Abstract
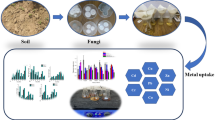
Trace metal (TM) pollution of soil is a worldwide problem threatening the quality of human life and a proper environment. We investigated fungal and bacterial diversity of trace metal-polluted site contaminated with paper mill effluent in India. Twelve fungal dominant isolates, viz. Aspergillus, Penicillium, Fusarium, Cunninghamella, Simplicillium, Trichoderma, Rhizomucor, Cladosporium and Hypocrea, were identified. Subsequent screening approach to assess their TM tolerance was performed in vitro cultures which revealed that the majority of the isolates were tolerant to Ni-, Cu-, Zn- and Cd-amended medium. The minimum inhibitory concentration (MIC) for Ni, Cu, Zn and Cd was also determined in isolated strains of Aspergillus, Penicillium, Rhizomucor, Trichoderma and Fusarium to study the concentration of growth against various trace metals. A total of 22 bacterial isolates was also isolated using 16S rRNA, and the dominant genera such as Bacillus, Rhizobium, Microbacterium, Arthrobacter, Kribbella and Chitinophaga were identified. The relative growth and LD−50 were also estimated against the different trace metals from concentration 0.1 to 4 mM. Thus, these fungal and bacterial isolates showed a high TM tolerance and would be a great interest for their use in bioremediation to clean up TM-polluted soil.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
1.1 Microbial Diversity in Extreme Habitats
Soil harbours a variety of fungi, bacteria and other soil microorganisms. Soil microorganisms are the living component of soil organic matter and are responsible for mineralization of nutrients, decomposition and degradation or transformation of toxic compounds. Metals and metal compounds are natural constituents of all ecosystems, moving between atmosphere, hydrosphere, lithosphere and biosphere (Bargagli 2000; Wuana and Okieimen 2011). One of the challenges facing the mankind in recent times is the degradation and pollution of soil. Since soil is a vital natural resource, its degradation threatens the basic life support system. The industrial influent’s sludge and solid waste are the sources of potentially harmful inorganic as well as organic contaminants. Microorganisms growing in such habitats evolved under conditions that permitted their survival and growth (Thakre and Shanware 2015). They multiplied in accordance with natural selection. For such adapted microorganisms, the conditions of these habitats are not ‘extreme’ but rather the normal physiological conditions for their growth in their natural habitats (Ali et al.2013; Akponah 2013; Kumar et al. 2014; Smith et al. 2015). In metal-contaminated soils, the siderophores and plant growth hormones are produced by plant-associated microbes (Pattus and Abdallah 2000; Wu et al. 2006; Schalk et al. 2011; Ullah et al. 2015). The secretion of siderophores by fungi and bacteria is dependent on several factors like soil pH, nutrient availability in soils and type and concentration of trace metals (Rajkumar et al. 2010; Sessitsch et al. 2013; Yu et al. 2014).
1.2 Heavy Metal Resistance in Fungi
Trace metals like Cu, Ni, Zn, Cd and Mn present in paper mill effluent can be removed by indigenous fungi isolated from effluent itself (Khan 2000, Karn and Reddy 2012). Biosorption of metal is carried out by (1) extracellular accumulation/precipitation, (2) cell surface sorption/precipitation and (3) intracellular accumulation through the cell wall of microorganisms (Volesky and Holan 1995; Valix et al. 2001; Madhaiyan et al. 2007; Ma et al. 2016). Penicillium, Aspergillus, Trichoderma, Cladosporium, etc., are found to be very useful for the removal of trace metals (Dursun 2008; Ezzourhi et al. 2009; de Lima et al. 2011). El-Morsy (2004) reported that Cunninghamella echinulata biomass could be employed as a biosorbent of metal ions in waste water. De Lima et al. (2013) and Bello and Abdullahi (2016) also studied the cadmium tolerance by Cunninghamella elegans by the polyphosphate metabolism. Trichoderma sp. produces organic acids like fumaric acid, citric acid and glycolic acid which can decrease the pH in alkaline soil and thus increase the solubility of macro- and micronutrients necessary for plant growth and metabolism (Malgorzata et al. 2014; Song et al. 2015).
1.3 Heavy Metal Resistance of Bacteria
Heavy metals can decrease carbon mineralization, nitrogen transformation and soil enzyme activities, microbial numbers (CFU), biomass (Borjesson et al. 2012) and frequency of trace metal-resistant bacteria (Wang et al. 2007, Kanmami et al. 2012). The molecular fingerprinting techniques are also useful to study the changes in the microbial community in trace metal stress conditions (Anyanwu et al. 2011; Andrew et al. 2013). Bacterial populations negatively affected by trace metals. Bacteria are found to develop five important mechanisms to detoxify the trace metals available in contaminated soils: (1) extracellular detoxification, (2) extracellular sequestration, (3) reduced permeability, (4) intracellular sequestration and (5) export. These resistant mechanisms are encoded in bacterial plasmids and transposons due to spontaneous mutation and gene transfer (Osborn et al. 1997; Karelová et al. 2011; Cetin et al. 2012; Zhou et al. 2013). Pal et al. (2004) reported Ni-resistant genes in Gram-positive and Gram-negative bacterial isolates from Ni-rich serpentine soil. In Gram-negative bacteria, the czc-genes encode for a cation-proton antiporter (CzcABC) which is responsible for the resistance against Cd, Zn and Co metals (Nies 1995; Harriso et al. 2007; Abdelatey et al. 2011; Mindlin et al. 2016).
The trace metal tolerance by a particular group of bacteria or isolate in artificial medium supplemented with trace metal showed high tolerance level as reported by Ahmed et al. (2001), Hayat et al. (2002) and Rajbanshi (2008). Olukoya et al. (1997) isolated 228 trace metal-resistant bacteria belonging to 9 genera, and the most common genera were Staphylococcus, Streptococcus and Bacillus found to be resistant to cobalt, zinc, copper, nickel and mercury. Temperature is also a determined factor that affects the growth of bacteria and bioaccumulation of trace metals (Lee et al. 2011a, b). The gene expression study revealed that mercuric ion (merA) and chromate (chrB) genes were downregulated in all the strains of bacteria, i.e. S. aureus, Bacillus subtilis, B. cereus, Pseudomonas sp. and Bordetella sp., when treated with Co and Cd. The expression level of genes merA, chrB, czc D and ncc A in these bacterial strains was measured by real-time PCR method (Abou-Shanab et al. 2007). Nies (1999) and Hirak and Das (2014) compared the metal resistance physiology in 63 species of bacteria and examined the protein-level similarities and suggested that these metal-resistant bacteria can be developed into metal pollution biosensors. Long et al. (2012) described the importance efflux transporters as a metal tolerance lactic by bacteria. Braud et al. (2010) reported a low level of toxicity of trace metals like Ni, Cu, Zn, Cd and Pb in Pseudomonas aeruginosa. Chitinophaga eiseniae was also reported as a trace metal tolerant by Yasir et al. (2011) and Gao et al. (2012); Stan et al. (2011) studied the significant increase of growth, abundance, genetic diversity, nodulation ability and efficacy in the diversity of Rhizobium sp. in the soil polluted with copper, zinc and lead. Hemida et al. (2012) and Hao et al. (2014) also discussed the potential role of legume-rhizobia symbiosis in aiding phytoremediation. Hijri et al. (2014) also studied the linkage between fungal and bacterial communities in rhizosphere in hydrocarbon-contaminated soil and their significant effect for plant productivity.
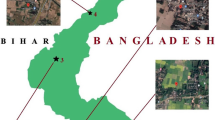
The present study was carried out to understand and evaluate the status of heavy metal-resistant fungi, bacteria and actinomycetes in the Hindustan Paper Corporation (HPC), Assam. Geographically the site is situated at longitude of 24°41′29.9˝N and latitude at 92°45′25.9˝E.
1.4 Characterization of Fungal and Metal-Resistant Bacteria Isolates
The fungal isolates were isolated and were identified to species level using colony diameter and spore measurement following references and monographs adopted by Gilman (1957) and Raper and Fennell (1965). The fungal DNA was isolated with help of nucleic acid and protein purification kit (Macherey-Nagel, USA). The fungal strains have been characterized by PCR with (forward) ITS1 5′-TCCGTAGGTGAACCTGCGG-3′ and (reverse) ITS43′TCCTCCGCTTATTGATATGC-5′ (White et al. 1990).
