Abstract
Reading is a heritable and biologically endowed ability that is enabled by the exaptation of genetic sub-skills that are readily available as language and object recognition abilities. The significant role of genes in biological processes such as the formation and plasticity of the specialized neural networks that accompany reading, the role of epigenetic modification of genes in reading, and the reciprocal effect of emotion and cognition in reading are discussed in this chapter. With due consideration of the genetic make-up of individuals, its effect on formation of the neural circuits corresponding to reading and the gene–environment interaction, the chapter proposes how knowledge of these aspects of a reader is pertinent to providing effective pedagogical solutions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Recent research on biology of reading calls for an understanding of the interplay between genes and neurons that affect language, cognition and emotion. Reading is a cognitive-linguistic process enabled by an interface between the predisposed areas of the human brain responsible for visual, auditory and linguistic capacities. This interface is a result of the ability of the brain to modify neural connections in response to environmental reading input (Pascual-Leone, Amedi, Fregni, & Merabet, 2005). The ability of the brain to form new cortical structures (widely known as neuroplasticity) is found to be regulated by a hereditary mechanism known as epigenetics (Allen, 2008; Borrelli, Nestler, Allis, & Sassone-Corsi, 2008; Felling & Song, 2015; Kennedy et al., 2016). Additionally, language acquisition, cognition and affective factors, which are also remarkably innate, can act both as cause and effect of reading. The above propositions suggest that reading overall is an outcome of simultaneous expressions of multiple genes with multiple functionalities which result in inter-individual differences in reading. This wider understanding of reading is necessary for an all-round development of reading pedagogy, considering learners with and without reading difficulties.
Reading as a Gene-Enabled Ability
Reading must have a high genetic association since it is a union of several sub-skills that are instinctively available to human beings, including in particular orthographic coding, phonological decoding and a set of language sub-skills (Dehaene, 2010). Orthographic coding is an extension of innate object recognition ability that aids letter recognition. Phonological decoding, the ability to decode letters to sounds, is found to be genetically correlated to orthographic coding (Gayán & Olson, 2001). This shows that genes play a major role in both the reading sub-skills. Moreover, other reading sub-skills corresponding to morphological, syntactic and semantic processing are part of language, indicating reading requires language ability. Since language sub-skills are biological and gene-rendered abilities (Berwick & Chomsky, 2016; Brzustowicz, 2014), it can be deduced that reading also makes use of those genetic abilities.
The afore-mentioned reading sub-skills are biologically encoded into dedicated areas of the human brain. Neuroimaging studies prove that these areas are universal among humans (therefore, genetic), with individual languages only effecting local changes in brain activation (Chen, Fu, Iversen, Smith, & Matthews, 2002; Dehaene, 2010; Fu, Chen, Smith, Iversen, & Matthews, 2002). The universal reading areas among humans can perhaps be attributed to the fact that many of these areas share their functions with innate capacities (language, for instance). Some of these universal areas are slightly modified from their genetic function to aid in reading. For example, visual word form area (VWFA) is one such universal area that underwent slight modification from face recognition ability to result in visual word-forming ability, an ability unique to reading (Dehaene & Cohen, 2011). Understandably, a functional MRI study has proved that literacy results in reduced neural activation of VWFA for faces as opposed to increased activation for letters (Dehaene et al., 2010). Another study conducted among subjects across the world found that illiterate individuals were better at face recognition than literate counterparts (Dehaene, Cohen, Morais, & Kolinsky, 2015), indicating that the innate face recognition ability is partly taken over by reading. Therefore, for effective reading ability, genes corresponding to all reading brain areas including VWFA must have normal expressions.
Another important aspect of reading is that it is defined by the biological capabilities of human beings. This means that human beings have developed reading by innovatively making use of available human capabilities. Similarities in letter patterns across languages (Changizi, Zhang, Ye, & Shimojo, 2006) indicate that reading could have evolved within the boundaries of the visual coding abilities of human beings (Dehaene, 2010). A study finds that the neural reorganization of the visual pathway in the brain corresponding to reading acquisition is genetically constrained (Chang et al., 2015). This evidence, along with the fact that object recognition and language abilities are prerequisites for reading, suggest that reading developed within the constraints posed by our genetic capacities. Thus, one would expect normal expression of genes related to vision and language for reading.
Reading Disability and Genes
Reading disability or dyslexia is a complex neurobehavioural disorder mainly affecting reading skill which is known to affect approximately 5–10% of individuals across the globe. Dyslexia provides an indirect gateway to understanding what enables reading in human beings.
Neuroimaging studies have revealed that dyslexia corresponds to reduced activation in the left temporo-parietal cortex (Gabrieli, 2009). Hyperlexia, an ability to read well ahead of age-mates, is characterized by higher activation in the left superior temporal cortex and the same area was characterized by hypoactivation among dyslexic individuals (Turkeltaub et al., 2004), suggesting similarity in the functioning of the reading brain across all individuals and hence a genetic basis for reading. Research has also confirmed that dyslexia is heritable and therefore genetic (Eicher & Gruen, 2013; Fisher & DeFries, 2002; Gialluisi, Newbury, Wilcutt, Consortium, & Luciano, 2014; Paracchini, Diaz, & Stein, 2016).
Genome-wide association studies (GWAS) provide insights into the genes responsible for the disorder. During the previous decade, several candidate genes for dyslexia, namely DYX1C1 (dyslexia susceptibility 1 candidate 1) on 15q21.3 (Taipale et al., 2003), ROBO1 (roundabout Drosophila homolog 1) on 3p12.3 (Hannula-Jouppi et al., 2005; Tran et al., 2014), DCDC2 (double cortin domain-containing protein 2) on 6p22.3 (Meng et al., 2005), KIAA0319 on 6p22.3 (Cope et al., 2005) and MRPL19/C2ORF3 on 2p12 (Scerri et al., 2012). More recently, another set of genes, namely CCDC136 (coiled-coil domain containing 136)/FLNC (filamin C) on 7q32.1 and RBFOX2 (RNA-binding protein, fox-1 homolog 2) were identified as candidate genes associated with both reading and language disabilities (Gialluisi et al., 2014). CMIP, a candidate gene for specific language impairment (SLI) is also found to be associated with reading (Scerri et al., 2011). So far, more than 20 dyslexia candidate genes have been identified by the Human Gene Nomenclature Committee (Raskind, Peter, Richards, Eckert, & Berninger, 2013).
