Abstract
Plant secretomics is an emerging subfield of proteomics studying proteins globally secreted into the extracellular space (apoplast) by plant cells at defined time under constitutive or induced conditions. Plant secretome has important biological functions in cell wall structure formation, cell-to-cell interaction, extracellular/intracellular signal relay and appropriate cellular response to environmental stimuli. It also regulates the ability or inability of the host to trigger the defence system against the invading pathogen. Defence proteins are secreted via a classical pathway involving N-terminal signal peptide which directs the protein to the ER for routing, modification and subsequent secretion involving the endoplasmic reticulum (ER)–Golgi–trans-Golgi network (TGN)–plasma membrane system. Plant secretome has an increasing number of proteins following unconventional, ER–Golgi-independent or ‘leaderless’ apoplastic protein secretion mechanisms. Nonconventional mechanisms would be necessary if the presence of a protein in the ER/Golgi disrupts ER functioning or has multiple functions, each occurring in different cellular compartment. A large number of apoplastic leaderless secretome proteins have been identified that play an important role under salinity, low temperature, ion homeostasis and pathogen invasion. Characterisation of secretome is a formidable task, and success can be obliged to the advancement in biochemical, proteomic techniques, mass spectroscopy and bioinformatics. Advanced proteomic technologies established detailed secretome profiles from normal and stressed cell types at a faster pace. Discrimination of the true secretome from those released under environmental stresses is a big challenge. It warrants improved strategies to investigate the secretomes with high sensitivity and reproducibility. The comprehensive mechanisms regulating constitutive and induced secretome of diverse plants and their habitat are future perspective.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Plant secretomics is an emerging field of proteomics studying the secreted proteins of plants called ‘secretome’. The term ‘secretome’ was first used to describe a genome-wide study of the signal peptide-dependent secreted proteins and the protein secretion machineries in Bacillus subtilis, a Gram-positive bacterium (Tjalsma et al. 2000). The term is more often limited to include only the secreted proteins (Greenbaum et al. 2001; Hathout 2007; Bouws et al. 2008). Thus, ‘secretomics’ is defined as the study of proteins globally secreted into the extracellular space (apoplast) of cell, tissue or organ at any given time under specific conditions through various secretory mechanisms under constitutive or induced conditions’ (Agrawal et al. 2010).
Plant secretome has important biological functions in the formation of cell wall structure, cell-to-cell interaction, appropriate response to environmental stimuli and defence against pathogens (Isaacson and Rose 2006; Kamoun 2009). The cell wall is a major interface between plant cells and its surrounding environment. Rapid and regulated secretion of specific proteins into this extracellular space (apoplast) is an important defence response (Grant and Lamb 2006).
Apoplastic fluid is a complex mixture of proteins secreted constitutively and proteins secreted in response to environmental stimuli. Secretion of defence proteins or exocytosis in both plants and animals is generally achieved through a conventional pathway involving the endoplasmic reticulum–Golgi–trans-Golgi network–plasma membrane in the plant endomembrane system. It required an N-terminal signal peptide directing the protein to the ER for routing, modification and subsequent secretion via the Golgi apparatus. However, the presence of an increasing number of proteins lacking signal peptide in the apoplastic fluid suggest the existence of unconventional protein secretion mechanism. Numerous ER–Golgi-independent or ‘leaderless’ eukaryotic secretion mechanisms have been reported. Proteins are secreted by nonconventional mechanisms for a number of reasons. For instance, non-Golgi secretion would be necessary if the presence of a protein in the ER/Golgi would disrupt ER functioning. Non-Golgi secretion could also be desirable if a protein has multiple functions, each occurring in different cellular compartment. The significance of these secretory pathways, particularly in response to stresses, is well studied in animals and yeast (Nickel and Rabouille 2009); however, our information related to the knowledge of the protein population of a plant secretome and related secretory mechanisms remains limited in plants.
Secretomics has increasingly been the focus of biological research, and it has now become an intricate subfield of proteomics. Moreover, the information regarding the number and types of proteins found in the secretome of a specific plant under normal growth and stress conditions is still unknown. Hence, the complete secretome profile has now become a prerequisite rather than an option before we begin to systematically understand the function of secretory processes and proteins. Improvement in sequencing technology has made the genome sequences of more plant species completely known. Currently there are 24 land plants having completed or draft genome sequences available and 72 land plant species with genome sequencing in progress (http://www.ncbi.nlm.nih.gov/genomes/static/gpstat.html). The improvement and automation in proteomic technology is proving increasingly helpful for a systematic identification, qualitative and quantitative profiling and functional characterisation of plant secretome. The parallel development in bioinformatics has multiplied our ability to predict the protein-coding genes and the subcellular topographic locations of the encoded proteins, which is essential for the functional annotation of the genomes. The combined analyses of secretome assisted with genomic and bioinformatic techniques can correlate large-scale plant secretome studies and unravel mechanisms of plant response to various internal and external stimuli. In the present chapter, we have discussed the mechanisms of protein secretion in apoplastic fluid and methods of secretome isolation, separation, identification and annotation of their role in plants successfully completing their life span.
General Pathway of Secretory Proteins
A general characteristic of all prokaryotic or eukaryotic cells is to export the proteins from the cytoplasm to intracellular or extracellular locations. The secreted proteins in the apoplast or the extracellular space mediate major defence responses (Grant and Lamb 2006). The proteins destined to be exported are synthesised with a signal peptide that guides its translocation. Generally, the precursor protein with amino acid sequences of signal peptides is initially recognised by soluble targeting factors for its transport to the target membrane, for its association with translocation machinery. Then polypeptide chain is transported through a proteinaceous channel. The secretion of proteins takes place through secretory pathways involving the endoplasmic reticulum and Golgi apparatus. Reports have shown that the secretion of protein also takes place without classical secretory pathway; in plants and animals, protein secretion is solely mediated by the endoplasmic reticulum and Golgi apparatus. Secreted proteins have a signal peptide at N-terminus to direct them into the ER for sorting, modification and further secretion through the Golgi network. The existence of an alternate secretion mechanism is known which takes place without signal peptide (Auron et al. 1987). Their mechanism of secretion is Golgi-independent or leaderless secretion and is called nonclassical or unconventional secretory pathways (Nickel and Rabouille 2009).
Unconventional protein secretion takes place by two major methods: proteins are either transported in a non-vesicular mode where they pass directly from the cytosol through the plasma membrane or by various vesicular modes with membrane-bounded structures fusing with the plasma membrane before release in the extracellular space (Ding et al. 2012). Recently, a plant-specific compartment named exocyst-positive organelle (EXPO), has been shown to mediate nonclassical protein secretion from cytosol to cellwall without passing proteins via the Golgi apparatus, trans-Golgi network or multivesicular body (Wang et al. 2010a).
Molecular Biology of Secretome
Delivery of proteins through the endomembrane system to plasma membrane or the extracellular space (apoplast) usually starts with cotranslational insertion of proteins into the endoplasmic reticulum and then the cleavage of the signal peptide. The classical secretory pathway is highly conserved in eukaryotes (Burgess and Kelly 1987; Jurgens and Geldner 2007; Marti et al. 2010; Cai et al. 2011, 2012). There are regulated and consecutive secretory pathways that diverge in the trans-Golgi network. The constitutive classical secretory pathway is highly complex and operates in all cells (Fig. 1).
The constitutive and regulated secretory pathways (Adapted from Alberts et al. 2008)
Many soluble proteins are continually secreted from the cell by this pathway and translocate newly synthesised lipids and proteins to the plasma membrane. Specialised secretory cells also have a regulated secretory pathway, by which selected proteins in the trans-Golgi network are diverted into secretory vesicles, where the proteins are stored until an extracellular signal stimulates their secretion.
The secretory pathway transports proteins from one organelle to another within transport vesicles. Vesicular transport mediates a continuous exchange of components between chemically distinct, membrane-enclosed compartments that collectively constitute the biosynthetic–secretory and endocytic pathways. Most transport vesicles form specialised, coated regions of membranes that bud off as coated vesicles, with a distinctive cage of proteins covering their cytosolic surface. Before the vesicles fuse with a target membrane, they discard their coat, as is required for the two cytosolic membrane surfaces to interact directly and fuse. The coat performs two main functions: First is the selection of appropriate molecule transport concentrating specific membrane proteins in a specialised patch, forming vesicle membrane. Second, the coat moulds the vesicle into a curved, basketlike lattice that deforms the membrane patch and thereby shapes the vesicle. Hence, vesicles with the similar type of coat often have relatively the same size and shape. The vesicular transport selectively uses various cytosolic proteins like coat proteins (clathrin, COPI, COPII and retromer), some GTPases (Sar1, Arf1 and Rabs) and the ESCRT complexes (Kirchhausen 2000; Nickel et al. 2002; Gabe Lee et al. 2009; Hurley and Hanson 2010; Gao et al. 2012). There are three well-characterised types of coated vesicles, distinguished by their coat proteins: clathrin-coated, COPl-coated and COPII-coated. Each type is used for different transport steps. Clathrin-coated vesicles mediate transport from the Golgi apparatus and from the plasma membrane, whereas COPI- and COPII-coated vesicles mostly mediate transport from the ER and Golgi cisternae. The correct targeting and fusion of these vesicles to destined organelle depends on organelle-specific tethering factors and SNARE complexes (Cai et al. 2007; Sztul and Lupashin 2009) (Fig. 2).
