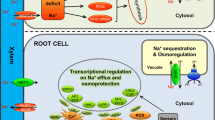
Abstract
Soil salinity is an unavoidable constraint in crop production globally. Soil salinization is often caused by improper soil management and/or crop production practices, which has made highly productive lands barren/unusable. Plant species have evolved several mechanisms to cope with salinity stress. Nutrient homeostasis is among the different mechanisms employed by plant species to withstand elevated salt levels in the root zone. Nutrients are the mediators of metabolism, so their cytoplasmic levels need to be effusively controlled both under stressful and benign environments. Several studies report the homeostasis of a single ion, i.e., sodium, potassium, or chloride. However, limited studies are available reporting the role of nutrient homeostasis (all nutrients together) under salinity stress. This chapter describes the role of nutrient homeostasis and ion channels and transporters in salt stress tolerance of plant species. The ion efflux at plasma membrane and vacuolar compartmentation in response to salinity stress has been described in detail. The impaired uptake of the nutrients is an obvious effect of salinity, mainly disturbing the sodium and potassium uptake. Much of the research has been done to test the role of different nutrients on salinity alleviation, and silicon is found to alleviate the negative effects of salinity. The nutrient homeostasis starts from ion sensing, uptake, transport, and activation of defense mechanisms as well as regulation of genes or gene networks to alleviate/withstand the adverse effects of salinity. Thus, the ion sensing, uptake, transport, and gene defense activation in response to salinity stress have also been described comprehensively.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
17.1 Introduction
Soil and water salinity stresses are unavoidable globally as around 830–950 million hectares of global soils are estimated to be affected by salinity (Rengasamy 2006, 2010; Ruan et al. 2010; Teakle and Tyerman 2010). Nearly half of the salt-affected soils are sodic, where 15% of cation exchange capacity is contributed by Na+ (Brady and Weil 2008; Rozema and Flowers 2008). The chloride (Cl−), sulfate (SO4 2−), carbonates (CO3 2−), and bicarbonates (HCO3 −) salts of different metals such as sodium (Na+), potassium (K+), magnesium (Mg+), and calcium (Ca2+) may give rise to soil salinity; however, sodium chloride (NaCl) is the most prevalent cause of salinity globally (Rengasamy 2002; Yadav et al. 2011).
Soil salinization is caused by improper management practices, which has made highly productive lands barren/unusable. The Mesopotamia, different regions of Indian and Pakistani Punjab, and the Fubei region in China are some of the examples of the areas where salinity has rendered productive lands barren (Hillel 2000, 2005; Swarajyalakshmi et al. 2003; Wang et al. 2008). The salts from deep in groundwater are moved upward by cultivation practices (Rengasamy 2006; Brady and Weil 2008; Yadav et al. 2011). The increased consumption of water by human has made salinity more acute today than before (Epstein and Bloom 2005; Brady and Weil 2008; Rozema and Flowers 2008). Out of the total available water for human consumption, 70% is consumed by agriculture, and this amount is rising with escalating population pressure (Hightower and Pierce 2008).
The noteworthy challenge for global agriculture is the production of an extra 70% food crops for another 2.3 billion people worldwide by the year 2050 (FAO 2009). However, the increased food demand is constrained by salinity. Over 20% of arable land around the world is influenced by salt stress, and the area is expanding steadily with each passing day. The crop plants could be divided into two major groups on the basis of their adaptive response to elevated salt levels in the soil. The first group comprises of the plants which can withstand salinity, known as halophytes (Hasanuzzaman et al. 2014), whereas the second group consists of the crop plants which are unable to tolerate the elevated salt levels and die, termed as glycophytes. Thus, salinity is regarded among the most ruthless abiotic stresses, which severely hamper the productivity of arable crops globally (Flowers 2004; Munns and Tester 2008).
Numerous physiological and metabolic processes are significantly altered by salinity stress. The extent of change in these processes depends on the nature of salts and the level and duration of stress, eventually hampering crop production (James et al. 2011; Rahnama et al. 2010; Munns 2005; Rozema and Flowers 2010). Salinity stress creates osmotic stress, which impair plant growths at the initial phases that is followed by ion toxicity at the later stages. Both osmotic stress and ion toxicity create unfavorable conditions for the normal growth and development of crop plants (James et al. 2011; Rahnama et al. 2010). High accumulation of salts at the initial phases of plant growth decreases the water absorption capacity of roots. Similarly, the osmotic stress caused by salinity stress increases water loss form the leaves. Thus salinity stress is also termed as hyperosmotic stress due to these reasons (Munns 2005).
Osmotic stress caused by elevated salt levels induces numerous physiological changes in crop plants. These physiological changes include membrane disruption, nutrient imbalance, lower detoxification of reactive oxygen species (ROS), diminished photosynthesis, opening of stomata, etc. (Munns and Tester 2005; Rahnama et al. 2010). Since NaCl is the most prevalent cause of salinity, accumulation of Na+ and Cl− ions in plant tissues is the most devastating effect on the crop plants exposed to elevated NaCl concentration in the soil. The K+ is a necessary element for normal plant growth; however, elevated levels of Na+ disrupt its uptake. The Na+-induced reduction in K+ uptake results in disturbed plant growth, low productivity, and even mortality based on the adaptive response of the crop plants to elevated salt levels in the soil (James et al. 2011). The ROS, such as singlet oxygen, superoxide, hydroxyl radical, and hydrogen peroxide, are produced in excess when plants are exposed to salinity stress (Apel and Hirt 2004; Mahajan and Tuteja 2005; Ahmad 2010; Ahmad et al. 2012; Ahmad and Umar 2011). Salinity-induced overproduction of ROS affects various cellular components (i.e., lipids, proteins, and DNA) through oxidative damage. Salinity induced oxidative damage eventually interrupts numerous functions at cell level in crop plants.
The physiology, development, and cellular metabolism in plants are driven by nutrient homeostasis (Clemens et al. 2002; Amtmann and Blatt 2009). The synthesis of organic macromolecules requires nutrient elements, which complete various functions in the key proteins. Moreover, they also act as cofactor of enzymes or as signaling molecules. The daily fluctuations posed by the environment on the plants have dramatic effects on the physiology and metabolism. The environmental fluctuations cause recurring fluctuations in the demands of essential nutrients required for the photosynthesis in chloroplasts. Moreover, nutrient transport pathways in xylem are also altered by the rhythmic changes in transpiration rates. Hence, continuous nutrient mobilization is required among organelles and tissues, particularly under nutrient deficit environments.
The identification of Na+ transport was a key gap in understanding the ionic homeostasis in plants under salinity stress (Niu et al. 1995). Generally, living cell under low or high salinity tend to balance passive Na+ inclusion with Na+ exclusion at two different levels. The balance is required at the plasma membrane and back to the apoplast, or across the tonoplast into the vacuole. Time is an imperative aspect needing consideration for salinity tolerance in addition to the considerable energy required for Na+ flow, as the Na + uptake rate determines the rate at which Na+ reaches toxic levels within the cell. Although there are number of studies available dealing with salinity tolerance and nutrient homeostasis under salinity stress, only a few aspects of nutrient homeostasis has been addressed. In this chapter, we will discuss all aspects of nutrient homeostasis and salinity tolerance in plant species. In summary, the chapter provides detailed information about how nutrients are mobilized under salinity stress to maintain normal growth.
17.2 Role of Nutrient Homeostasis in Salt Tolerance
Nutrients are the mediators of metabolism, so their cytoplasmic levels need to be effusively controlled both under stressful and benign environments. The reestablishment of nutrient homeostasis under stressful environments is the key strategy of plant species to improve their resistance against abiotic stresses, particularly salinity. Both types of nutrient homeostasis, i.e., ionic and osmotic homeostasis, need to be restored for salinity tolerance. The ionic homeostasis is determined/mediated by various ion transporters. Two nutrients, i.e., Na+ and Cl−, are the most detrimental to plant health under salinity stress; therefore both need to be under control in order to withstand higher salt levels (particular of NaCl) in growth medium. Many enzyme activities are inhibited by elevated levels of Na+, therefore Na+ accumulation in the cytoplasm or in organelles other than the vacuole needs to be prevented to reach at higher levels. This can be accomplished by either preventing Na+ entry or reducing it to safe or lower levels. The Na+ entry into plant cells is known to be controlled by nonselective cation channels (NCC) (Amtmann and Sanders 1998). The NCC is a channel, independent of voltage and serves as Na+ entry gate into the plant cells. Furthermore, there is the K+ outward-rectifying channel, opened by depolarization of plasma membrane and enables K+ exclusion and Na+ entry, which leads to the accumulation of Na+ in cytosol. The excess Na+ is pushed into vacuole with the help of vacuolar Na+/H+ exchanger. Another pump, the H+/Ca2+ antiporter, also aids toward Ca2+ homeostasis (Zhu 2002; Mahajan et al. 2006).
Na+ homeostasis is imbalanced by high salinity, while collective action of numerous pumps, ions, Ca2+ sensors, and their downstream interacting partners tends to normalize it. The organized action of these pumps, ions, and Ca2+ sensors ultimately causes efflux of excess Na+ ions. Certain channels tend to exhibit higher K+ sensitivity/selection over Na+, and K inward-rectifying channel is one of them. The excessive NaCl hyperpolarizes the plasma membrane, thus K-rectifying channel mediates K+ influx in response to higher NaCl, thus results in selective accumulation of K+ over Na+. On the other hand, there are some channels restrict the Na+ influx into the cytosol rather than selective accumulation of K+ over Na+. The histidine kinase transporter (HKT) is such an ion transporter, which restricts Na+ entry into the cytosol (Platten et al. 2006). There are numerous voltage-dependent anions, which are upregulated in response to elevated salinity, thus normalizing the Na+ homeostasis. The upregulation of a voltage-dependent anion has been observed in Pennisetum glaucum under salinity stress (Desai et al. 2006).