Specific variants of these candidate genes are also identified to have associations with reading sub-skills such as phonological awareness, phonological decoding, orthographic coding and rapid serial naming. Substantially, the presence of genetic variants for dyslexia could indicate that each reading sub-skill involves specific genes. SNPs of KIAA0319, namely rs761100 and rs2038137 are found to be associated with rapid naming and rs6935076, which again is an SNP of KIAA0319, is associated with word reading fluency. Similarly, rs17236239, an SNP of CNTNAP2, is associated with non-word repetition. Population dyslexia studies show that the above abilities differ among individuals, resulting in inter-individual differences in brain circuitry (McGrath et al., 2011; Peterson & Pennington, 2012). These differences are also found to be genetic (Willcutt et al., 2010). Research in this direction could further highlight the genetic associations with the key reading sub-skills.
The candidate genes for reading, apart from DYX1C1, are responsible for brain development as well, implying multiple functions of the candidate genes (McGrath, Smith, & Pennington, 2006). As in the case of general language ability (Centanni, Green, Iuzzini-seigel, Bartlett, & Hogan, 2015), reading ability is manifested through multiple genes, with no specialist gene corresponding to reading disability identified. Also, the above-mentioned candidate genes are not yet known to have been mutated among dyslexic subjects (McGrath et al., 2006). However, involvement of multiple genes and their variants, and the absence of mutation in dyslexia does not undermine the heritability of the disorder, for even a simple genetic trait like eye colour may involve multiple genes and the absence of mutation (White & Rabago-Smith, 2011).
The discussion so far suggests that genes play an important role in making people dyslexic, but the role of genes and environment cannot be quantified unless twin or family studies are employed. Twin studies provide a natural setting for the genetic etiology estimations. Monozygotic (MZ) twins, who share all their genes, and dizygotic (DZ) twins, who share half the genes, are studied to show how a certain trait or disease is genetic or environmental. In such studies, people with a trait or disease (known as probands), their co-twins and co-siblings of the twins are studied for the corresponding variance from expected results which is: shared trait/disease for all MZ twins and shared trait or disease in 50% of the DZ twin population. Then, based upon the comorbidity of a condition among MZ and DZ twins, further analysis is conducted to find the attribution (in percentage) to genetic and environmental causes (Olson, 2006). One such twin study of dyslexic children conducted by Gayán & Olson (2001) with a sample of 215 MZ and 159 DZ twins throws light on three reading sub-skills: word recognition, phonological decoding and orthographic coding. The genetic etiology estimation (with 95% confidence level) for word recognition attributed 54% to genetic influence, 40% to shared environment and 6% to non-shared environment. The percentages for phonological decoding were estimated to be 71, 18 and 11% for genetic influence, shared environment and non-shared environments respectively. For orthographic coding, it was 67% genetic, 17% shared environment effects and 16% non-shared environmental effects (Gayán & Olson, 2001; Olson, 2006). Several other twin and family studies point towards the dominant genetic determination of the disorder along with environmental causes (Bates et al., 2007; Gayán & Olson, 2003; Ho, Wong, Chow, Waye, & Bishop, 2017; Little & Hart, 2016; Olson, 2006; Swagerman et al., 2017).
Having fairly concluded that dyslexia has a major genetic etiology, a question arises as to how the disability is passed on to future generations in the absence of genetic mutations. As it is known that we inherit genetic diseases not only through mutation but also through epigenetic processes such as DNA methylation and histone modification, it could well be the case that genes responsible for reading are impaired through epigenetic processes. The fact that reading and writing could be restored to a limited extent to dyslexic children through interventions points to the possibility of an extremely conducive environment modifying the genetic disability. The role of environmental impact on gene expression in dyslexia could be understood by turning our focus towards the acquisition of reading.
Biology of Reading Acquisition and Fluent Reading
Genetic and neural mechanisms play a causal role in successful reading acquisition. As human biology is an enabler of the ability, it cannot be neglected in seeking an understanding of reading, both in health and disability. The biological process of acquiring reading involves simultaneous expression of genes that enable reading along with support from genes related to cognition and emotion. As these gene expressions trigger the formation of neural networks, reading is materialized in the human brain through the establishment of neuronal connections among various brain areas corresponding to the innate capacities involved in reading. This formation of neural networks of brain areas requires neuroplasticity, the ability to alter the interconnection of neurons located in different areas in response to environmental input. For example, as a child learns to read, letter–sound–meaning correspondences take place through intricate brain structures that connect visual-auditory areas and language areas in the brain (Dehaene, 2010; Norton, Beach, & Gabrieli, 2015; Wandell & Le, 2017).
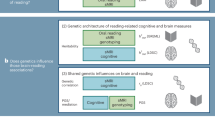
A study suggests that the functional networking of the brain is found to be associated with polymorphisms of a set of genes promoting individual differences in functional connectivity of the brain (Richiardi, Altmann, & Jonas, 2015). Since reading is a result of a complex network of functional areas of the brain, the above finding can be extended to reading ability as well. It can thus be predicted that individual differences in reading ability result from the influence of genetic polymorphisms on functional brain circuits. Similarly, variants of genes also contribute significantly to inter-individual differences in reading achievement among non-dyslexic children. For example, the gene variants, i.e., SNPs, of CCDC136/FLNC and RBFOX2 were found to be associated with reading scores and restructuring of brain tissues (Gialluisi, Guadalupe, Francks, & Fisher, 2017; Gialluisi et al., 2014). Thus, functional networks corresponding to reading have molecular genetic associations and the gene variants correspond to individual differences in reading.