The isolation and purification of chromosomal DNA as well as the amplification and sequencing of partial 16S rRNA gene of potential metal-resistant bacteria isolate was carried out. Bacterial 16S rDNA sequences were amplified using the 27F Lane (1991) and 1492R Turner et al. (1999) primer sets.
The selected bacterial isolate was tested for their resistance to different trace metals by their growth in nutrient broth tubes containing various concentrations of trace metals (0.1, 0.5, 2.0, 4.0 mM). The metals selected for the present investigation included Ni, Cu, Zn and Cd. The bacterial growth was determined by measuring the optical density using spectrophotometer at 540 nm. Relative growth of the isolate was expressed as the percentage of those obtained in untreated control. Lethal dose (LD-50) was estimated for all the tested bacterial isolates (Essam et al. 2013; Anderson and Hughes 2014). DNA sequencing was performed on an Applied Biosystems 3730xl DNA Analyzer (Applied Biosystems, Carlsbad, CA). The nucleotide sequence of bacterial isolate is thus obtained by the use of database using the ‘NCBI BLAST’ (Altschul et al. 1990).
1.5 Metal-Resistant Fungal Isolates
Twelve fungal strains were isolated from polluted soil contaminated with trace metals in paper mill effluents and tested with different trace metals in different concentrations. The 12 genera like Aspergillus, Penicillium, Cladosporium, Cunninghamella, Trichoderma, Fusarium and Hypocrea showed significant tolerance against various trace metals (Table 8.1). Minimum inhibitory concentration (MIC) of the isolated fungal strains against the different concentration of trace metals was estimated and found that, at higher metal ion concentrations, most of the tested fungal strains were found tolerant and showed strong growth (Table 8.2).
In the presence of various concentrations of nickel, the fungal strains which were able to grow in 15–20 mM were Trichoderma sp., Penicillium sp., Rhizomucor sp., Cladosporium sp. and Hypocrea sp. The other tested strains like Penicillium, Aspergillus and Cunninghamella were also to grow in MIC of 10–15 mM (Plates 8.1 and 8.2).
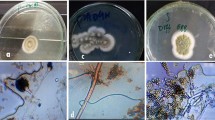
In the presence of various concentrations of copper, most of the tested strains showed a very low MIC except Cunninghamella and Cladosporium where MIC range was 5–10 mM. Their mycelia became diffused compared with the control. All strains studied could not grow in higher concentrations except Rhizomucor sp. (KC602345) which showed the highest MIC of 15–25 mM. The white colour of the mycelium became blue green due to accumulation of Cu ions inside the cell wall of the tested fungi (Plate 8.3). The growth rate of fungi tested was reduced, and their conidiogenesis was also slowed down. Addition of copper sulphate to the PDA resulted in the growth of the isolated fungal strains and changed the colour and morphology of the mycelium. The mycelium of Cladosporium sp. (KC602374) secreted a deep brown substance (Plate 8.4), and the Fusarium isolate (KC602349) (Plate 8.5) secreted violet pigment due to the response to the metal stress.
In the presence of various concentrations of cadmium, the isolates Trichoderma, Aspergillus sp. and Penicillium sp. showed a high MIC with 15–20 mM. When the concentration of cadmium increased in the media, the absorbance of the fungal culture was found to be decreased. The most tolerant fungi which were found to grow in high concentration of the trace metals were Penicillium sp. (KC602310), Trichoderma sp. (KC602314), Aspergillus sp. (KC602350), Fusarium sp. (KC602349), Hypocrea sp. (KC602373), Penicillium janthinellum (KC602344) and Cladosporium (KC602374). The value of tolerance index of Penicillium sp. (KC602310 and KC602359), Aspergillus sp. (KC602350 and KC602371), Rhizomucor sp. (KC602345), Fusarium sp. (KC602349) and Trichoderma sp. (KC602314 and KC602331) showed a maximum value of 0.9 tested against all the metals, i.e. Ni, Cu, Zn and Cd (Figs. 8.1, 8.2, 8.3 and 8.4 ).
1.6 Identification and Characterization of Metal-Resistant Bacteria Isolates
Twenty-two bacterial isolates showed resistance to different trace metals, and the molecular characterization for these isolates was carried out (Table 8.3). The trace metals like Ni, Cu, Zn and Cd were selected in a concentration ranged from 0.1 to 4.0 mM for identification. Among the various genera, Bacillus, Agromyces, Microbacterium, Arthrobacter, Chitinophaga, Rhizobium and Kribbella were showing a range of 30–40% relative growth at the higher concentrations of all heavy metals tested. These bacterial isolates are capable to grow at higher concentrations of trace metals, and thus they were resistant to Ni, Cu, Zn and Cd. The species of Agromyces, Bacillus, Chittinophaga and Kribbella (isolates 1, 3, 4, 6, 10) showed significant relative growth values ranging from 40 to 70% at 2 mM and 4 mM concentrations of zinc. The species of Rhizobium, Bacillus and Arthrobacter showed a range of 20–60% of relative growth at 2 mM and 4 mM concentrations of nickel, copper and cadmium. The species of Bacillus and Microbacterium (isolates 18 and 19) showed a range of 20–30% of relative growth at 2 mM and 4 mM concentrations of cadmium.
The isolated trace metal-resistant bacterial strains that were identified with their accession numbers are Bacillus cereus (KC602258), Bacillus sp. (KC602265), Chitinophaga sp. (KC602266), Chitinophaga bacter (KC602269), Rhizobium sp. (KC602276), Microbacterium sp. (KC602277), Bacillus sp. (KC602282), Kribbella sp. (KC602294), Arthrobacter sp. (KC602298), Arthrobacter oryzae (KC602305) and Arthrobacter nicotinovorans (KC602306) and were found to show positive test as tested against the 15 sugars, i.e. glucose, sucrose, xylose, maltose, rhamnose, rafffinose, cellobiose, dextrose, galactose, arabinose, lactose, sorbitol, melibiose, saccharose and trehalose. All the bacterial strains were tested for antibiotic sensitivity (Bauer 1996). Most of the isolates of Bacillus, Agromyces, Microbacterium, Arthrobacter, Chitinophaga, Rhizobium, Brachybacterium and Kribbella appeared to be inhibited by eight antibiotics and resistant to ampicillin, while Chitinophaga sp. (KC602269) was resistant to chloramphenicol (Adesoji et al. 2015). Among all the strains tested, the isolates (KC602240, KC602277, KC602301, KC602283 and KC602286) showed resistance to ampicillin, whereas the rest showed no inhibition. The antibiotics like streptomycin, polymyxin B, vancomycin, tetracycline, gentamicin, amikacin, ciprofloxacin and levofloxacin were found to be susceptible to all the 22 tested strains.
Lethal dose (LD-50) was estimated for all the tested bacterial isolates. The species of Arthrobacter, Chitinophaga, Kribbella, Microbacterium, Bacillus, Agromyces and Rhizobium showed a significant range of LD-50 values (0.2–1.8) tested against zinc, (0.3–1.6) for copper, (0.6–1.5) for nickel and (0.1–0.8) for cadmium. The highest LD-50 value of 1.8 was showed by the Chitinophaga sp. (KC602266), while the highest LD-50 value of 1.6 was showed by Chitinophaga bacter sp. (KC602269) and Microbacterium sp. (KC602277) against Ni. The highest LD-50 value of 0.8 was showed by the Chitinophaga sp. (KC602266), Arthrobacter (KC602303) and Agromyces (KC602270) against the metal Cd (Fig. 8.5 ).
The fungal and bacterial sequences were analysed by the Basic Local Alignment Tool (BLAST) for finding the closest homologous sequences. These sequences were saved in a fasta format (*.fasta or *.txt) and aligned with CLUSTAL X2. The output of CLUSTAL (i.e. *.aln file) was saved for the output of MEGA version 5. Phylogenetic and molecular evolutionary analysis was carried out by MEGA version 5 (Tamura et al. 2011). A distance matrix was made based on nucleotide sequence homology, and neighbour joining (NJ) consensus trees were obtained using Kimura-2 parameter substitution model (MEGA 5) (Saitou and Nie, 1987). The bootstrap values above 50% and the genetic distance scale are shown for the relationship of the isolated fungal (Figs. 8.6, 8.7, 8.8, 8.9 and 8.10) and bacterial strains (Figs. 8.11, 8.12, 8.13, 8.14, 8.15, 8.16 and 8.17) with their closely related neighbouring species.
