Abstract
Salinity, recognized as a major threat in agriculture, causes 4.0–6.3% yield loss annually across the world. The problem is aggravated due to increasing irrigation with suboptimal quality of irrigation water and more salinization of coastal area due to the rise in sea level because of climate change. In saline soil, excessive concentrations of Na+ and Cl− impair absorption of other beneficial ions such as K+ and Ca2+ that in turn inhibit plant growth and productivity. Maintenance of cellular K+ level and K+/Na+ ratio is still considered the most important factor for salt tolerance. Under high-Na+ environment, excess Na+ competes with K+ thereby hindering its uptake. Tolerant plants by employing a number of strategies restrict Na+ movement to young meristematic tissues and allow greater movement and/or tissue retention of K+ to physiologically more active tissues. Under salt stress different K+- and Na+-specific transporters, viz. SOS, NHX, and HKT family transporters (regulate cellular Na+ movement) and HAK, AKT, KT, and KUP (regulate K+ movement), either by upregulation or downregulation, control the cellular ion homeostasis and salt tolerance in plants. SOS1, a plasma membrane-bound Na+/H+ antiporter, mostly active in root tissue, removes the excess salt from the plant body by pumping them back to the rhizosphere in an energy-dependent process. Tonoplast-bound vacuolar Na+/H+ antiporters (NHX family transporters) play crucial role in Na+ compartmentalization inside the vacuole in mature cell in both root and leaf tissues. Storing excess salts in vacuole imparts tolerance in multifaceted manner, viz. imparting tissue and osmo-tolerance. Biosynthesis of organic osmolytes, a more energy-expensive process, is sometimes substituted by the accumulation of excess Na+ in non-active tissues under salt stress. Improved Ca2+ status inside the plant tissue is another important factor associated with salt tolerance and acts as a key signalling molecule to initiate Na+ exclusion. Several QTLs and miRNAs were reported to impart salt tolerance in several crops. Managing salinity beyond crop improvement strategies was also deliberated, e.g. lowering salt effect through K+ supplementation and phytohormones, etc. In this compilation, emphasis has been given on how nutrient/ionic imbalance causes deleterious effects on plants under saline conditions and what are the possible adaptive strategies plants employ to maintain the ionic homeostasis in saline environment.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Salinity
- Na+-K+ transporter
- Osmolytes
- Tissue tolerance
- ROS detoxification
- Salt overly sensitive (SOS) pathway
14.1 Introduction
In the last few decades, we witnessed substantial increase in productivity of food grains, oilseeds, pulses and cash crops mostly through adoption of intensive agriculture, viz. high-yielding varieties, precise fertilizer and nutrient management practices, more efficient crop protection measures, etc. But, with the continuous increase in global population by every passing year, there is an estimated need to produce 87% more food crops such as rice, wheat and maize by 2050 over that we are producing today (FAO 2017). As the horizontal area expansion in fertile agricultural land almost came to a saturation in most countries, there is a current shift in encompassing more and more nontraditional areas under cultivation to cater the global food demand. Despite the much advancement in agricultural science in all over the world, abiotic stresses still cause havoc on cultivation due to its widespread and unmanageable nature, including salinity, drought, heat and cold, critically threaten crop production and result in substantial yield loss in large arable land worldwide. Among these, soil salinity is one of the prime environmental constraints to crop production and is further expected to increase due to global climate changes (increase in coastal salinity mainly due to the rise in mean sea level) and as a consequence of injudicious and/or faulty irrigation practices. As per the recent estimate, about 800 million hectares of land globally is affected by salinity (FAO 2017). On an average 2,000 ha of irrigated land across 75 countries has been degraded by excess salt annually owing an estimated economic loss in the tune of US$ 12 billion (Ghassemi et al. 1995).
Soil salinization is a worldwide problem for agriculture affecting 6% of total Earth’s land, as a result of natural accumulation over long periods of time (Rengasamy 2002). However, agricultural activity contributes to secondary salinization: 2% of all dry land is becoming salinized, and more than 20% of irrigated soils are affected, mostly because of irrigation water containing small amounts of sodium chloride (Tester and Davenport 2003). Saline soils in general affect plant growth negatively and may even have a lethal effect causing programmed cell death upon extended exposure to high salinity. Based on the ability to tolerate NaCl concentrations, plants can be classified into two groups: glycophytes or salt-sensitive species (which are unable to tolerate even mild levels of salinity for longer periods of time) and halophytes or salt-tolerant species (which are capable of growing and thriving under high salinity). Primarily, excess salt in the soil decreases the water potential in the rhizosphere region, rendering plants unable to absorb water even in the absence of actual limitation of water quantity, a soil condition termed as physiological drought. As a result, many plant processes, viz. at the cellular level including cell enlargement, cell division, cell wall properties, etc., as well as various leaf parameters such as colour, succulence, necrosis, etc., and at whole-plant level, shoot/root ratio, growth and yield get affected (Hasegawa et al. 2000).
For most of the glycophytes, highly saline growing environment adversely affects the germination process, plant growth and metabolism as well as the overall physiology by causing ionic and osmotic stresses (Iterbe-Ormaetxe et al. 1998). Salt stress is often noted as a causal factor for increased respiration rate and ion toxicity while subsequently altering the C and N metabolism in plant cell (Kim et al. 2004). Additionally, mineral distribution and membrane instability (Marschner 1986) along with permeability (Gupta et al. 2002) and decreased biosynthesis of chlorophyll pigments and photosynthetic inefficiency (Munns 2002), all of which are caused by salt stress, collectively lead to impaired economic crop productivity. Stress-induced build-up of sugars and other compatible organic solutes is a common phenomenon for most of the abiotic stresses including soil salinity. They can serve as osmoprotectants, thus helping in stabilizing biomolecules under stress conditions. Although accumulation of ions for osmotic adjustment is energetically more preferable, many plants accumulate organic osmolytes (proline, betaine, polyols, sugar alcohols and soluble sugars) to counteract osmotic stresses. Both glycine betaine and trehalose serve as major osmoprotectants which stabilize the quaternary structures of proteins and highly ordered cellular and intracellular membranes. Proline acts as a sink for carbon and nitrogen and scavenger of free radical, which stabilizes subcellular structures (membranes and proteins) and maintains cellular redox potential (Reviewed in Chakraborty et al. 2013).
Salt stress also induces accumulation of reactive oxygen species (ROS), causing oxidative damage to cellular macromolecules, viz. proteins, membrane lipids and nucleic acids. Detoxification of these ROS is absolutely essential for plants to survive under salinity stress. To counterbalance the ROS production and oxidative stress, plants produce either molecular antioxidants for direct scavenging of these ROS or detoxify them via coordinated network of antioxidant enzymes, viz. superoxide dismutase, catalases, peroxidases and enzymes of ascorbate-glutathione cycle. Under various abiotic stresses, the activity and expression levels of genes encoding ROS-detoxifying enzymes were reported to be enhanced by oxidative stress (Abogadallah 2010; Chakraborty et al. 2016a).
Ionic homeostasis or regulation of Na+/K+ balance inside metabolically active tissue is still considered to be the most important criteria for salt tolerance (Munns and Tester 2008). Exposure to higher levels of salt stress, particularly NaCl, affects uptake of water and dissolved nutrients resulting in impaired plant water status and creates ionic imbalance by means of the cellular accumulation of toxic Na+ and Cl− ions. Sodium ions if accumulated in the cytoplasm can become extremely toxic to living cells showing their adverse effects on K+ nutrition and other pivotal plant physiological mechanisms like activity of cytosolic enzymes, photosynthesis and metabolism (Shabala and Cuin 2008; Degl’Innocenti et al. 2009). Besides, salt stress heavily tolls on the ionic homeostasis of other complementary ions such as Ca2+, Mg2+ and NO3 − , and therefore, further investigation requires altered transport and compartmentation mechanism of these nutrients under salinity stress. In plants, predominantly three distinct but complementary mechanisms operate cooperatively that selectively inhibit the accumulation of Na+ in the cytoplasm following one or other processes, i.e. checking of Na+ influx, promotion of active Na+ efflux and sequestration of Na+ in the vacuole, which will be discussed categorically in this compilation. But before that we need to understand various soil-related factors affecting availability of nutrients under salt stress.
14.2 Soil-Driven Factors Affecting Nutrient Availability Under Salt Stress
14.2.1 Salinity: Origin and Extent
Salinity is predominant in two major forms over the land surface, (1) dry land salinity and (2) irrigation salinity, and arid and semiarid climatic regions throughout the world are suffering due to salinity in one form or another. The dry land salinity is often detected as primary or secondary salting, either occurring naturally (naturally occurring saline wet and dry lands including salt lakes, salt pans, salt marshes and salt flats) or via secondary salting that is induced by human activities such as agriculture (Fig. 14.1). Secondary salinity is majorly caused by anthropogenic activities, while practising land development and agriculture may play some role (Queensland Government 1995–2017), and the common forms are:
-
Irrigation: prevalent in irrigated agricultural lands, due to excessive irrigation (results into rising groundwater tables) or the application of poor-quality water.
-
Dry land: prevalent in rainfed or nonirrigated landscapes, generally as a result of deforestation and land-use changes. Irrigation salinity often resembles dry land salinity, except that inclined level of groundwater that also resulted in deposition of salt layers in the plant root zone or on the soil surface.
-
Sea water intrusion: In coastal aquifer systems, it is commonly found that fresh groundwater is gradually replaced by sea water.
-
Point source: originated from the high concentration of diluted salt in effluent either released from intensive agriculture loaded with pesticide/chemical residues or from polluted wastewater stream from industries.
Over the years, several estimations have been published regarding the extent of salinity. Oldeman et al. (1991) reported that the total area affected by waterlogging was over 10 m ha and that affected by salinity was over 76 m ha. They counted both irrigated and rainfed areas. Dregne et al. (1991) published that about 43 m ha of irrigated land in dry lands was affected by several forms of degradation, including waterlogging, salinization and sodicity. Umali (1993) estimated that 1–1.5 m ha of lands were lost to salinization every year. Further it had been reported that nearly 12 m ha of irrigated land may have phased out from production due to salinization (Nelson and Maredia 2001). An approximate area of 7 m ha of land is estimated to be under saline soil in India (Patel et al. 2011). These lands are classified in Table 14.1.
14.2.2 Salinity Impacts on Crop Production
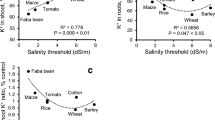
Agriculture is one of prime importance as far as salinity hazard is concerned. Crops may differ in their tolerance to salinity, and some of them are extremely sensitive, while few perform better even after crossing the threshold of marginal salinity and emerge as tolerant crops in terms of salinity stress. Even varietal differences are also prominent in major field crops. A report published from World Bank showed the degree of loss and major impacts due to salinity or other forms of soil degradation as a whole (Table 14.2; Fig. 14.2).
The most promising impacts of soil salinity (FAO 2017)
14.2.3 Salinity Impacts on Nutrient Mobility in Soil and Plants
Besides the crop yield perspective, soil salinity has its own demerits creating problems like nutrient loss, nutrient imbalance, poor soil structure and health, and soil degradation. Dry land salinity is often considered as a major soil degradation issue, including soil erosion. Salinity is often associated with prolonged wetness, sparse vegetation and lack of surface cover and therefore increases the vulnerability of soils to erosion. Salt concentration in the soil solution (salinity) which governs the osmotic potential and the concentration of sodium on the exchange complex sites (sodicity) further determines soil structural stability. Thus, salinity slowly turns into sodicity. The major soluble salts in soils are the cations like Na+ (sodium), Ca2+ (calcium), Mg2+ (magnesium) and K+ (potassium) and the anions like Cl− (chloride), SO4 2− (sulphate), HCO3 − (bicarbonate), CO3 2− (carbonate) and NO3 − (nitrate) (Shi et al. 2005). These are the basic ions that dominate in the exchange sites under salinity and thus compromise the places of other essential nutrients like PO4 3− (phosphate) and micronutrients (Zn, Fe, Mn, Cu). This imbalance of nutrients created in soil is often measured by nutrient concentration or uptake by plants comparing the plants grown under normal and saline conditions. Similar occurrence was reported by Bhaduri et al. (2016) where P uptake of eight groundnut cultivars were studied and observed that the P uptake of groundnut cultivars is affected at irrigation salinity level of 6.0 ECiw. Salinity stress lowered down the N content in Brassica leaves as well as seed protein content; moreover the reduced accumulation of micronutrients (Fe, Mn, Zn) was also noticed in the leaf, stem and root at flowering and post-flowering stages (Chakraborty et al. 2016b).