The discussion so far suggests that genetic make-up of individuals influences reading. However, reading can also influence gene–environment interactions, which leads to modification of the genes corresponding to the innate sub-skills of reading by epigenetic mechanisms. Recent breakthrough evidence suggests that an epigenetic process (DNA methylation) in reading-related gene KIAA0319 plays an important role in language lateralization (Schmitz, Kumsta, Moser, Güntürkün, & Ocklenburg, 2018). We can reasonably predict that environmental modification of KIAA0319, which helps language lateralization, comes to the aid of reading acquisition as well. This prediction is in line with a study that finds left hemisphere lateralization of object recognition is observed among children as they acquire reading skills (Caffarra et al., 2017). Additionally, since language sub-skills such as phonological decoding and meaning associations are shared between reading and spoken language (Olson, 2006), one could expect epigenetic effects of language genes passed on to reading skill as well.
Emotion and Cognition in Reading
Affective factors such as pleasure, motivation and self-determination in reading are important factors in reading achievement (McKenna & Kear, 1990; Ölmez, 2015). Self-determined learning is found to be fruitful for struggling learners of reading (Wehmeyer, Shogren, Toste, & Mahal, 2016). Reading for pleasure is linked to a positive attitude towards reading and greater self-confidence (Guthrie & Alvermann, 1999). When reading is pleasurable, it motivates people to read more and it also results in better cognition and language acquisition. The increased cognition, improved language and the pleasure derived drive them to read more, resulting in a cyclic relationship. Positive affective factors have a positive correlation to reading achievement in addition to various cognitive, psychological and physiological benefits. They also have genetic pathways which underlie inter-individual differences. Rs322931, an SNP of LINC01221 on chromosome 1, was found to be associated significantly with positive emotions (Wingo et al., 2017). The study also shows that the minor allele of rs322931 predicts the expression of microRNAs, namely miR-181a and miR-181b and it also postulates that the miRNA is associated with positive emotional stimuli and inhibition of fear. Thus, positive emotions are gene regulated and could positively affect reading acquisition, suggesting the possibility of reading achievement being heritable.
Conversely, if reading is forceful or stressful, it will have a detrimental effect on reading and language acquisition. Studies on reading have confirmed that anxiety is negatively correlated to reading comprehension (Blicher, Feingold, & Shany, 2017; Hewitt & Stephenson, 2012; Knickerbocker, Johnson, & Altarriba, 2015). To illustrate this mechanism in a typical classroom environment, imagine a second-language reading session where a teacher assigns a common text with the intention of enhancing comprehension and acquisition of vocabulary and grammar. The teacher may have chosen the text based on her preference or based on what she perceives as interesting to students. However, in reality, people have a wide variety of preferences such as novels and short stories of different genres, gadget reviews, cooking recipes, newspaper articles, magazines, investment tips, etc. The choice of such texts is dependent on the innate drive to seek knowledge and pleasure through reading. Therefore, when students read a text that is not of their choosing, it may not relate to their innate drive to read and they may not be motivated to continue reading. In effect, they will be demotivated to read. Krashen’s second-language acquisition theories too point out that such anxiety in learning negatively affects language acquisition (Krashen, 1982). This could well be the case in first-language (L1) reading if the L1 reading session is dealt with in a similar manner.
The effects of stress in reading can be explained in terms of biology. As seen earlier, the reading process changes the structure of the brain and new cortical networks are established as a result of its acquisition. Stress acts in a detrimental way in establishing the change in brain connection since it restricts learning and physical growth by inhibiting the required neuronal plasticity (McEwen, Eiland, Hunter, & Miller, 2012). In addition, stress is also epigenetically heritable (McEwen, 2016; Nieto, Patriquin, Nielsen, & Kosten, 2016; Palmisano & Pandey, 2017), raising concerns over stressful methods of language learning.
In addition to the affective factors, human cognition is also influenced by reading. As discussed earlier, reading itself is a complex cognitive process that involves several sub-skills. As we learn to read, these reading sub-skills get better. Apart from this, reading is favourably correlated with several cognitive dimensions. For example, acquisition of grammar (Cox & Guthrie, 2001; Lee, Schallert, & Kim, 2015), and incidental vocabulary enhancement and retention (Ghanbari & Marzban, 2014; Wasik, Hindman, & Snell, 2016) are highly influenced by extensive reading. Moreover, reading enhances social cognitive abilities such as interpersonal sensitivity (Fong, Mullin, & Mar, 2013), and general knowledge and understanding of other cultures (Clark & Rumbold, 2006). Reading is also positively correlated with the theory of mind (Black & Barnes, 2015; Kidd & Castano, 2013), which is the ability to predict or infer our own and other people’s beliefs, desires and intentions (Malle, 2005). Conversely, improvements in several constructs of cognition result in better reading ability (Cox & Guthrie, 2001).
Cognition developed while reading is influenced by human biology. A study suggests that the amygdala, a part of the brain implicated in cognitive mechanisms of learning and memory, releases higher dopamine during reading tasks (Fried et al., 2001). A variety of cognitive processes are linked to genes responsible for dopamine, a neurotransmitter that is a critical modulator of those cognitive processes (Elvevåg & Weinberger, 2009). Another study proposes that cognitive processes such as selective attention, working memory and decision making are genetically associated, leading to individual differences in cognition (Parasuraman, 2009). In addition, human intelligence has been found to have neurobiological and genetic associations (Deary, Penke, & Johnson, 2010). Moreover, cognition is found to be regulated by epigenetic mechanisms (Day & Sweatt, 2011). These findings point towards reading as a complex cognitive process that is genetically enabled and regulated.
Interaction Between Emotion and Cognition in Reading
Emotion and cognition work in tandem in almost all communication situations. For example, if a person wants to express happiness, cognition helps readily by fetching the necessary words to use in the context. Similarly, our emotional experiences help us learn, accentuate thinking processes and solve problems by making decisions (Zambo & Brem, 2004). It is also known that emotion affects cognitive processes such as perception, attention, long-term memory, working memory and decision making; and cognition also affects emotional factors (Dolcos, Iordan, & Dolcos, 2011).