The present experimental findings revealed the effects of trace metals on microbial diversity, i.e. fungi, bacteria and actinomycetes, in the polluted site of Hindustan Paper Corporation (HPC) paper mill. The diversity and abundance of soil microorganisms were found to be affected by naturally occurring environmental variables, including soil types, soil pH, moisture content and natural availability. Carson et al. (2010) and Stefanowicz et al. (2010) also reported that the soil microorganisms are affected positively by environmental factors. All the isolated strains of fungi, bacteria and actinomycetes were found to be resistant to various trace metals at higher concentrations. Similar observations were observed by Freitas et al. (2009) and Appenroth (2010). Soil microbial populations were found to multiply even under metal-contaminated soil which in turn maintains the diversity of fungi and bacteria (Chen et al. 2014). The resistance of the selected strains to Cr6+, Pb2+, Zn2+ and Cu2+ was determined by the dilution method to calculate the tolerance index for all the tested fungi. Penicillium sp. (KC602310), Trichoderma sp. (KC602314), Aspergillus sp. (KC602350), Fusarium sp. (KC602349), Hypocrea sp. (KC602373), Penicillium janthinellum (KC602344) and Cladosporium (KC602374) were reported for their great importance in removal of trace metals from contaminated site. Some deuteromycetes have been reported by Ghorbani et al. (2007) and Zafar et al. (2007). Metals such as copper and zinc are essential to bioactivities; however, they tend to show toxicity after a certain level.
The fungal strains which were able to grow in 15–20 mM were Trichoderma sp., Penicillium sp., Rhizomucor sp., Cladosporium sp. and Hypocrea sp. The other tested strains like Penicillium, Aspergillus and Cunninghamella were also to grow in MIC of 10–15 mM (Table 8.2). Rao et al. (2005) and Sun and Shah (2007) also observed that with the increasing metal concentration of trace metals, the fungi Aspergillus niger and Cunninghamella echinulata can increase the rate of metal removal by saturation adsorbent concentrations by increasing mobilization of metal ions (Burford et al. 2003, Thippeswamy et al. 2012, 2014). Penicillium and Aspergillus showed a higher metal tolerance against nickel. Similar effects were also observed by Shivkumar et al. (2011) who discussed the high tolerance and bioaccumulation ability in Penicillium sp. and Rhizopus sp. against the various trace metal like Cu2+, Zn2+, Cd2+, Ni2+ and Pb2+. The growth of all fungi tested was decreased after addition of copper in high concentration in comparison with zinc, nickel and cadmium. All strains studied could not grow in higher concentrations except Rhizomucor sp. (KC602345) which showed the highest MIC of 15–25 mM. Van and Christov (2002) and Tripathi et al. (2007) also observed that Rhizomucor pusillus adsorption capacity was isolated from effluent plant. Rouhollahi et al. (2014) studied the nickel biosorption capacity of Rhizomucor pusillus by enzymatic and alkali treatments. The white colour of the mycelium became blue green due to accumulation of Cu ions inside the cell wall of the tested fungi. Copper tolerance in fungi ascribed to diverse mechanisms also described by Cervantes and Gutierrez (1994). The most of the tested strains showed a very low MIC except Cunninghamella and Cladosporium where MIC range was 5–10 mM. The morphology of strains was highly affected by the presence of Cu. Their mycelia became diffused compared with the control. The growth rate of fungi tested was reduced, and their conidiogenesis was also slowed down. In Cladosporium sp. (KC602374), the mycelia changed into deep brown colour in the high concentration of Cu. The tolerance of the tested fungi to high copper concentrations could be related to metallothioneins and other thiol compounds which may be promising detoxifying agents for copper as reported by Malik (2004) and Dusrun (2008). Similar biosorption mechanisms were also reported by Juliana et al. (2013) who discussed the biomass of Cladosporium as an efficient biosorbent of copper.
The fungal colour and morphology were both affected by high Zn concentrations in Fusarium sp. as the mycelium changed to violet pigment which is probably due to the stress imposed by the Zn. The zinc MIC was in the range 20–25 mM, 15–20 mM, 10–15 mM and 5–10 mM for the Fusarium sp. Biosorption of various trace metals by Fusarium sp. was also reported earlier by Sen (2011), Zhang et al. (2012) and Verma et al. (2016).
The isolates Trichoderma, Aspergillus niger, Cunninghamella sp. and Penicillium sp. showed a high MIC with 15–20 mM in Cd-amended media. DeLima et al. (2011, 2013) also reported a higher potential of cadmium tolerance in the fungi Trichoderma harzianum and Cunninghamella elegans. The dominant genus of fungi identified and characterized were Aspergillus, Penicillium, Fusarium, Cunninghamella, Trichoderma, Rhizomucor, Cladosporium and Hypocrea by PCR with (forward) ITS1 and (reverse) ITS4 from the polluted soil. This may be due to the processes of valence transformation, active uptake, complexation, crystallization and biosorption of trace metals to the fungal cell walls (Jaeckel et al. 2005; Willie et al. 2007; Palanivel et al. 2010; Anahid et al. 2011; Iram et al. 2012; Do Carmo et al. 2013; Rhodes 2013; Akhtar et al. 2013). Yazdani et al. (2009) and Malgorzata et al. (2014) found the application of Trichoderma sp. on various plant and found that this fungus has positive effects on increasing the biomass, soil parameters (C, N and P) and solubility of trace metals in soil, thereby enhancing phytoextraction in the plants. Copper tolerance of various Trichoderma sp. is also studied by Petrovic et al. (2014). Teng et al. (2015) also studied the phytoremediation in Cd-contaminated soil by Trichoderma reesei FS10-C strain.
A total of 22 bacterial isolates exhibited resistance to different trace metals. These bacterial isolates were capable to grow at higher concentrations of trace metals and showing different degree of resistance to Ni, Cu, Zn and Cd. The major bacterial genera were identified as Arthrobacter, Brachybacterium, Chitinophaga, Kribbella, Microbacterium, Bacillus, Agromyces and Rhizobium. The resistance of these bacterial strains towards trace metal could be a result of the interaction between the metals and amphoteric groups such as the carboxyl and phosphoryl groups. In the present study, Gram-positive bacteria showed a major group for absorption capacity than the Gram-negative isolates as tested against different trace metals as Gram-positive bacteria have high chemisorption sites (Tunali et al. 2006, Long et al. 2012). The glycoproteins present on the outer site of Gram-positive bacterial cells have more potential binding sites than the Gram-negative bacteria having an outer layer of lipopolysaccharide (LPS), phospholipids and proteins (Gupta et al. 2012, 2016; Issazadeh et al. 2013).
The isolates of Bacillus, Agromyces, Microbacterium, Arthrobacter, Chitinophaga, Rhizobium, Actinobacterium and Kribbella showed positive activity towards urease, nitrate, H2S production, citrate utilization, methyl red, malonate utilization, oxidase production, starch amylase and catalase activity. The Gram-positive isolates found to be positive against catalase and negative against oxidase activity were identified as Brachybacterium, Agromyces, Arthrobacter, Kribbella and Microbacterium. Similar observations were reported by different workers for these Gram-positive strains of same bacterial strain: Agromyces sp. (Chen et al. 2011; Thawai et al. 2011), Arthrobacter sp. (Elanvogvan et al. 2010; Rosales et al. 2012; Santa et al. 2013; Sahoo et al. 2014; Swer et al. 2016), Chitinophaga sp. (Gao et al., 2015), Kribbella sp. (Clara et al. 2008) and Microbacterium sp. (Mondani et al. 2012; Brown et al. 2012; Tappe et al.2013).