Soil microorganisms, an integral component of soil ecosystem, are largely involved in an array of important soil nutrient cycling processes. Their roles in nitrification, ammonification, nitrogen fixation, P mineralization, S oxidation, decomposition of soil organic matter and transformation of all primary and secondary nutrients (Amato and Ladd 1994) are already established. Microbes also act in formation of humic substances which makes stable forms of organic C and contribute in C sequestration in soils. The high concentration of soluble salts affects the microbes by increasing the osmotic potential (more negative) of the soil water, promotes exosmosis and dehydrates the microbial cell. Thus, it makes difficult for microbes to survive and perform their basic functions in a saline soil. Even if they can survive under such stressful situation that needs more investment of energy for producing osmolytes. Till date, only few halophytic microbes and some endophytes are reported to tolerate such extreme saline conditions. Moreover, soil microbial community structure also differs significantly since fungi are more salt sensitive over bacteria, and thus bacteria/fungi ratio can be increased under saline soil environment (Wichern et al. 2006; Yan et al. 2015). All these phenomena either singly or collectively influence the soil nutrient availability.
14.3 Uptake of Regulatory Ions in Plant Cells: An Interplay of Nutrient Balance/Imbalance
Salinity is a much complex phenomenon rather than a simple escalation in the concentrations of sodium and chloride ions inside the plant tissue (Nouri et al. 2017). Apart from Na+ and Cl−, number of other cations and anions, viz. calcium, carbonates and sulphates, may be present in disproportionate amounts and play crucial role in negatively affecting plant growth (Gorham 1992). Simultaneously, certain nutrients (particularly potassium, nitrogen and phosphorus) may be available or present in such low amounts under saline condition that they might hamper proper growth (Chakraborty et al. 2016c).
Saline conditions affect plant growth and metabolism in many different ways. These harmful effects are generally associated with (1) reduced osmotic potential of the soil solution in plants (water stress), (2) nutritional imbalance, (3) effect exerted by a specific salt (salt stress) or (4) a combination of all of these factors (Ashraf and Foolad 2007; HanumanthaRao et al. 2016). These factors act in an adverse way affecting growth and development in plants at both physiological and biochemical levels (Munns 2002; Munns and Tester 2008) and also at the molecular level (Tester and Davenport 2003). Tolerance to saline conditions involves a myriad number of physiological processes manifested in numerous levels of organization, viz. alterations in gross morphology, tissue partitioning and coordinated control of transport, biological change for maintenance of protein structure and regulated transcriptome level changes (Tester and Davenport 2003).
14.3.1 Scenario of K+ vs Na+ and K+/Na+ Homeostasis Under Salt Stress
Sodium, an integral constituent of our Earth’s crust, is naturally present in all soil types. At lower concentration Na+ helps in supporting growth and development for some plants, but at higher concentration in soil or other growing medium, it eventually turns out to be toxic to even glycophytes (Flowers and Colmer 2008). Both Na+ and K+ share high similarity in ionic as well as its chemical and structural properties, but unlike Na+, K+ are integral part of plant’s life and play essential role in growth and development (Schachtman and Liu 1999). Many core physiological processes, primarily dependent on K+, show impairment due to hindrances in specific transport and interactions of K+ with enzymes and membrane proteins (Britto and Kronzucker 2008). This may be manifested as transient maintenance of membrane potential for stomatal movement and development of pollen tube in plants (Dietrich et al. 2001). Under saline condition, due to prolonged exposure to salt stress, plants often exhibit K+ deficiency symptoms majorly because of reduced uptake by the root tissue and/or lesser K+ retention in different plant parts coupled with a concomitant accumulation of tissue Na+ concentration (Munns et al. 2002). Under salt stress, plants with hindered growth and metabolism are observed due to the skewed K+/Na+ ratio in metabolically active plant tissues (Shabala and Cuin 2007; Degl’Innocenti et al. 2009). Because of such ionic imbalances, hindrances in various physiological and biochemical processes are observed in plants.
Under high external Na+ concentrations, Na+ enters through K+ pathway altering the ion ratios in plants. The similarity of the ionic radii of the hydrated molecule of Na+ and K+ renders the capability of discrimination between them much difficult and hence forms the basis of Na+ toxicity. Cellular K+ concentrations in the range of 100–150 mM are essential for in vitro protein biosynthesis. Moreover, at higher concentrations, Na+ competes for K+ sites and inhibits the whole process when Na+ concentrations is >100 mM (Cheeseman 2013). The similar level of sensitivity of cytosolic enzymes of halophytes and glycophytes towards saline conditions hinders the adaption of halophytes to high salt concentration (Flowers et al. 2014). Maintenance of low cytosolic Na+ concentrations and a high cytosolic K+/Na+ ratio is a key strategy adapted by plants to respond to elevated external Na+ concentrations (Blumwald et al. 2000). The approach towards such maintenance involves extrusion of Na+ or its compartmentalization mainly in the vacuoles for metabolism (Zhu 2003) and is critical for the detoxification of excess Na+ present in cytosol and the osmotic adjustment necessary to endure salt stress (Blumwald et al. 2000; Chakraborty et al. 2016d).
14.3.2 Regulation of Tissue Na+ Concentrations
14.3.2.1 Sodium Uptake
On exposure to salt stress, maintenance of low concentrations of Na+ and high concentrations of K+ in the cytosol becomes crucial and is achieved by controlled expression and activity of K+ and Na+ transporters (Shabala et al. 2015). Na+ enters the plant cells passively through the high-affinity K+ transporter HKT1 (Rus et al. 2001; Maser et al. 2002) and non-selective cation channels (NSCCs). Due to non-selectivity of a few transporters and/or ion channels, under highly saline conditions, Na+ ions compete with K+ ions for uptake and enter inside the plant through normal rhizospheric nutrient uptake process. At transcriptional level, these K+/Na+ transporter genes are either up- or downregulated as a response to salt stress (Chakraborty et al. 2016e). It has been reported that the transcript level of Arabidopsis root K+ transporter AtKC1 increases under salt stress (Pilot et al. 2003). As reported by Zhu (2003), upregulation in the expression level of KMT1 (a AKT/KAT family member) and various HAK/KUP (high-affinity K+ transporter/K+ uptake transporter)-type genes was observed, whereas for MKT1 (another AKT/KAT family member), the expression level was found to be downregulated for common ice plant.
14.3.2.2 Sodium Efflux
The primary mechanism of Na+ extrusion in case of plants is mediated by energy-driven active pumping out of Na+ by plasma membrane-bound Na+/H+ transporter and H+-ATPases (Zhu 2001). The H+-ATPase acts to pump H+ out of the cell using the energy of ATP hydrolysis, thus generating an electrochemical proton gradient. The proton-motive force thus generated is further required for the Na+/H+ antiporter operation as the inward movement of H+ along with the electrochemical gradient is coupled to the outward exclusion of Na+ against the electrochemical gradient. Confirmation of the existence of such biochemical mechanism has been documented for various plant species (Blumwald et al. 2000). Identification of a putative Na+/H+ antiporter with substantial similarity in sequence with plasma membrane Na+/H+ antiporters from bacteria and fungi has further strengthened the views. The SOS1 (salt overly sensitive 1) locus encoding a putative Na+/H+ antiporter having considerable sequence similarity to plasma membrane Na+/H+ antiporters from bacterial and fungal species has been identified in Arabidopsis (Shi et al. 2000), rice (Martinez-Atienza et al. 2007), wheat (Yang et al. 2009) and in Brassica (Chakraborty et al. 2012a).
14.3.2.3 Vacuolar Sodium Compartmentation
It has been observed that both halophytes and glycophytes regardless of the high influx of Na+ maintain the cytosolic concentration of the ion at non-toxic levels (Blumwald et al. 2000). The compartmentalization of Na+ into vacuoles has been found to be the primary mechanism of evading the harmful effects Na+ exerts in the cytosol. Additionally, the vacuolar compartmentalization of Na+ (and Cl−) allows the usage of NaCl as an osmoticum thereby contributing in maintenance of an osmotic potential for the process of water uptake into cells (Blumwald et al. 2000). Tonoplast-bound vacuolar Na+/H+ antiporters (NHX family transporters) play crucial role in Na+ compartmentalization inside the cell (Yokoi et al. 2002). Identification and characterization of several plant transporters have been made possible by detection of the higher degree of homology between several plant and yeast genes, and the detailed genetic information is available in the public domain (Halfter et al. 2000; Ji et al. 2013). Evidences suggest that Na+ detoxification mechanisms employed in yeast cells may be found to be quite similar to that existing in plant cells. This similarity mostly holds true for the role played by the Ca2+-dependent signal transduction mechanism which becomes operational under salinity stress (Halfter et al. 2000). Putative Na+/H+ antiporters (both SOS1 and NHX family) from both organisms are also similar (Apse et al. 1999; Fukuda et al. 1999).
14.3.3 Interaction Between Na+ and Ca2+
An important inorganic nutrient, calcium, plays a vital role in salt detoxification, in addition to its well-known metabolic and structural functions (Jin et al. 2007). This response stems from the fact that increasing Na+ concentrations may not only reduce Ca2+ availability but may also displace Ca2+ from its extracellular binding sites within the plant organs and further disrupt Ca2+ acquisition (Hadi and Karimi 2012). The interaction between Na+ and Ca2+ in salt-stressed plants has been the focus of several research agendas over the years (Cramer 2002; Nedjimi and Daoud 2009). It has been advocated that Na+ tolerance of plants is determined to a larger extent by interactions of Ca2+ and Na+ ions (Buschmann et al. 2000). It is reported that high NaCl induces calcium deficiencies in different plants such as Vigna unguiculata (Murillo-Amador et al. 2006) and tomato (Tuna et al. 2007). Allen et al. (1995) reported that Na+ influx on durum wheat cells can also be inhibited by calcium. According to Jin et al. (2007), saline conditions restricted Ca2+ uptake by the roots of Aloe vera plants and its subsequent transport to shoots resulting in a marked decrease in Ca2+ contents of all plant parts. It was further reported that in plants under salt stress, the Ca2+ contents of the leaves and stems show a noticeable decrease; salt-tolerant genotypes were found to exhibit three times higher Ca2+ concentrations as compared to salt-sensitive ones.
High Na+ concentration in the root zone was found to inhibit Ca2+ uptake and its transport resulting in lower Ca2+/Na+ ratios in salt-stressed plants (Hadi et al. 2008). Additionally, Jin et al. (2007) showed that salt-tolerant genotypes of Aloe vera maintained a significantly low Na+/Ca2+ ratio and experienced least membrane damage. The rapid Na+ uptake process across the plasma membrane in excess salt condition diminishes the binding capability of Ca2+ to the plasma membrane thus inhibiting its influx. High Na+ concentrations can displace Ca2+ in membrane thus disrupting the integrity of it (Janicka-Russak and Kłobus 2007). Hasegawa et al. (2000) in their report indicated that the increase in intercellular Ca2+ content could cause a decline in Na+ influx and in turn increase the K+ selectivity for absorption thus alleviating the damaging effects of salinity stress. They also pointed out that under salinity stress, Na+ can compete with Ca2+ by entering the cell through the same channels. The excess intercellular sodium can then displace the Ca2+ in the membranes causing membrane damage. Membrane-bound catalase activity could also be inhibited by excess Na+ which can be reversed by excess Ca2+ (Arbona et al. 2003). It has been proposed that Ca2+ plays a central role in plants exposed to NaCl salinity because of its active participation in reducing Na+ absorption and increasing potassium (K+) and Ca2+ uptake, resulting in an increase in plant growth (Caines and Shennan 1999). Additionally, Ca2+ may compete with Na+ for membrane-binding sites thereby shielding the cell membrane from the unfavourable saline conditions (Shabala et al. 2006).
The [Ca2+]ext augments salt tolerance by eliciting a transient increase in [Ca2+]ext either from a peripheral or an internal source (Knight et al. 1997). Experiments conducted on yeast have generated preliminary views of Ca2+-mediated activation of signalling pathways for regulation of ionic homeostasis and tolerance mechanisms in response to salt stress conditions. A suggestive model for salt-induced Ca2+ signalling and activated SOS pathway includes components of the SOS pathway; the SOS3 or other upstream elements might become connected with the osmotically responsive channel triggering Ca2+ influx which might possibly initiate signalling through the pathway (Chakraborty et al. 2016e). Reports suggests that salt-induced [Ca2+]ext transient as well as the new [Ca2+]ext steady state may be influenced by the ECA and ACA Ca2+-ATPases as well as the CAX1 and CAX2 transporters, the orthologs of VCX1P (Sze et al. 2000). Ca2+ plays two vital roles in conferring tolerance towards salinity, the fundamental signalling function leading to adaptation during salt stress conditions and a direct inhibitory effect on the entry of Na+ ions.