Research in neuroscience reports that the dynamic interaction between cognition and emotion emerges from brain activation (Deary et al., 2010). Cognition- and emotion-based neural circuits are so intertwined that it is difficult to differentiate them (Gray, 1990). It has been argued that there are no different systems in the brain for cognition and emotion because both cognitive and emotional behaviour emerge from a complex, dynamic interaction between brain networks (Pessoa, 2008). In fact, both cognition and emotion are regulated by the amygdala (Dolcos et al., 2011; Fried et al., 2001; Gallagher & Chiba, 1996; Laeger et al., 2012).
Further, research on this topic has pointed to a genetic link in cognition–emotion interactions. It has been found that PCDH17 (Protocadherin 17) and its polymorphisms are associated with cognition, emotion (mood disorders) and the functioning of amygdala (Chang et al., 2017). Genetic association studies point to the fact that interaction between cognition and emotion is highly heritable (ranging from 30–80%) (Scult & Hariri, 2018). A branch named integrative neuroscience calls for the neuroscience of cognition and emotion, and their brain and genetic correlates, to be brought together (Williams, Tsang, Clarke, & Kohn, 2010).
Despite overwhelming evidence of the reciprocal relationship between cognition and emotion, most reading sessions in academic contexts discard the importance of emotion. In a typical reading-based classroom setting, learners experience a diverse set of emotions from the same reading passage. Some may be anxious because they cannot comprehend the text, some may find it uninteresting, some may find it informative, and so on. However, most learning environments tend to ignore emotion and focus on cognition. Emotion is more basic than cognition and can be expected to have a greater impact on learner outcomes.
The Need for Personalized Learning
Classroom reading/writing is one of the important ways people gain knowledge, enhance first-language skills and acquire a second language. However, formal classrooms that still follow one-size-fits-all policy of education are nightmares for many language learners (Hashemi, 2011). This policy goes against the genetic and epigenetic aspects of reading which perpetrate diversity among learners. A large-sample study postulates that 40% of variance in educational attainment is due to differences in the genetic make-up of individuals (Rietveld et al., 2013), suggesting reading-based training modules should be in tune with genetic variations in learning among individuals. Most importantly, people with learning difficulties such as reading disability and specific language impairment must have access to specialized teaching aids and personnel. Healthy learners also require personalized training since they differ greatly owing to inter-individual differences in learning methods and capacity. Moreover, the biological view of reading has shown the adverse effects of stress and positive effects of self-motivation in reading. This calls for a revisit of the formal classroom environment to make classrooms inclusive, with provision for stress-free, personalized and self-motivated learning.
A basic knowledge of neurological functions and epigenetics leading to inter-individual learning differences is as important to teachers as medical examination of a patient is important to doctors. However, the formal language teaching environment does not take into account the variety in brain functions that are due to genetic and epigenetic factors. An analogy from the field of medicine can provide a parallel insight into how personalized teaching makes the environment conducive to natural reading/language acquisition. The emerging field of pharmacogenomics studies the role of genes in inter-individual differences in drug response (Weinshilboum & Wang, 2006). Patients are tested for their neural, genetic and epigenetic markers and the physiological effect is predicted in advance. A suitable drug is then prescribed as a response to the genetic make-up of each patient. Similarly, in the pursuit of personalizing reading/language acquisition, a personalized teaching strategy that takes into account genetic make-up and an individual’s response to learning methods, might be proposed.
A review of the topic (Wong, Vuong, & Liu, 2017) has suggested that the emerging personalized medicine can be extended to personalized learning as well. Personalized learning would aim to use genetic, neuronal and behavioural predictors to identify learning ability and needs of individuals, thereby arriving at an optimal teaching-learning solution. Neuroimaging studies (Bach, Richardson, Brandeis, Martin, & Brem, 2013; Hoeft et al., 2007, 2011; Molfese, 2000) have shown that neuromarkers (initial brain measures such as cortical volume and thickness) of reading have successfully predicted future reading gains in dyslexic patients. Since epigenetic mechanisms influence neuromarkers, their role in neurodevelopment of reading can also be understood. A study confirms this view and suggests that genetic variants can be linked to neuroimaging of the brain which in turn is connected to reading impairments (Eicher & Gruen, 2013). This could be the future of personalized learning in reading classrooms.
Conclusion
This chapter has demonstrated that several dimensions of reading such as reading sub-skills, reading brain areas, functional neuroplasticity (of reading), the dyslexic brain, positive affect and cognition are effected in the brain as neurobiological dispensations caused by genetic underpinnings of individuals. Therefore, reading must be viewed in its totality as a genetic ability that causes inter-individual difference in learning ability. Such a view is a prerequisite to understanding differing reading environment needs for dyslexics and healthy learners with differences in gene expression. A great deal of evidence pointing to neurobiology and genetics of reading ability suggests the direction of personalized learning that reading classrooms must take. Only a holistic approach to reading and switching to personalized learning methods will help us effectively tackle the inherent learning gaps between individuals.
References
Allen, N. D. (2008). Temporal and epigenetic regulation of neurodevelopmental plasticity. Philosophical Transactions: Biological Sciences, 363(1489), 23–38. https://doi.org/10.1098/rstb.2006.2010.
Bach, S., Richardson, U., Brandeis, D., Martin, E., & Brem, S. (2013). Print-specific multimodal brain activation in kindergarten improves prediction of reading skills in second grade. Neuroimage, 82, 605–615. https://doi.org/10.1016/j.neuroimage.2013.05.062.
Bates, T. C., Castles, A., Luciano, M., Wright, M. J., Coltheart, M., & Martin, N. G. (2007). Genetic and environmental bases of reading and spelling: A unified genetic dual route model. Reading and Writing, 20(1–2), 147–171. https://doi.org/10.1007/s11145-006-9022-1.