The strains of Microbacterium sp. showed a positive catalase activity and negative oxidase and H2S production. Piccirillo et al. (2013) also observed similar biochemical activities in Microbacterium oxydans to be tolerant against Zn (II) and Cd (II). The isolated strains of Brachybacterium sp. were observed negative for catalase and oxidase activities and positive against starch hydrolysis and found to have relative growth range of 40–80% on higher concentrations of Zn, Cu, Ni and Cd at 2 mM and 4 mM (Wang et al. 2009; Park et al. 2011). Various strains of Arthrobacter sp. were isolated and found to be resistant against different trace metals (Paris and Blondeau 1999; Bafana et al. 2010; Inga 2013). The genus Kribbella sp. was isolated and showed 40–90% of relative growth on different concentrations of trace metals. Biochemical tests showed positive against catalase activity, nitrate reduction and H2S production and negative against oxidase production. Similar chemotaxonomic characteristics were reported earlier by Carlsohn et al. (2007) who also reported the greater accumulation capacity of Kribbella aluminosa against the metal Pb, Fe, Zn and Cu when grown in medium with 200 ppm of Pb, Fe and Zn and 100 ppm of Cu.
Bacillus sp. was found to be resistant with a relative growth of 30–40% on higher concentrations of all the trace metals. The isolates were found to be positive against nitrate reduction, citrate utilization, oxidase production, starch amylase, methyl red test and catalase activity. Similar biochemical activities and multi-tolerance and bioremediation of trace metals in Bacillus strains were observed earlier by various workers (Rathnayake et al. 2009; Elsilk et al. 2014). Chitinophaga sp. was negative against oxidase, catalase and starch amylase tests. The similar biochemical characteristics were observed in Chitinophaga sp. by Lee et al. (2009) and Wang et al. (2014). Rhizobium sp. is a Gram-negative, aerobic, non-endospore-forming rods, showed positive results against nitrate and catalase test and negative against oxidase, indole, VP test and urease test (Kuykendall et al. 2005; Grison et al. 2015). Rhizobium sp. was found to grow with 20–40 % of relative growth of copper and nickel at higher concentrations. The resistance of Rhizobium towards trace metals can produce huge amount of extracellular polysaccharide (EPS) and lipopolysaccharide (LPS), which can attach most of the metals extracellularly, acting a first-defence barrier against trace metal stress (Mohamed et al. 2012; Mandal and Bhattacharyya 2012). Our results were supported by Reeve et al. (2002), Hemida et al. (2012) and Hao et al. (2014) who also observed that Rhizobium played a very important role of legume-rhizobia symbiosis in aiding phytoremediation of polluted site contaminated with trace metals (Mergeay et al. 2003; Piotrowska-Seget et al. 2005; Zhang et al. 2011; Aafi et al. 2012; Rajkumar et al. 2012; Yang et al. 2012; Adel et al. 2014).
2 Conclusion
The present study focused on the effect of trace metal on the diversity of microorganisms (fungi, bacteria, actinomycetes) in the Hindustan Paper Corporation (HPC), Cachar. The most tolerant fungi grown in high concentration of the trace metals were identified as Penicillium sp. (KC602310), Trichoderma sp. (KC602314), Aspergillus sp. (KC602350), Fusarium sp. (KC602349), Hypocrea sp. (KC602373), Penicillium janthinellum (KC602344) and Cladosporium sp. (KC602374).
The most tolerant bacteria grown in high concentration of the trace metals were identified as Bacillus cereus (KC602258), Bacillus sp. (KC602265), Chitinophaga sp. (KC602266), Chitinophaga bacter (KC602269), Rhizobium sp. (KC602276), Microbacterium sp. (KC602277), Bacillus sp. (KC602282), Kribbella sp. (KC602294), Arthrobacter sp. (KC602298), Arthrobacter oryzae (KC602305) and Arthrobacter nicotinovorans (KC602306).
From the results of the present investigation, it can be concluded that biotic and abiotic stress in trace metal-polluted soil of the paper mill greatly influenced the enzyme activity, composition and function of the indigenous microorganisms (fungi, bacteria, actinomycetes). The current study clearly showed that the native dominant resistant indigenous fungal, bacterial isolates can be used as a biosensor to assess the trace metal toxicity in the polluted environment. Thus, future research may be proposed for further advances in microbial genetics by studying the mechanism of metal-microbe-plant interactions and their potential use as metal-resistant microbial inoculants in microbial-assisted phytoremediation.
2.1 Future Prospective
With the increased demand of paper, the treatment of effluents emerges as most pressing problem in environmental protection. The current study clearly showed that the native dominant resistant indigenous fungal, bacterial isolates can be used as a biosensor to assess the heavy metal toxicity in the polluted environment contaminated with paper mill effluents. A further understanding of metal-microbe-plant interactions will increase our knowledge to design microbial-assisted phytoremediation in the trace metal-contaminated sites.
References
Aafi NE, Brhada F, Dary M, Maltouf AF, Pajuelo E (2012) Rhizostabilization of metals in soils using Lupinus luteus inoculated with the metal resistant rhizobacterium Serratia sp. MSMC 541. Int J Phytoremediation 14:261–274. doi:10.1080/15226514.2011.604693
Abdelatey LM, Khalil WKB, Ali TH, Mahrous KF (2011) Heavy metal resistance and gene expression analysis of metal resistance genes in Gram-positive and Gram-negative bacteria present in Egyptian soils. J Appl Sci Env San 6:201–211
Abou-Shanab RAI, Van Berkum P, Angle JS (2007) Heavy metal resistance and genotypic analysis of metal resistance genes in gram-positive and gram-negative bacteria present in Ni-rich serpentine soil and in the rhizosphere of Alyssum murale. Chemosphere 68:360–367. doi:10.1016/j.chemosphere.2006.12.051
Adel Al-Gheethi AS, Noril I, Lalung J, Azan AM, Nurfarehah ZA, Kadir AB (2014) Biosorption of heavy metals and cephalexin from secondary effluents by tolerant bacteria. Clean Technol Envir 16(1):137–148
Adesoji AT, Ogunjobi AA, Olatoye IO, Douglas DR (2015) Prevalence of tetracycline resistance genes among multi-drug resistant bacteria from selected water distribution systems in southwestern Nigeria. Ann Clin Microbiol Antimicrob 14:35. doi:10.1186/s12941-015-0093-1
Ahmed I, Hayat S, Ahmed A, Samiullah IA (2001) Metal and antibiotic resistance traits in Bradyrhizobium sp (cajanus) isolated from soil receiving oil refinery wastewater. World J Microbiol Biotechnol 17:379–384
Akhtar S, Mahmood-ul-Hassan M, Ahmad R, Suthor V, Yasin M (2013) Metal tolerance potential of filamentous fungi isolated from soils irrigated with untreated municipal effluent. Soil Environ 32:55–62
Akponah E (2013) Bacterial and fungal biosorbents of heavy metals associated with hydrocarbon contaminated rainforest soils of the Niger delta region of Nigeria. Eur J Exp Biol 3(3):95–100
