Abstract
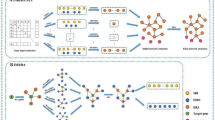
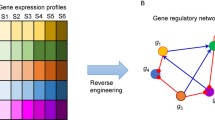
Gene regulatory network is a model of a network that describes the relationships among genes in a given condition. However, constructing gene regulatory network is a complicated task as high-throughput technologies generate large-scale of data compared to number of sample. In addition, the data involves a substantial amount of noise and false positive results that hinder the downstream analysis performance. To address these problems Bayesian network model has attracted the most attention. However, the key challenge in using Bayesian network to model GRN is related to its learning structure. Bayesian network structure learning is NP-hard and computationally complex. Therefore, this research aims to address the issue related to Bayesian network structure learning by proposing a low-order conditional independence method. In addition we revised the gene regulatory relationships by integrating biological heterogeneous dataset to extract transcription factors for regulator and target genes. The empirical results indicate that proposed method works better with biological knowledge processing with a precision of 83.3% in comparison to a network that rely on microarray only, which achieved correctness of 80.85%.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
References
Yavari, F., Towhidkhah, F., Gharibzadeh, S.: Gene regulatory network modeling using Bayesian networks and cross correlation. Biomedical Engineering Conference, CIBEC, Cairo (2008)
Gevaert, O., Van Vooren, S., Moor, B.D.: A framework for elucidating regulatory networks based on prior information and expression data. In: Eklund, P., Mann, G.A., Ellis, G. (eds.) ICCS 1996. LNCS, vol. 1115, pp. 240–248. Springer, Heidelberg (1996)
Huang, Z., Li, J., Su, H., Watts, G.S., Chen, H.: Large-scale regulatory network analysis from microarray data: Modified Bayesian network learning and association rule mining. Decision Support Systems 43, 1207–1225 (2007)
Friedman, N., Linial, M., Nachman, I., Pe’er, D.: Using bayesian networks to analyze expression data. Journal of Computational Biology 7(3), 601–620 (2000)
Zou, M., Conzen, S.D.: A new dynamic Bayesian network (DBN) approach for identifying gene regulatory networks from time course microarray data. Bioinformatics 21(1), 71–79 (2005)
Ahmad, F.K., Deris, S., Othman, N.H.: The Inference of Breast Cancer Metastasis through Gene Regulatory Networks. Journal of Biomedical Informatics (JBI) 45(2), 350–362 (2012)
Rhodes, D.R., Yu, J., Shanker, K., Deshpande, N., Varambally, R., Ghosh, D., et al.: Large-scale meta-analysis of cancer microarray data identifies common transcriptional profiles of neoplastic transformation and progression. Proceedings of the National Academy of Sciences of the United States of America 101(25), 9309–9314 (2004)
Zhang, Y., Zha, H., Wang, J.Z., Chu, C.H.: Gene co-regulation vs. co-expression: The Pennsylvania State University, University Park, PA (2004)
Yeung, K.Y., Medvedovic, M., Bumgarner, R.E.: From co-expression to co-regulation: how many microarray experiments do we need? Genome Biology 5, R48 (2004)
Zhao, W., Serpedin, E., Dougherty, E.R.: Recovering genetic regulatory networks from chromatin immunoprecipitation and steady-state microarray data. Eurasip. J. Bioinform. Syst. Biol. (2008)
Kaleta, C., Göhler, A., Schuster, S., Jahreis, K., Guthke, R., Nikolajewa, S.: Integrative inference of gene-regulatory networks in Escherichia coli using information theoretic concepts and sequence analysis. BMC Systems Biology 4(116) (2010)
Pearl, J.: Probabilistic reasoning in intelligent systems: Networks of plausible inference. Morgan Kaufmann Publishers Inc., Francisco (1988)
Wille, A., Buhlmann, P.: Low-order conditional independence graphs for inferring genetic networks. Statistical Applications in Genetics and Molecular Biology 4(32) (2006)
van’t Veer, L.J., Dai, H., van de Vijver, M.J., He, Y.D., Hart, A.A.M., Mao, M., Peterse, H.L., van de Kooy, K., Marton, M.J., Witteveen, A.T., Schreiber, G.J., Kerkhoven, R.M., Roberts, C., Linsley, P.S., Bernards, R., Friend, S.H.: Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002)
Mi, Z., Guo, H., Wai, P.Y., Gao, C., Wei, J., Kuo, P.C.: Differential Osteopontin expression in phenotypically distinct subclones of murine breast cancer cells mediates metastatic behavior*. Journal of Biological Chemistry 279(45), 46659–46667 (2004)
Kato, T., Katabami, K., Takatsuki, H., Han, S.A., Takeuchi, K., Irimura, T., et al.: Characterization of the promoter for the mouse a3 integrin gene Involvement of the Ets-family of transcription factors in the promoter activity. Eur. J. Biochem. 269, 4524–4532 (2002)
Fang, S.H., Chen, Y., Weigel, R.J.: GATA-3 as a marker of hormone response in breast cancer. Journal of Surgical Research 157(2), 290–295 (2009)
Xiao, X., Li, B., Mitton, B., Ikeda, A., Sakamoto, K.: Targeting CREB for cancer therapy: friend or foe. Curr. Cancer Drug Targets 10(4), 384–391 (2010)
Gordon, S., Akopyan, G., Garban, H., Bonavida, B.: Transcription factor YY1: structure, function, and therapeutic implications in cancer biology. Oncogene 25, 1125–1142 (2006)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Kabir Ahmad, F., Yusoff, N. (2013). Reconstructing Gene Regulatory Network Using Heterogeneous Biological Data. In: Ramanna, S., Lingras, P., Sombattheera, C., Krishna, A. (eds) Multi-disciplinary Trends in Artificial Intelligence. MIWAI 2013. Lecture Notes in Computer Science(), vol 8271. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-44949-9_10
Download citation
DOI: https://doi.org/10.1007/978-3-642-44949-9_10
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-44948-2
Online ISBN: 978-3-642-44949-9
eBook Packages: Computer ScienceComputer Science (R0)