Abstract
Our understanding of the viral world changed just after the first structures of icosahedral viral particles were unveiled. The structural similarities between capsid proteins of distant viral groups were not anticipated, and the findings suggested the existence of common ancestors for viruses with different host range, genomic structure and multiplication strategies. This way, diverse viruses with icosahedral particles can now be grouped based on the structural homology between their capsid proteins. In the last years, the presence of conserved folds between viral proteins in non-icosahedral viruses has also emerged. Viral particles with radically different morphologies, ranging from naked and filamentous to enveloped and pleomorphic, have shown structural homology between the nucleoproteins that bind directly to their genomes. This chapter overviews recent findings regarding the similar structure found between nucleoproteins of eukaryotic ssRNA viruses. The structural homology includes the coat proteins from all known families of flexible filamentous plant viruses, a group with monopartite (+)ssRNA genomes. Their coat proteins share a core domain with nucleoproteins of previously unrelated families of enveloped viruses that have segmented (−)ssRNA genomes. This last group consists of mostly animals viruses, including influenza virus.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- ssRNA viruses
- Nucleoproteins
- Icosahedral particles
- Non-icosahedral particles
- Capsid proteins
- Structural homology
Introduction
Almost 40 years ago, pioneering crystallographic studies on two icosahedral viruses (Abad-Zapatero et al. 1980; Harrison et al. 1978) revealed a clear structural similarity between their capsid proteins. The resemblance was not expected since the proteins that build the capsids of the two viruses have no sequence homology between them; however they display the same so-called jelly roll fold. Additional early works on viral structures supported the notion of structural homology between capsid proteins and an evolutionary divergence of the viruses from common ancestors (Rossmann et al. 1983). Currently, the classical taxonomy of viruses by their genomic features (Baltimore 1971) is challenged by a structure-based classification (Abrescia et al. 2012). In the latter, the viral universe is segregated into four major lineages (PRD1/Adenovirus-like, Picornavirus-like, HK97-like, and BTV-like), and new groups have been recently proposed (Nasir and Caetano-Anolles 2017). A drawback of these new structure-based classifications is that only icosahedral viruses are clearly grouped, and helical and non-icosahedral enveloped viruses lie outside the described lineages. The strong spatial restrictions for capsid proteins in icosahedral arrangement seem to limit their structural variation; thus, the similarities between them are kept at recognition levels. Nucleoproteins from viruses that do not construct well-ordered icosahedral particles exhibit larger structural variability, and their relationships are harder to reveal. Nevertheless, the latest structural studies on nucleoproteins and virions from non-icosahedral viruses substantiate new homologies between viral groups with different morphologies. This chapter is focused on a recently described structural homology between the nucleoproteins from several families of ssRNA viruses that infect eukaryotes.
Eukaryotic ssRNA Viruses
Among viruses that infect eukaryotic organisms, RNA viruses are the most abundant and diverse, especially the ones with (+)ssRNA genomes. It seems that the compartments in the cytoplasm provide a rich niche where RNA replication complexes are constructed via interactions with proteins and membranes from the hosts (Nagy and Pogany 2011). In the last ICTV release of virus classification (Adams et al. 2017), (+)ssRNA viruses are distributed in 3 orders and 22 unassigned families (Fig. 6.1, that includes only viral families relevant for this chapter), while the less populated group of (−)ssRNA viruses contains 1 order and 4 unassigned families (in this last ICTV release, the previously unassigned family Bunyaviridae is redistributed in several families within the new order Bunyavirales, but this chapter keeps the name of this family to easily refer to previous works). A tentative phylogeny of eukaryotic (+)ssRNA viruses has been proposed based on the sequence homology between their RNA-dependent RNA polymerases (RdRp), the only common gene to all the families, and the structure of their viral genomes (Koonin 1991). This phylogeny distinguishes three superfamilies: alphavirus-like, picornavirus-like, and flavivirus-like. On the other hand, (−)ssRNA viruses, whose RdRp differ significantly from the ones of the (+)ssRNA groups, are segregated in the order Mononegavirales (which includes eight families with monopartite genomes) and in several unassigned families with segmented genomes (Fig. 6.1) (Koonin et al. 2015).
Groups of ssRNA eukaryotic viruses. Some of the orders and families of eukaryotic ssRNA viruses are shown grouped accordingly to the polarity of their ssRNA. Only the viral families relevant for this chapter are included, together with cartoons that represent the architecture of their virions or infective particles. The names of the different families are seen in green (plant infecting viruses), red (animal infecting viruses), or orange (family with plant and animal viruses). (*) Bunyaviridae family is currently reassigned in the order Bunyavirales (see main text)
Flexible Filamentous Plant Viruses
Flexible filamentous plant viruses are plant pathogens that contain a monopartite (+)ssRNA genome protected by hundreds of copies of their coat protein (CP) arranged in helical mode (Kendall et al. 2008). Their infective particles are long (several hundreds of nm) and thin (10–15 nm diameter) flexible filaments. They are transmitted by mechanical contact or by arthropod vectors and cause severe economic impact in agriculture. Currently there are more than 380 species (Adams et al. 2017) grouped in four families: Alphaflexiviridae (50 species, where genus Potexvirus has 35 representatives), Betaflexiviridae (89 species, genus Carlavirus includes 47 different viruses), Closteroviridae (49 species), and Potyviridae (195 species, and genus Potyvirus includes 160). All those viruses display a very similar architecture for their non-enveloped virions, although some genus within family Closteroviridae have segmented genomes.