Berwick, R. C., & Chomsky, N. (2016). Why only us: Language and evolution. Cambridge, MA: MIT Press.
Black, J. E., & Barnes, J. L. (2015). The effects of reading material on social and non-social cognition. Poetics, 52, 32–43. https://doi.org/10.1016/j.poetic.2015.07.001.
Blicher, S., Feingold, L., & Shany, M. (2017). The role of trait anxiety and preoccupation with reading disabilities of children and their mothers in predicting children’s reading comprehension. Journal of Learning Disabilities, 50(3), 309–321. https://doi.org/10.1177/0022219415624101.
Borrelli, E., Nestler, E. J., Allis, C. D., & Sassone-Corsi, P. (2008). Decoding the epigenetic language of neuronal plasticity. Neuron, 60(6), 961–974. https://doi.org/10.1016/j.neuron.2008.10.012.
Brzustowicz, L. M. (2014). Molecular genetic approaches to the study of language. Human Biology, 70(2): 199–213. Retrieved from http://www.jstor.org/stable/41465641.
Caffarra, S., Martin, C. D., Lizarazu, M., Lallier, M., Zarraga, A., Molinaro, N., et al. (2017). Word and object recognition during reading acquisition: MEG evidence. Developmental Cognitive Neuroscience, 24(16), 21–32. https://doi.org/10.1016/j.dcn.2017.01.002.
Centanni, T. M., Green, J. R., Iuzzini-seigel, J., Bartlett, C. W., & Hogan, T. P. (2015). Evidence for the multiple hits genetic theory for inherited language impairment: A case study. Frontiers in Genetics, 6(August), 6–11. https://doi.org/10.3389/fgene.2015.00272.
Chang, C. H. C., Pallier, C., Wu, D. H., Nakamura, K., Jobert, A., Kuo, W. J., et al. (2015). Adaptation of the human visual system to the statistics of letters and line configurations. Neuroimage, 120, 428–440. https://doi.org/10.1016/j.neuroimage.2015.07.028.
Chang, H., Hoshina, N., Zhang, C., Ma, Y., Cao, H., Wang, Y., et al. (2017). The protocadherin 17 gene affects cognition, personality, amygdala structure and function, synapse development and risk of major mood disorders. Molecular Psychiatry, 23, 1–13. https://doi.org/10.1038/mp.2016.231.
Changizi, M., Zhang, Q., Ye, H., & Shimojo, S. (2006). The structures of letters and symbols throughout human history are selected to match those found in objects in natural scenes. The American Naturalist, 167(5), E117–E139. https://doi.org/10.1086/502806.
Chen, Y., Fu, S., Iversen, S. D., Smith, S. M., & Matthews, P. M. (2002). Testing for dual brain processing routes in reading: A direct contrast of Chinese character and pinyin reading using fMRI. Journal of Cognitive Neuroscience, 14(7), 1088–1098. https://doi.org/10.1162/089892902320474535.
Clark, C., and Rumbold, K. (2006). Reading for pleasure: A research overview. National Literacy Trust. London. Retrieved from http://www.scholastic.com/teachers/article/collateral_resources/pdf/i/Reading_for_pleasure.pdf.
Cope, N., Harold, D., Hill, G., Moskvina, V., Stevenson, J., Holmans, P., et al. (2005). Strong evidence that KIAA0319 on chromosome 6p is a susceptibility gene for developmental dyslexia. American Journal of Human Genetics, 76(4), 581–591. https://doi.org/10.1086/429131.
Cox, K. E., & Guthrie, J. T. (2001). Motivational and cognitive contributions to students’ amount of reading. Contemporary Educational Psychology, 26, 116–131. https://doi.org/10.1006/ceps.1999.1044.
Day, J. J., & Sweatt, J. D. (2011). Epigenetic mechanisms in cognition. Neuron, 70(5), 813–829. https://doi.org/10.1016/j.neuron.2011.05.019.
Deary, I. J., Penke, L., & Johnson, W. (2010). The neuroscience of human intelligence differences. Nature Reviews Neuroscience, 11(3), 201. https://doi.org/10.1038/nrn2793.
Dehaene, S. (2010). Reading in the brain: The science and evolution of a human invention. New York, NY: Viking.
Dehaene, S., & Cohen, L. (2011). The unique role of the visual word form area in reading. Trends in Cognitive Sciences, 15(6), 254–262. https://doi.org/10.1016/j.tics.2011.04.003.
Dehaene, S., Cohen, L., Morais, J., & Kolinsky, R. (2015). Illiterate to literate: Behavioural and cerebral changes induced by reading acquisition. Nature Reviews Neuroscience, 16(4), 234–244. https://doi.org/10.1038/nrn3924.
Dehaene, S., Pegado, F., Braga, L. W., Ventura, P., Filho, G. N., Jobert, A., et al. (2010). How learning to read changes the cortical networks for vision and language. Science, 330(6009), 1359–1364. https://doi.org/10.1126/science.1194140.
Dolcos, F., Iordan, A. D., & Dolcos, S. (2011). Neural correlates of emotion—cognition interactions: A review of evidence from brain imaging investigations. Journal of Cognitive Psychology, 23(6), 669–694. https://doi.org/10.1080/20445911.2011.594433.
Eicher, J. D., & Gruen, J. R. (2013). Imaging-genetics in dyslexia: Connecting risk genetic variants to brain neuroimaging and ultimately to reading impairments. Molecular Genetics and Metabolism, 110(3), 201–212. https://doi.org/10.1016/j.ymgme.2013.07.001.
Elvevåg, B., & Weinberger, D. R. (2009). Introduction: Genes, cognition and neuropsychiatry. Cognitive Neuropsychiatry, 14(4–5), 261–276. https://doi.org/10.1080/13546800903126016.
Felling, R. J., & Song, H. (2015). Epigenetic mechanisms of neuroplasticity and the implications for stroke recovery. Experimental Neurology, 268, 37–45. https://doi.org/10.1016/j.expneurol.2014.09.017.