The Ca2+ has a significant role in salinity tolerance of crop plants by keeping the pivotal role in nutrient signaling under elevated salt levels. The cytosolic Ca2+ is increased under salinity stress, which initiates stress signal transduction pathways for tolerance to increased salinity levels. In addition, Ca2+ binding proteins may deliver an extra regulation level of Ca2+ signaling. The information provided by Ca2+ signaling is recognized and decoded by sensor proteins, which communicate this information to start a phosphorylation cascade regulating gene expressions.
Any Na+ entering the cells can be stored in the vacuole or transferred outside the cell. Na+ compartmentation is an inexpensive way to prevent Na+ toxicity in the cytoplasm, as Na+ can be used as a vacuolar osmolyte to provide osmotic homeostasis. Many salt-tolerant plants (halophytes) use this strategy to withstand the negative effects of salinity stress (Flowers et al. 1977).
The nutrient homeostasis prevents the Na+ entry into the cell, reduces it or pushes the excess Na+ into the vacuoles. There are a lot of channels and transporters involved in this process, which ultimately reduce the brutal effects of Na+ on plant growth. The most commonly observed homeostasis is for K+ and Ca2+ ions, which mitigate the adverse effects of Na+. The channels and transporters involved will be discussed in the coming sections. It can be briefly concluded that nutrient homeostasis is inevitable to achieve salinity tolerance for plants, which is accomplished by several ways as described under.
17.3 Types of Homeostasis
The homeostasis in plant species under salinity stress is divided into two categories, i.e., osmotic and ionic homeostasis. The osmotic homeostasis comprises of accumulation of compatible solutes, while ionic homeostasis consists of ionic influx and efflux and compartmentation at different levels in plant cells. Since the scope of the chapter is nutrient homeostasis, we will only discuss osmotic homeostasis in this chapter.
17.3.1 Ionic Homeostasis and Salt Tolerance
The maintenance of ionic homeostasis by partitioning and absorption of ions is not only required for the normal growth of plants under salinity-free conditions but also for highly saline environments (Niu et al. 1995; Serrano et al. 1999; Hasegawa 2013). Both glycophytes and halophytes species are unable to tolerate elevated salt levels in the cytoplasm. Therefore, the extra salt is elated to vacuole or reserved in old tissues, which are then for protection against salinity stress (Reddy et al. 1992; Zhu 2003). The main salt in the soil is NaCl, so this chapter focused on the transport mechanism and Na+ and its compartmentation.
The Na+/H+ antiporter transport the excess Na+ entering to the vacuole. There are two different kinds of H+ pump in the vacuolar membrane, i.e., H+-ATPase (V-ATPase) type vacuole and vacuolar pyrophosphatase (V-PPase) (Dietz et al. 2001; Otoch et al. 2001; Wang et al. 2001). Among these two pumps, V-ATPase dominates plant cells. These pumps have a vital role in the preservation of solute homeostasis, ensure secondary transfer energy, and facilitate vesicle fusion in stress-free conditions. The viability of the plant species under stress conditions is dependent on V-ATPase activity (Dietz et al. 2001). It has been observed that the efficacy of the VATPaz pump was increased in hypocotyls of Vigna unguiculata when exposed to elevated salt levels (Otoch et al. 2001). However, V-PPase activity was retarded under same conditions. The activity of V-ATPase was increased in halophytic species Suaeda salsa, while V-PPase had a negligible role (Wang et al. 2001).
The role of “Salt Overly Sensitive” (SOS) stress signaling pathway has increasingly been suggested in ionic homeostasis as well as tolerance to elevated salinity (Hasegawa et al. 2000; Sanders 2000). The SOS signaling pathway includes three main proteins, which are SOS1, SOS2, and SOS3. The first protein, i.e., SOS1, encodes the Na+/H+ antiporter of plasma membrane and is vital for the regulation of Na+ flux at the cellular level. At the same time, the Na+ transport from roots to the shoot is eased by this protein; thus overexpression of the protein imparts tolerance to the crop plants against elevated salt levels (Shi et al. 2000, 2002). The second protein, i.e., SOS2, encrypts a serine/threonine kinase and is activated by the Ca2+ signals resulting from salinity stress. This protein comprises of well-developed N-terminal catalytic domain and a C-terminal regulatory domain (Liu et al. 2000). The third protein, i.e., SOS3, is a myristoylated protein that binds Ca2+ and contains a myristoylation site in its N-terminus. This site has an important role in imparting tolerance to crop plants against elevated salinity (Ishitani et al. 2000). Therefore, a strong increase in intracellular Ca2+ level is observed with increasing Na+ concentration, which eases the binding with SOS3 protein. Intracellular Na+ homeostasis is moderated by Ca2+ along with SOS proteins. The SOS2 protein is activated by SOS3 protein thus releasing the spontaneous inhibition. The SOS3-SOS2 complex is then laden onto the plasma membrane, where SOS1 is phosphorylated.
Phosphorylated SOS1 increases Na+ flux, thus reducing the toxicity of Na+ (Martınez-Atienza et al. 2007). Many plant species have established an effective method to maintain ion concentration at a low level in the cytoplasm. Membranes, together with their associated apparatuses, regulate the uptake and transport of ions under elevated salinity, thus play an essential role in maintaining the ions concentration of in the cytosol in response to increased salt levels (Sairam and Tyagi 2004). Different carrier and proteins, antiporters, and symporters carry out the transplantation phenomenon.
The maintenance of cellular Na+/K+ homeostasis is vital for plant survival under highly saline conditions. Ma et al. (2011) reported that Arabidopsis NADPH oxidases function under salt stress in the ROS-dependent regulation of Na+/K+ homeostasis in Arabidopsis by AtrbohD and AtrbohF genes. Plants maintain high K+ level in cytoplasm (approximately 100 mM) for cytoplasmic enzyme activities. The K+ concentration in the vacuole ranges from 10 to 200 mM. The vacuole is the largest K+ pool within the plant cell. K+ has a vital role in maintaining the turgor within the cell. The K+ transporter and membrane channels transport K+ to the plant cells against the concentration. The K+ intake mechanism is strongly mediated by its amount present in the soil. When extracellular concentration of K+ is low, high affinity K+ carriers uptake K+ from the soil. However, the low affinity K+ channels get activated for restricted uptake when extracellular concentration of K+ is high. On the other hand, a very low Na+ ion concentration is maintained in the cytosol. Since both K+ and Na+ share the same transport mechanism, Na+ competes K+ for the carrier, under salinity stress due to the increased Na+ concentration in the soil, thus reducing K+ absorption (Sairam and Tyagi 2004; Munns and Tester 2008).
Numerous genes and proteins encoding K+ carriers and channels such as Na+ influx transporter (HKT) and the tonoplast Na+/H+ antiporter (NHX) have been recognized and duplicated in several crop plants. The expression of low abundance transcripts increases the uptake of K+ under salinity stress. This expression of low abundance transcripts has been recorded in a halophyte species, i.e., Mesembryanthemum crystallinum (Yen et al. 2000).
The HKT family transporters found on the plasma membrane have a significant role in salinity tolerance through the regulating the transport of Na+ and K+. HKT Class 1 (HKT1) transporters, classified in Arabidopsis, prevent excessive buildup of Na+ in leaves, thus protecting plants form the adverse effects of salinity stress. The HKT1 rice transporter removed extra Na+ from xylem, thus protecting the photosynthetic tissue of the rice leaves from the Na+ toxicity (Schroeder et al. 2013). Barragán et al. (2012) indicated that two localized tonoplast NHX proteins (NHX1 and NHX2) are vital for active K+ uptake at the tonoplast. These proteins are required for regulation of turgor regulation and normal functioning of stomata. Several NHX isoforms have been proved to play an imperative role in ionic homeostasis (Na+, K+, H+) in different crop species (Gálvez et al. 2012).
17.4 Cation Uptake, Mechanisms, Transporters Involved, and Role of Ion Channels
Crop plants grown under elevated salt levels are exposed to explicit ionic effects, which damage the enzymes structure as well as some macromolecules including Na+ and Clˉ. The ionic effects also exert damages to the cell organelles as well as resulted in impaired photosynthesis and respiration. Salinity stress also results in the physiological drought resulting in impaired uptake and transport of nutrients. The disturbed uptake and transport of nutrients leads to imbalanced nutrition in crop plants under elevated salt levels (Munns and Tester 2008; Ruiz-Lozano et al. 2012). Higher accumulation and degradation of Na+ and reduced K+ uptake are the obvious detrimental effects of elevated salt levels on plant growth, while these mechanisms are still unclear or understudied (Chen et al. 2007). Na+ is an important cation prevailing in the soils affected by salinity. The K+ and Na+ activate and inhibit numerous cytosolic enzymes, respectively (Shi et al. 2002). Under natural physiological circumstances, the plants hold 1–10 mM Na+ and 60–100 mM K+ in the cytosol (Bassil et al. 2012).