Ali H, Khan E, Sajad MA (2013) Phytoremediation of metals-concepts and applications. Chemosphere 91:869–881. doi:10.1016/j.chemosphere.2013.01.075
Altschul SF, Gish W, Miller W, Meyer EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Anahid S, Yaghmaei S, Ghobadinejad Z (2011) Heavy metal tolerance of fungi. Scientia Iranica 18(3):502–508
Anderson DI, Hughes D (2014) Microbiological effects of sublethal levels of antibiotics. Nature Rev Microbiol 12:465–478. doi:10.1038/nrmicro3270
Andrew B, Brown M, Steven DS, Peter HT (2013) Microbial community responses to anthropogenically induced environmental change: towards a systems approach. Ecol Lett 16(Supplement S1):128–139. doi:10.1111/ele.12109
Anyanwu CU, Nwankwo SC, Moneke AN (2011) Soil bacterial response to introduced metal stress. Int J Basic Appl Sci 11(1):109–115
Appenroth, K-J (2010) Defination of heavy metals and their role in biological systems. In: Sheramati I, Varma A (eds) Soil heavy metals. Soil biology, vol 19. Springer, Berlin, pp 19–29
Bafana A, Krishnamurthi K, Patil M, Chakrabarti T (2010) Heavy metal resistance in Arthrobacter ramosus strain G2 isolated from mercuric salt-contaminated soil. J Hazard Mater 177(1–3):481–486. doi:10.1016/j.saa.2009.12.058
Bargagli R (2000) Trace metals in Antarctica related to climate change and increasing human impact. Rev Environ Contam Toxicol 166:129–173
Bauer AW, Kirby WMM, Sherris JC (1996) Antibiotic susceptibility testing by a standardized single disc method. Am J Clin Pathol 45(4):493–496
Bello OA, Abdullahi OA (2016) Tolerance to heavy metals by some fungal isolates from petroleum refinery effluent in Kaduna, Nigeria. Br Microbiol Res J 12(6):1–8. doi:10.9734/BMRJ/2016/22728
Borjesson G, Menichetti L, Kirchmann H, Kätterer T (2012) Soil microbial community structure affected by 53 years of nitrogen fertilization and different organic amendments. Biol Fert Soils 48(3):245–257
Braud A, Jézéquel K, Bazot S, Lebeau T (2010) Enhanced phytoextraction of an agricultural Cr, hg− and Pb−contaminated soil by bioaugmentation with siderophore producing bacteria. Chemosphere 74:280–286. doi:10.1016/j.chemosphere.2008.09.013
Brown SD, Palumbo AV, Panikov N, Ariyawansa T, Klingeman DM, Johnson KM, Land ML, Utturkar SM, Epstein SL (2012) Draft genome sequence for Microbacterium laevaniformans strain OR221, a bacterium tolerant to metals, nitrate, and low pH. J Bacteriol 194(12):3279–3280. doi:10.1128/JB.00474-12
Burford EP, Fomina M, Gadd GM (2003) Fungal involvement in bioweathering and biotransformation of rocks and minerals. Mineral Mag 67:1127–1155. doi:10.1180/0026461036760154
Carlsohn MR, Groth I, Spror C, Schutze B, Saluz HP, Munder T, Stackebrandt E (2007) Kribbella aluminosa sp. nov., isolated from a medieval alum slate mine. Int J Syst Evol Microbiol 57:1943–1947
Carson JK, Gonzalez-Quinones V, Murphy DV, Hinz C, Shaw JA, Gleeson DB (2010) Low poreconnectivity increases bacterial diversity in soil. Appl Environ Microbiol 76:3936–3942. https://doi.org/10.1128/AEM.03085-09
Cervantes C, Gutierrez-Corona F (1994) Copper resistance mechanisms in bacteria and fungi. FEMS Microbiol Rev 14:121–137. doi:10.1111/j.1574- 6976.1994.tb00083.x
Cetin SC, Karaca A, Kizilkaya R, Turgay OC (2012) Role of plant growth promoting bacteria and fungi in heavy metal detoxification. In: Sheramati VA (ed) Detoxification of heavy metals. Soil Biology, vol 30. Springer-Verlag, Berlin, pp 369–388
Chen J, Chen HM, Zhang YQ, Wei YZ, Li QP, Liu HY, Jing S, Zhang YQ, Yu LY (2011) Agromyces flavus sp. nov., an actinomycete isolated from soil. Int J Syst Evol Microbiol 61:1705–1709. doi:10.1099/ijs.0.023242-0
Chen J, He F, Zhang X, Sun X, Zheng J, Zheng J (2014) Heavy metal pollution decreases microbial abundance, diversity and activity within particle-size fractions of a paddy soil. FEMS Microbiol Ecol 87:164–181. https://doi.org/10.1111/1574-6941.12212
Clara U, Filomena DL, Schumann P (2008) Kribbella catacumbae sp. nov. and Kribbella sancticallisti sp. nov., isolated from whitish-grey patinas in the catacombs of St Callistus in Rome, Italy. Int J Syst Evol Microbiol 58:2090–2097. doi:10.1099/ijs.0.65613-0
de Lima AF, de Moura GF, de Lima MA, de Souza PM, da sliva CA, De Campos GM, Do Nascimento AE (2011) Role of the morphology and polyphosphate in Trichoderma harzianum related to cadmium removal. Molecules 16(3):2486–2500. doi:10.3390/molecules16032486
de Lima MAB, de Franco, Luciana O, de Souza, Patrícia M, do Nascimento, Aline E, da Silva, Carlos AA, de Maia, Rita CC, Rolim, Hercília ML, Galba, MC (2013). Cadmium tolerance and removal from Cunninghamella elegans related to the polyphosphate metabolism. Int J Mol Sci 14(4):7180–7192. doi:https://doi.org/10.3390/ijms14047180
Do Carmo JR, Pimenta CJ, da Silva JF, de Souza SMC (2013) Recovery of copper (II) absorbed in biomass of Cladosporium cladosporioides. SciAgric 70:3. org:10.1590/S0103-90162013000300002
Dursun AY (2008) A comparative study on determination of the equilibrium, kinetic and thermodynamic parameters of biosorption of copper (II) and lead (II) ions onto pretreated Aspergillus niger. Biochem. Eng J 28:187–195. doi:org/10.1016/j.bej.2005.11.003
Elanvogvan R, Philip L, Chandraraj K (2010) Hexavalent chromium reduction by free and immobilized cell-free extract of Arthrobacter rhombi-RE. Appl Biochem Biotechnol 160(1):81–97. doi:org/10.1007/s12010-008-8515-6
El-Morsy EM (2004) Cunninghamella echinulata a new biosorbent of metal ions from polluted water in Egypt. Mycologia 96:1183–1189
Elsilk SE, Abd El-Raheem R, El-Shanshoury APS (2014) Accumulation of some heavy metal resistant avirulent Bacillus anthracis PS2010 isolated from Egypt. Afr J Micro Res 8(12):1266–1276. doi:10.5897/AJMR2013.6551
Essam F, Jumaily BH, Saleh BH, Shalal M, Hussain SM (2013) Determination of lethal dose (LD50)of exotoxin a from Pseudomonasaeruginosa26A in mice histopathology. IOSR J Pharm Biol Sci 7(6):49–54
Ezzourhi L, Castro E, Moya M (2009) Heavy metal tolerance of filamentous fungi isolated from polluted sites in tangier, Morocco. Afr J Microbiol Res 3:35–48. doi.org/10.7745/KJSSF.2012.45.4.565
Freitas AC, Ferreira F, Costa AM, Pereira R, Antunes SC, Gonçalves F, Rocha-Santos TAP, Diniz MS, Castro L, Peres I, Duarte AC (2009) Biological treatment of the effluent from a bleached kraft pulp mill using basidiomycete and zygomycete fungi. Sci Total Environ 407:3282–3289. doi:org/10.1016/j.scitotenv.2009.01.054
Gao Y, Miao C, Xia J, Mao L, Wang Y, Zhou P (2012) Plant diversity reduces the effect of multiple heavy metal pollution on soil enzyme activities and microbial community structure. Front Environ Sci En 6(2):213–223. doi:10.1007/s11783-011-0345-z
Gao S, Zhang WB, Sheng XF, He LY, Huang Z (2015) Chitinophaga longshanensis sp. nov., a mineral-weathering bacterium isolated from weathered rock. Int J Syst Evol Microbiol 65:418–423. doi:10.1099/ijs.0.067249-0
Ghorbani Y, Oliazadeh M, Shahvedi A (2007) Use of some isolated fungi in biological leaching of aluminum from low grade bauxite. Afr J Biotech 11:1284–1288. doi:10.5897/AJB2007.000-2178