Most of the flexuous filamentous plant virus groups belong to the alphavirus-like superfamily (Fig. 6.1). They share a closely related RdRp, a capping enzyme, and the superfamily 1 helicase gene (Koonin and Dolja 1993). However, family Potyviridae fits in the picornavirus-like superfamily following the RdRp-based phylogeny, the expression and processing of a polyprotein, and the presence of a genome-linked VPg protein. Potyviruses are clear outsiders within the picornavirus-like superfamily where the icosahedral capsid made of proteins with the jelly roll fold is abundant; however potyviruses display helical and filamentous virions. It is thought that a common CP gene for flexible filamentous viruses has been transferred and finally shared by all the families (Koonin et al. 2015).
Enveloped and Segmented (−)ssRNA Viruses
Families Orthomyxoviridae (e.g., influenza virus), Bunyaviridae (e.g., Rift Valley fever virus or RVFV), and Arenaviridae (e.g., Lassa fever virus) have been sometimes grouped within the order Multinegavirales, i.e., enveloped viruses with segmented (−)ssRNA genomes or sNSV (segmented negative-strand viruses). They present genomes divided into two (Arenaviridae), three (Bunyaviridae), and six to eight (Orthomyxoviridae) fragments. These subgenomic segments have complementary ends and form circular nucleocapsids (Raju and Kolakofsky 1989; Hsu et al. 1987) together with nucleoproteins and the viral polymerase. The ribonucleoprotein complexes of arenaviruses and bunyaviruses are rather flexible and unstructured, but in influenza they construct double-helical nucleoproteins (Arranz et al. 2012). For all the representatives of this tentative order, the genomic material is protected inside an envelope coming from the membrane of the infected cell. Most of the viruses within these three families infect animals, but the genus Tospovirus (e.g., tomato spotted wilt virus or TSWV, family Bunyaviridae) is a plant-infecting group that can multiplicate within the arthropod vector (usually thrips) leading to persistent vector transmission (Kormelink et al. 2011).
Structure of Flexible Filamentous Plant Viruses
Initial structural studies of flexuous filamentous plant viruses revealed their common overall architecture (Kendall et al. 2008). Apart from possible differences at their ends (for instance, the presence of VPg linked to the 5′ genomic end in potyviruses), low-resolution X-ray fiber diffraction data and cryoEM 3D maps showed filaments of 120–130 Å diameter constructed by CPs arranged in helical mode, with about nine subunits per turn. The studies were carried out with soyben mosaic virus (SMV), a potyvirus, and three potexviruses (family Alphaflexiviridae), potato virus X (PVX), papaya mosaic virus (PapMV), and narcissus mosaic virus (NMV) (Kendall et al. 2013; Yang et al. 2012; Kendall et al. 2008). The flexible nature of the virions precluded atomic resolved data, and the virions were depicted following a right-handed helical arrangement, as observed for rod-shaped rigid viruses such as tobacco mosaic virus or TMV (Namba and Stubbs 1986). In recent years, by using single-particle based helical reconstruction of cryoEM data, several virions have been characterized at higher structural detail: bamboo mosaic virus (BaMV), a potexvirus resolved at 5.6 Å resolution (DiMaio et al. 2015); pepino mosaic virus (PepMV), another potexvirus solved at 3.9 Å resolution (Agirrezabala et al. 2015); and watermelon mosaic virus (WMV), a potyvirus solved at 4.0 Å resolution (Zamora et al. 2017). The availability of structures for flexible filamentous plant viruses from different families (Alphaflexiviridae and Potyviridae) allows for direct comparison (Fig. 6.2).