The Na+ is a cytotoxin which severely disrupts proteins and membranes. Moreover, several physiological processes including cell expansion and cell division, metabolism (primary and secondary) and Na+ severely impacts nutrient homeostasis (Hasegawa et al. 2000; White and Broadley 2001; Munns and Tester 2008; Teakle and Tyerman 2010). The presence of excessive Na+ in the soil restricts K+, which decreases intracellular K+ ultimately disturbing K+/Na+ imbalance (Hauser and Horie 2010; Leidi et al. 2010; Alemán et al. 2011; Pardo and Rubio 2011). Excessive presence of Na+ offers strong competition to K+ for even in the presence of high-affinity K+ transport systems (Rus et al. 2004; Kronzucker et al. 2008; Alemán et al. 2009, 2011; Pardo and Rubio 2011). The conductance of K+ is reduced by Na+ through AKT1 (Qi and Spalding 2004) which suppresses the expression of AtHAK5 (Nieves-Cordones et al. 2008; Alemán et al. 2011; Pardo and Rubio 2011).
The synthesis of proteins and activities of cytosolic enzymes are inhibited under abnormally high cytosolic Na+/K+ ratio (Shabala and Cuin 2008). Thus, plants are equipped with numerous mechanisms (biochemical and molecular) to withstand the brutal impacts of salt stress. Regulating salinity uptake genes and Na+ and/or K+ transport or compartmentation are the mechanisms developed by plants for adequate ionic homeostasis. The undue Na+ accrual in cytosol is prohibited through a number of mechanisms by glycophyte plant species. The first mechanism is the selective uptake of ions to restrict Na+ for the regulation of ionic homeostasis under salinity stress. The second mechanism is to maximize the Na+ efflux to growth medium or to apoplastic spaces. The plant species finally can impound Na+ into vacuoles for restricting the Na+ transfer to the shoot (Cuin et al. 2011). The two mechanisms described above are vital and usually used by crop plants to control undue Na+ buildup (Cuin et al. 2011; Cabot et al. 2014). The vacuolar Na+ and K+/H+ antiporters NHXs catalyze the impounding of Na+ into vacuoles (Cuin et al. 2011). Four different genes (OsNHX1–4) relating to these antiporters have recently been identified in rice crop (Fukuda et al. 2011; Kumar et al. 2013).
Plasma membrane Na+/H+ antiporter (SOS1) catalyzes the Na+ efflux from cytosol to the growth medium or to apoplastic spaces in numerous crop plants (Kumar et al. 2013). The SOS1 is preferentially expressed in the cells which surround xylem, thus suggesting that this transporter plays a vital role in the redistribution of Na+ between roots and shoots. The transporter has also been suggested to have ability of preventing Na+ to reach the photosynthetic tissues (Shi et al. 2002; Olias et al. 2009). The unloading of Na+ from photosynthetic organs and its recirculation to roots have also suggested as a mechanism imparting salinity tolerance to crop plants (Davenport et al. 2007). The high-affinity HKT transporters are reported to be involved in this mechanism from several crop plants (Garciadeblás et al. 2003; Ren et al. 2005).
Different studies relating to physiological and molecular mechanisms/processes have identified the channels and transporters involved in the tolerance mechanisms to elevated salt levels in various plant species. These channels and transporters are the passages for the ions required for cellular function related to ionic toxicity.
It is now known that HKT1 transporter-mediated salinity tolerance is too much complex than expected. To infer the role of these transporters, Chen et al. (2017) evaluated the correlation between the activity of two Mg2+ transporter, i.e., OsMGT1 and OsHKT1. It was observed that OsMGT1 mutants accumulated excessive amount of Na+ with OsHKT1 mutants. The expression of OsMGT1 different plant parts such as parenchyma cells in xylem and phloem and leaf sheath tissue spatially overlapped with the expressions OsHKT1 (Kobayashi et al. 2017). These findings regarding the co-expression of these transporters added valuable information toward understanding complex Na+ regulation mechanisms at the organismal level. The recent findings of Kobayashi et al. (2017) and Chen et al. (2017) cleared that nodes and sheaths have an imperative role in avoiding ion toxicity in the reproductive organs of plant species. The restricted transport of Na+ to older leaves is a typical mechanism of salinity tolerance in plants (Cotsaftis et al. 2012); however, this mechanism vanished in OsMGT1 mutants (Chen et al. 2017). These findings suggest that the fine-tuning of HKT1 activity is mediated by OsMGT1 mutants. Thus, it is noteworthy to determine the cell-specific expression and functions/roles of OsMGT1 mutant at later stages, particularly reproductive development of the crop plants. The OsHKT1 is overexpressed during reproductive developmental stage of crop plants, particularly in node I (Kobayashi et al. 2017). It is obvious from the abovementioned both studies (Chen et al. 2017; Kobayashi et al. 2017) that nodes function as fences where OsMGT1 regulates the gating of OsHKT1 in a spatiotemporal manner to limit the ion toxicity.
Different cation channels which are nonselective (NSCC) are indulged in the unidirectional intracellular Na+ influx. The members of NSCC, including HAK/KUP/KT and AKT1, are the channels having higher affinity for K+ acquisition, whereas the low-affinity acquisition is mediated by different cation transporters including cation-Cl– cotransporter and high-affinity K+ transporter (HKT1 and HKT2) (Plett and Møller 2010; Zhang et al. 2010; Kronzucker and Britto 2011). Although NSCCs and HKT1 transporters are main mediators of Na+ uptake, their comparative contribution is ambiguous (Roberts and Tester 1997; Amtmann and Sanders 1998; Demidchik and Maathuis 2007). Among two classes of HKT proteins, HKT1 is more selective for Na+, while HKT2 proteins exhibit higher K+ selectivity than Na+ or remain nonselective (Hauser and Horie 2010; Lan et al. 2010; Mian et al. 2011). Ali et al. (2012) have recently identified a HKT1 family transporter having higher K+ selectivity than Na+ from Thellungiella sp. The SOS1 Na+/H+ antiporter controls the Na+ efflux across the plasma membrane. The antiporter has phylogenetic similarity with mammalian NHE and bacterial NhaP Na+/H+ antiporters (Zhu 2002, 2003; Pardo and Rubio 2011; Kronzucker and Britto 2011). The Na+ efflux to the apoplast is mediated by SOS1 against the electrochemical potential by secondary active transport driven by the H+ gradient across the plasma membrane.
17.5 Anion Uptake, Mechanisms, Transporters Involved, and Role of Ion Channels
The anions are more prevalent than cations in the soil; however, the transport mechanisms of anions are less understood than cation transport mechanisms both under normal soil conditions and elevated salt levels. The Cl− transport mechanisms are also less discussed than cations for plant mineral nutrition as well. The Cl− performs several mechanisms such as enzyme activities’ regulation in cytoplasm acting as an important cofactor in photosynthesis and counteranion stabilizing membrane potential and also regulates turgor and pH. Thus it is considered as an important micronutrient for plant growth (Tyerman 1992; Marschner 1995; Teodoro et al. 1998; Xu et al. 1999; White and Broadley 2001). The higher amounts of Cl− are toxic for crop plants. The critical toxicity level of Cl− is known to be 4–7 and 15–50 mg g−1 dry weight for Cl−-sensitive and Cl−-tolerant species, respectively (Xu et al. 2000). The most dominant ions in NaCl-affected soils are Na+ and Cl−, which at higher concentrations are toxic for plant species. However, some of the plant species are able to better tolerate Na+ transport than Cl−, while others have better control on Cl− transport (Munns and Tester 2008). The differential salt tolerance level of plant species has strong correlation with Cl− transport and exclusion from shoots. Different legume species such as Trifolium pratense (Winter 1982; Rogers et al. 1997), Medicago sativa (Sibole et al. 2003), Glycine max (Luo et al. 2005), and Lotus (Teakle et al. 2006, 2007) have higher tolerance level to salinity and thus can better exclude Cl− than nonleguminous species. Similarly, some woody species, such as Pinus banksiana (Franklin and Zwiazek 2004), Citrus reticulata, and Vitis vinifera (Sykes 1992; Romero-Aranda et al. 1998; Moya et al. 2003), also have better control over Cl− transport and exclusion.
The focus of research dealing with salinity tolerance has been to maintain favorable K+/Na+ ratio through establishing selectivity between Na+ and K+. Several specific transport systems, or combination of selective transporters at different membranes as well as different cells alongside the transport pathways through root to shoot, are related to Na+ and K+ selectivity (Hua et al. 2003; Horie et al. 2005; Volkov and Amtmann 2006; Apse and Blumwald 2007; Byrt et al. 2007). A resembling mechanism might be present for the exclusion or selectivity of Cl− at least for major macronutrient anions (NO3 −, SO4 2−) and organic anions. However, the mechanisms and transporters mediating the Cl− exclusion and or the pathways involved are not well-studied until now. Several micronutrients impart salinity tolerance to crop plants, which might be attributed to the activities of transporters and selective channels (Grattan and Grieve 1998). The NO3 − is the most prevalent univalent anion in soils under salinity-free conditions; therefore it must be focused to understand Cl− and salinity tolerance (Fricke et al. 1994; Frachisse et al. 1999). There are numerous anion channels having higher affinity and more selective to NO3 − than Cl− (Roberts 2006). Thus the NO3 − in growth media could decrease the concentration of Cl−concentration in (Abdolzadeh et al. 2008; Gimeno et al. 2009; Song et al. 2009). The NO3 −/Cl− balance is akin of K+/Na+ balance or interactions as well as selectivity for Cl− efflux mechanisms.