Fisher, S. E., & DeFries, J. C. (2002). Developmental dyslexia: Genetic dissection of a complex cognitive trait. Nature Reviews Neuroscience, 3(10), 767–780. https://doi.org/10.1038/nrn936.
Fong, K., Mullin, J. B., & Mar, R. A. (2013). What you read matters: The role of fiction genre in predicting interpersonal sensitivity. Psychology of Aesthetics, Creativity, and the Arts, 7(4), 370–376. https://doi.org/10.1037/a0034084.
Fried, I., Wilson, C. L., Morrow, J. W., Cameron, K. A., Behnke, E. D., Ackerson, L. C., et al. (2001). Increased dopamine release in the human amygdala during performance of cognitive tasks. Nature Neuroscience, 4(2), 201–206. https://doi.org/10.1038/84041.
Fu, S., Chen, Y., Smith, S., Iversen, S., & Matthews, P. M. (2002). Effects of word form on brain processing of written Chinese. Neuroimage, 17(3), 1538–1548. https://doi.org/10.1006/nimg.2002.1155.
Gabrieli, J. D. E. (2009). Dyslexia: A new synergy between education and cognitive neuroscience. Science, 325(5938), 280–283. https://doi.org/10.1126/science.1171999.
Gallagher, M., & Chiba, A. A. (1996). The amygdala and emotion. Current Opinion in Neurobiology, 6(2), 221–227. https://doi.org/10.1016/S0959-4388(96)80076-6.
Gayán, J., & Olson, R. K. (2001). Genetic and environmental influences on orthographic and phonological skills in children with reading disabilities. Developmental Neuropsychology. Developmental Neuropsychology, 20(2), 483–507. https://doi.org/10.1207/S15326942DN2002_3.
Gayán, J., & Olson, R. K. (2003). Genetic and environmental influences on individual differences in printed word recognition. Journal of Experimental Child Psychology, 84(2), 97–123. https://doi.org/10.1016/S0022-0965(02)00181-9.
Ghanbari, M., & Marzban, A. (2014). Effect of extensive reading on incidental vocabulary retention. Procedia—Social and Behavioral Sciences, 116, 3854–3858. https://doi.org/10.1016/j.sbspro.2014.01.854.
Gialluisi, A., Guadalupe, T., Francks, C., & Fisher, S. E. (2017). Neuroimaging genetic analyses of novel candidate genes associated with reading and language. Brain and Language, 172, 9–15. https://doi.org/10.1016/j.bandl.2016.07.002.
Gialluisi, A., Newbury, D. F., Wilcutt, E. G., Consortium, T. S. L. I., & Luciano, M. (2014). Genome-wide screening for DNA variants associated with reading and language traits. Genes, Brain and Behavior, 13(7), 686–701. https://doi.org/10.1111/gbb.12158.
Gray, J. A. (1990). Brain systems that mediate both emotion and cognition. Cognition and Emotion, 4(3), 269–288. https://doi.org/10.1080/02699939008410799.
Guthrie, J. T., & Alvermann, D. E. (1999). Engaged reading: Processes, practices and policy implications. New York: Teachers College Press.
Hannula-Jouppi, K., Kaminen-Ahola, N., Taipale, M., Eklund, R., Nopola-Hemmi, J., Kääriäinen, H., et al. (2005). The axon guidance receptor gene ROBO1 is a candidate gene for developmental dyslexia. PLoS Genetics, 1(4), 0467–0474. https://doi.org/10.1371/journal.pgen.0010050.
Hashemi, M. (2011). Language stress and anxiety among the English language learners. Procedia—Social and Behavioral Sciences, 30, 1811–1816. https://doi.org/10.1016/j.sbspro.2011.10.349.
Hewitt, E., & Stephenson, J. (2012). Foreign language anxiety and oral exam performance: A replication of Phillips’s MLJ study. Modern Language Journal, 96(2), 170–189. https://doi.org/10.1111/j.1540-4781.2011.01174.x.
Ho, C. S. H., Wong, S. W. L., Chow, B. W. Y., Waye, M. M. Y., & Bishop, D. V. M. (2017). Genetic and environmental etiology of speech and word reading in Chinese. Learning and Individual Differences, 56, 49–58. https://doi.org/10.1016/j.lindif.2017.04.001.
Hoeft, F., McCandliss, B. D., Black, J. M., Gantman, A., Zakerani, N., Hulme, C., et al. (2011). Neural systems predicting long-term outcome in dyslexia. Proceedings of the National Academy of Sciences, 108(1), 361–366. https://doi.org/10.1073/pnas.1008950108.
Hoeft, F., Meyler, A., Hernandez, A., Juel, C., Taylor-Hill, H., Martindale, J. L., et al. (2007). Functional and morphometric brain dissociation between dyslexia and reading ability. Proceedings of the National Academy of Sciences of the United States of America, 104(10), 4234–4239. https://doi.org/10.1073/pnas.0609399104.
Kennedy, A. J., Rahn, E. J., Paulukaitis, B. S., Michael, T. P., Day, J. J., David, J., et al. (2016). Tcf4 regulates synaptic plasticity, DNA methylation, and memory function. Cell Reports, 16, 2666–2685. https://doi.org/10.1016/j.celrep.2016.08.004.
Kidd, D. C., & Castano, E. (2013). Reading literary fiction improves theory of mind. Science, 342(6156), 377–380. https://doi.org/10.1126/science.1239918.
Knickerbocker, H., Johnson, R. L., & Altarriba, J. (2015). Emotion effects during reading: Influence of an emotion target word on eye movements and processing. Cognition and Emotion, 29(5), 784–806. https://doi.org/10.1080/02699931.2014.938023.
Krashen, S. D. (1982). Priniciples and practice in second language acquisition (1st ed.). London: Penguin Press Inc.
Laeger, I., Dobel, C., Dannlowski, U., Kugel, H., Grotegerd, D., Kissler, J., et al. (2012). Amygdala responsiveness to emotional words is modulated by subclinical anxiety and depression. Behavioural Brain Research, 233(2), 508–516. https://doi.org/10.1016/j.bbr.2012.05.036.