Vesicle tethering to target membrane: Rab effector proteins interact via active Rab proteins (Rab-GTPs) on the target membrane, vesicle membrane or both to establish the first connection between the two membranes going to fuse. Rab effector is shown here as a filamentous tethering protein; SNARE proteins on the two membranes pair to dock the vesicle to the target membrane and catalyse the fusion of the two apposed lipid bilayer (Adapted from Alberts et al. 2008)
Many of the secretory proteins from mammals and yeasts are known to follow an unconventional secretory pathway (Auron et al. 1987); such processes are less reported in plants. In plants, more than 50 % of secretory proteins from total known plant secretome lack a signal peptide sequence and follow a leaderless secretory pathway (Agrawal et al. 2010). Studies performed using methods that cause least contamination of cytoplasmic proteins during secretome preparation and their analyses using highly sensitive enzymatic, immunoblotting and microarray showed the presence of high percentage of leaderless secretory protein in the plant secretome ruling out the possibility of contaminating nonsecretory proteins (Jung et al. 2007; Tran and Plaxton 2008; Cho et al. 2009).
Classical Secretome with Leader Peptide
The Classical Secretory Pathway for Protein Translocations Across Membrane
Proteins are the workhorses, which are synthesised in the cytoplasm. They ought to transport the entire polypeptide chain across one or two membranes in a unidirectional manner from the site of synthesis to the site of its biological function through the secretory pathway. The classical secretory pathway is a series of steps a cell follows to translocate a protein across a membrane bilayer or out of the cell via the endoplasmic reticulum through a process known as secretion. Translocation of nascent proteins across the membrane of the ER is known to occur in two ways: cotranslational translocation, in which translocation is concurrent with peptide synthesis by the ribosome, or posttranslational translocation, in which the protein is first completely synthesised in the cytosol and released from its polysomal complex and, thereafter, is transported into the ER. Both the methods of translocation are mediated by the same protein channel, known as Sec61 in eukaryotes and SecY in prokaryotes and archaea.
Cotranslational Translocation or Signal Recognition Particle (SRP)-Dependent Pathway
Proteins that follow a secretory pathway are destined for translocation across the ER membrane. The first stretch of the amino acids synthesised, called a signal/leader/transit peptide, allows a series of interactions starting with the recognition and its binding with SRP (Fig. 3).
Mechanism of cotranslational translocation of newly synthesised protein across the membrane (Adapted from Corsi and Schekman 1996)
The amino acid sequences of signal peptides are not conserved. ER targeting is specified by a central stretch of 7–20 hydrophobic amino acids. The extent of hydrophobicity of this region dictates cotranslational import into the ER. Eukaryotic SRP is a complex of six associated polypeptides and an RNA component which target substrates for cotranslational translocation into the ER. The SRP54 binds to the hydrophobic core of signal sequence as it emerges from the ribosome. The SRP complex, when bound to the ribosome and the signal sequence of the nascent peptide, pauses the elongation of the polypeptide by the subcomplex SRP9 and SRP14 by blocking the tRNA (Walter and Johnson 1994; Lutcke 1995). This translational arrest is to ensure proper targeting to the ER membrane before significant portions of the polypeptide emerge from the ribosome and begin to fold.
The ribosome along with its transit peptide–SRP complex is then attached to a docking protein. The docking protein is a heterodimeric SRP receptor (SR) composed of SRα and SRβ subunits. The SRP–nascent chain–ribosome complex binds to the docking protein and transfers the SRP–nascent chain–ribosome complex to the translocon, the Sec61, and then recycles back to the cytosol. The SRP is released from the SRP receptor after receptor-induced GTP hydrolysis by SRP54 component and completes the cycle (Miller et al. 1993). As the SRP and SRP receptor dissociate from the ribosome, the ribosome is able to bind directly with docking protein, Sec61. The Sec61 translocation channel (called SecY in prokaryotes) is a highly conserved heterotrimeric complex composed of α-, β- and γ-subunits. The pore of the channel, formed by the α-subunit, is blocked by a short helical segment which is thought to become unstructured during the beginning of protein translocation, allowing the peptide to pass through the channel. Completion of the synthesis of prepeptide resumes once the nascent signal peptide translocates across the channel into ER lumen. As the synthesis of prepeptide continues, it progressively penetrates into the ER lumen.
During translocation, the signal sequence is cleaved off by a signal peptidase present specifically in the ER lumen, freeing the amino terminus of the growing peptide. Translocated protein undergoes specific posttranslational modifications such as glycosylation or insertion of specific cofactor and is eventually stabilised by attaining a stable functional conformation. The stable functional protein can then be secreted via retrograde/anterograde pathways involving the Golgi apparatus to its final destination. If the secreted protein lacks any secondary signal sequence, they are secreted in the apoplast.
Posttranslational Translocation or SRP-Independent Pathway in Eukaryotes
Unlike cotranslational translocation, posttranslational or SRP-independent translocation of secretory proteins occurs independently of SRP in eukaryotes. The secretory precursor protein is completely translated and released from the ribosomal translation machinery (Fig. 4).
Mechanism of posttranslational translocation of newly synthesised protein in yeast (Adapted from Corsi and Schekman 1996)
Posttranslational targeting of secretory proteins requires cytosolic components, viz. cytosolic heat shock proteins (Hsp70), to maintain the polypeptide in an incompletely folded state. The posttranslational modes of translocation of secretory/membrane proteins prominently involve Sec translocon pathway, and Sec61p is a significant candidate subunit of the translocation channel.
In addition to the Sec61p complex, a second set of proteins called the Sec62p/Sec63p complex is required for posttranslational translocation in yeast. The Sec61 translocon associates with oligomeric membrane protein complex (Rapoport et al. 1999). This oligomeric membrane protein complex includes three integral membrane proteins, Sec62p, Sec63p and Sec71p, as well as Sec72p, which is peripherally associated with the cytosolic face of the ER, probably through association with Sec71p. Sec63p has been shown to form a subcomplex with Sec71p, Sec72p and BiP (Brodsky and Schekman 1993). BiP is a member of the Hsp70 family of ATPases, a group which is characterised as having an N-terminal nucleotide-binding domain and a C-terminal substrate-binding domain, which binds to peptides. Studies have proposed that Sec62p, Sec71p and Sec72p, together, create a surface for secretory precursors to bind before crossing the ER membrane.
Translocation apparatus for posttranslational translocation into a reconstituted proteoliposome consists of Sec61p and Sec62p/Sec63p complexes (Panzner et al. 1995). The Sec62p/Sec63p complex contains a cytoplasmic signal sequence receptor site that binds newly synthesised secretory proteins. The substrates are maintained in an unfolded, translocation-competent conformation with the aid of cytoplasmic chaperones (Chirico et al. 1988). Subsequent to binding, the signal is transferred from Sec62/Sec63 to the signal sequence receptor of the Sec61 translocon, and translocation occurs via the Sec61p channel. The primary role of the membrane protein complex Sec62/Sec63 is to activate the ATPase activity of BiP via Sec63p. The final step in the completion of translocation of precursor secretory proteins is full transfer from the pore into the ER lumen and requires functional Sec63p and BiP. The association of substrate-binding domain of BiP through Sec63p binds nonspecifically to the precursor peptide as it enters the ER lumen and allows the BiP to act as a translocation motor (Glick 1995; Brodsky 1996) and keeps the peptide from sliding backwards in a ratchet-type mechanism.
Unconventional Secretome with Leaderless Peptide or Without Leader Peptide
Secretion of defence proteins in both plants and animals was originally thought to be solely via an endoplasmic reticulum (ER)/Golgi-mediated pathway, with the help of an N-terminal signal peptide directing the protein to the ER.
Leaderless secretory proteins are modified in response to stress, thereafter enabling its interaction with relevant secretory pathways and subsequently resulting in its movement across the membrane (Denny et al. 2000; Backhaus et al. 2004). Multivesicular bodies (MBVs) in plants are prevacuolar compartments (Tse et al. 2004; Miao et al. 2008) and normally considered as endosomes of plants (Lam et al. 2007; Otegui and Spitzer 2008; Wang et al. 2009; Niemes et al. 2010; Robinson et al. 2012). They have been reported in the cytoplasm underlying the invasion papillae surrounding the fungal haustorium. The paramural bodies or lomasome is frequently observed at these sites and considered as the fusion profiles of MVBs with the plasma membrane. Callose is known to accumulate in the papillae and in the multivesicular bodies transported through endocytosis (An et al. 2006; Xu and Mendgen 1994).
Ding et al. (2012) described the three possible pathways for the leaderless secretory proteins (LSP) or nonclassical secretion of proteins.
The first LSP pathway is based on the fusion of multivesicular bodies with the plasma membrane to release the intraluminal vesicles to the apoplast, as exosomes. The release of exosomes depends on the behaviour of cytoplasmic domains of the two plasma membrane-localised SNAREs (syntaxin PEN1 and SNAP33), as their integration into the membrane of early endosome or trans-Golgi network in plants has been shown (Lam et al. 2007, 2008; Robinson et al. 2008, 2012; Meyer et al. 2009; Bednarek et al. 2010; Wang et al. 2010b). After maturation into the multivesicular bodies, these SNAREs are on the intraluminal vesicles within the multivesicular body (Robinson et al. 2012; Scheuring et al. 2011). These exosomal intraluminal vesicles accumulate SNAREs in the matrix of the papilla after fusion of MBVs to plasma membrane (Fig. 5).