Charge balance is another subject which has not been sufficiently addressed and correlated with the comparative roles of Cl− and Na+ in salinity tolerance. For salinity tolerance the movement and net charge must be in equiliburium by the opposite ions in each compartment to reach the similar charge in each section. If Cl− uptake is higher, then a cation (e.g., Na+) uptake or exclusion of anion must balance this charge from Cl− perspective. The potential anions which could balance this charge are not readily moved out as these are related to nutrition (e.g., NO3 −) or carbon balance (e.g., malate). However, from Na+ viewpoint, addition to the opposite of abovementioned mechanisms, either Cl− or K+ might be excluded (Shabala et al. 2006). This exclusion is true in a sense if Na+ substitutes K+ role in vacuole and concentration of K+ in the cytoplasm is retained below or equal to adequate limits (Carden et al. 2003). The voltage changes across the membrane eventually regulate charge balance, which then impacts the driving force on counterions, effectually as a self-regulating system. The type, nature, and selectivity of transporters present at the membrane barriers as well as their response to voltage changes in the membrane eventually determine the nature of the charge balance. These responses are also important for the energy needs of transport (Britto and Kronzucker 2009). Plett and Møller (2009) are referred for the further readings on Na+ transport mechanisms.
Various aspects of anion transport have been covered in detail by several reviews (Roberts 2006; De Angeli et al. 2009). However, there has been lacking information on the Cl− transport mechanisms and their role in salinity tolerance (White and Broadley 2001). Salinity tolerance of different plant species is imparted by a number of traits including osmotic stress tolerance, compatible solute accumulation, and tolerance to oxidative stress. Several reviews have described the traits imparting salinity tolerance in various crop species (Bartels and Sunkar 2005; Munns 2005; Flowers and Colmer 2008; Munns and Tester 2008). In summary, not only cation uptake and balance is essential for salinity tolerance, but anions also play an integral role. Thus studying the anion uptake mechanisms, regulation of anions, transporters, and channels involved in transport of anions should be the focus of salinity tolerance studies to be conducted in the future.
17.6 Ion Efflux at the Plasma Membrane
Maintenance of high tissue K+/Na+ ratio through regulating Na+ uptake and transport in plant species has been interpreted in various studies for salinity tolerance. Thus, a high K+/Na+ has been considered as an important trait imparting salinity tolerance to crop plants (Shabala and Pottosin 2014). Since K+ participates in various physiological processes; thus, this interpretation is logical. Moreover, higher concentration of Na+ often competitively disrupts K+ uptake, and increased K+ deficiency under elevated salt levels severely impedes plant growth and development. Thus, it could be concluded that sensitivity of crop plants to salinity stress is because of K+ deficiency, especially keeping in view that concentration of K+ in soil is typically in the micromolar range (Very and Sentenac 2003). Several recent studies have focused the plant adaptation mechanisms to low K+ under elevated salts or salinity-free conditions. Hence, these mechanisms must be explored to better understand salinity tolerance in crop plants. The studies focusing on these mechanisms have concluded that sufficient or higher K+ availability imparts salinity tolerance to crop plants, while low K+ availability in soils makes salinity stress highly devastating for crop plants.
The Na+-induced K+ exclusion form root and leaf cells is the most obvious effect of salinity stress on K+ homeostasis (Wang et al. 2009; Demidchik et al. 2014). The K+ exclusion has been concluded to be the exclusive result of Na+ entering into the cytoplasm. The Na+ inclusion depolarizes membrane potential below the resting potential. This depolarization consequently activates K+ outward rectifier channels, such as GORK (guard cell outward rectifying K+ channel), through which K+ is excluded from the plant cells and tissues. Thus, maintenance of higher inside negative potential through prevention of membrane depolarization for enhancing intracellular K+ retention (inhibition of K+ efflux) (Falhof et al. 2016) would be a salinity tolerance mechanism of crop plants. Recent studies have identified that retention capacity of intracellular K+ is a crucial mechanism for salinity stress tolerance (Janicka-Russak and Kabała 2015). The PM H+-ATPases and tonoplast H+-ATPases/H+-PPases, K+ transporters, NHX antiporter, and SOS1 proteins have been considered as vital players in the process of subsiding salinity stress and negative effects of low K+ availability on crop plants (Pottosin and Dobrovinskaya 2014; Janicka-Russak and Kabała 2015; Falhof et al. 2016). It is concluded that K+ efflux and influx mechanism are really crucial for salinity tolerance in crop plants. Thus, equipping crops with these mechanism will assure survival and sustained production under elevated salt levels in the soil.
17.7 Vacuolar Compartmentation and Ion Homeostasis
The ability of different plant species to tolerate elevated salinity levels is determined by the Cl− partitioning between different types of roots and shoots cells. Some evidence report that Cl− is accrued in the epidermis cells of leaves, which reduces the Cl− toxicity photosynthetically important mesophyll cells. The differential ability to exclude Cl− from mesophyll cells has been observed in two barley cultivars where salt-tolerant genotype was better able to exclude Cl− than salt-sensitive genotype (Huang and Van Steveninck 1989). A more sophisticated analyses conducted by using single-cell-sampling techniques revealed that Cl− accumulation increased in both epidermal and mesophyll cells with rising NaCl (Fricke et al. 1996). It was also concluded that photosynthesis in salt-stressed plants was mildly affected, which gives rise to the doubts that Cl− accumulation in epidermal cells is linked with salinity tolerance. Fricke et al. (1996) also revealed that the Cl− accumulation in epidermal cells was three times higher than the Cl− accumulation in mesophyll cells. However, regardless of this fact, Na+ and K+ accumulation was similar in both types of cells. James et al. (2006) studied the accumulation of Cl− in two barley cultivars differing in a salinity tolerance. It was concluded that Cl− preferred epidermis cells over mesophyll cells for accumulation. Similar pattern of Cl− accumulation was observed in both sensitive and tolerant cultivar, which indicates that Cl− accumulation in a specific type of cells could not be concluded a major trait conferring salinity tolerance in crop plants.
The Cl− accumulation in salt glands or bladders is one other form of intercellular compartmentation. Halophyte group of crop plants possesses these special structures, which can store Cl− (and even Na+), helping plants to withstand the negative effects of elevated salt levels. The secretion of Cl− through salt glands can be concluded as an important salinity tolerance trait as >20% Cl− in the leaves of Leptochloa fusca under 100 mm NaCl level was secreted by salt glands (Jeschke et al. 1995). Different studies have focused the salt glands possessed by some halophytic species, i.e., Bienertia sinuspersici and Limonium sinense (Ding et al. 2009; Park et al. 2009). A cation-chloride cotransporter (CCC) was localized to leaf trichomes and hydathodes in Arabidopsis (Colmenero-Flores et al. 2007). The ultimate role of CCC is still unclear, however, investigating whether CCC are also present in the slat glands of halophyte species to infer the possible Cl− efflux (and Na+) from leaves.
The salinity tolerance of the crop plants could not always be attributed to the low concentrations of Cl− and Na+ in the shoots. The Cl− tolerance largely varies among genotypes of a same species. Therefore, salinity tolerance could not be attributed to the shoot Cl− concentration. The Cl− and Na+ are excluded by most of the plant species up to a certain level (90–98%; Munns 2005); however, effective Cl− and Na+ sequestration in the vacuole to prevent them to accumulate at toxic levels will impart ultimate salinity tolerances to crop plants. Even the halophyte plant species are unable to elevate levels of cytoplasmic Cl−; thus they have evolved strategies which effectively sequester Cl− into vacuoles to control the accumulation of Cl− and other ions through turgor-driven growth (Flowers et al. 1977; Glenn et al. 1999).
The direct measurements of Cl− fluxes and its concentration in the vacuoles of the plants intact from salinity through experimental studies are difficult; however, there exist some estimates conducted through X-ray microanalysis, intracellular ion-sensitive microelectrodes, tracer compartmental analysis, or Cl−-sensitive fluorescent probes (Hajibagheri and Flowers 1989; Felle 1994; Britto et al. 2004; Lorenzen et al. 2004). All these estimates have concluded that the vacuole of plant cells are able to accumulate Cl− up to 500 mm (Cram 1973).
There have been some indirect confirmation that efficient Cl− sequestration in the vacuole imparts salinity tolerance to some plant species. The salinity-tolerant genotypes of citrus, grapevine, and Lotus had low Cl− in shoots, while accumulated more Cl− in the roots compared to their respective sensitive genotypes (Storey and Walker 1998; Storey et al. 2003). These findings suggest that the tolerant genotypes efficiently compartmentalized the Cl− in root vacuoles. Some avocado rootstocks have been found to possess high amount of Cl− in the leaves, which has been linked to their salinity tolerance (Xu et al. 2000). Similarly, some lupin cultivars also accumulated more Cl− in the leaves for salinity tolerance (Van Steveninck et al. 1982).