Structure of two flexible filamentous plant viruses belonging to different families. (a) CryoEM micrograph field of a WMV (family Potyviridae) sample, together with the rendering of the cryoEM map (EMD-3785) for the virion depicted blue (Zamora et al. 2017). (b) An electron micrograph field for a sample with PepMV (family Alphaflexiviridae) virions is shown, and the corresponding cryoEM map (EMD-3236) rendered in red (Agirrezabala et al. 2015). (c) Representation of the atomic models (pdb code 5ODV) of several CPs from WMV as seen in the virion. The atomic coordinates are seen in ribbons with different blue colors for each subunit. One of the CP monomers is depicted as a solid surface. (d) Similar depiction for the atomic models of CPs subunits of PepMV (pdb code 5FN1) shown in red. (e) Two views of the atomic model of the CP from WMV including a fragment of ssRNA. (f) The atomic coordinates for the CP from PepMV are depicted in similar orientations as in (e)
The three described virions (BaMV, PepMV, and WMV) display almost identical helical arrangement, with about 34.5–35 Å of helical pitch and 8.8 subunits per turn in left-handed helices (Fig. 6.2a–b). The CPs show a core domain rich in alpha helices and two long arms at both ends of the protein. The assembly of the CPs is mostly mediated by flexible N- and C-terminal arms, in a way that slight relative movements between CP subunits are allowed (Fig. 6.2c–d), and this is the structural basis for the flexible nature of the virions (Zamora et al. 2017; DiMaio et al. 2015; Agirrezabala et al. 2015). Essentially, the C-terminal arm contributes to the oligomerization between CP subunits at different turns of the helix, i.e., the axial or longitudinal assembly. In all the cases, a final segment of the N-terminal end of the CPs is missing in the atomic models due to its high flexibility. A significant difference is seen in the role of this N-terminal arm. While in potexviruses the N-terminal of one CP interacts with the next subunit in side-by-side contact (Fig. 6.2d) (DiMaio et al. 2015; Agirrezabala et al. 2015), in the potyvirus, a longer N-terminal segment bridges the next subunit in the helix, and by a sharp turn, also interacts with another CP copy at adjacent turn (Fig. 6.2c) (Zamora et al. 2017), displaying a dual role supporting side-by-side and axial polymerization.
Structural Homology Between CPs from Flexible Filamentous Plant Viruses
Remarkably, despite the low sequence identity between CPs from two different families (Potyviridae and Alphaflexiviridae), their 3D fold is almost identical (Fig. 6.2e–f) with rmsd values at the core of the protein (excluding flexible N- and C-terminal arms) bellow 3 Å (Zamora et al. 2017), and all the essential alpha-helical elements of their structure superimpose (Fig. 6.3). Thus, at least for these two families, their CPs are clear structural homologues, which suggest that a gene transfer occurred at some time between families that are distant with regard to other genetic elements and characteristics.
Comparison of the CP structure from WMV and PepMV. The ribbon representation for WMV CP (pdb code 5ODV (Zamora et al. 2017)) is depicted in rainbow colors. The atomic structure for PepMV CP (pdb code 5FN1, (Agirrezabala et al. 2015)) is seen in gray ribbons. The 3D alignment between both structures was performed in Matras (Kawabata 2003). The numbers indicate the residue number at the N- and C-terminal ends of both atomic coordinates
Conserved RNA-Binding Site
In both, potexviruses and potyviruses, the CP in the virion binds to five nucleotides of the ssRNA in a very similar mode (Zamora et al. 2017; DiMaio et al. 2015; Agirrezabala et al. 2015). Although the density for the ssRNA in those cryoEM maps of virions is an average of RNA segments with different compositions, the signal attributed to the nucleic acid is alike in the three available density maps. The higher-resolution studies (Zamora et al. 2017; Agirrezabala et al. 2015) showed that one out of the five nucleotides bound by each CP fits in a binding pocket (nucleotide labeled as U4 in Fig. 6.4). Essentially, several residues from the CP interact with consecutive phosphates backbone groups, and the nucleoside in between goes deep into the binding pocket (Fig. 6.4b).
Conserved ssRNA-binding pocket. (a) Semi transparent depiction of one CP subunit from WMV (gray) segmented from the cryoEM map of the virion (EMD-3785 (Zamora et al. 2017)), together with the density attributed to the path of the ssRNA (in red) and the derived atomic model (pdb code 5ODV). (b) Close-up view of the ssRNA-binding pocket in the CP of WMV with some of the amino acids highlighted. (c), (d), and (e) show the regions that participate in the RNA binding pockets of the CP from WMV (c), PepMV (d), and a comparison between them (e). Three key and conserved amino acids are seen
Direct comparison of the atomic models for WMV (Fig. 6.4c) and PepMV (Fig. 6.4d) reveals that three amino acids that participate in the ssRNA-binding pocket are at the same position in the CP of both viruses (Zamora et al. 2017) (Fig. 6.4e). Furthermore, these serine (S), arginine (R), and aspartic (D) acid residues are universally conserved along the four families of flexible filamentous plant viruses (Fig. 6.5) (Zamora et al. 2017; Dolja et al. 1991), with the exception of two potexviruses (bamboo mosaic virus and foxtail mosaic virus) where the conserved arginine is substituted by histidine. Despite the lack of structures for CPs from other families, the high conservation of invariant amino acids suggests that the CPs from flexuous filamentous plant viruses display the same fold and contain a highly conserved RNA-binding site.