Gilman JC (1957) A manual of soil fungi. Iowa State Univ Press, Ames, p 450
Grison CM, Jackson S, Merlot S, Dobson A, Grison C (2015) Rhizobium metallidurans sp. nov., a symbiotic heavy metal resistant bacterium isolated from the Anthyllis vulneraria Zn-hyperaccumulator. Int J Syst Evol Microbiol 65:1525–1530. doi:10.1099/ijs.0.000130
Gupta K, Chatterjee C, Gupta B (2012) Isolation and characterization of heavy metal tolerant Gram-positive bacteria with bioremedial properties from municipal waste rich soil of Kestopur canal (Kolkata)est Bengal, India. Biologia 67(5):827–836. doi:10.2478/s11756-012-0099-5
Gupta R, Kumar T, Mittal A (2016) Isolation, identification and characterization of heavy metal resistant bacteria from soil of an iron industry, Haryana (India). IJPSR 7(3):1308–1313. doi:10.13040/IJPSR.0975-8232.7(3).1308-13
Hao X, Taghavi S, Xie P, Orbach J, Alwathnani HA, Rensing C, Wei G (2014) Phytoremediation of heavy and transition metals aided by legume-rhizobia symbiosis. Int J Phytoremed 16(2):179–202. https://doi.org/10.1080/15226514.2013.773273
Harriso JJ, Ceri H, Raymond JT (2007) Multimetal resistance and tolerance in microbial biofilms. Nat Rev Microbiol 5:928–938. doi:10.1038/nrmicro1774
Hayat S, Ahmed I, Ahmed A, Samiullah IA (2002) Effect of long-term application of oil refinery wastewater on soil health with special reference to microbiological characteristics. Bioresour Technol 84:159–163. doi:10.1016/S0960-8524(02)00027-5
Hemida M, Morsy FM, El-Enany AE, Ohyama T (2012) Isolation and characterization of a heavy-metal-resistant isolate of Rhizobium leguminosarum viciae potentially applicable for biosorption of Cd2+ and Co2+. IntBiodeterBiodegr 67:48–55. doi:10.1016/j.ibiod.2011.10.008
Hijri M, Bell TH, Hassan S, Lauron-Moreau A, Al-Otaibi F (2014) Linkage between bacterial and fungal rhizosphere communities in hydrocarbon-contaminated soils is related to plant phylogeny. ISME J 8:331–334. doi:10.1038/ismej.2013.149
Hirak RD, Das S (2014). Bioremediation potential of mercury by Bacillus species isolated from marine environment and wastes of steel industry. Biorem J.18(3):204–212. doi.org/10.1080/10889868.2014.899555
Inga Z (2013) Study of the interaction of metals (Hg, Cr, Zn, Ag, Au) with Arthrobacter genera and Spirulina platensis. PhD thesis
Iram S, Parveen K, Usman J, Nasir K, Akhtar N, Arouj S, Ahmed I (2012) Heavy metal tolerance of filamentous fungal strains isolated from soil irrigated with industrial wastewater. Biologija 58(3):107–116. doi:org/10.6001/biologija.v58i3.2527
Issazadeh K, Jahanpour N, Pourghorbanali F, Raeisi G, Faekhondeh J (2013) Heavy metals resistance by bacterial strains. Ann Biol Res 4(2):60–63
Jaeckel P, Krauss GJ, Krauss G (2005) Cadmium and zinc response of the fungi Heliscus lugdunensis and Verticillium cf. alboatrum isolated from highly polluted water. Sci. Total Environ 346:274–279. doi:10.1016/j.scitotenv.2004.12.082
Juliana RDC, Pimenta CJ, Silva JF, Souza SMC (2013) Recovery of copper (II) absorbed in biomass of Cladosporium cladosporioides. Sci Agric 70(3):147–151. doi:org/10.1590/S0103-90162013000300002
Kanmami P, Aravind J, Preston D (2012) Remediation of chromium contaminants using bacteria. Int J Environ Sci Technol 9(1):183–193. doi:10.1007/s13762-011-0013-7
Karelová E, Harichová J, Stojnev T, Pangallo D, Ferianc P (2011) The isolation of heavy-metal resistant culturable bacteria and resistance determinants from a heavy-metal contaminated site. Biologia 66:18–26. doi:10.2478/s11756-010-0145-0
Karn SK, Reddy S (2012) Degradation of 2, 4,6-trichlorophenol by bacteria isolated from secondary sludge of a pulp and paper mill. J Gen Appl Microbiol 58:413–420. doi:org/10.2323/jgam.58.413
Khan AG (2000) Role of soil microbes in the rhizospheres of plants growing on trace metal contaminated soils in phytoremediation. JTraceElemMedBiol 18:355–364. doi:10.1007/s11356-015-4471-1
Kumar R, Singh P, Dhir B, Sharma AK, Mehta D (2014) Potential of some fungal and bacterial species in bioremediation of heavy metals. Journal of Nuclear Physics, Material Sciences, Radiation and Application 1(2):212–223
Kuykendall LD, Young J, Martinez-Romero E, Kerr A, Sawada H (2005) Genus I. Rhizobium. In: Garrity GM, Brenner DJ, Krieg NR, Staley J (eds) Bergey’s manual of systematic bacteriology, vol 2. Springer: Verlag, New York, pp 325–340
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics (modern microbiological methods). Wiley, New York, pp 115–175
Lee DW, Lee JE, Lee SD (2009) Chitinophaga rupis sp. nov., isolated from soil. Int J Syst Evol Microbiol 59:2830–2833. doi:10.1099/ijs.0.011163-0
Lee S, Cho K, Lim J, Kim W, Hwang S (2011a) Acclimation and activity of ammonia-oxidizing bacteria with respect to variations in zinc concentration, temperature, and microbial population. Bioresour Technol 102:4196–4203. doi:10.1016/j.biortech.2010.12.035
Lee M, Ten LN, Woo SG, Park J (2011b) Agromyces soli sp. nov., isolated from farm soil. Int. J Syst Evol Microbiol 61:1286–1292. doi:10.1155/2013/831308
Long F, Su CC, Lei HT, Bolla JR, Do SV, Yu EW (2012) Structure and mechanism of the tripartite CusCBA heavy-metal efflux complex. Philos Trans R Soc Lond Ser B Biol Sci 367:1047–1058. doi:10.1098/rstb.2011.0203
Ma Y, Oliveira RS, Freitas H, Zhang C (2016) Biochemical and molecular mechanisms of plant-microbe-metal interactions: relevance for phytoremediation. FrontPlant Sci 7:918. doi:org/10.3389/fpls.2016.00918
Madhaiyan M, Poonguzhali S, Sa T (2007) Metal tolerating methylotrophic bacteria reduces nickel and cadmium toxicity and promotes plant growth of tomato (Lycopersicon esculentum L.) Chemosphere 69:220–228. doi:10.1016/j.chemosphere.2007.04.017
Malgorzata JK, Rosikon K., Fijalkowski K, Grobelak A (2014).The effect of Trichoderma on heavy metal mobilityand uptake by Miscanthus giganteus, Salix sp., Phalaris arundinaceaand Panicumvirgatum.Appl Environ Soil Sci.1–10. doi.org/10.1155/2014/506142
Malik A (2004) Metal bioremediation through growing cells. Environ Int 30:261–278. doi:10.1016/j.envint.2003.08.001
Mandal SM, Bhattacharyya R (2012) Rhizobium-legume Symbiosis: a model system for the recovery of metal-contaminated agricultural land. In: Zaidi A, Wani PA, Khan MS (eds) Toxicity of heavy metals to legumes and bioremediationol.12.Springer, New York, pp 115–127
Mergeay M, Monchy S, Vallaeys T, Auquier V, Benotmane A, Bertin P, Taghavi S, Dunn J, van der Lelie D, Wattiez R (2003) Ralstonia metallidurans, a bacterium specifically adapted to toxic metals: towards a catalogue of metal-responsive genes. FEMS Microbiol 9:1181–1191. doi:10.1016/S0168-6445(03)00045-7
Mindlin S, Petrenko A, Kurakov A, Beletsky A, Mardanov A, Petrova M (2016) Resistance of permafrost and modern Acinetobacter lwoffii strains to heavy metals and arsenic revealed by genome analysis. Biomed Res Int. doi:org/10.1155/2016/3970831