Some direct comparison among different genotypes has concluded that effective compartmentation of intracellular Cl− imparts salinity tolerance to crop plants. Hajibagheri et al. (1989) compared two maize genotypes differing salinity tolerance for cytoplasmic Cl− concentrations. It was concluded that sensitive genotypes accumulated more root cytoplasmic Cl− concentrations than tolerant genotype. The accumulated amount of Cl− was comparable to accumulated amount of Cl− (350 mm for barley at 100 mm NaCl) in barley genotypes (Britto et al. 2004). Flowers and Hajibagheri (2001) compared two barley genotypes having differential ability of salt tolerance through X-ray microanalysis. It was concluded that cytoplasmic and vacuolar concentrations of Cl− (and Na+) were similar; however, the tolerant barley cultivar accumulated half of the Cl− concentration accumulated by the sensitive genotype. The higher concentration of ions in the cell wall of sensitive genotype reduced turgor, which ultimately resulted poor growth. To assess whether vacuolar Cl− sequestration is involved in salinity tolerance, further studies are needed focusing on the comparison of genotypes having differential salt tolerance level, but having similar leaf or root Cl− concentrations.
17.8 Ion Sensing and Gene Defense Activation
Several genes imparting salinity tolerance to crop plants have been identified through the use of genetic damage or gain of function methods (Zhu 2002, 2003; Apse and Blumwald 2007; Pardo and Rubio 2011; Peleg et al. 2011). The identified genes encode transport determinants mediating Na+ homeostasis (Plett and Møller 2010; Pardo and Rubio 2011; Peleg et al. 2011). These identified determinants mainly from glycophyte plant species enhance the salinity tolerance capabilities of halophytic species as well as crop plants (Oh et al. 2010, 2012; Dassanayake et al. 2011a, b; Munns et al. 2012). There has been a lot of conversation on the underlying mechanisms and determinants imparting salinity tolerance to halophytes and glycophytes. There are emerging evidence which provide a degree of systematic clarification that why halophytes are better able to tolerate elevated salt levels (Amtmann 2009; Dassanayake et al. 2011b; Oh et al. 2012).
The halophytic plant species possess “superior” alleles and novel loci involved in Na+ homeostasis and salinity tolerance (Edelist et al. 2009; Dassanayake et al. 2011a, b; Oh et al. 2012). There have been several critical questions relating to Na+ homeostasis in halophytic plant species. These critical questions include the following: Do halophytes have unique transport determinants? Do halophytes have orthologous determinants with diverse actions? And do halophytes differentially control transport protein action or expression of encoding genes to increase salinity tolerance (Oh et al. 2009, 2012; Plett and Møller 2010; Dassanayake et al. 2011b)? The rapidly advancing omics technologies such as massively parallel sequencing, whole genome sequencing, and sequencing by genotyping as well as phenotyping could facilitate to find the answers of all these critical questions providing valuable insights on the salinity tolerance mechanisms of halophytic plant species (Lin et al. 2004; Takeda and Matsuoka 2008; Dassanayake et al. 2011b; Oh et al. 2012).
The well-known and suggested salinity tolerance signaling pathway includes facilitation of ionic and osmotic homeostasis, growth regulation, and regulation of development (Zhu 2002). The SOS proteins, phospholipid, ROS, ABA, cytokinin, Ca2+, hyperosmotic and osmotic solute, and kinase/phosphatase pathways are the probable pathways integrated to cope salinity tolerance/acclimation in plant species, which are mainly concluded from the studies conducted on Arabidopsis (Gong et al. 2001; Zhu 2002; Qin et al. 2011; Reddy et al. 2011; Suzuki et al. 2012). It has been described that several determinants are governed and regulated by abovementioned networks and various transcriptomic and proteomic analyses have identified that these determinants play a crucial role in salinity tolerance of crop plants (Zhu 2002; Golldack et al. 2011; Pérez-Alfocea et al. 2011; Singh et al. 2011; Zhang et al. 2012).
The Na+ homeostasis, as described above, is mediated by a highly defined SOS Ca2+ signaling pathway (Zhu 2002, 2003). The Na+ signal perception is unclear; however, Ca2+ is known to be a secondary messenger in signal transduction (Zhu 2002, 2003; Conde et al. 2011). The activates of SOS1 Na+/H+ antiporter are regulated by SOS pathway, which facilitates Na+ efflux across the plasma membrane (Zhu 2002, 2003; Pardo and Rubio 2011). Calcineurin B and neuronal Ca2+ sensor-like protein SOS3 decode the NaCl-induced cytosolic Ca2+ increase (CBL4). The CBL4 is a myristoylated protein with EF-hand Ca2+-binding places (Zhu 2002, 2003; Gong et al. 2004; Sánchez-Barrena et al. 2005; Tracy et al. 2008; Pardo and Rubio 2011). Ca2+-activated SOS3 interacts with the auto-inhibitory domain of SOS2 (CIPK24), a member of the SnRK family (Zhu 2002, 2003; Gong et al. 2004; Cosello et al. 2011; Kulik et al. 2011; Pardo and Rubio 2011). SOS3 binding to the SOS2 auto-inhibitory domain triggers kinase action and enables localization of the SOS2-SOS3 complex (Zhu 2002, 2003; Sánchez-Barrena et al. 2007; Pardo and Rubio 2011). SOS2 then acquaintances with SOS1 Na+/H+ in the plasma membrane, phosphorylating the transporter and triggering Na+ exclusion (Zhu 2002, 2003; Martínez-Atienza et al. 2007; Pardo and Rubio 2011; Quintero et al. 2011). Several genes and gene networks are being identified with each passing day with technological advancements. Thus a comprehensive understanding of nutrient homeostasis requires sound knowledge of different mechanisms involved. Although, several mechanisms are discussed in this chapter, still gaps exist which need to be filled through comprehensive studies on salt tolerance and nutrient homeostasis in plants. Moreover, the studies focusing on mediating nutrient homeostasis could provide valuable insights on inducing salinity tolerance in plant species through the manipulation of nutrient homeostasis.
References
Abdolzadeh A, Shima K, Lambers H, Chiba K (2008) Change in uptake, transport and accumulation of ions in Nerium oleander (Rosebay) as affected by different nitrogen sources and salinity. Ann Bot 102:735–746
Ahmad P (2010) Growth and antioxidant responses in mustard (Brassica juncea L.) plants subjected to combined effect of gibberellic acid and salinity. Arch Agron Soil Sci 56:575–588
Ahmad P, Umar S (2011) Oxidative stress: role of antioxidants in plants. Studium Press, New Delhi
Alemán F, Nieves-Cordones M, Martínez V, Rubio F (2009) Potassium/sodium steady-state homeostasis in Thellungiella halophila and Arabidopsis thaliana under long-term salinity conditions. Plant Sci 176:768–774
Alemán F, Nieves-Cordones M, Martínez V, Rubio F (2011) Root K+ acquisition in plants: the Arabidopsis thaliana model. Plant Cell Physiol 52:1603–1612
Ali Z, Park HC, Ali A, Oh DH, Aman R, Kropornicka A, Hong H, Choi W, Chung WS, Kim WY, Bressan RA, Bohnert HJ, Lee SY, Yun DJ (2012) TsHKT1; 2, a HKT1 homolog from the extremophile Arabidopsis relative Thellungiella salsuginea, shows K(+) specificity in the presence of NaCl. Plant Physiol 158:1463–1474
Amtmann A (2009) Learning from evolution: Thellungiella generates new knowledge on essential and critical components of abiotic stress tolerance in plants. Mol Plant 2:3–12
Amtmann A, Blatt MR (2009) Regulation of macronutrient transport. New Phytol 181:35–52
Amtmann A, Sanders D (1998) Mechanisms of Na+ uptake by plant cells. Adv Bot Res 29:75–112
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Apse MP, Blumwald E (2007) Na+ transport in plants. FEBS Lett 581:2247–2254
Barragán V, Leidi EO, Andrés Z, Rubio L, De Luca A, Fernández JA, Cubero B, Pardo JM (2012) Ion exchangers NHX1 and NHX2 mediate active potassium uptake into vacuoles to regulate cell turgor and stomatal function in Arabidopsis. Plant Cell 24:1127–1142
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58
Bassil E, Coku A, Blumwald E (2012) Cellular ion homeostasis: emerging roles of intracellular NHX Na+/H+ antiporters in plant growth and development. J Exp Bot 63:5727–5740
Brady NC, Weil RR (2008) The nature and properties of soils, 14th edn. Pearson Prentice Hall, Upper Saddle River
Britto DT, Kronzucker HJ (2009) Ussing’s conundrum and the search for transport mechanisms in plants. New Phytol 183:243–246