Unconventional protein secretion pathways in plants (Adapted from Ding et al. 2012)
The second LSP pathway is based on the vacuolar fusion to plasma membrane. This was established by the pathogen-induced localised apoptosis at the site pathogen invasion. The localised apoptosis was due to the fusion of vacuole with the plasma membrane and the releasing of hydrolytic vacuolar enzymes with caspase-3-like activity into the apoplast, resulting in the lysis of bacterial and plant cells (Hatsugai and Hara-Nishimura 2010). It suggests that these vacuolar enzymes were originally delivered to the vacuolar lumen through conventional secretory organelles, but their secretion to apoplast is an unconventional secretion (Fig. 5).
The third LSP pathway is mediated by exocyst-positive organelles (EXPOs) discovered from suspension culture of Arabidopsis and tobacco BY-2 cells (Wang et al. 2010a). These organelles have also been reported from root tips, root hair cells and pollen grains. EXPOs are double-membrane in the cytoplasm but are single-membrane vesicles outside the plasma membrane. EXPOs are like autophagosomes being double-membrane-bound vesicles. But they are different from autophagosomes because their number does not change much under starvation, they do not fuse with endosomes and also they do not localise with autophagosome using standard marker.
EXPOs are not influenced by Brefeldin A (BFA, a fungal metabolite that reversibly inhibits the anterograde transport from ER to Golgi apparatus) or wortmannin (a specific inhibitor of phosphatidylinositol 3-kinase used to study protein trafficking and identifies organelles of plant secretory and endocytic pathways). It shows that the EXPOs do not follow the conventional secretory or endocytic pathways of the cell (Fig. 5). All vesicle carriers, irrespective of being involved in conventional or unconventional protein secretion, interact with the plasma membrane through the tethering factor called exocyst, a hetero-octameric complex made of Sec3, Sec5, Sec6, Sec8, Sec10, Sec15, Exo70 and Exo84 subunits (Chong et al. 2010). Each exocyst protein subunit is a single-gene product in yeasts and mammals, while in plants, Sec3, Sec5, Sec10 and Sec15 subunits are encoded by two genes, Exo84 by three genes and Exo70 by 23 genes (Zhang et al. 2010). The tethering factor exocyst is involved in conventional secretory processes, self-incompatibility response and pathogen response (Samuel et al. 2009; Zhang et al. 2010; Pecenkova et al. 2011).
The markers, pathways and regulators of unconventional protein secretory pathways which are reported from plants have been listed in Table 1. However, unconventional protein secretion still requires the omic studies utilising biochemical, cellular, molecular and genetic approaches to portray better understanding of nonconventional protein secretion.
Characterisation of Global Secretome
Plants produce metabolic responses against received stress signals to regulate entire plant growth and development. Plants have continuity of symplast and apoplast that helps to establish communication within the physiological system (Sakurai 1998). The apoplast is a dynamic compartment and helps to perceive and transduce signals from the external environment to the intracellular symplast. Hence, proteins secreted into the apoplastic fluid play an important role in biotic and abiotic stress responses. Various apoplast-secreted proteins are identified, which play important biological roles in cell wall structure, cellular communication and the responses to host–pathogen relationships (Masuda et al. 2001; Rep et al. 2003; Boudart et al. 2005; Alvarez et al. 2006; Djordjevic et al. 2007; Floerl et al. 2008). Germination of barley seed showed α-amylase synthesis in the aleurone layer and its secretion into the endosperm to break down starch (Ranki and Sopanen 1984). Apoplastic secretome of apple, peach, pear and plum including xylem sap and leaf apoplast is known to have antioxidative system in response to pox virus (Diaz-Vivancos et al. 2006). In poplar, nearly 300 unique apoplastic proteins have been identified, among which ~144 are from leaf apoplast and ~135 are from stem apoplast (Pechanova et al. 2010), whereas ~97 were root apoplast protein (Dafoe and Constabel 2009). The leaf apoplast proteins have major roles in cell wall metabolism and stress/defence response, whereas root apoplast proteins have the major function of stress/defence with cell wall metabolism as the secondary function (Pechanova et al. 2010).
Detailed studies of secreted proteins under normal, biotic/abiotic stress conditions revealed several types of novel secreted proteins, including the leaderless secretory proteins. These leaderless secretory proteins account for more than 50 % of the total identified secretome from eukaryotes. Presently, about 24 terrestrial plant genomes have been completely sequenced, whereas many are under progress.
The analyses of the different components of cells have revealed that >80 % of the curated secreted proteins are present in the apoplast or exterior to the cell wall. Approximately 1,700 secreted proteins have been manually curated from more than 150 plant species in the UniProtKB/Swiss-Prot database. Their subcellular locations are yet to be verified experimentally. Arabidopsis thaliana, being the most extensively studied plant model system, has 941, while Oryza sativa (japonica) has 226 curated secreted proteins in the database (Table 2).
Gene ontology analyses based on molecular functions showed that ~40 % of the total plant-secreted proteins and ~50 % of Arabidopsis-secreted proteins show hydrolase activity, with almost one third showing binding activity and ~15 % showing the catalytic activity (Lum and Min 2011).
The functional genome annotation requires prediction of protein-coding sequences as well as their subcellular locations. The UniProt Consortium (2010) has a database of plant secretomes allowing better and efficient prediction and analyses of curated and annotated secreted proteins, thus ultimately leading to enhancement of database by accurate prediction of plant secretomes and thus enhancing understanding about the response or action of secreted protein to a variety of internal and external environments.
Secretome Under Stresses
The plant apoplast research is lagging due to an obsolete concept of apoplast function and difficulties in the extraction and analysis of apoplastic proteins. Apoplast consisting compartments beyond the plasmalemma has a variety of functions during plant growth and development as well as in plant defence responses to stresses (Tian et al. 2009; Pennell 1998). During signal transduction, plant cells transport ligand like ions and other metabolites from the apoplast; therefore, a signal must cross the apoplast and plasmalemma (Sakurai 1998). Stress conditions significantly affect both quantity and quality of apoplastic proteins (Dani et al. 2005). Some stress conditions evidenced to alter the apoplast proteins in response to them include salt (Zhang et al. 2009), low temperature (Marentes et al. 1993), salicylic acid (Cheng et al. 2009a), metal toxicity (Fecht-Christoffers et al. 2003) and pathogen invasion (Oh et al. 2005). The roles of plant apoplastic proteins have been obviously ignored in analysing the plant stress response in comparison to the intracellular signalling pathway components and effectors.
Studies on ex planta (suspension cultured cells) and in planta systems identified a large number of leaderless secretory proteins in plants (Tran and Plaxton 2008; Cho et al. 2009; Agrawal et al. 2010), accounting for more than 50 % of total secretome, identified under biotic and abiotic stress conditions, exhibiting the existence of signal peptide-independent secretory mechanism.
Secretome Under Abiotic Stresses
Plants have evolved sophisticated systems to cope with adverse environmental conditions such as cold, drought and salinity. Although a number of stress response networks have been proposed, the role of plant apoplast protein stress response is less known.
The monocot model plant rice has salinity as one of the major environmental factors limiting growth and productivity. The rice root apoplastic proteins in response to salt stress have been deciphered (Zhang et al. 2009). The differential expression of rice secretome compared to untreated control revealed its role in response to salt stress and identified approximately 40 proteins, mainly involved in carbohydrate metabolism, oxidoreduction, protein processing and degradation (Song et al. 2011) (Fig. 6).
Salinity stress-responsive apoplastic protein network in rice seedlings (Song et al. 2011)
Low temperature stress decreases the productivity and limits the distribution of crop. Plants have different responses to freezing stress; some are freezing tolerant to withstand extracellular ice formation, and others prevent freezing by supercooling their sap. Apoplast proteome components prevent the lethal cell damage by ice-interacting proteins. The molecular mechanisms and components of the low temperature signalling in the apoplast are little known. The antifreezing proteins in the apoplast bind to the ice crystals, thus inhibiting growth of ice crystal rather than ice formation in plants. Some of the plant antifreezing proteins are homologous to pathogenesis-related proteins (chitinase and glucanase) which have hydrolytic activity in addition to antifreezing protein activity (Hon et al. 1995; Yaish et al. 2006).
Apoplast acts as the mediator of cell communication with the environment and is altered by the freezing stress and the analyses of secretome give better understanding to the mechanism of freezing tolerance. Secretome of Hippophae rhamnoides (sea buckthorn) identified 60 low-temperature-responsive (LTR) proteins, of which 50 % were upregulated LTR proteins (Gupta and Deswal 2012). SignalP and SecretomeP analysis showed that 76 % of the proteins identified were apoplastic proteins, among which 24 % were following classical and 52 % following the nonclassical secretory pathway (Table 3). Also, the nonsecretory proteins identified were the non-resident apoplast proteins and might be imported in response to any stimulus like low temperature.