Mohamed HA, Morsy FM, Enamy AWE, Ohyama (2012) Isolation and characterization of a heavy-metal-resistant isolate of Rhizobium leguminosarum bv. viciae potentially applicable for biosorption of Cd2+ and Co2+. Int Biodeter Biodegr 67:48–55. doi:10.1016/j.ibiod.2011.10.008
Mondani L, Piette L, Christen R, Bachar D, Berthomieu C, Chapon V (2012) Microbacterium lemovicicum sp. nov., a bacterium isolated from a natural uranium-rich soil. IntJSystEvolMicrobiol 63:2600–2606. doi:10.1007/s00203-015-1183-3
Nies DH (1995) The cobalt, zinc, and cadmium efflux system CzcABC from Alcaligenes eutrophus functions as a cation-proton antiporter in Escherichia coli. J Bacteriol 177:2707–2712
Nies DH (1999) Microbial heavy metal resistance. ApplMicrobial Biotechnol 51:730–750. doi:10.1007/s002530051457
Olukoya DK, Smith SI, Hori MO (1997) Isolation and characterization of heavy metal resistant bacteria from Lagos lagoon folia. Microbiol 42(5):441–444. doi:10.1007/BF02826550
Osborn AM, Bruce KD, Strike P, Ritchie DA (1997) Distribution, diversity and evolution of the bacterial mercury resistance (mer) operon. FEMS Microbiol Rev 19:239–262. doi:http://dx.doi.org/10.1111/j.1574-6976
Pal A, Choudhuri P, Dutta S, Mukherjee PK, Paul AK (2004) Isolation and characterization of nickel-resistant microflora from serpentine soils of Andaman. World J Microbiol Biotech 20:881–886. doi:10.1007/s11274-004-2776-1
Palanivel VP, Shim J, You Y, Choi S, Seralathan KK, Lee KJ, Kim HJ, Oh BT(2010). Removal of zinc by live, dead, and dried biomass of Fusarium spp. isolated from the abandoned-metal mine in South Korea and its perspective of producing nanocrystals. J Haz Mat. 182(1–3):317–324.doi:org/10.1016/j.jhazmat.2010.06.032
Paris D, Blondeau R (1999) Isolation and characterization of Arthrobacter sp. from activated sludge of a pulp and paper mill. Water Res 33(4):947–950. doi:org/10.1016/S0043-1354(98)00298-X
Park SK, Kim MS, Jung MJ, Nam YD, Park EJ, Roh SW, Bae JW (2011) Brachybacterium squillarum sp. nov., isolated from salt-fermented seafood. Int J Syst Evol Microbiol 61:1118–1122. https://doi.org/10.1099/ijs.0.022517-0
Pattus F, Abdallah M (2000) Siderophores and iron-transport in microorganisms: review. J Chin Chem Soc 47:1–20. doi:10.1002/jccs.200000001
Petrovic JJ, Danilovic G, Curcic N, Milinkovic M, Stosic N, Pankovic D, Raicevic V (2014) Coppertolerance of Trichoderma species. Arch Biol Sci Belgrade 66(1):137–142. https://doi.org/10.2298/ABS1401137J
Piccirillo C, Pereira SIA, Marques APGC, Pullar RC, Tobaldi DM, Pintado ME, Castro PML (2013) Bacteria immobilization on hydroxyapatite surface for heavy metals removal. J Environ Manag 121:87–95. doi:org/10.1016/j.jenvman.2013.02.036
Piotrowska-Seget Z, Cycon, M, Kozdroj J (2005) Metal tolerant bacteria occurring in heavily polluted soil and mine spoil. Appl Soil Ecol. 28:237–246. doi:org/10.1016/j.apsoil.2004.08.001
Rajbanshi A (2008) Study on heavy metal resistant bacteria in Guheswori sewage treatment plant. Our Nature 6:52–57. doi:10.3126/on.v6i1.1655
Rajkumar M, Ae N, Prasad MNV, Freitas H (2010) Potentialof siderophore-producing bacteria for improving heavy metal phytoextraction. Trends Biotechnol. 28:142–149. doi:org/10.1016/j.tibtech.2009.12.002
Rajkumar M, Sandhya S, Prasad MNV, Freitas H (2012) Perspectives of plant-associated microbes in heavy metal phytoremediation. Biotechnol Adv. 30(6):1562–1574.doi:org/10.1016/j.biotechadv.2012.04.011
Rao KR, Rashmi K, Latha JNL (2005) Bioremediation of toxic metal ions using biomass of Aspergillus fumigates from fermentative waste. Indian J Biotechnol 4:139–143. http://hdl.handle.net/123456789/5627
Raper KB, Fennell DJ (1965) The genus Aspergillus. Williams and Wilkins, Baltimore, p 686
Rathnayake VN, Megharaj M, Bolan N, Naidu R (2009) Tolerance of heavy metals by gram positive soil bacteria. WASET 53:1185–1189
Reeve WG, Tiwari RP, Kale NB, Dilworth MJ, Glenn AR (2002) ActP controls copper homeostasis in Rhizobium leguminosarum bv. viciae and Sinorhizobium meliloti preventing low pH-induced copper toxicity. Mol Biol 43(4):981–991. doi:10.1046/j.1365-2958.2002.02791.x
Rhodes CJ (2013) Applications of bioremediation and phytoremediation. Sci Prog. 96(4): 417–427.doi:org/10.3184/003685013X13818570960538
Rosales E, Pazos M, Sanroman MA, Tavares T (2012) Application of zeolite-Arthrobacter viscosus system for the removal of heavy metal and dye: chromium and azure B. Desalination. 284:150–156. doi:org/10.1016/j.desal.2011.08.049
Rouhollahi F, Zamani A, Karimi K, Etesami N (2014) Enhancement of nickel biosorption on fungal biomass by enzymatic and alkali pretreatments. Int J Environ Sci Technol. doi:10.1007/s13762-014-0624-x
Sahoo KN, Pakshirajan K, Ghosh PK (2014) Evaluation of 4-bromophenol biodegradation in mixed pollutants system by Arthrobacter chlorophenolicus A6 in an upflow packed bed reactor. Biodegradation 25(5):705–718. doi:10.1007/s10532-014-9693-2
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstruction phylogenetic trees. Mol Biol Evol 4:406–425
Santa CL, Gonzalez-Lopez J, Manzanera M (2013) Arthrobacter siccitolerans sp. nov., a highly desiccation-tolerant, xeroprotectant-producing strain isolated from dry soil. Int J Syst Evol Microbiol 63:4174–4180. doi:10.1099/ijs.0.052902-0
Schalk IJ, Hannauer M, Braud A (2011) New roles for bacterial siderophores in metal transport and tolerance. Environ Microbiol 13:2844–2854. doi:10.1111/j.1462-2920.2011.02556.x
Sen M (2011) Biosorption of Cr (VI) by resting cells of Fusarium solani Iran. J Environ Health SciEng 8(2):117–120. doi:10.4236/ojapps.2012.23021
Shivkumar CK, Thippeswamy B, Krishnappa M, Ananthamurthy KS (2011) Bioaccumulation potency of ecotoxic heavy metals by indigenous Rhizopus and Penicillium sp: a comparative study. J Pure. Appl Microbiol 6(1):231–239
Sessitsch A, Kuffner M, Kidd P, Vangronsveld J, Wenzel WW, Fallmann K, Puschenreiter M (2013) The role of plant-associated bacteria in the mobilization and phytoextraction of trace elements in contaminated soils. Soil Biol Biochem 60(100):182–194. doi:10.3389/fpls.2016.00918
Smith KS, Balistrieri LS, Todd AS (2015) Using biotic ligand models to predict metal toxicity in mineralized systems. Appl Geochem 57:55–72. doi:10/1016/j.apgeochem.2014.07.005
Song N, Ma Y, Zhao Y, Tang S (2015) Elevated ambient carbon dioxide and Trichoderma inoculum could enhance cadmium uptake of Lolium perenne explained by changes of soil pH, cadmium availability and microbial biomass. Appl Soil Ecol 85:56–64. doi:10.1016/j.apsoil.2014.09.007
Stan V, Gament E, Cornea CP, Voaideş C, Duşa M, Plopeanu G (2011) Effects of heavy metal from polluted soils on the Rhizobium diversity. Not Bot Hort Agrobot Cluj 39(1):88–95. doi:http://dx.doi.org/10.15835/nbha3916081
Stefanowicz AM, Niklińska M, Kapusta P, SzarekŁukaszewska G, Grodzińska K (2010).Metal pollution, soil properties and plant diversity as determinants of bacterial and fungal performance in soils. In: 15th ICHMET, pp 708–712
Sun F, Shao Z (2007) Biosorption and bioaccumulation of lead by Penicillium sp. Psf-2 isolated from the deep sea sediment of the Pacific Ocean. Extremophiles 11:853–858. doi:10.1007/s00792-007-0097-7