Conservation of amino acids in the RNA-binding pocket along the families of flexible filamentous plant viruses. Consensus sequence logos (Crooks et al. 2004) for CPs from different families of flexuous filamentous plant viruses. The conserved invariant amino acids (Ser or S, Arg or R, and Asp or D) are highlighted in red boxes
Architecture of Enveloped Viruses with Segmented (−)ssRNA Genomes
The members of sNSV have a common overall design that includes the presence of an envelope that protects a variable number of ribonucleoproteins (RNPs) inside (Fig. 6.6). The envelope is taken from the host cell membrane by budding (Lyles 2013) and contains viral glycoproteins that have different roles during the viral cycle. The shape of the virions ranges from pleomorphic (Arenaviridae) to spherical and/or elongated (Orthomyxoviridae and Bunyaviridae). Most of the representatives infect animals, with the exception of plant-infecting tospoviruses (Bunyaviridae). Several bunyaviruses and arenaviruses are present in rodents and arthropods and occasionally infect humans in outbreaks of hemorrhagic fever and encephalitis-related diseases. Family Orthomyxoviridae includes well-known influenza viruses that have a large impact in human health. Influenza representatives infect birds and mammals and are transmitted by aerosols between humans. The nucleoproteins of sNSV are associated to the genomic segments and the RdRp in nucleocapsids of dissimilar morphologies (Ruigrok et al. 2011). These nucleoproteins are mainly helical globular with a positively charged groove for RNA binding (Reguera et al. 2014), but no structural homology has been described between nucleoproteins of different families.
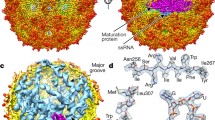
Morphology and organization of segmented (−)ssRNA viruses. (a) Rendering of a segmented cryoelectron tomogram for influenza A virus (image courtesy of J. Martín-Benito) (Arranz et al. 2012) and a cartoon that summarizes the general features of the virion. (b) Semi transparent view of the cryoEM map for native influenza RNPs (EMD2205) with fitted atomic coordinates for its nucleoproteins (pdb code 4BBL) (Arranz et al. 2012). (c) Display that includes the representation of the cryoEM map for RVFV (EMD-5124 (Sherman et al. 2009)) and a schematic cartoon of the viral architecture. (d) Crystallographic structure of the hexameric form of the nucleoprotein from RVFV assembled with ssRNA (pdb code 4H5O (Raymond et al. 2012))
Large part of the structural information in Orthomyxoviridae family has been obtained for influenza A virus. Nucleoproteins of influenza virus polymerize through the insertion of a loop into the neighboring subunit (Ye et al. 2006). In the constructed RNPs, the ssRNA is in closed conformation, and the viral RdRp binds both RNA ends. CryoET analysis of influenza virions showed the presence of helical RNPs inside the virus (Fig. 6.6a), and cryoEM of isolated RNPs revealed a double-helical architecture with two antiparallel strands of nucleoproteins (Fig. 6.6b) (Arranz et al. 2012). For bunyaviruses (RVFV is used as a representative), loose and flexible RNPs (Raymond et al. 2012) are seen protected inside a spherical shell of glycoproteins (Fig. 6.6c) inserted in the enveloping membrane (Huiskonen et al. 2009; Freiberg et al. 2008). Crystallographic structures of nucleoproteins from bunyaviruses with and without RNA have shown several oligomeric states, from tetramers to hexamers (Fig. 6.6d), where side-by-side interaction between subunits is mediated by N- and/or C-terminal arms (Zhou et al. 2013). The number of ssRNA nucleotides bound by each nucleoprotein subunit can vary from 7 as seen for RVFV (Raymond et al. 2012) up to 11 for orthobunyaviruses (Reguera et al. 2013; Niu et al. 2013; Dong et al. 2013; Ariza et al. 2013), one of the genus in the family Bunyaviridae. The structure of their native RNPs is not well known, but it seems to be rather flexible, ranging from loose and unstructured as in RVFV (Raymond et al. 2012) to different levels of helical arrangement as in La Crosse orthobunyavirus (Reguera et al. 2013). The members of the family Arenaviridae present unique nucleoproteins. This way, the nucleoprotein of Lassa virus (Arenaviridae) displays an additional C-terminal domain with exonuclease activity that seems to be involved in immune suppression (Hastie et al. 2011; Qi et al. 2010).
Structural Homology between Nucleoproteins of Eukaryotic ssRNA Viruses
It is clear that flexible filamentous plant viruses (at least two of the families) display high structural homology between their CPs, which are also nucleoproteins (Zamora et al. 2017). Despite the abundant structural information about nucleoproteins of sNSV, no structural homology was detected by direct comparison between atomic coordinates of nucleoproteins from different families (Ruigrok et al. 2011). However, using the atomic models of CPs from flexuous filamentous plant viruses as structural targets, structural similarities with several nucleoproteins of sNSV emerged (Zamora et al. 2017; Agirrezabala et al. 2015). The core region of CPs shows structural homology with nucleoproteins of representatives from families Bunyaviridae (Zamora et al. 2017; Agirrezabala et al. 2015) and Orthomyxoviridae (Zamora et al. 2017) (Fig. 6.7). The structures share similar topology where alpha-helical secondary structure elements are easily aligned. Both N- and C-terminal ends are in similar positions, and the grooves for ssRNA binding are at the same location within the nucleoproteins. For the nucleoprotein of influenza virus, there is not any available atomic structure in complex with ssRNA, but the proposed binding site (Ye et al. 2006) aligns well with that of the other nucleoproteins (indicated by an arrow in Fig. 6.7c). The estimated probability that all these nucleoproteins (the ones displayed in Fig. 6.7 and related) belong to the same fold is above 90% (Zamora et al. 2017; Kawabata 2003). In essence, there is a clear fragment of about 150 residues that shares the same fold in nucleoproteins of different ssRNA viruses, and each of these proteins has additional regions of variable length at N- and C-terminal ends. There is no significant structural homology with nucleoproteins from family Arenaviridae, although as mentioned earlier, their nucleoproteins have adopted an additional domain and might have diverged from a similar fold.