Phosphate is a macronutrient important for plant growth and metabolism. The excessive use of phosphate fertilisers results in phosphate deficiency in soil. Plants respond to phosphate deficiency by increased root growth, lateral roots to increase surface area of absorption and reduced shoot growth (Vance et al. 2003). Differentially expressed secretome of Arabidopsis thaliana suspension cell cultures under phosphate-sufficient and phosphate-deficient conditions was analysed by proteomic approach which identified 37 unique proteins (Tran and Plaxton 2008). Among them, 24 secreted proteins were phosphorus-deficiency-responsive proteins, while 18 of them were upregulated and six downregulated secretory proteins (Table 4).
The mannitol dehydrogenase (MDH) is a cytoplasmic enzyme which is secreted in a Golgi-independent manner by tobacco in response to salicylic acid (Cheng et al. 2009b). The mannitol acts as a metabolite as well as an osmoprotectant due to its regulated conversion to mannose by MDH in the cytosol of plants like celery (Stoop et al. 1996). Thus, mannitol and MDH play an important role in plant–pathogen interactions. Golgi-independent secretion of MDH catabolises fungal mannitol in the extracellular space. The apoplastic peroxidases and high levels of leaderless secretory antioxidant protein Cu/Zn superoxide dismutase in response to biotic or salicylic acid stress lead to oxidative burst (Bindschedler et al. 2006; Cheng et al. 2009a).
Arabidopsis secretome induced by 1 mM salicylic acid (SA) showed a number of secretory proteins into the apoplast through classical or nonclassical secretory pathway (Cheng et al. 2009a).
Poplar (Populus spp) plants growing in riverine ecosystems, characterised by rapid environmental changes, have evolved as multistress response in the apoplast. Apoplast secretome of poplar constitutes a potential adaptive mechanism to inhabit successfully in dynamic riverine ecosystem (Pechanova et al. 2010).
Secretome Under Biotic Stresses
The plants’ cell wall acts as a barrier separating plant cells from the external environment. Therefore, proteins that are secreted in the extracellular space or apoplast play an important role in defence response. They reinforce the cell wall and antimicrobial activity via defence proteins such as chitinases, β-1, 3 glucanases, thionins, and defensins and lipid transfer proteins (Grant and Lamb 2006). These secreted proteins include various hydrolytic enzymes that are secreted in the apoplast as self-defence response to pathogen (bacteria, fungi and viruses) attack. Such pathogenesis-related proteins (PRPs) mediate cell-to-cell communication, and many of them follow the nonclassical secretory pathway (Bowles 1990; Agrawal et al. 2010). Thus, plant secretome might play a key role in the early recognition and defence against pathogen attack.
There are three types of plant responses to pathogen invasion: (1) by sensing the presence of elicitors through the cell surface receptors, (2) by producing localised oxidative burst by release of reactive oxygen species or (3) by releasing different types of antimicrobial compounds. The elicitors arise from the cell wall of pathogens or fragments of the host cell wall released by pathogen activity such as chitin fragments and detection of the fungal attack by release of chitinases like PR3 (Verburg and Huynh 1991; Kaku et al. 2006).
Studies on plant secretome improves our insight of defence mechanism during plant–pathogen interactions. Identification of secreted proteins in Arabidopsis suspension-cultured cells (Ndimba et al. 2003; Oh et al. 2005), maize (Chivasa et al. 2005) and tobacco BY2 cell (Okushima et al. 2000) have been reported in response to fungal pathogens. The various pathogen elicitor-responsive proteins identified from plants include lectin receptor-like kinase, endochitinase, xyloglucan endo-1, 4-β-D-glucanases and peroxidise from cell suspension culture filtrates. Also, there are reports from whole-protein extracts that furthers our understanding on plant–pathogen interactions and defence signalling in wheat (Rampitsch et al. 2006), maize (Chivasa et al. 2005), pea (Curto et al. 2006), Arabidopsis (Jones et al. 2006) and rice (Ventelon-Debout et al. 2004; Lee et al. 2006).
Kim et al. (2009) identified 21 differentially expressed proteins in the secretome in response to rice blast fungus (Magnaporthe grisea) and its elicitor in rice suspension culture. These secreted proteins include chitinases, expansins and germins/oxalate oxidases. The secretory proteins expressed in elicitor-treated suspension cell culture were nearly similar to the M. grisea-infected rice leaves. It established the early recognition of pathogens via secreted proteins in resistant rice plants.
Pathogen attack in plants initiates a signalling cascade that leads to the synthesis of salicylic acid, which induces the expression or secretion of various PRPs that play a key role in systemic acquired resistance of plants.
In planta secretome analyses of Phytophthora capsici-infected pepper (Capsicum annuum) identified 75 secretory proteins (Yeom et al. 2011). Majority of the secreted proteins were defence- and stress-related proteins, proteases, protease inhibitors or cell wall structural proteins.
Water and pathogen stress-mediating PRPs in apoplast are effective in suppressing growth of Melampsora causing leaf fungal rust in poplar (Rinaldi et al. 2007). The leaf apoplast secretome showed the presence of proteins like acidic class III chitinase, thaumatin-like protein, blight-associated protein p12, cationic peroxidase 1 and cysteine-rich repeat secretory protein 38 in response to Melampsora infection. The acidic class III chitinase expression increased under pathogen challenge with M. larici-populina as well as under drought condition. It suggests a broad spectrum of role of apoplastic PRPs under stresses. A group of cysteine-rich peptides, defensins, are conserved in plants, and animals possess antimicrobial activity against a variety of fungus (Thomma et al. 2002). Some of them in plants are antibacterial or even few have a role in anti-insect activity. The defensins have different mechanisms for antifungal activity, such as (1) by interacting fungal cell wall components and causing localised apoptosis (Thevissen et al. 2012), (2) by binding to fungal ion channels to block it (Spelbrink et al. 2004) or (3) by modulating permeability of fungal plasma membrane leading to fungal cell death (Mello et al. 2011). It reveals that apoplast secretome-based defence mechanism works effectively against pathogens by activating pathogenesis-related proteins under biotic stress (Pechanova et al. 2010).
Secretome of Developmental Stages
Multicellular organisms have evolved an efficient system for cellular communication to ensure the ordered development, growth, maintenance and reproduction to successfully complete their life span. It requires cells to coordinate response to environmental stimuli as well as to each other by integrating the wide array of extracellular and intercellular signals. The cellular secretion in the apoplast by plant is an important biological process and serves as an interface between the environment and organism. Cell-to-cell communication during developmental stages predominantly involves secreted peptide ligands and interacts with their receptors present on the plasma membrane on the target cells. Apoplastic secretome contains proteins secreted through the classical ER–Golgi–TGN pathway or secreted by unconventional protein secretion mechanisms. The apoplast is not an empty space bordering a cell but rather participates in functions in plants (Lippmann et al. 2009) including nutrients and growth regulation, water regulation, osmoregulatory homeostasis of solutes, tissue structure, defence against biotic/abiotic stress, transport, osmosis, cell adhesion and gas exchange (Floerl et al. 2012).
The secretome of a tobacco cell suspension culture identified proteins mainly involved in stress defence and cell regeneration processes (Lippmann et al. 2009). Secretome analysis of chickpea (Cicer arietinum) callus suspension culture revealed 773 proteins in the extracellular medium (Gupta et al. 2011). Peroxidases, chitinases and other pathogenesis-related proteins were identified in cell cultures of tomato and grapevine after application of elicitors like cyclodextrins and methyl jasmonate (Briceno et al. 2012; Martinez-Esteso et al. 2009). Functional studies have revealed a multitude of secreted peptides involved in diverse biological processes (Table 5). Secreted peptides are categorised into two main groups distinguished on the basis of their biogenesis and overall structure: the CLEs (CLAVATA3/embryo-surrounding region, CLV3/ESR) including related peptide family and the CRPs (cysteine-rich peptides).
In conclusion, secreted peptides play a much more important role in cell–cell communication through relaying signals via ligand–receptor machinery of diverse biological contexts. These recent findings defy the traditional view of plant cells being unable to communicate by ligand–receptor interaction on the surface because of the surrounding cell wall. It is highly likely that better understanding of secretomics of plants growing in diverse environment will reveal more biological processes in which interaction of apoplastic fluid proteins with receptors presents on the plasma membrane of one cell with another cell.
Current Strategies to Study Plant Secretome
Plant cell secretes many proteins through exocytosis to the apoplastic fluid to maintain cell structure and regulate the external environment and as a part of signalling and defence mechanisms.
In recent years, there has been an increased interest in plant and microbe secretomes as the secreted proteins have shown to play an important role in normal growth, stress biology, infection and progression of diseases and subsequent response in plant protection (Agrawal et al. 2010; Stassen et al. 2012). Advancement in the proteomic profiling of plant secretome can be owed to the advancement in biochemical, proteomic techniques, mass spectroscopy and bioinformatics approaches. The complete characterisation of a proteome/secretome is a formidable task, and the degree of success achieved depends on the methods available and their amenability to automation and high-throughput formats. Parameters such as the complexity of the protein mixture, levels of expression and modification and intracellular localisation all impact the choice of proteomics technology to be used. It had established several in-depth secretome profiles from different cell types, apoplastic fluids from normal and diseased conditions at a faster pace. The biggest challenge in secretomic studies is the discrimination between the proteins that are truly secreted from those that are released as the result of non-physiological environmental stresses. Hence, strategies and techniques are being continuously modified to best adapt and to investigate the plant secretomes with high reproducibility. It is therefore important to establish a simple, reproducible and economic procedure for systematic secretome analysis in plants.