Swer PB, Joshi SR, Acharya C (2016) Cesium and strontium tolerant Arthrobacter sp. strain KMSZP6 isolated from a pristine uranium ore deposit. AMB Expr 6:69. doi:10.1186/s13568-016-0247-3
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121.pmid: 21546353
Tappe W, Herbst M, Hofmann D, Koeppchen S, Kummer S, Thiele B (2013) Degradation of sulfadiazine by Microbacterium lacus strain SDZm4, isolated from lysimeters previously manured with slurry from sulfadiazine-medicated pigs. Appl Environ Microb 79(8):2572–2577. doi:10.1128/AEM.03636-12 pmid:23396336
Teng Y, Luo Y, Ma W, Zhu L, Ren W, Luo Y. Li, Z (2015). Trichoderma reesei FS10-C enhances phytoremediation of cd-contaminated soil by Sedum plumbizincicola and associated soil microbial activities. Front Plant Sci. 9: 220. doi.org/10.3389/fpls.2015.00438
Thakre NA, Shanware AS (2015) Promising biological indicator of heavy metal pollution: bioluminescent bacterial strains isolated and characterized from marine niches of Goa, India. Indian J Microbiol 55(3):327–332. doi:10.1007/s12088-015-0531-y
Thawai C, Tanasupawat S, Suwanborirux K, Kudo T (2011) Agromyces tropicus sp. nov., isolated from soil. Int J Syst Evol Microbiol 61:605–609. doi:10.1099/ijs.0.021774-0
Thippeswamy B, Shivakumar CK, Krishnappa M (2012) Accumulation potency of heavy metals by Saccharomyces sp. indigenous to paper mill effluent. J Env Res Develop 6(3):439–445
Thippeswamy B, Shivakumar CK, Krishnappa M (2014) Optimization of heavy metals bioaccumulation in Aspergillus niger and Aspergillus flavus. Int J Environ Biol 4(2):188–195
Tripathi AK, Harsh NSK, Gupta N (2007) Fungal treatment of industrial effluents: a mini-review. Life Science Journal 4(2):78–81
Tunali S, Cabuk A, Akar T (2006) Removal of lead and copper ions from aqueous solutions by bacterial strain isolated from soil. ChemEngJ 115(3):203–211. doi:10.1016/j.cej.2005.09.023
Turner S, Pryer KM, Miao VPW (1999) Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol 46:327–338. doi:10.1111/j.1550-7408.1999.tb04612.x
Ullah A, Heng S, Munis MFH, Fahad S, Yang X (2015) Phytoremediation of heavy metals assisted by plant growth promoting (PGP) bacteria: a review. Environ Exp Bot 117:28–40. doi:10.1016/j.envexpbot.2015.05.001
Valix M, Tang JY, Malik R (2001) Heavy metal tolerance of fungi. Miner Eng 14:499–505. doi:10.1016/S0892-6875(01)00037-1
Van DB, Christov L (2002) Adsorption of colour from bleach plant effluent using biomass and cell wall fractions from Rhizomucor pusillus. J Chem Technol Biotechnol 77:155–158. doi:10.1002/jctb.535
Verma P, Singh S, Verma RK (2016) Heavy metal biosorption by Fusarium strains isolated from iron ore mines overburden soil. IntJEnvironSci Toxicol Res 4(4):61–69
Volesky B, Holan ZR (1995) Biosorption of heavy metals. Biotechnol Prog 11:235–250. doi:10.1021/bp00033a001
Wang YP, Shi JY, Wang H, Lin Q, Chen XC, Chen YX (2007) The influence of soil heavy metals pollution on soil microbial biomass, enzyme activity and community composition near a coppersmelter. Ecotoxicol Environ Saf 67:75–81. doi:10.1016/j.ecoenv.2006.03.007
Wang W, Shao Z, Liu Y, Wang G (2009) Removal of multi-heavy metals using biogenic manganese oxides generated by a deep-sea sedimentary bacterium - Brachybacterium sp. strain Mn32. Microbiology 155:1989–1996. doi:10.1099/mic.0.024141-0
Wang Q, Cheng C, He LY, Huang Z (2014) Chitinophaga jiangningensis sp. nov., a mineral-weathering bacterium. Int J Syst Evol Microbiol 64:260–265. doi:10.1099/ijs.0.067249-0
White TJ, Bruns T, LeeS TJ (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for Phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, New York, pp 315–322
Willie JG, Peijnenburg M, Zablotskaja M, Martina GV (2007) Monitoring metals in terrestrial environments within a bioavailability framework and a focus on soil extraction. Ecotox Environmen Safe 67(2):163–179. doi:10.1016/j.ecoenv.2007.02.008
Wu SC, Cheung KC, Luo YM, Wong MH (2006) Effects of inoculation of plant growth-promoting rhizobacteria on metal uptake by Brassica juncea. Environ Pollut 140:124–135. doi:10.1016/j.envpol.2005.06.023
Wuana RA, Okieimen FE (2011) Heavy metals in contaminated soils: a review of sources, chemistry, risks and best available strategies for remediation. ISRN Ecology. doi: org/10.5402/2011/402647
Yang HB, Tan N, Wu FJ, Liu HJ, Sun M, She ZG, Lin YC (2012) Biosorption of uranium (VI) by mangrove endophytic fungus Fusarium sp. ZZF51 from the South China Sea. J Radioanal Nucl Chem 292(3):1011–1016. doi:10.1007/s10967-011-1552-6
Yasir M, Chung EJ, Song GC, Bibi F, Jeon CK, Chung YR (2011) Chitinophaga eiseniae sp. nov., isolated from vermicompost. Int J Syst Evol Microbiol 61:2373–2378. doi:10.1099/ijs.0.040014-0
Yazdani MF, Abdullah CKY, Tan SG (2009) Trichoderma atroviride as a bioremediatorof cu pollution: an in vitro study. Toxicol Environ Chem 91(7):1305–1314. doi:10.1080/02772240802616510
Yu X, Li Y, Zhang C, Liu H, Liu J, Zheng W, Kang X, Qiang C (2014) Culturable heavy metal-resistant and plant growth promoting bacteria in V-Ti magnetite mine tailing soil from Panzhihua, China. PLoS One 9(9):e106618. doi:10.1371/journal.pone.0106618
Zafar S, Aqil F, Ahmad I (2007) Metal tolerance and biosorption potential of filamentous fungi isolated from metal contaminated agriculture soil. Bioresour Technol 98:2557–2561. doi:10.1016/j.biortech.2006.09.051
Zhang YF, He LY, Chen ZJ, Zhang WH, Wang QY, Qian M (2011) Characterization of lead-resistant and ACC deaminase-producing endophytic bacteria and their potential in promoting lead accumulation of rape. J Hazard Mater 186:720–725. doi:10.1016/j.jhazmat.2010.12.069
Zhang X, Lin L, Chen M, Zhu Z, Yang W, Chen B, Yang X, An Q (2012) A nonpathogenic Fusarium oxysporum strain enhances phytoextraction of heavy metals by the hyperaccumulator sedum alfredii Hance. J Hazard Mater 229-230:361–370. doi:10.1016/j.jhazmat.2012.06.013
Zhou Q, Chen Y, Yang M, Li W, Deng L (2013) Enhanced bioremediation of heavy metal from effluent by sulfate-reducing bacteria with copper-iron bimetallic particles support. Bioresour Technol 136:413–417. doi:10.1016/j.biortech.2013.03.047
Acknowledgements
The authors are grateful to the Commonwealth Scholarship grant, Canada, for carrying out the present work in the Department of Biological Science, University De Montreal, Montreal, and Genome Quebec Innovation Centre, Canada, for getting the accession numbers of all isolated trace metal-tolerant microbial strains.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Chanda, D., Sharma, G.D., Jha, D.K., Hijri, M. (2017). Tolerance of Microorganisms in Soil Contaminated with Trace Metals: An Overview. In: Shukla, P. (eds) Recent advances in Applied Microbiology . Springer, Singapore. https://doi.org/10.1007/978-981-10-5275-0_8
Download citation
DOI: https://doi.org/10.1007/978-981-10-5275-0_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-5274-3
Online ISBN: 978-981-10-5275-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)