Structural homology between nucleoproteins of eukaryotic ssRNA viruses. The panels show ribbon representations for the core regions of nucleoproteins from different viruses in rainbow color mode (left side). At the right side, the core of the nucleoprotein subunit is seen green, and the N- and C-terminal extensions are red and yellow, respectively. Other subunits that interact with the colored ones are depicted gray to illustrate their oligomerization. The atomic coordinates are (a) WMV CP and ssRNA (pdb code 5ODV (Zamora et al. 2017)); (b) PepMV CP and ssRNA, (pdb code 5FN1 (Agirrezabala et al. 2015)); (c) in the left side, a single influenza virus A nucleoprotein subunit, (pdb code 3ZDP (Chenavas et al. 2013)), and at the right side, two interacting nucleoproteins (pdb code 2IQH (Ye et al. 2006)); (d) RVFV nucleoprotein in complex with ssRNA, (pdb code 4H5O (Raymond et al. 2012)); (e) La Crosse virus nucleoprotein and ssRNA, (pdb code 4BHH(Reguera et al. 2013); (f) and TSWV nucleoprotein in complex with ssRNA, (pdb code 5IP2 (Komoda et al. n.d.). The numbers indicate the residue number at the N- and C-terminal ends of the atomic coordinates
The oligomerization of these nucleoproteins takes place through interactions via N- and C-terminal arms (right panels in Fig. 6.7), although the final multimeric RNPs have different arrangements (from loose to full helical). Remarkably, the N-terminal arm oligomerization modes between nucleoproteins in potexviruses (family Alphaflexiviridae) and phlebovirues (family Bunyaviridae) are comparable and use the same groove in the neighboring subunit to receive the N-terminal arm (Agirrezabala et al. 2015). Nucleoproteins of influenza viruses are a clear exception, since a folded and large C-terminal region (depicted yellow in the right panel of Fig. 6.7c) contributes to oligomerization by the insertion of an internal loop in the adjacent subunit (Arranz et al. 2012; Ye et al. 2006).
Evolutionary Implications
Structural homology between proteins is usually understood as an indication of common evolutionary origin (homology) rather than the product of convergent evolution (analogy). This is based on the observation that the structure of proteins is more conserved than their sequence of amino acids (Illergard et al. 2009) and that convergent evolution of protein domains is a rare event (Gough 2005). In the current matter, the structural similarity between nucleoproteins is further supported by their role as viral proteins that bind and protect ssRNA genomes. It can be presumed that the genes of nucleoproteins from flexible filamentous plant viruses and from at least two families of sNSV share a common ancestor (Fig. 6.8). The homology between these two sets of viruses was not anticipated until atomic structures for the first group (the flexuous plant viruses) were available (Zamora et al. 2017; Agirrezabala et al. 2015). This suggests that the structure of flexible filamentous plant virus nucleoproteins displays a fold closer to a common ancestor protein and that their homology with nucleoproteins of sNSV is still recognized. However, nucleoproteins from sNSV are diverse and show lower levels of structural homology between them. This is an indication of a higher structural divergence within sNSV nucleoproteins.
Landscape for putative evolution of nucleoproteins in ssRNA viruses. The two sets of viruses are segregated in two main groups, naked and plant viruses (left) and enveloped and animal viruses (right). Their nucleoproteins are represented by a green circle, and the N- and C-terminal extensions are depicted red and yellow, respectively. In the nucleoprotein (or CP) of flexible filamentous plant virus, the star indicates the conserved RNA-binding site. LCA: last common ancestor
The evolution of RNA viruses is hard to unveil, and in the current scenario, we do not known how the nucleoprotein gene has spread along several families of ssRNA viruses. Some viral evolutionary mechanism such as cross-species transmission (Geoghegan et al. 2017) and transfer of genes between virus and host (Aiewsakun and Katzourakis 2015) have recently been acknowledged as frequent events. For instance, CP sequences from potato virus Y (PVY, a potyvirus) have been found in the genomes of grapevines, probably after nonhomologous recombination with retrotransposable elements (Tanne and Sela 2005). In the same line, genomic sequences from bunyaviruses and orthomyxoviruses have been found as endogenous viral elements in insects and crustaceans (Theze et al. 2014; Ballinger et al. 2013; Katzourakis and Gifford 2010). Importantly, RNA sequencing works have found a large genomic diversity of RNA viruses and related sequences in invertebrates (Shi et al. 2016; Li et al. 2015). The phylogenetic analysis of these viromes reveals frequent recombination, gene transfer events, and genetic reassortments. Invertebrates, specially insects, play a central role as vectors for several of the ssRNA viruses discussed in this chapter and might have provided a niche for an evolutionary explosion of eukaryotic RNA viruses (Koonin et al. 2015).