Sample Preparation
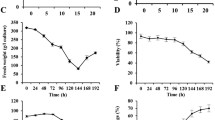
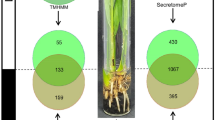
The sample preparation of secreted proteins devoid of host-plant proteins is one of the most critical and challenging aspect in the secretome analyses. Most secretome studies to date are performed both on ex vivo suspension cell cultures (SCCs) and in planta systems (Fig. 7). Ex planta SCCs may not sufficiently complement to the in planta environment, thus reducing the correlation to the true physiological secretome.
A general overview of the ex planta and in planta systems for secretome analysis (Agrawal et al. 2010)
Ex Planta System
Plant suspension cell cultures are widely used as a convenient tool for secretome analyses of the plant bypassing the structural complexity of the plant. The homogeneity of SCC cell population; the large availability of material; the easiness to maintain, handle and scale up/down; the high rate of cell growth; and the good reproducibility of conditions make it suitable for the analysis of plant secretome. Secreted proteins into the culture medium are used to prepare the secretome.
Different strategies and techniques have been applied to isolate pure secreted proteins suitable for proteomic analysis (Fig. 8).
Workflow showing preparation of secreted proteins using the ex planta SCC (Agrawal et al. 2010)
Particulate free cell culture fluid containing pure secreted proteins can be obtained by filtration and/or high-speed centrifugation. Basically, filtration and centrifugation are a good combination to obtain clear cell culture fluid. Isolated secretome is either snap frozen immediately and stored at −80 °C or freeze-dried in a vacuum lyophiliser followed by dialysis against a suitable buffer. The lyophilised protein sample can be subjected to TCA precipitation to concentrate and effectively remove the salts, small peptides, water-soluble medium components, secondary metabolites and polysaccharides. The secreted protein pellet can be immediately processed for biochemical and proteomic analysis.
In Planta System
The apoplastic fluid between the middle lamella and primary wall is isolated using biochemical methods. The classical vacuum infiltration–centrifugation (VIC) method and the newly introduced gravity extraction method (GEM) are suitable and well-established methods for apoplastic secretome isolation (Fig. 9).
Workflow for in planta preparation of secretome using VIC and GEM (Agrawal et al. 2010)
The vacuum infiltration–centrifugation method involves two critical steps: (1) vacuum infiltration with or without appropriate extraction buffer and (2) centrifugation speed and time. The suitability of the classical vacuum infiltration–centrifugation method (Terry and Bonner 1980) for apoplastic secretome collection was further supported by critically evaluating it on intact leaves from different plant species – Vicia faba L., Phaseolus vulgaris L., Pisum sativum L., Hordeum vulgare L, Spinacia oleracea L., Beta vulgaris L. and Zea mays L. (Lohaus et al. 2001). Strength of infiltration buffer, incubation time and processing time showed relatively little impact on composition of the apoplastic fluid. In contrast, the pH of infiltrated solution highly influenced the concentration of sucrose and hexoses.
Separation of secretome from the culture medium or extracellular fluid can be easily done by filtration without cell disruption or by low-speed centrifugation. Moreover, the fraction of dead cells can be determined by staining the culture with trypane blue to identify any nonsecreted cytoplasmic proteins as being contaminants. Ex planta SCCs in model plants like tobacco, Arabidopsis, rice and Medicago have been used for secretome studies.
In gravity extraction method, the apoplastic fluid is obtained in a single step. It is a simple, reproducible and novel method for the extraction and preparation of pure secreted proteins (Cottingham 2008).
Two types of biochemical analyses are generally used to assess sample free from contamination of soluble cytoplasmic proteins before starting the proteomic analyses. The enzyme activity and western blotting of soluble cytoplasmic marker proteins, e.g. exclusively cytoplasmic enzymes like glucose 6-phosphate dehydrogenase (GAPDH) (Oh et al. 2005), phosphoenolpyruvate carboxylase (PEP-carboxylase) (Tran and Plaxton 2008) and cytosolic aldolase (Tran and Plaxton 2008), are assessed. The absence of the enzymatic activity and reference band on western blot confirms the purity of apoplastic fluid secretome preparation.
Protein Separation
Proteins are extremely diverse molecules and differ by mass, charge, hydrophobicity, tertiary shape and their affinity for other molecules. Several approaches are being employed for the separation of complex mixtures of protein. It is crucial to obtain a protein sample which contains only the molecule of interest. One-dimensional electrophoresis (SDS-PAGE) and 2D gel electrophoresis are the most popular gel-based protein separation methods. In 1DE, proteins are separated on the basis of molecular mass. Moreover, 1DE is simple to perform, is reproducible and can be used to resolve proteins with molecular masses of 10–300 kDa. However, limited resolving power of a 1DE is a limitation, if a more complex protein mixture, such as a crude cell lysate, is to be separated. Limitation of resolving power can be conquered by the use of 2-dimensional gel electrophoresis.
Two-dimensional (2D) gel electrophoresis is a powerful gel-based method commonly used for ‘global’ analyses of complex samples, i.e. when we are interested to characterise the entire spectrum of proteins in a sample. One of the greatest advantages of 2DE is the ability to resolve proteins that have undergone some form of posttranslational modification. This resolution is possible in 2DE because many types of protein modifications confer a difference in charge as well as a change in mass on the protein. In 2D, proteins are separated by two distinct physical properties of protein. In the first dimension, proteins are resolved according to their net charge and in the second dimension, according to their molecular mass. The combination of these two methods produces resolution far exceeding that obtained in 1DE using a crude protein sample.
In this technique, the proteins are focused first in an immobilised pH gradient strip on the basis of their isoelectric point (acid vs. basic character). The isoelectric focusing gel is then placed over an SDS-PAGE gel and runs in the perpendicular dimension, and proteins are resolved on the basis of mass (Fig. 10). Thus, proteins separate not as bands but as spots, and the position of each protein spot depends on both the size and charge of protein.
Identification of Proteome Make-up of Secretome
After separation of secretome contents on 2D array, its proteome profile is identified by deducing the primary structure of each spot arrayed. Presently, mass spectrometry (MS), coupled with other preparative and analytical methods, is the main method in proteomics for achieving sensitivity and reproducibility. The major advantage of MS analysis is that it can process posttranslationally modified protein sample of any size required in a picomole quantities. Modification of N-terminus of protein cannot be sequenced using Edman’s degradation method, and its sensitivity is up to 30 residues. Normally, the combination of 2D gel electrophoresis and MS is the most widely used method to study proteome. The protein spots are in-gel digested with proteolytic enzymes like pepsin, trypsin, chymotrypsin, papain, bromelain and subtilisin. After proteolysis, the peptides are separated on in-line LC-MS instrument. The data obtained from MS analysis of peptides can be taken directly for comparison to protein sequences derived from protein and nucleotide sequence databases.
Recently, a modified version called differential in-gel electrophoresis (DIGE) has improved performance at the gel-based part (Knowles et al. 2003; Marouga et al. 2005; Ye et al. 2010). Multidimensional protein identification technology (MudPIT) (Link et al. 1999) is a gel-free method to analyse the highly complex samples necessary for large-scale proteome analysis by ESI-MS/MS and database searching. As it is most frequently used, MudPIT couples a two-dimensional liquid chromatography (2D-LC) separation of peptides on a microcapillary column with detection in a tandem mass spectrometer. In the MudPIT experiment, a protein or mixture of proteins is reduced, alkylated and digested into a complex mixture of peptides. The digested peptide sample is pressure-loaded directly onto a microcapillary column where they are separated on the basis of their size and hydrophobicity. Once peptides are separated and eluted from the microcapillary column, they are ionised and enter the mass spectrometer, where they are isolated based on their mass-to-charge ratio (m/z). Tandem mass spectra are generated and are searched against a protein database. With the advent of proteomic techniques and mass spectrometry instrumentation, the efficiency of identifying and quantifying proteins in biological samples, including secretomes, has greatly improved.
Bioinformatic Analysis of Secretome Data from 2D and MS Array
High level of precision, sensitivity and resolution of MS have significantly increased high throughput of proteomics. Such high-precision instruments are capable of generating huge volume of high-quality data. Thus, comparison of secretome data obtained from different experiments/laboratories on specific cell/tissue types in defined conditions and specific diseases is warranted. For validation and extraction of significant outcomes has become feasible only by the development of multiple proteomic database and related software. A growing number of prediction tools for the plant secretome enable prediction of SPs, TMDs, GPI anchors or conserved domains in novel secretory proteins. Proteins secreted via the classical ER–Golgi–TGN pathway can be identified by their signal peptide using the SignalP 4.0 server (http://www.cbs.dtu.dk/services/SignalP) (Petersen et al. 2011), Phobius (http://phobius.binf.ku.dk/) (Kall et al. 2004) and TargetP (http://www.cbs.dtu.dk/services/TargetP) (Emanuelsson et al. 2007). The SecretomeP software helps to find LSPs by searching for certain LSP typical protein features apart from the lack of a signal sequence (Bendtsen et al. 2004). The recently developed bioinformatics tool LocTree2 uses a hierarchic, decision tree-like structure imitating the cellular protein sorting cascade to predict the subcellular localisation of proteins and thus also their secretion into the apoplast (Goldberg et al. 2012).