14.3.4 Transport and Xylem Loading
Na+ transport across the root and into the xylem occurs both symplastically and apoplastically from the epidermis to the xylem (Maathuis et al. 2014). Na+ export to the xylem is supposed to be an active process, given that the electric membrane potential of xylem parenchyma has been found to be negative. In Arabidopsis, under the conditions of salinity, xylem loading of Na+ was found to be mediated by SOS1 (salt overly sensitive1), while its unloading, on the other hand, was found to be a passive process, involving transportation through the Na+-permeable channels (Apse and Blumwald 2007). High-affinity K+ transporters or HKTs, classified in class I and class II types, were one of the most studied Na+-permeable transporters in plants (Horie et al. 2009). These HKT transporters, often located in the xylem parenchyma and root epidermal cells of many plants, exhibit a crucial role in adapting the plant to saline conditions for both mono- and dicotyledonous species (Møller et al. 2009; Munns et al. 2012). The class I HKT transporters showing specificity for mostly Na+ ions are characterized as low-affinity transporters (Munns and Tester 2008). Among the different subtypes of HKT1 transporter, a few are reported to be located in the plasma membrane of root stele cells, particularly in the xylem parenchyma cells (XPC), where their main function is to regain the Na+ ion from the xylem sap thereby avoiding transport and accumulation of toxic Na+ in the above ground plant parts and preventing damage to the more sensitive and photosynthetically active tissues (Ren et al. 2005).
14.4 Mechanisms of Nutrient Homeostasis: A Balancing Approach of Plants Facing Salt Stress
Physiological studies carried out in many crops during salt stress at vegetative stage indicated that stress tolerance trait inversely correlates with shoot Na+ concentration and Na+/K+ ratio (Ashraf 2004; Negrão et al. 2011). Different mechanisms associated with salt tolerance in crop plants include (1) maintenance of a more negative membrane potential, (2) intrinsically higher H+-ATPase activity, (3) extrusion of Na+ from the cytosol to the external medium, (4) maintenance of mineral nutrient homeostasis, particularly, higher selectivity to K+ and Ca2+ over Na+, (5) scavenging of ROS, (6) accumulation of compatible solutes for osmotic adjustment, etc. At the physiological level, salt tolerance and ion homeostasis are mostly governed by three major strategies in crop plants: (I) Na+ exclusion, (II) K+ retention and (III) tissue tolerance/Na+ sequestration (Munns and Tester 2008).
14.4.1 Electrophysiological Basis of Salt Tolerance: Role of Transporter/Pumps/Ion Channels
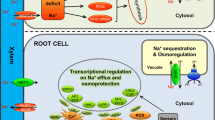
Plant salinity stress signalling is a complex phenomenon involving the interplay of many biomolecules ranging from receptor molecules, ion fluxes that serve as signals, transcription factors, hormones, reactive oxygen species (ROS) and numerous downstream proteins. In the cyanobacterium, Synechocystis sp., Marin et al. (2003) identified sensory histidine kinases, namely, HIK16, HIK33, HIK34 and HIK41, involved in the perception and transduction of salt stress. In plants, there is less clarity about the proteins that perceive salt stress. Salt overly sensitive (SOS) genes (SOS1-SOS4), first identified in Arabidopsis thaliana through positional cloning, are potential candidates for detecting elevated Na+ concentrations in intracellular and extracellular sites. The AtSOS1 protein is a putative plasma membrane Na+/H+ antiporter that regulates plant Na+ homeostasis by extrusion and is aided in its function by two other proteins SOS2 and SOS3 (Qiu et al. 2002; Zhu 2003). SOS-mediated salt stress signalling is represented in Fig. 14.3. The sos1, sos2 and sos3 mutants show salt sensitive phenotype, and their genetic analysis has helped to improve our understanding of the mechanism of salt stress tolerance in plants (Zhu et al. 1998). Yeast mutants lacking endogenous Na+ transporters were used to investigate the role of the three SOS proteins in salt-stress response pathway (Quintero et al. 2002). Perception of salt stress is followed by subtle changes in Ca2+ concentration in cytosol of root cells that triggers the SOS pathway (Guo et al. 2004; Chinnusamy et al. 2005). SOS3 is a myristoylated Ca2+-binding protein that recruits SOS2 serine-threonine protein kinase to the plasma membrane after binding of Ca2+ (Ishitani et al. 2000; Halfter et al. 2000). An alternative regulator of SOS2 activity, SOS3-like calcium-binding protein 8 (SCaBP8, a.k.a. calcineurin B-like CBL10), has been shown to function primarily in the shoots of Arabidopsis, while SOS3 expresses predominantly in roots (Quan et al. 2007). SOS2-mediated phosphorylation of SCaBP8 or SOS3-like proteins increases their stability (Lin et al. 2009). The SOS3-SOS2 or SCaBP8-SOS2 complex then recruits SOS2 to plasma membrane to activate downstream SOS1, which functions to extrude excess Na+ from the cytosol (Shi et al. 2000; Qiu et al. 2002; Quintero et al. 2002, 2011; Quan et al. 2007). SOS4 and SOS5 also play important roles in salt stress tolerance. While SOS4 encodes a pyridoxal kinase that is involved in regulation of Na+ and K+ homeostasis (Shi et al. 2002), SOS5 aids in the maintenance of normal cell expansion during stress (Shi et al. 2003).
Other Na+ transporters functioning in salinity tolerance include those involved in intracellular compartmentalization of Na+ into vacuoles, older leaves or leaf sheath, extrusion outside the cell and recirculation of Na+ out of the shoots to be stored elsewhere, for example, in roots or stem cell vacuoles. Vacuolar Na+ sequestration is one of the most energetically efficient mechanisms by which plants achieve turgor maintenance and cell expansion in saline conditions. The NHX-type intracellular Na+/H+ exchangers that mediate this process are driven by the differential proton (H+) gradient generated by vacuolar H+-translocating enzymes such as H+-ATPase and H+-PPase. Plant NHX family can be divided into two groups, class I and class II, based on protein sequence and subcellular localization (Rodriguez-Rosales et al. 2009; Pardo et al. 2006). The class I NHX proteins are located on the tonoplast, where they function as (Na+, K+)/H+ antiporters (Venema et al. 2002), while the class II NHX proteins are located in endosomal vesicles of plants (Bassil et al. 2011). These proteins maintain K+ homeostasis and function in aiding normal plant growth and development as well as tolerance to salt stress (Pardo et al. 2006). The AtNHX1 gene, the first plant member of the NHX subfamily of intracellular Na+/H+ antiporters from Arabidopsis thaliana, was identified based on its homology to animal plasma membrane Na+/H+ antiporters of the NHE family and the yeast ScNHX1 gene (Gaxiola et al. 1999). Overexpression of AtNHX1 in other plant systems led to improved salt stress tolerance (Zhang and Blumwald 2001; Zhang et al. 2001). A different model for the role of NHX transporters has been proposed by Jiang et al. (2010), which states that the NHX proteins function mainly to prevent toxic Na+/K+ ratios in the cytosol and for maintaining osmotic balance which is achieved by the vacuolar compartmentalization of Na+ and, in some cases, of other cations as well. A wheat NHX antiporter, TaNHX2, having significant sequence homology to NHX sodium exchangers as reported from Arabidopsis, was found to suppress the salt sensitivity of a yeast mutant strain by improving its K+ content when faced the salt stress (Xu et al. 2013). Here an attempt had been made to compile reported transporters/ion channels/ pumps associated with movement of Na+ and K+ in plants (Table 14.3).
14.4.2 Transcription Factor (TFs) Involved in Salinity Stress Tolerance and Ion Homeostasis
In order to impart enhanced salt tolerance, it is essential to develop a basic understanding of biochemical, physiological and gene regulatory networks of stress response pathways. Transcription factors (TFs) play a critical role in signal transduction network starting from the perception of stress signal to the expression of stress-responsive genes. Unlike the structural genes, TFs tend to control several complex pathways (master regulator) making them one of the ideal candidates for pathway manipulation. Several TFs (OsRAB1, MYC/MYB, OsNAC/SNAC, etc.) have been identified which are differentially expressed during adaptation to salt stress; interestingly, many of these TFs are also differentially expressed during other stresses. In particular, there are many common TFs that control gene expression both during salt and drought stress. A comprehensive database of rice TFs involved in adaptation of salt and drought is available at Rice Stress-Responsive Transcription Factor Database (RiceSRTFDB; http://www.nipgr.res.in/RiceSRTFDB.html) (Priya and Jain 2013). A recent study on transcriptome analysis of common bean (Phaseolus vulgaris L.) under salt stress has reported differential expression of 59 different families of TFs among which 10 TF families, viz. AP2-EREBP, bHLH, PHD, HB, (R1)R2R3_Myb, WRKY_Zn, NAC, bZIP, C3H-TypeI and Myb_related, were most abundant (Hiz et al. 2014). A comparison of transcriptome of salt-sensitive (Hua 30) and salt-tolerant (Hua 11) barley varieties under salt stress has also shown differential expression of several transcription factors belonging to different families, such as TIFY (earlier known as ZIM), WRKY, zinc finger, MYB, bHLH, CBF, NAC, bZIP, AP2, whirly, HD-ZIP, etc. Two interesting observations of this study were that (i) the number of differentially expressed genes was more in shoots as compared to roots and (ii), compared to control, more number of genes were found to be upregulated in response to salt stress than downregulated (Gao et al. 2013). Cloning and validation of such salt stress-responsive TFs is a step in the right direction for improvement of salinity tolerance in crops. For example, stress-specific NAC1 (SNAC1) cloned from rice landrace Pokkali (Hu et al. 2006) was found to confer salt stress tolerance in rice by working downstream to ABA-induced salt and drought tolerance pathway (Khong et al. 2008). Similarly, OsMYB48-1 conferred tolerance to salt stress along with drought in rice (Xiong et al. 2014). On the other hand, cold-induced MYB 1 (CMYB1) which is involved in cold tolerance and circadian rhythm maintenance in rice is negatively correlated with salt stress tolerance. Many of these cloned TFs are trans-acting, i.e. they can impart salt tolerance in different (non-native) backgrounds. For example, OsDREB2A, a transcription factor of AP2/ERF family in rice is capable of imparting salt tolerance in transgenic soybean by accumulation of higher level of osmolytes (Zhang et al. 2013). Some of the other such transacting TFs are OsMYB3R-2 (Dai et al. 2007), HvCBF4 (Oh et al. 2007), DREB1A (Oh et al. 2005), NAC (Tran et al. 2004), etc. If regulated properly in the transgenic background, these trans-acting TFs can work to regulate pathways in any of the desired crops and thus be ideal candidates for engineering salt stress tolerance in crop plants.