Lee, J., Schallert, D. L., & Kim, E. (2015). Effects of extensive reading and translation activities on grammar knowledge and attitudes for EFL adolescents. System, 52, 38–50. https://doi.org/10.1016/j.system.2015.04.016.
Little, C. W., & Hart, S. A. (2016). Examining the genetic and environmental associations among spelling, reading fluency, reading comprehension and a high stakes reading test in a combined sample of third and fourth grade students. Learning and Individual Differences, 45, 25–32. https://doi.org/10.1016/j.lindif.2015.11.008.
Malle, B. F. (2005). Folk theory of mind: Conceptual foundations of human social cognition. In R. Hassin, S. J. Uleman, & J. A. Bargh (Eds.), The new unconscious (pp. 225–255). New York: Oxford University Press. https://doi.org/10.1093/acprof:oso/9780195307696.003.0010
McEwen, B. S. (2016). In pursuit of resilience: Stress, epigenetics, and brain plasticity. Annals of the New York Academy of Sciences, 1373(1), 56–64. https://doi.org/10.1111/nyas.13020.
McEwen, B. S., Eiland, L., Hunter, R. G., & Miller, M. M. (2012). Stress and anxiety: Structural plasticity and epigenetic regulation as a consequence of stress. Neuropharmacology, 62(1), 3–12. https://doi.org/10.1016/j.neuropharm.2011.07.014.
McGrath, L. M., Pennington, B. F., Shanahan, M. A., Santerre-Lemmon, L. E., Barnard, H. D., Willcutt, E. G., et al. (2011). A multiple deficit model of reading disability and attention-deficit/ hyperactivity disorder: Searching for shared cognitive deficits. Journal of Child Psychology and Psychiatry and Allied Disciplines, 52(5), 547–557. https://doi.org/10.1111/j.1469-7610.2010.02346.x.
McGrath, L. M., Smith, S. D., & Pennington, B. F. (2006). Breakthroughs in the search for dyslexia candidate genes. Trends in Molecular Medicine, 12(7), 333–341. https://doi.org/10.1016/j.molmed.2006.05.007.
McKenna, M. C., & Kear, D. J. (1990). A new tool for teachers. The Reading Teacher, 43(8), 626–639. https://doi.org/10.1598/RT.43.8.3.
Meng, H., Smith, S. D., Hager, K., Held, M., Liu, J., Olson, R. K., et al. (2005). DCDC2 is associated with reading disability and modulates neuronal development in the brain. Proceedings of the National Academy of Sciences, 102(47), 17053–17058. https://doi.org/10.1073/pnas.0508591102.
Molfese, D. L. (2000). Predicting dyslexia at 8 years of age using neonatal brain responses. Brain and Language, 72(3), 238–245. https://doi.org/10.1006/brln.2000.2287.
Nieto, S. J., Patriquin, M. A., Nielsen, D. A., & Kosten, T. A. (2016). Don’t worry: Be informed about the epigenetics of anxiety. Pharmacology, Biochemistry and Behavior, 146–147, 60–72. https://doi.org/10.1016/j.pbb.2016.05.006.
Norton, E. S., Beach, S. D., & Gabrieli, J. D. E. (2015). Neurobiology of dyslexia. Current Opinion in Neurobiology, 30, 73–78. https://doi.org/10.1016/j.conb.2014.09.007.
Ölmez, F. (2015). An investigation into the relationship between L2 reading motivation and reading achievement. Procedia—Social and Behavioral Sciences, 199, 597–603. https://doi.org/10.1016/j.sbspro.2015.07.561.
Olson, R. K. (2006). Genes, environment, and dyslexia the 2005 Norman Geschwind Memorial Lecture. Annals of Dyslexia, 56(2), 205–238. https://doi.org/10.1007/s11881-006-0010-6.
Palmisano, M., & Pandey, S. C. (2017). Epigenetic mechanisms of alcoholism and stress-related disorders. Alcohol, 60, 46. https://doi.org/10.1016/j.alcohol.2017.01.001.
Paracchini, S., Diaz, R., and Stein, J. (2016). Advances in dyslexia genetics—new insights into the role of brain asymmetries. In T. Friedmann, J. Dunlap, & G. S. F. (Eds.), Advances in genetics (1st ed., Vol. 96, pp. 53–97). Cambridge, MA: Elsevier Inc. https://doi.org/10.1016/bs.adgen.2016.08.003.
Parasuraman, R. (2009). Assaying individual differences in cognition with molecular genetics: Theory and application. Theoretical Issues in Ergonomics Science, 10(5), 399–416. https://doi.org/10.1080/14639220903106403.
Pascual-Leone, A., Amedi, A., Fregni, F., & Merabet, L. B. (2005). The plastic human brain cortex. Annual Review of Neuroscience, 28(1), 377–401. https://doi.org/10.1146/annurev.neuro.27.070203.144216.
Pessoa, L. (2008). On the relationship between emotion and cognition. Nature Reviews Neuroscience, 9(2), 148–158. https://doi.org/10.1038/nrn2317.
Peterson, R. L., & Pennington, B. F. (2012). Developmental dyslexia. The Lancet, 379(9830), 1997–2007. https://doi.org/10.1016/S0140-6736(12)60198-6.
Raskind, W. H., Peter, B., Richards, T., Eckert, M. M., & Berninger, V. W. (2013). The genetics of reading disabilities: From phenotypes to candidate genes. Frontiers in Psychology, 3(1), 1–20. https://doi.org/10.3389/fpsyg.2012.00601.
Richiardi, J., Altmann, A., & Jonas, R. (2015). Correlated gene expression supports synchronous activity in brain networks. Science, 348(6240), 11–14. https://doi.org/10.1126/science.1255905.