Regardless of the evolutionary mechanisms that transferred the nucleoprotein gene, there are some general trends that might explain the current diversity of morphologies in these viral families. Naked and filamentous forms are linked to plant- infecting viruses (Fig. 6.8), while enveloped viruses are essentially animal pathogens, with the exception of tospoviruses (within the groups discussed in this chapter). There is a clear relationship between the presence of a cell wall in the host cell and the lack of viral envelope (Buchmann and Holmes 2015). Also, the need to cross the narrow plasmodesmata between plant cells during infection favors filamentous versus spherical virions in plant viruses (Hong and Ju 2017). It is possible that the naked nucleoproteins from flexuous plant viruses undergo functional restrictions that limit their structural variation and they conserve a very close fold and a specific RNA-binding site. Nucleoproteins from sNSV, however, are protected inside the membranous envelope and have explored a wider structural landscape and several oligomerization strategies.
References
Abad-Zapatero C, Abdel-Meguid SS, Johnson JE, Leslie AG, Rayment I, Rossmann MG, Suck D, Tsukihara T (1980) Structure of southern bean mosaic virus at 2.8 A resolution. Nature 286(5768):33–39
Abrescia NG, Bamford DH, Grimes JM, Stuart DI (2012) Structure unifies the viral universe. Annu Rev Biochem 81:795–822. https://doi.org/10.1146/annurev-biochem-060910-095130
Adams MJ, Lefkowitz EJ, King AMQ, Harrach B, Harrison RL, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Mushegian AR, Nibert M, Sabanadzovic S, Sanfacon H, Siddell SG, Simmonds P, Varsani A, Zerbini FM, Gorbalenya AE, Davison AJ (2017) Changes to taxonomy and the international code of virus classification and nomenclature ratified by the international committee on taxonomy of viruses (2017). Arch Virol 162:2505. https://doi.org/10.1007/s00705-017-3358-5
Agirrezabala X, Mendez-Lopez E, Lasso G, Sanchez-Pina MA, Aranda M, Valle M (2015) The near-atomic cryoEM structure of a flexible filamentous plant virus shows homology of its coat protein with nucleoproteins of animal viruses. Elife 4:e11795. https://doi.org/10.7554/eLife.11795
Aiewsakun P, Katzourakis A (2015) Endogenous viruses: connecting recent and ancient viral evolution. Virology 479-480:26–37. https://doi.org/10.1016/j.virol.2015.02.011
Ariza A, Tanner SJ, Walter CT, Dent KC, Shepherd DA, Wu W, Matthews SV, Hiscox JA, Green TJ, Luo M, Elliott RM, Fooks AR, Ashcroft AE, Stonehouse NJ, Ranson NA, Barr JN, Edwards TA (2013) Nucleocapsid protein structures from orthobunyaviruses reveal insight into ribonucleoprotein architecture and RNA polymerization. Nucleic Acids Res 41(11):5912–5926. https://doi.org/10.1093/nar/gkt268
Arranz R, Coloma R, Chichon FJ, Conesa JJ, Carrascosa JL, Valpuesta JM, Ortin J, Martin-Benito J (2012) The structure of native influenza virion ribonucleoproteins. Science 338(6114):1634–1637. https://doi.org/10.1126/science.1228172
Ballinger MJ, Bruenn JA, Kotov AA, Taylor DJ (2013) Selectively maintained paleoviruses in Holarctic water fleas reveal an ancient origin for phleboviruses. Virology 446(1–2):276–282. https://doi.org/10.1016/j.virol.2013.07.032
Baltimore D (1971) Expression of animal virus genomes. Bacteriol Rev 35(3):235–241
Buchmann JP, Holmes EC (2015) Cell walls and the convergent evolution of the viral envelope. Microbiol Mol Biol Rev 79(4):403–418. https://doi.org/10.1128/MMBR.00017-15
Chenavas S, Estrozi LF, Slama-Schwok A, Delmas B, Di Primo C, Baudin F, Li X, Crepin T, Ruigrok RW (2013) Monomeric nucleoprotein of influenza A virus. PLoS Pathog 9(3):e1003275. https://doi.org/10.1371/journal.ppat.1003275
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6):1188–1190. https://doi.org/10.1101/gr.849004
DiMaio F, Chen CC, Yu X, Frenz B, Hsu YH, Lin NS, Egelman EH (2015) The molecular basis for flexibility in the flexible filamentous plant viruses. Nat Struct Mol Biol 22(8):642–644. https://doi.org/10.1038/nsmb.3054
Dolja VV, Boyko VP, Agranovsky AA, Koonin EV (1991) Phylogeny of capsid proteins of rod-shaped and filamentous RNA plant viruses: two families with distinct patterns of sequence and probably structure conservation. Virology 184(1):79–86