Britto DT, Ruth TJ, Lapi S, Kronzucker HJ (2004) Cellular and whole-plant chloride dynamics in barley: insights into chloride–nitrogen interactions and salinity responses. Planta 218:615–622
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, Tester M, Munns R (2007) HKT1;5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1. Plant Physiol 143:1918–1928
Cabot C, Sibole JV, Barceló J, Poschenrieder C (2014) Lessons from crop plants struggling with salinity. Plant Sci 226:2–13
Carden DE, Walker DJ, Flowers TJ, Miller AJ (2003) Single-cell measurements of the contributions of cytosolic Na+ and K+ to salt tolerance. Plant Physiol 131:676–683
Chen Z-H, Zhou M, Newman IA, Mendham NJ, Zhang GP, Shabala H (2007) Potassium and sodium relations in salinised barley tissues as a basis of differential salt tolerance. Funct Plant Biol 34:150–162
Chen ZC, Yamaji N, Horie T, Che J, Li J, An G, Ma JF (2017) A magnesium transporter OsMGT1 plays a critical role in salt tolerance in rice. Plant Physiol 174:1837–1849
Clemens S, Palmgren MG, Krämer U (2002) A long way ahead: understanding and engineering plant metal accumulation. Trends Plant Sci 7:309–315
Cosello P, Hey SJ, Halford NG (2011) The sucrose non-fermenting-1-related (SnRK) family of protein kinases: potential for manipulation to improve stress tolerance and increase yield. J Exp Bot 62:883–893
Colmenero-Flores JM, Martinez G, Gamba G, Vázquez N, Iglesias DJ, Brumós J, Talón M (2007) Identification and functional characterization of cation-chloride cotransporters in plants. Plant J 50:278–292
Conde A, Chaves MM, Gerós H (2011) Membrane transport, sensing and signaling in plant adaptation to environmental stress. Plant Cell Physiol 52:1583–1602
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS One 7(7):e39865
Cram WJ (1973) Internal factors regulating nitrate and chloride influx in plant cells. J Exp Bot 24:328–341
Cuin TA, Bose J, Stefano G, Jha D, Tester M, Mancuso S, Shabala S (2011) Assessing the role of root plasma membrane and tonoplast Na+/H+ exchangers in salinity tolerance in wheat: in planta quantification methods. Plant Cell Environ 34:947–961
Dassanayake M, Oh D-H, Haas JS, Hernandez A, Hong H, Ali S, Yun D-J, Bressan RA, Zhu J-K, Bohnert HJ, Cheeseman JM (2011a) The genome of the extremophile crucifer Thellungiella parvula. Nat Genet 43:913–918
Dassanayake M, Oh D-H, Hong H, Bohnert HJ, Cheeseman JM (2011b) Transcription strength and halophytic lifestyle. Trends Plant Sci 16:1–3
Davenport RJ, Muñoz-Mayor A, Jha D, Essah PA, Rus A, Tester M (2007) The Na+ transporter AtHKT1; 1 controls retrieval of Na+ from the xylem in Arabidopsis. Plant Cell Environ 30:497–507
De Angeli A, Monachello D, Ephritikhine G, Frachisse J-M, Thomine S, Gambale F, Barbier-Brygoo H (2009) CLC mediated anion transport in plant cells. Philos Trans R Soc Lond Ser B Biol Sci 364:195–201
Demidchik V, Maathuis FJ (2007) Physiological roles of nonselective cation channels in plants: from salt stress to signalling and development. New Phytol 175:387–404
Demidchik V, Straltsova D, Medvedev SS, Pozhvanov GA, Sokolik A, Yurin V (2014) Stress-induced electrolyte leakage: the role of K+-permeable channels and involvement in programmed cell death and metabolic adjustment. J Exp Bot 65:1259–1270
Desai MK, Mishra RN, Verma D, Nair S, Sopory SK, Reddy MK (2006) Structural and functional analysis of a salt stress inducible gene encoding voltage dependent anion channel (VDAC) from pearl millet Pannisetum glaucum. Plant Physiol Biochem 44:483–493
Dietz KJ, Tavakoli N, Kluge C, Mimura T, Sharma SS, Harris GC, Chardonnens AN, Golldack D (2001) Significance of the V-type ATPase for the adaptation to stressful growth conditions and its regulation on the molecular and biochemical level. J Exp Bot 52:1969–1980
Ding F, Song J, Ruan Y, Wang B (2009) Comparison of the effects of NaCl and KCl at the roots on seedling growth, cell death and the size, frequency and secretion rate of salt glands in leaves of Limonium sinense. Acta Physiol Plant 31:343–350
Edelist C, Raffoux X, Falque M, Dillmann C, Sicard D, Rieseberg LH, Karrenberg S (2009) Differential expression of candidate salt-tolerance genes in the halophyte Helianthus paradoxus and its glycophyte progenitors H. annuus and H. petiolaris (Asteraceae). Am J Bot 96:1830–1838
Epstein E, Bloom AJ (2005) Mineral nutrition of plants: principles and perspectives, 2nd edn. Sinauer Associates, Sunderland
Falhof J, Pedersen JT, Fuglsang AT, Palmqren M (2016) Plasma membrane H+-ATPase regulation in the center of plant physiology. Mol Plant 9:323–337
FAO (2009) High level expert forum—how to feed the world in 2050, economic and social development. Food and Agricultural Organization of the United Nations, Rome
Felle HH (1994) The H+/Cl- symporter in root hair cells of Sinapis alba (an electrophysiological study using ion-selective microelectrodes). Plant Physiol 106:1131–1136
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55:307–319
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179:945–963
Flowers TJ, Hajibagheri MA (2001) Salinity tolerance in Hordeum vulgare: ion concentrations in root cells of cultivars differing in salt tolerance. Plant Soil 231:1–9
Flowers TJ, Troke PF, Yeo AR (1977) The mechanism of salt tolerance in halophytes. Annu Rev Plant Physiol 28:89–121
Frachisse JM, Thomine S, Colcombet J, Guern J, Barbier-Bryqoo H (1999) Sulfate is both a substrate and an activator of the voltage-dependent anion channel of Arabidopsis hypocotyl cells. Plant Physiol 121:253–262
Franklin JA, Zwiazek JJ (2004) Ion uptake in Pinus banksiana treated with sodium chloride and sodium sulphate. Physiol Plant 120:482–490
Fricke W, Leigh RA, Tomos AD (1994) Epidermal solute concentrations and osmolality in barley leaves studied at the single-cell level – changes along the leaf blade, during leaf aging and NaCl stress. Planta 192:317–323
Fricke W, Leigh RA, Tomos AD (1996) The intercellular distribution of vacuolar solutes in the epidermis and mesophyll of barley leaves changes in response to NaCl. J Exp Bot 47:1413–1426
Fukuda A, Nakamura A, Hara N, Toki S, Tanaka Y (2011) Molecular and functional analyses of rice NHX-type Na+/H+ antiporter genes. Planta 233:175–188
Gálvez FJ, Baghour M, Hao G, Cagnac O, Rodríguez-Rosales MP, Venema K (2012) Expression of LeNHX isoforms in response to salt stress in salt sensitive and salt tolerant tomato species. Plant Physiol Biochem 51:109–115
Garciadeblás B, Senn ME, Banuelos MA, Rodríguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801
Gimeno V, Syvertsen JP, Nieves M, Simón I, Martínez V, García-Sánchez F (2009) Additional nitrogen fertilization affects salt tolerance of lemon trees on different rootstocks. Sci Hortic 121:298–305
Glenn EP, Brown JJ, Blumwald E (1999) Salt tolerance and crop potential of halophytes. Crit Rev Plant Sci 18:227–255
Golldack D, Lüking I, Yang O (2011) Plant tolerance to drought and salinity: stress regulating transcription factors and their functional significance in the cellular transcriptional network. Plant Cell Rep 30:1383–1391
Gong Z, Koiwa H, Cushman MA, Ray A, Bufford D, Kore-eda S, Matsumoto TK, Zhu J, Cushman JC, Bressan RA, Hasegawa PM (2001) Genes that are uniquely stress regulated in salt overly sensitive (sos) mutants. Plant Physiol 126:363–375
Gong D, Guo Y, Schumaker KS, Zhu J-K (2004) The SOS3 family of calcium sensors and SOS2 family of protein kinases in Arabidopsis. Plant Physiol 134:919–926
Grattan SR, Grieveb CM (1998) Salinity-mineral nutrient relations in horticultural crops. Sci Hortic 78:127–157
Hajibagheri MA, Flowers TJ (1989) X-ray microanalysis of ion distribution within root cortical cells of the halophyte Suaeda maritima (L.) Dum. Planta 177:131–134
Hajibagheri MA, Yeo AR, Flowers TJ, Collins JC (1989) Salinity resistance in Zea mays: fluxes of potassium, sodium and chloride, cytoplasmic concentrations and microsomal membrane lipids. Plant Cell Environ 12:753–757
Hasanuzzaman M, Nahar K, Alam MM, Bhowmi, Hossain MA, Rahman MM, Prasad MNV, Ozturk M, Fujita M (2014) Potential use of halophytes to remediate saline soils. Bio Med Resh Int. https://doi.org/10.1155/2014/589341
Hasegawa PM (2013) Sodium (Na+) homeostasis and salt tolerance of plants. Environ Exp Bot 92:19–31
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Biol 51:463–499
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K+/Na+ ratio in leaves during salinity stress. Plant Cell Environ 33:552–565
Hightower M, Pierce SA (2008) The energy challenge. Nature 452:285–286
Hillel D (2000) Salinity management for sustainable irrigation: integrating science, environment, and economics. World Bank Publications