14.4.3 Post-transcription Gene Regulation and Adaption to Salt Stress
Apart from the transcription factors, which play a role as master switches to control and coordinate transcription of several genes, a plethora of genes are also controlled post-transcriptionally under salinity stress. This post-transcriptional regulation is achieved through a group of salt-responsive microRNAs (miRNAs), a class of small non-coding RNAs of ~21 nucleotide length, which exerts an additional level of control over plant gene expression under stress. In fact, miRNAs are now considered as one of the major players in gene regulation which downregulate expression of their target genes by mRNA cleavage or translation-arrest mechanisms based on the perfect or near-perfect complementary pairing, respectively (Ambros 2004). A plethora of miRNAs have been demonstrated to play a role in several stress tolerance pathways; some of them are found to be involved in multiple stresses and across species (Dugas and Bartel 2004; Zhang and Wang 2015). Till date, about 40 different families of miRNAs have been shown to play a role in abiotic stress response among which many are associated with salt stress (Covarrubias and Reyes 2010; Sunkar 2010; Wang et al. 2013). Arabidopsis and rice have come up as a model system in recent times to study molecular biology of dicotyledonous and monocotyledonous plant systems, respectively. Numerous studies on these two plants have revealed the importance miRNAs in salt responses. Apart from these, the role of miRNA under salt stress has also been elucidated in several other crop plants like cotton, soybean, Populus, tobacco, Medicago, etc. About 217 miRNAs have been reported till date in different plant species which are involved in salinity stress. A detailed list of the salt-responsive miRNA and their target genes is given in Mittal et al. (2016). The unifying themes which emerge from the studies investigating the role of miRNA in response to salinity are summarized here. Firstly, it has been observed that miRNAs, grossly, target master switches of gene regulation such as transcription factors (e.g. MYB, NAC1, homeodomain-leucine zipper, etc.) or phytohormones (auxin, GA, ethylene and ABA signalling) which in turn regulate expression of several downstream genes in the expression cascade ultimately governing the plant development and physiology (Jones-Rhoades and Bartel 2004). Besides, some of these miRNAs regulate enzymes such NADP-dependent malic enzyme, cytochrome oxidase, laccase, etc. which are of broad spectrum and are involved not only in salt stress but also in several other abiotic and even in biotic stresses (Yan et al. 2005). Hence, most of the miRNAs are not specific to salt but are involved in multiple stresses; especially, several miRNA are commonly regulated in salt and drought stress (Kong et al. 2010; Xie et al. 2014). Secondly, it has been found that the stress-responsive miRNAs are more or less conserved in plant kingdom. For example, miR393 was found to be upregulated in rice, cotton and Arabidopsis under salt stress (Sunkar and Zhu 2004) and subsequently regulates auxin signalling in those plants (Xia et al. 2012). Similarly, salt-induced upregulation of miR156 is observed in seven different species, viz. Arabidopsis thaliana (Liu et al. 2008), Zea mays (Ding et al. 2009), Populus euphratica (Qin et al. 2011), Vigna unguiculata (Paul et al. 2011), Panicum virgatum (Sun et al. 2012), Populus trichocarpa (Li et al. 2013) and Gossypium raimondii (Xie et al. 2014). Several other miRNAs like miR159, miR160, miR162, miR164, miR166, miR167, miR168, miR169, miR395, miR397, etc. are also differentially expressed in multiple species under salt stress (Mittal et al. 2016). Combining these two facts, i.e. the evolutionary conserved nature and the overlapping expression pattern of miRNA in different stresses, it is intimidating to speculate that the miRNA-mediated gene regulation is an ancient phenomenon (relics of RNA world hypothesis?) which has probably originated as early as the plant kingdom made their existence in the world. And this regulation mechanism is not stress specific in most of the cases; rather it has been placed on top of the specific stress regulation mechanisms in order to combine and coordinate plants’ response under multiple stresses.
14.4.4 Important Genes and/or QTLs Associated with Salt Tolerance and Ion Homeostasis
Genetically, salinity tolerance is a complex quantitative trait (Foolad and Jones 1993) which makes it difficult for plant breeders to select for improved genotypes due to low expressivity, heritability and large effects of environment on the trait. Still, genetic resources are vital for any trait, and intraspecific selection has contributed to improved tolerance in rice (Akbar and Yabuno 1977) and barley (Epstein et al. 1980). Over the past decade, research efforts have focused on the mapping and identification of QTLs contributing to salt stress tolerance through marker-assisted selection (Singh et al. 2007; Haq et al. 2010; Table 14.4). For example, the Saltol QTL in rice was identified by employing a RIL population between the tolerant landrace Pokkali and the highly sensitive IR 29 by AFLP genotyping (Gregorio 1997). Further, it was shown that the Saltol QTL contributed to 43% of variation for seedling shoot Na+/K+ ratio (Bonilla et al. 2002). Lin et al. (2004) identified a total of 11 QTLs from an F2 population including major QTLs for shoot K+ concentration on chromosome 1 (qSKC-1) and shoot Na+ concentration on chromosome 7 (qSNC-7) as derived from a cross between tolerant indica rice (Nona Bokra) and sensitive japonica (Koshihikari). These QTLs were found to influence the root and shoot Na+ and K+ accumulation as well as survival under salt stress. Subsequently, a single QTL, qSKC1 or OsHKT1.5 was fine mapped and successfully cloned (Ren et al. 2005). A list of some of the identified QTLs governing salt stress tolerance in different crop species is provided in Table 14.4, and a list of some experimentally validated gene involved in salt-induced response across plant species is given in Table 14.5. In rice alone, about 70 QTLs for salt stress have been mapped (Hu et al. 2012); however, cloning of QTLs is still a rate-limiting step, mainly due to difficulties in fine mapping and defining precise QTL limits. Hence, there is a need to direct research efforts towards identification of genes governing tolerance to salt stress which in turn would aid in development of perfect gene-based markers and pyramiding of multiple QTLs in a single genetic background so as to provide tolerance under diverse stress environments.
14.5 Cellular Defence Network and Plant’s Adaptive Strategy
14.5.1 Role of Reactive Oxygen Species (ROS) in Salinity Tolerance
Salt stress disrupts metabolic coordination between different biochemical pathways, leading to formation of high-energy electrons which, when donated to molecular oxygen, result in the formation of different reactive oxygen species (ROS) such as 1O2, H2O2, O2 •− and HO•. The plant organelles, chloroplast, mitochondria and peroxisomes, are the sites of production of ROS in plants; however, detailed biochemical reactions leading to their production are beyond the scope of this chapter. Both ROS formation and associated injury during salt stress in plants have been previously reported (Gomez et al. 2004; Rubio et al. 2009; Chen et al. 2012). ROS-induced damage in plants depends on the nature and severity of stress, the duration of exposure and even environmental conditions. However, it is well known that while ROS accumulation causes intracellular damage to lipids, proteins and DNA (Bi et al. 2009), it also functions as a signalling molecule in plant-pathogen interaction and abiotic stresses (Mittler et al. 2004; Torres and Dangl 2005). The plasma membrane-located NADP oxidase (NOX) genes or the respiratory burst oxidases (RBOH) which catalyse the synthesis of the superoxide radical are important constituents of ROS-mediated signalling system (Desikan et al. 2001; Mittler et al. 2004; Torres and Dangl 2005). Ma et al. (2012) reported that double mutants atrbohD1/F1 and atrbohD2/F2 of Arabidopsis disrupted Na+/K+ homeostasis therefore showing increased sensitivity to NaCl treatments than wild-type or single null mutants. AtrbohF, apart from increasing ROS levels in response to increased soil salinity, also reduced Na+ concentrations in xylem sap and prevented accumulation of excess Na+ in shoot cells through transpiration (Jiang et al. 2012).
Reactive oxygen species (ROS) scavenging is also extremely important for salt tolerance. The main defence against ROS includes enzymes such as ascorbate peroxidase (APX), superoxide dismutase (SOD), catalase (CAT), glutathione peroxidase (GPX), polyphenol oxidase (PPO) and monodehydroascorbate reductase (MDAR) along with low-molecular weight antioxidants such as ascorbate, glutathione, glycine betaine, trehalose, alpha-tocopherol and proline (Foyer and Noctor 2005; Abogadallah 2010). Overexpression of a rice ascorbate peroxidase gene in alfalfa led to improved salt stress tolerance. In rice transgenic overexpressing pea DEAD-box helicase gene PDH45 which also showed improved salt stress tolerance, it was found that the protein PDH45 physically interacts with Cu/Zn SOD, adenosine-5′-phosphosulphate-kinase, cysteine proteinase and eIF(4G) thus implicating the role of ROS-scavenging machinery in stress tolerance (Gill et al. 2013). Transgenic tobacco plants overexpressing cotton type 3 metallothionein gene GhMT3a also showed increased tolerance against different abiotic stresses including salinity stress (Xue et al. 2009). Improved ROS scavenging may also be attained by manipulation of certain master regulator genes. For instance, Schmidt et al. (2013) identified a rice transcription factor, salt-responsive ERF1 (SERF1), that showed increased expression upon salt and H2O2 treatment. SERF1 showed direct binding to promoters of genes like MAPK kinase kinase 6 (MAP3K6), MAPK5, dehydration-responsive element binding 2A (DREB2A) and zinc finger protein 179 (ZFP179) thus suggesting that it may be the master regulator of ROS-activated MAPK cascade during the initial phase of salt stress making way for downstream gene expression changes resulting in salt stress tolerance.
14.5.2 Osmolytes or Compatible Solute-Mediated Adaptation to Salt Stress
Since osmotic imbalance is one of the most prominent effects of salt stress, adaptation to this stress, to a great extent, depends on the ability to mediate quick osmotic adjustment by accumulation of organic osmolytes like proline, mannitol, fructans, trehalose, glycine betaine, ononitol, etc. In fact, due to the importance of osmotic adjustment in salinity stress adaptation and in many other abiotic stresses as well, it has been regarded as the central dogma of stress physiology (Hare et al. 1998). Not many direct evidences, however, confirm this hypothesis, and most of the evidences are largely correlative. Exposed to salt stress, plants start accumulating organic osmolytes, most of which belong to the class of polyhydroxylic compounds (carbohydrates and sugar alcohols) and zwitterionic alkylamines (amino acids and quaternary amines), as an adaptive response. Unlike ROS, which can be potentially damaging to the cell itself, these organic osmolytes are non-toxic in nature and hence are also termed as ‘compatible solutes’. Cumulatively, these compatible solutes decrease the water potential of cell making them osmotically more competent for water uptake. Several reviews are available which discuss about osmolyte accumulation in plant (Bohnert and Jensen 1996; Serrano et al. 1998; Chakraborty et al. 2012b). Given their immense importance, osmolytes have emerged as tempting candidates to engineer stress resistance in crop plants. Several efforts have been made to develop transgenic plants containing osmolyte-synthesizing genes with an ultimate aim to engineer salt and/or drought stress tolerance. Few of such efforts have been successful (Hayashi et al. 1997), while many of them did not (Smart and Flores 1997). Even where some success has been achieved, the improvement was marginal. The limited success of the osmolyte-overexpressing transgenics is not a reason to dismiss their potential in engineering stress adaptation; rather it indicates the fact that the relative proportion of different osmolytes and the spatiotemporal expression of osmolyte-synthesizing genes are more important as compared to the absolute amount (Hare et al. 1998). Untimely, out of place and/or excessive expression of a particular osmolyte can be associated with yield penalty because of metabolite diversion from primary metabolism (which favours growth and yield) to secondary metabolism (which favours defence). Hence, it is imperative that future research needs to focus more on pathway engineering and devising controlled gene regulation machineries to achieve success in this area.
14.6 Possible Management Options for Alleviation of Salinity Stress
Apart from our traditional effort to breed salt-tolerant crop varieties, sometimes improved crop management practices also play important role in counteracting ill effect of salt stress. Hence, we should consider different external management approaches, viz. maintenance of K+ homeostasis and the use of phytohormones for the growing plants in saline environment.
14.6.1 Exogenous Application of Potassium (K+)
Around the world, researchers have attempted to alleviate the salinity stress applying potassium by and large. However, the mode of application varied over the experiments, either by soil or foliar application, while varying doses of sole potassium or in combination with some soil amendments (like FYM) or with external phytohormones. But some of the results obtained during the course of study are indeed exciting and hence addressing to solve the salt stress in crop plants by suitable crop management.
Salt stress is often noticed by the skewed K+/Na+ ratio in actively growing plant tissues along with stunted growth and metabolic activity of plant tissues (Shabala and Cuin 2008; Degl’Innocenti et al. 2009). Excess build-up of tissue Na+ along with reduced uptake and tissue retention of K+ in plant parts has been conspicuous under saline environment (Munns et al. 2002). Several basic physiological processes in plants, like stomatal closure, destruction of chlorophyll pigment system, etc., have been observed to be hampered under salinity (Gama et al. 2009; Parida et al. 2004). The role of K+ is established in regulation of stomatal movement of plant tissue; thus, better maintenance of water storage and cell turgidity can be assured under osmotic stress (Marschner 2012).
Both soil and foliar application of K+ supplemented the growth, yield and fruit quality of tolerant and sensitive cultivars of tomato grown under salinity. It ensured the role of external K+ application apart from genetic tolerance ability to manage the stress (Amjad et al. 2014). Basal application of potassium improved the overall performance of contrasting peanut varieties at defined salinity levels, while TG 37A, the susceptible one, responded better over GG 2, the tolerant one (Chakraborty et al. 2016c). Similarly, Arshadullah et al. (2014) conducted a hydroponic study on sunflower crop and confirmed that 2% K+ foliar application (as K2SO4 solution) revived the tissue K+ concentration after imposition of salt stress, thus resulting in more biomass production.
In other study, Khan et al. (2016) reported that K+ application along with FYM and other nutrients (N, P, Zn) enhanced the growth, yield and fibre quality of cotton plants by reducing the Na+ uptake and Na+/K+ ratio under the salinity. While in most of the cases potassium sources have been restricted to KCl (muriate of potash) or K2SO4 (potassium sulphate) salts, a recent study tested the K2SO4 nanoparticles on growth and physiological responses of forage crop, alfalfa (Medicago sativa L.), under salt stress, and subsequently better performance of nano-fertilizer was revealed by lower electrolyte leakage, higher proline and relative water content, along with higher antioxidant enzyme activities (superoxide dismutase and catalase), and other growth and yield parameters (El-Sharkawy et al. 2017).