Dong H, Li P, Bottcher B, Elliott RM, Dong C (2013) Crystal structure of Schmallenberg orthobunyavirus nucleoprotein-RNA complex reveals a novel RNA sequestration mechanism. RNA 19(8):1129–1136. https://doi.org/10.1261/rna.039057.113
Freiberg AN, Sherman MB, Morais MC, Holbrook MR, Watowich SJ (2008) Three-dimensional organization of Rift Valley fever virus revealed by cryoelectron tomography. J Virol 82(21):10341–10348. https://doi.org/10.1128/JVI.01191-08
Geoghegan JL, Duchene S, Holmes EC (2017) Comparative analysis estimates the relative frequencies of co-divergence and cross-species transmission within viral families. PLoS Pathog 13(2):e1006215. https://doi.org/10.1371/journal.ppat.1006215
Gough J (2005) Convergent evolution of domain architectures (is rare). Bioinformatics 21(8):1464–1471. https://doi.org/10.1093/bioinformatics/bti204
Harrison SC, Olson AJ, Schutt CE, Winkler FK, Bricogne G (1978) Tomato bushy stunt virus at 2.9 A resolution. Nature 276(5686):368–373
Hastie KM, Kimberlin CR, Zandonatti MA, MacRae IJ, Saphire EO (2011) Structure of the Lassa virus nucleoprotein reveals a dsRNA-specific 3′ to 5′ exonuclease activity essential for immune suppression. Proc Natl Acad Sci U S A 108(6):2396–2401. https://doi.org/10.1073/pnas.1016404108
Hong JS, Ju HJ (2017) The plant cellular systems for plant virus movement. Plant Pathol J 33(3):213–228. https://doi.org/10.5423/PPJ.RW.09.2016.0198
Hsu MT, Parvin JD, Gupta S, Krystal M, Palese P (1987) Genomic RNAs of influenza viruses are held in a circular conformation in virions and in infected cells by a terminal panhandle. Proc Natl Acad Sci U S A 84(22):8140–8144
Huiskonen JT, Overby AK, Weber F, Grunewald K (2009) Electron cryo-microscopy and single-particle averaging of Rift Valley fever virus: evidence for GN-GC glycoprotein heterodimers. J Virol 83(8):3762–3769. https://doi.org/10.1128/JVI.02483-08
Illergard K, Ardell DH, Elofsson A (2009) Structure is three to ten times more conserved than sequence--a study of structural response in protein cores. Proteins 77(3):499–508. https://doi.org/10.1002/prot.22458
Katzourakis A, Gifford RJ (2010) Endogenous viral elements in animal genomes. PLoS Genet 6(11):e1001191. https://doi.org/10.1371/journal.pgen.1001191
Kawabata T (2003) MATRAS: A program for protein 3D structure comparison. Nucleic Acids Res 31(13):3367–3369
Kendall A, McDonald M, Bian W, Bowles T, Baumgarten SC, Shi J, Stewart PL, Bullitt E, Gore D, Irving TC, Havens WM, Ghabrial SA, Wall JS, Stubbs G (2008) Structure of flexible filamentous plant viruses. J Virol 82(19):9546–9554. https://doi.org/10.1128/JVI.00895-08
Kendall A, Bian W, Maris A, Azzo C, Groom J, Williams D, Shi J, Stewart PL, Wall JS, Stubbs G (2013) A common structure for the potexviruses. Virology 436(1):173–178. https://doi.org/10.1016/j.virol.2012.11.008
Komoda K, Narita M, Yamashita K, Tanaka I, Yao M (n.d.) Tomato spotted wilt tospovirus nucleocapsid protein-ssRNA complex. pdb code 5IP2
Koonin EV (1991) The phylogeny of RNA-dependent RNA polymerases of positive-strand RNA viruses. J Gen Virol 72(Pt 9):2197–2206. https://doi.org/10.1099/0022-1317-72-9-2197
Koonin EV, Dolja VV (1993) Evolution and taxonomy of positive-strand RNA viruses: implications of comparative analysis of amino acid sequences. Crit Rev Biochem Mol Biol 28(5):375–430. https://doi.org/10.3109/10409239309078440
Koonin EV, Dolja VV, Krupovic M (2015) Origins and evolution of viruses of eukaryotes: the ultimate modularity. Virology 479-480C:2–25. https://doi.org/10.1016/j.virol.2015.02.039
Kormelink R, Garcia ML, Goodin M, Sasaya T, Haenni AL (2011) Negative-strand RNA viruses: the plant-infecting counterparts. Virus Res 162(1–2):184–202. https://doi.org/10.1016/j.virusres.2011.09.028
Li CX, Shi M, Tian JH, Lin XD, Kang YJ, Chen LJ, Qin XC, Xu J, Holmes EC, Zhang YZ (2015) Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. Elife 4:e05378. https://doi.org/10.7554/eLife.05378
Lyles DS (2013) Assembly and budding of negative-strand RNA viruses. Adv Virus Res 85:57–90. https://doi.org/10.1016/B978-0-12-408116-1.00003-3
Nagy PD, Pogany J (2011) The dependence of viral RNA replication on co-opted host factors. Nat Rev Microbiol 10(2):137–149. https://doi.org/10.1038/nrmicro2692
Namba K, Stubbs G (1986) Structure of tobacco mosaic virus at 3.6 A resolution: implications for assembly. Science 231(4744):1401–1406