Hillel D (2005) Soil salinity: historical and contemporary perspectives. In: Proceedings of the International Salinity Forum, Riverside, p. 235–240
Horie ST, Motoda J, Kubo M, Yang H, Yoda K, Horie R, Wai-Yin C, Ho-Yin L, Hattori K, Konomi M, Osumi M, Osumi M, Yamagami M, Schroeder JI, Uozumi N (2005) Enhanced salt tolerance mediated by AtHKT1 transporter-induced Na+ unloading from xylem vessels to xylem parenchyma cells. Plant J 44:928–938
Hua BG, Mercier RW, Leng Q, Berkowitz GA (2003) Plants do it differently. A new basis for potassium/sodium selectivity in the pore of an ion channel. Plant Physiol 132:1353–1361
Huang CX, Van Steveninck RF (1989) Maintenance of low Cl concentrations in mesophyll cells of leaf blades of barley seedlings exposed to salt stress. Plant Physiol 90:1440–1443
Ishitani M, Liu J, Halfter U, Kim CS, Shi W, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12:1667–1678
James RA, Davenport RJ, Munns R (2006) Physiological characterization of two genes for Na+ exclusion in durum wheat, Nax1 and Nax2. Plant Physiol 142:1537–1547
James RA, Blake C, Byrt CSm Munns R (2011) Major genes for Na+ exclusion, Nax1 and Nax2 (wheat HKT1; 4 and HKT1; 5), decrease Na+ accumulation in bread wheat leaves under saline and waterlogged conditions. J Exp Bot 62:2939–2947
Janicka-Russak M, Kabała K (2015) The role of plasma membrane H+-ATPase in salinity stress of plants. In: Progress in Botany. Springer, Cham/Switzerland, pp 77–92
Jeschke WD, Klagges S, Hilpert A, Bhatti AS, Sarwar G (1995) Partitioning and flows of ions and nutrients in salt-treated plants of Leptochloa fusca L. Kunth. I. Cations and chloride. New Phytol 130:23–35
Kobayashi NI, Yamaji N, Yamamoto H, Okubo K, Ueno H, Costa A, Tanoi K, Matsumura H, Fujii-Kashino M, Horiuchi T, Al Nayef M, Shabala S, An G, Ma JF, Horie T (2017) OsHKT1;5 mediates Na+ exclusion in the vasculature to protect leaf blades and reproductive tissues from salt toxicity in rice. Plant J 91:657–670
Kronzucker HJ, Britto DT (2011) Sodium transport in plants: a critical review. New Phytol 189:54–81
Kronzucker HJ, Szczerba MW, Schulze LM, Britto DT (2008) Non-reciprocal interactions between K+ and Na+ ions in barley (Hordeum vulgare L.) J Exp Bot 59:2793–2801
Kulik A, Wawer I, Krzywińska E, Bucholc M, Dobrowolska G (2011) SnRK2 protein kinases—key regulators of plant response to abiotic stresses. OMICS 15:859–872
Kumar K, Kumar M, Kim S-R, Ryu H, Cho Y-G (2013) Insights into genomics of salt stress response in rice. Rice 6:27
Lan W-Z, Wang W, Wang S-M, Li L-G, Buchanan BB, Lin H-X, Gao J-P, Luan S (2010) A rice high-affinity potassium transporter (HKT) conceals a calcium-permeable cation channel. Proc Natl Acad Sci 107:7089–7094
Leidi EO, Barragán V, Rubio L, El-Hamdaoui A, Ruiz MT, Cubero B, Fernández JA, Bressan RA, Hasegawa PM, Quintero FJ, Pardo JM (2010) The AtNHX1 exchanger mediates potassium compartmentation in vacuoles of transgenic tomato. Plant J 61(3):495–506
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108:253–260
Liu J, Ishitani M, Halfter U, Kim CS, Zhu JK (2000) The Arabidopsis thaliana SOS2 gene encodes a protein kinase that is required for salt tolerance. Proc Natl Acad Sci 97:3730–3734
Lorenzen I, Aberle T, Plieth C (2004) Salt stress-induced chloride flux: a study using transgenic Arabidopsis expressing a fluorescent anion probe. Plant J 38:539–544
Luo Q, Yu B, Liu Y (2005) Differential selectivity to chloride and sodium ions in seedlings of Glycine max and G. soja under NaCl stress. J Plant Physiol 162:1003–1012
Ma L, Zhang H, Sun L, Jiao Y, Zhang G, Miao C, Hao F (2011) NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na+/K+ homeostasis in Arabidopsis under salt stress. J Exp Bot 63:305–317
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444:139–158
Mahajan S, Sopoy SK, Tuteja N (2006) CBL–CIPK paradigm: role in calcium and stress signaling in plants. Proc Indian Natl Sci Acad U S A 72:63–78
Marschner H (1995) Mineral nutrition of higher plants, 2nd edn. Academic, London
Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu J-K, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012
Mian A, Oomen RJ, Isayenkov S, Sentenac H, Maathuis FJ, Véry AA (2011) Over-expression of an Na+-and K+-permeable HKT transporter in barley improves salt tolerance. Plant J 68:468–479
Moya JL, Gomez-Cadenas A, Primo-Millo E, Talon M (2003) Chloride absorption in salt-sensitive Carrizo citrange and salt tolerant Cleopatra mandarin citrus rootstocks is linked to water use. J Exp Bot 54:825–833
Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167:645–663
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364
Nieves-Cordones M, Miller AJ, Alemán F, Martinez V, Rubio F (2008) A putative role for the plasma membrane potential in the control of the expression of the gene encoding the tomato high-affinity potassium transporter HAK5. Plant Mol Biol 68:521–532
Niu X, Bressan RA, Hasegawa PM, Pardo JM (1995) Ion homeostasis in NaCl stress environments. Plant Physiol 109:735–742
Oh DH, Leidi E, Zhang Q, Hwang SM, Li Y, Quintero FJ, Jiang X, D'Urzo MP, Lee SY, Zhao Y, Bahk JD, Bressan RA, Yun DJ, Pardo JM, Bohnert HJ (2009) Loss of halophytism by interference with SOS1 expression. Plant Physiol 151:210–222
Oh DH, Dassanayake M, Haas JS, Kropornika A, Wright C, d'Urzo MP, Hong H, Ali S, Hernandez A, Lambert GM, Inan G, Galbraith DW, Bressan RA, Yun DJ, Zhu JK, Cheeseman JM, Bohnert HJ (2010) Genome structures and halophyte-specific gene expression of the extremophile Thellungiella parvula in comparison with Thellungiella salsuginea (Thellungiella halophila) and Arabidopsis. Plant Physiol 154:1040–1052
Oh DH, Dassanayake M, Bohnert HJ, Cheeseman JM (2012) Life at the extreme: lessons from the genome. Genome Biol 13:241
Olias R, Eljakaoui Z, Li J, De Morales PA, Marín-Manzano MC, Pardo JM, Belver A (2009) The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs. Plant Cell Environ 32:904–916
Otoch MLO, Sobreira ACM, de Aragão MEF, Orellano EG, Lima MGS, de Melo DF (2001) Salt modulation of vacuolar H+-ATPase and H+-pyrophosphatase activities in Vigna unguiculata. J Plant Physiol 158:545–551
Pardo JM, Rubio F (2011) Na+ and K+ transporters in plant signaling. In: Transporters and pumps in plant signaling. Springer, Berlin, pp 65–98
Park J, Okita TW, Edwards GE (2009) Salt tolerant mechanisms in single-cell C4 species Bienertia sinuspersici and Suaeda aralocaspica (Chenopodiaceae). Plant Sci 176:616–626
Peleg Z, Apse MP, Blumwald E (2011) Engineering salinity and water-stress tolerance in crop plants: getting closer to the field. Adv Bot Res 57:405–443
Pérez-Alfocea F, Ghanem ME, Gómez-Cadenas A, Dodd IC (2011) Omics of root-to-shoot signaling under salt stress and water deficit. OMICS 15:893–901
Platten DJ, Cotsaftis O, Berthomieu P, Bohnert H, Davenport RJ, Fairbairn DJ, Horie T, Leigh RA, Lin H-X, Luan S, Maser P, Pantoja O, RodriguezNavarro A, Schachtman DP, Schroeder JI, Sentenac H, Uozumi N, Very AA, Zhu JK, Dennis ES, Tester M (2006) Nomenclature for HKT transporters, key determinants of plant salinity tolerance. Trends Plant Sci 11:372–374
Plett DC, Møller IS (2009) Na+ transport in plants: what we know and would like to know. Plant Cell Environ 33:612–626
Plett CD, Møller IS (2010) Na+ transport in glycophytic plants: what we know and would like to know. Plant Cell Environ 33:612–626
Pottosin I, Dobrovinskaya O (2014) Non-selective cation channels in plasma and vacuolar membranes and their contribution to K+ transport. J Plant Physiol 171:732–742
Qi Z, Spalding EP (2004) Protection of plasma membrane K+ transport by the salt overly sensitive1 Na+-H+ antiporter during salinity stress. Plant Physiol 136:2548–2555
Qin F, Shinozaki K, Yamaguchi-Shinozaki K (2011) Achievements and challenges in understanding plant abiotic stress responses and tolerance. Plant Cell Physiol 52:1569–1582
Quintero FJ, Martinez-Atienza J, Villalta I, Jiang X, Kim W-Y, Ali Z, Fujii H, Mendoza I, Yun D-J, Zhu J-K, Pardo JM (2011) Activation of the plasma membrane Na/H antiporter salt-overly-sensitive 1 (SOS1) by phosphorylation of an auto-inhibitory C-terminal domain. Proc Natl Acad Sci 108:2611–2616
Rahnama A, James RA, Poustini K, Munns R (2010) Stomatal conductance as a screen for osmotic stress tolerance in durum wheat growing in saline soil. Funct Plant Biol 37:255–263
Reddy MP, Sanish S, Iyengar ERR (1992) Photosynthetic studies and compartmentation of ions in different tissues of Salicornia brachiata under saline conditions. Photosynthetica (CSFR)