Currently there is an urgent need to deposit both raw mass spectrometry data and the corresponding list of identified proteins in public domains such as PRIDE (www.ebi.ac.uk/pride) and ProteomeXchange (www.proteomexchange.org) for proteomics-related studies.
References
Agrawal GK, Jwa NS, Lebrun MH, Job D, Rakwal R (2010) Plant secretome: unlocking secrets of the secreted proteins. Proteomics 10(4):799–827
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2008) Molecular biology of the cell. Garland Science, Taylor & Francis Group, New York
Alvarez S, Goodger JQD, Marsh EL, Chen S, Asirvatham VS, Schachtman DP (2006) Characterization of the maize xylem sap proteome. J Proteome Res 5:963–972
An Q, Hückelhoven R, Kogel KH, Van Bel AJE (2006) Multivesicular bodies participate in a cell wall associated defence response in barley leaves attacked by the pathogenic powdery mildew fungus. Cell Microbiol 8:1009–1019
Auron PE, Warner SJC, Webb AC, Cannon JG, Bernheim HA, McAdam KJPW, Rosenwasser LJ, LoPreste G, Mucci SF, Dinarello CA (1987) Studies on the molecular nature of human interleukin 1. J Immunol 138:1447–1456
Backhaus R, Zehe C, Wegehingel S, Kehlenbach A, Schwappach B, Nickel W (2004) Unconventional protein secretion: membrane translocation of FGF-2 does not require protein unfolding. J Cell Sci 117:1727–1736
Bednarek P, Kwon C, Schulze-Lefert P (2010) Not a peripheral issue: secretion in plant–microbe interactions. Curr Opin Plant Biol 13:378–387
Bendtsen JD, Jensen LJ, Blom N, Von Heijne G, Brunak S (2004) Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng Des Sel 17:349–356
Bindschedler LV, Laurence V, Dewdney J, Blee KA, Stone JM, Asai T, Plotnikov J, Denoux C, Hayes T, Gerrish C, Davies DR, Ausubel FM, Bolwell GP (2006) Peroxidase-dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. Plant J 47:851–863
Boudart G, Jamet E, Rossignol M, Lafitte C, Borderies G, Jauneau A, Esquerre-Tugaye M-T, Pont-Lezica R (2005) Cell wall proteins in apoplastic fluids of Arabidopsis thaliana rosettes: identification by mass spectrometry and bioinformatics. Proteomics 5:212–221
Bouws H, Wattenberg A, Zorn H (2008) Fungal secretomes-nature’s toolbox for white biotechnology. Appl Microbiol Biotechnol 80:381–388
Bowles DJ (1990) Defence-related proteins in higher plants. Annu Rev Biochem 59:873–907
Briceno Z, Almagro L, Sabater-Jara AB, Calderon AA, Pedreno MA, Ferrer MA (2012) Enhancement of phytosterols, taraxasterol and induction of extracellular pathogenesis-related proteins in cell cultures of Solanum lycopersicum cv Micro-Tom elicited with cyclodextrins and methyl jasmonate. J Plant Physiol 169:1050–1058
Brodsky JL (1996) Post-translational protein translocation: not all hsc70s are created equal. Trends Biochem Sci 21:122–126
Brodsky JL, Schekman R (1993) A Sec63p-BiP complex from yeast is required for protein translocation in a reconstituted proteoliposome. J Cell Biol 123:1355–1363
Burgess TL, Kelly RB (1987) Constitutive and regulated secretion of proteins. Annu Rev Cell Biol 3:243–293
Cai H, Reinisch K, Ferro-Novick S (2007) Coats, tethers, Rabs, and SNAREs work together to mediate the intracellular destination of a transport vesicle. Dev Cell 12:671–682
Cai Y, Jia T, Lam SK, Ding Y, Gao C, San MWY, Pimpl P, Jiang L (2011) Multiple cytosolic and transmembrane determinants are required for the trafficking of SCAMP1 via an ER-Golgi-TGN-PM pathway. Plant J 65:882–896
Cai Y, Zhuang X, Wang J, Wang H, Lam SK, Gao C, Wang X, Jiang L (2012) Vacuolar degradation of two integral plasma membrane proteins, AtLRR84A and OsSCAMP1, is cargo ubiquitination-independent and prevacuolar compartment mediated in plant cells. Traffic 13:1023–1040
Cheng FY, Blackburn K, Lin YM, Goshe MB, Williamson JD (2009a) Absolute protein quantification by LC/MS(E) for global analysis of salicylic acid-induced plant protein secretion responses. J Proteome Res 8:82–93
Cheng FY, Zamski E, Guo W, Pharr DM, Williamson JD (2009b) Salicylic acid stimulates secretion of the normally symplastic enzyme mannitol dehydrogenase (MTD): a possible defense against mannitol secreting fungal pathogens. Planta 230:1093–1103
Chirico WJ, Waters MG, Blobel G (1988) 70 K heat shock related proteins stimulate protein translocation into microsomes. Nature 332:805–810
Chivasa S, Simon WJ, Yu XL, Yalpani N, Slabas AR (2005) Pathogen elicitor–induced changes in the maize extracellular matrix proteome. Proteomics 5:4894–4904
Cho WK, Chen XY, Chu H, Rim Y, Kim S, Kim ST, Kim SW, Park ZY, Kim JY (2009) The proteomic analysis of the secretome of rice calli. Plant Physiol 135:331–341
Chong YT, Gidda SK, Sanford C, Parkinson J, Mullen RT, Goring DR (2010) Characterization of the Arabidopsis thaliana exocyst complex gene families by phylogenetic, expression profiling, and subcellular localization studies. New Phytol 185:401–419
Corsi AK, Schekman R (1996) Mechanism of polypeptide translocation into the endoplasmic reticulum. J Biol Chem 271:30299–30302
Cottingham K (2008) Unlocking the secrets of the rice secretome. J Proteome Res 7:5072
Curto M, Camafeita E, Lopez JA, Maldonado AM, Rubiales D, Jorrin JV (2006) A proteomic approach to study pea (Pisum sativum) responses to powdery mildew (Erysiphe pisi). Proteomics 6:163–174
Dafoe NJ, Constabel CP (2009) Proteomic analysis of hybrid poplar xylem sap. Phytochemistry 70:856–863
Dani V, Simon WJ, Duranti M, Croy RR (2005) Changes in the tobacco leaf apoplast proteome in response to salt stress. Proteomics 5:737–745
Denny PW, Gokool S, Russell DG, Field MC, Smith DF (2000) Acylation-dependent protein export in Leishmania. J Biol Chem 275:11017–11025
Diaz-Vivancos P, Rubio M, Mesonero V, Periago PM, Ros-Barcelo A, Martinez-Gomez P, Hernandez JA (2006) The apoplastic antioxidant system in Prunus: response to long-term plum pox virus infection. J Exp Bot 57:3813–3824
Ding Y, Wang J, Wang J, Stierhof YD, Robinson DG, Jiang L (2012) Unconventional protein secretion. Trends Plant Sci 17(10):606–615
Djordjevic MA, Oakes M, Li DX, Hwang CH, Hocart CH, Gresshoff PM (2007) The Glycine max xylem sap and apoplast proteome. J Proteome Res 6:3771–3779
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2(4):953–971
Fecht-Christoffers MM, Braun HP, Lemaitre-Guillier C, VanDorsselaer A, Horst WJ (2003) Effect of manganese toxicity on the proteome of the leaf apoplast in cowpea. Plant Physiol 133:1935–1946
Floerl S, Druebert C, Majcherczyk A, Karlovsky P, Kues U, Polle A (2008) Defence reactions in the apoplastic proteome of oilseed rape (Brassica napus var. napus) attenuate Verticillium longisporum growth but not disease symptoms. BMC Plant Biol 8:129. doi:10.1186/1471-2229-8-129
Floerl S, Majcherczyk A, Possienke M, Feussner K, Tappe H, Gatz C, Feussner I, Kues U, Polle A (2012) Verticillium longisporum infection affects the leaf apoplastic proteome, metabolome, and cell wall properties in Arabidopsis thaliana. PLoS One 7:e31435
Gabe Lee MT, Mishra A, Lambright DG (2009) Structural mechanisms for regulation of membrane traffic by Rab GTPases. Traffic 10:1377–1389
Gao C, Yu CK, Qu S, San MW, Li KY, Lo SW, Jiang L (2012) The Golgi-localized Arabidopsis endomembrane protein12 contains both endoplasmic reticulum export and Golgi retention signals at its C terminus. Plant Cell 24:2086–2104
Glick BS (1995) Can Hsp70 proteins act as force-generating motors? Cell 80:11–14
Goldberg T, Hamp T, Rost B (2012) LocTree2 predicts localization for all domains of life. Bioinformatics 28:i458–i465
Grant M, Lamb C (2006) Systemic immunity. Curr Opin Plant Biol 9:414–420
Greenbaum D, Luscombe NM, Jansen R, Qian J, Gerstein M (2001) Interrelating different types of genomic data, from proteome to secretome: ’oming in on function. Genome Res 11:1463–1468
Gupta R, Deswal R (2012) Low temperature stress modulated secretome analysis and purification of antifreeze protein from Hippophae rhamnoides, a Himalayan wonder plant. J Proteome Res 11:2684–2696. dx.doi.org/10.1021/pr200944z