14.6.2 Use of Phytohormones
Phytohormones, synonymously used as plant growth regulators, refer to the compounds originated from plant biosynthetic processes that can act either locally (at the site of their synthesis) or transported to some other sites within the plant in order to promote growth and development responses both under normal and adverse/stressful environment (Peleg and Blumwald 2011). A large array of phytohormones like abscisic acid (ABA), gibberellins (GA), ethylene, auxins (IAA), cytokinin (CKs), and brassinosteroids (BRs), has established their role in abiotic stress management (reviewed in Fahad et al. 2015). Moreover, salinity tolerance mechanism via proline biosynthesis as influenced by regulatory role of phytohormones under salinity stress has also been highlighted (reviewed in Iqbal et al. 2014). Kanmani et al. (2017) conducted a pot experiment with contrasting rice varieties (Pokkali and CO51) treated with foliar application of four different plant growth regulators for mitigating the stress. The response of brassinolide (1.0 ppm) was found interesting for photosynthetic rate and chlorophyll fluorescence, and gibberellic acid (50 ppm) increased the chlorophyll content, while enhanced transpiration rate was observed at kinetin application (20 ppm). In other instances, pretreatment with phytohormones (NAA and BAP) in pineapple (cv. MD Gold) minimized the salt stress effects suffered by the plant by maintaining optimum biomass, increasing tissue K+ concentration, reducing the damage to cell membranes and increasing total soluble sugars (Melo et al. 2017).
14.7 Conclusion and Future Research Strategies
Salinity stress is second most important abiotic stress for cultivated crop plants. Salinity-affected area is gradually increasing under the scenario of global climate change. The importance of properly understanding and combating this threat to world agriculture is, therefore, more apprehended by the researchers. Soil salinity adversely affects plant growth and development accompanied by an increase in uptake of Na+ and Cl− ions and a decrease in uptake of K+, Ca2+ and Mg2+ resulting in ionic imbalance, sodium ion injury and disturbed metabolic processes, changed concentration of biomolecules, photosynthetic activity and poor productivity. Other most detrimental effect faced by the plants is sudden outburst of reactive oxygen species produced due to salinity stress, which disrupts the cellular structure and damages subcellular organelles, leading to cell death. At molecular level, efficient operation of different signal proteins and various symporters and antiporters lying either in the plasma membrane or tonoplast plays important role in salinity tolerance. Activity of different Na+/H+ antiporters, viz. SOS1 and NHX1, depends upon the activation of other signal proteins like SOS2, SOS3 and other calcium-binding proteins.
The genotypes having superior antioxidant defence capacity in terms of either accumulation of antioxidants like ascorbic acid, glutathione, malondialdehyde, etc. or higher activity of the enzymes are more capable of withstanding salinity stress. Salinity stress causes osmotic and oxidative stress; hence, genetic modifications in these areas could yield beneficial result in bringing salinity tolerance in crop plants. Incorporation of genes facilitating biosynthesis of compatible solutes whose accumulation will help in osmotic adjustment in the plant cell and thereby maintaining better water balance inside the plant tissues when facing osmotic pressure from outside. Though there is ample opportunity for research in this area, it needs multidisciplinary approaches to address all the component of the problem of salinity.
Adaptation to salinity stress involves osmotic homeostasis, ionic homeostasis, ROS detoxification as well as tissue adaptation mechanisms. These adaptation strategies are governed by a network of several interacting pathways which are controlled by both genetic and epigenetic regulations. In this context, many cultivated accessions were identified with tolerance to salinity stress. QTLs for salt tolerance have been identified from those cultivated accessions. A few of these have been identified, mapped, cloned and introgressed into elite varieties using molecular breeding approaches. Satisfactory progress in transferring tolerance to high-yielding cultivars for better survivability is made. But yield penalty under salinity stress can’t be reduced significantly. In this context, some of the wild relatives with better tolerance are being utilized in breeding. The major setback faced by the breeders in this approach is that most of wild tolerant genotypes are often cross-incompatible with the cultivated species. There is a need to go for mining of the differentially expressed genes and subsequent transfer of those to cultivable species.
The search for novel salt-tolerant genes or protein is presently extended to some of the halophytic plants such as mangroves which can thrive well under extremely saline environment. The successful transfer of the important genes imparting tolerance to soil salinity from mangrove gene pool to some of important crop plants is being attempted. Besides this, mining of the genes is also possible from a wide range of microbial gene pool as well as from Archaea. Apart from the commonly known pathways that impart tolerance to eukaryotes including higher plants, there may be some other mechanisms operating in these organisms which help them to survive in the extreme environment of sea or saline hot spring. During the past few decades, ‘omics’ approaches have opened possibilities of understanding interaction dynamics between genes, proteins, metabolites and small RNA in salinity stress tolerance both under acute and chronic salinity stress. Recent advent of ‘phenomics’ or large-scale phenotyping is expected to further facilitate efficient identification of promising germplasms for enhancing salt stress tolerance. But the progress made so far is still at the infant stage, and mechanisms of salinity tolerance at the morphological, physiological and molecular level are not very well deciphered in all the crops. It is understood that salinity tolerance like other complex abiotic stress tolerance is controlled by many environmentally responsive genes. Many of them are affected by the occasional post-transcriptional modifications due to extreme climatic fluctuation. Under global climate change, ‘envirotyping’ is emerging as a new concept which will predict multiple genes more precisely along with genotyping and phenotyping and haplotypes interacting with environments across developmental stages. This will help in understanding and genetic manipulation for achieving better salinity tolerance. Besides the potential crosstalk of salinity stress regulatory circuit with other pathways governing the overall physiology of the plants is still mostly under cover. Hence, in-depth studies by means of real-time and cutting-edge technologies at cellular and at the whole-plant level is essentially required in the days to come to have a complete understanding for developing salt-tolerant and environment-resilient varieties in the future.
References
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought-and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868
Abogadallah GM (2010) Insights into the significance of antioxidative defense under salt stress. Plant Signal Behav 5:369–374
Agarwal PK, Agarwal P, Reddy MK, Sopory SK (2006) Role of DREB transcription factors in abiotic and biotic stress tolerance in plants. Plant Cell Rep 25:1263–1274
Akbar M, Yabuno T (1977) Breeding saline-resistant varieties of rice. IV. Inheritance of delayed type panicle sterility induced by salinity. Jpn J Breed 27:237–240
Allen GJ, Wyn-Jones RG, Leigh RA (1995) Sodium transport in plasma membrane vesicles isolated from wheat genotypes with differing K /Na discrimination traits. Plant Cell Environ 18:105–115
Almeida P, Katschnig D, de Boer AH (2013) HKT transporters—state of the art. Int J Mol Sci 14:20359–20385. https://doi.org/10.3390/ijms141020359
Almeida DM, Oliveira MM, Saibo NJ (2017) Regulation of Na+ and K+ homeostasis in plants: towards improved salt stress tolerance in crop plants. Genet Mol Biol. https://doi.org/10.1590/1678-4685-gmb-2016-0106
Amato M, Ladd JN (1994) Application of the ninhydrin reactive N assay for microbial biomass in acid soils. Soil Biol Biochem 26:1109–1115
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355
Amjad M, Akhtar J, Haq MAU, Imran S, Jacobsen SE (2014) Soil and foliar application of potassium enhances fruit yield and quality of tomato under salinity. Turk J Biol 38:208–218
Apse MP, Blumwald E (2007) Na+ transport in plants. FEBS Lett 581:2247–2254
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Arbona V, Flors V, Jacas J, García-Agustín P, Gómez-Cadenas A (2003) Enzymatic and non-enzymatic antioxidant responses of Carrizo citrange, a salt-sensitive citrus rootstock, to different levels of salinity. Plant Cell Physiol 44:388–394
Arshadullah M, Ali A, Hyder SI, Mahmood IA, Zaman BU (2014) Effect of different levels of foliar application of potassium on Hysun-33 and Ausigold-4 sunflower (Helianthus annuus L.) cultivars under salt stress. Pak J Sci Indust Res Series B: Biol Sci 57:1–4
Ashraf M (2004) Some important physiological selection criteria for salt tolerance in plants. Flora 199:361–376
Ashraf M, Foolad M (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216
Bassil E, Tajima H, Liang YC, Ohto M, Ushijima K, Nakano R, Esumi T, Coku A, Belmonte M, Blumwald E (2011) The Arabidopsis Na+/H+ antiporters NHX1 and NHX2 control vacuolar pH and K+ homeostasis to regulate growth, flower development, and reproduction. Plant Cell 23:3482–3497
Bassil E, Coku A, Blumwald E (2012) Cellular ion homeostasis: emerging roles of intracellular NHX Na+/H+antiporters in plant growth and development. J Exp Bot 63:5727–5740
Bhaduri D, Meena HN, Chakraborty K (2016) Variation in phosphorus accumulation in groundnut cultivars as influenced by water salinity. Legum Res 39:215–220
Bi YH, Chen WL, Zhang WN, Zhou Q, Yun LJ, Xing D (2009) Production of reactive oxygen species, impairment of photosynthetic function and dynamic changes in mitochondria are early events in cadmium- induced cell death in Arabidopsis thaliana. Biol Cell 101:629–643
Blumwald E, Aharon GS, Apse MP (2000) Sodium transport in plant cells. Biochim Biophys Acta 1465:140–151
Bohnert HJ, Jensen RG (1996) Strategies for engineering water-stress tolerance in plants. Trends Biotechnol 14:89–97
Bonilla P, Dvorak J, Mackill D, Deal K, Gregorio G (2002) RLFP and SSLP mapping of salinity tolerance genes in chromosome 1 of rice (Oryza sativa L.) using recombinant inbred lines. Philipp Agric Sci 85:68–76
Brandon C, Homman K (1995) The cost of inaction: valuing the economy-wide cost of environmental degradation in India. The World Bank, New Delhi
Brini F, Masmoudi K (2012) Ion transporters and abiotic stress tolerance in plants. ISRN Mol Biol. https://doi.org/10.5402/2012/927436
Britto DT, Kronzucker HJ (2008) Cellular mechanisms of potassium transport in plants. Physiol Plant 133:637–650
Buschmann PH, Vaidyanathan R, Gassmann W, Schroeder JI (2000) Enhancement of Na+ uptake currents, time-dependent inward-rectifying K+ channel currents, and K+ channel transcripts by K+ starvation in wheat root cells. Plant Physiol 122:1387–1398