Reddy AS, Ali GS, Celesnik H, Day IS (2011) Coping with stresses: roles of calcium-and calcium/calmodulin-regulated gene expression. Plant Cell 23:2010–2032
Ren Z-H, Gao J-P, Li L-G, Cai X-L, Huang W, Chao D-Y, Zhu M-Z, Wang Z-Y, Luan S, Lin H-X (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1147
Rengasamy P (2002) Transient salinity and subsoil constraints to dryland farming in Australian sodic soils: an overview. Aust J Exp Agric 42:351–361
Rengasamy P (2006) World salinization with emphasis on Australia. J Exp Bot 57:1017–1023
Rengasamy P (2010) Soil processes affecting crop production in salt-affected soils. Funct Plant Biol 37:613–620
Roberts SK, Tester M (1997) Permeation of Ca2+ and monovalent cations through an outwardly rectifying channel in maize root stelar cells. J Exp Bot 48:839–846
Roberts SK (2006) Plasma membrane anion channels in higher plants and their putative functions in roots. New Phytol 169:647–666
Rogers ME, Noble CL, Pederick RJ (1997) Identifying suitable forage legume species for saline areas. Aus J Exp Agric 37:639–645
Romero-Aranda R, Moya JL, Tadeo FR, Legaz F, Primo-Millo E, Talon M (1998) Physiological and anatomical disturbances induced by chloride salts in sensitive and tolerant citrus: beneficial and detrimental effects of cations. Plant Cell Environ 21:1243–1253
Rozema J, Flowers T (2008) Crops for a salinized world. Science 322:1478–1480
Ruan CJ, da Silva JAT, Mopper S, Qin P, Lutts S (2010) Halophyte improvement for a salinized world. Crit Rev Plant Sci 29:329–359
Ruiz-Lozano JM, Porcel R, Azcón C, Aroca R (2012) Regulation by arbuscular mycorrhizae of the integrated physiological response to salinity in plants: new challenges in physiological and molecular studies. J Exp Bot 63:4033–4044
Rus A, Lee B-h, Muñoz-Mayor A, Sharkhuu A, Miura K, Zhu J-K, Bressan RA, Hasegawa PM (2004) AtHKT1 facilitates Na+ homeostasis and K+ nutrition in planta. Plant Physiol 136:2500–2511
Sánchez-Barrena MJ, Martínez-Ripoll M, Zhu JK, Albert A (2005) The structure of the Arabidopsis thaliana SOS3: molecular mechanism of sensing calcium for salt stress response. J Mol Biol 345:1253–1264
Sánchez-Barrena M, Fujii H, Angulo I, Martinez-Ripoll M, Zhu J-K, Albert A (2007) The structure of the C-terminal domain of the protein kinase AtSOS2 bound to the calcium sensor AtSOS3. Mol Cell 26:427–435
Sanders D (2000) Plant biology: the salty tale of Arabidopsis. Curr Biol 10:486–488
Sairam RK, Tyagi A (2004) Physiology and molecular biology of salinity stress tolerance in plants. Curr Sci 86:407–421
Schroeder JI, Delhaize E, Frommer WB, Guerinot ML, Harrison MJ, Herrera-Estrella L, Horie T, Kochian LV, Munns R, Nishizawa NK, Tsay Y-F, Sanders D (2013) Using membrane transporters to improve crops for sustainable food production. Nature 497:60–66
Serrano R, Mulet JM, Rios G, Marquez JA, de Larrinoa IF, Leube MP, Mendizabal I, Pascual-Ahuir A, Proft M, Ros R, Montesinos C (1999) A glimpse of the mechanisms of ion homeostasis during salt stress. J Exp Bot 50:1023–1036
Shabala S, Cuin TA (2008) Potassium transport and plant salt tolerance. Physiol Plant 133:651–669
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279
Shabala S, Demidchik V, Shabala L, Cuin TA, Smith SJ, Miller AJ, Davies JM, Newman IA (2006) Extracellular Ca2+ ameliorates NaCl-induced K+ loss from Arabidopsis root and leaf cells by controlling plasma membrane K+-permeable channels. Plant Physiol 141:1653–1665
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci 97:6896–6901
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477
Sibole JV, Cabot C, Poschenrieder C, Barcelo J (2003) Efficient leaf partitioning, an overriding condition for abscisic acidcontrolled stomatal and leaf growth responses to NaCl salinization in two legumes. J Exp Bot 54:2111–2119
Singh K, Singla-Pareek SL, Pareek A (2011) Dissecting out the crosstalk between salinity and hormones in roots of Arabidopsis. OMICS 15:913–924
Song J, Chen M, Feng G, Jia Y, Wang B, Zhang F (2009) Effect of salinity on growth, ion accumulation and the roles of ions in osmotic adjustment of two populations of Suaeda salsa. Plant Soil 314:133–141
Storey R, Walker RR (1998) Citrus and salinity. Sci Hortic 78:39–81
Storey R, Schachtman DP, Thomas MR (2003) Root structure and cellular chloride, sodium and potassium distribution in salinised grapevines. Plant Cell Environ 26:789–800
Suzuki N, Koussevitzky S, Mittler R, Miller G (2012) ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ 35:259–270
Swarajyalakshmi G, Gurumurthy P, Subbaiah GV (2003) Soil salinity in South India: problems and solutions. J Crop Prod 7:247–275
Sykes SR (1992) The inheritance of salt exclusion in woody perennial fruit species. Plant Soil 146:123–129
Takeda S, Matsuoka M (2008) Genetic approaches to crop improvement: responding to environmental and population changes. Nat Rev Genet 9:444–457
Teakle NL, Tyerman SD (2010) Mechanisms of Cl-transport contributing to salt tolerance. Plant Cell Environ 33:566–589
Teakle NL, Real D, Colmer TD (2006) Growth and ion relations in response to combined salinity and waterlogging in the perennial forage legumes Lotus corniculatus and Lotus tenuis. Plant Soil 289:369–383
Teakle N, Flowers T, Real D, Colmer T (2007) Lotus tenuis tolerates the interactive effects of salinity and waterlogging by ‘excluding’ Na+ and Cl- from the xylem. J Exp Bot 58:2169–2180
Teodoro AE, Zingarelli L, Lado P (1998) Early changes of Cl− efflux and H+ extrusion induced by osmotic stress in Arabidopsis thaliana cells. Physiol Plant 102:29–37
Tracy FE, Gilliham M, Dodd AN, Webb AA, Tester M (2008) NaCl-induced changes in cytosolic free Ca2+ in Arabidopsis thaliana are heterogeneous and modified by external ionic composition. Plant Cell Environ 31:1063–1073
Van Steveninck RFM, Van Steveninck ME, Stelzer R, Läuchli A (1982) Studies on the distribution of Na and Cl in two species of lupin (Lupinus luteus and Lupinus angustifolius) differing in salt tolerance. Physiol Plant 56:465–473
Very AA, Sentenac H (2003) Molecular mechanisms and regulation of K+ transport in higher plants. Annu Rev Plant Biol 54:575–603
Volkov V, Amtmann A (2006) Thellungiella halophila, a salt tolerant relative of Arabidopsis thaliana, has specific root ionchannel features supporting K+/Na+ homeostasis under salinity stress. Plant J 48:342–353
Wang B, Lüttge U, Ratajczak R (2001) Effects of salt treatment and osmotic stress on V-ATPase and V-PPase in leaves of the halophyte Suaeda salsa. J Exp Bot 52:2355–2365
Wang Y, Xiao D, Li Y, Li X (2008) Soil salinity evolution and its relationship with dynamics of groundwater in the oasis of inland river basins: case study from the Fubei region of Xinjiang Province, China. Environ Monit Assess 140:291–302
Wang CM, Zhang JL, Liu XS, Li Z, Wu GQ, Cai JY, Flowers TJ, Wang SM (2009) Puccinellia tenuiflora maintains a low Na+ level under salinity by limiting unidirectional Na+ influx resulting in a high selectivity for K+ over Na+. Plant Cell Environ 32:486–496
White PJ, Broadley MR (2001) Chloride in soils and its uptake and movement within the plant: a review. Ann Bot 88:967–988
Winter E (1982) Salt tolerance of Trifolium alexandrinum L. II. Ion balance in relation to its salt tolerance. Aus J Plant Physiol 9:227–237
Xu G, Magen H, Tarchitzky J, Kafkafi U (1999) Advances in chloride nutrition of plants. Adv Agron 68:97–150
Yadav S, Irfan M, Ahmad A, Hayat S (2011) Causes of salinity and plant manifestations to salt stress: a review. J Environ Biol 32:667–685
Yen HE, Wu S-M, Hung Y-H, Yen S-K (2000) Isolation of 3 salt-induced low-abundance cDNAs from light-grown callus of Mesembryanthemum crystallinum by suppression subtractive hybridization. Physiol Plant 110:402–409
Zhang J-L, Flowers TJ, Wang S-M (2010) Mechanisms of sodium uptake by roots of higher plants. Plant Soil 326:45–60
Zhang H, Han B, Wang T, Chen S, Li H, Zhang Y, Dai S (2012) Mechanisms of plant salt response: insights from proteomics. J Proteome Res 11:49–67
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zhu JK (2003) Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol 6:441–445
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Farooq, S., Ahmad, S., Hussain, S., Hussain, M. (2018). Nutrient Homeostasis and Salt Stress Tolerance. In: Hasanuzzaman, M., Fujita, M., Oku, H., Nahar, K., Hawrylak-Nowak, B. (eds) Plant Nutrients and Abiotic Stress Tolerance. Springer, Singapore. https://doi.org/10.1007/978-981-10-9044-8_17
Download citation
DOI: https://doi.org/10.1007/978-981-10-9044-8_17
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-9043-1
Online ISBN: 978-981-10-9044-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)