Gupta S, Wardhan V, Verma S, Gayali S, Rajamani U, Datta A, Chakraborty S, Chakraborty N (2011) Characterization of the secretome of chickpea suspension culture reveals pathway abundance and the expected and unexpected secreted proteins. J Proteome Res 10:5006–5015
Hathout Y (2007) Approaches to the study of the cell secretome. Expert Rev Proteomics 4:239–248
Hatsugai N, Hara-Nishimura I (2010) Two vacuole-mediated defence strategies in plants. Plant Signal Behav 5:1568–1570
Hon WC, Griffith M, Mlynarz A, Kwok YC, Yang DSC (1995) Antifreeze proteins in winter rye are similar to pathogenesis related proteins. Plant Physiol 109:878–889
Hurley JH, Hanson PI (2010) Membrane budding and scission by the ESCRT machinery: it’s all in the neck. Nat Rev Mol Cell Biol 11:556–566
Isaacson T, Rose JKC (2006) The plant cell wall proteome, or secretome. In: Finnie C (ed) Plant proteomics, vol 28, Annual Plant Reviews Series. Blackwell Publishing, Oxford, pp 185–209
Jones AM, Thomas V, Bennett MH, Mansfield J, Grant M (2006) Modifications to the Arabidopsis defense proteome occur prior to significant transcriptional change in response to inoculation with Pseudomonas syringae. Plant Physiol 142:1603–1620
Jung C, Lyou SH, Yeu SY, Kim MA, Lee JS, Choi YD, Cheong JJ (2007) Microarray-based screening of jasmonate responsive genes in Arabidopsis thaliana. Plant Cell Rep 26:1053–1063
Jurgens G, Geldner N (2007) The high road and the low road: trafficking choices in plants. Cell 130:977–979
Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, Takio K, Minami E, Shibuya N (2006) Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc Natl Acad Sci U S A 103:11086–11091
Kall L, Krogh A, Sonnhammer EL (2004) A combined transmembrane topology and signal peptide prediction method. J Mol Biol 338(5):1027–1036
Kamoun S (2009) The secretome of plant-associated fungi and oomycetes. In: Deising VH (ed) Plant relationships, 2nd edn, The Mycota. Springer, Berlin/Heidelberg, pp 173–180
Kim ST, Kang YH, Wang Y, Wu J, Park ZY, Rakwal R, Agrawal GK, Lee SY, Kang KY (2009) Secretome analysis of differentially induced proteins in rice suspension-cultured cells triggered by rice blast fungus and elicitor. Proteomics 9:1302–1313
Kirchhausen T (2000) Three ways to make a vesicle. Nat Rev Mol Cell Biol 1:187–198
Knowles MR, Cervino S, Skynner HA, Hunt SP, de Felipe C, Salim K, Meneses-Lorente G, McAllister G, Guest PC (2003) Multiplex proteomic analysis by two-dimensional differential in-gel electrophoresis. Proteomics 3:1162–1171
Krause C, Richter S, Knöll C, Jürgens G (2013) Plant secretome – from cellular process to biological activity. Biochim et Biophys Acta http://dx.doi.org/10.1016/j.bbapap.2013.03.024
Lam SK, Tse YC, Robinson DG, Jiang L (2007) Tracking down the elusive early endosome. Trends Plant Sci 12:497–505
Lam SK, Cai Y, Hillmer S, Robinson DR, Jiang L (2008) SCAMPs highlight the developing cell plate during cytokinesis in tobacco BY-2 cells. Plant Physiol 147:1637–1645
Lee J, Bricker TM, Lefevre M, Pinson SRM, Oard JH (2006) Proteomic and genetic approaches to identifying defence-related proteins in rice challenged with the fungal pathogen Rhizoctonia solani. Mol Plant Pathol 7:405–416
Link AJ, Eng J, Schieltz DM, Carmack E, Mize GJ, Morris DR, Garvik BM, Yates JR III (1999) Direct analysis of protein complexes using mass spectrometry. Nat Biotechnol 17:676–682
Lippmann R, Kaspar S, Rutten T, Melzer M, Kumlehn MA, Mock HP (2009) Protein and metabolite analysis reveals permanent induction of stress defense and cell regeneration processes in a tobacco cell suspension culture. Int J Mol Sci 10:3012–3032
Lohaus G, Pennewiss K, Sattelmacher B, Hussmann M, Mühling KH (2001) Is the infiltration-centrifugation technique appropriate for the isolation of apoplastic fluid? A critical evaluation with different plant species. Physiol Plant 111:457–465
Lum G, Min XJ (2011) Plant secretomics: current status and future perspectives. Plant Omics J 4(2):114–119
Lutcke H (1995) Signal recognition particle (SRP), a ubiquitous initiator of protein translocation. Eur J Biochem 228:531–550
Marentes E, Griffith M, Mlynarz A, Brush RA (1993) Proteins accumulate in the apoplast of winter rye leaves during cold acclimation. Physiol Plant 87:499–507
Marouga R, David S, Hawkins E (2005) The development of the DIGE system: 2D fluorescence difference gel analysis technology. Anal Bioanal Chem 382:669–678
Marti L, Fornaciari S, Renna L, Stefano G, Brandizzi F (2010) COPII-mediated traffic in plants. Trends Plant Sci 15:522–528
Martinez-Esteso MJ, Selles-Marchart S, Vera-Urbina JC, Pedreno MA, Bru-Martinez R (2009) Changes of defense proteins in the extracellular proteome of grapevine (Vitis vinifera cv. Gamay) cell cultures in response to elicitors. J Proteomics 73:331–341
Masuda S, Kamada H, Satoh S (2001) Chitinase in cucumber xylem sap. Biosci Biotechnol Biochem 65:1883–1885
Mello EO, Ribeiro SF, Carvalho AO, Santos IS, Da Cunha D, Santa-Catarina C, Gomes VM (2011) Antifungal activity of PvD1 defensin involves plasma membrane permeabilization, inhibition of medium acidification, and induction of ROS in fungi cells. Curr Microbiol 62:1209–1211
Meyer D, Paionk S, Micali C, O’Connell R, Schulze-Lefert P (2009) Extracellular transport and integration of plant secretory proteins into pathogen-induced cell wall compartments. Plant J 57:986–999
Miao Y, Li KY, Yao X, Jiang L (2008) The vacuolar transport of aleurain-GFP and 2S albumin-GFP fusions is mediated by the same pre-vacuolar compartments in tobacco BY-2 and Arabidopsis suspension cultured cells. Plant J 56:824–839
Miller JD, Wilhelm H, Gierasch L, Gilmore R, Walter P (1993) GTP binding and hydrolysis by the signal recognition particle during initiation of protein translocation. Nature 366:351–354
Ndimba BK, Chivasa S, Hamilton JM, Simon WJ, Slabas AR (2003) Proteomic analysis of changes in the extracellular matrix of Arabidopsis cell suspension cultures induced by fungal elicitors. Proteomics 3:1047–1059
Nickel W, Rabouille C (2009) Mechanisms of regulated unconventional protein secretion. Nat Rev Mol Cell Biol 10:148–155
Nickel W, Briigger B, Wieland FT (2002) Vesicular transport: the core machinery of COPI recruitment and budding. J Cell Sci 115:3235–3240
Niemes S, Langhans M, Viotti C, Scheuring D, Yan MSY, Jiang L, Hilmer S, Robinson DG, Pimpl P (2010) Retromer recycles vacuolar sorting receptors from the trans-Golgi network. Plant J 61:107–121
Oh IS, Park AR, Bae MS, Kwon SJ, Kim YS, Lee JE, Kang NY, Lee S, Cheong H, Park OK (2005) Secretome analysis reveals an Arabidopsis lipase involved in defense against Alternaria brassicicola. Plant Cell 17:2832–2847
Okushima Y, Koizumi N, Kusano T, Sano H (2000) Secreted proteins of tobacco cultured BY2 cells: identification of a new member of pathogenesis-related proteins. Plant Mol Biol 42:479–488
Otegui MS, Spitzer C (2008) Endosomal functions in plants. Traffic 9:1589–1598
Panzner S, Dreier L, Hartmann E, Kostka S, Rapoport TA (1995) Post-translational protein transport in yeast reconstituted with a purified complex of Sec proteins and Kar2p. Cell 81:561–570
Pecenkova T, Hála M, Kulich I, Kocourková D, Drdová E, Fendrych M, Toupalová H, Žárskýý V (2011) The role for the exocyst complex subunits Exo70B2 and Exo70H1 in the plant–pathogen interaction. J Exp Bot 62:2107–2116
Pechanova O, Hsu CY, Adams JP, Pechan T, Vandervelde L, Drnevich J, Jawdy S, Adeli A, Suttle JC, Lawrence AM, Tschaplinski TJ, Séguin A, Yuceer C (2010) Apoplast proteome reveals that extracellular matrix contributes to multistress response in poplar. BMC Genomics 11:674. doi:10.1186/1471-2164-11-674
Pennell R (1998) Cell walls: structures and signals. Curr Opin Plant Biol 1:504–510
