films. Sensors Actuat B-Chem 139:592–597
Niu S, Zhao M, Hu L et al (2008) Carbon nanotube-enhanced DNA biosensor for DNA hybridization detection using rutin-Mn as electrochemical indicator. Sensors Actuat B-Chem 135:200–205
Niu S, Han B, Cao W et al (2009) Sensitive DNA biosensor improved by Luteolin copper(II) as indicator based on silver nanoparticles and carbon nanotubes modified electrode. Anal Chim Acta 651:42–47
Notomi T, Okayama H, Masubuchi H et al (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28:e63
Patterson AS, Hsieh K, Soh HT et al (2013) Electrochemical real-time nucleic acid amplification: towards point-of-care quantification of pathogens. Trends Biotechnol 31:704–712
Pedrero M, Campuzano S, Pingarrón JM (2011) Electrochemical genosensors based on PCR strategies for microorganisms detection and quantification. Anal Methods 3:780–789
Pemov A, Modi H, Chandler DP et al (2005) DNA analysis with multiplex microarray-enhanced PCR. Nucleic Acids Res 33:e11
Qi et al (2018) Isothermal exponential amplification techniques: from basic principles to applications in electrochemical biosensors. Biosens Bioelectron 110:207–217
Radoi A, Compagnone D (2009) Recent advances in NADH electrochemical sensing design. Bioelectrochemistry 76:126–134
Reid GD, Whittaker DJ, Day MA et al (2002) Femtosecond electron-transfer reactions in mono- and polynucleotides and in DNA. J Am Chem Soc 124:5518–5527
Shi P, Zhang Y, Yu Z et al (2017) Label-free electrochemical detection of ATP based on amino-functionalized metal-organic framework. Sci Rep 7:6500
Song T, Guo X, Li X et al (2016) Label-free electrochemical detection of RNA based on “Y” junction structure and restriction endonuclease-aided target recycling strategy. J Electroanal Chem 781:251–256
Steel AB, Herne TM, Tarlov MJ (1998) Electrochemical quantitation of DNA immobilized on gold. Anal Chem 70:4670–4677
Vincent M, Xu Y, Kong H (2004) Helicase-dependent isothermal DNA amplification. EMBO Rep 5:795–800
Wang W, Liu S, Ma C et al (2008) Determination of physiological thiols by electrochemical detection with piazselenole and its application in rat breast cancer cells 4T-1. J Am Chem Soc 130:10846–10847
Wang Q, Zheng H, Gao X et al (2013) A label-free ultrasensitive electrochemical aptameric recognition system for protein assay based on hyperbranched rolling circle amplification. Chem Commun 49:11418–11420
White RJ, Phares N, Lubin AA et al (2008) Optimization of electrochemical aptamer-based sensors via optimization of probe packing density and surface chemistry. Langmuir 24:10513–10518
Wilhelm J, Pingoud A (2003) Real-time polymerase chain reaction. ChemBioChem 4:1120–1128
Willner I, Zayats M (2007) Electronic aptamer-based sensors. Angew Chem Int Ed 46:6408–6418
Won BY, Shin S, Baek S et al (2011) Investigation of the signaling mechanism and verification of the performance of an electrochemical real-time PCR system based on the interaction of methylene blue with DNA. Analyst 136:1573–1579
Xuan F, Hsing IM (2014) Triggering hairpin-free chain-branching growth of fluorescent DNA dendrimers for nonlinear hybridization chain reaction. J Am Chem Soc 136:9810–9813
Xuan F, Fan TW, Hsing IM (2015) Electrochemical interrogation of kinetically-controlled dendritic DNA/PNA assembly for immobilization-free and enzyme-free nucleic acids sensing. ACS Nano 9:5027–5033
Yeung SS, Lee TM, Hsing IM (2006) Electrochemical real-time polymerase chain reaction. J Am Chem Soc 128:13374–13375
Yeung SS, Lee TM, Hsing IM (2008) Electrochemistry-based real-time PCR on a microchip. Anal Chem 80:363–368
Yu N, Wang Z, Wang C et al (2017) Combining padlock exponential rolling circle amplification with CoFe2O4 magnetic nanoparticles for microRNA detection by nanoelectrocatalysis without a substrate. Anal Chim Acta 962:24–31
Yurke B, Turberfield AJ, Mills Jr AP et al (2000) A DNA-fuelled molecular machine made of DNA. Nature 406:605–608
Zhang DY, Seelig G (2011) Dynamic DNA nanotechnology using strand-displacement reactions. Nat Chem 3:103–113
Zhang S, Tan Q, Li F et al (2007) Hybridization biosensor using diaquabis[N-(2-pyridinylmethyl)benzamide-κ2N, O]-cadmium(II) dinitrate as a new electroactive indicator for detection of human hepatitis B virus DNA. Sensors Actuat B-Chem 124:290–296
Zhang S, Xia J, Li X (2008) Electrochemical biosensor for detection of adenosine based on structure-switching aptamer and amplification with reporter probe DNA modified Au nanoparticles. Anal Chem 80:8382–8388
Zhang X, Qi B, Li Y et al (2009a) Amplified electrochemical aptasensor for thrombin based on bio-barcode method. Biosens Bioelectron 25:259–262
Zhang X, Su H, Bi S et al (2009b) DNA-based amplified electrical bio-barcode assay for one-pot detection of two target DNAs. Biosens Bioelectron 24:2730–2734
Zhang H, Fang C, Zhang S (2011) Ultrasensitive electrochemical analysis of two analytes by using an autonomous DNA machine that works in a two-cycle mode. Chem Eur J 17:7531–7537
Zhong H, Lei X, Hun X et al (2009) Design of one-to-one recognition triple Au nanoparticles DNA probe and its application in the electrochemical DNA biosensor. Chem Commun 6958–6960
Zhou LY, Zhang XY, Wang GL et al (2012) A simple and label-free electrochemical biosensor for DNA detection based on the super-sandwich assay. Analyst 137:5071–5075
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Gao, Z. (2019). Application of Nucleic Acid Amplification Strategies in Electrochemical DNA Sensors. In: Zhang, S., Bi, S., Song, X. (eds) Nucleic Acid Amplification Strategies for Biosensing, Bioimaging and Biomedicine. Springer, Singapore. https://doi.org/10.1007/978-981-13-7044-1_7
Download citation
DOI: https://doi.org/10.1007/978-981-13-7044-1_7
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-7043-4
Online ISBN: 978-981-13-7044-1
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)