Rietveld, C. A., Medland, S. E., Derringer, J., Yang, J., Esko, T., Martin, N. W., … Koellinger, P. D. (2013). GWAS of 126,559 Individuals identifies genetic variants associated with educational attainment. Science, 340(6139): 1467–1471. https://doi.org/10.1126/science.1235488.
Scerri, T. S., Darki, F., Newbury, D. F., Whitehouse, A. J. O., Peyrard-Janvid, M., Matsson, H., et al. (2012). The dyslexia candidate locus on 2p12 is associated with general cognitive ability and white matter structure. PLoS ONE, 7(11), 50321. https://doi.org/10.1371/journal.pone.0050321.
Scerri, T. S., Morris, A. P., Buckingham, L., Newbury, D. F., Miller, L. L., Bishop, D. V. M., et al. (2011). DCDC2, KIAA0319 and CMIP are associated with reading-related traits. Biological Psychiatry, 70(3), 237–245. https://doi.org/10.1016/j.biopsych.2011.02.005.
Schmitz, J., Kumsta, R., Moser, D., Güntürkün, O., & Ocklenburg, S. (2018). KIAA0319 promoter DNA methylation predicts dichotic listening performance in forced-attention conditions. Behavioural Brain Research, in press.. https://doi.org/10.1016/J.BBR.2017.09.035.
Scult, M. A., & Hariri, A. R. (2018). A brief introduction to the neurogenetics of cognition-emotion interactions. Current Opinion in Behavioral Sciences, 19, 50–54. https://doi.org/10.1016/j.cobeha.2017.09.014.
Swagerman, S. C., van Bergen, E., Dolan, C., de Geus, E. J. C. C., Koenis, M. M. G. G., Hulshoff Pol, H. E., et al. (2017). Genetic transmission of reading ability. Brain and Language, 172, 3–8. https://doi.org/10.1016/j.bandl.2015.07.008.
Taipale, M., Kaminen, N., Nopola-Hemmi, J., Haltia, T., Myllyluoma, B., Lyytinen, H., et al. (2003). A candidate gene for developmental dyslexia encodes a nuclear tetratricopeptide repeat domain protein dynamically regulated in brain. Proceedings of the National Academy of Sciences of the United States of America, 100(20), 11553–11558. https://doi.org/10.1073/pnas.1833911100.
Tran, C., Wigg, K. G., Zhang, K., Cate-Carter, T. D., Kerr, E., Field, L. L., et al. (2014). Association of the ROBO1 gene with reading disabilities in a family-based analysis. Genes, Brain and Behavior, 13(4), 430–438. https://doi.org/10.1111/gbb.12126.
Turkeltaub, P. E., Flowers, D. L., Verbalis, A., Miranda, M., Gareau, L., & Eden, G. F. (2004). The neural basis of hyperlexic reading: An fMRI case study. Neuron, 41(1), 11–25. https://doi.org/10.1016/S0896-6273(03)00803-1.
Wandell, B. A., & Le, R. K. (2017). Diagnosing the neural circuitry of reading. Neuron, 96(2), 298–311. https://doi.org/10.1016/j.neuron.2017.08.007.
Wasik, B. A., Hindman, A. H., & Snell, E. K. (2016). Book reading and vocabulary development: A systematic review. Early Childhood Research Quarterly, 37, 39–57. https://doi.org/10.1016/j.ecresq.2016.04.003.
Wehmeyer, M. L., Shogren, K. A., Toste, J., & Mahal, S. (2016). Self-determined learning to motivate struggling learners in reading and writing. Intervention in School and Clinic. https://doi.org/10.1177/1053451216676800.
Weinshilboum, R. M., & Wang, L. (2006). Pharmacogenetics and pharmacogenomics: Development, science, and translation. Annual Review of Genomics and Human Genetics, 7, 223–245. https://doi.org/10.1146/annurev.genom.6.080604.162315.
White, D., & Rabago-Smith, M. (2011). Genotype–phenotype associations and human eye color. Journal of Human Genetics, 56(1), 5–7. https://doi.org/10.1038/jhg.2010.126.
Willcutt, E. G., Betjemann, R. S., McGrath, L. M., Chhabildas, N. A., Olson, R. K., DeFries, J. C., et al. (2010). Etiology and neuropsychology of comorbidity between RD and ADHD: The case for multiple-deficit models. Cortex, 46(10), 1345–1361. https://doi.org/10.1016/j.cortex.2010.06.009.
Williams, L. M., Tsang, T. W., Clarke, S., & Kohn, M. (2010). An “integrative neuroscience” perspective on ADHD: Linking cognition, emotion, brain and genetic measures with implications for clinical support. Expert Review of Neurotherapeutics, 10(10), 1607–1621. https://doi.org/10.1586/ern.10.140.
Wingo, A. P., Almli, L. M., Stevens, J. S., Jovanovic, T., Wingo, T. S., Tharp, G., et al. (2017). Genome-wide association study of positive emotion identifies a genetic variant and a role for microRNAs. Molecular Psychiatry, 22(5), 774–783. https://doi.org/10.1038/mp.2016.143.
Wong, P. C. M., Vuong, L. C., & Liu, K. (2017). Personalized learning: From neurogenetics of behaviors to designing optimal language training. Neuropsychologia, 98, 192–200. https://doi.org/10.1016/j.neuropsychologia.2016.10.002.
Zambo, D., & Brem, S. K. (2004). Emotion and cognition in students who struggle to read: New insights and ideas. Reading Psychology, 25(3), 189–204. https://doi.org/10.1080/02702710490489881.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Sriganesh, R., Rahul, D.R., Ponniah, R.J. (2018). Genetics of Reading Ability and Its Role in Solving Reading Difficulties. In: Ponniah, R., Venkatesan, S. (eds) The Idea and Practice of Reading. Springer, Singapore. https://doi.org/10.1007/978-981-10-8572-7_8
Download citation
DOI: https://doi.org/10.1007/978-981-10-8572-7_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8571-0
Online ISBN: 978-981-10-8572-7
eBook Packages: Social SciencesSocial Sciences (R0)