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, Tester M, Munns R (2007) HKT1;5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1. Plant Physiol 143:1918–1928
Caines AM, Shennan C (1999) Interactive effects of Ca2+ and NaCl salinity on the growth of two tomato genotypes differing in Ca2+ use efficiency. Plant Physiol Biochem 37:569–576
Chakraborty K, Sairam RK, Bhattacharya RC (2012a) Differential expression of salt overly sensitive pathway genes determines salinity stress tolerance in Brassica genotypes. Plant Physiol Biochem 51:90–101
Chakraborty K, Sairam RK, Bhattacharya RC (2012b) Salinity induced expression of pyrrolline-5-carboxylate synthetase determine salinity tolerance in Brassica spp. Acta Physiol Plant 34:1935–1941
Chakraborty K, Singh AL, Bhaduri D, Sairam RK (2013) Mechanism of salinity stress tolerance in crop plants and recent developments. In: Hemantaranjan A (ed) Advances in plant physiology, vol 14. Scientific Publishers, Jodhpur, pp 466–496
Chakraborty K, Bishi SK, Goswami N, Singh AL, Zala PV (2016a) Differential fine-regulation of enzyme driven ROS detoxification network imparts salt tolerance in contrasting peanut genotypes. Environ Exp Bot 128:79–90
Chakraborty K, Sairam RK, Bhaduri D (2016b) Effects of different levels of soil salinity on yield attributes, accumulation of nitrogen, and micronutrients in Brassica spp. J Plant Nutr 39:1026–1037
Chakraborty K, Bhaduri D, Meena HN, Kalariya K (2016c) External potassium (K+) application improves salinity tolerance by promoting Na+-exclusion, K+-accumulation and osmotic adjustment in contrasting peanut cultivars. Plant Physiol Biochem 103:143–153
Chakraborty K, Bose J, Shabala L, Eyles A, Shabala S (2016d) Evaluating relative contribution of osmo- and tissue-tolerance mechanisms towards salinity stress tolerance in three Brassica species. Physiol Plant 158:135–151
Chakraborty K, Bose J, Shabala L, Shabala S (2016e) Difference in root K+ retention ability and reduced sensitivity of K+-permeable channels to reactive oxygen species confer differential salt tolerance in three Brassica species. J Exp Bot 67:4611–4625
Cheeseman JM (2013) The integration of activity in saline environments: problems and perspectives. Funct Plant Biol 40:759–774
Chen L, Song Y, Li S, Zhang L, Zou C, Yu D (2012) The role of WRKY transcription factors in plant abiotic stresses. Biochim Biophys Acta-Gene Regul Mech 1819:120–128
Cheng NH, Pittman JK, Barkla BJ, Shigaki T, Hirschi KD (2003) The Arabidopsis cax1 mutant exhibits impaired ion homeostasis, development, and hormonal responses and reveals interplay among vacuolar transporters. Plant Cell 15:347–364
Chinnusamy V, Jagendorf A, Zhu JK (2005) Understanding and improving salt tolerance in plants. Crop Sci 45:437–448
Coello P, Hey SJ, Halford NG (2010) The sucrose non-fermenting-1-related (SnRK) family of protein kinases: potential for manipulation to improve stress tolerance and increase yield. J Exp Bot 62:883–893
Cotsaftis O, Plett D, Shirley N, Tester M, Hrmova M (2012) A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS One 7:e39865
Covarrubias AA, Reyes JL (2010) Post-transcriptional gene regulation of salinity and drought responses by plant microRNAs. Plant Cell Environ 33:481–489
Cramer GR (2002) Sodium-calcium interactions under salinity stress. In: Salinity: environment-plants-molecules. Springer, Dordrecht, pp 205–227
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143:1739–1751
Degl’Innocenti E, Hafsi C, Guidi L, Navari-Izzo F (2009) The effect of salinity on photosynthetic activity in potassium-deficient barley species. J Plant Physiol 166:1968–1981
Desikan R, Mackerness SAH, Hancock JT, Neill SJ (2001) Regulation of the Arabidopsis transcriptome by oxidative stress. Plant Physiol 127:159–172
Dietrich P, Sanders D, Hedrich R (2001) The role of ion channels in light-dependent stomatal opening. J Exp Bot 52:1959–1967
Ding D, Zhang L, Wang H, Liu Z, Zhang Z, Zheng Y (2009) Differential expression of miRNAs in response to salt stress in maize roots. Ann Bot 103:29–38
Dregne H, Kassas M, Rosanov B (1991) A new assessment of the world status of desertification. Desertification Control Bull 20:6–18
Dubcovsky J, María GS, Epstein E, Luo MC, Dvořák J (1996) Mapping of the K+/Na+ discrimination locus Kna1 in wheat. Theor Appl Genet 92:448–454
Dugas DV, Bartel B (2004) MicroRNA regulation of gene expression in plants. Curr Opin Plant Biol 7:512–520
El-Sharkawy MS, El-Beshsbeshy TR, Mahmoud EK, Abdelkader NI, Al-Shal RM, Missaoui AM (2017) Response of Alfalfa under salt stress to the application of potassium sulfate nanoparticles. Am J Plant Sci 8:1751–1773
Epstein E, Norlyn JD, Rush DW, Kingsbury R, Kelley DB, Wrana AF (1980) Saline culture of crops: a genetic approach. Science 210:399–404
Fahad S, Hussain S, Matloob A, Khan FA, Khaliq A, Saud S, Hassan S, Shan D, Khan F, Ullah N, Faiq M (2015) Phytohormones and plant responses to salinity stress: a review. Plant Growth Regul 75:391–404
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179:945–963
Flowers TJ, Munns R, Colmer TD (2014) Sodium chloride toxicity and the cellular basis of salt tolerance in halophytes. Ann Bot 115:419–431
Foolad MR, Jones RA (1993) Mapping salt-tolerance genes in tomato (Lycopersicon esculentum) using trait-based marker analysis. Theor Appl Genet 87:184–192
Foyer CH, Noctor G (2005) Redox homeostasis and antioxidant signaling: a metabolic interface between stress perception and physiological responses. Plant Cell 17:1866–1875
Fuchs I, Stölzle S, Ivashikina N, Hedrich R (2005) Rice K+ uptake channel OsAKT1 is sensitive to salt stress. Planta 221:212–221
Fukuda A, Nakamura A, Tanaka Y (1999) Molecular cloning and expression of the Na+/H+ exchanger gene in Oryza sativa. Biochim Biophys Acta-Gene Str Expr 1446:149–155
Fukuda A, Nakamura A, Tagiri A, Tanaka H, Miyao A, Hirochika H, Tanaka Y (2004) Function, intracellular localization and the importance in salt tolerance of a vacuolar Na+/H+ antiporter from rice. Plant Cell Physiol 45:146–159
Fukuda A, Nakamura A, Hara N, Toki S, Tanaka Y (2011) Molecular and functional analyses of rice NHX-type Na+/H+ antiporter genes. Planta 233:175–188
Gama PBS, Tanaka K, Eneji A, Eltayeb AE, Elsiddig K (2009) Salt induced stress effects on biomass, photosynthetic rate and reactive oxygen species scavenging enzyme accumulation in common bean. J Plant Nutr 32:837–854
Gao R, Duan K, Guo G, Du Z, Chen Z, Li L, He T, Lu R, Huang J (2013) Comparative transcriptional profiling of two contrasting barley genotypes under salinity stress during the seedling stage. Int J Genomics 2013:1–19. https://doi.org/10.1155/2013/972852
Garciadeblas B, Senn ME, Banuelos MA, Rodriguez-Navarro A (2003) Sodium transport and HKT transporters: the rice model. Plant J 34:788–801
Gaxiola RA, Rao R, Sherman A, Grifasi P, Alpier SL, Fink GR (1999) The Arabidopsis thaliana proton transporters, AtNHX1 and Avp1, can function in cation detoxification in yeast. Proc Natl Acad Sci U S A 96:1480–1485
Ghassemi F, Jakeman AJ, Nix HA (1995) Salinization of land and water resources. Univ. of New South Wales Press, Ltd., Canberra
Gill SS, Tajrishi M, Madan M, Tuteja N (2013) A DESD-box helicase functions in salinity stress tolerance by improving photosynthesis and antioxidant machinery in rice (Oryza sativa L. cv. PB1). Plant Mol Biol 82:1–22
Gomez LD, Noctor G, Knight M, Foyer CH (2004) Regulation of calcium signaling and gene expression by glutathione. J Exp Bot 55:1851–1859
Gong Q, Li P, Ma S, InduRupassara S, Bohnert HJ (2005) Salinity stress adaptation competence in the extremophile Thellungiella halophila in comparison with its relative Arabidopsis thaliana. Plant J 44:826–839
Gorham J (1992) Salt tolerance of plants. Sci Prog (1933-) 76:273–285
Gregorio GB (1997) Tagging salinity tolerance genes in rice using amplified fragment length polymorphism (AFLP). Dissertation, University of the Philippines, Los Baños
Guo Y, Qiu QS, Quintero FJ, Pardo JM, Ohta M, Zhang C, Schumaker KS, Zhu JK (2004) Transgenic evaluation of activated mutant alleles of SOS2 reveals a critical requirement for its kinase activity and C-terminal regulatory domain for salt tolerance in Arabidopsis thaliana. Plant Cell 16:435–449
Gupta NK, Meena SK, Gupta S, Khandelwal SK (2002) Gas exchange, membrane permeability, and ion uptake in two species of Indian jujube differing in salt tolerance. Photo-Dermatology 40:535–539
Hadi MR, Karimi N (2012) The role of calcium in plants’ salt tolerance. J Plant Nutr 35:2037–2054
Hadi MR, Khiyam-Nekoie SM, Khavarinejad R, Khosh Kholgh Sima NA, Yavari P (2008) Accumulation and role of ions (Ca2+, Mg2+, SO4 −2) on salt tolerance in Triticum turgidum L. J Biol Sci 8:143–148
Halfter U, Ishitani M, Zhu JK (2000) The Arabidopsis SOS2 protein kinase physically interacts with and is activated by the calcium-binding protein SOS3. Proc Natl Acad Sci U S A 97:3735–3740
Handbook of Agriculture (2011) Indian Council of Agricultural Research (ICAR), New Delhi. 1617 p. ISBN: 978-8171640966
HanumanthaRao B, Nair RM, Nayyar H (2016) Salinity and high temperature tolerance in mungbean [Vigna radiata (L.) Wilczek] from a physiological perspective. Front Plant Sci 7:957. https://doi.org/10.3389/fpls.2016.00957
Haq TU, Gorham J, Akhtar J, Akhtar N, Steele KA (2010) Dynamic quantitative trait loci for salt stress components on chromosome 1 of rice. Funct Plant Biol 37:634–645
Hare PD, Cress WA, Van Staden J (1998) Dissecting the roles of osmolyte accumulation during stress. Plant Cell Environ 21:535–553
Haro R, Banuelos MA, Senn MAE, Barrero-Gil J, Rodriguez-Navarro A (2005) HKT1 mediates sodium uniport in roots. Pitfalls in the expression of HKT1 in yeast. Plant Physiol 139:1495–1506
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Ann Rev Plant Bio 51:463–499
Hayashi H, Alia Mustardy L, Deshnium P, Ida M, Murata N (1997) Transformation of Arabidopsis thaliana with the codA gene for choline oxidase; accumulation of glycine betaine and enhanced tolerance to salt and cold stress. Plant J 12:133–142
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome analysis of salt tolerant common bean (Phaseolus vulgaris L.) under saline conditions. PLoS ONE 9(3):e92598. https://doi.org/10.1371/journal.pone.0092598
Horie T, Costa A, Kim TH, Han MJ, Horie R, Leung HY, Miyao A, Hirochika H, An G, Schroeder JI (2007) Rice OsHKT2;1 transporter mediates large Na+ influx components into K+-starved roots for growth. EMBO J 26:3003–3014
Horie T, Hauser F, Schroeder JI (2009) HKT transporter-mediated salinity resistance mechanisms in Arabidopsis and monocot crop plants. Trends Plant Sci 14:660–668
Horie T, Brodsky DE, Costa A, Kaneko T, Lo Schiavo F, Katsuhara M, Schroeder JI (2011) K+ transport by the OsHKT2;4 transporter from rice with atypical Na+ transport properties and competition in permeation of K+ over Mg2+ and Ca2+ ions. Plant Physiol 156:1493–1507
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC ) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci U S A 103:12987–12992
Hu S, Tao H, Qian Q, Guo L (2012) Genetics and molecular breeding for salt-tolerance in rice. Rice Genomics Genet 3:38–39
Huang SB, Spielmeyer W, Lagudah ES, James RA, Platten JD, Dennis ES, Munns RA (2006) Sodium transporter (HKT7) is a candidate for Nax1, a gene for salt tolerance in durum wheat. Plant Physiol 142:1718–1727
Iqbal N, Umar S, Khan NA, Khan MIR (2014) A new perspective of phytohormones in salinity tolerance: regulation of proline metabolism. Environ Exp Bot 100:34–42
Ishitani M, Liu J, Halfter U, Kim CS, Shi W, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12:1667–1678