Nasir A, Caetano-Anolles G (2017) Identification of capsid/coat related protein folds and their utility for virus classification. Front Microbiol 8:380. https://doi.org/10.3389/fmicb.2017.00380
Niu F, Shaw N, Wang YE, Jiao L, Ding W, Li X, Zhu P, Upur H, Ouyang S, Cheng G, Liu ZJ (2013) Structure of the Leanyer orthobunyavirus nucleoprotein-RNA complex reveals unique architecture for RNA encapsidation. Proc Natl Acad Sci U S A 110(22):9054–9059. https://doi.org/10.1073/pnas.1300035110
Qi X, Lan S, Wang W, Schelde LM, Dong H, Wallat GD, Ly H, Liang Y, Dong C (2010) Cap binding and immune evasion revealed by Lassa nucleoprotein structure. Nature 468(7325):779–783. https://doi.org/10.1038/nature09605
Raju R, Kolakofsky D (1989) The ends of La Crosse virus genome and antigenome RNAs within nucleocapsids are base paired. J Virol 63(1):122–128
Raymond DD, Piper ME, Gerrard SR, Skiniotis G, Smith JL (2012) Phleboviruses encapsidate their genomes by sequestering RNA bases. Proc Natl Acad Sci U S A 109(47):19208–19213. https://doi.org/10.1073/pnas.1213553109
Reguera J, Malet H, Weber F, Cusack S (2013) Structural basis for encapsidation of genomic RNA by La Crosse Orthobunyavirus nucleoprotein. Proc Natl Acad Sci U S A 110(18):7246–7251. https://doi.org/10.1073/pnas.1302298110
Reguera J, Cusack S, Kolakofsky D (2014) Segmented negative strand RNA virus nucleoprotein structure. Curr Opin Virol 5:7–15. https://doi.org/10.1016/j.coviro.2014.01.003
Rossmann MG, Abad-Zapatero C, Murthy MR, Liljas L, Jones TA, Strandberg B (1983) Structural comparisons of some small spherical plant viruses. J Mol Biol 165(4):711–736
Ruigrok RW, Crepin T, Kolakofsky D (2011) Nucleoproteins and nucleocapsids of negative-strand RNA viruses. Curr Opin Microbiol 14(4):504–510. https://doi.org/10.1016/j.mib.2011.07.011
Sherman MB, Freiberg AN, Holbrook MR, Watowich SJ (2009) Single-particle cryo-electron microscopy of Rift Valley fever virus. Virology 387(1):11–15. https://doi.org/10.1016/j.virol.2009.02.038
Shi M, Lin XD, Tian JH, Chen LJ, Chen X, Li CX, Qin XC, Li J, Cao JP, Eden JS, Buchmann J, Wang W, Xu J, Holmes EC, Zhang YZ (2016) Redefining the invertebrate RNA virosphere. Nature 540:539. https://doi.org/10.1038/nature20167
Tanne E, Sela I (2005) Occurrence of a DNA sequence of a non-retro RNA virus in a host plant genome and its expression: evidence for recombination between viral and host RNAs. Virology 332(2):614–622. https://doi.org/10.1016/j.virol.2004.11.007
Theze J, Leclercq S, Moumen B, Cordaux R, Gilbert C (2014) Remarkable diversity of endogenous viruses in a crustacean genome. Genome Biol Evol 6(8):2129–2140. https://doi.org/10.1093/gbe/evu163
Yang S, Wang T, Bohon J, Gagne ME, Bolduc M, Leclerc D, Li H (2012) Crystal structure of the coat protein of the flexible filamentous papaya mosaic virus. J Mol Biol 422(2):263–273. https://doi.org/10.1016/j.jmb.2012.05.032
Ye Q, Krug RM, Tao YJ (2006) The mechanism by which influenza a virus nucleoprotein forms oligomers and binds RNA. Nature 444(7122):1078–1082. https://doi.org/10.1038/nature05379
Zamora M, Mendez-Lopez E, Agirrezabala X, Cuesta R, Lavin JL, Sanchez-Pina MA, Aranda M, Valle M (2017) Potyvirus virion structure shows conserved protein fold and RNA binding site in ssRNA viruses. Sci Adv 3(9):eaao2182. https://doi.org/10.1126/sciadv.aao2182
Zhou H, Sun Y, Guo Y, Lou Z (2013) Structural perspective on the formation of ribonucleoprotein complex in negative-sense single-stranded RNA viruses. Trends Microbiol 21(9):475–484. https://doi.org/10.1016/j.tim.2013.07.006
Acknowledgments
This work was supported by grant BFU2015-66326-P from the Spanish Ministry of Economy and Competitiveness (MINECO). Also the Severo Ochoa Excellence Accreditation (SEV-2016-0644) by MINECO is acknowledged.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Valle, M. (2018). Structural Homology Between Nucleoproteins of ssRNA Viruses. In: Harris, J., Bhella, D. (eds) Virus Protein and Nucleoprotein Complexes. Subcellular Biochemistry, vol 88. Springer, Singapore. https://doi.org/10.1007/978-981-10-8456-0_6
Download citation
DOI: https://doi.org/10.1007/978-981-10-8456-0_6
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8455-3
Online ISBN: 978-981-10-8456-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)