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Rampitsch C, Bykova NV, McCallum B, Beimcik E, Ens W (2006) Analysis of the wheat and Puccinia triticina (leaf rust) proteomes during a susceptible host-pathogen interaction. Proteomics 6:1897–1907
Ranki H, Sopanen T (1984) Secretion of α-amylase by the aleurone layer and the scutellum of germinating barley grain. Plant Physiol 75:710–715
Rapoport TA, Matlack KE, Plath K, Misselwitz B, Staeck O (1999) Post-translational protein translocation across the membrane of the endoplasmic reticulum. Biol Chem 380:1143–1150
Rep M, Dekker HL, Vossen JH, de Boer AD, Houterman PM, de Koster CG, Cornelissen BJC (2003) A tomato xylem sap protein represents a new family of small cysteine-rich proteins with structural similarity to lipid transfer proteins. FEBS Lett 534:82–86
Rinaldi C, Kohler A, Frey P, Duchaussoy F, Ningre N, Couloux A, Wincker P, Thiec DL, Fluch S, Martin F, Duplessis S (2007) Transcript profiling of poplar leaves upon infection with compatible and incompatible strains of the foliar rust Melampsora larici-populina. Plant Physiol 144:347–366
Robinson DG, Langhans M, Saint-Jore-Dupas C, Hawes C (2008) BFA effects are tissue and not just plant specific. Trends Plant Sci 13:405–408
Robinson DG, Pimpl P, Scheuring D, Stierhof YD, Sturm S, Viotti C (2012) Trying to make sense of retromer. Trends Plant Sci 17:431–439
Sakurai N (1998) Dynamic function and regulation of apoplast in the plant body. J Plant Res 111:133–148
Samuel MA, Chong YT, Haasen KE, Aldea-Brydges MG, Stone SL, Goring DR (2009) Cellular pathways regulating responses to compatible and self-Incompatible pollen in Brassica and Arabidopsis stigmas intersect at Exo70A1, a putative component of the exocyst complex. Plant Cell 21:2655–2671
Scheuring D, Viotti C, Krüger F, Künzl F, Sturm S, Bubeck J, Hillmer S, Frigerio L, Robinson DG, Pimpl P, Schumacher K (2011) Multivesicular bodies mature from the trans-Golgi network/early endosome in Arabidopsis. Plant Cell 23:3463–3481
Song Y, Zhang C, Ge W, Zhang Y, Burlingame AL, Guo Y (2011) Identification of NaCl stress-responsive apoplastic proteins in rice shoot stems by 2D-DIGE. J Proteomics 74(7):1045–1067
Spelbrink RG, Dilmac N, Allen A, Smith TJ, Shah DM, Hockerman GH (2004) Differential antifungal and calcium channel-blocking activity among structurally related plant defensins. Plant Physiol 135:2055–2067
Stassen JH, Seidl MF, Vergeer PW, Nijman IJ, Snel B, Cuppen E, Van den Ackervenken G (2012) Effector identification in the lettuce downy mildew Bremia lactucae by massively parallel transcriptome sequencing. Mol Plant Pathol 13(7):719–731
Stoop JMH, Williamson JD, Pharr DM (1996) Mannitol metabolism in plants: a method for coping with stress. Trends Plant Sci 1:139–144
Sztul E, Lupashin V (2009) Role of vesicle tethering factors in the ER–Golgi membrane traffic. FEBS Lett 583:3770–3783
Terry M, Bonner B (1980) An examination of centrifugation as a method of extracting an extracellular solution from peas, and its use for the study of indoleacetic acid-induced growth. Plant Physiol 66:321–325
The UniProt Consortium (2010) The universal protein resource (UniProt) in 2010. Nucleic Acids Res 38:D142–D148. doi:10.1093/nar/gkp846
Thevissen K, de Mello Tavares P, Xu D, Blankenship J, Vandenbosch D, Idkowiak-Baldys J, Govaert G, Bink A, Rozental S, de Groot PW, Davis TR, Kumamoto CA, Vargas G, Nimrichter L, Coenye T, Mitchell A, Roemer T, Hannun YA, Cammue BP (2012) The plant defensin RsAFP2 induces cell wall stress, septin mislocalization and accumulation of ceramides in Candida albicans. Mol Microbiol 84:166–180
Thomma BP, Cammue BP, Thevissen K (2002) Plant defensins. Planta 216:193–202
Tian L, Zhang L, Zhang J, Song Y, Guo Y (2009) Differential proteomic analysis of soluble extracellular proteins reveals the cysteine protease and cystatin involved in suspension-cultured cell proliferation in rice. Biochim Biophys Acta 1794:459–467
Tjalsma H, Bolhuis A, Jongbloed JDH, Bron S, van Dijl JM (2000) Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome. Microbiol Mol Biol Rev 64:515–547
Tran HT, Plaxton WC (2008) Proteomic analysis of alterations in the secretome of Arabidopsis thaliana suspension cells subjected to nutritional phosphate deficiency. Proteomics 8:4317–4326. doi:10.1002/pmic.200800292
Tse YC, Mo B, Hilmer S, Zhao M, Lo SW, Robinson DG, Jiang L (2004) Identification of multivesicular bodies as prevacuolar compartments in Nicotiana tabacum BY-2 Cells. Plant Cell 16:672–693
Vance CP, Uhde-Stone C, Allan DL (2003) Phosphorus acquisition and use: critical adaptations by plants for securing a nonrenewable resource. New Phytol 157:427–447
Ventelon-Debout M, Delalande F, Brizard JP, Diemer H, Van Dorselaer A, Brugiduo C (2004) Proteome analysis of cultivar-specific deregulations of Oryza sativa indica and O. sativa japonica cellular suspensions undergoing rice yellow mottle virus infection. Proteomics 4:216–225
Verburg JG, Huynh QK (1991) Purification and characterization of an antifungal chitinase from Arabidopsis thaliana. Plant Physiol 95:450–455
Walter P, Johnson AE (1994) Signal sequence recognition and protein targeting to the endoplasmic reticulum membrane. Annu Rev Cell Biol 10:87–119
Wang J, Cai Y, Miao Y, Lam SK, Jiang L (2009) Wortmannin induces homotypic fusion of plant prevacuolar compartments. J Exp Bot 60:3075–3083
Wang J, Ding Y, Wang J, Hillmer S, Miao Y, Lo SW, Wang X, Robinson DG, Jiang L (2010a) EXPO, an exocyst-positive organelle distinct from multivesicular endosomes and autophagosomes, mediates cytosol to cell wall exocytosis in Arabidopsis and tobacco cells. Plant Cell 22:4009–4030
Wang H, Tse YC, Law AHY, Sun SSM, Sun YB, Xu ZF, Hillmer S, Robinson DG, Jiang L (2010b) Vacuolar sorting receptors (VSRs) and secretory carrier membrane proteins (SCAMPs) are essential for pollen tube growth. Plant J 61:826–838
Xu H, Mendgen K (1994) Endocytosis of 1,3-β-glucans by broad bean cells at the penetration site of the cowpea rust fungus (haploid stage). Planta 195:282–290
Yaish MWF, Doxey AC, McConkey BJ, Moffatt BA, Griffith M (2006) Cold-active winter rye glucanases with ice-binding capacity. Plant Physiol 141:1459–1472
Ye Y, Mar E-C, Tong S, Sammons S, Fang S, Anderson JL, Wang D (2010) Application of proteomics methods for pathogen discovery. J Virol Methods 163:87–95. doi:10.1016/j.jviromet.2009.09.002
Yeom SI, Baek HK, Oh SK, Kang WH, Lee SJ, Lee JM, Seo E, Rose JK, Kim BD, Choi D (2011) Use of a secretion trap screen in pepper following Phytophthora capsici infection reveals novel functions of secreted plant proteins in modulating cell death. Mol Plant Microbe Interact 24:671–684
Zhang L, Tian LH, Zhao JF, Song Y, Zhang CJ, Guo Y (2009) Identification of an apoplastic protein involved in the initial phase of salt stress response in rice root by two-dimensional electrophoresis. Plant Physiol 149:916–928
Zhang Y, Liu CM, Emons AMC, Ketelaar T (2010) The plant exocyst. J Integr Plant Biol 52:138–146
Acknowledgements
We are thankful to Dr. Javed Ahmed, Assistant Professor, King Saud University, and Dr. Abhishek Ojha, Postdoctoral Fellow, International Centre for Genetic Engineering and Biotechnology, New Delhi, for providing us the literatures for consultation and their timely and useful suggestions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer India
About this chapter
Cite this chapter
Yadav, N., Khurana, S.M.P., Yadav, D.K. (2015). Plant Secretomics: Unique Initiatives. In: Barh, D., Khan, M., Davies, E. (eds) PlantOmics: The Omics of Plant Science. Springer, New Delhi. https://doi.org/10.1007/978-81-322-2172-2_12
Download citation
DOI: https://doi.org/10.1007/978-81-322-2172-2_12
Published:
Publisher Name: Springer, New Delhi
Print ISBN: 978-81-322-2171-5
Online ISBN: 978-81-322-2172-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)