Iterbe-Ormaetxe I, Escuredo PR, Arrese-Igor C, Becana M (1998) Oxidative damage in pea plants exposed to water deficit of paraquat. Plant Physiol 161:173–181
Jabnoune M, Espeout S, Mieulet D, Fizames C, Verdeil JL, Conejero G, Rodriguez-Navarro A, Sentenac H, Guiderdoni E, Abdelly C et al (2009) Diversity in expression patterns and functional properties in the rice HKT transporter family. Plant Physiol 150:1955–1971
Janicka-Russak M, Kłobus G (2007) Modification of plasma membrane and vacuolar H+-ATPases in response to NaCl and ABA. J Plant Physiol 164:295–302
Ji H, Pardo JM, Batelli G, Van Oosten MJ, Bressan RA, Li X (2013) The salt overly sensitive (SOS) pathway: established and emerging roles. Mol Plant 6:275–286
Jiang XY, Leidi EO, Pardo JM (2010) How do vacuolar NHX exchangers function in plant salt tolerance? Plant Signal Behav 5:792–795
Jiang C, Belfield EJ, Mithani A, Visscher A, Ragoussis J, Mott R, Smith JA, Harberd NP (2012) ROS-mediated vascular homeostatic control of root-to-shoot soil Na delivery in Arabidopsis. EMBO J 31:4359–4370
Jin ZM, Wang CH, Liu ZP, Gong WJ (2007) Physiological and ecological characters studies on Aloe vera under soil salinity and seawater irrigation. Process Biochem 42:710–714
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Kamiya T, Akahori T, Ashikari M, Maesshima M (2005) Expression of the vacuolar Ca2+/H+ exchanger, OsCAX1a, in rice: cell and age specificity of expression and enhancement by Ca2+. Plant Cell Physiol 47:96–106. https://doi.org/10.1093/pcp/pci227
Kanmani E, Ravichandran V, Sivakumar R, Senthil A, Surendar KK, Boominathan P (2017) Influence of plant growth regulators on physiological traits under salinity stress in contrasting rice varieties (Oryza sativa L.) Int J Curr Microbiol App Sci 6:1654–1661
Khan HR, Ashraf M, Shahzad SM, Imtiaz M, Aziz A, Piracha MA, Siddiqui AR (2016) Additional application of plant nutrients with farm yard manure for improving the adaptation of cotton crop to salinity stress. J Appl Agric Biotechnol 1:48–57
Khong GN, Richaud F, Coudert Y, Pati PK, Santi C, Périn C, Breitler JC, Meynard D, Vinh DN, Guiderdoni E, Gantet P (2008) Modulating rice stress tolerance by transcription factors. Biotechnol Genet Eng Rev 25:381–404
Kim Y, Arihara J, Nakayama T, Nakayama N, Shimada S, Usui K (2004) Antioxidative responses and their relation to salt tolerance in Echinochloa oryzicola vasing and Steraia viridis (L.) Beauv. Plant Growth Regul 44:87–92
Knight H, Trewavas AJ, Knight MR (1997) Calcium signaling in Arabidopsis thaliana responding to drought and salinity. Plant J 12:1067–1078
Kong Y, Elling AA, Chen B, Deng X (2010) Differential expression of microRNAs in maize inbred and hybrid lines during salt and drought stress. Am J Plant Sci 1:69
Koyama ML, Levesley A, Koebner RM, Flowers TJ, Yeo AR (2001) Quantitative trait loci for component physiological traits determining salt tolerance in rice. Plant Physiol 125:406–422
Kronzucker HJ, Britto DT (2011) Sodium transport in plants: a critical review. New Phytol 189:54–81
Kumar K, Mosa K (2015) Ion transporters: a decisive component of salt stress tolerance in plants. In: Wani SH, Hossain MA (eds) Managing salt tolerance in plants: molecular and genomic perspectives. CRC Press, Boca Raton, pp 373–390
Lan WZ, Wang W, Wang SM, Li LG, Buchanan BB, Lin HX, Gao JP, Luan S (2010) A rice high-affinity potassium transporter (HKT) conceals a calcium-permeable cation channel. Proc Natl Acad Sci U S A 107:7089–7094
Laurie S, Feeney KA, Maathuis FJM, Heard PJ, Brown SJ, Leigh RA (2002) A role for HKT1 in sodium uptake by wheat roots. Plant J 32:139–149
Li B, Duan H, Li J, Deng XW, Yin W, Xia X (2013) Global identification of miRNAs and targets in Populus euphratica under salt stress. Plant Mol Biol 81:525–539
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108:253–260
Lin H, Yang Y, Quan R, Mendoza I, Wu Y, Du W, Zhao S, Schumaker KS, Pardo JM, Guo Y (2009) Phosphorylation of SOS3-LIKE CALCIUM BINDING PROTEIN8 by SOS2 protein kinase stabilizes their protein complex and regulates salt tolerance in Arabidopsis. Plant Cell 21:1607–1619
Liu HH, Tian X, Li YJ, Wu CA, Zheng CC (2008) Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 14:836–843
Ma L, Zhang H, Sun L, Jiao Y, Zhang G, Miao C, Hao F (2012) NADPH oxidase AtrbohD and AtrbohF function in ROS-dependent regulation of Na+/K+ homeostasis in Arabidopsis under salt stress. J Exp Bot 63:305–317
Maathuis F (2006) The role of monovalent cation transporters in plant responses to salinity. J Exp Bot 57:1137–1147
Maathuis FJ, Ahmad I, Patishtan J (2014) Regulation of Na+ fluxes in plants. Front Plant Sci 5:467. https://doi.org/10.3389/fpls.2014.00467
Marin K, Suzuki I, Yamaguchi K, Ribbeck K, Yamamoto H, Kanesaki Y, Hagemann M, Murata N (2003) Identification of histidine kinases that act as sensors in the perception of salt stress in Synechocystis sp., PCC 6803. Proc Natl Acad Sci U S A 100:9061–9066
Marschner H (1986) Mineral nutrition in higher plants. Academic, London, pp 477–542
Marschner P (2012) Marschner’s mineral nutrition of higher plants (3rd). ISBN: 978-0-12-384905-2. Academic, Cambridge, MA
Martinez-Atienza J, Jiang X, Garciablades B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012
Maser P, Eckelman B, Vaidyanathan R, Horie T, Fairbairn DJ, Kubo M, Yamagami M, Yamaguchi K, Nishimura M, Uozumi N, Robertson W, Sussman MR, Schroeder JI (2002) Altered shoot/root Na+ distribution and bifurcating salt sensitivity in Arabidopsis by genetic disruption of the Na+ transporter AtHKTI1. FEBS Lett 531:57–161
Melo YL, Dantas CVS, Lima-Melo Y, Maia JM, Macêdo CECD (2017) Changes in osmotic and ionic indicators in Ananas comosus (L.) cv. MD gold pre-treated with phytohormones and submitted to saline medium. Rev Bras Frutic 39:e-155
Mian A, Oomen RJ, Isayenkow S, Sentenac H, Maathuis FJ, Very AA (2011) Overexpression of a Na+ and K+-permeable HKT transporter in barley improves salt tolerance. Plant J 68:468–479
Mittal D, Sharma N, Sharma V, Sopory SK, Sanan-Mishra N (2016) Role of microRNAs in rice plant under salt stress. Ann Appl Biol 168:2–18
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Møller IS, Gilliham M, Jha D, Mayo GM, Roy SJ, Coates JC, Haseloff J, Tester M (2009) Shoot Na+ exclusion and increased salinity tolerance engineered by cell type–specific alteration of Na+ transport in Arabidopsis. Plant Cell 21:2163–2178
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, Husain S, Rivelli AR, James RA, Condon AT, Lindsay MP, Lagudah ES, Schachtman DP, Hare RA (2002) Avenues for increasing salt tolerance of crops, and the role of physiologically based selection traits. In: Progress in plant nutrition: plenary lectures of the XIV international plant nutrition colloquium. Springer, Dordrecht, pp 93–105
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotechnol 30:360–364
Murillo-Amador B, Jones HG, Kaya C, Aguilar RL, García-Hernández JL, Troyo-Diéguez E, Ávila-Serrano NY, Rueda-Puente E (2006) Effects of foliar application of calcium nitrate on growth and physiological attributes of cowpea (Vigna unguiculata L. Walp.) grown under salt stress. Environ Exp Bot 58:188–196
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta- Gene Regul Mech 1819:97–103
Nedjimi B, Daoud Y (2009) Ameliorative effect of CaCl2 on growth, membrane permeability and nutrient uptake in Atriplex halimus subsp. schweinfurthii grown at high (NaCl) salinity. Desalination 249:163–166
Negrão S, Courtois B, Ahmadi N, Abreu I, Saibo N, Oliveira MM (2011) Recent updates on salinity stress in rice: from physiological to molecular responses. Crit Rev Plant Sci 30:329–377
Nelson M, Maredia M (2001) Environmental impacts of the CGIAR: an assessment
Nouri H, Borujeni SC, Nirola R, Hassanli A, Beecham S, Alaghmand S, Saint C, Mulcahy D (2017) Application of green remediation on soil salinity treatment; a review on halophyte remediation. Process Saf Environ Prot 107:94–107
Obata T, Kitamoto HK, Nakamura A, Fukuda A, Tanaka Y (2007) Rice shaker potassium channel OsKAT1 confers tolerance to salinity stress on yeast and rice cells. Plant Physiol 144:1978–1985
Oh SJ, Song SI, Kim YS, Jang HJ, Kim M, Kim YK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138:341–351
Oh SJ, Kwon CW, Choi DW, Song SIK, Kim JK (2007) Expression of barley HvCBF4 enhances tolerance to abiotic stress in transgenic rice. J Plant Biotechnol 5:646–656
Oldeman LR, Hakkeling TA, Sombroek WG (1991) World map of the status of human induced soil degradation: an explanatory note. International Centre and United Nations Environment Programme, Wageningen
Oomen RJ, Benito B, Sentenac H, Rodríguez-Navarro A, Talón M, Véry AA, Domingo C (2012) HKT2; 2/1, a K -permeable transporter identified in a salt-tolerant rice cultivar through surveys of natural genetic polymorphism. Plant J 71:750–762
Pardo JM, Cubero B, Leidi EO, Quintero FJ (2006) Alcali cation exchangers: roles in cellular homeostasis and stress tolerance. J Exp Bot 57:1181–1199
Parida AK, Das AB, Mohanty P (2004) Defense potentials to NaCl in a mangrove, Bruguiera parviflora: differential changes of isoforms of some antioxidative enzymes. J Plant Physiol 161:531–542
Patel BB, Patel BB, Dave RS (2011) Studies on infiltration of saline–alkali soils of several parts of Mehsana and Patan districts of North Gujarat. J Appl Technol Environ Sanitation 1:87–92
Paul S, Kundu A, Pal A (2011) Identification and validation of conserved microRNAs along with their differential expression in roots of Vigna unguiculata grown under salt stress. Plant Cell Tissue Organ Cult 105:233–242
Peleg Z, Blumwald E (2011) Hormone balance and abiotic stress tolerance in crop plants. Curr Opin Plant Biol 14:290–295
Phuc DT, Minh NV, Yen HH (2016) Assessment of natural variation in OsHKT1;2 gene in rice (Oryza sativa). VNU J Sci Nat Sci Technol 32:189–193
Pilot G, Gaymard F, Mouline K, Chérel I, Sentenac H (2003) Regulated expression of Arabidopsis Shaker K+ channel genes involved in K+ uptake and distribution in the plant. Plant Mol Biol 51:773–787
Priya P, Jain M (2013) RiceSRTFDB: a database of rice transcription factors containing comprehensive expression, cis-regulatory element and mutant information to facilitate gene function analysis. Database 2013:bat027
Qi Z, Spalding EP (2004) Protection of plasma membrane K+ transport by the salt overly sensitive1Na+/H+ antiporter during salinity stress. Plant Physiol 136:2548–2555
Qin Y, Duan Z, Xia X, Yin W (2011) Expression profiles of precursor and mature microRNAs under dehydration and high salinity shock in Populus euphratica. Plant Cell Rep 30:1893
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu JK (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci U S A 99:8436–8441
Quan R, Lin H, Mendoza I, Zhang Y, Cao W, Yang Y, Shang M, Chen S, Pardo JM, Guo Y (2007) SCABP8/CBL10, a putative calcium sensor, interacts with the protein kinase SOS2 to protect Arabidopsis shoots from salt stress. Plant Cell 19:1415–1431
Queensland Government (1995–2017) The state of Queensland, Australia. https://www.qld.gov.au/environment/land/soil/salinity
Quintero FJ, Ohta M, Shi H, Zhu JK, Pardo JM (2002) Reconstitution in yeast of the Arabidopsis SOS signaling pathway for Na+ homeostasis. Proc Natl Acad Sci U S A 99(13):9061–9066
Quintero FJ, Martinez-Atienza J, Villalta I, Jiang X, Kim WY, Ali Z, Fujii H, Mendoza I, Yun DJ, Zhu JK, Pardo JM (2011) Activation of the plasma membrane Na/H antiporter Salt-Overly-Sensitive 1 (SOS1) by phosphorylation of an auto-inhibitory C-terminal domain. Proc Natl Acad Sci U S A 108:2611–2616
Reguera M, Bassil E, Blumwald E (2014) Intracellular NHX-type cation/H+ antiporters in plants. Mol Plant 7:261–263
Ren ZH, Gao JP, Li LG, Cai XL, Wei H, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37:1141–1146
Rengasamy P (2002) Transient salinity and subsoil constraints to dryland farming in Australian sodic soils: an overview. Aust J Exp Agric 42:351–361
Rivandi J, Miyazaki J, Hrmova M, Pallotta M, Tester M et al (2011) A SOS3 homologue maps to HvNax4, a barley locus controlling an environmentally sensitive Na (+) exclusion trait. J Exp Bot 62:1201–1216
Rodriguez-Rosales MP, Galvez FJ, Huertas R, Aranda MN, Baghour M, Cagnac O, Venema K (2009) Plant NHX cation/proton antiporters. Plant Signal Behav 4:265–276
Rubio F, Gassmann W, Schroeder JI (1995) Sodium-driven potassium uptake by the plant potassium transporter HKT1 and mutations conferring salt tolerance. Science 2701:1660–1663
Rubio MC, Bustos-Sammamed P, Clemente MR, Becana M (2009) Effects of salt stress on expression of antioxidant genes and proteins in the model legume Lotus japonicus. New Phytol 181:851–859
Rus AM, Estan MT, Gisbert C, Garcia-Sogo B, Serrano R, Caro M et al (2001) Expressing the yeast HAL1 gene in tomato increases fruit yield and enhances K+/Na+ selectivity under salt stress. Plant Cell Environ 24:875–880
Schachtman D, Liu W (1999) Molecular pieces to the puzzle of the interaction between potassium and sodium uptake in plants. Trends Plant Sci 4:281–287
Schmidt R, Mieulet D, Hubberten HM, Obata T, Hoefgen R, Fernie AR, Fisahn J, San Segundo B, Guiderdoni E, Schippers JH, Mueller-Roeber B (2013) SALT-RESPONSIVE ERF1 regulates reactive oxygen species–dependent signaling during the initial response to salt stress in rice. Plant Cell 25:2115–2131
Schulz P, Herde M, Romeis T (2013) Calcium-dependent protein kinases: hubs in plant stress signaling and development. Plant Physiol 163:523–530
Serrano R, Culiañz-Maciá FA, Moreno V (1998) Genetic engineering of salt and drought tolerance with yeast regulatory genes. Sci Hortic 78:261–269
Shabala S, Cuin TA (2007) Potassium transport and plant salt tolerance. Physiol Plant 133:651–669. https://doi.org/10.1111/j.1399-3054.2007.01008.x
Shabala S, Cuin TA (2008) Potassium transport and plant salt tolerance. Physiol Plant 133:651–669
Shabala S, Pottosin II (2010) Potassium and potassium-permeable channels in plant salt tolerance. In: Demidchik V, Maathuis F (eds) Ion channels and plant stress responses. Springer, Heidelberg, pp 87–110
Shabala S, Demidchik V, Shabala L, Cuin TA, Smith SJ, Miller AJ, Davies JM, Newman IA (2006) Extracellular Ca2+ ameliorates NaCl-induced K+ loss from Arabidopsis root and leaf cells by controlling plasma membrane K+-permeable channels. Plant Physiol 141:1653–1665
Shabala S, Bose J, Fuglsang AT, Pottosin I (2015) On a quest for stress tolerance genes: membrane transporters in sensing and adapting to hostile soils. J Exp Bot 67:1015–1031
Shavrukov Y, Gupta NK, Miyazaki J, Baho MN, Chalmers KJ et al (2010) HvNax3-a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp spontaneum). Funct Integr Genomics 10:277–291
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci U S A 97:6896–6901
Shi H, Xiong L, Stevenson B, Lu T, Zhu JK (2002) The Arabidopsis salt overly sensitive 4 mutants uncover a critical role for vitamin B6 in plant salt tolerance. Plant Cell 14:575–588
Shi H, Kim Y, Guo Y, Stevenson B, Zhu JK (2003) The Arabidopsis SOS5 locus encodes a putative cell surface adhesion protein and is required for normal cell expansion. Plant Cell 15:19–32
Shi Z, Li Y, Wang RC, Makeschine F (2005) Assessment of temporal and spatial variability of soil salinity in a coastal saline field. Environ Geol 48(2):171–178
Singh RK, Gregorio GB, Jain RK (2007) QTL mapping for salinity tolerance in rice. Physiol Mol Biol Plants 13:87–99
Smart CC, Flores S (1997) Overexpression of d-myo-inositol-3-phosphate synthase leads to elevated levels of inositol in Arabidopsis. Plant Mol Biol 33:811–820
Su H, Golldack D, Katsuhara M, Zhao CS, Bohnert HJ (2001) Expression and stress-dependent induction of potassium channel transcripts in the common ice plant. Plant Physiol 125:604–614
Sun G, Stewart CN Jr, Xiao P, Zhang B (2012) MicroRNA expression analysis in the cellulosic biofuel crop switchgrass (Panicum virgatum) under abiotic stress. PLoS One 7:e32017
Sunarpi HT, Motoda J, Kubo M, Yang H, Yoda K, Horie R, Chan WY, Leung HY, Hattori K et al (2005) Enhanced salt tolerance mediated by AtHKT1 transporter-induced Na unloading from xylem vessels to xylem parenchyma cells. Plant J 44:928–938
Sunkar R (2010) MicroRNAs with macro-effects on plant stress responses. Semin Cell Dev Biol 21:805–811
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Sze H, Liang F, Hwang I, Curran AC, Harper JF (2000) Diversity and regulation of plant Ca2+ pumps: insights from expression in yeast. Annu Rev Plant Physiol Plant Mol Biol 51:433–462
Szyroki A, Ivashikina N, Dietrich P, Roelfsema MRG, Ache P, Reintanz B, Deeken R, Godde M, Felle H, Steinmeyer R, Palme K, Hedrich R (2001) KAT1 is not essential for stomatal opening. Proc Natl Acad Sci U S A 98:2917–2921
Teige M, Scheikl E, Eulgem T, Dóczi R, Ichimura K, Shinozaki K et al (2004) The MKK2 pathway mediates cold and salt stress signaling in Arabidopsis. Mol Cell 15:141–152
Tester M, Davenport R (2003) Na+ tolerant and Na+ transport in higher plants. Ann Bot 91:503–527
Thomson MJ, Ocampo DM, Egdane J, Katimbang M, Singh RK, Gregorio G, Ismail M (2007) QTL mapping and marker assisted backcrossing for improved salinity tolerance in rice. In: Plant and animal genomes XV conference, San Diego, CA, pp 13–17
Torres MA, Dangl JL (2005) Functions of the respiratory burst oxidase in biotic interactions, abiotic stress and development. Curr Opin Plant Biol 8:397–403
Tran LSP, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498
Tran LSP, Urao T, Qin F, Maruyama K, Kakimoto T, Shinozaki K, Yamaguchi-Shinozaki K (2007) Functional analysis of AHK1/ATHK1 and cytokinin receptor histidine kinases in response to abscisic acid, drought, and salt stress in Arabidopsis. Proc Natl Acad Sci U S A 104:20623–20628
Tuna AL, Kaya C, Ashraf M, Altunlu H, Yokas I, Yagmur B (2007) The effects of calcium sulphate on growth, membrane stability and nutrient uptake of tomato plants grown under salt stress. Environ Exp Bot 59:173–178
Umali DL (1993) Irrigation-induced salinity: a growing problem for development and the environment, vol 215. World Bank Publications, Washington, DC
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci U S A 97:11632–11637
Urao T, Yakubov B, Satoh R, Yamaguchi-Shinozaki K, Seki M, Hirayama T, Shinozaki K (1999) A transmembrane hybrid-type histidine kinase in Arabidopsis functions as an osmosensor. Plant Cell 11:1743–1754
Venema K, Quintero FJ, Pardo JM, Donaire JP (2002) The Arabidopsis Na+/H+exchanger AtNHX1 catalyzes low affinity Na+ and K+ transport in reconstituted liposomes. J Biol Chem 277:2413–2418
Wang XC, Chang LL, Wang BC, Wang D, Li PH, Wang L, Yi X, Huang Q, Peng M, Guo A (2013) Comparative proteomics of Thellungiella halophila leaves from plants subjected to salinity reveals the importance of chloroplastic starch and soluble sugars in halophyte salt tolerance. Mol Cell Proteomics 12:2174–2195
Wang R, Jing W, Xiao L, Jin Y, Shen L, Zhang W (2015) The Rice high-affinity potassium transporter1;1 is involved in salt tolerance and regulated by an MYB-type transcription factor. Plant Physiol 168:1076–1090. https://doi.org/10.1104/pp.15.00298
Wichern J, Wichern F, Joergensen RG (2006) Impact of salinity on soil microbial communities and the decomposition of maize in acidic soils. Geoderma 137:100–108
Wu YS, Hu YB, Xu GH (2009) Interactive effects of potassium and sodium on root growth and expression of K+/Na+ transporter genes in rice. Plant Growth Regul 57:271–280
Xia K, Wang R, Ou X, Fang Z, Tian C, Duan J et al (2012) OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One 7:e30039
Xie F, Wang Q, Sun R, Zhang B (2014) Deep sequencing reveals important roles of microRNAs in response to drought and salinity stress in cotton. J Exp Bot 66:789–804
Xiong H, Li J, Liu P, Duan J, Zhao Y, Guo X, Li Y, Zhang H, Ali J, Li Z (2014) Overexpression of OsMYB48-1, a novel MYB-related transcription factor, enhances drought and salinity tolerance in rice. PLoS One 9:e92913. https://doi.org/10.1371/journal.pone.0092913
Xu R, Wang J, Li C, Johnson P, Lu C, Zhou M (2012) A single locus is responsible for salinity tolerance in a Chinese landrace barley (Hordeum vulgare L.) PLoS One 7:e43079.59
Xu Y, Zhou Y, Hong S, Xia Z, Cui D, Guo J, Xu H, Jiang X (2013) Functional characterization of a wheat NHX antiporter gene TaNHX2 that encodes a K+/H+ exchanger. PLoS One 8:e78098. https://doi.org/10.1371/journal.pone.0078098
Xue T, Li X, Zhu W, Wu C, Yang G, Zheng C (2009) Cotton metallothionein GhMT3a, a reactive oxygen species scavenger, increased tolerance against abiotic stress in transgenic tobacco and yeast. J Exp Bot 60:339–349
Yan S, Tang Z, Su W, Sun W (2005) Proteomic analysis of salt stress-responsive proteins in rice root. Proteomics 5:235–244
Yan N, Marschner P, Cao W, Zuo C, Qin W (2015) Influence of salinity and water content on soil microorganisms. Int Soil Water Conserv Res 3:316–323
Yang Q, Chen ZZ, Zhou XF, Yin HB, Li X, Xin XF, Hong XH, Zhu JK, Gong ZZ (2009) Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2:22–31
Yao X, Horie T, Xue SW, Leung HY, Katsuhara M, Brodsky DE, Wu Y, Schroeder JI (2010) Differential sodium and potassium transport selectivities of the rice OsHKT2;1 and OsHKT2;2 transporters in plant cells. Plant Physiol 152:341–355
Yokoi S, Quintero FJ, Cubero B, Ruiz MT, Bressan RA, Hasegawa PM, Pardo JM (2002) Differential expression and function of Arabidopsis thaliana NHX Na+/H+ antiporters in the salt stress response. Plant J 30:529–539
Zhang HX, Blumwald E (2001) Transgenic salt-tolerant tomato plants accumulate salt in foliage but not in fruit. Nat Biotechnol 19:765–768
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230:1–15
Zhang HX, Hodson JN, Williams JP, Blumwald E (2001) Engineering salt-tolerant Brassica plants: characterization of yield and seed oil quality in transgenic plants with increased vacuolar sodium accumulation. Proc Natl Acad Sci U S A 98:12832–12836
Zhang XX, Tang YJ, Ma QB, Yang CY, Mu YH, Suo HC, Luo LH, Nian H (2013) OsDREB2A, a rice transcription factor, significantly affects salt tolerance in transgenic soybean. PLoS One 8:e83011. https://doi.org/10.1371/journal.pone.0083011
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Zhu JK (2003) Regulation of ion homeostasis under salt stress. Curr Opin Plant Biol 6:441–445
Zhu JK, Liu J, Xiong L (1998) Genetic analysis of salt tolerance in Arabidopsis: evidence for a critical role of potassium nutrition. Plant Cell 10:1181–1191
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Chakraborty, K. et al. (2018). Ionic Basis of Salt Tolerance in Plants: Nutrient Homeostasis and Oxidative Stress Tolerance. In: Hasanuzzaman, M., Fujita, M., Oku, H., Nahar, K., Hawrylak-Nowak, B. (eds) Plant Nutrients and Abiotic Stress Tolerance. Springer, Singapore. https://doi.org/10.1007/978-981-10-9044-8_14
Download citation
DOI: https://doi.org/10.1007/978-981-10-9044-8_14
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-9043-1
Online ISBN: 978-981-10-9044-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)